Genetics lab exam final
1/57
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
58 Terms
population genetics
study of genetic variation within a population.
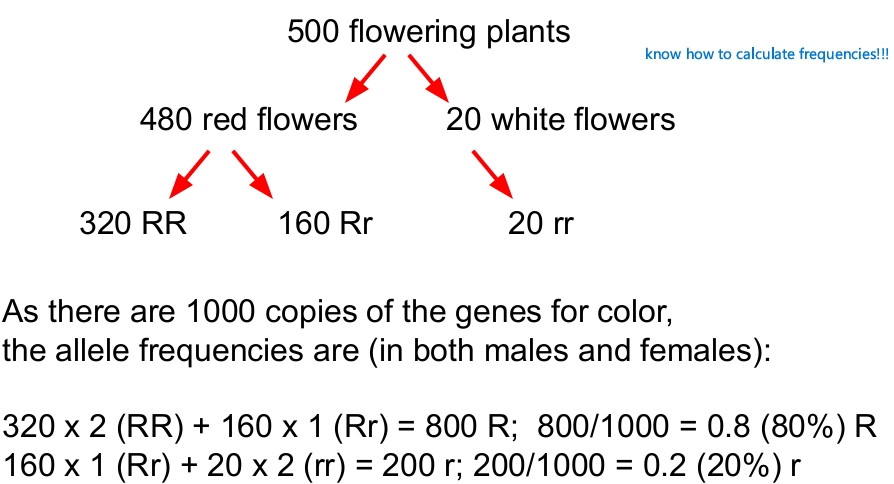
Allele frequencies define gene pools
how to calculate per allele

what is a test cross?
crossing an unknown genotype with a homozygous recessive to determine if the genotype is homozygous dominant or heterozygous
rr x Rr or rr x RR
the ratio for a test cross is always 1:1
what is the difference between genotype, phenotype, and allele?
Genotype: The genetic makeup of an organism, consisting of the specific alleles found on each chromosome.
Phenotype: The observable traits or characteristics of an organism, determined by the genotype.
Allele: A particular form of a gene, which can be either dominant or recessive.
Population
a localized group of individuals of the same species.
Species
a group of populations whose individuals have the ability to breed and produce fertile offspring
gene pool
total of all genes in the population at any one time
what is The Hardy-Weinberg Theorem?
non-evolving population, not natural. deviation from this theorem indicates evolution and how it occurs.
The gene pool of a non-evolving population remains constant over multiple generations; the allele frequency does not change over generations of time.
what are assumptions of the hardy-Weinberg theorem
large population size- small populations can have chance fluctuations in allele frequencies
No migration- immigrants can change the frequency of an allele by bringing in new alleles to a population
No net mutations- if alleles change from one to another, this will change the frequency of those alleles
Random mating- if certain traits are more desirable, then individuals with those traits will be selected and this will not allow for random mixing of alleles
No natural selection- some individuals survive and reproduce at a higher rate than others, then their offspring will carry those genes and the frequency will change for the next generation.
