Protein Quality Control
1/26
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
27 Terms
Chaperone proteins
Specialized proteins that assist in protein folding, refolding, and assembly into multimeric complexes
What occurs if folding fails
Unfolded or misfolded proteins might form disordered aggregates or highly ordered amyloid fibers
What results from persistent misfolded proteins
Apoptosis (programmed cell death) will be triggered if the protein quality control systems fail as well
Ubiquitin proteasome complex (UPS)
The mechanism for degrading misfolded proteins from the cytoplasm and the ER via the endoplasmic reticulum-associated protein degradation (ERAD)
Autophoagy
Degrades larger protein aggregates that cannot be process by UPS
Lon and Clp Proteases
ATP-dependent proteases that degrade defective or short-lived proteins, found in mitochondria (eukaryotes) and bacteria
How does the half life differ of proteins (ex: hemoglobin vs defective proteins)
Half-lives can range from seconds to months:
Hemoglobin is long-lived whereas defective proteins are short-lived,
Ubiquitin
A small, highly conserved protein that tags substrates for degradation via the proteosome
How is ubiquitin prepared before the degradation process
Ubiquitin is covalently attached to proteins slated for destruction; the free carboxy terminus of ubiquitin is linked to a Lys sidechain in a target protein
The first step of the ubiquitination process
Activation: The free carboxyl group of ubiquitin’s carboxyl-terminal Gly residue first becomes linked to an E1-class activating enzyme via a thioester
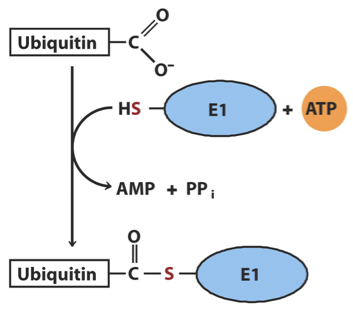
The second step of the ubiquitination process
Conjugation: Ubiquitin is transferred to an E2 conjugating enzyme
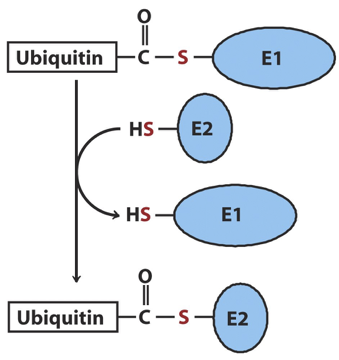
The third step of the ubiquitination process
Ligation: An E3 ligase catalyzes the transfer of ubiquitin from E2 to the target, linking ubiquitin through an amide (isopeptide) bond to an ε-amino group of a Lys residue of the target protein
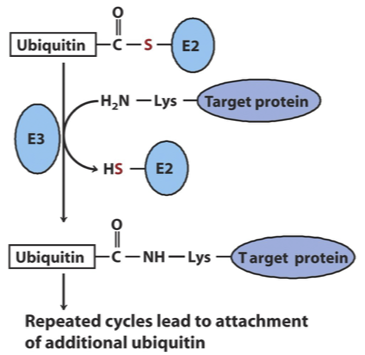
What do additional cycles of ubiquitination produce and what is its function
Polyubiquitin, a covalent polymer of ubiquitin subunits that targets the attached protein for destruction in eukaryotes
Proteasome function
A large, ATP-dependent molecular machine that degrades unneeded or damaged proteins into short peptides
Proteasome structure
A large 4-ringed barrel-shaped complex made of many polypeptide chains with several proteases with different substrate specificities inside
N-end rule
The N-terminal amino acid of a protein influences its half-life by serving as a degradation signal recognized by ubiquitin ligases
N-end rule importance
It provides an estimate of protein stability but vary due to factors such as -terminal modifications or organismal differences
Lon (long form) proteases
Highly conserved, ATP-dependent serine proteases that hydrolyze defective or short-term peptides
Where are lon proteases found
They are found in archaea, bacteria, and eukaryotes (in the mitochondrial matrix)
Clp proteases function in Bacteria
They degrade proteins marked for destruction, by recognizing specific degradation signals
Clp proteases function in Bacteria process:
Adaptor proteins (ClpS) recognize and bind to exposed N-end rule residues in substrates, delivering them to Clp proteases
SsrA-tagged proteins resulting from stalled ribosomes during translation are rescued by the rmRNA pathway
The ssrA peptide ag directs these proteins to ClpXP protease via the SspB adaptor for degradation
The GroEL/GroES system
A bacterial chaperonin complex that facilitates protein folding, preventing aggregation under stress
The GroEL/GroES steps (1)
Substrate proteins in a kinetically trapped but aggregated state are delivered to GroEL by the DnaK/DnaJ (Hsp 70) system
The GroEL/GroES steps (2)
Upon binding, substrate proteins undergo partial unfolding into expanded and compact conformations
The GroEL/GroES steps (3)
ATP-driven conformational changes in GroEL apical domains stretch tightly bound substrate regions and release less stable parts
The GroEL/GroES steps (4)
GroES caps the GroEL chamber, encapsulating the substrate and allowing it to fold within a protected environment
The GroEL/GroES steps (5)
Substrate proteins are released upon GroES dissociation; incompletely folded proteins may rebind for further folding cycles, helping proteins reach their native state