Lecture 6: Enzyme Mechanisms and Kinetics
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
example of an enzyme mechanism
chymotrypsin
example of “serine protease” → enzyme protein
hydrolysis cleaves polypeptides on C-terminal side of aromatic amino acids
mechanims → both acid/base catalysis and covalent intermediate
describe the chymotrypsin mechanims: free enzyme
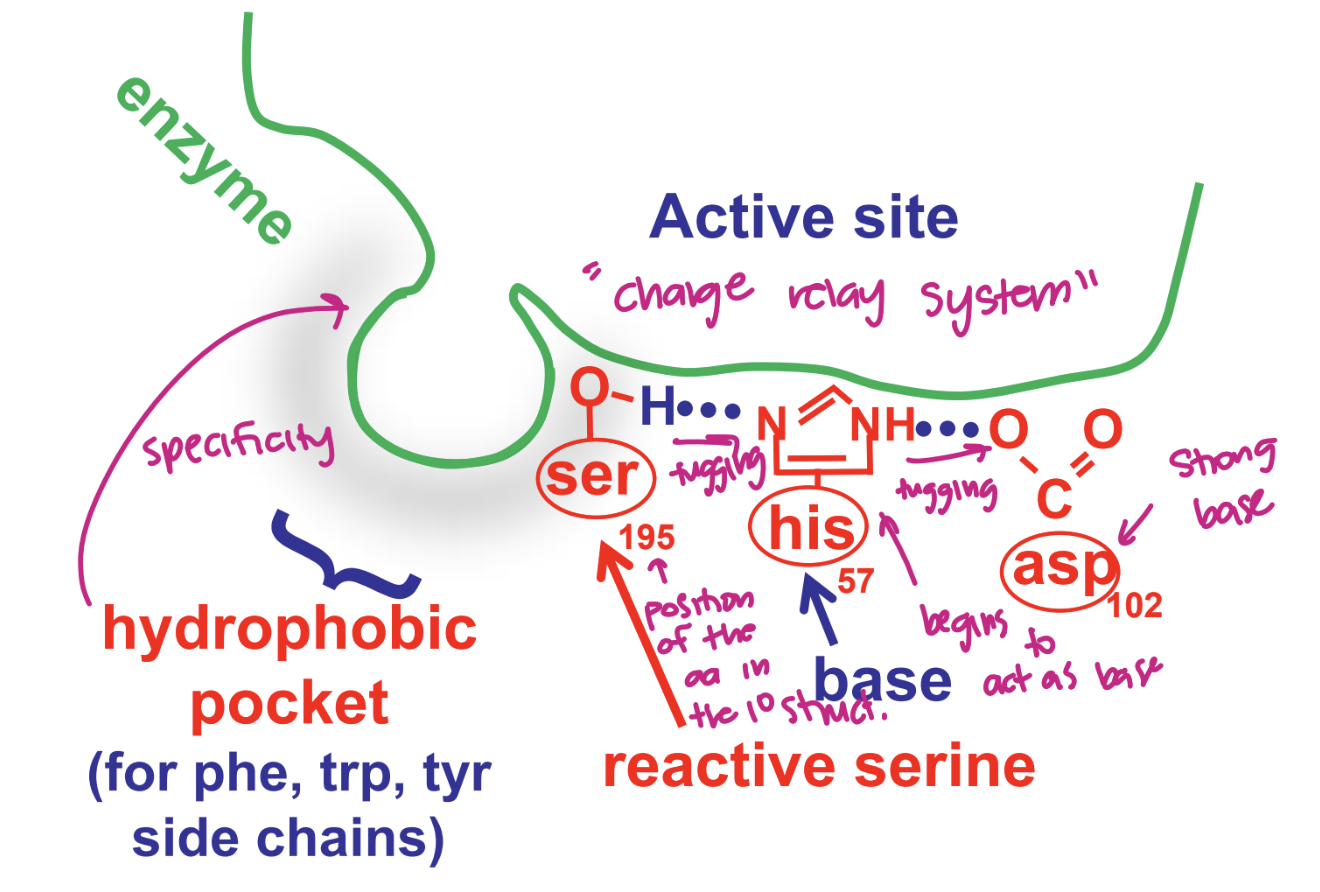
asp acts as strong base → O tugs NH on his (interaction makes his a stronger base) → which tugs H on ser (making ser a stronger acid) → making it reactive
change relay system
describe the covalent transition state
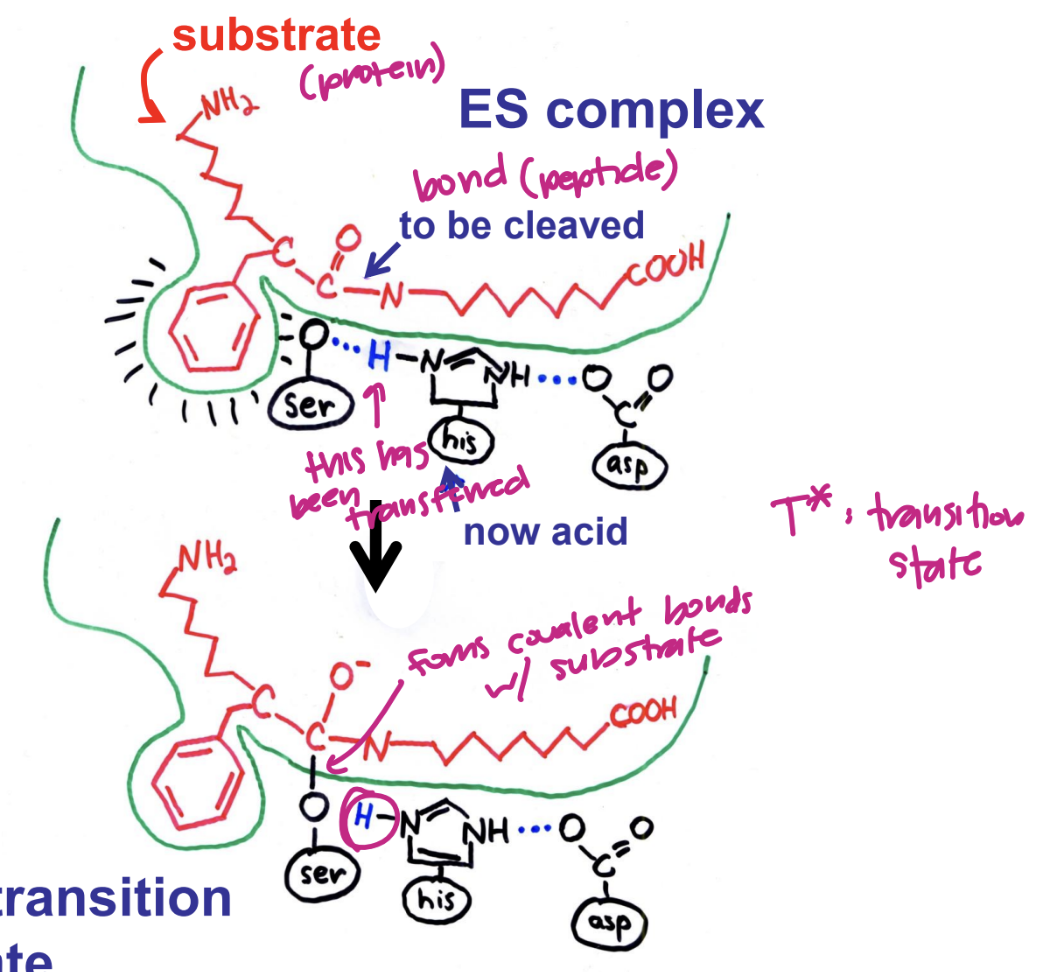
bond cleaves
H on ser has been transformed
forms covalent bonds with substrate
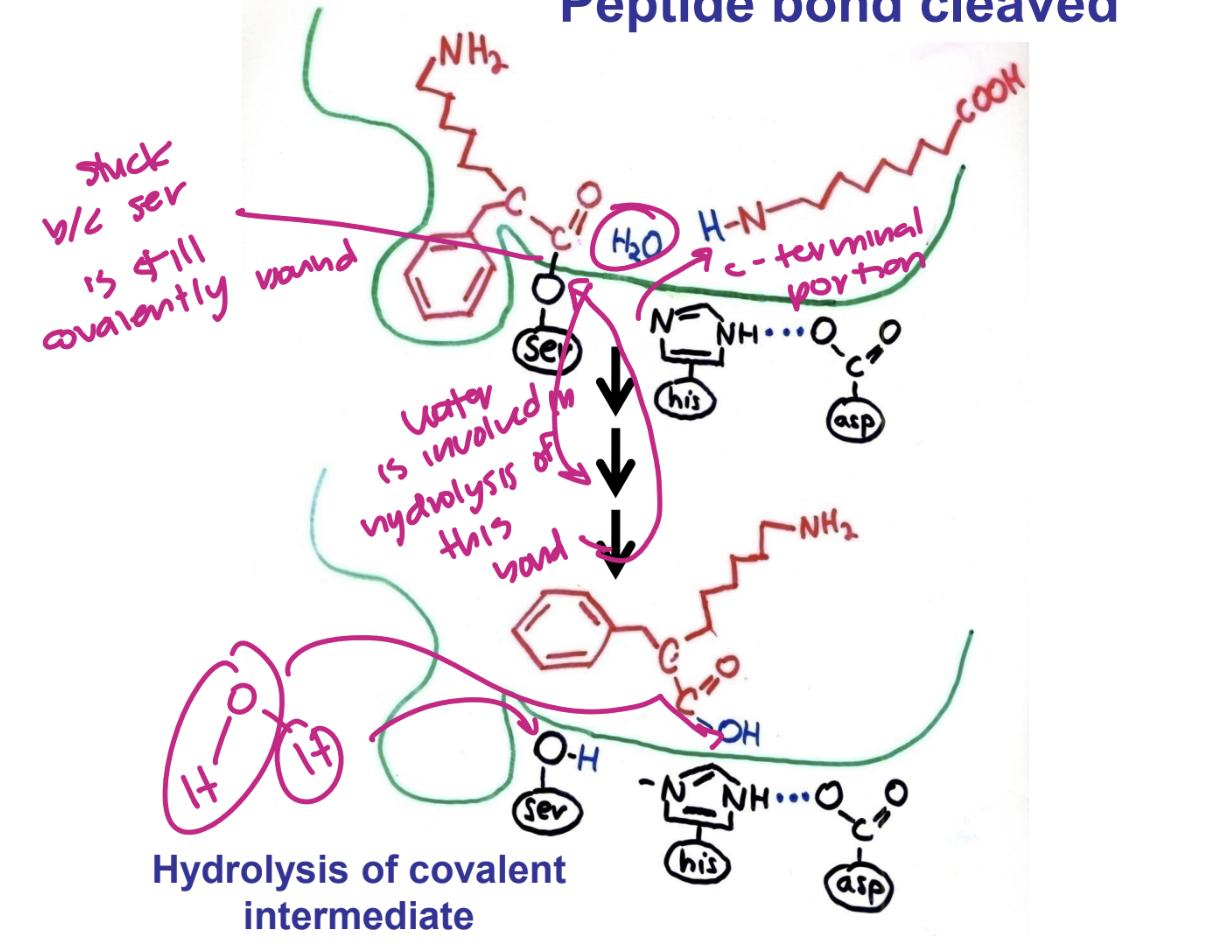