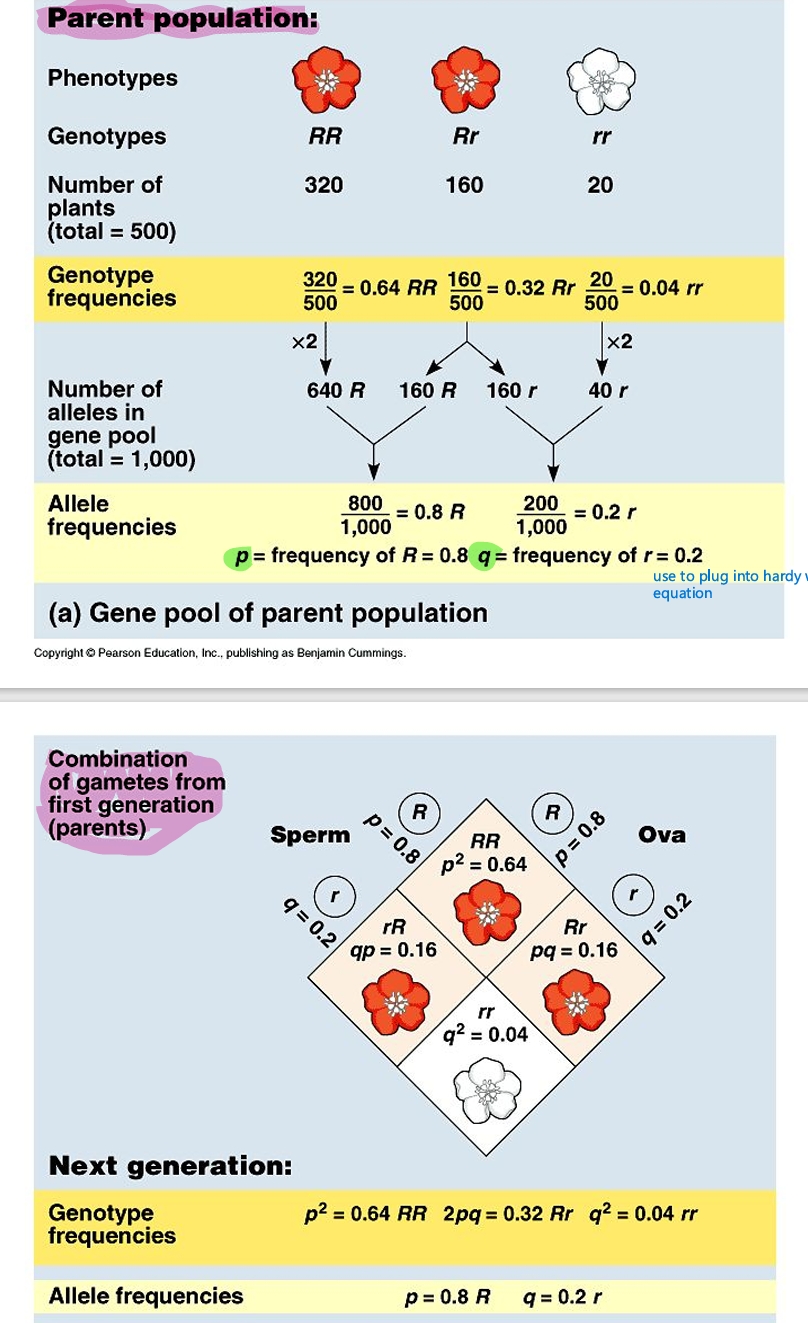
Hardy-Weinberg equation
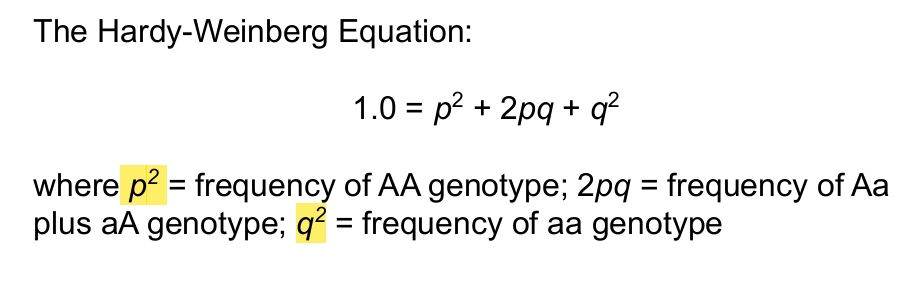

Hardy Weinberg equation example how to solve
microevolution
Evolution within a species/population
what are the 4 causes of microevolution? (remember 1&2)
genetic drift
natural selection
gene flow
mutation
what is genetic drift?
alteration of a small populations gene pool due to chance.
can be caused by:
bottle neck effect
founder effect
Bottleneck effect
a disturbance that removed a large part of the population that carried the original allele frequency resulting in the surviving population not having it.
Founder effect
may lead to reduced variability when a few individuals from a large population colonize an isolated habitat
what is Natural selection
differential success in reproduction based on heritable traits results in selected alleles being passed to relatively more offspring (Darwinian inheritance). The only agent that results in adaptation to environment
defined by
Darwinian fitness
relative fitness
what is gene flow
-is genetic exchange due to the migration of fertile individuals or gametes between populations.
what are mutations
change in an organism’s DNA and is represented by changing alleles. Mutations can be transmitted in gametes to offspring, and immediately affect the composition of the gene pool
Polymorphism
is the existence of two or more forms of a character, in high frequencies, within a population. Applies only to discrete characters
Geographic variations
differences between gene pools affected by environmental factors. could be affected by natural selection
Diploidy
hides genetic variation from selection in the form of recessive alleles..
Balanced polymorphism
the ability of natural selection to maintain stable frequencies of at least two phenotypes.
Heterozygote advantage
example of a balanced polymorphism, where the heterozygote has greater survival and reproductive success than either homozygote
Frequency-dependent selection
survival of one phenotype declines if that form becomes too common.
