Northwestern University Bio 201 Exam 3
1/68
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
69 Terms
Central Dogma
DNA -> RNA -> Protein
Chemical differences between DNA and RNA
1) DNA contains the sugar deoxyribose, RNA contains the sugar ribose.
Ribose has one more -OH group than deoxyribose
2) Base pairing is different
In DNA, adenine (A) pairs with thymine (T). In RNA, adenine (A) pairs with uracil (U).
3) RNA is single stranded
5' to 3' direction for DNA
the only direction that DNA polymerase can synthesize DNA; it does so by adding nucleotides to the 3' end of a DNA strand.
RNA polymerase
Enzyme similar to DNA polymerase that binds to DNA and separates the DNA strands during transcription. Moves along template 3' to 5' beginning at +1 position. If moving in a different direction, always read 3' to 5' (mRNA strand is equivalent to DNA coding strand with U=T)
template strand of DNA
the DNA strand that is copied into mRNA and has the complementary sequence to the mRNA
coding strand of DNA
the DNA that is being mimicked by the RNA
Promoter
DNA sequences upstream of start of transcription that orient RNA polymerase in correct direction.
Bacterial Promoters
-35 and -10 from start site (+1). Promoter not found in transcript.
Sigma factor
A bacterial protein that recognizes the -10 and -35 regions of bacterial promoters positioning the RNA polymerase to initiate transcription at the correct start site (+1). It dissociates before transcription starts.
Eukaryotes have 3 types of RNA polymerases
RNA polymerase 1: transcribes most rRNA genes
RNA Polymerase 2: most important because it generates mRNA. all protein coding genes, miRNA genes, and genes for noncoding RNA like spliceosomes
RNA polymerase 3: tRNA genes, 5s rRNA genes, genes for other small RNAs.
Initiation
requires general transcription factors, regulatory DNA sequences far from the promoter regulate transcription, DNA packing into high order structures can alter transcription.
Eukaryotic Promoters
Also dictate RNA polmerase bindings. generally focus on TATA box. Sequences of the DNA bring in the right proteins
and orient the RNA polymerase so it goes in the right direction.
TATA-binding protein (TBP)
distorts DNA allowing a surface for other proteins can assemble called general transcription factors to recruit RNA polymerase 2.
initiation, elongation, termination
3 stages of transcription
Initiation of Eukaryotic Transcription
tata binding protein, recruits general transcription factors and eventually RNA polymerase 2 to the TATA-box containing promoter
Prokaryotic and Eukaryotic Transcription Similarities
Transcription occurs in initiation, elongation and termination.
RNA polymerases generases 5' to 3' direction.
RNA polymerase moves 3' to 5' on the template strand.
Transcription starts at +1 marker.
Genes can be transcribed from either DNA strand.
Promoter sequence direct the RNA polymerase in the proper orientation so a gene will be transcribed from the same strand every time.
multiple RNA polymerases can transcribe a gene simultaneously.
Prokaryote transcription differences
only has one RNA polymerase
sigma Factor binds promoter to recruit RNA polymerase during initiation
promoter is -35/-10
Eukaryote transcription differences
has 3 types of RNA polymerase, most important is RNA polymerase 2 which generates mRNAs
uses TBP and general transcription factors at promoter during initiation..
promoter is TATA box
RNA polymerase 2 has a C-terminal domain with important regulator functions.
RNA capping and polyadenylation (eukaryotic specific)
2 methods used to process primary transcripts to increase the stability of mRNA being exported to the cytoplasm. 5' cap and poly-A tail. Quality control because both together means you have a completed transcript.
5' Cap
Atypical guanine added to 5; nucleotide through 5' to 5' triphosphate linking. done by capping enzymes associated with RNA polymerase. recruits proteins that facilitate nuclear exit and protects mRNA from degradation. marks the message as mRNA and serves as landing pad for ribosome.
3' end polyadenylation
end of transcript is cleaved and add a string of adenine nucleotides ussign poly(A) polymerase. recruits proteins that facilitate nuclear exit and protects mRNA from degradation. marks the message as mRNA.
Introns
Noncoding segments of nucleic acid that lie between coding sequences in Eukaryotic transcripts (specific). Useful so mutations may not have an effect if they occur at introns, also allow for alternativesplicing and protein evolution.
