3.2 PHOTOSYNTHESIS USES LIGHT ENERGY
1/43
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
44 Terms
Photosynthesis equation
6CO2+ 6H2O→ C6H12O6 +6O2
Distribution of chloroplasts (light trapping)
Main → Palisade mesophyll
Location → Below upper epidermis→ High light intensity
Maximises light absorption
Spongy mesophyll fewer
Chloroplasts movement why ?
Move and rotate within palisade cell depending on light intensity
Low→ Surface maximum absorption of light intensity
High→ Vertical against cell wall prevent over exposure pigment bleaching
Chloroplast adaptions
Large surface area - max absorption of light
Pigments in thylakoids single layer membrane surface - Maximise light absorption
Grana - large surface area for light dependent
Starch grains - store energy
Leaf structure (top→bottom)
Cuticle
Upper epidermis
Palisade mesophyll
Spongy mesophyll
Lower epidermis
Stoma
Leaf adaptions (light trapping)
Orientate perpendicular to light
Large surface area→ capture more light
Thin leaf→ light penetrate short co2 diffusion distance
Transparent upper epidermis - light penetrate
Palisade near top- lots chloroplasts
Air space spongy mesophyll - rapid gas diffusion
Stomata → co2 entry o2 exit
Chloroplasts as tranducer
Change energy from one form to another
Light photon energy → chemical energy (ATP)
Light absorbed by pigments (in thylakoid membranes ) drive photosynthesis
Photosynthetic pigments
Absorb light energy at particular wavelengths of light
Different pigments absorb different wavelengths
Chlorophyll a, b carotene xanthophylls
Where are pigments found
Thylakoid membranes of chloroplasts
Advantage of having lots of pigments
Greater rage of wavelengths absorbed
