Microbial Physiology Final
1/74
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
75 Terms
Oxidative Phosphorylation
used in respiration (oxygen present), makes ATP, need PMF
3H+ per ATP
Substrate-Level Phosphorylation
Universal, anaerobic, produce ATP (PEP —> Pyruvate)
Photophosphorylation
phototrophs, rxn centers, uses PMF
chemotrophs
use chemical energy
phototrophs
use light energy
autotrophs
CO2 —> reduced Carbon
heterotrophs
reduced carbon —> CO2
ChemoLITHOtrophs
inorganic electron donors
ChemHETEROtrophs
organic electron donor
Allosteric Regulation
small protein impacts enzyme activity
post-translational
sRNA
determine if ribosomal binding site is open or not
post-transcriptional
feedback inhibition
end product stops earlier activity/ translation
post-translational
Covalent Modification
methylation or phosphorylation to change activity
post-translational
RNA Polymerase
aka sigma factors
in bacteria, catalyzes transcription
Activator
Transcription factors, bind upstream of promoter to increase transcription
Repressor
transcription factor, binds downstream promoter/ operator to decrease transcription by blocking RNAP
Gm+ bacteria
thick PG cell wall, no outer membrane
Gm- bacteria
thin cell wall, two membranes- outermembranes contains LPS, porins, and lipoproteins
Archaea
Pseudo-PG/ S-layer cell wall, no outer membrane
Sec-dependent transport system
dominant system, can transport proteins: to the periplasm, into the inner membrane, or the outer membrane
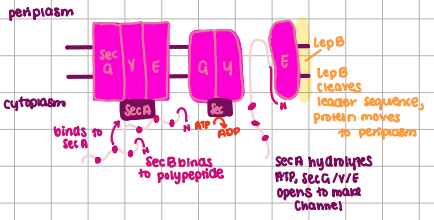
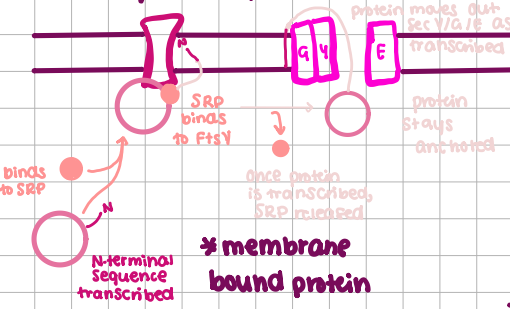
SRP-Dependent Transport
Co-Translational
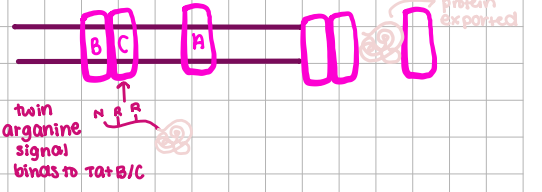
TAT-System
twin arganine signals, move folded proteins, sec-independent
Transmembrane Spanning Domain

Stop Transfer Sequence
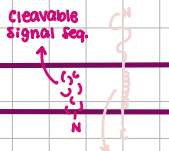
Non-Cleavable Signal Sequence
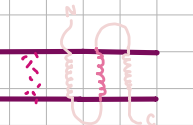
Type II Transport
major secretion route, SEC-dependent