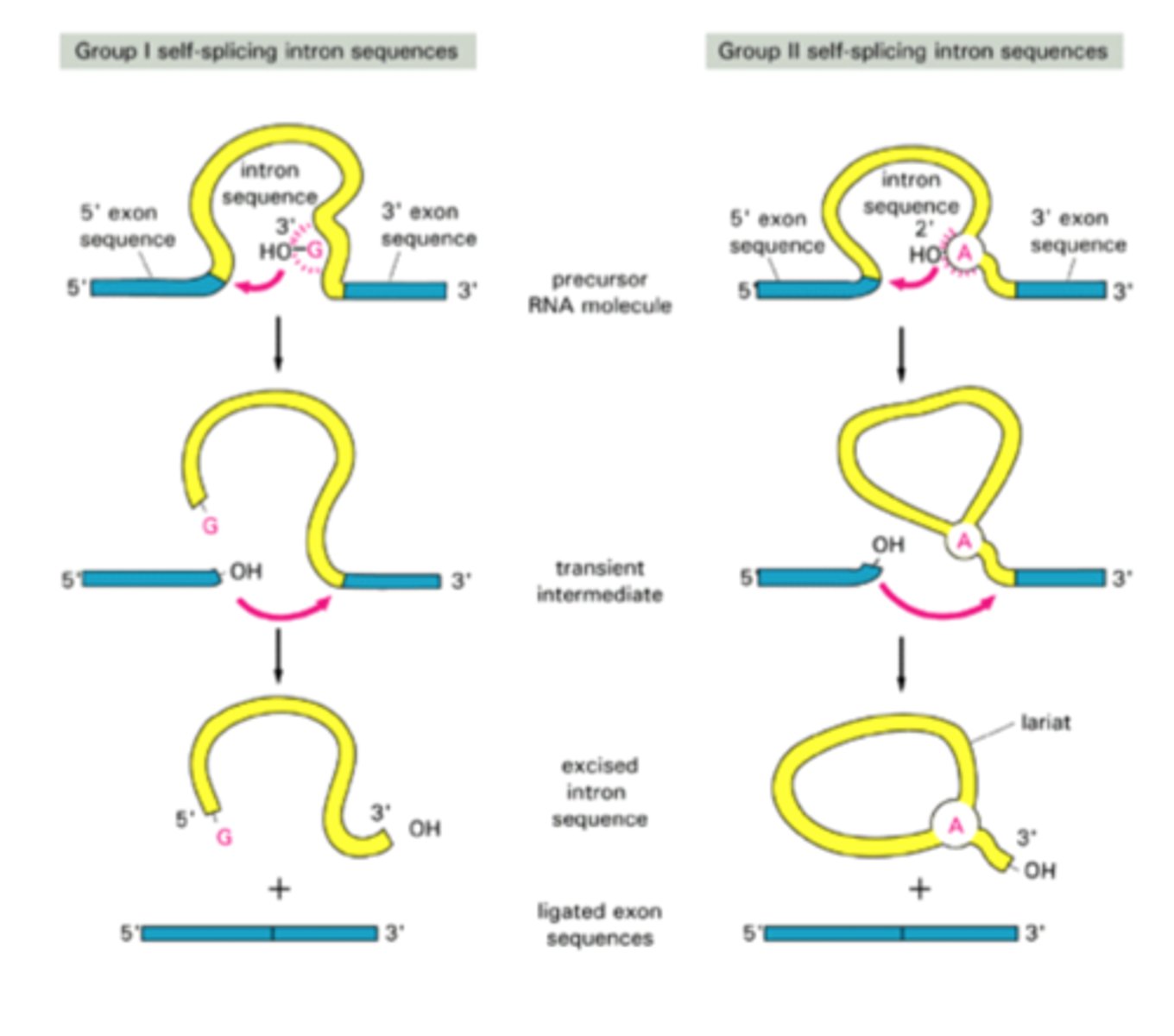
Splicing process of introns
Eukaryotic specific. like group II image.
1. 2'-OH in adenine (called the
branch-point) attacks the 5'
splice site, breaking the
sugar-phosphate backbone
2. It becomes covalently attached
to intron adenine
3. Frees a 3'-OH end of exon
reacts with start of next exon
joining the pieces together
4. The intron is released as a lariat
structure
How to find Splice Site
Use complementary base pairing of snRNPs.