Neutral variation
genetic variation that results in no competitive advantage to any individual
sickle cell anemia vs sickle cell trait
sickle cell anemia: severe blood disorder caused by inheriting two sickle cell genes
a recessive disease where misshapen sticky C-shaped stiff blood cells that are short lived. (SS)
sickle cell trait: an individual carries only one sickle cell gene and the normal dominant beta-globin gene can take over so the individual has normal blood cell production. (AS)
normal dominant gene: HbA
sickle cell recessive gene: HbS
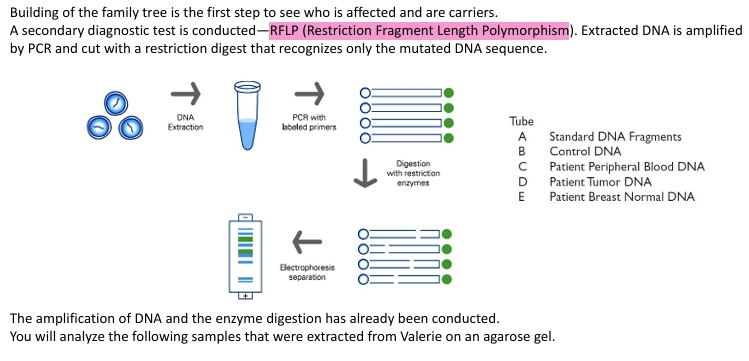
how is an individual tested for sickl cell?
