Midterm 2
1/128
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
129 Terms
What is haplosuffiency?What is haplosuffiency and an example?
one copt of WT allele sufficient for activity
normal Mendellian genetics
dominant allele gets expressed
What is haploinsufiency and an example?
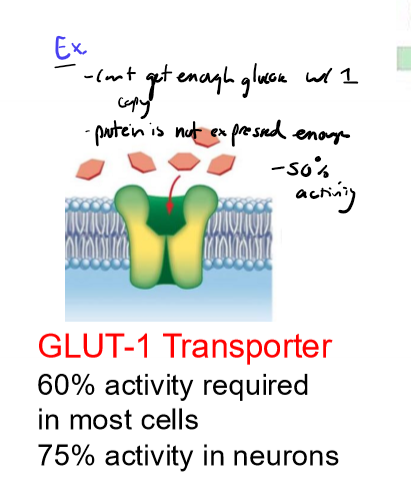
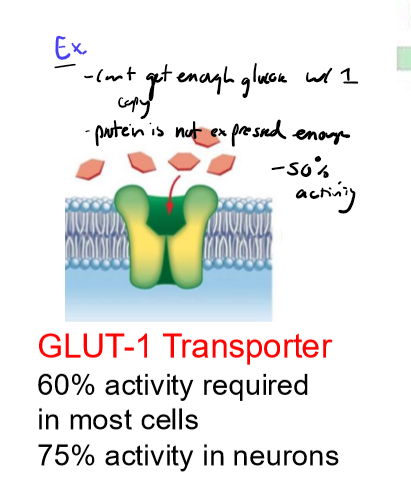
one copy of WT allele not sufficient for activirty
recessive mendialian genetics
ex. GLUT-1 Transporter, 60% activity required in most cells to get enough glucose in, with 50% activity doesn’t work
What is a dominant negative mutation?
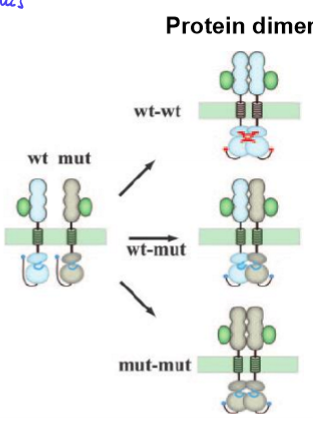
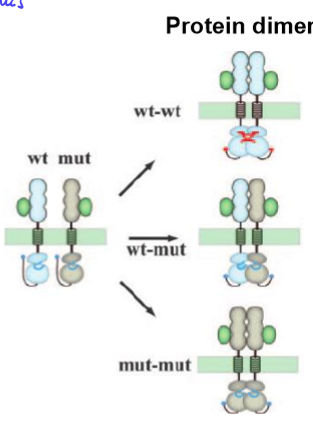
heterozygous has NO function
mutant allele interferes with normal WT functioning
ex. WT-mut disrupts protein shape, protein can’t form, so no function
What is an allelic series?
A set of more than two alleles at a single gene locus that show a range of phenotypic effects or dominance relationships.
ex. Rabbit coat color (C > c^ch > c^h > c), producing full color, chinchilla, Himalayan, albino
Q: What determines whether an allele is dominant or recessive at the molecular level?
Dominance depends on gene dosage and protein function
Example: A normal enzyme allele (A) makes enough enzyme for function → dominant over a nonfunctional allele (a)
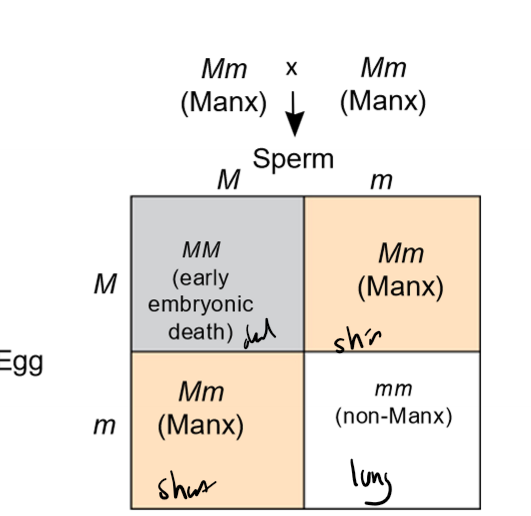
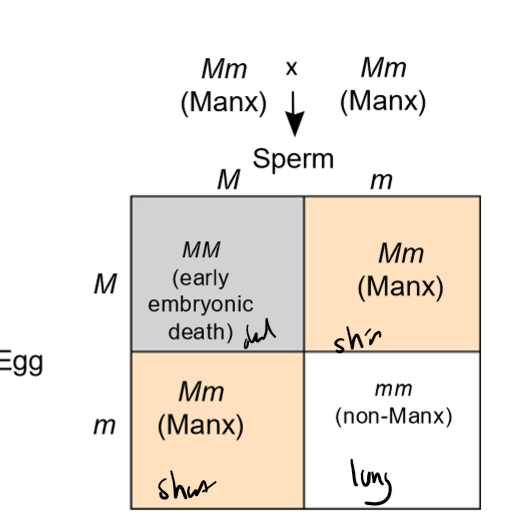
How do lethal alleles produce modified Mendelian ratios?
lethal alleles cause death prenatally when homozoygous, removing that genotypic option
ex. Aa x Aa; aa produces death, 2:1 genotypic ratio in offspring
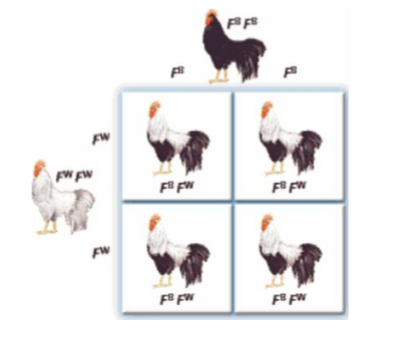
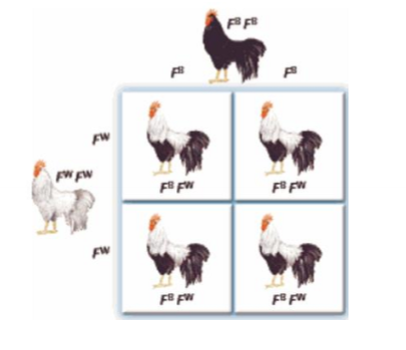
What is codominance and an example?
the expression of both alleles in a heterozygote
ex. allele for white and allele for black, express both black and white (not grey)

What is incomplete dominance and an example?
heterozygote has a unique phenotype compared to both types of homozygotes
heterozygote has an intermediate phenotype
ex. AA red father, aa white mother: Aa pink child

What is polygenic inheritance?
when a single trait is controlled 2+ genes, each contributng a small and equal portion to the overall phenotype
if 2 independently assorting genes contribute to a phenotype, ratios in 16ths when 2 heterozygote parents are crossed
produces a wide, continuous range of traits
ex. height, skin pigmentation
What are multiple dominance relationships? What are the types?
dominance rules that arent just dom/recessive
complete dominance - one allele masks the other (mendellian genetics)
incomplete dominance - neitehr allele fully dominates, heterozygote is a blend of 2 alleles ex. A and B antigens in blood
codominance - expression of both alleles in a heterozygote
multiple dominance
How do you solve problems with incomplete dominance or codominance?