what is the specificity of Serine proteases determined by
binding pocket
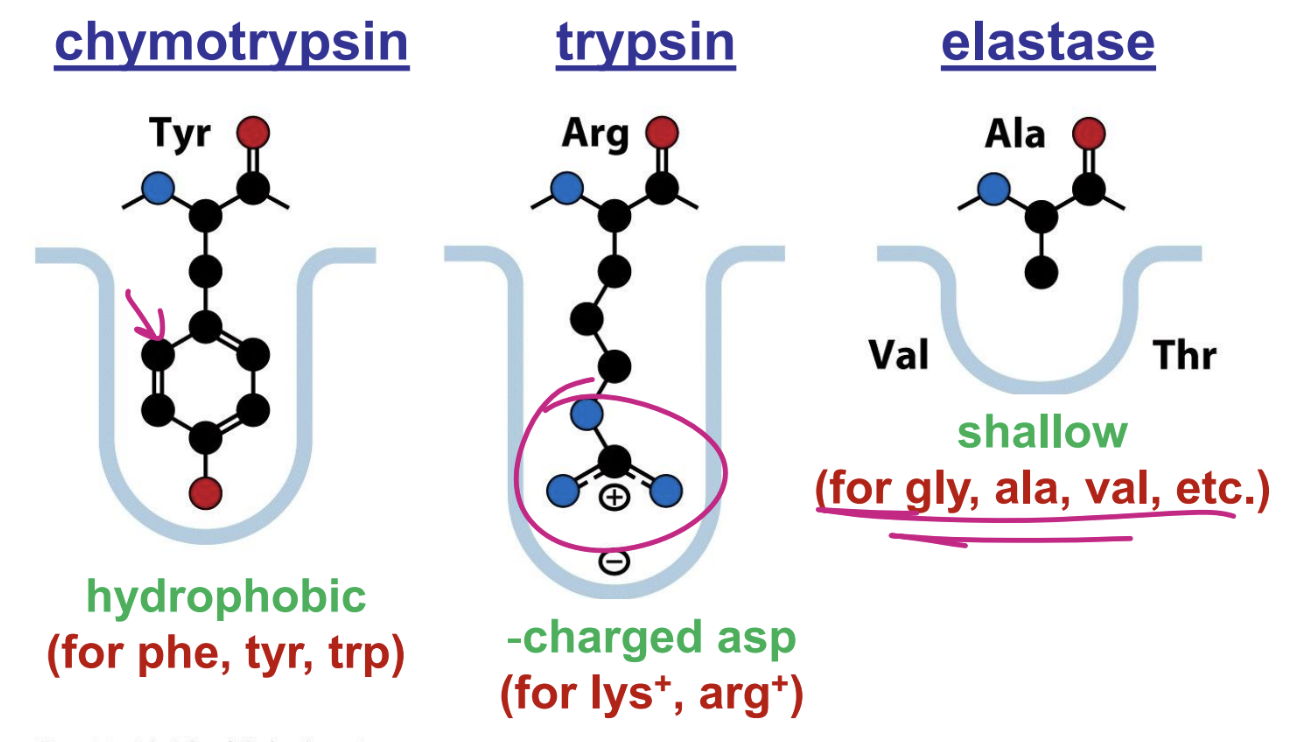
discuss enzyme kinetics
allows one to predict rates, given [S]
as [S] increase so does the rate
reaction mechanisms
reveals regulation
as initial stages, [S] >> [P] → rate is ~constant
velocity (delta P/ delta t) is dependent on [S] at low [S]; subtsrates to Vmax at high [S]
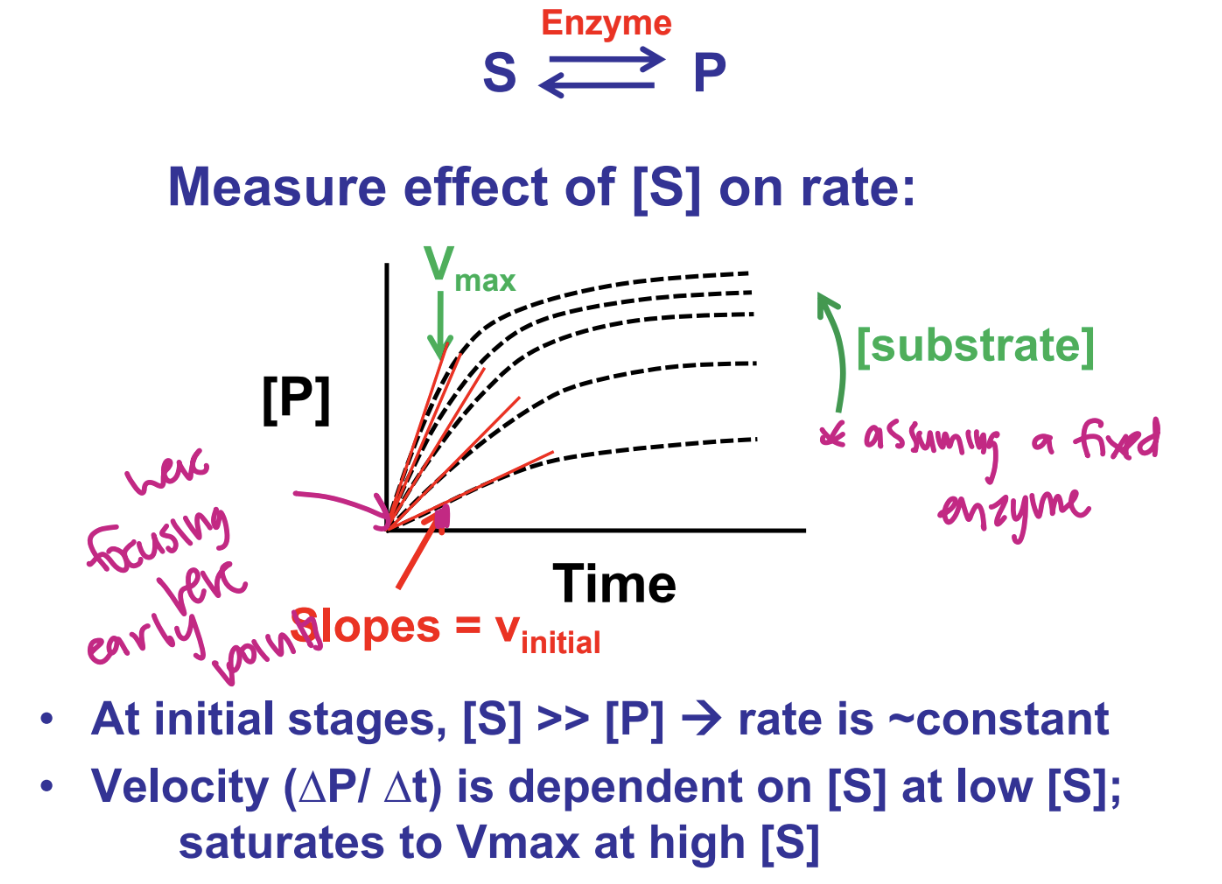
describe reaction velicity vs [S] what is the relationship between vinitial and [S]
Michaelis-Menten rate law

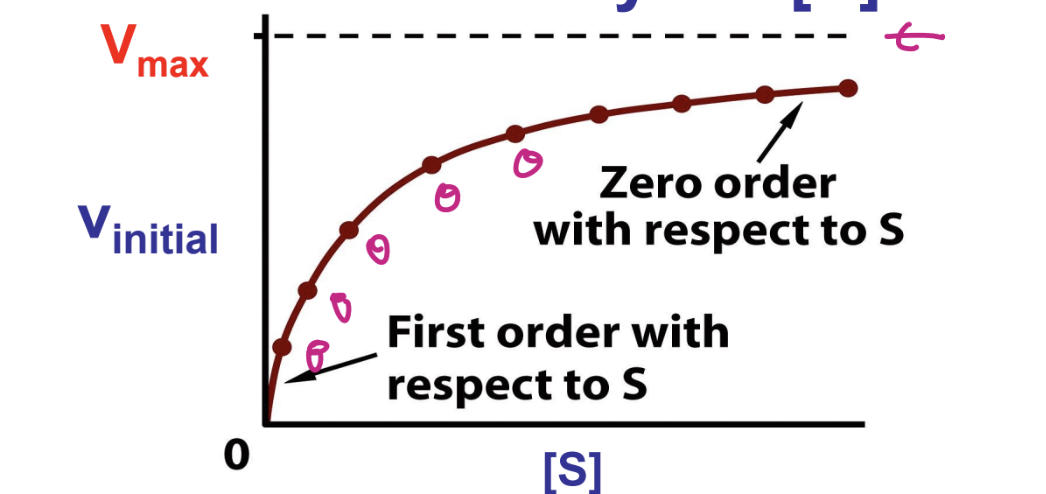
think about the conceptual view of enzyme saturation
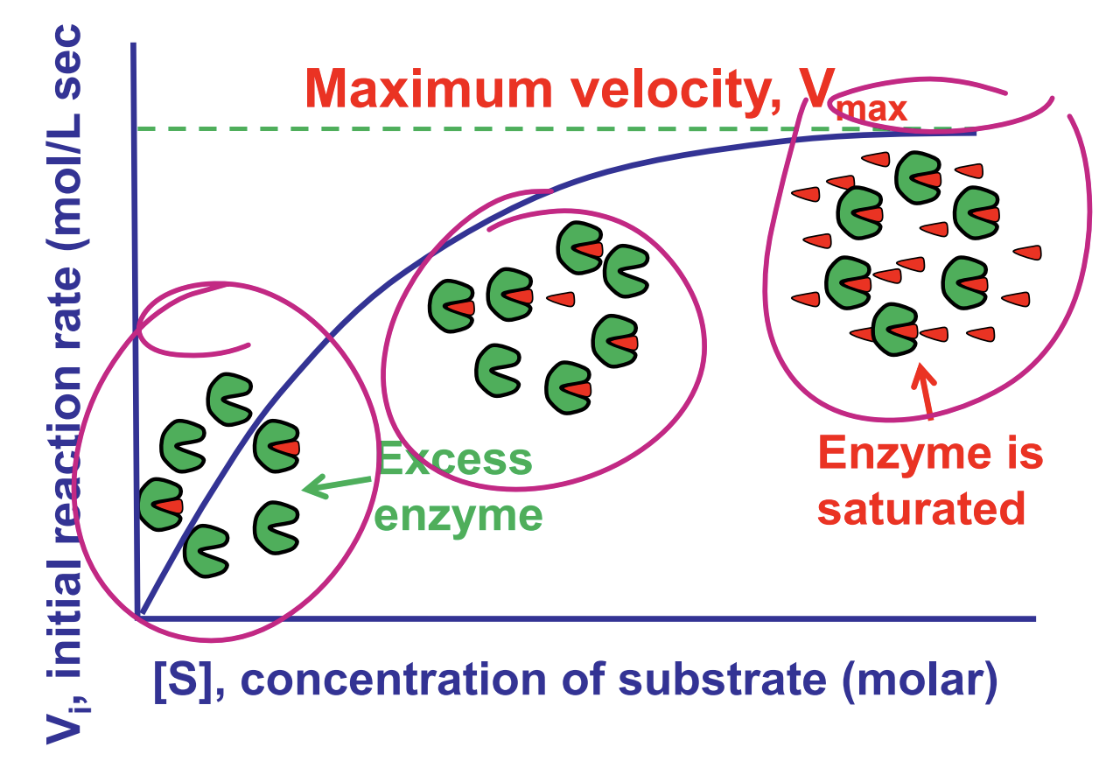
what does the michaelis mentain equation describe
observed behavior
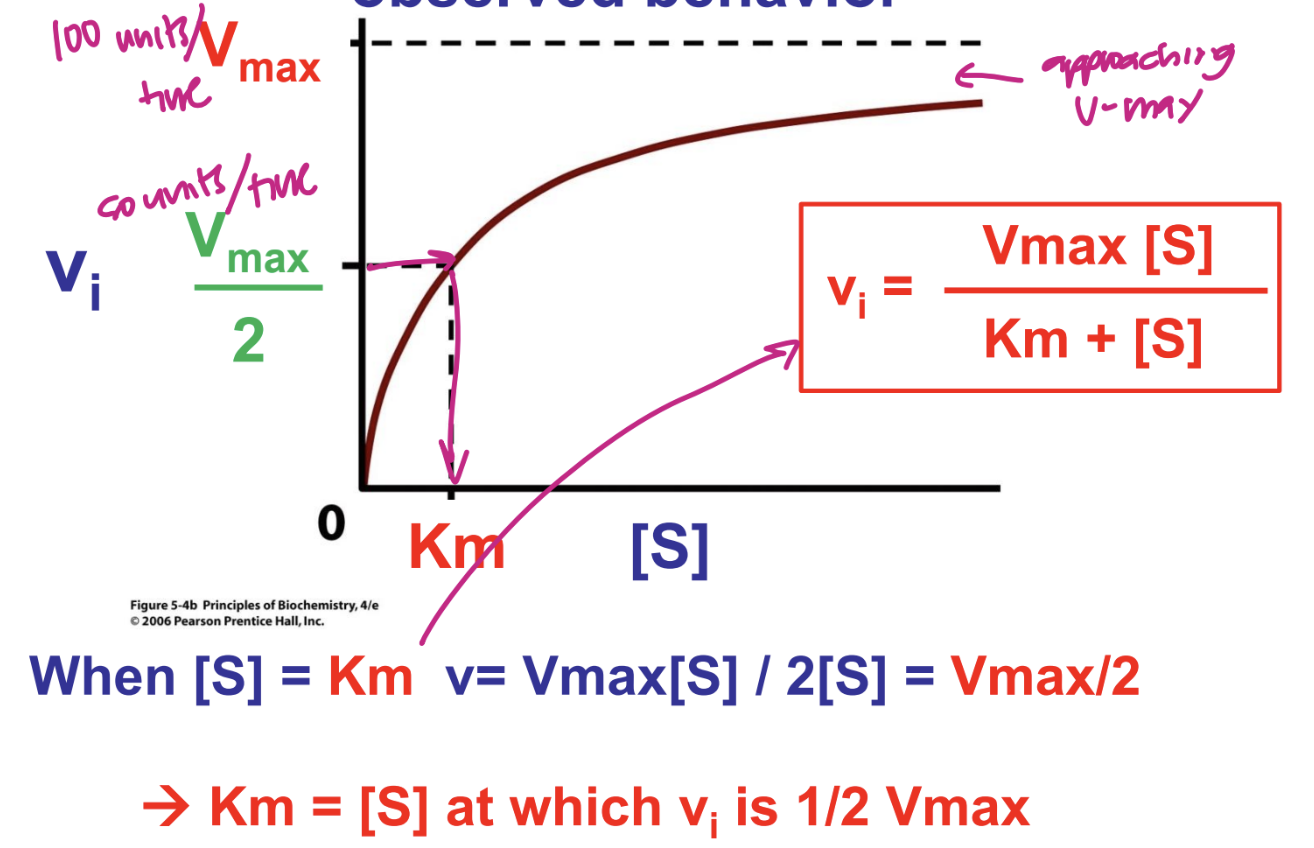