More photons absorbed more products of light dependent made
Greater rate of photosynthesis
Chromatography of leaf pigments
Separate pigments to identify
Principle: Pigments move at different rates depending on solubility in solvent (higher Rf more soluble)
Rf Value
Distance moved by pigment / Distance moved by solvent
Compare calculated Rf with known to identify
Always <1
Light harvesting
pigments absorb light - antenna complex
Accessory (broad spectrum)
Chlorophyll a reaction centre - energy transferred maximise light capture efficiency
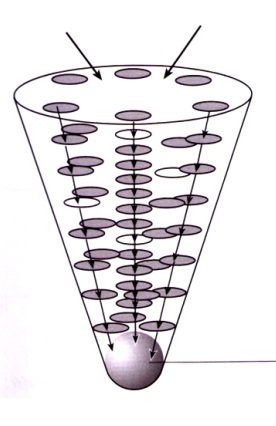
Reaction centre
2 Chlorophyll a pigments (primary pigments)
Excited chlorophyll emits high energy electrons → start light dependent
Photosystems
Photosystem 1 absorption peak 700nm (far red)
Photosystem 2 absorption peak 680nm (red)
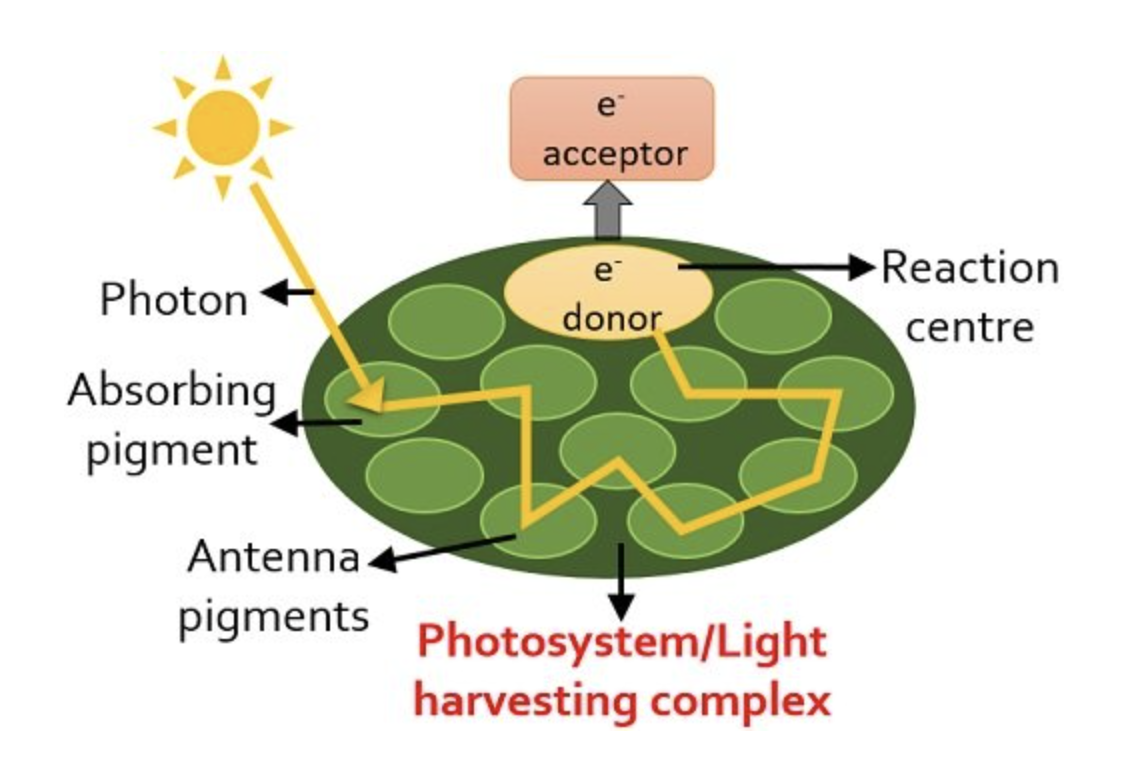