identify heterozygote phenotype and see if an allele fully dominates (complete), if heterozygote is a blend (incomplete) or if both alleles are expressed (codominances)
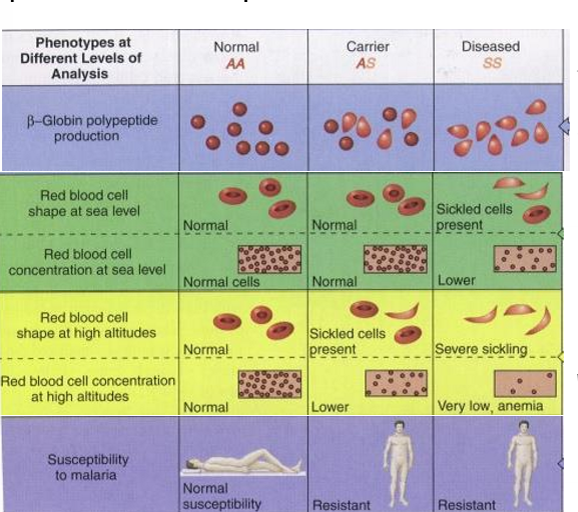
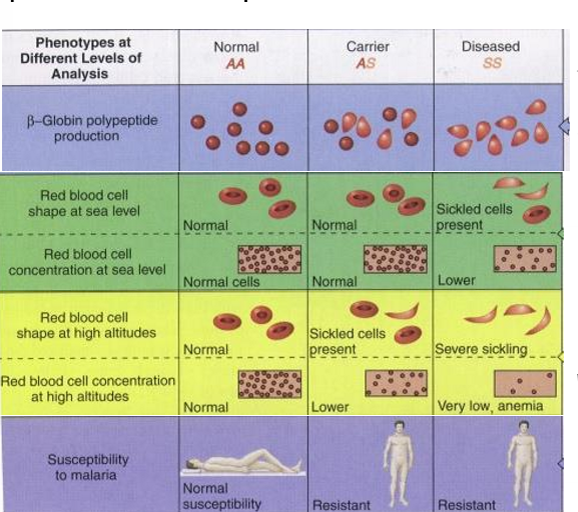
How does sickle-cell anemia illustrate dominance?
mixed dominance depending on level you’re looking at
What is pleiotropy?
single gene has 2+ unrelated phenotypes
ex. death or short tail
What is penetrance?
describes if a genotype shows up as a phenotype
measures percentage of individuals with given genotype who actually show the expected trait
ex. If a dominant allele has 80% penetrance, 20% of people who carry it show no signs of the condition whatsoever. They have the allele but look completely normal.
What is expressivity?
describes how severely a trait is expressed, from severe phenotype to mild phenotype
What is a temperature-sensitive (ts) mutation and what is its molecular mechanism? Why are ts mutations useful in reserach?
ts mutation produces a protein that folds normally at the normal temperature but misfolds at the restrictive temperature and doesn’t work
can switch phenotype on or off by changing expressivity
What are molecular causes of incomplete penetrance and variable expressivity, and why are they a challenge?
modifier genes, environmental factors
challenging because carriers may appear unaffected and make pedigree analysis difficult
What is complementation in genes? How do you know its occcuring?
where two mutations on different genes affecting the same phenotype restore a WT phenotype when present together
Each mutation knocks out a different gene, so when combined, each gene still has one functional copy (from the other parent/mutation set).
The organism regains normal function because the mutations “complement” each other.
If two mutants have WT offspring, the mutations complement and are on different genes
What are mutations in 2 different genes that produce different phenotypes of the same trait called? Why does this happen?
complementation
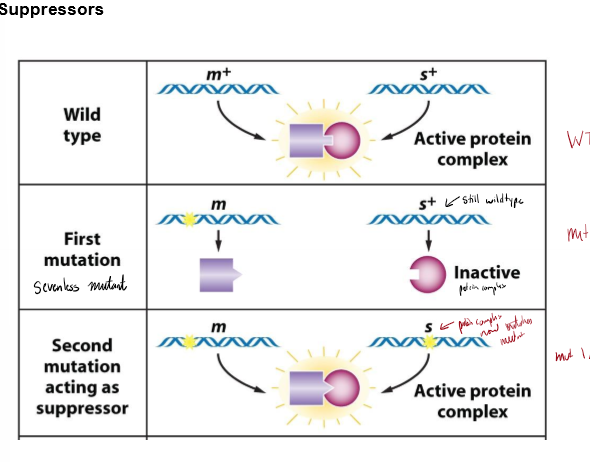
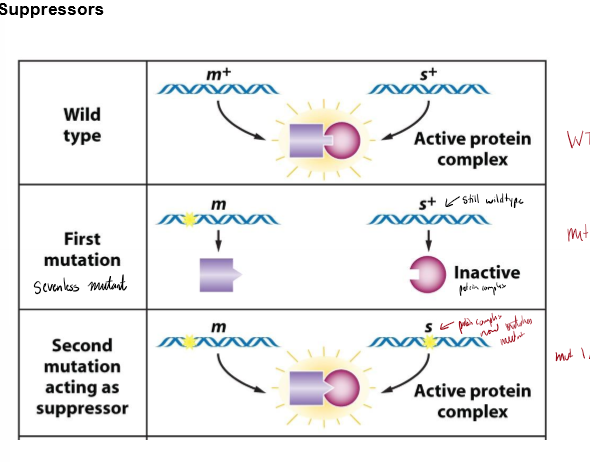
What is a suppressor? How can this work?
a mutant allele at one gene locus that reverses the effect of a mutation at a different gene locus, restoring WT
if one gene is mutated, protein won’t activate. if both genes are mutated, proteins fit and can activate again

What is epistatis? Give an example
alleles at one gene locus covering up alleles at a different locus
Ex. E gene controls deposition of pigment, and B gene controls whether dog is brown or black.
If ee, no pigment can be deposited so dog is yellow, regardless of whats at B locus.
If E-, B can present as black or brown

What is recessive and dominant epistasis, including their phenotypic ratios?
recessive - homozygous recessive genotype covers up the other gene - 9:3:4
dominant - dominant genotype covers up the other gene - 12:3:1
Describe the following gene interactions, including the number of genes involved, number of traits controlled, and F2 phenotypic ratieos
mendelian genetics
codominance
incomplete dominance
lethal
dihybrid
complementation
epistasis
mendelian genetics - 1 gene, 1 trait, 3:1
codominance - 1 gene, 1 trait, 1:2:1 (black: black and white: white)
incomplete dominance - 1 gene, 1 trait, 1:2:1 (black: grey: white)
lethal - 1 gene, 1 trait, 2:1 ( 1 died)
dihybrid - 2 genes, 2 traits, 9:3:3:1 (ex. yellow and wrinkled)
complementation - 2+ genes, 1 trait
epistasis - 2 genes, 1 trait - recessive: 9:3:4; dominant: 12:3:1
Describe phosphorylation as a post-translational modification
addition of phosphate group through a kinase to R group of amino acid
kinase adds phosphate and phosphatase removes phosphate
Which way are mRNAs read?
5’ to 3’
Describe codons
3 letters
read 5’ to 3’
1 codon always encodes 1 specific amino acid (unambiguous, degenerate)
1 amino acid can be encoded by multiple codons
What is a silent mutation? How is redundancy related to these?
nucleotide change that doesn’t affect the amino acid produced
redudancy in codon decreases likelihood of a mutation affecting amino acid sequence negatively
What is a frameshift mutation?
insertion or deletion of NTs not in multiples of 3
shifts reading frame of amino acids
can be a non-sense mutation (introduces stop codon early)
What is a non-sense mutation?
mutation that converts a codon to a stop codon
causes premature termination of translation
can be from a frameshift mutation
What is a missense mutation?
mutation that converts amino acid to a different one
can be neutral, harmful, or beneficial
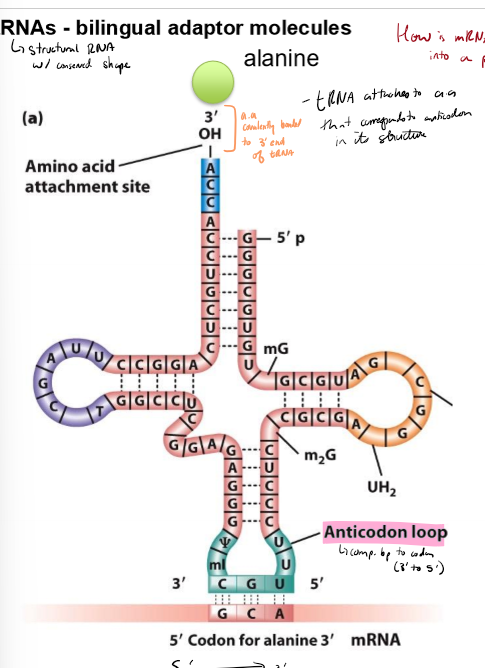
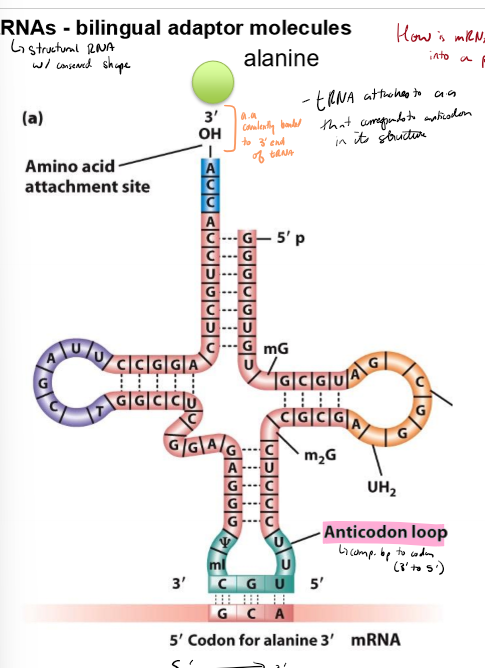
Describe tRNAs
structural RNAs with conserved shape that translate mRNAs into protons
have amino acid covalently bonded to 3’ end
have anticodon loops that complementary basepairs to codon (3’ to 5;)
What is wobble pairing?