restriction length polymorphism- through karyotyping from amniocentesis through PCR and restriction digest of amplified gene. The palindrome recognition site of the restriction site of the restriction enzyme Mstll. Mstll can be used to recognize normal bet-globin (CCTGAGG) but cannot recognize the mutated form. these are cut into fragments and then analyzed using gel electrophoresis to determine the presence of sickle trait. a person with sickle cell has their Mstll restriction site altered.
what is the restriction enzyme used in sickle cell testing restriction length polymorphism (RFLP)?
Mstll is CCTNAGGG where N=G

sickle cell trait practice problem
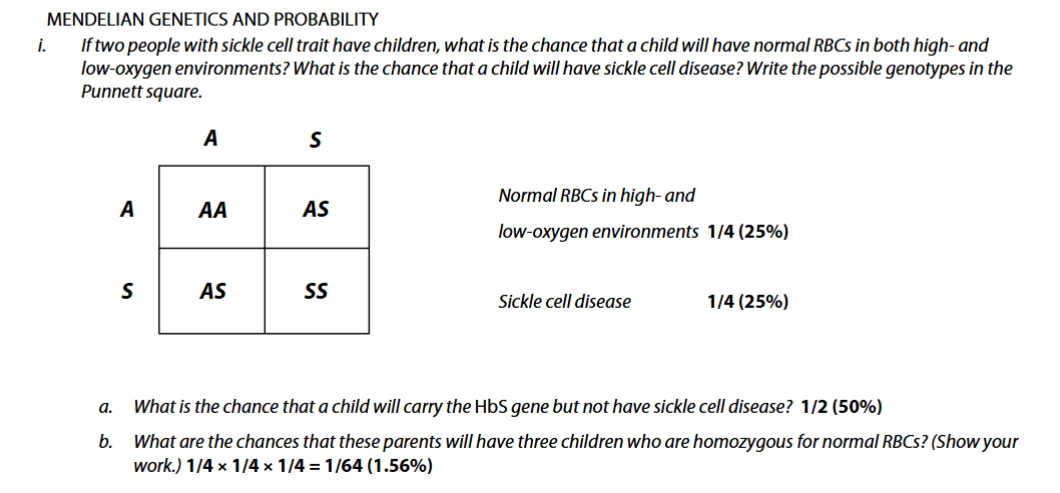
sickle cell practice problem
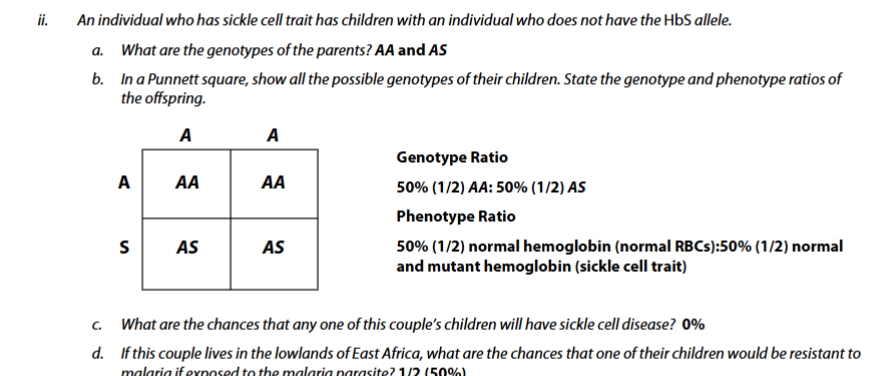

sickle cell practice problem
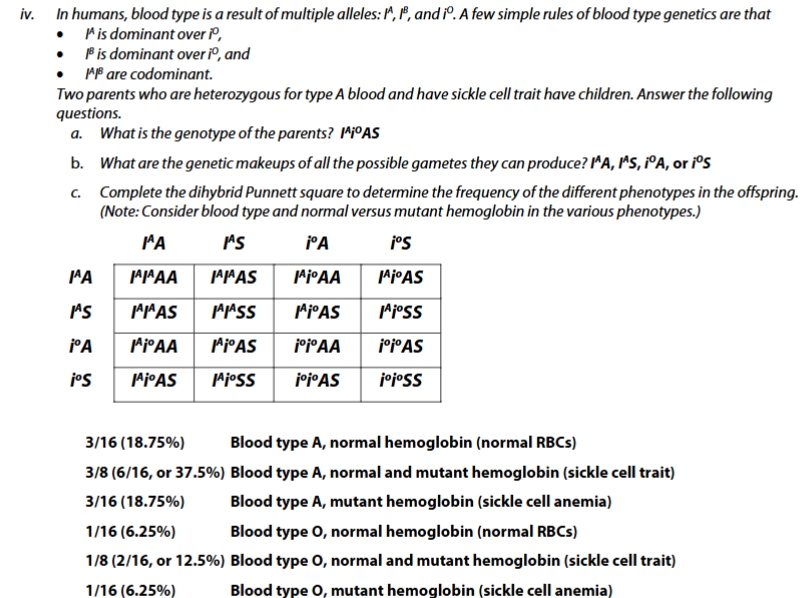

sickle cell practice problem
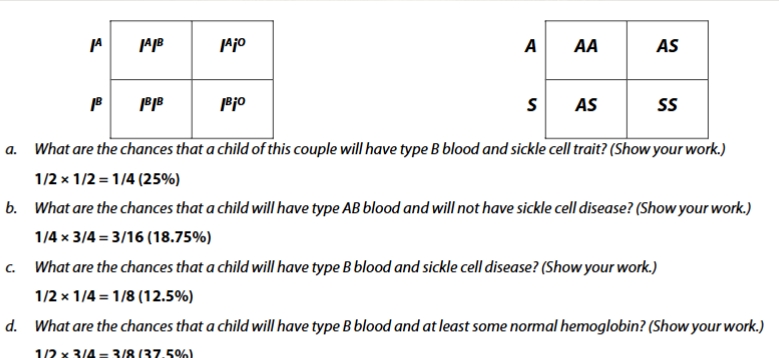

sickle cell pedigree practice problem
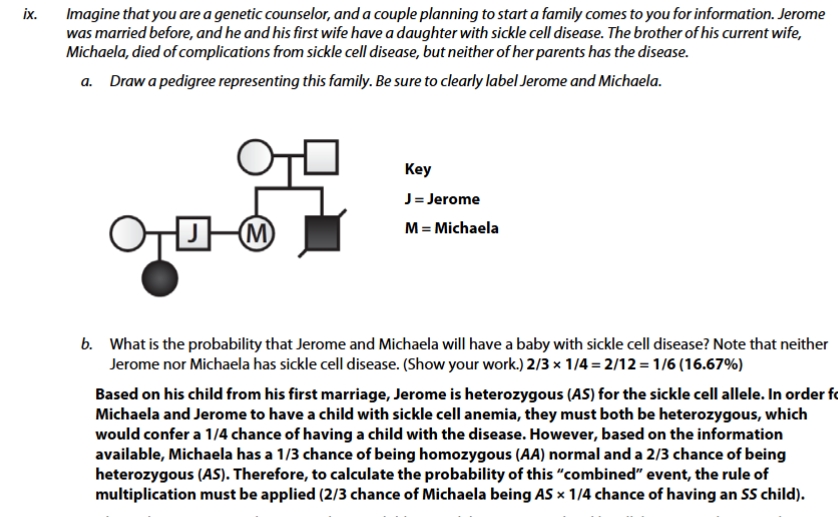
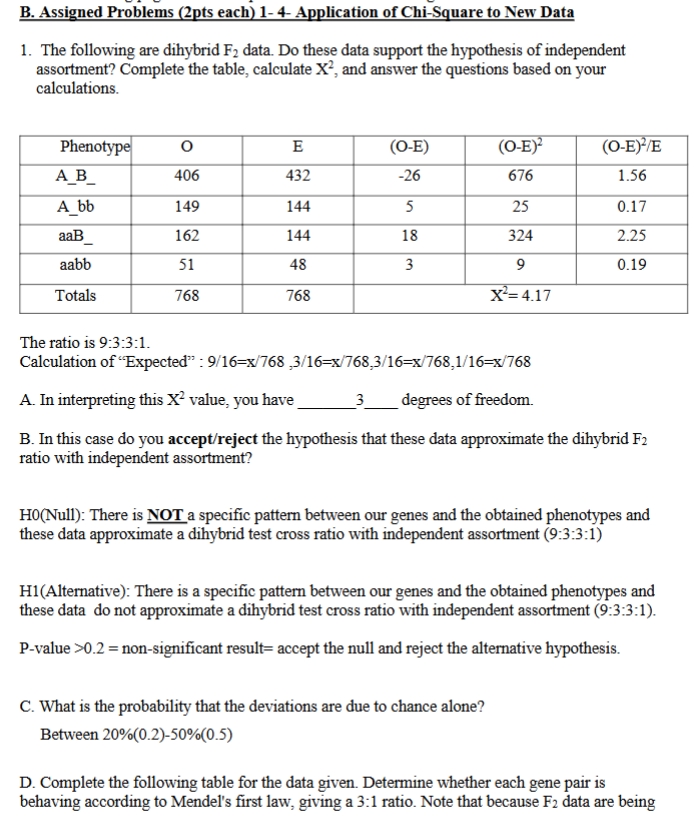
chi sqrd practice problem sickle cell
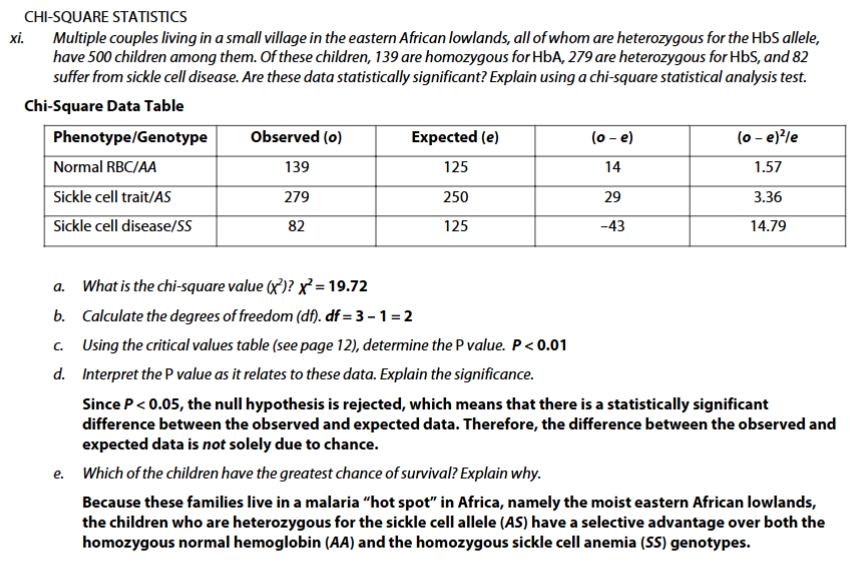
chi sqrd practice
accept null hyothesis : p-value > 0.05 = no significant difference between observed and expected values
reject null hypothesis: p-value < 0.05
expected value = total/ 16 x ratio
ratio=
df= n (number of columns) -1