what is the meaning of Km
always [S] at which velocity is half maximal
often a measure of affinity of E for S
if k2<< k-1 then:
Km = (k-1+
k2)/k1 = (k-1)/k1 = Ks
where Ks is the dissociation constant for ES → E+S
low Km = high affinity
what is the meaning of Vmax
always maximum velocity when E is saturated with S (depends on amount of E)
often measure of rate constant for catalytic step
when k2<k1 then
Vmax = k2[Etotal]
for complex reactions kcat = rate constant for overall reaction
kcat = turnover number = # molecules S → P per second per E molecule
how fast an enzyme can complete one round of rxn
describe enzyme activation by proteolysis (protein breakdown)
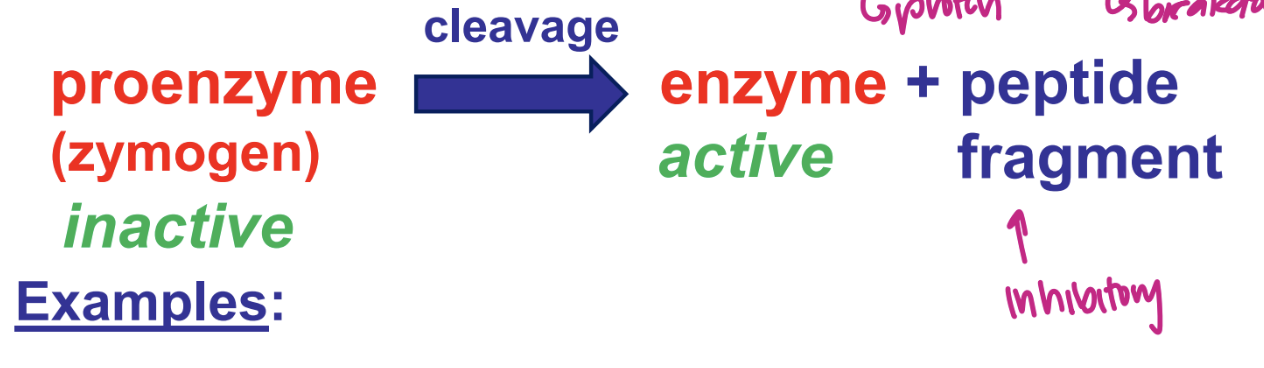
examples
digestive enzymes
blood clotting factors
programmed cell death
peptide hormones
