KIN217 20-25CH
1/89
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
90 Terms
Oxidative phosphorylation location
Mitochondria
Mitochondria evolutionary endosymbiosis
-Was once and independent organism that was engulfed by another cell
-Has some DNA
Mitochondrial outer membrane
-defines intermembrane space
-permeable to small ions and molecules through mitochondrial porin
Mitochondrial inner membrane
-defines matrix
-folded into cristae
-high protein content
-high amounts of cariolipin
Heart vs liver mitochondrion
Heart mitochondrion have many more cristae allowing for more energy production
Electron transfer potential
Measure of as molecule’s tendency to donate or accept electrons
Negative E0’
-reducing agent that donates electrons
-low e- affinity
Positive E0’
-oxidizing agents that accept electrons
-high e- affinity
Electron movement in the chain
From low E0’ to high E0’
NADH-Q oxidoreductase
-complex I
-accepts 2 e- from NADH and trasfers to ubiquinone
-pumps 4 protons
Succinate-Q reductase
-complex II
-accepts e- from FADH and transfers to ubiquinone
-not a proton pump
Q-cytochrome c oxidoreductase
-complex III
-accepts e- from QH2 and transfers to cytochrome c
-pumps 4 protons
Cytochrome c oxidase
-Complex IV
-accepts 4 e- from cytochrome c and transfers to oxygen
-pumps 4 protons
Respirasome
-complexes of the ETC associated with one another
-mainly complexes I, III, IV
QH2
Reduced form of ubiquinone, carries electrons
Succinate dehydrogenase
-Kerb’s cycle enzyme, reduces FAD to FADH2
-converts succinate to fumarate
coenzyme Q (ubiquinone)
Serves as a shuttle for electrons between complexes I/II, and III
Process of Q-cytochrome c oxidoreductase
-Cyt C can only receive 1e- at a time
-1st QH2 gives Cyt C 1e- (pumps 2 protons)
-2nd QH2 gives 1e- to CYT C and other to make Q
Cytochrome C
-protein with heme group
-carries 1 e- on a heme iron
-transfers electrons from complex III to complex IV
Protons pumped per NADH
-Total of 10
-4 at complex I
-4 at complex III
-4 at complex IV
Protons pumped per FADH2
-Total of 6
-4 at complex III
-2 at complex IV
reactive oxygen species
-ROS
-result of partial reduction of oxygen
-can oxidize other compounds spontaneously
ROS defence
-Superoxide dismutase
-catalase
-exercise increases the expression of these enzymes
Proton gradient
-the respirasome pumping of electrons results in an unequal distribution of protons
-electrochemical gradient
Proton motive force
-use the energy of the proton gradient to drive ATP synthesis
-protons flow through ATP synthase like a waterwheel
energy of ATP synthesis
