Cell Biology Chapter 11: Protein Sorting and Transport 낱말 카드 | Quizlet
1/60
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
61 Terms
Endomembrane System
Network of communicating membrane with related functions in the cell
-Communicate via vesicular transport
-Function in protein processing and lipid synthesis
Where does protein synthesis take place?
On ribosomes in the cytoplasm (i.e. either free in the cytosol of on the RER)
Where does protein synthesis begin?
On ribosomes that are free in the cytosol
Are proteins ready to function once synthesized?
No; proteins need to fold and typically go through a variety of processing events as well
Events "After" Protein Synthesis
Folding
Protein cleavage
Glycosylation
Lipid attachment
Protein phosphorylation
Protein Targeting
Bold=Protein Modification
Protein Targeting
Involves:
Specific signal sequence
Receptors & recognition protein
Chaperones (in some cases)
Endoplasmic Reticulum (ER)
Network of membrane enclosed
Two sides:
Lumenal (lumen) vs. Cytosolic (cisternal space)
Smooth ER
No ribosomes
Lipid Metabolism
Tubules
Rough ER
Ribosomes
Protein synthesis and processing
Flat sacs
Protein Sorting
Some proteins can synthesis free in the cytosol or on membrane bound ribosomes and release into either in the cytosol or ER lumen where they are later taken to an organelles to fulfill its function.
Secretory Pathway
RER > Golgi > Secretory Vesicle > Cell Exterior
Pulse Chase Experiment
Studies the pathway of secreted proteins by labeling new synthesized proteins with radioactive amino acids
1. Pancreatic cells were exposed to AA, then were found in the RER
2. Incubated with nonradioactive AA the cells were found in the Golgi Apparatus
3. After longer chase period the cells went from the Golgi Apparatus to secretory vesicles.
4. Secretory vesicles fuse with plasma membrane to be released outside the cell
Protein Targeting
How protein are targeted to ER
Signal Sequence
At AA N-terminus of the polypeptide chain (polymer of AA)
Short stretch of hydrophobic AA that are cleaved from the polypeptide chain during its transfer into the ER lumen
Span about 20 AA
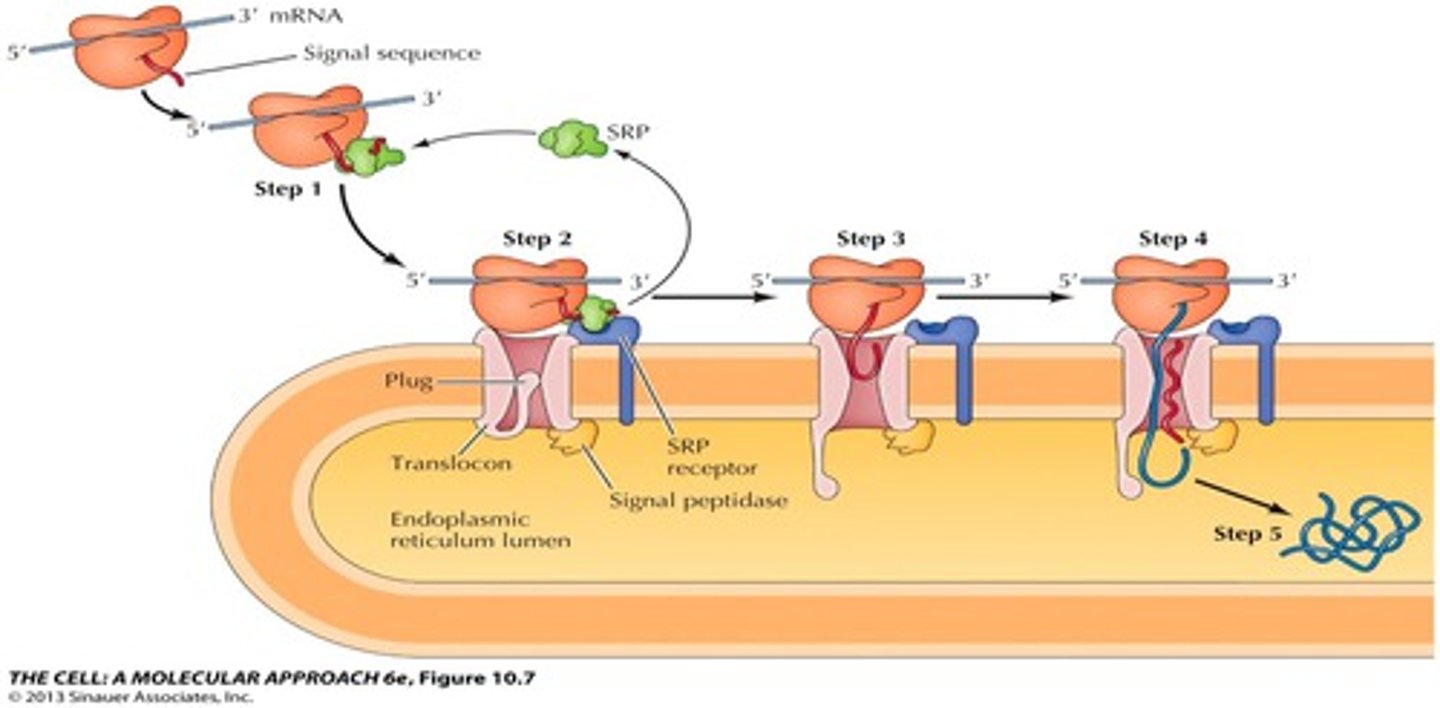
Cotranslational Targeting