Spliceosome contains both protein and RNA.
The RNA portion of snRNP base-pairs with sequences that signal splicing at each end of the intron to excise the intron as a lariat. Example of a Ribozyme because only RNA provides catalytic activity. A protein is then deposited to mark the splice site.
Alternative Splicing Process
generates related but different proteins from the same gene called isoforms. Changes such that internal exons can be removed, but start and end must be maintained and must be ordered in ascending order. ex 1234, 124, 134 or 14.
Alternative Splicing can retain/bypass certain functional units to
change protein binding properties, cellular localization, enzymatic activity, protein stability, and post-translational modifications.
Titin Alternative Splicing
when low levels of RBM20 are present, titin is predominantly spliced as longer springy isoform. if high levels are present, titin is short and stiff because it leaves out exons.
Fetal heart and titin
low RBM20: long springy titin protein. high ratio of fetal to adult titin leads to healthy fetal heart. high ratio of fetal to adult titin results in dilated cardiomyopathy.
high RBM20: short, stiff titin protein. low ratio of fetal to adult titin is a healthy adult heart.
Eukaryotic mRNA quality control overview
only complete mRNAs with a 5' cap and poly-A tail can be exported from the nucleus.
Elements of complete Eukaryotic mRNA are: 5' cap, 5' UTR, coding sequence with introns removed, 3' UTR, and poly-A tail.
Eukaryotic mRNA coordination
Phosphorylation of the C-terminal Tail Domain (CTD) of RNA polymerase recruits capping, splicing and polyadenylation factors. These processing events occur as the RNA polymerase moves along the DNA generating the mRNA.
C-terminal tail phosphorylation
Serines and Threonine amino acids near the