-endergonic process
- +30.5kJ/mol
F0 component
-embedded in the inner mitochondrial membrane
-contains the proton channel
F1 component
-protrudes in the mitochondrial matrix
-contains 3 catalytic beta subunits, each in a different form
Connection of F0 and F1 subunits
-gamma subunit
-external column
Beta subunits
contains active sites that generates ATP
Alpha subunits
isolate beta subunits from each other
ATP synthase and shape of mitochondria
-form dimers and cluster together to form curvatures in the cristae of the mitochondria
-clustering stabilizes rotational forces, increasing efficiency
O (open) form
nuleotides can bind to or be released from the beta subunit
L(loose form)
nucleotides are trapped in the beta subunit
T(tight) form
ATP is synthesized from ADP and Pi
Subunit a
-2 half channels for proton flow in F0
-one opens into intermembrane space, one into matrix
Ring of c subunits spinning
-glutamate residue at spot on c subunit is exposed to each half channel of subunit a
-protons bind to gluamate to form glutamic acid
-subunit c with no charge can then move into nonpolar region of phospholipids
What dictates direction of spin
-proton gradient
-entry of protons into half channel
protons and ATP
-nned 4 protons per ATP
-8 protons per spin
-3 ATP per spin
-1 proton for substrate availability/ATP export
ATP-ADP translocase enzyme
-antiporter
-ATP export is coupled to ADP import
Pi into matrix
-phosphate carrier
- -OH antiporter
ATP synthasome
ATP synthase + ATP-ADP translocase + phosphate carrier
NADH from glycolysis
cytoplasmic NADH needs to get into matrix before being used by ETC
glycerol-3-phosphate shuttle
-prominent in muscle
-cytoplasmic NADH transfers e- to DHAP to form glycerol-3-phosphate
-G-3-P transfers e- to FAD in mitochondrial G-3-P dehydrogenase
-FADH2 transfers e- to Q
glycerol-3-phosphate shuttle control
shuttle functions regardless of matrix [NADH]
malate-asparate shuttle
-prominent in heart and liver
-cytoplasmic NADH transfers e- to oxaloacetate to form malate
-malate transfers e- to NAD+ in matrix to go back to oxaloacetate
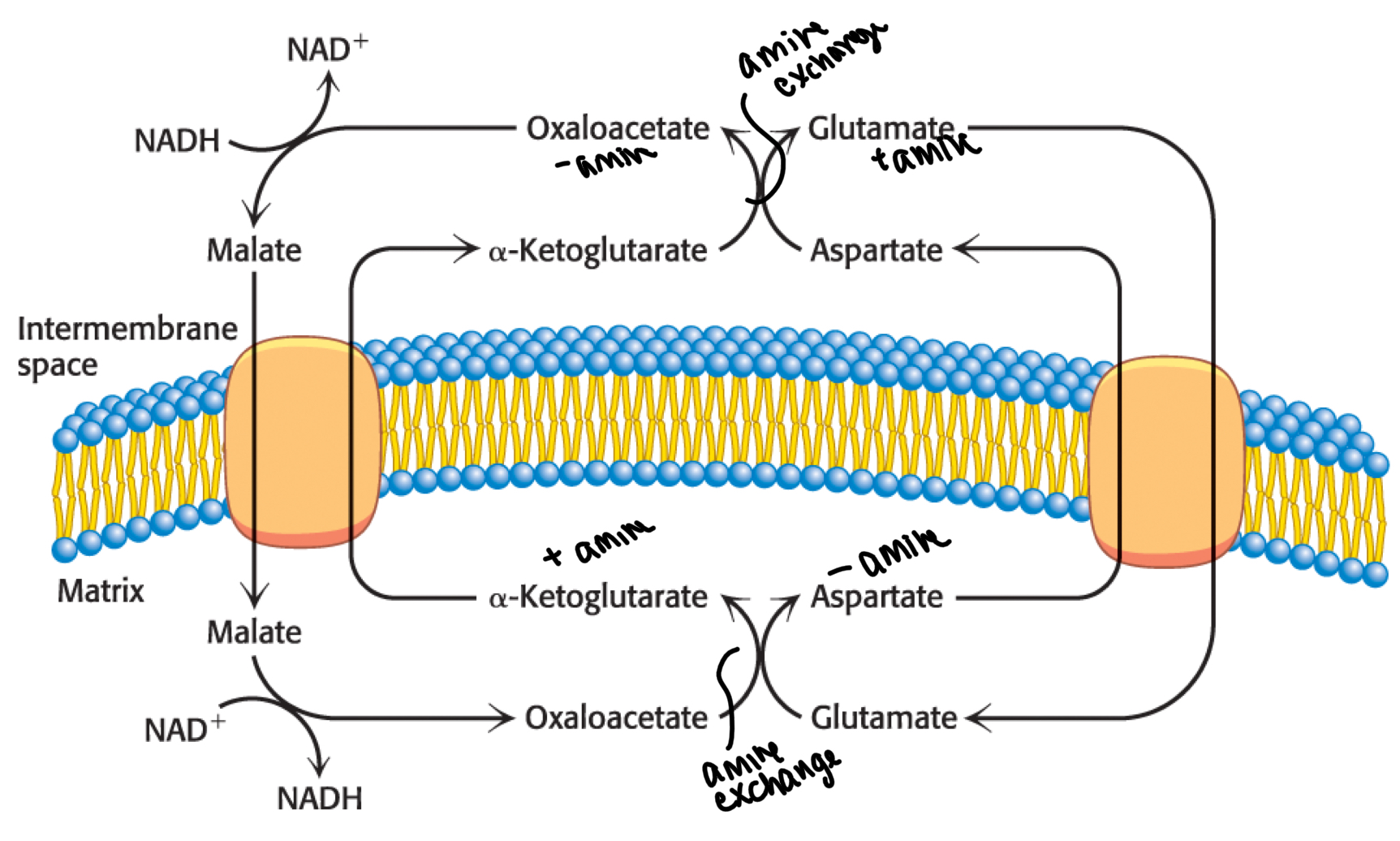
High martix NADH can inhibit shuttle
malate-aspartate shuttle control
ATP from matrix NADH (from ETC)
2.5
ATP from FADH2 (from ETC)
1.5 (FADH2 doesn’t pump as many protons, not in complex I)
ATP from cytoplasmic NADH
-G-3-P dehydrogenase transport
~1.5 ATP
-malate-aspartate transport ~2.5 ATP
total ATP per molecule of glucose
30-32 ATP
control of oxidative phosphorylation
-dictated by [ADP]
-electrons do not flow from fuel to O2 unless there is a need to make ATP