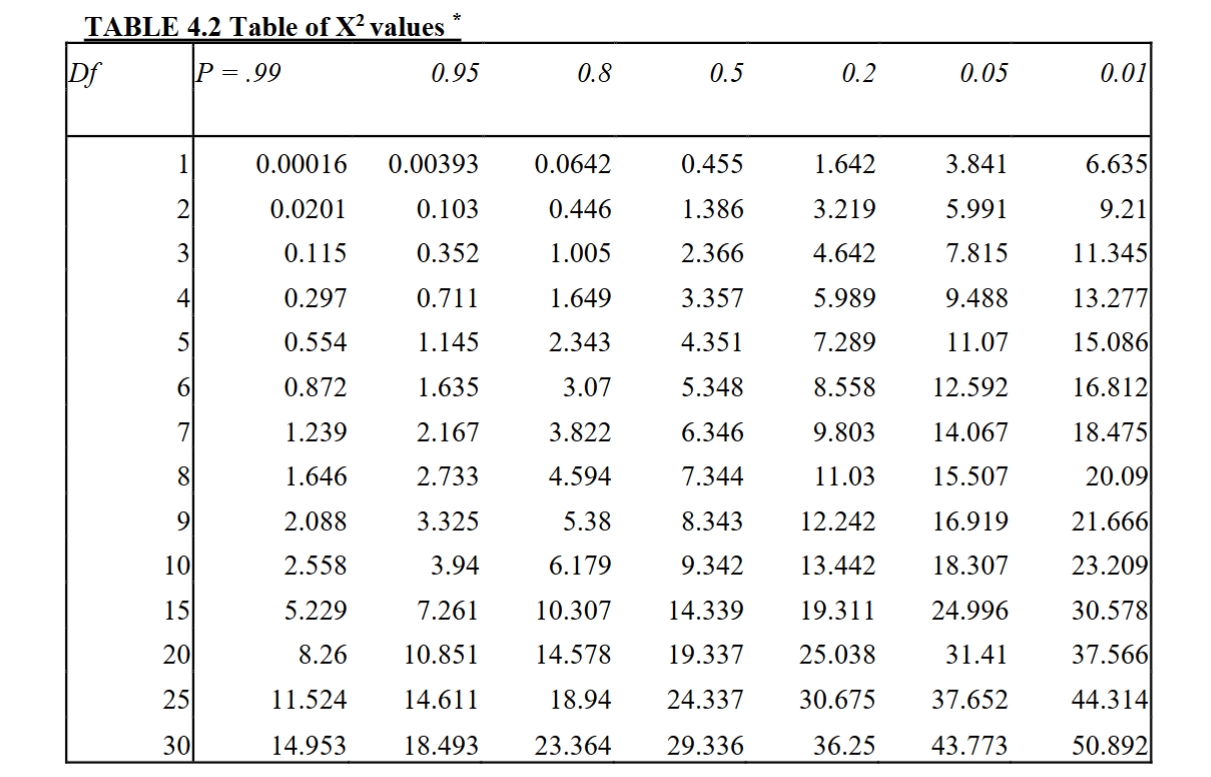
chi squared practice for expected value
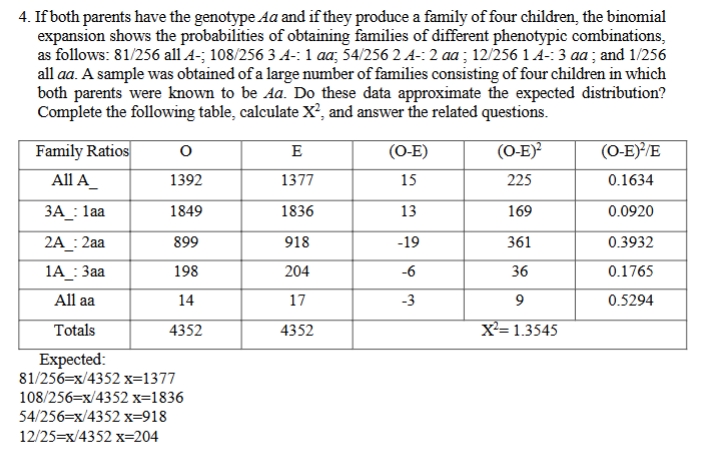
chi sqrd practice problem
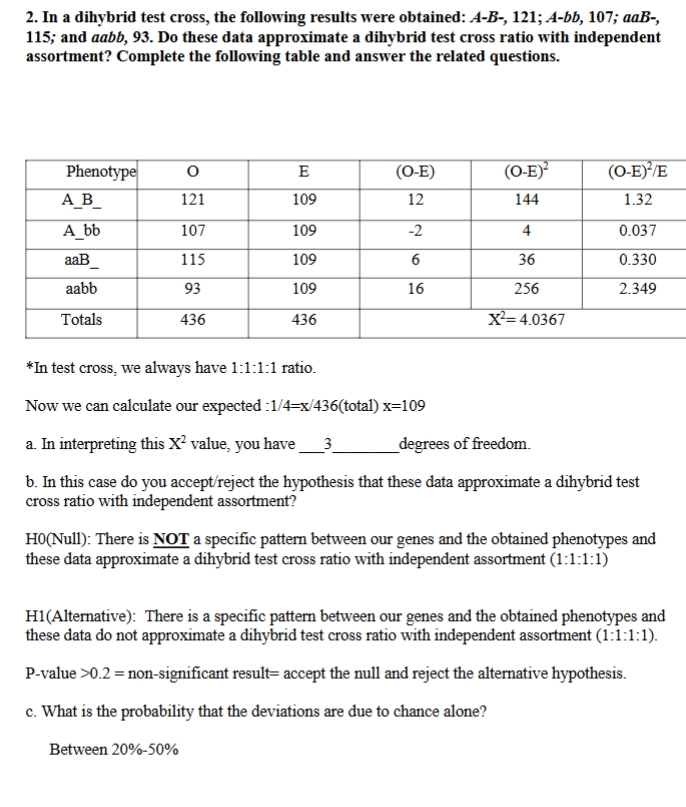
chi sprd practice problem

Acquired (somatic) mutations:
Exposure to mutagens that affect the DNA Errors during replication
Germline mutations:
Directly inherited through generations
p53, tumor suppressor protein:
• Gene located on the short arm of chromosome 17.
• Mutations to the gene causes the protein to loses its ability to bind to DNA.
• p53 that have mutations in specific hot spots promote uncontrolled cell growth and therefore function as oncogenes.