proteolytic activation is irreversible
example: proteolytic activation of chymotrypsin
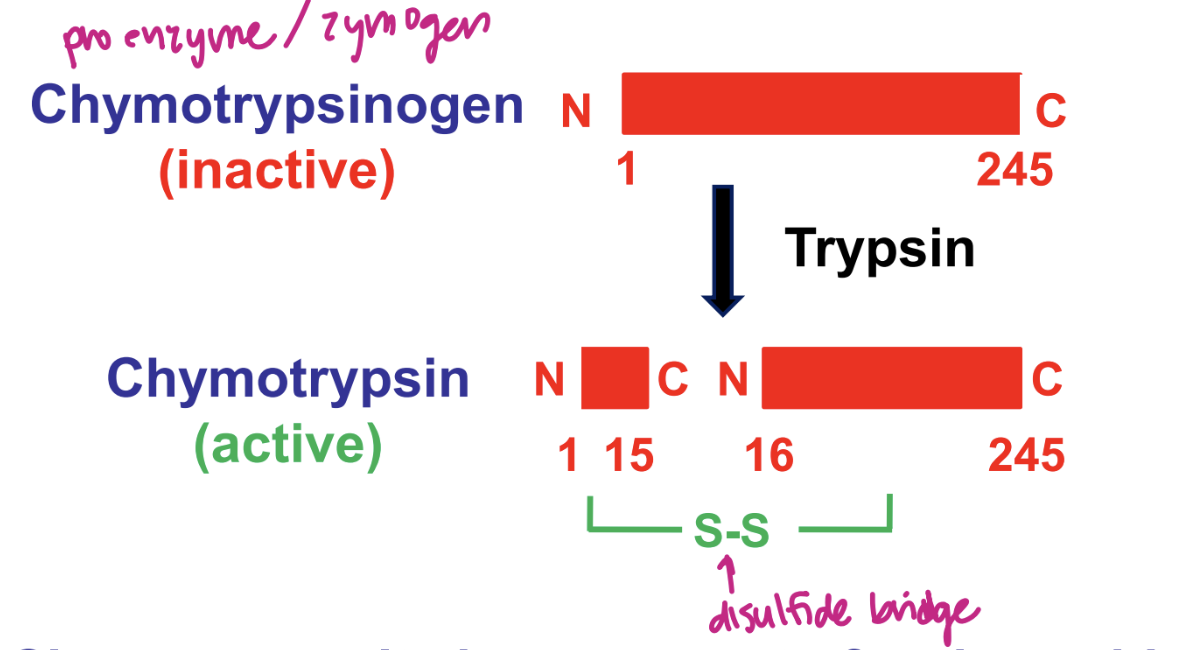
cleavage results in movement of amino acid that “plugs” active site away from active site
describe regulation of proteases by inhibitor peptides
small peptides that bind to the active site and inhibit protease function
example
anti-elastase, produced by the lungs, blocks connective tissue breakdown by elastase (emphysema)
describe the activation cascade in blood clotting
clotting factors (except fibrin) are serine proteases
activated by proteolytic cleavage
enormous amplification of very small signal
describe regulation of enzymes by covalent modification
usually reversible
can activate or inhibit enzyme
types
phosphorylation
adenylation/uridylylation