C-terminus of RNA polymerase II are phosphorylated by serine/threonine protein kinases. this allows promoter escape of RNA polymerase 2 and initiates transcription. it also brings capping, splicing and polyadenylation factors.
What determines level of gene expression? (how much protein is made by coding gene)
for both eukaryotes and prokaryootes, amount generated ddepnds on rates of transcription, translation, and degradation.
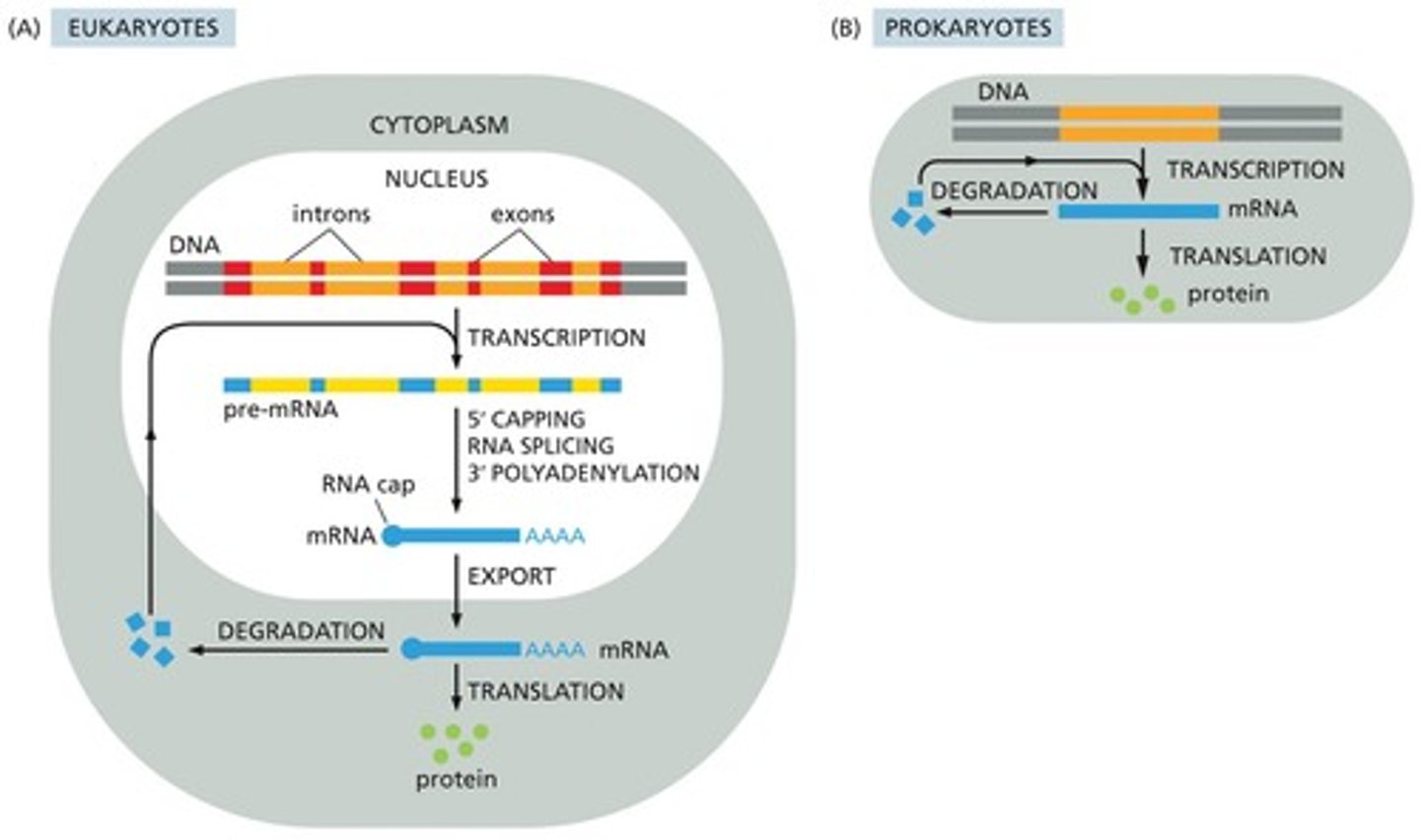
mRNA stability and degradation in Eukaryotes (NA for prokaryotes)
removal of 5' cap and poly-A tail. exonucleases will destroy the RNA breaking it down to ribonucleoted to eb recycled. Prokaryotic mRNAs are so short lived so there aren't many mecahnisms to control stability.
Converting nucleotides into amino acids/proteins
tRNAs act as adaptors. tRNAs use complementary base pairing to form cloverleaf structure. Anti-codon (3' to 5') base pairs with codon in mRNA (5' to 3') in the ribosome and amino acid covalently attached at 3' end.
aminoacyl-tRNA synthetase
An enzyme that joins each amino acid to the appropriate tRNA. Serves as bindng pocket for amino acid, anti-codon, amino-acid accepting arm. uses energy of ATP to conjugate amino acid to tRNA leaving a high energy bondto drive polypeptide chain formation in the ribosome.
Genetic code is degenerate
multiple codons encode the same amino acid (wobble), except for Met which is just AUG
Bacterial Ribosome Differences
60% rRNA and 40% protein by mass, simultaneous transcription and translation. free in cytosol. less proteins and rRNAs in both subunits than eukaryotic.
Eukaryotic Ribosome Differences
40% rRNA and 60% protein by mass. free in cytosol or bound to rough ER for secreted proteins. More proteins and rRNAs in both subunits than prokaryotic/bacterial.