Thomas Englemann - spirogyra
Determine wavelengths most used
Algae spirogyra suspension motile aerobic bacteria
Prism refract white light → rainbow`
Wavelengths making most oxygen corespond
Bacteria migrate towards region with highest oxygen concentration (blue and red) most photosynthetic activity
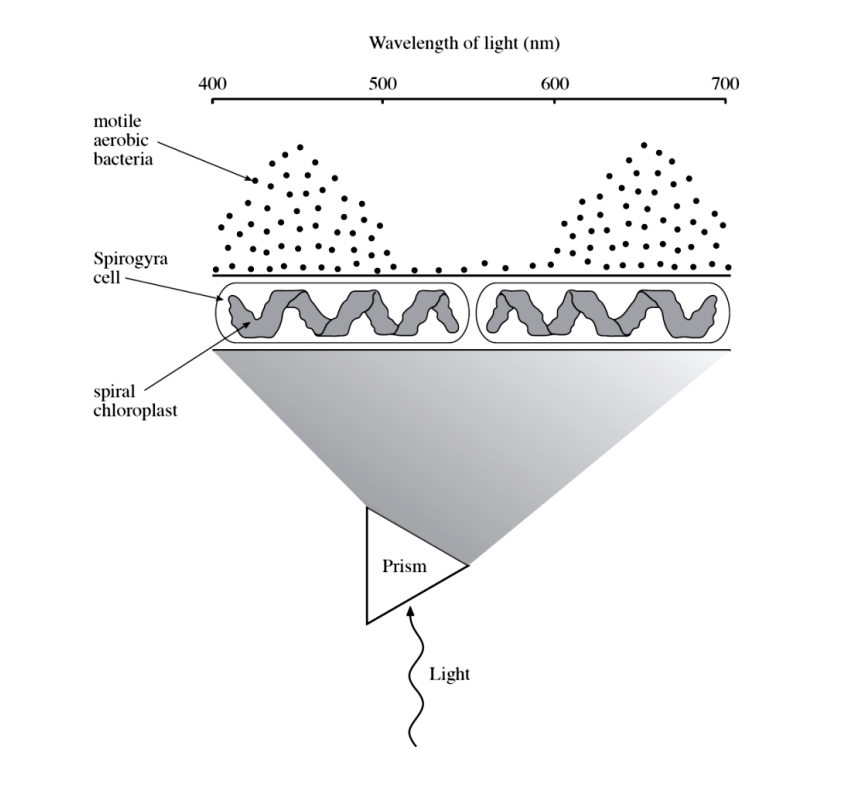
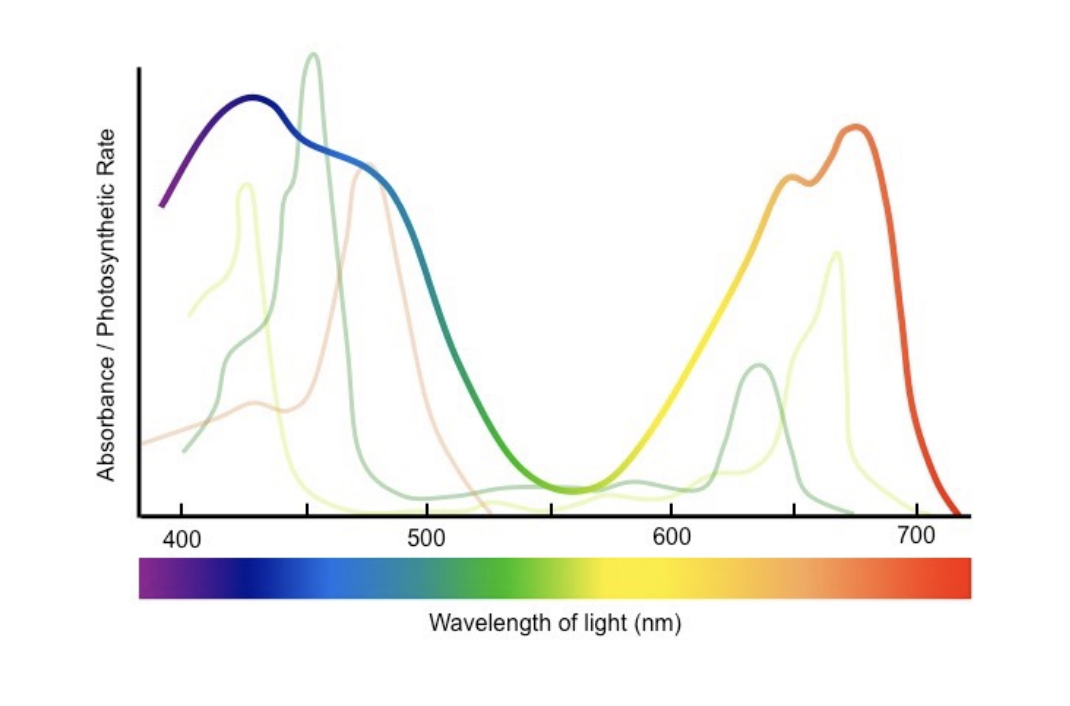
Absorption spectrum
Amount of light absorbed (by pigment) against different wavelengths
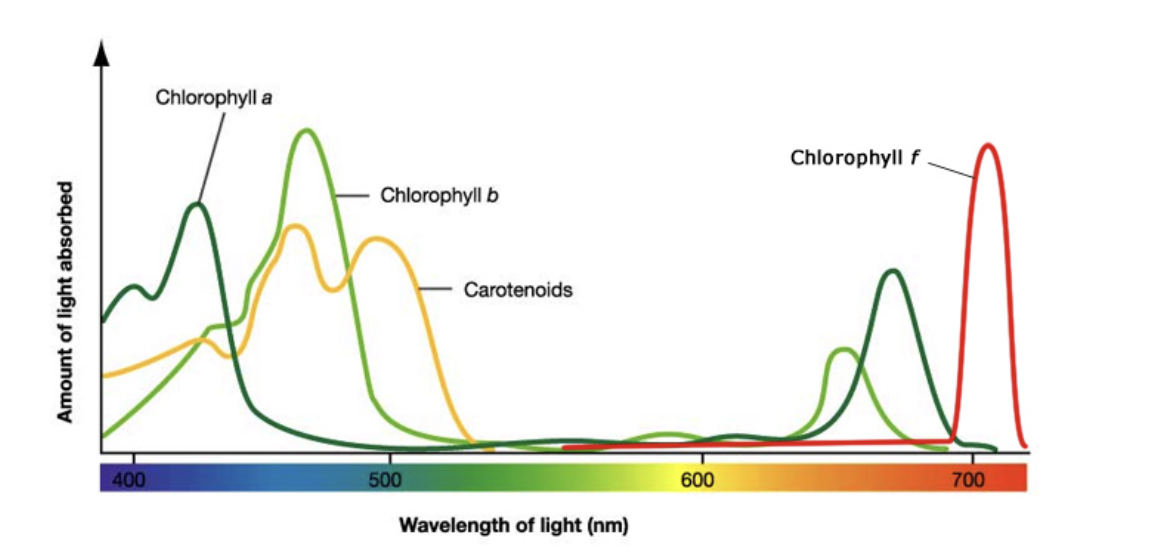
Action spectrum
Rate of photosynthesis at different wavelengths of light
Absorption and action close correlation
Suggest wavelengths absorbed by pigments used for photosynthesis
What prevents light escaping when absorbed
Proteins in antenna complex prevent light energy escaping
Light dependent photophosphorylation - Thylakoid membrane