property of codon and anticodon
last anticodon (5’ end) known as wobble position
wobble position doesn’t need to perfectly basepair to still get correct amino acid since multiple codons can specifiy the same amino acid (degenerate code)
Are there more codons or different tRNAs? why?
way more codons that tRNAs
because of wobble pairing, multiple codons can encode for same amino acid
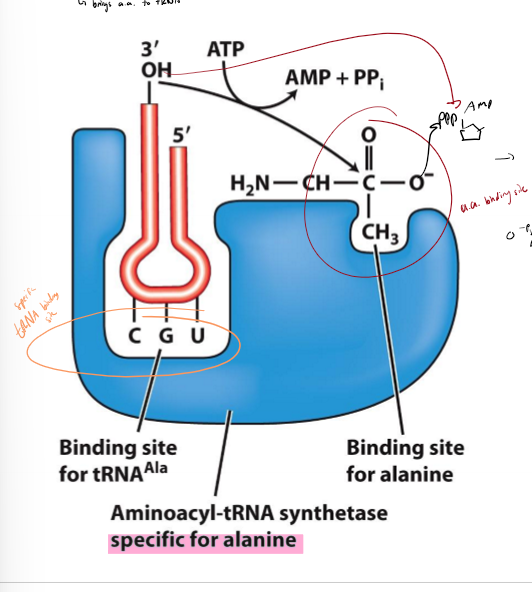
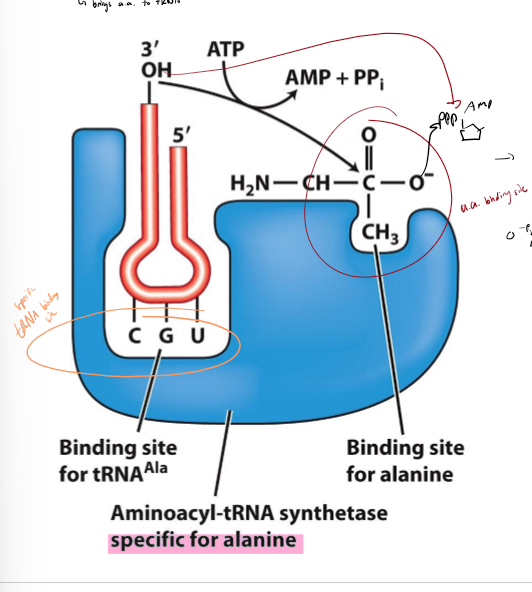
What is aminoacyl-tRNA synthetase?
brings and loads loads correct amino acid on to tRNA
1 type of aminoacyl tRNA per amino acid to increase specificity and ensure tRNAs get correct amino acid
has binding site for amino acid and binding site for tRNA
1. loads tRNA and amin oacids into binding site
2. activation - amino acid carboxylate group (3’ end) attacks phosphate of amino acid with energy from ATP, creating AMP as a byproduct
3. transfer - amino acid transferred to 3’ end of tRNA, creating charged/loaded tRNA
What happens to tRNA after its finished with translation?
stable, so reused to translate again
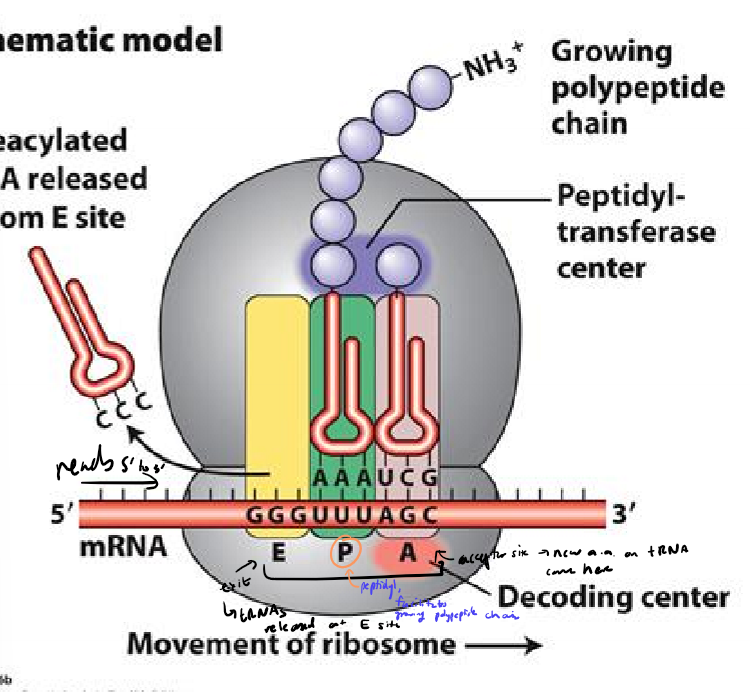
Describe the general structure of ribosomes
large and small sub unit
built around mRNA, with mRNa read 5’ to 3’
tRNA enters at A site, forms peptide bond at P site, and exits at E site
amino acid chain grows out with N terminus out
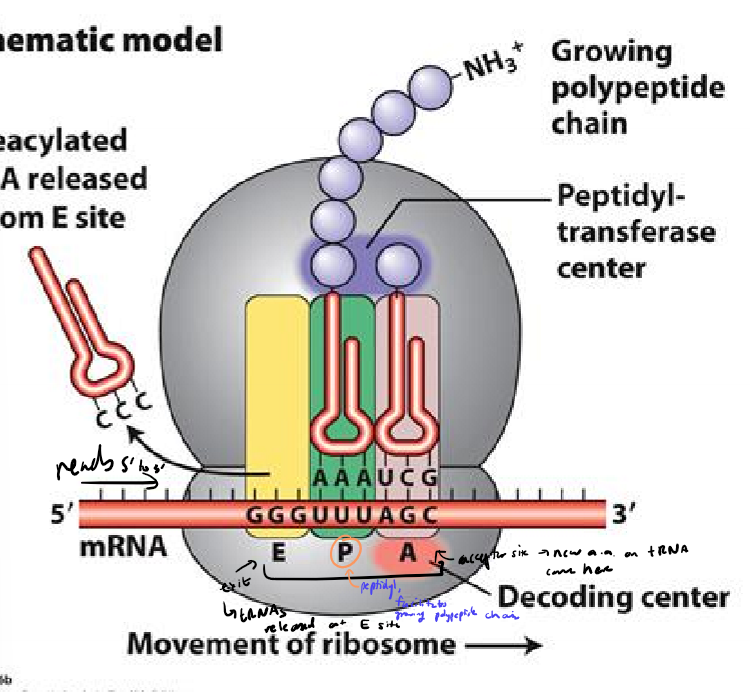
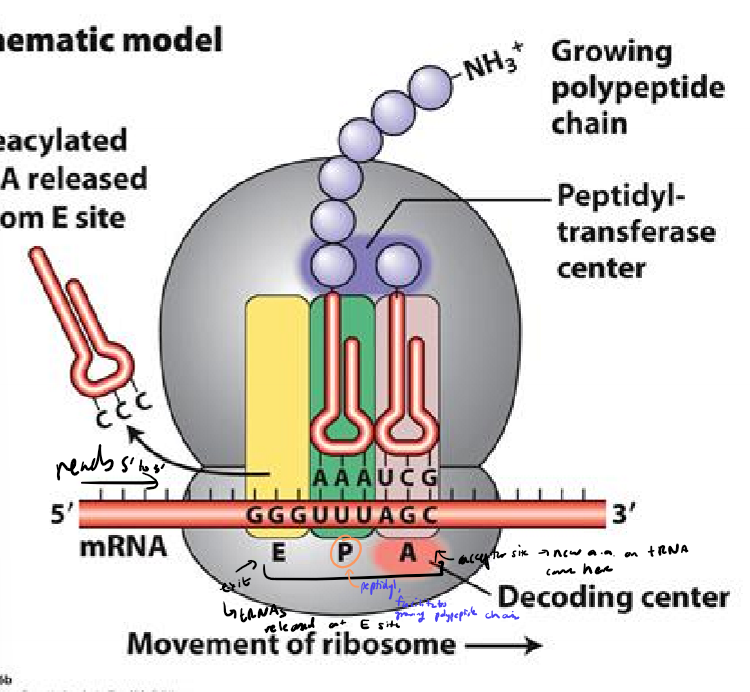
What are ribosomes made of?