• For p53 to play a role in cancer, both alleles need to be altered.
what is karyotyping
Chromosomes are generally classified using multiple criteria. They are matched and numbered from largest to smallest, G-banding, and centromere location. homologous chromosomes 1-22 and then the 23rd pair of sex chromosomes.
sex chromosomes are labeled appropriately as either XX in females or XY in males. Note that the X-chromosome is much larger than the Y-chromosome. Therefore, the X- and Y- chromosomes are considered nonhomologous (although there are a few homologous regions.
pedigree symbols
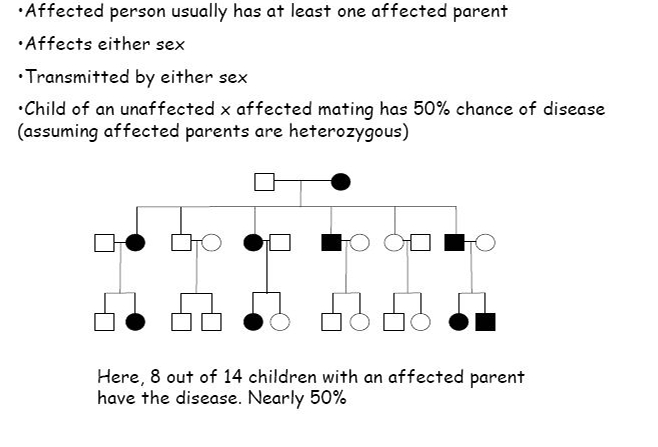
autosomal dominant disease
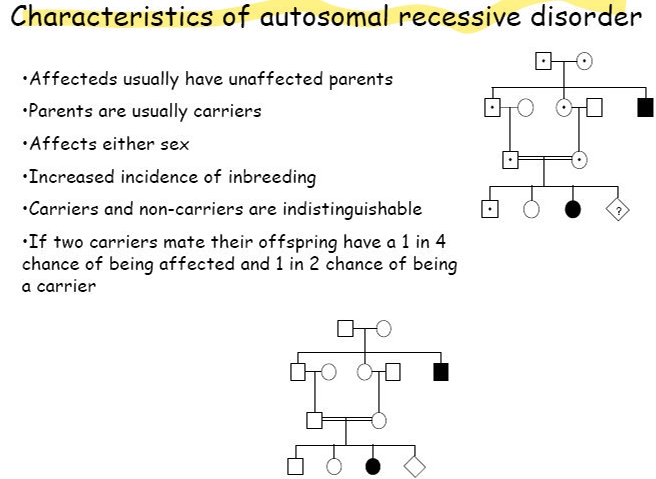
autosomal recessive disease
X-linked dominant disorders
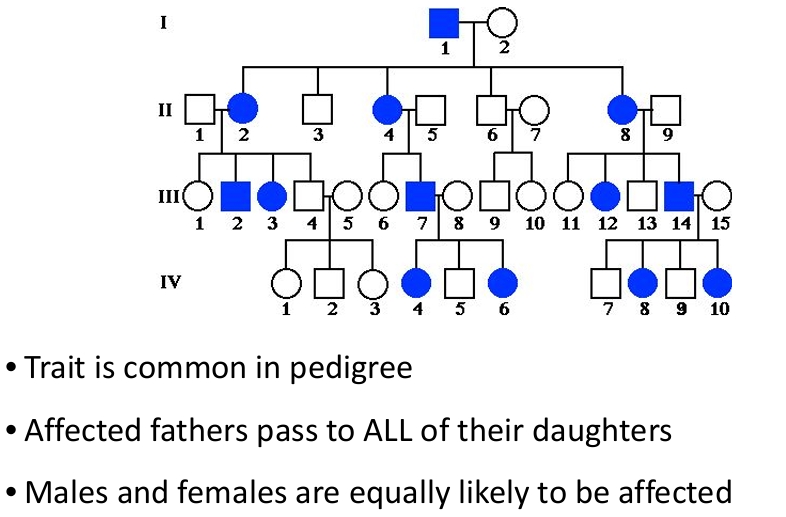
X-linked recessive disorders
Trait is much more common in males than females
• An affected man passes the gene to all of his daughters
• A son of a carrier mother has a 50 % chance of inheriting the trait
• Male-to-male transmission never occurs
• Carrier females are usually asymptomatic, but some may express the condition with variable severity because of Lyonization, or X-inactivation.
