Prokaryotic Transcription
1/324
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
325 Terms
After replication fo DNA what is thenext step in the central dogma theoryu
transcription

what RNA molecule contains a sequence of bases that encodes the primary amino acid sequence for a protein while serving as the template for translation by a ribosome
Messenger RNA (mRNA)
what RNA molecule carries an amino acid into the catalytic site of a ribosome.
it base pairsa to mRNA to ensure the selection of the correct amino acid for incoportation into a nascent polypeptide
transfer RNA (tRNA)
what RNA molecule are structural components of a ribosome, the enzyme that cataclyzes translation
ribosomal RNA (rRNA)
what RNA molecule does nt encode proteins but have a variety of catalyticx structural, and regulatory function
Noncoding RNAs (ncRNAs)
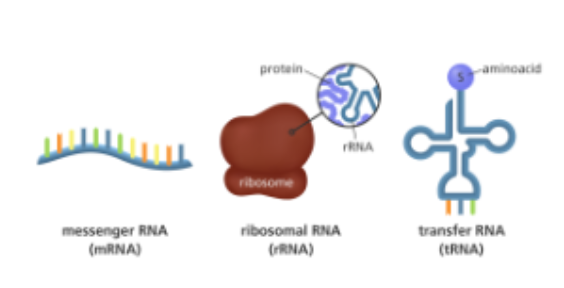
what are central to how geentic information stored in DNA is actually turned into functional proteins and each type plays a distinct by highly coordinated role in this process
RNA molecules

Messenger RNA (mRNA) is essentially the “working copy” of a gene: it is synthesized from DNA during transcription and carries a specific sequence of nucleotide bases that directly encodes the order of amino acids in a protein
How is this sequence read?
in groups of 3 bases called codons, each of which respondd to a specific amino acid
Once produced, what happens to mRNA
mRNA travels to a ribosome where it serves as the template for protein synthesis ( translation)
Once produced, the mRNA travels to a ribosome, where it serves as the template for protein synthesis, a process known as translation. However, mRNA cannot build a protein on its own
What becomes essential?
transfer RNA (tRNA)
What is each tRNA molecule attached to
a specific amino acid
What regions do tRNAs contain
a region called an anticodon that is complimentary to a codon on the mRNA
Through precise base pairing, what does the tRNA ensure?
that the correct amino acid is delievered to the growing polypeptide chain maintaing the fidelity of protein synthesis
Through precise base pairing, the tRNA ensures that the correct amino acid is delivered to the growing polypeptide chain, maintaining the fidelity of protein synthesis.
Where is thr actual site where this assembly occurs
the ribosome
what is the ribosom elargely composed of
ribsomal RNA (rRNA) and proteins
What is important to know about rRBA
it is not just structural— it also has catalytic activity
Through precise base pairing, the tRNA ensures that the correct amino acid is delivered to the growing polypeptide chain, maintaining the fidelity of protein synthesis.
The actual site where this assembly occurs is the ribosome, which is composed largely of ribosomal RNA (rRNA) along with proteins. Importantly, rRNA is not just structural—it also has catalytic activity…
What does this mean?
it directly participates in forming peptide bonds between amino acids, making the ribosome a ribozyme (an RNA enzyme)
Cells also contain a wide variety of noncoding RNAs (ncRNAs)
What are ncRNAs essential for if they are not translated into proteins
crucial for regulating gene expression and maintaing cellular function
Beyond these classic roles, cells also contain a wide variety of noncoding RNAs (ncRNAs), which are not translated into proteins but are crucial for regulating gene expression and maintaining cellular function.
What molecules are included in this
molecules that can modify other RNAs
molecules that regualte trancxcription or translation
catalyze biochemical reactions
Altogether, RNA is not just an intermediary between DNA and protein
What is it?
it is a dynamic and multifunctional set of molecules that drive and regulate nearly every step of gene expression.
What is the DNA encoding the protein + DNA regulatory elements needed for transcription
Gene
What is a gene used to create an RNA primariy transcript
Transcription
what is the significance of bateria transcripton
the primary transcript is mrna ready fro translation
what us the significance of eukarytoe trancrption
the primary transcript is NOT the translation- ready mRNA
What is a continous sequence from start cidin to stop codon
an open reading frame (ORF)
What is a single unit that procduces a macrocolecule like a protein
cistron
what tytpe of mrna do prokaryotes have
polycistronic mRNA
what type of mRNA do eukaryotes have
monocistronic mRNA
What is a sequence directly correlating to protein primary sequence
coding sequence
True or False: All CDS are ORFs, but not all ORFs are CDS
True
A gene is not just the portion of DNA that directly codes for a protein, what does it also include
important regulatory DNA sequences ( like promoters and enchancer) that control when, where, and how much of that protein is made
A gene is not just the portion of DNA that directly codes for a protein—it also includes important regulatory DNA sequences (like promoters and enhancers) that control when, where, and how much of that protein is made.