large and small subunit
proteins and rRNAS
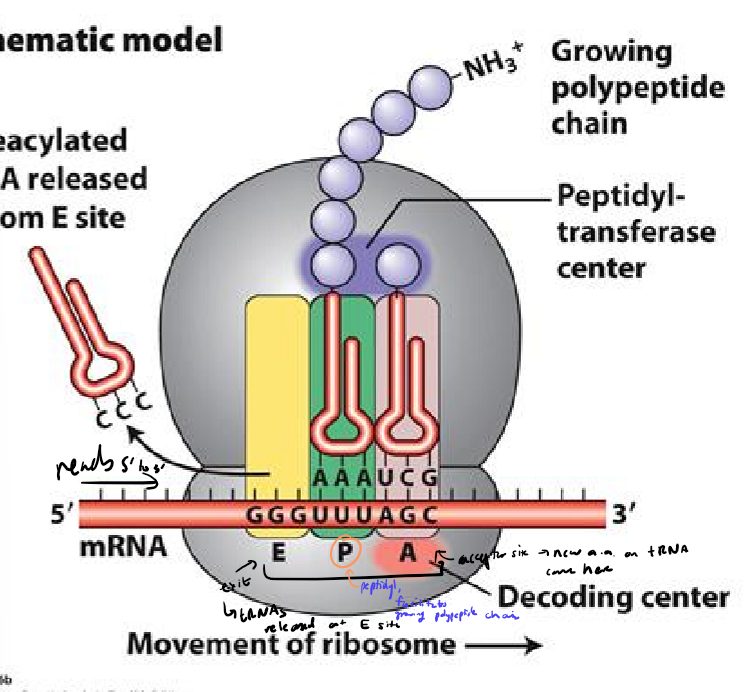
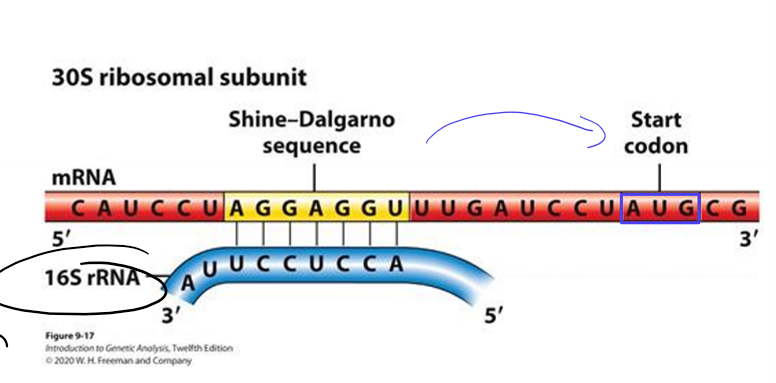
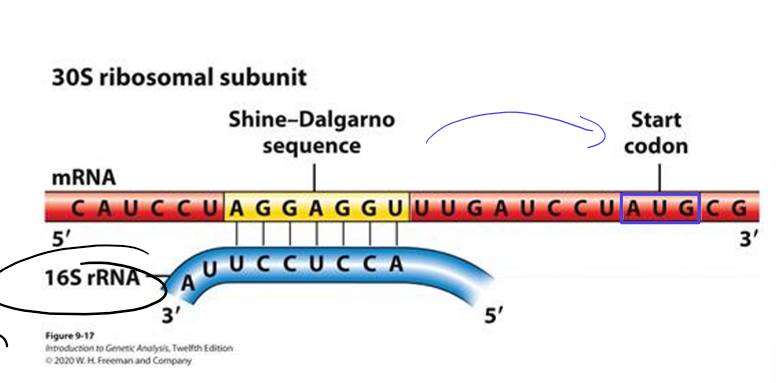
In bacteria, how do ribosomes know which AUG is the start codon?
using Shine-Delgarno sequence on mRNA to align ribosome small subunit upstream of translation start codon
aligns first AUG with P site
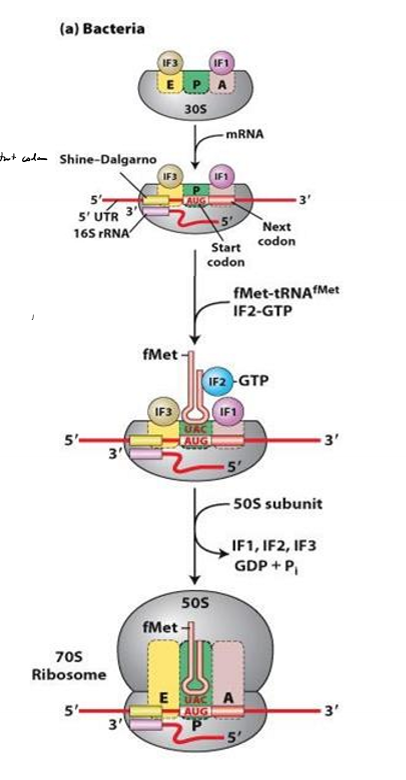
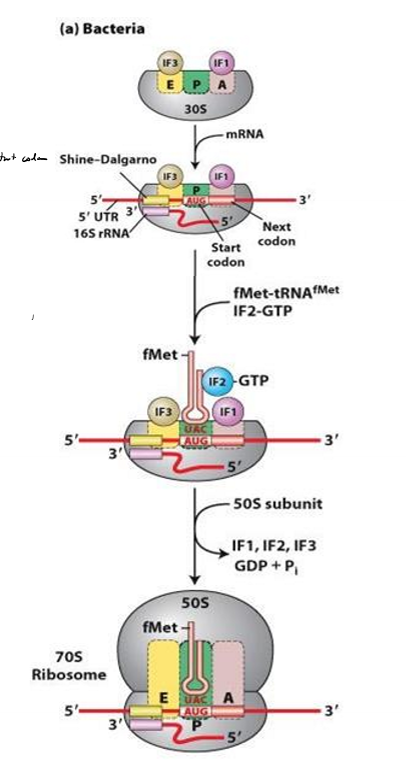
Describe translation initiation in bacteria
1. ribosome small sub units positions at shine-dalgarno sequence, with P site at start codon
2. IF1 and If3 block E and A site to prevent large subunit from binding
3. IF2 brings fMet tRNA to P site
4. GTP on IF2 hydrolyzed
5. IFs kicked off
6. large subunit binds and elongation can begin
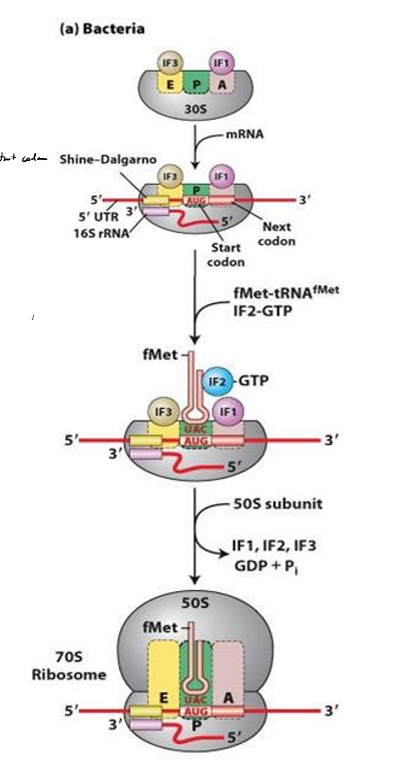
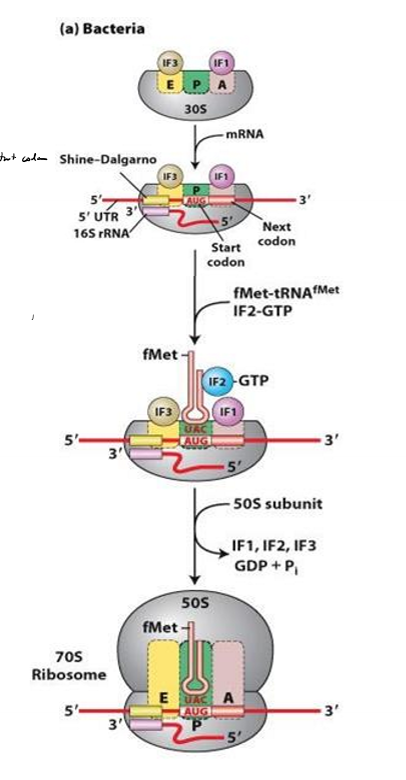
What do the initiation factors do in bacteria?