1. As the signal sequence emerges from the ribosome, it is recognized and bound by the signal recognition particle (SRP)
2. The SRP escorts the complex to the ER membrane where it binds to the SRP receptor
3. The SRP, is released, the ribosome binds to the translocon and insertion of the signal sequence opens the translocon.
4.Translation resumes and the signal sequence is cleaved by signal peptidase
5. Continued translation drive the translocation of the growing polypeptide chain across the membrane.
6. The complete polypeptide chain is released within the ER lumen.
Signal Recognition Particle (SRP)
Composed of:
6 polypeptide
srpRNA (a 7s RNA)
Post-Translational Targeting
NO SRP
1. Synthesized on free
ribsomes
2. Cytosolic chaperones maintain unfolded conformation
3. Recognized by the Sec 62/63 complex and SS in inserted into the translocon in the ER membrane.
4. The Sec 63 protein is associated with a chaperone protein called BiP.
5. The binding of multiple BiP molecules and the their release gives off ATP hydrolysis which drives the translocation of the polypeptide chain into the ER lumen.
Stop Transfer Sequence
Terminal signal sequence cleaved
Internal Signal Sequence
Signal sequence is not cleaved
Processing and other events in the ER
Protein folding
-chaperones: catalyze protein folding (e.g. BiP, hsp's, calnexin, calreticulin)
-Assembly of multisubunit proteins
-Disulfide bond formation
protein disulfide isomerase
-Initial stages of glycosylation
-Glycolipid anchors
Chaperones
Essential in many processes in cell
Increase the rate of processes related to protein structure, conformation, folding
Lipid Synthesis
Happens in the SER
Membrane lipids such as phospholipids are synthesized by using:
-on pre-existing membranes
-using water-soluble cytosolic precursors (hydrophobic)
Lipids get to other membranes in the cell by: Transporting in the vesicles or by carrier proteins
Phospholipid Synthesis
1. Glycerol-3-Phosphate is synthesized on the cytosolic side of ER membrane from water soluble precursors.
2. Fatty acids are first transferred from coenzyme A carrier to glycerol-3-phosphate by membrane-bound enzymes, and the result of phosphatidic in inserted into the membrane.