ADP ribosylation
methylation
acetylation
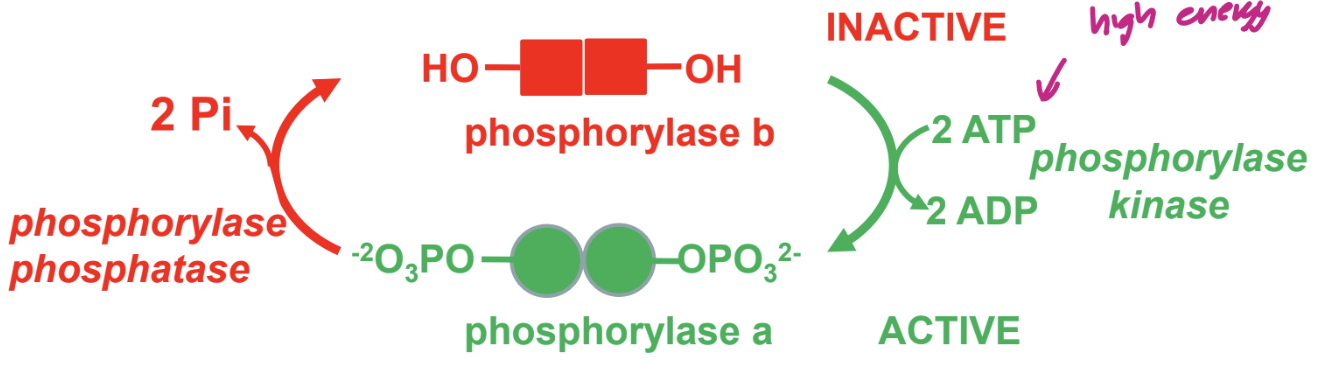
describe regulation of glycoge phosphorylase by phosporylation
glycogen phosporylase cleaves glycogen
→ releases glucose for energy
reegulated by phosphorylation
also regulated by hormone triggered phosphorylation cascafe and allosteric regulation
enzyme inhibitors (2 types)
non-specific: urea, detergents etc
specific: interact with one enzyme
antibodies (inibits enzyme only in bacteria)
drugs (lovistatin inhibits HMG-CoA redutase → insecta)
herbicides (target ezymes unique to plants or insects)
pesticides ^^
metabolites (feedback regulation)
specific inhibitors often resemble substrate or transition state
can be reversible or irreversible
discuss reviersible enzyme inhibition