block E and A site so large subunit can’t bind early
IF2 brings first tRNA to P site and hydrolyzes it to kick off IFs
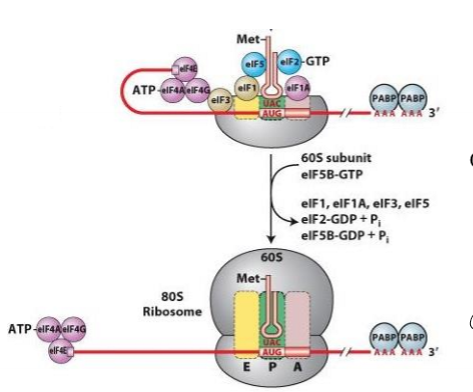
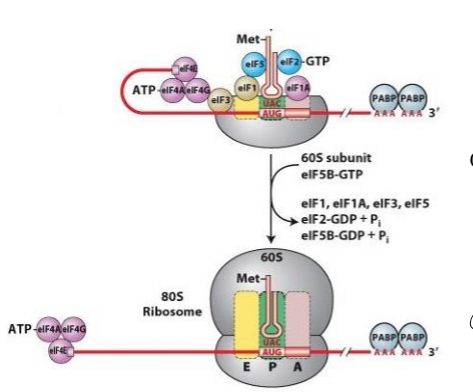
Describe translation initiation in eukaryotes
1. preassembly of eIFs and 40s small ribosomal subunit before interating with mRNA, forming ternary complex with small subunit, fMet tRNA at P site , and eif-tu-GTP
2. eIFs block A and E site
3. eIF-4f binds to 5’ cap of transcript
4. eif-4F binds to ternary complex and holds complex to cap at 5’ UTR
5. anticodon of fMet tRNA and start codon of mRNa basepair, and eIF-tu hydrolyzes GTP, releasing eIFs
6. 60s large subunit joins and fMet tRNA positioned at P site
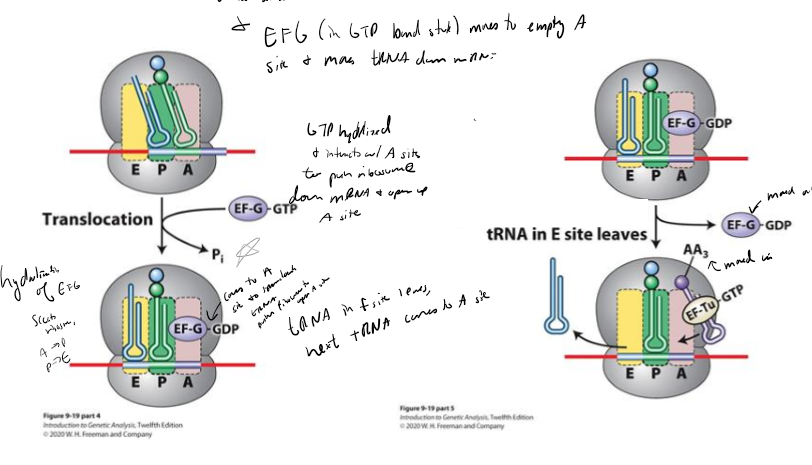
Describe translation elongation
1. charged tRNA deliver to A site by Ef-tu-GTP
2. anticodon of tRNA baseparirs with mRNA codon, and GTP hydrolyzed, releasing EF
3. peptide bond formed between amino acids by rRNA, growing polypeptide chain transferred from tRNA in the P site to the amino acid on the tRNA in the A site
4. ribosome moves down mRNA one codon using EFG hydrolysis, putting tRNA from A→ P and P → E, with E tRNA leaving
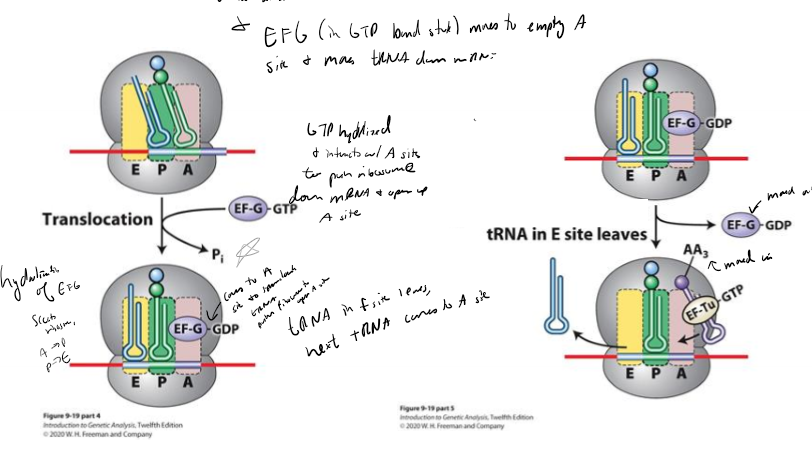
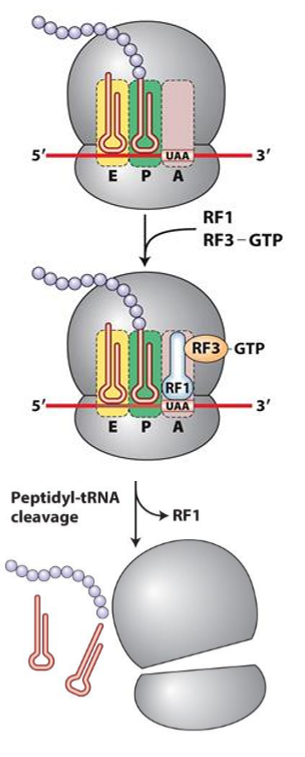
Describe translation termination
1. A site of ribosome interacts with stop codon
2. RF 1 or RF2 binds at A site, mimicing shape of tRNA
3. RF3 hydrolyzes GTp to cleave peptide chain from P site, releasing chain
4. hydrolysis destabilizes ribosomal subunit, disassembling it
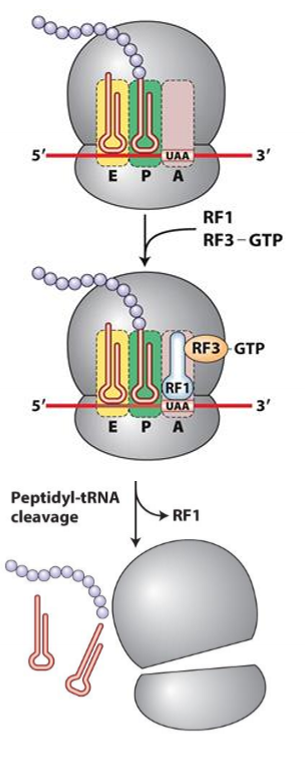
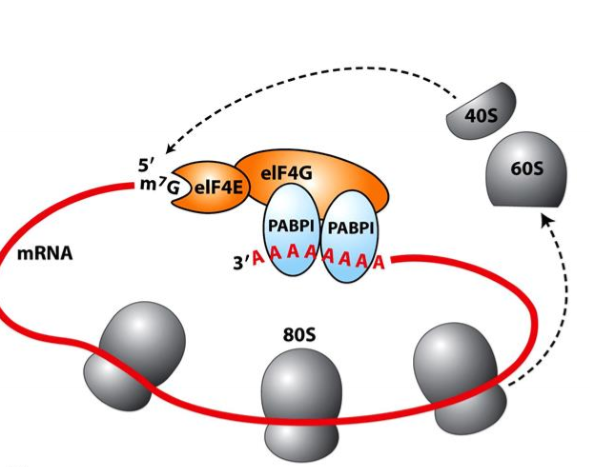
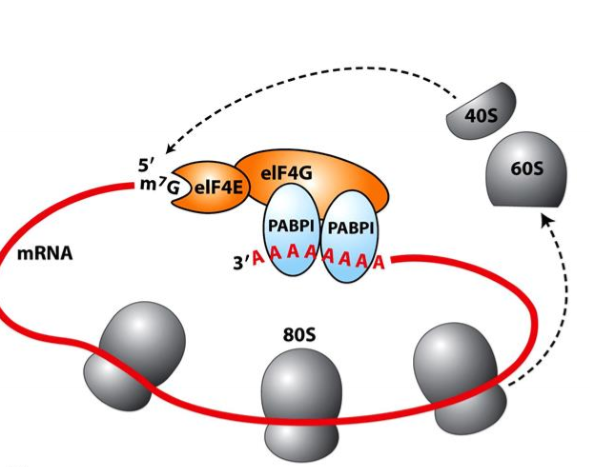
How do eukaryotic mRNAs enhance translation efficiency?