The process begins with transcription.
What happens during transcription
the information in DNA is copied into an RNA molecule called the primary transcript
The process begins with transcription, where the information in DNA is copied into an RNA molecule called the primary transcript
What is this process like in bacteria
his process is very streamlined: the primary RNA transcript is already essentially a finished mRNA, meaning it can be immediately used by ribosomes for protein synthesis (translation)
What happens to the primary transcript in eukaryotic cells
the primary transcript pre-mRNA must undergor several processing steps
What are the several processing steps that the pre mRNA in eukaryotic cells go through beopre it becomes a mature, translation-ready mRNA ?
splicing
capping
polyadenylation
Within any mRNA, what is the region that can actually be translated into a protein?
open reading frame (ORF)
what is an open reading frame?
a cnotinuous stretch of nucleotides that begins with a start codon, usually (AUG) and ends at a stop codon without interruption
Not every ORF is necessarily used by the cell to make a protein. What refers specifically to the ORF that is actually translated into a protein
the coding sequence
The coding sequence (CDS) refers specifically to the ORF that is actually translated into a protein, so while all coding sequences are ORFs, what does that mean for others?
some ORFs may never be expressed and therefore not true coding sequences
What is a functional gene unit that produces a single macrmolecule, typically a protein
cistron
Another important concept is the cistron, which is essentially a functional gene unit that produces a single macromolecule, typically a protein. This distinction becomes especially important when comparing organisms
What is this in relation to prokaryotes
prokaryotes often produce polycistronic mRNA
what is polycistronic mRNA
a single mRNA molecules contains multiple cistrons and can encode severak different proteins (usuallyt related in fucntion, like enzymes in the same pathway)
Another important concept is the cistron, which is essentially a functional gene unit that produces a single macromolecule, typically a protein. This distinction becomes especially important when comparing organisms
What is this in relation to eukaryotes?
eukaryotic mRNA is typically monocistronic
what does it mean to be monocistronic
each mRNA encodes only one protein
Another important concept is the cistron, which is essentially a functional gene unit that produces a single macromolecule, typically a protein.
This distinction becomes especially important when comparing organisms: prokaryotes often produce polycistronic mRNA, meaning a single mRNA molecule contains multiple cistrons and can encode several different proteins (usually related in function, like enzymes in the same pathway).
In contrast, eukaryotic mRNA is typically monocistronic, meaning each mRNA encodes only one protein. Altogether, this reflects a major difference in gene organization and regulation between prokaryotes and eukaryotes
What is the relevence with prokaryotes?
prokaryotes favor efficiency and coordination
Another important concept is the cistron, which is essentially a functional gene unit that produces a single macromolecule, typically a protein.
This distinction becomes especially important when comparing organisms: prokaryotes often produce polycistronic mRNA, meaning a single mRNA molecule contains multiple cistrons and can encode several different proteins (usually related in function, like enzymes in the same pathway).
In contrast, eukaryotic mRNA is typically monocistronic, meaning each mRNA encodes only one protein. Altogether, this reflects a major difference in gene organization and regulation between prokaryotes and eukaryotes
What is the relevence with eukaryotes
they favor more complex and tightly controlled gene expression
What are coordinately regulated gene clusters that are functionally related and regulated as a single unit
Operons
What is prokarytic gene organization built for?
efficiency and coordination,
Prokarytic gene organization built for efficiency and coordination what does this allow for cells to do?
allowing cells to rapidly respond to environmental changes by expressing groups of related genes together
What is one of the most important features of the prokaryotic gene organization
operon
Instead of each gene having its own promoter, what does an operon use?
one shared promoter and one terminator
Instead of each gene having its own promoter, an operon uses one shared promoter and one terminator, what does this mean
the entire group of genes is transcribed together into a single RNA molecule
Instead of each gene having its own promoter, an operon uses one shared promoter and one terminator, meaning the entire group of genes is transcribed together into a single RNA molecule.
Within this transcription unit, what are arranged sequentially in the 5’ →3’ direction each eachi corresponding to a different protein
open reading frames (ORFs)
Within this transcription unit, multiple open reading frames (ORFs) are arranged sequentially in the 5′ → 3′ direction, and each ORF corresponds to a different protein.