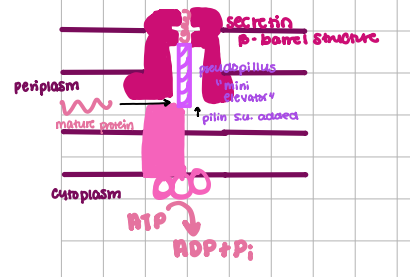
Type V Transport
“autotransportation”, SEC-Dependent
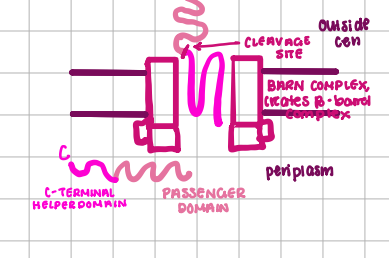
Type IV Transport
transport protein, virulence and exoenzymes, SEC-Dependent
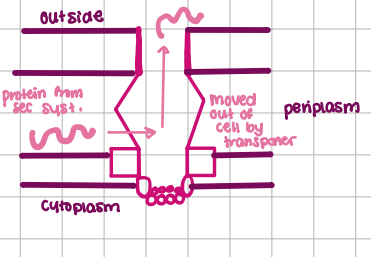
Type I transport
3 linked proteins, move protein outside cell, ATP hydrolysis, SEC-Independent
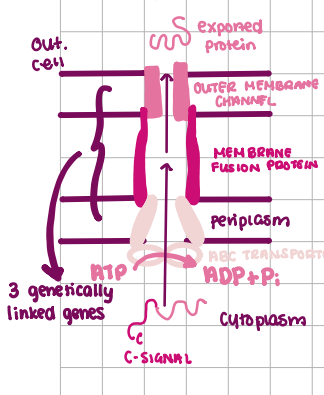
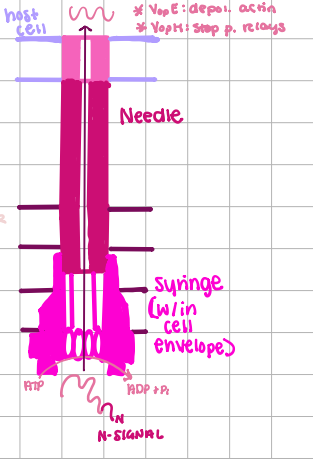
Type III Transport
many accessory proteins, transport directly to host, SEC-Independent
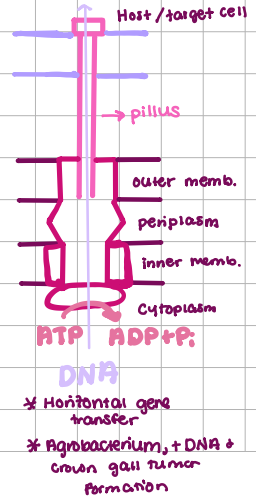
Type IV Transport (SEC-Independent)
DNA Transport/ Conjugation
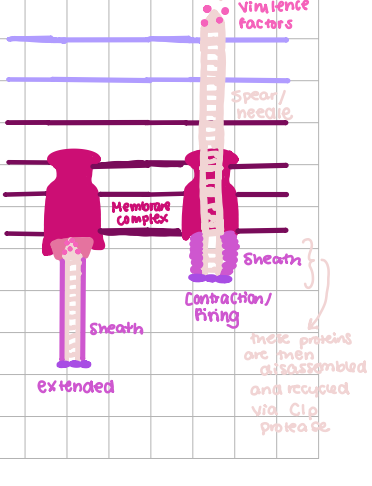
Type VI Transportation
phage-like, inject toxins into host, sheath contracts to push needle w/ virulence factors
Type IV pili
built from subunits of pilin at base, use ATP
Gliding Motility
slime extrusion, A-type motility (myxobacteria)
Twitching Motility
polyermize/ extend pillus, attach to surface of cell, retract and depolymerize
Flagella: Early Gene Expression
σ70 helps express FlnDC —> master regulator
Flagella: Middle Genes
Master regulator + σ70 —> components of basal body, FlgM (anti-σ factor) unbound from σ28 when hook is finished
Flagella: Late Genes
master regulator + σ28 work together to express flagellin, motor, and chemotaxis genes
Repellent Present
MCPs —> CheW —> CheA —> CheB demethylates MCPs and CheY-P causes CW tumble
Attractant Present
MCPs —> CheW —> CheA kinase —> CheY causes CCW movement, CheB unphosphorylated, CheR resets system
Smooth Run Mutant
CCW movement of C-ring, no CheY or Che A
Tumble Mutant
CW movement of C-ring, no CheB or CheZ
Luciferase
mixed function oxidase, needs ATP, participates in redox reactions and reduces coenzymes
Gm- Autoinducer
Acyl Homoserine Lactone (AHL)
Gm- Autoinducer Synthesis
LuxI, transcribed when LuxR forms a complex with AHL
LuxR
acts as signal sensor and transcription factor, bind to AHL then DNA, in cytoplasm.