circularization
cap binding proteins interact with PolyA binding proteins (PABP), forming a circle, so ribosomes that have disassembled can immediately erassable at start site of translation
multiple ribosomes can translate 1 mRNA at a time
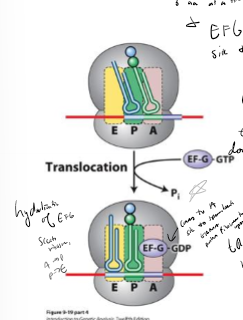
In bacteria, what facilitates translocation of the ribosome and how?
Ef-G by hydrolyzing GTP
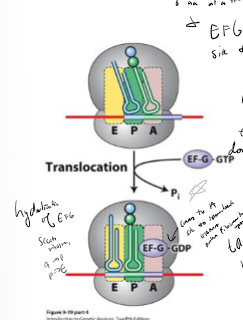
What moves new tRNA into the A site in translation elongation?
EF-Tu
What hydrolyzes GTP to cleave peptide chains from the P site in translation termination?
RF 3
What causes peptide bond formation between amino acids during translation elongation?
peptidal transferase activity from rRNA
What releases Ef-Tu from the ribosome after it brings in tRNA?
hydrolysis of its GTP
What binds to the 5’ cap of transcript and ternary complex during eukaryotic translation initation?
eIF-4F
Where does translation occur in prokaryotes and eukaryotes?
prokaryotes - cytoplasm, coupled to transcription
eukaryotes - cytoplasm, has to be brought out of nucleus first
Where does mRNA binding occur during translation initiation in prokaryotes and eukaryotes?
prokaryotes - Shine-Dalgarno sequence upstream of start codon
eukaryotes - 5’ cap binding
What IFs are involved in translation initiation in prokaryotes and eukaryotes?
prokaryotes - IFs 1, 2, and 3
eukaryotes - 9+ eIFs
What are the translation ribosome subunits in prokaryotes and eukaryotes?
prokaryotes - 70s, 30s + 50s
eukaryotes - 80s, 40s + 60s
What is transcription?
creation of RNA from DNA template
What causes a gene to be expressed?
DNA info is transcribed and translated so encoded function is present in cell
each cell types expresses different genes at different times and locations under different condition
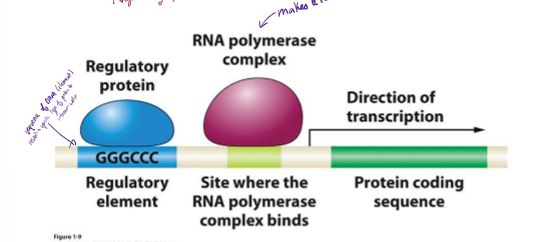
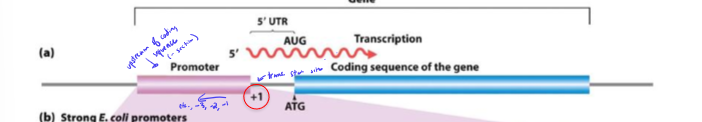
Where do regulatory and promoter regions sit relative to the beginning of transcription?
upstream
negative numbering
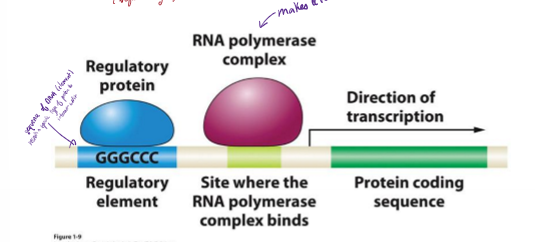
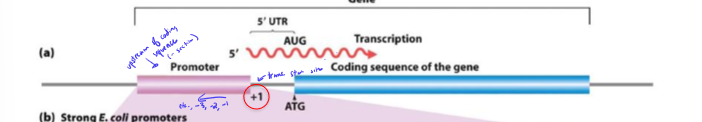
In the transcription number system, where does transcription start? where is the promoter?
+1
promoter is immediately upstream (-1)
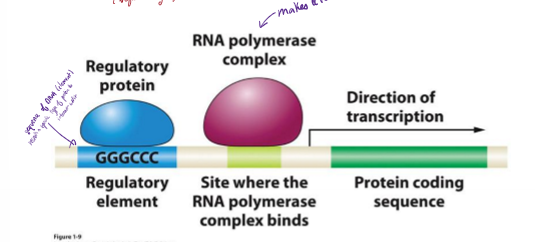
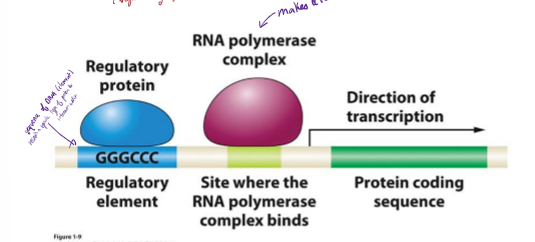
What is a regulatory element?
sequence of DNA (element) that recruits a specific type of protein to interact with it and tells whether or not a protein should be expressed
G and C rich
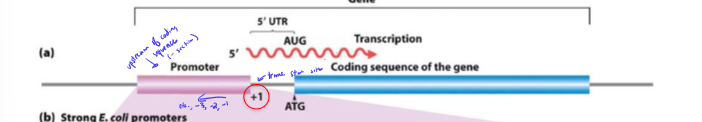
What is a cis-element?
promoter sequence nearby a gene it regulates
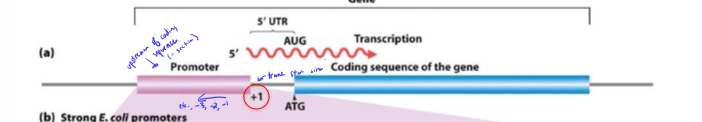
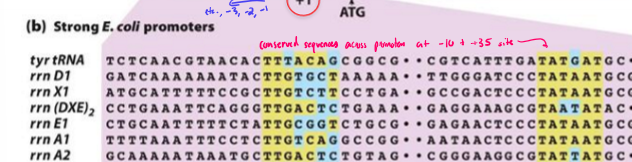
What is the consensus sequence?
most common nucleotide at each position of promoter
conserved sequence across promoters from -10 to -35 site of transcription
+1 = transcritpion start site
the more similar a promoter sequence is to the consensus sequence, the stronger binding affinity RNA poly has to the promoter, increasing transcription
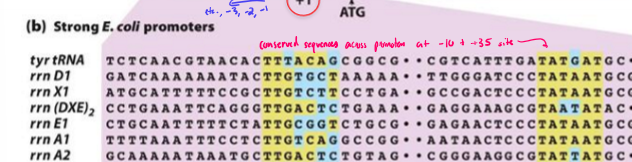
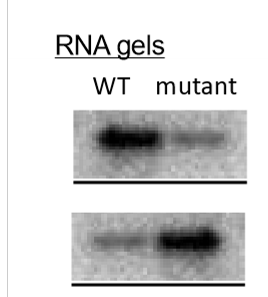
How does the promoter sequence of a gene alter gene expression?
the more similar a sequence is to the consensus, the better binding affinity RNA has and the more transcription is done
the less similar a sequence is to the consensus, the worse binding affinity RNA has and less transcription is done
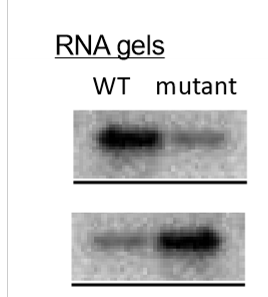
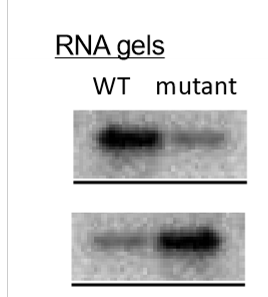
what promoter mutations can cause gene LOF? what can cause gene GOF?