Light absorbed Photosystem II (PSII) Electrons excited (gain energy) leave
Pass ETC Energy released → ATP (photophosphorylation)
Electrons lost from PSII are replaced by photolysis of water
Water → O₂ + H⁺ + e⁻
Photophosphorylation
Addition of phosphate + ADP → ATP by light energy
Cyclic photophosphorylation
PSI absorbs light → electrons excited (chlorophyll a)
Electrons → electron acceptor → ETC
Energy released → proton gradient → ATP (chemiosmosis)
Electrons return to PSI (lower energy)
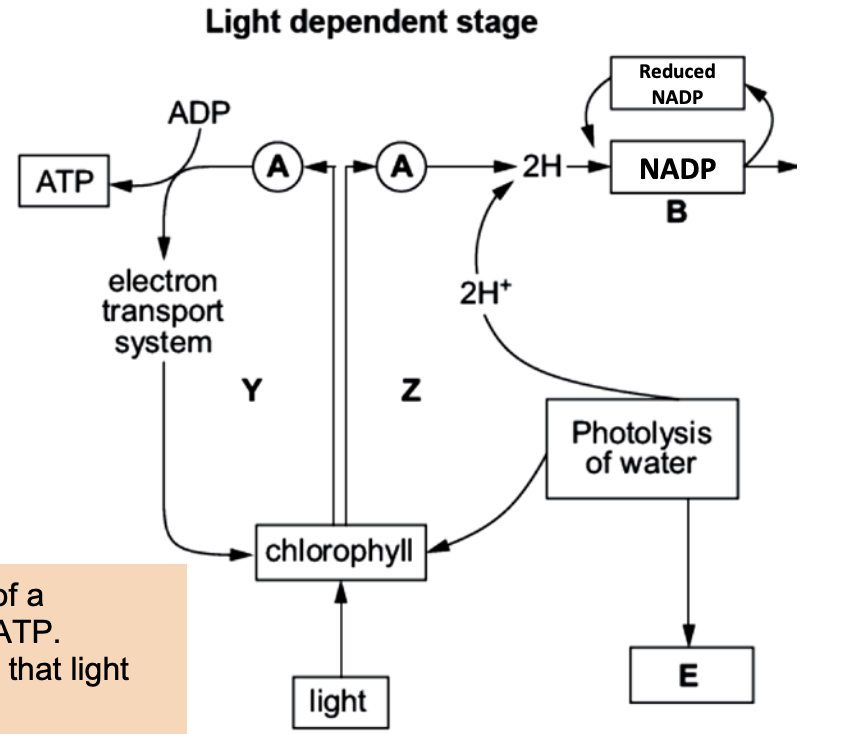
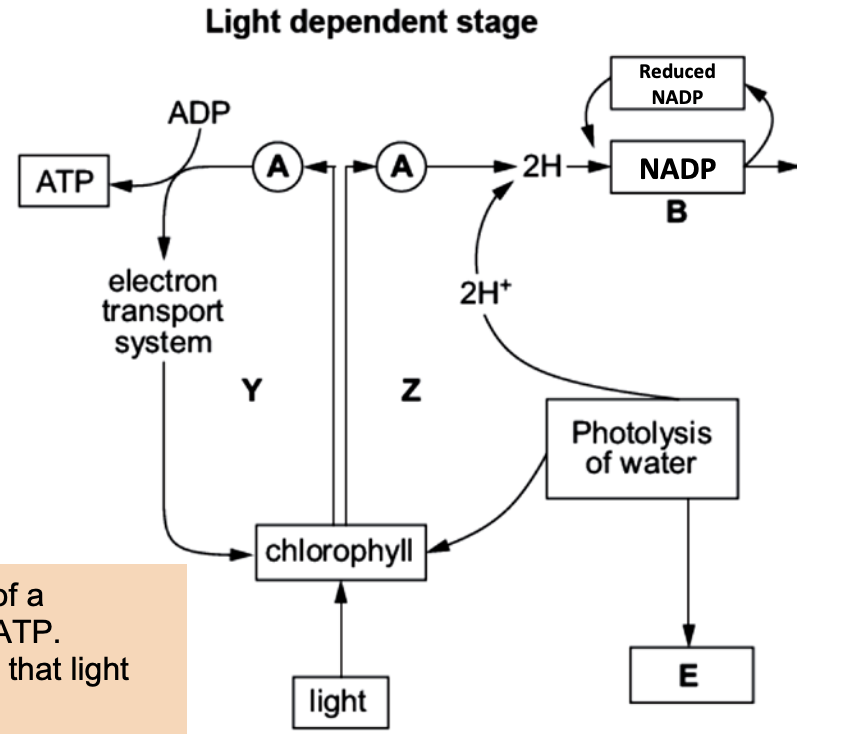
Non cyclic z scheme
Light absorbed by PSII → electrons excited (chlorophyll a)
Electrons → electron acceptors → ETC
Energy → proton pumping (stroma → thylakoid) → ATP
Photolysis of water replaces lost electrons
H₂O → 2H⁺ + 2e⁻ + ½O₂ (O₂ released)