3. A phosphate then converts phosphatidic acid to diacylglycerol
4. Catalyzation of a additional different polar heads occur via changing of glycerol by enzymes.
Bilayer
Since the membrane is a bilayer more phospholipids are produced on one side than the other
Flippases
By catalyzing the rapid translocation of phospholipids across the ER membrane, the flippases ensure even growth of both halves of the bilayer
Endomembrane System: Regulation of Transport
Nature of signals:
Peptide sequence
-short stretch of AAs in linear sequence (e.g. KDEL)
Signal patch or Conformation
-series of AA's brought together by 3D strc vs. linear sequence
Carbohydrate
Types of signals:
Export
Retention/Retrieval
Default
ER Export Signal
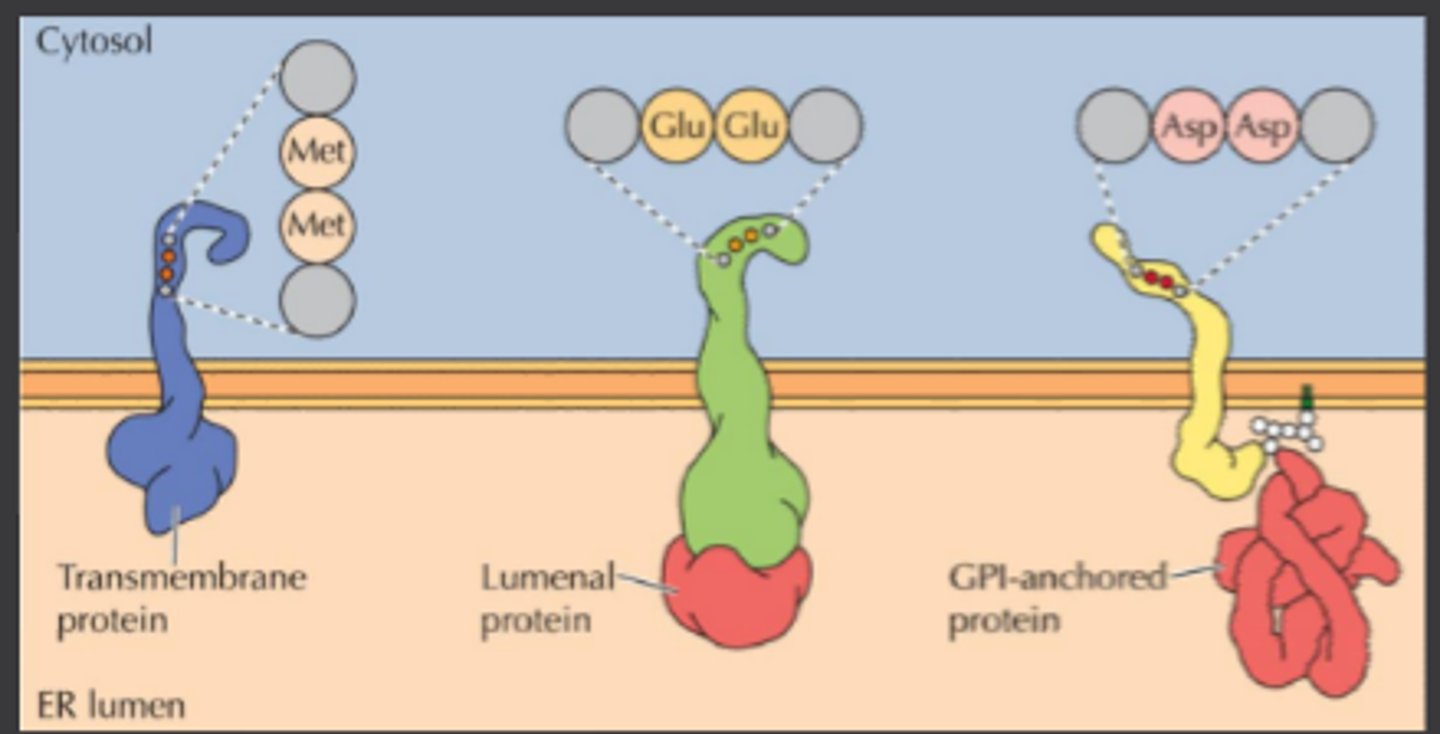
Transmembrane proteins:
-Di-acidic or di-hydrophobic signals bind to cytosolic adaptor proteins
-Also act as receptors for lumenal and GPI-anchored proteins
Lumenal proteins:
-Signal patch, conformation or default pathway
GPI anchors:
-Recognized by the GPI anchors
Di-acidic = Asp-Asp or Glu-Glu
Di-hydrophobic = Met-Met
Asp = aspartic acid
Glu = glutamic acid
Met = methionine
GPI=glycosylphosphatidylinositol
ER Retention/Retrieval Signals
-Also called retrograde transport
-Targets to specific location through specific recycling pathway
-Signal recognized by recycling receptor
-Some proteins retrieved by binding to proteins w/
such signals
KDEL
Soluble in ER
K=lysine D=aspartic acid E=glutamic acid L=leucine
KKXX
Transmembrane ER
Default Signals
Lack of signal?
Bulk flow through secretory pathway?
Unsure if may be an active signal
Secreted or cell surface
Polarized Cells
Plasma membrane has two domains:
Apical
Basolateral
Signals direct to membrane proteins to one membrane surface or other
Variety of signals involved
Golgi
Golgi Apparatus
Golgi Complex
What is the Golgi function?