LOF - make promoter sequence more different from consensus, deletion of consensus sequence
GOF - change promoter sequence to better match consensus
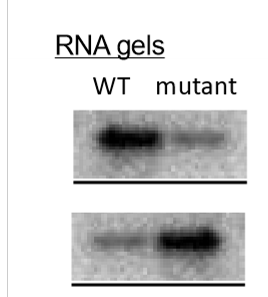
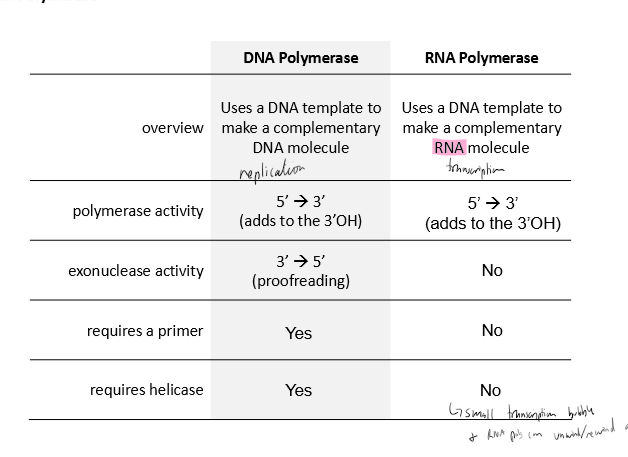
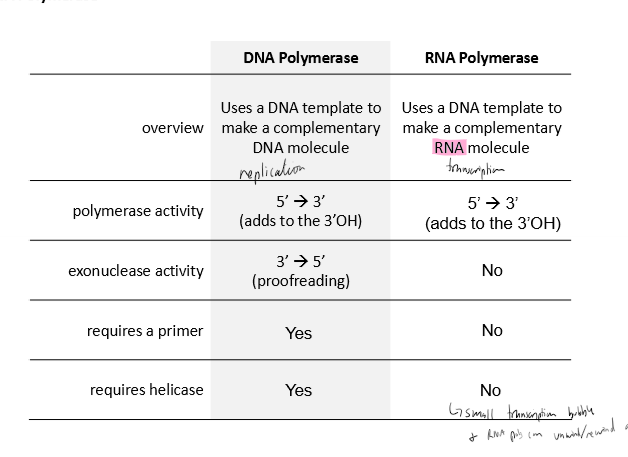
Describe RNA polymerase’s
polymerase activity
exonuclease activity
requires a primer
requires helicase
copying
5’ to 3’ polymerization
no proofreading (3’ to 5’)
does not require a primer
does not require helicase; can unwind/rewing itself
only 1 part of strand copied, thousands or more copies made a time
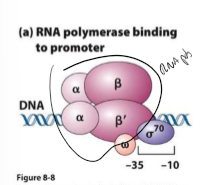
What is a holoenzyme?
RNA polymerase core enzyme
can synthesize RNA but needs sigma factors to recognize promoters and start transcription
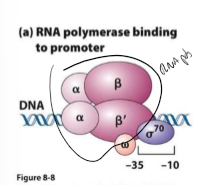
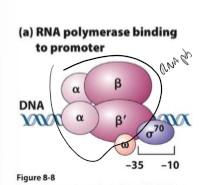
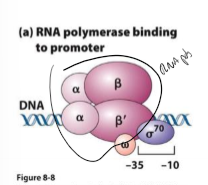
What is sigma factor?
binds to one strand of DNA at consensus sequence (-10 to -35 sites) to help RNA polymerase bind and copy
induces bend in DNA at AT ri h region to start destabilizing DNA
dissociates from DNA once RNA polymerase holoenzyme binds
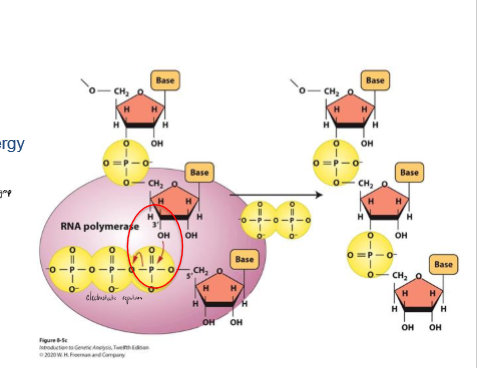
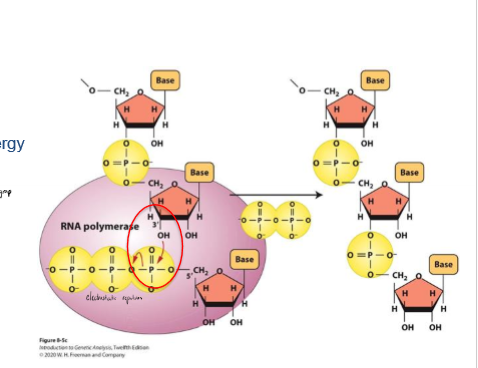
What provides the energy for the RNA polymerization reaction?
high energy phosphate bond (3 Pi together) on nucleotides from electrostatic repulsion
phosphate bond broken as new nucleotide is added at 3’, providing energy for the reaction
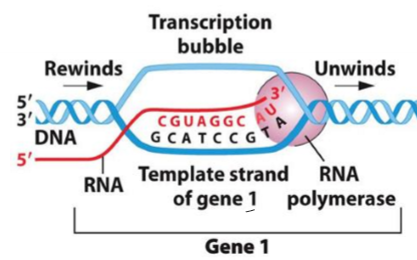
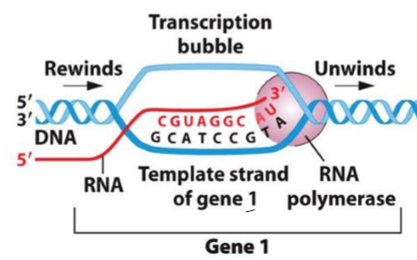
What are the template and coding strand in transcription?
template strand - side of DNA RNA Is copying from
coding strand - non-template strand, matches new RNA molecule made from template strand
either top or bottom strand of DNA can be the template strand; depends on where promoter sequence is
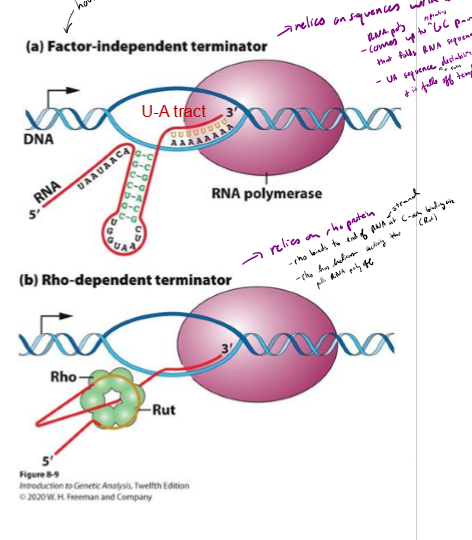
How can transcription be terminated in bacteria? When is each method used?