Light absorbed by PSI → electrons re-excited
Electrons → electron acceptor → NADP
NADP + 2e⁻ + H⁺ → NADPH
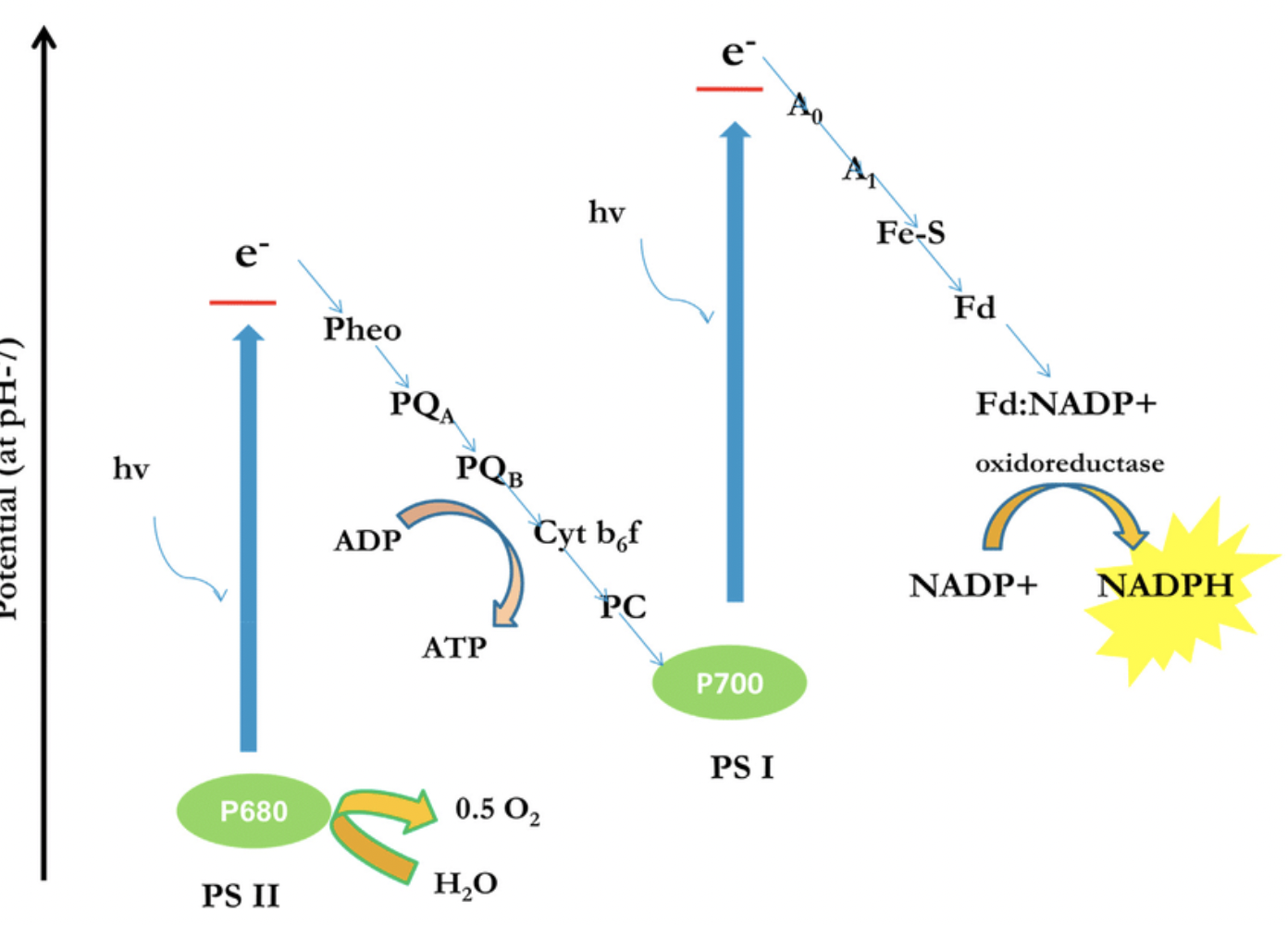
Photolysis of water
Water in thylakoid space split
H20 → 2H+ 2e- ½O2
electron removed replace lost by the chlorophyll photosystem 2
light is responsible only indirectly for splitting water.
Light independent - Stroma
Consume Co2 energy from ATP and reduced NADP and organic chemicals, (carbohydrates are produced)
Calvin cycle- light independent stage
Uptake CO2 by 5c Ribulose bisphosphate - Rubisco → unstable 6c
Splitting occur 2× 3c glycerate-3 phosphate
ATP and reduced NADP (from light dependent used) reducing
2x Triose phosphate 3c (glucose→ starch)
Regeneration ribulose phosphate 5c → ribulose bisphosphate (require ATP)
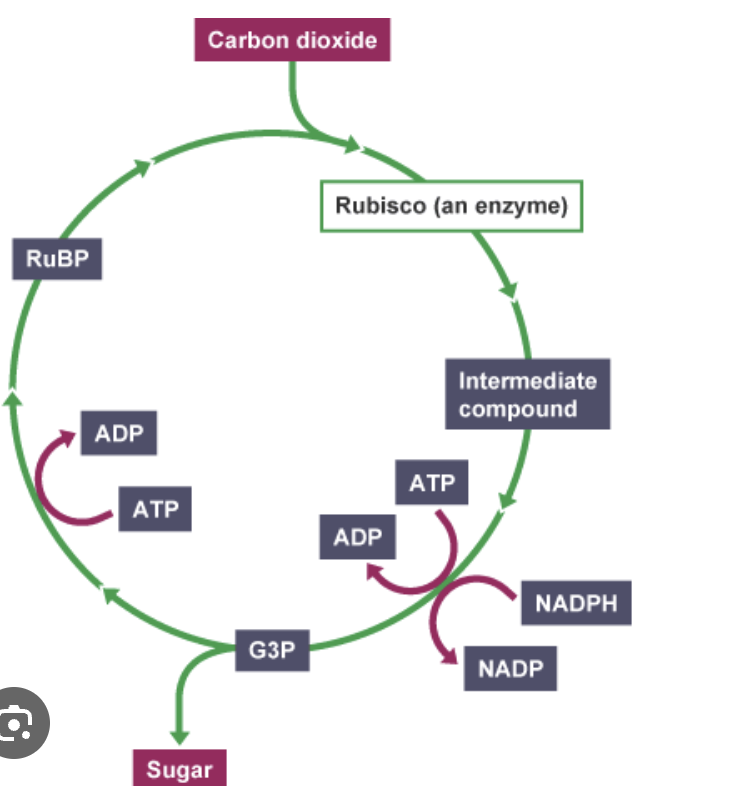
What is needed for the calvin cycle?