Further processing of protein and sorting them to their final destination
Targets to: intracellular destinations
Cell Surface
-via recycling endosome
-Constitutive pathway
-Regulated secretory pathway (stored in secretory vesicles)
What is more of the Golgi functions?
Synthesis of lipids
-Glycolipids & sphingomyelin
-Ceramide precursor synthesized in ER
Plants: complex cell wall polysaccharide
-Cellulos, straight chain, synthesis at cell surface by plasma membrane enzyme
-Hemicelluloses & pectins, branched chain, synthesized in Golgi
Structure of the Golgi
Series of separate flattened membrane-enclosed sacs (cisternae)
Distinct polarity in both structure and function
Structural Polarity:
Cis Face
-entry of molecules from ER, usually side closer to nucleus
Trans Face
- Exit golgi on route to final destination, usually side closer to cell surface
Polarity Function in Golgi
Proteins move within compartments that mature:
-Cisternal migration/progression
-Golgi components return to previous sections by retrograde flow within vesicles
Compartments:
-ER-golgi IC
-Cis golgi network
-Trans golgi network
Orientation of Membrane Surface
Topologically lumen is equivalent to cell exterior
What is glycosylation?
Modification of the carbohydrate portion of glycolipids
What is protein glycosylation?
It is the addition of a carbohydrate to protein by a covalently attached of an amino acid side chain
What are the two types of protein glycosylation?
N-linked and O-linked
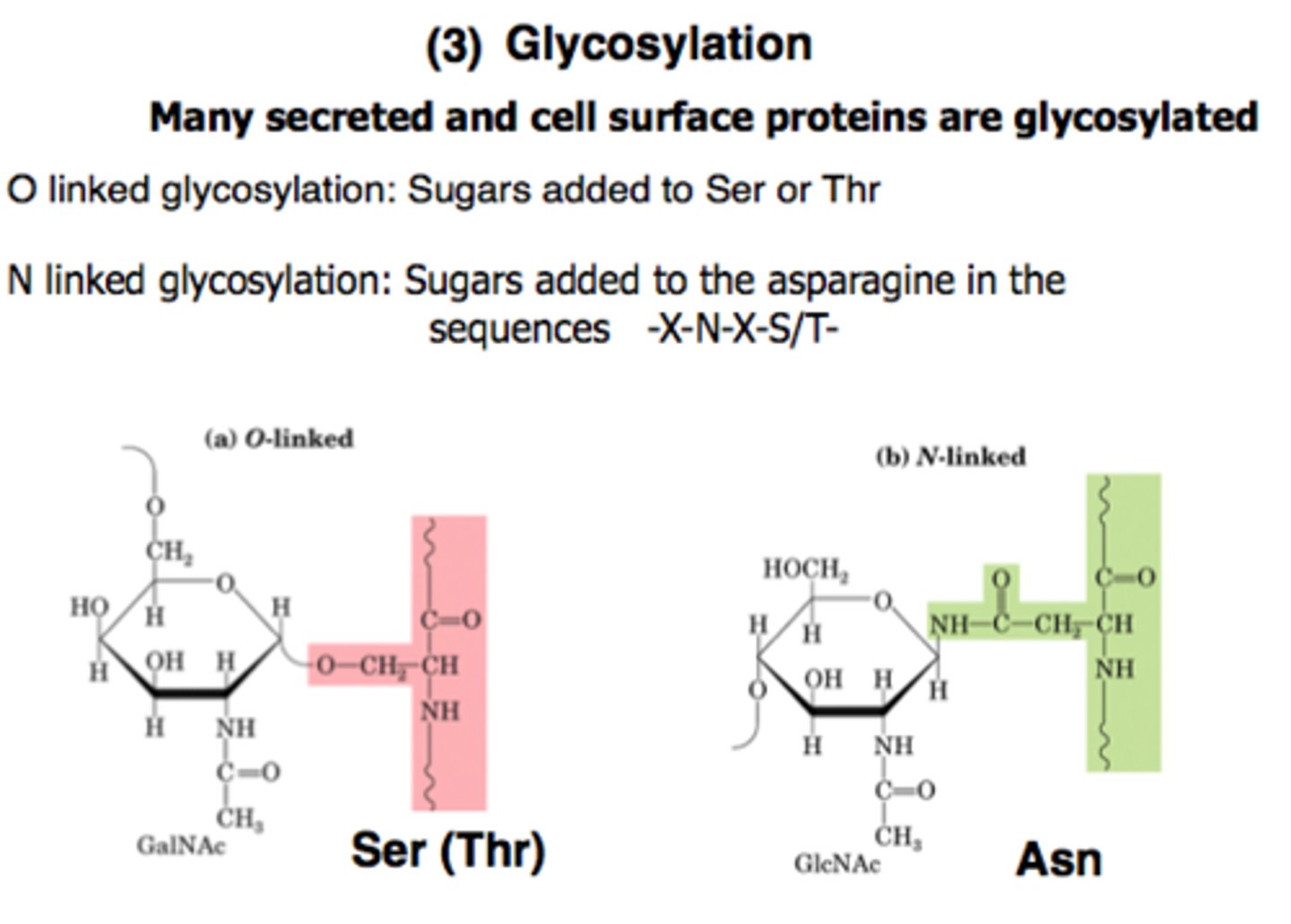
N-linked
-CH2On attached to N of asparagine
-Begun in the ER further processed in Golgi
O-linked
-CH2On attached to O of Serine or Threonine
-Occur in Golgi
N-linked vs O-linked
-CH2On attachment point
-Location where addition begins
-Mechanism: addition and modification of performed structure vs sequential addition
What determines glycosylation pattern of a protein?