Eukaryote and Prokaryote Ribosome Similarities
Composed of a small and large subunit
Ribosomal RNAs give the complex its structure
RNA catalyzes the peptide bond reaction, so it is considered a ribozyme
Small subunit matches tRNAs to the codons of the mRNA
Large subunit RNAs catalyze formation of the peptide bonds that covalently link the amino acids together into a polypeptide chain
Large ribosomal subunit function
formation of peptide bonds by covalently linking amino acids
Small ribosomal subunit function
matches tRNA to mRNA with complementary base pairing
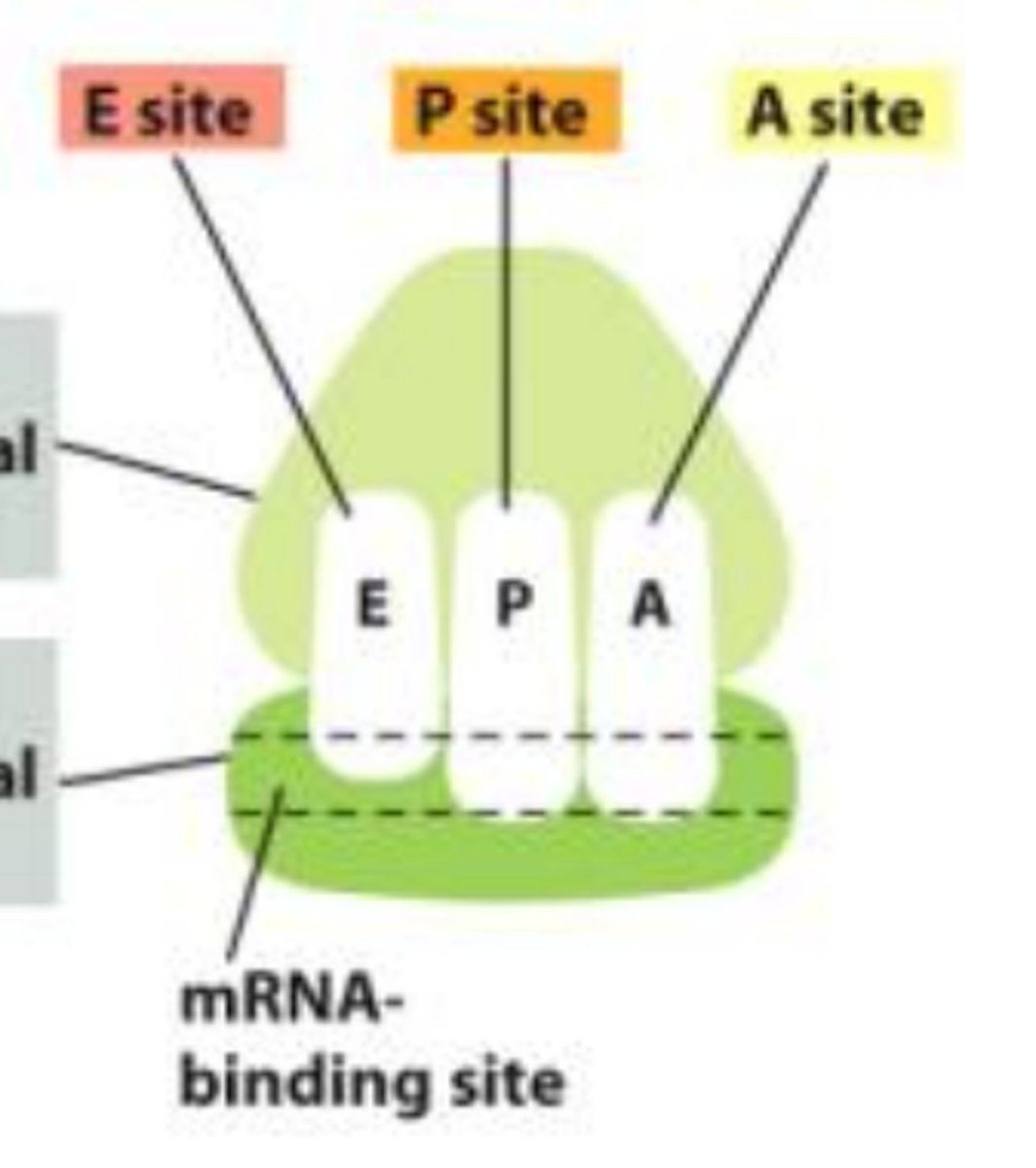
EPA
E=exit site (tRNA leaves), P=Peptidyl site, A=aminoacyl site. Progresses A>P>E right to left.
Eukaryotic translation initiation
initiator tRNA with Methionine binds in P-site of small subunit with translation initiation factors. recognizes the 5' cap on the mRNA and scans until it finds the first AUG (Met). Large subunit binds and translation begins.
Eukaryotic translation termination condition
When the ribosome encounters a UAA, UAG or UGA stop codon and the release factor hydrolyzes releases of polypeptide chain.
Bacterial translation initiation
begins with Shine-Dalgarno sequence in mRNA. specific seqeunces in mRNA base-pair with RNA in the small subunit to position the intiator formyl-Methionine tRNA over the AUG.
Shine-Dalgarno Sequence
AGGAGGU used in bacterial translation initiation
Protein production EPA step 1
growing polypeptide chain binds to P site. newly bound charged tRNA binds to A site.