inhibitor - a compound that binds E and interferes with its activity
can act by preventing formation of the ES complex or by blocking the chemical reaction that leads to the formation of product
what are the types of inhibitors
competetive: binds only to free E not bound to S (most common
non-competetive: can bind E or ES; not substrate analogs and do not bind same site as S
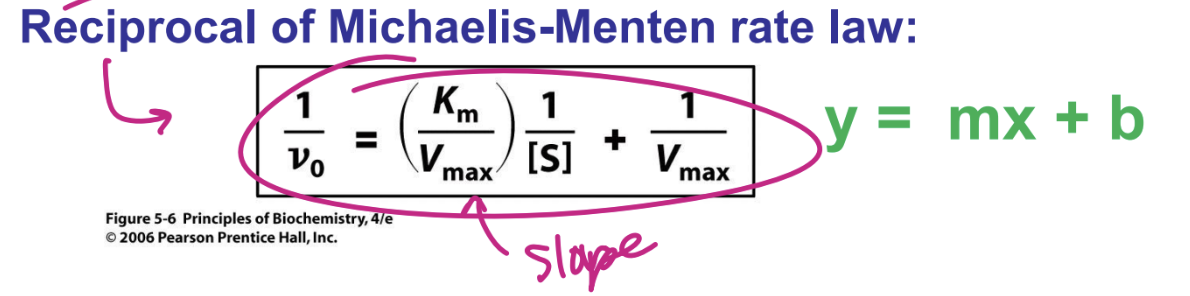
Determining Km and Vmax: Line weaver-Burk Linear plot
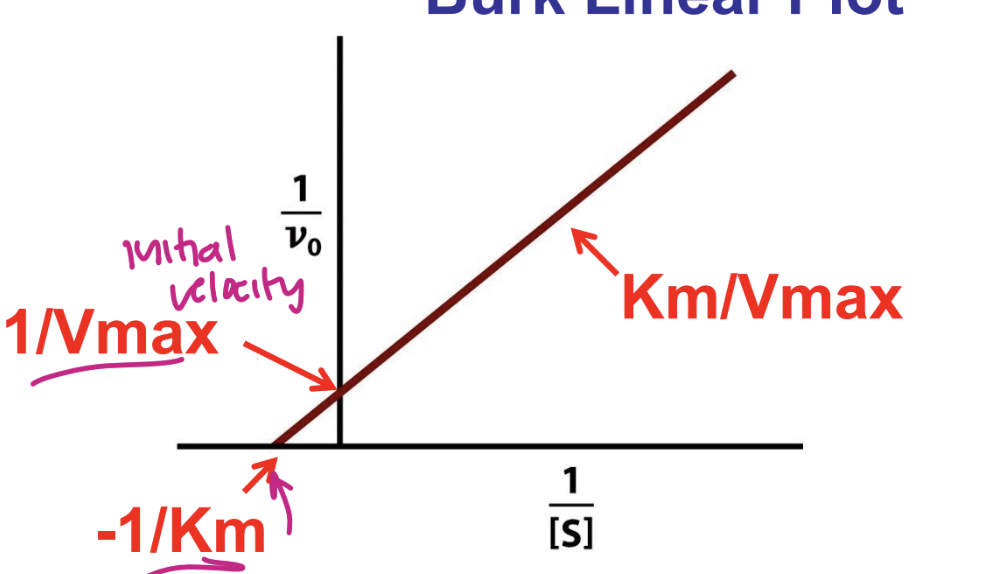
x intercept = -1/Km
y intercept = 1/Vmax
reciprocal of Michaelis Menten rate law
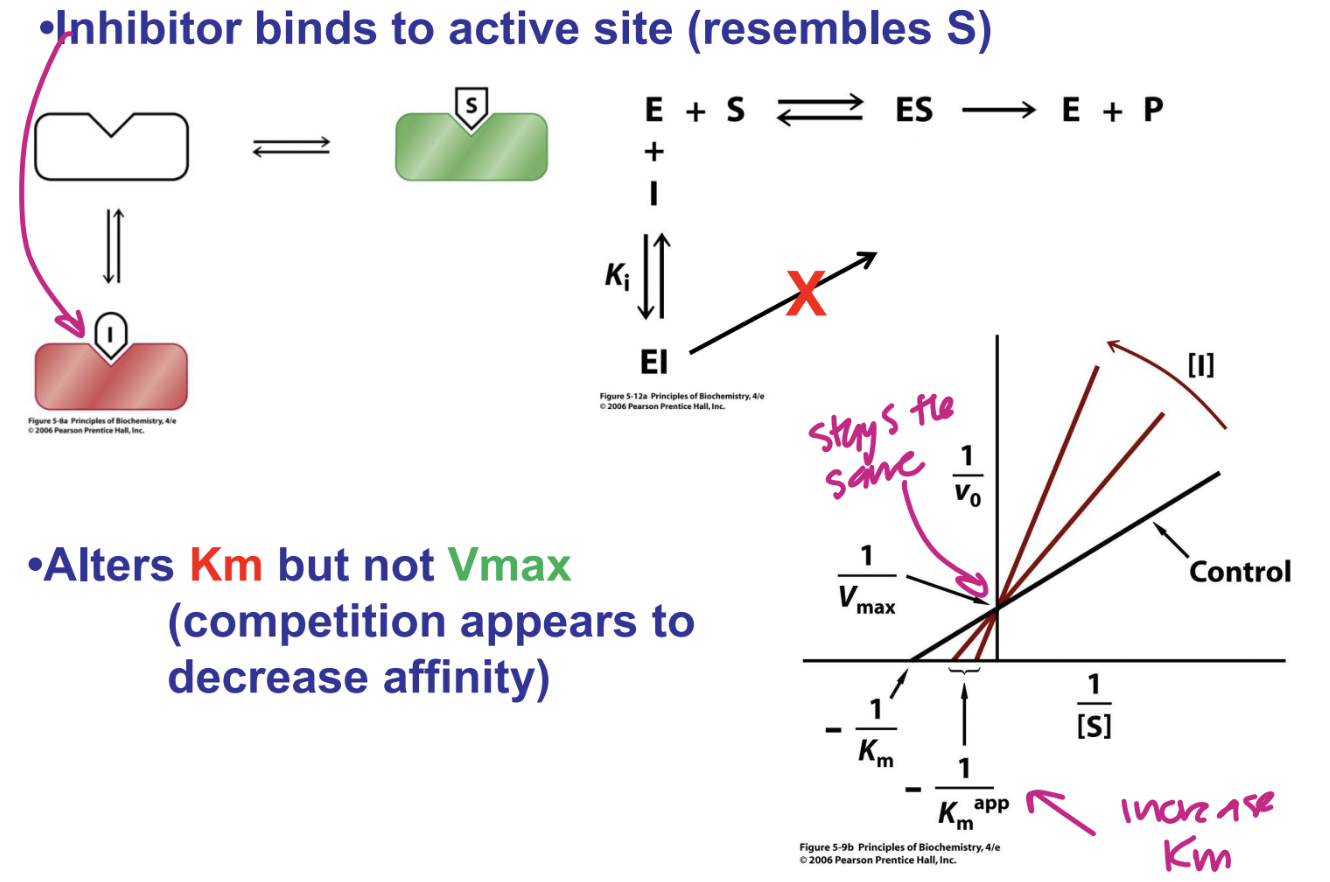
Describe competetive inhibitors
inhibitor binds to active site (resembles S)
Alters Km but not Vmac
competition appears to decrease affinity
ex. used to treat poisonings