proton gradient making heat
-uncoupling proton gradient from ATP synthesis
-protons flow through uncoupling proteins
-energy not captured chemically and released as heat
brown fat
-specializes in uncoupling
-high in hibernating animals
-in humans high in infants, higher in females
-can be increased with cold exposure
thermogenesis and weight loss
-increased heat production and less ATP prdocution
-not efficient metabolism
-dangerous
Glycogen
-a highly branched homopolymer of glucose present in all tisues
-formed and stored in the cytoplasm
liver glycogen
-10% by weight
-maintains blood glucose when fasting
muscle glycogen
-2% by weight but larger store than liver due to higher muscle mass
-provides glucose for sudden strenuous exercise
glycogen stucture
-straight chains have alpha-1,4,-glycosidic bonds
-branch chains(every tenth residue) have alpha-1,6-glycosidic bonds
-non reducing ends (OH) on exterior
glycogenin
protein at the core of glycogen
glycogen breakdown steps
-degraded glycogen
-remodel glycogen
-convert breakdown product into usable product
use of G-6-P after release
-enter glycolysis
-in liver it can be converted into free glucose
-processed in the pentose phosphate pathway
cleaving glycogen
-glycogen phosphorylase degrades glycogen from the nonreducing ends
-catalyzes a reaction that yields glucose 1-phosphate
remodeling glycogen problems
-glycogen phosphorylase cannot cleave the 4 glucose residues near branch points
-glycogen phosphorylase can only cleave alpha-1,4-glycosidic bonds
remodeling glycogen solutions
-bifunctional debranching enzyme
-transferase and alpha1,6-glucosidase activity
debranching transferase activity
-shifts small oligosaccarchides near the branch point to a nearby chain
-exchange of alpha-1,4-glycosidic bonds
-shifted glucose moieties become accessible to phosphorylase
debranching alpha-1,6-glucosidase activity
-cleaves alpha-1,6 bond at the branch point using H2O
-releases a free glucose (no phosphate)
making glucose-6-phosphate
-phosphoglucomutase
-enzyme has a phosphorylated serine residue, adds phosphate onto C-6
-removes phosphate from C-1
glycogen phosphorylase
-key enzyme for regulating glycogenolysis
-has a more active ‘a’ form and less active ‘b’ form
-active ‘a’ form has phosphorylated serine
glycogen phosphorylase a and b
-both have R and T state
-in the ‘a’ form, R state is favoured
-in the ‘b’ form, T state is favours
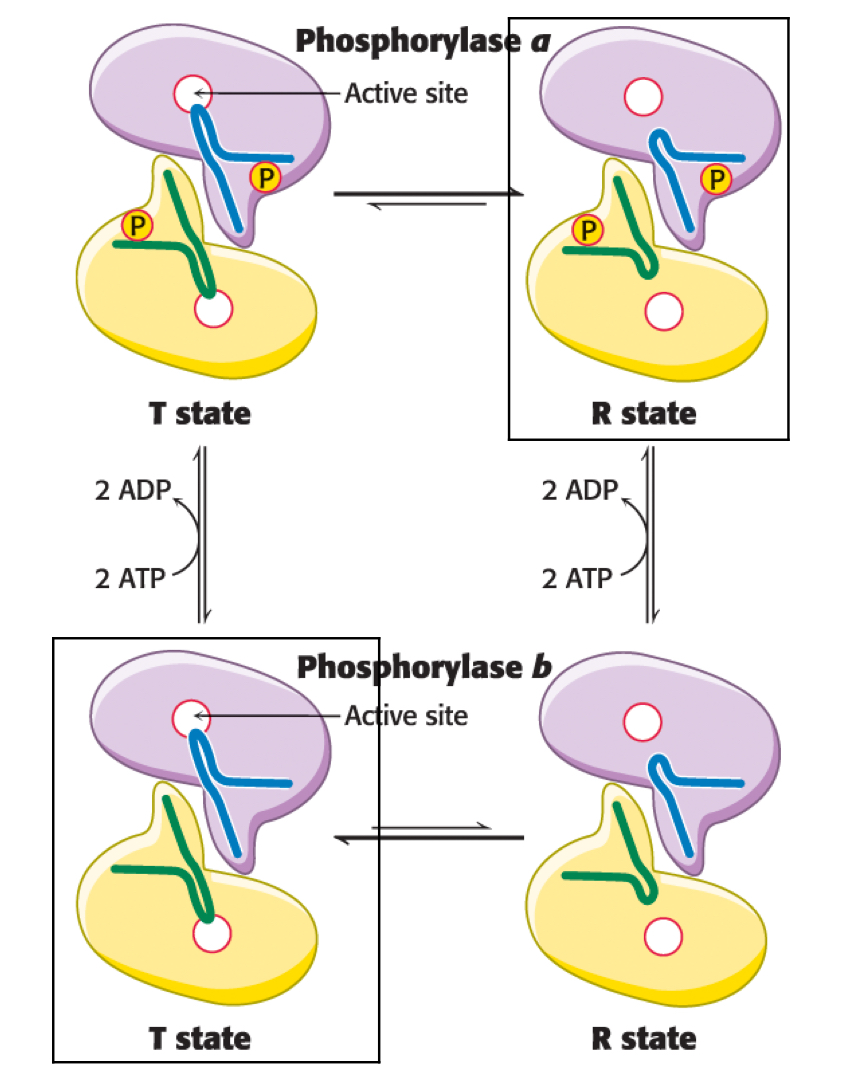
muscle glycogen phosphorylase
-default is the ‘b’ form in T state
-when energy is needed shifts to R state by binding of AMP