Because of this arrangement, what happens the resultigng mRNA
polycistronic mRNA
Within this transcription unit, multiple open reading frames (ORFs) are arranged sequentially in the 5′ → 3′ direction, and each ORF corresponds to a different protein. Because of this arrangement, the resulting mRNA is called polycistronic mRNA
What does that mean?
it contains multiple coding reguons (cistrons) , each of which can be translated into a seperate protein
Within this transcription unit, multiple open reading frames (ORFs) are arranged sequentially in the 5′ → 3′ direction, and each ORF corresponds to a different protein. Because of this arrangement, the resulting mRNA is called polycistronic mRNA, meaning it contains multiple coding regions (cistrons), each of which can be translated into a separate protein.
How is this is a major distinction from eukaryotic systems
where each mRNA typically encodes only one protein
Within this transcription unit, multiple open reading frames (ORFs) are arranged sequentially in the 5′ → 3′ direction, and each ORF corresponds to a different protein.
Because of this arrangement, the resulting mRNA is called polycistronic mRNA, meaning it contains multiple coding regions (cistrons), each of which can be translated into a separate protein.
This is a major distinction from eukaryotic systems, where each mRNA typically encodes only one protein.
What is a classic example of this organization
the e coli ATP operon
What makes the ATP operon an example of different organization within different species
which contains nine ORFs that collectively encode the different subunits of the F₁F₀ ATP synthase complex.
Within this transcription unit, multiple open reading frames (ORFs) are arranged sequentially in the 5′ → 3′ direction, and each ORF corresponds to a different protein.
Because of this arrangement, the resulting mRNA is called polycistronic mRNA, meaning it contains multiple coding regions (cistrons), each of which can be translated into a separate protein.
This is a major distinction from eukaryotic systems, where each mRNA typically encodes only one protein.
A classic example of this organization is the E. coli atp operon, which contains nine ORFs that collectively encode the different subunits of the F₁F₀ ATP synthase complex.
What do propkarytoic cells ensuyre by organizing it this way
that all components of a multi-subunit protein complex or metabolic pathway are produced simultaneously and in the correct proportions.
Within this transcription unit, multiple open reading frames (ORFs) are arranged sequentially in the 5′ → 3′ direction, and each ORF corresponds to a different protein.
Because of this arrangement, the resulting mRNA is called polycistronic mRNA, meaning it contains multiple coding regions (cistrons), each of which can be translated into a separate protein.
This is a major distinction from eukaryotic systems, where each mRNA typically encodes only one protein.
A classic example of this organization is the E. coli atp operon, which contains nine ORFs that collectively encode the different subunits of the F₁F₀ ATP synthase complex.
By organizing genes this way, prokaryotic cells ensure that all components of a multi-subunit protein complex or metabolic pathway are produced simultaneously and in the correct proportions
How is this coordinated regulation crucial for efficiency?
since it prevents the wasteful production of incomplete or unbalanced protein systems and allows the cell to quickly adapt its metabolism to changing conditions
Understanding ________ is key to grasping how gene expression is controlled at a molecular level, especially in systems like operons
cis and trans acting factors
wht do cis and trans acting factors describe
how regulatory elements influcence a gene based on their location and mobility
What are typically diffusible molecules- meaning that they are produces in one location in the cell but can move through the cytoplasm or nucleus to act on multiple target genes
trans-acting factors
What type of proteins are trans-acting factors that are most common?
DNA binding proteins
trans-acting factors are typically diffusible molecules, meaning they are produced in one location in the cell but can move (diffuse) through the cytoplasm or nucleus to act on multiple target genes. Most commonly, these are DNA-binding proteins such as…
transcripting factors that recognize specific DNA sequences and either activate or repress transcription
Because transacting factors are not physically attached to the protein they regulate, what can a single transacting protein do?
they can control many different genes across the genome, making them powerful regulators of gene expression
What are nondiffusible DNA sequences that must be located on the same moecule of DNA as the gene they regulate
cis-acting elements
In contrast, cis-acting elements are non-diffusible DNA sequences that must be located on the same molecule of DNA as the gene they regulate. These elements do not move or produce a product
what do they do instead?
they serve as binding sites for trans acting factors
What are common examples of cis-acting elements?
promoters
operators
enhancers
Why is the positon of cis-acting elements critical
if a cis acting element is moved to a different location ( for example, onto another DNA molecules) it will no longer properly regulate the original gene
What is the distinction between cis-acting elements and trans-acting elements
cis elements are physical DNA sequences tied to a specific gene, while trans factors are mobile molecules (usually proteins) that interact with those sequences.
the key distinction is that cis elements are physical DNA sequences tied to a specific gene, while trans factors are mobile molecules (usually proteins) that interact with those sequences.
What do they form together?
a coordinated regulatory system:
cis elements provide the “addresses” on the DNA,
trans-acting factors are the “messengers” that bind those addresses to control whether a gene is turned on or off.