2 ATP ( glycerate3phosphate → triose phosphate ) (ribulose phosphate → ribulose bisphosphate)
Reduced NADP ( glycerate 3 phosphate → triose phosphate)
What happens to ADP and NADP after the calvin cycle?
Return back to the light dependent stage (thylakoid membrane ) to reform ATP and reduced NADP
Product synthesis carb lipids proteins (calvin cycle products)
Carbohydrates
Glucose (fructose bisphosphate) → starch (alpha) cellulose cell walls
Lipids
Acetyl coenzyme A (from Glycerate 3phosphate) → fatty acids → triglycerides
Proteins
GP → amino acids Nitrogen derived from NH4+ and NO3- absorbed by root hair cells active transport
Calvin 14Co2 experiment
Chlorella exposed 14CO2
Samples at time intervals
Hot ethanol stop enzymatic reactions
Radioactive compounds separated by chromatography
Dark large spot - more present GP TP carbon fixation
Interpreting autoradiographs
Describe -Spot size and darkness
Compare
5s: Mainly GP (early product) small TP
30s More GP and TP amino acids and sucrose
Explain: Gp before TP (Amino acids require GP) Sugars
4.Starch lipids proteins
Limiting factor definition
Factor limit rate of physical process by being short supplly
Carbon dioxide as limiting factor
Carbon source for calvin cycle
Low Co2 → RuBP cant fix Co2 → Less GP
TP and glucose slow
Light intensity as limiting factor
Darkness light independent reactions can’t occur no oxygen evolved
If light intensity higher than optimum, rate of photosynthesis decrease pigments damaged, wont absorb light efficiently
Low light → Less ATP and reduced NADP → Slow calvin cycle
Need to excite electrons
Light compensation point
Light intensity at which plant has no net gas exchange volume produced (respiration) released (photosynthesis) equal
Sun and shade plants
Shade plants
Low LCP → photosynthesis efficiently under low light
Sun plants
Higher LCP → need more light
Temp
Optimum around 25 after enzyme denature rate of photosynthesis decreases
Effect enzyme activity
Rubisco and ATP synthase low temp too slow
Minerals
Inorganic ions
Macronutrients needed in substantial quantities (magnesium copper)
Micronutrients needed in tiny amounts (manganese copper)
Nitrogen
Absorbed root hair cells active transport as NO3-
In xylem as amino acids
Nucleic acids, amino acids, nucleotides
Deficiency
Reduced growth of all organs
Chlorosis yellowing
Magnesium
Absorbed ad Mg2+ → xylem
Need for chlorophyll (avoid chlorosis)
How many times does the calvin cycle need to occur to form one molecule of glucose?
6 times
1/6 carbon leave for carbohydrate
If inhibitor blocks electrons enter ETC after ps2 what happens ?
Stop electrons ps2→ ps1
Block nadp reduction
Cyclic only ps1 can occur as ps1 → ps1 carrier involved not affected
Calvin cycle in dark and light
Initally Co2 and RuBO continues
GP cant be converted to TP
No ATP and reduced nadp only made in light dependent
Less TP for regeneration and glucose
Rate of co2 and RuBp decreas
Sweetness when photosynthesis rate higher than respiration
Higher rate of photosynthesis
More sugar is produced by photosynthesis than used in respiration
More sugar stored sweeter