Protein production EPA step 2
New peptide bond forms such that chain at P site attaches to new tRNA at A site.
what catalyzes peptide bond formation in protein production?
the ribosome
Protein production EPA step 3
large subunit translocates three nucleotides to the right leaving small subunit behind such that P->E and A->P leaving A open.
Protein production EPA step 4
Ejects tRNA from E site, and small subunit translocate three nucleotides in the 3' direction to match large subunit, restarting the process with a polypeptide chain at P, with A and E open.
Read-through mutations
mutations that alter a stop codon
release factor protein
catalyzes hydrolysis reaction which releases polypeptide chain from the P-site tRNA
Polycistronic
Many bacterial mRNAs have one mRNA with more than one polypeptide chain. Ribosome can initiate translation from anywhere in the transcript that has a Shine-Dalgarno sequence followed by an AUG start codon.
Monocistronic
Eukaryotic mRNA has one gene, one mRNA, and one polypeptide chain. Unlike in prokaryotes, ribosome cannot enter the mRNA anywhere except by scanning from the 5' cap into the mRNA.
polyribosome (or polysome)
many ribosomes can bind to one mRNA. number of ribosomes on a mRNA can be used to determine efficeincy of translation.
Silent Mutation
A mutation that changes a single nucleotide, but does not change the amino acid coded for. Example is DNA AAG, mRNA AAG which codes for Lys being mutated to DNA AAA, mRNA AAA which still codes for Lys.
Nonsense mutation
Mutation that changes a single nucleotide such that at the mRNA level it codes a stop codon. Example is DNA AAG, mRNA AAG which codes for Lys changing to DNA TAG, mRNA UAG which is a stop codon.
missense mutation (conservative)
Mutation that changes a single nucleotide such that new AA is similar in chemical structure. A mutation that changes a Lysine to an Arginine may be considered a conservative missense mutation because both of these amino acids are amine, hydrophobic and have similar structure so the mutation is not likely to have a significant impact on the function of the protein.
missense mutation (nonconservative)
Mutation that changes a single nucleotide such that new AA is different in chemical structure. A mutation that changes a Lysine to an Threonine may be considered a non-conservative missense mutation because these amino acids are have different structures so the mutation may have a significant impact on the function of the protein.
Frameshift mutation
mutation that shifts the "reading" frame of the genetic message by inserting or deleting a nucleotide. can change the amino acids coded for in the RNA.
Prokaryotic and eukaryotic transcription differences
Bacteria use sigma factor and RNA polymerase to initiate transcription. requires a bacterial promoter -35/10 in the plasmid.
Eukaryotes have introns that must be spliced from the final transcript using cDNA as template in PCR.
Generate DNA without introns - making a copy of mRNA
Use cDNA to get exons only, no introns. by bonding poly-A tail with poly-T primer. Complete reverse transcriptase starting at tail to form RNA/cDNA helix. partially degrade RNA with RNAse, then synthesize a complementary DNA strand using DNA polymerase to get completed double stranded cDNA molecule.