Describe non-competitive inhibitors
alters Vmax but not km
cant drive to same vmac even with excess S
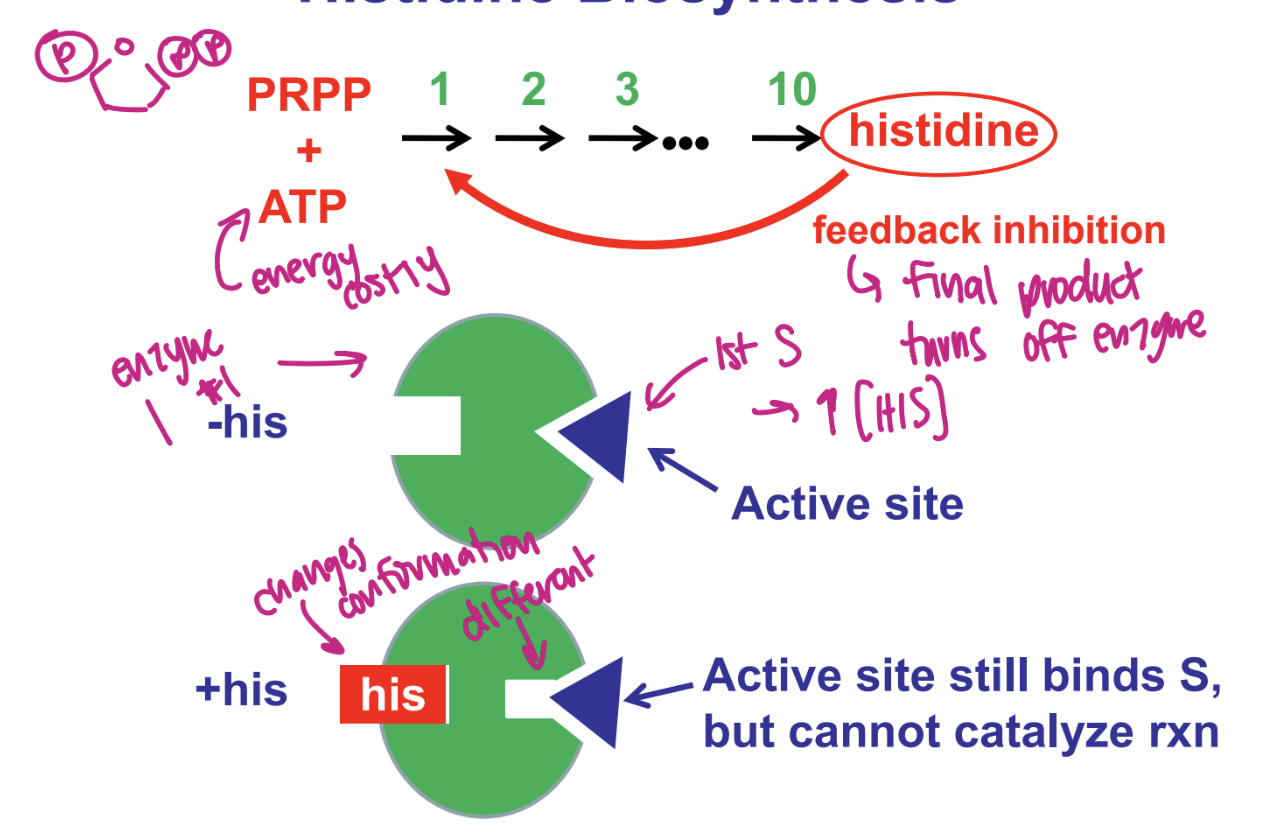
ex. histidine biosynthesis
describe allosteric regulation
modulators (activators or inhibitors) bind to allosteric site of enzyme ( seperate from actove site)
binding of modulator alters enzyme activity
can affect either Km or Vmax
allostery generally results from interactions among subunits of multimeric proteins → quaternary structure
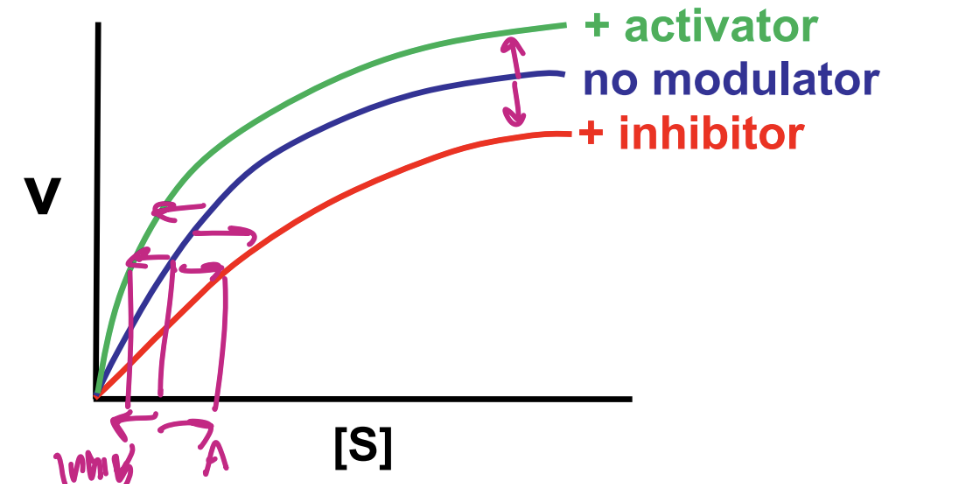
describe cooperative allosteric regulation
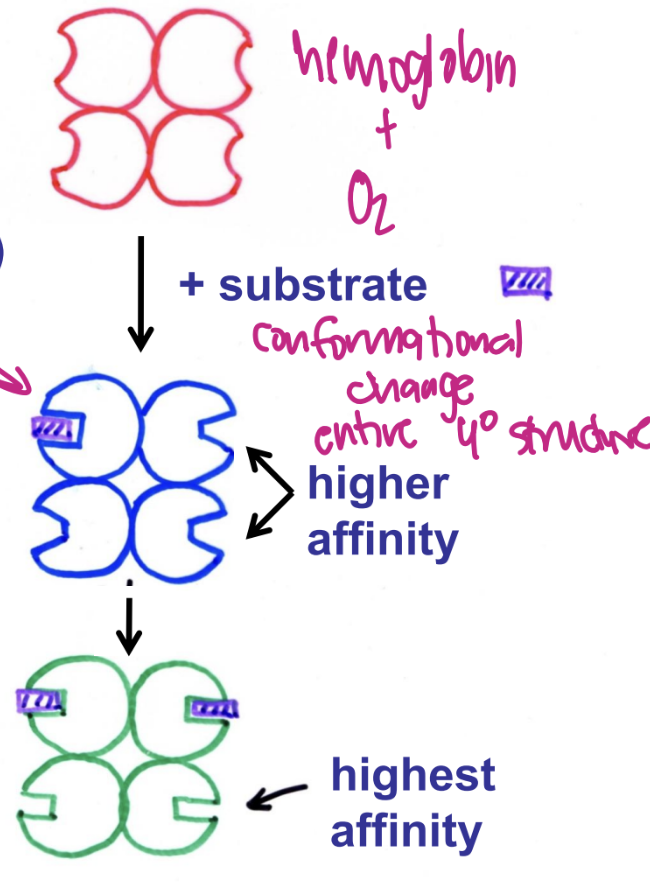
multimeric proteins
binding of effectir (modulator) to one subunit alters activity of others
ex. homotrophic allostery: effector = S
heterotrophic: effector and S are different