What is a cis acting element typically?
a DNA sequence
what is the process where a segment of DNA is used as a template to synthesyze a complementary RNA molecule
transcription
what is transcripton seen as
the foundation for how genetic inforamtion in DNA is selectively used to produce RNA- the first step in gene regulation
What enzyme carries out transcription
RNA polymerase
Unlike DNA replication—which copies the entire genome every time a cell divides… what is different about transcription
transcription is highly selective, meaning only specific genes are transcribed at a given time depending on the cell’s needs
Why is the high selectivity of transcripton cruciakl
because different cell types and conditions require different sets of protiens
Transcription begins at a specific DNA sequence called…
the promoter
what does the promoter act as?
the binding site for RNA polymerase ans associated factors
What happens once RNA polymerease is bound to the promoter?
RNA polymerase starts synthesizing RNA at a precise location
Once bound, RNA polymerase starts synthesizing RNA at a precise location.
What is this precise location known as?
the transcription start site (TSS)
What is the transcription start site (TSS) always designated as?
+1
Once bound, RNA polymerase starts synthesizing RNA at a precise location known as the transcription start site (TSS), which is always designated as +1.
From this reference point, how are positions along the DNA described
relative to the TSS
Once bound, RNA polymerase starts synthesizing RNA at a precise location known as the transcription start site (TSS), which is always designated as +1. From this reference point, positions along the DNA are described relative to the TSS
How are sequences upstream described?
(toward the 5′ direction of the coding strand) are given negative numbers (e.g., −10, −35)
they are given negative numbers
Once bound, RNA polymerase starts synthesizing RNA at a precise location known as the transcription start site (TSS), which is always designated as +1. From this reference point, positions along the DNA are described relative to the TSS
How are sequences downstream described?
(toward the region being transcribed)
they are given positive numbers
As RNA polymerase moves along the DNA, it continues synthesizing RNA until it reaches a certain sequence
What is this certain sequence?
terminator sequence
What does the terminator sequence signal
the end of transcription
As RNA polymerase moves along the DNA, it continues synthesizing RNA until it reaches a terminator sequence, which signals the end of transcription
What is the RNA molecules produced at thi sstage called
the primary transcript
As RNA polymerase moves along the DNA, it continues synthesizing RNA until it reaches a terminator sequence, which signals the end of transcription. The RNA molecule produced at this stage is called the primary transcript
What does this allow in prokaryotes?
it may be ready for translation immediately in prokaryotes
As RNA polymerase moves along the DNA, it continues synthesizing RNA until it reaches a terminator sequence, which signals the end of transcription. The RNA molecule produced at this stage is called the primary transcript
What does this allow in eukaryotes?
it may require further processing
What other terms are used to describe how close or far certain DNA elements are releatice to the gene or TSS
proximal and distal
The RNA molecule produced at this stage is called the primary transcript, which may be ready for translation immediately in prokaryotes or require further processing in eukaryotes. Additionally, terms like proximal and distal are used to describe how close or far certain DNA elements are relative to the gene or TSS—
What does proximal mean?
neraby and often within a few dozen base pairs
The RNA molecule produced at this stage is called the primary transcript, which may be ready for translation immediately in prokaryotes or require further processing in eukaryotes. Additionally, terms like proximal and distal are used to describe how close or far certain DNA elements are relative to the gene or TSS—
What does distal mean
farther away often thousands of base pairs
The mechanism of RNA polymerase explains how cells physically build an RNA strand from a DNA template, and it highlights some important differences from DNA replication.
What is one key feature of RNA polyjmerase
RNA polymerase can initiate RNA synthesis de novo
what does de novo mean?
it does not require a primer to get started
The mechanism of RNA polymerase explains how cells physically build an RNA strand from a DNA template, and it highlights some important differences from DNA replication. One key feature is that RNA polymerase can initiate RNA synthesis de novo, meaning it does not require a primer to get started.
How is this different from DNA polymerase
it always needs a pre-exsisting strand to extend
The mechanism of RNA polymerase explains how cells physically build an RNA strand from a DNA template, and it highlights some important differences from DNA replication.
One key feature is that RNA polymerase can initiate RNA synthesis de novo, meaning it does not require a primer to get started.
This is unlike DNA polymerase, which always needs a pre-existing strand to extend.
What does RNA polymerase do?
it brings together the first two ribonucleoside triphosphates (NTPs)
What are ribonucleoside triphosphates (NTPs)
the building blocks of RNA (ATP, GTP,CTP, and UTP)
RNA polymerase brings together the first two ribonucleoside triphosphates (NTPs)—the building blocks of RNA (ATP, GTP, CTP, UTP)
What follows this?
they directly catalyzes the formation of the first phosphodiester bond
what does the formation of the first phosphodiester bond establish?
the beginning of the RNA chain