-in high energy shifts to T state with binding or ATP or glucose-6-phosphate
epinephrine and muscle glycogen phosphorylase
-phosphorylates enzyme to active a form
-active regardless of ATP, AMP or glucose-6-phosphate
liver glycogen phosphorylase
-default is the ‘a’ form in R state
-glucose as a negative feedback inhibitor
hormonal regulation of liver glycogen phosphorylase
-glucagon/epinephrine activates
-insulin deactivates
phosphorylating glycogen phosphorylase
-adding phosphate shifts from ‘b’ state to ‘a’ state
-maximally active when phosphorylated and bound to calcium
glycogen and fatigue
-fatigue is associated with depletion of glycogen stores
-low glycogen may result in increased ADP
UDP glucose pyrophosphorylase
-substrates are UTP and G-1-P
-products are UDP-glucose and pyrophosphate
-reversible
pyrophosphates
-hydrolyzes pyrophosphate to make 2Pi
-irreversible
glycogen synthase
-key regulatory enzyme
-transfers glucose from UDP glucose to C-4 of the terminal residue of the glycogen chain
-can only add glucose to chain of 4 or more residues
glycogenin
-glycogen priming enzyme
-makes alpha-1,4 chain
-remains attached to glucose via tyrosine residue
branching enzyme
-makes alpha-1,6 linkage
-breaks off alpha-1,4 linkage with ~7 glucose
branching enzyme specifics
-chain tha twas broken must have at least 11 glucose
-new alpha-1,6 linkage is at least 4 residue inwards
glycogen synthase and regulation
-active in unphosphorylated ‘a’ form
-inactive in phophorylated ‘b’ form
-G-6-P stabilizes active R state of b form
reciprocal regulation of glycogen breakdown and synthesis
-glycogen synthesis is inhibited by glucagon and epinephrine
-activated with insulin
-same signalling pathways that stimulate glycogen breakdown
PP1
-protein phosphatase 1
-shifts glycogen metabolism to synthesis
-removes phosphates from glycogen synthase b
epinephrine/glucagon and PP1
-need glucose signal
-activates pkA, phosphorylates regulatory subunit Gm
-allows inhibitor to bind
insulin and PP1
-store glucose signal
-phosphorylates and inactivates glycogen synthase kinase
-PP1 can dephosphorylate glycogen synthase to activate
type 1 diabetes
-autoimmune attack on beta cells
-no insulin production
-poor uptake of glucose, high blood glucose
type 2 diabetes
-more common
-insulin resistance
-glucose uptake impaired, high blood glucose
-insulin still inhibits hormone sensitive lipase in adipose so low risk of acidosis
natural selection and type 2 diabetes
-polygenic
-people with ancestry that had a history of surviving starvation/ famine are at higher risk