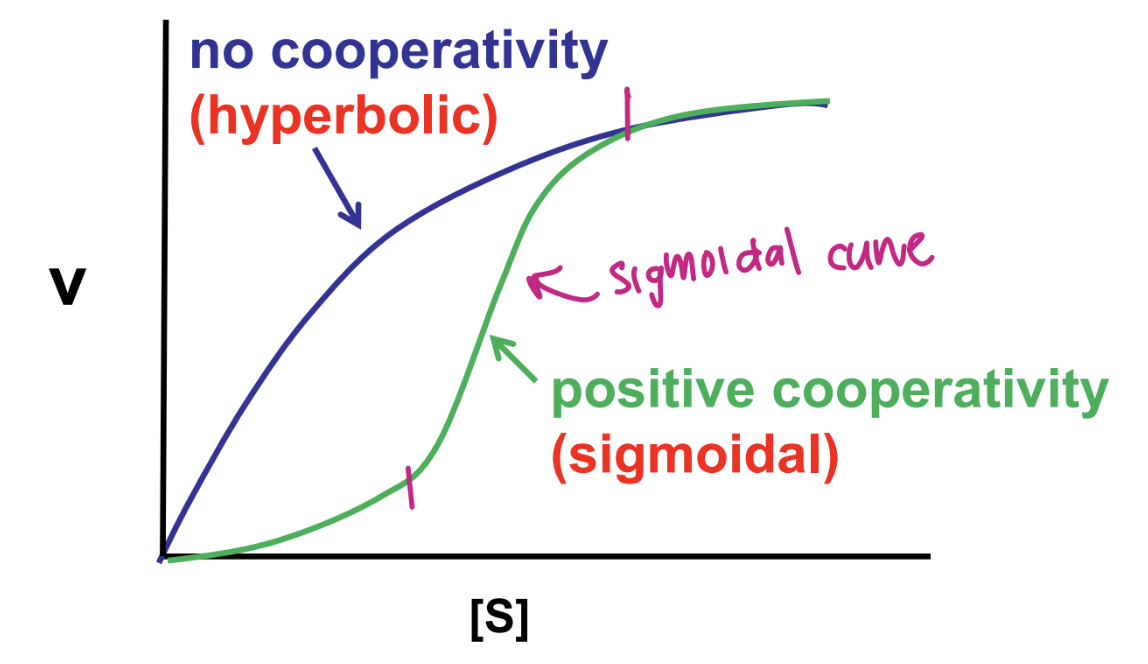
discuss cooperativity and kinetics
coopertivity does not follow michaels-menten kinetics
still can experimentally determine Km and Vmax
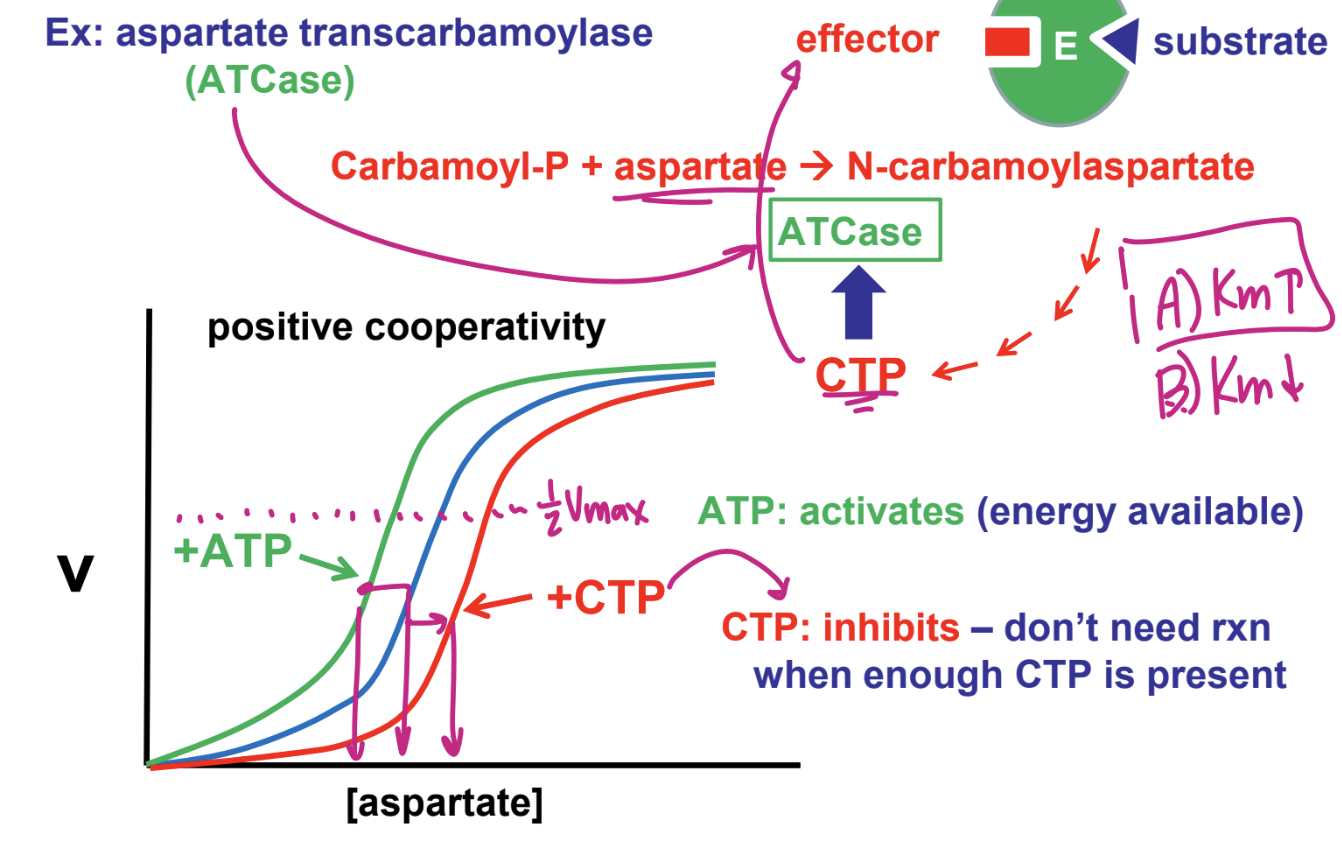
Discuss Allosteric regulation by heterotrophic (non-S) effectors (modulators)
ATP: activates (energy available)
CTP: inhibits (dont need rxn when enough CTP is present)
what is feedback inhibition
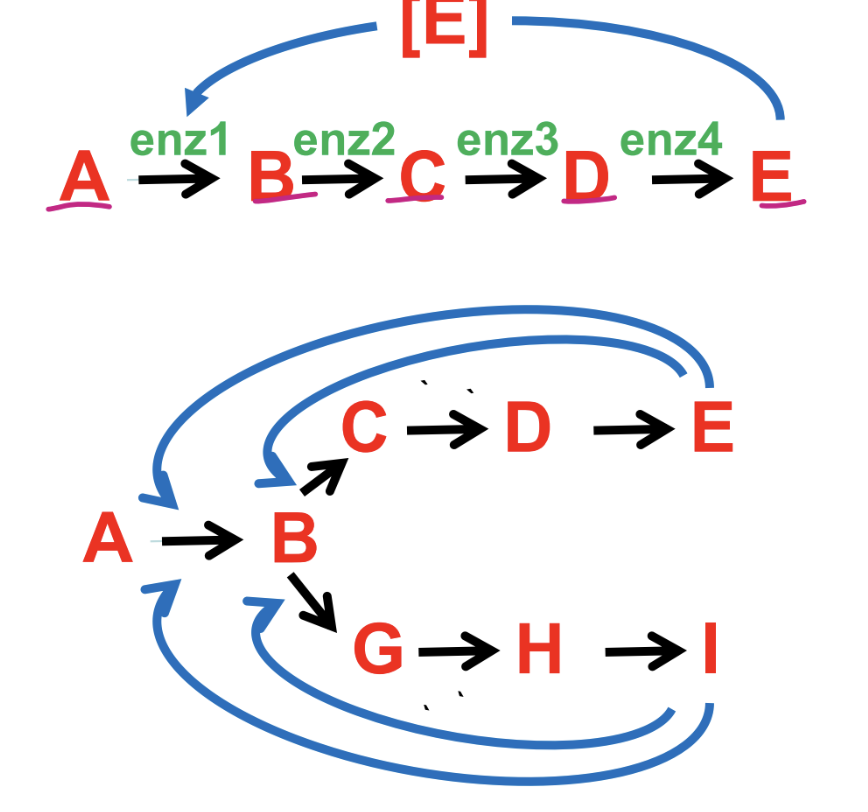
what is cumulatove feedback inhibition
cumulative effect of allosteric regulators decreases G.S. activity in stepwide fashion
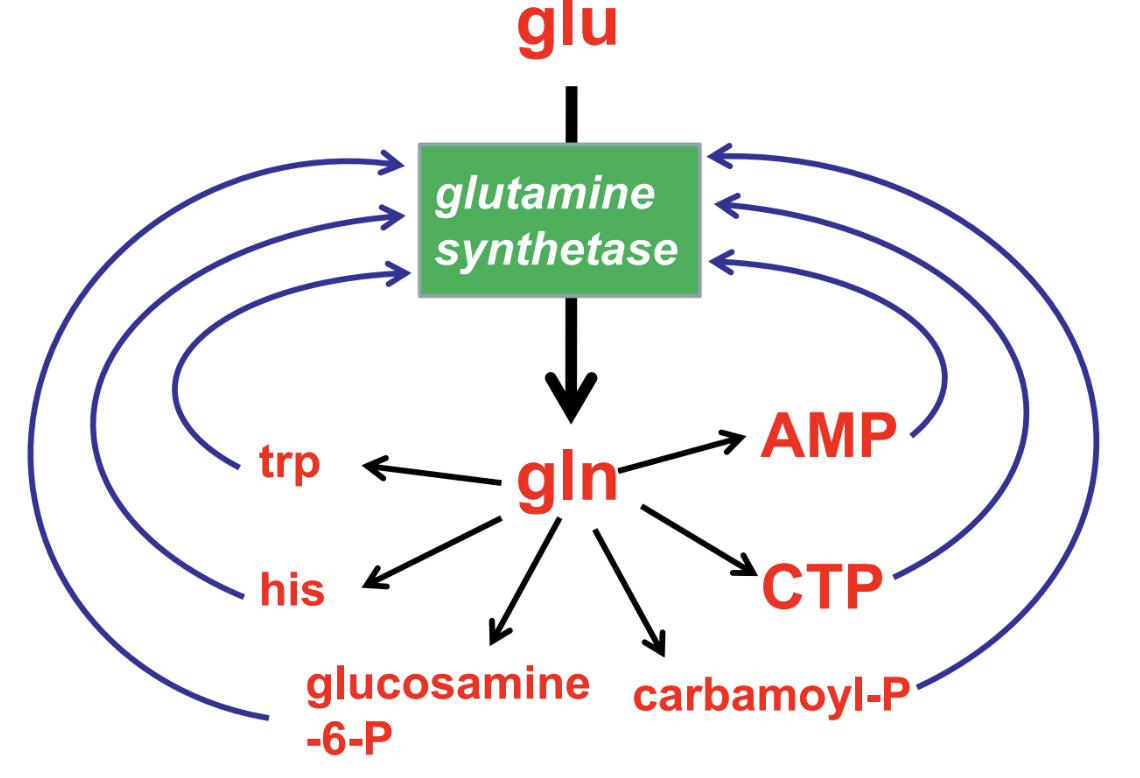
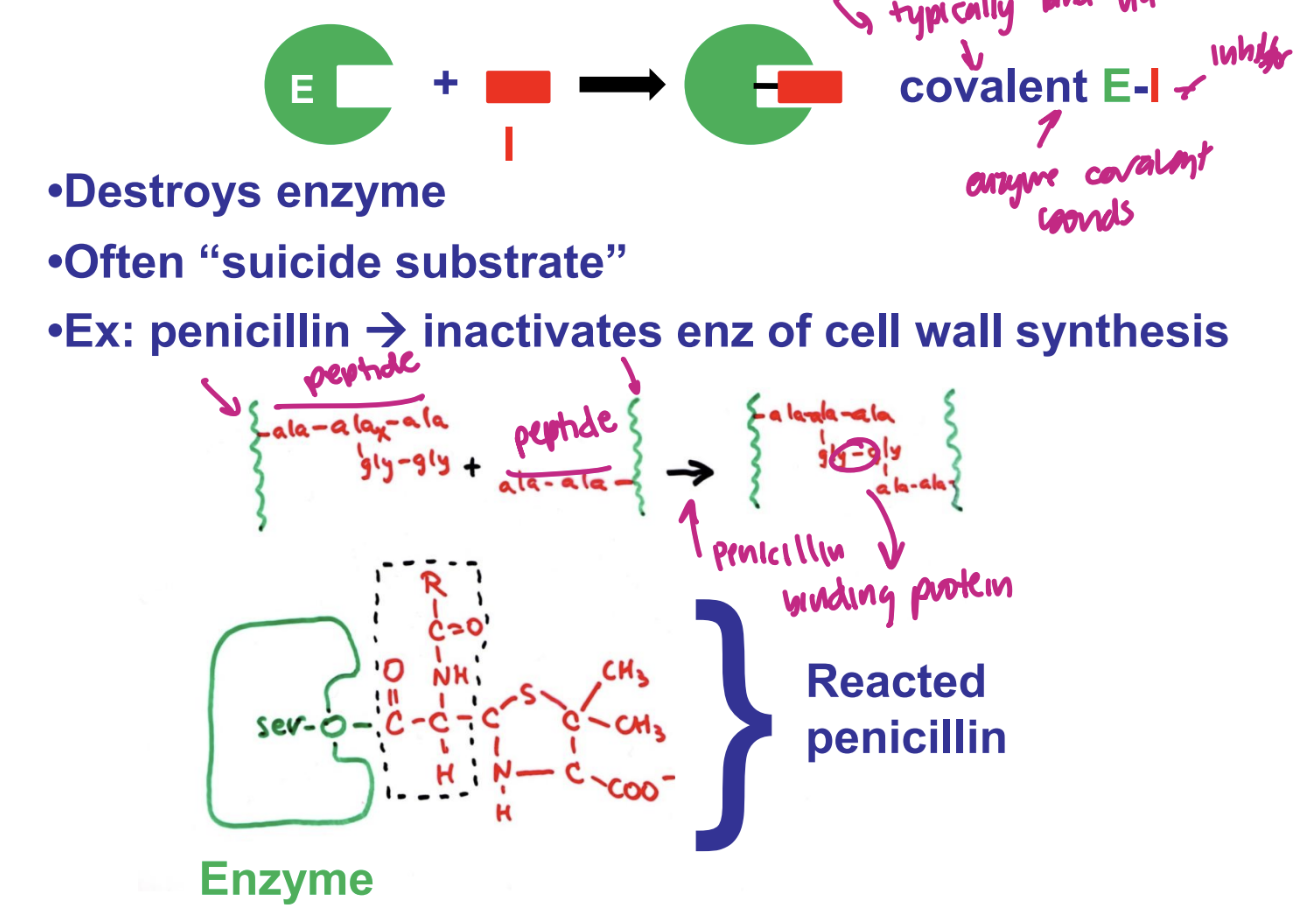
what are irreversible inhibitors
destroys enzyme
“suicide substrate”
ex. penicillin → inactivates enx of cell wall synthesis