• Trait is rare in pedigree/ skips generations
• Males are more often affected than females
females only affected if father affected and mother is a carrier
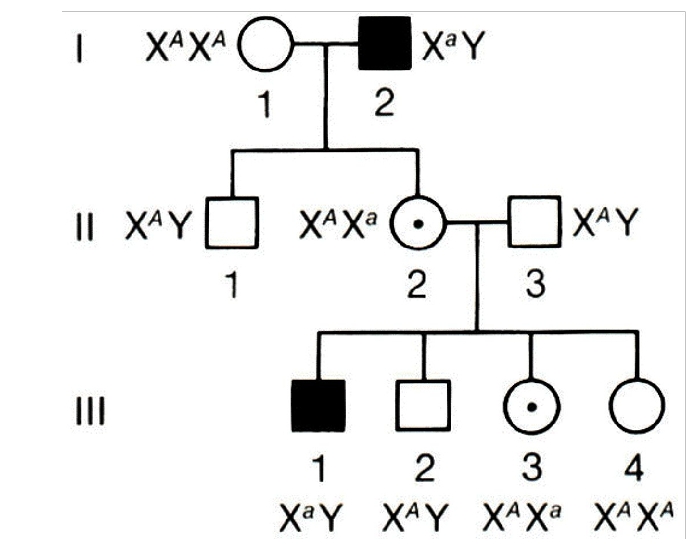
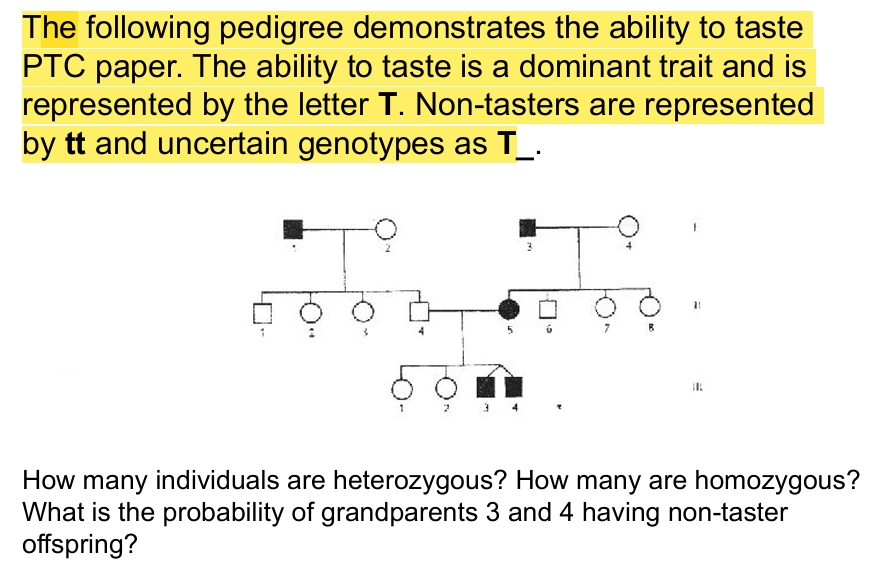
PTC paper practice pedigree problem

corn genetics!
purple kernels are dominant (P) and yellow kernels are recessive (p)
round (starchy) kernels are dominant (R) and wrinkled (sweet) kernels are recessive (r )
what makes the kernels purple?
anthocyanin coloring aleurone wherase in yellow kernels the aleurone is colorless due to defective enzyme
PTC
ptc- bitter if taster (T)
ptc- no taste if not (t)
PTC is present in about 70% o

Polymerase chain reaction
Three major steps in a PCR, which are repeated for 20 to 40 cycles. This is done on an automated Thermocycler, which can heat and cool the reaction tubes in a very short time.
•Denaturation at around 94°C : the double strand melts open to single stranded DNA
•Annealing between 54-55°C : Hydrogen bonds are constantly formed and broken between the single stranded primer and the single stranded template. When the primers find their complement, they attach. Researchers use specific primers to determine the region to be amplified
•Extension at around 72-75°C : Taq polymerase extends the primer by adding bases (A,T,C,G) complementary to the template. A new strand is built complementary to the template; A-T, C-G.
what are the requirements for successful PCR?
Template DNA
Primers
DNA nucleotides
DNA Taq Polymerase- found from Thermus aquaticus bacterium in hot springs
Buffer solution
how to find number of DNA copies made
every PCR cycle doubles number of DNA strands made.
Equation: 2(n ) ^# cycles = number copies made
what is restriction enzyme digest?
Each enzyme digests (cuts) DNA at a specific sequence = restriction site
Enzymes recognize 4- or 6- base pair, palindromic sequences (eg GAATTC)
Restriction enzyme digestion Restriction Fragment Length Polymorphism RFLP which is used to determine PTC taster from non PTC taster
GEL electrophoresis
separates DNA fragments by size and charge where negatively charged DNA fragments migrate towards the positive (anode) in a conductive buffer solution
the dna is loaded into troughs in the agarose gel and the results are compared to a standard
smaller dna fragments migrate farther than larger ones
what were the specific enzymes used for the restiction enzyme digest?
the PCR-RFLP method to examine the presence of an amino acid coding SNP. Students will use the PCR to amplify a polymorphic region of the TAS2R38 gene. The amplified DNA will be digested with the restriction enzyme HaeIII to determine their genotype at position 145, which correlates with the ability to taste PTC. Agarose gel electrophoresis of the restriction-digestion PCR products will reveal the 2 alleles of the TAS2R38 gene, indicating whether a student is homozygous or heterozygous for the taster phenotype, or a homozygous non-taster. In the final module, students will test their ability to taste the bitter PTC and correlate their genotype with their phenotype