Gm+ Quorum Sensing Signal
peptide (binds to sensor histidine kinase, creates HTH, response regulator phosphorylated)
Gm+ signal transport
ABC transport
Lactonase
cleaves lactone ring of AHL
Furanone
blocks the formation of biofilms
Anaerobic Response to Oxidative Stress
superoxide reductase (makes hyd. peroxide) → peroxidase
Aerobic Response to Oxidative Stress
superoxide dismustase (makes hyd. peroxide) —> catalase
Superoxide Present
SoxR (transcription factor) → SoxS (transcription factor) → SodA encodes sup. dismutase & nFO endcodes for endonuclease (DNA repair)
Glutathinone
tripeptide, senses RED/OX state of cytoplasm
forms a disulfide bond and bonds with a molecule when under stress conditions, kefB/C antiporter activated to acidify cytoplasm, detoxifies ROS
Hydrogen Peroxide Present
OxyR → gorA → Glutathianone reductase
σ32 + chaperone proteins
universal heat shock response
mRNA unfolds, activates σ factor involved, makes chaperones (refold proteins and regulate σ) and proteases (degrade what cannot be saved)
σ24E + RseA
extreme temperature/ stress response
heat denatures outer membrane protein, degrades inner membrane protein to release σ, transcribes cell envelope repair proteins
Spore Properties
SASP (aid in resistance), dehydrated, NO metabolic activity, restive layers (spore coat, etc.)
Spo0A
master regulator of spore formation, activated via phosphorelay
activates endospore formation, turns off gene inhibition
σ-factor regulation of spore formation
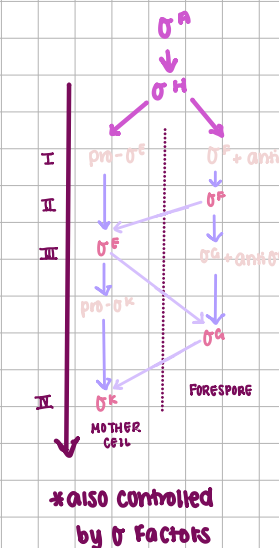
Mother Cell σ factors
E → K, must be modified before active
Forespore σ factors
F → G, must remove anti σ-factor
Germination
occurs when spores sense nutrients → water enters, cortex breaks, and vegetative growth resumes
EPS Matrix
found in biofilms
adherence, prevent dehydration, provide protection
eDNA
extracellular DNA
promotes HGT, help increase antibiotic resistance
NOSTOC cell differentiation
practices nitrogen fixation
no more RubisCO, no pigments, no PS II; glutamine and nitrogenase expressed, obtain sugar from neighbors
Streptomyces
germ tube grows into substrate then aerial hyphae forms

Myxococcus and Fruiting Body Formation
Swarm (group feeding, type A motility) → aggregate → Mound forms → Myxospores and Peripheral Rod Cells → Myxospores released → become vegetative cells again
FruA(-P)
induces early gene development, encourages aggregation and sporulation
Signaling Fruiting Body Formation
starvation → PPGTP → protease (exported) → A-Signal → 2 comp. reg → FruA-P
C-Signal
cell surface signal, cell density
binds to surface → Kinase → FruA-P
A-Motility (Myxococcus)
gliding motility (slime trails), single cells at edge of colony
S-Motility (Myxococcus)
social, group movement of cells, mediated by type IV pili
Swarmer Cells
C. Crescentus
no cell division, motile, high conc. of CtrA-P, colonizers
Stalked Cells
C. Cresentus
attach to surface w/ Holdfast, high cell division, low conc. of CtrA-P