Requirements to replicate DNA
1. Template sequence (cDNA)
2. Method to separate double stranded template DNA. (heat replaces helicase in PCR)
3. Primers. use synthesized DNA primers to eliminate need for primase, nuclease and ligase.
4. Enzymes. DNA polymerase.
5. Nucleotides.
For recombinant DNA, plasmid must have
ORI, -35/-10 promoter to recruit RNA polymerase, Shine-Dalgarno Sequence, Multiple cloning site, selectable marker
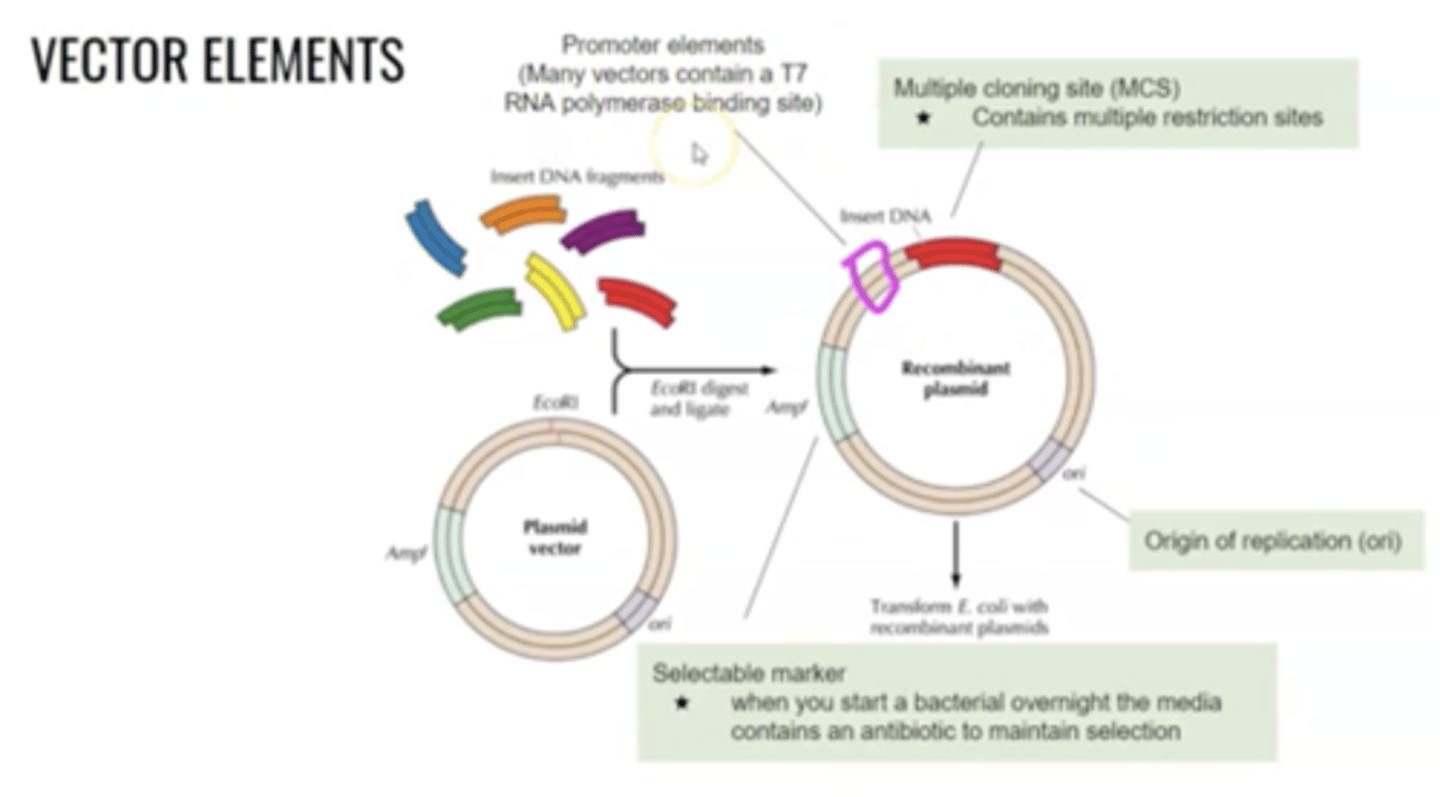
For recombinant DNA, bacteria must
copy the plasmid, promote expression of inserted gene, allow for insertion of new genes and retain the plasmid.
Selectable marker
all bacteria don't take up the plasmid, so we select the ones that did by first letting bacteria grow, and then because bacteria with plasmid express drug resistance gene selectable marker, we plate them onto a surface with the drug and the surviving cells will form a colony.
Recombinant DNA steps
PCR, digest, ligate, transform, plate.
1. PCR sequence of interest containing
proper restriction enzyme sites at the 5'
and 3' end (cDNA template)
2. DIGEST the backbone/vector with the
same restriction enzymes
3. LIGATE the insert with the vector in a test
tube (outside of the cell)
4. TRANSFORM into bacteria cells
5. PLATE onto petri plates with agar and drug
to kill any cells that did not take up
plasmid
Gel electrophoresis in recombinant DNA
Since DNA sugar-phosphate backbone is negatively charged the DNA will run toward the positive electrode The agarose in gel forms "pores" that the DNA must travel through. Large molecules can not travel as far as shorter molecules. DNA is then visualized using a Dye.