Structure of protein
-Appropriate AA and signal
-Accessibility of region
Amount processing enzymes present with in Golgi of different types of cells
Carbohydrates modification is important in targeting of proteins to lysosomes because the chain carries information
N-linked glycosylation (1)
Occurs in golgi
The N-linked oligosaccharides of glycoproteins transported from the ER and are further modified by an ordered sequence of reaction catalyzed by enzymes in different compartments of the Golgi.
N-linked glycosylation (2)
For lysosomal targeting occurs in the cis golgi
Proteins destined for incorporation into lysosomes are specifically recognized and modified by the addition of phosphate groups to the number 6 position of mannose residues. In the first step of the reaction, N-acetylglucosmine phosphates are transferred to mannose residues from UDP-N-acetylglucosamine group is then removed, leaving mannose-6-phosphate.
Vesicular Transport
Targeting
Establishes and maintains the functional organization of the cell
Selectivity of transport
-is the key to appropriate targeting
-determined by combination of vesicle coat proteins, GTP binding proteins and other associated proteins
Vesical coat proteins
are involved in specific trafficking routes
What are coat proteins and GTP-binding proteins?
Vesicle Coat Proteins:
Clathrin
COP I
COP II
Small GTP-binding proteins:
ARF 1-3
Snar 1
Rab protein family
Formation of Transport Vesicle
Activities:
-sorting of cargo
-budding of vesicle
Players:
-Coat proteins=Clathrin
-Adaptor proteins=GGA, AP1
-GTP-binding protein=ARF1
Clathrin
-Structural Role
-Basket-like lattice
-Distorts membrane
Adaptor Proteins
-Medites clathrin binding
-Selects vesicle contents
Transport of lysosomal proteins by clathrin-coated vesicles
-Lysosomal protein bound on lumenal face to R
-R=transmembrane proteins
-Adaptors (GGA & AP1) binds cytosolic portion of R
-Designates clathrin... lysosomal target
GTP-binding protein
ARF1 initiates vesicle budding
-ARF is activated by ARF-GEF: ARF(GDP) > ARF(GTP)
-ARF (GTP) recuits adaptor proteins: GGA
-GGA recruits receptor which carries the lysosomal protein (a lysosomal hydrolase)
-GGA also recruits AP1 which is a binding site for clathrin
Movement of Transport Vesicle
Activities
-Recycle ARF, coat proteins and adapters
-Travel along cytoskeleton
Players
-GTP hydrolysis on ARF weakens coat
-Hsp 70s remove some of coat proteins
-Remaining coat proteins bind tubular and molecular motors
Membrane Fusion
involves protein-protein interactions, the assembly of protein complexes, and GTP hydrolysis
Lysosomes
-Membrane enclosed organelle
-Vary size & shape
-Enzyme digest all biological polymers
Functions of Lysosome
1. Digest material taken up by endocytosis
2. Phagocytosis
-Cleaning up debris via macrophages
3. Autophagy
-Programmed cell death
-The turnover of the cell's own components
Organization of Lysosome
-Acid hydrolyses (ph5)
-Expend energy maintain internal pH.
The acidic internal pH of lysosomes results from the action of a proton pump in the lysosomal membrane, which imports protons from cytosol coupled to ATP hydrolysis.