Intrinsic termination (Factor-Independent termination) - RNA secondary structure slows down RNA polymerase; usually house-keeping genes
rho-dependent terminatino - RNA binding protein slows down RNA polymerase
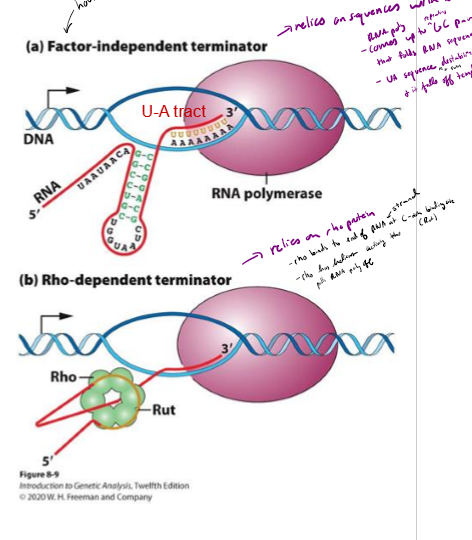
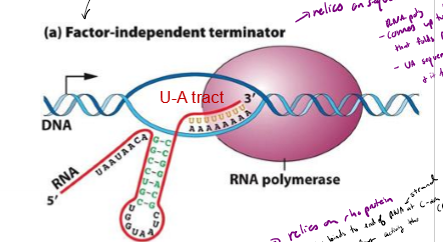
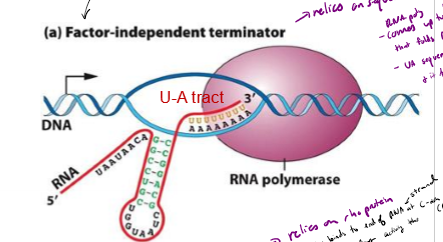
Describe factor-independent termination of transcription in bacteria
releies on sequence of nucleotides within DNA Itself
- RNA poly comes up to GC rich region on end of gene, followed by a long stretch of As
GC rich strand createdbinds to each other and folds up on itself
UA rich region and fold destabilizies RNA poly and it falls off the template
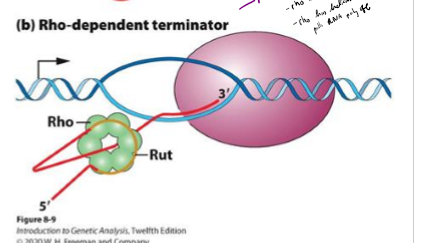
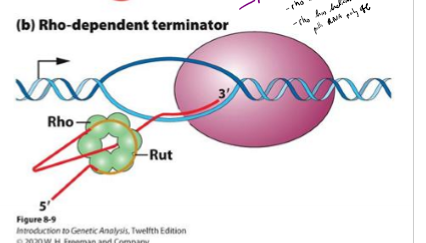
Describe rho-dependent termination of transcription in bacteria
relies on rho binding protein to slow down RNA polymerase
rho binds to end of RNA strand at C- rich region called rut
rho has helicase activity that pulls RNA poly off strand
Describe mRNA decay in bacteria. Why does it happen?
mRNA created decays/degrades after making proteins
used to regulate gene expression because mRNA only needs to exist long enough to make proteins while the cell needs them
RNA strand can be cute in the middle (endonuclease) or end (exonuclease)
What are the RNA polymerases in eukaryotes and what does each do?
RNA poly 1 - makes rRNA
RNA poly 2 - makes mRNA and helps make snRNA, and non-coding RNAs; done in nuceloplasm
RNA poly 3 - makes tRNA
What is the Preinitation Complex?
made of general transcription factors with different roles, such as binding to TATA box or recruiting later proteins, and eventually RNA poly 2
recognizes promoter sequences for RNA poly 2
recruits and helps bind RNA poly 2 to DNA
not gene specific
bidns to all promoters at consesus sequence
is phosphorylaed by TF2H at the Carboxy-terminal domain of RNA poly2 to being elongation, releasing RNA poly 2
Describe the intiation of transcripition in eukaryotes
TF2D binds to TATA box and bends DNA, destabilizing it for transcription
TF2D recruits other GTFs (general transcription factors), forming the Preiniitation complex (PIC)
GTF recruits RNA poly 2, helping stabilizing it
TF2H phosphorylated RNA poly 2, allowing Poly 2 to break free and being elongating mRNA
Describe transcription termination in eukaryotes, including the 2 models described
RNA Poly 2 transcribes past the termination site
The CP (cleavage) complex recognizes a sequence in the 3’ UTR (untranslated region) of the mRNA and cuts it
RNA is cut and a polyA tail is added to the 3’ end
Torpedo model - Exonuclease protein digests RNA until it reaches RNA poly 2, dissociating RNA poly 2
Allosteric model - CP binding modifies RNA poly 2’s configuration, causing mRNA and poly 2 to dissociate
downstream mRNA is destroyed
Compare prokaryotic and eukaryotic promoters in terms of sequences and proteins binding:
prokaryotic - -10 and -35 boxes for promoter; consensus sequence; needs sigma factor to find gene, bend DNA, and recruit RNA poly
eukaryotic - downstream TATA box, binded by TF2D on PIC recognize promoter sequences and PIC recruits and positions RNA poly 2
What are the functions of the GTFs in the Preinitation complex?
promoter recognition and DNA bending - TF2D binds to TATA box and bends DNA, destabilizing it for transcription
Recruiting other enzymes - TF2D recruits other GTFs that can recruit even more enzymes
Recruiting RNA pol 2 - GTF brings RNA poly 2 and positions it on DNA
Phosphorylation - TF2H phosphorylates the CTD of RNA pol2 to allow it to release from the complex and elongate
What is the RNA poly 2 Carboxy-Terminal Domain? (CTD) Why is it important?
on end of RNA pol 2 where TF2H phosphorylates it to release RNA pol 2 from the PIC and allow it to elongate mRNA
What is the torpedo model of transcription termination?
polyA signal is transcribed
CP complex cuts mRNA
exonuclease enzyme digest RNA until it reaches RNA pol2, degrading mRNA faster than RNA pol 2 can syntehsize it
RNA pol 2 dissociates
What is the allosteric termination model of transcription termination?
proposes that termination is caused by a conformational change in RNA Pol 2
pol 2 transcribes poly A signal
CP complex cuts mRNA
Cp binding alters pol 2 shape, causing it to dissociate and fall off
What post-transcriptional modificatins to mRNA can happen in eukaryotes?
all = 5’ cap, 3’ poly A tail, splicing out of introns
some = editing of bases
What is the TATA box and what binds it?
eukaryoric promoter element with consensus TATAAA
bound by TF2D
What is the key difference between prokaryotic and eukaryotic transcription termination?
prokaryotes use secondary structure (factor-independent) or rho-depdendent termination
eukaryotes cleave at the polyA site, and RNA poly either changes shape to dissociate or is knocked off by and enzyme
What differs between pre-mRNA and mRNA
pre-mRNA - introns and exons, no 5’ cap or 3’ Poly A Tail
mRNA - introns removed, 5’ cap and 3’ Poly A tail
Why is the 5’ cap added to mRNA in eukaryotes?
protective -signals to cell that the mRNA is human and stabilizes mRNA
aids in export out of nucleus
ensures initiation of translation
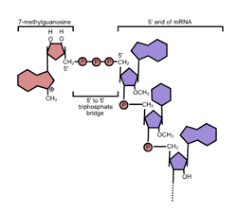
How is the 5’ cap added to mRNA?
1. PIC removes phospharte from first nucleotide
2. capping enzyme associated with CTD adds guanine monophosphate to open spot
3. methyl group added to end of 5’ cap
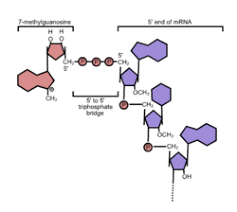
Why is the 5’ capping enzyme associated with CTD (C-Terminal Domain of RNA poly 2)?
so that they are positioned to the 5’ cap right as transcripion begins to stabilize the mRNA
How is the Poly A tail added to the 3’ end of mRNA?
after enzyme cleaved
5. poly A polymerase enzyme add A’s to end of sequence without a template
6. poly A binding protein binds to As added to increase mRNA stability and increase translatability of mRNA
What is the PolyA tail?
stretch of A nucleotides added to the 3’ end of mRNA after transcription
What is the Poly A signal? What is the poly A site?
Poly A signal - consensus UTR region of mRNA where enzymes bind to poisition CF enzyme at polyA site and add poly A tail to 3’ end of RNA
Poly A site - specific nucleotide position where cleavage of mRNA occurs
What order does post transcriptional modification occur in?
5’ cap
poly A tail
splicing
What is alternative splicing?
same pre-mRNA can be spliced in different ways to produce multiple distinct mRNA and therefore multiple different proteins, from a single gene
cuts out introns and pieces together exons (in order)
On a genome browser, what is the narrow portion at the end of the exons?
untranslated region that gets transcribed but NOT translated
Where are UTR’s found in genome browsers of RNA?
exons
What are the benefits of alternative splicing?
1 gene can encode for different but related genes (like binding to some of the same ligands but not all)
distinct phenotypes between sexes
How does mRNA splicing work? What pieces are involved?
cis-elements in the pre-mRNA introns help regulate a gene within the mRNA
each intron conatins invariable nucleotides: GU at 5’ splice site, AG at 3’ splice site, and A at the branch site where the mRNA folds up
5' splice site - cut between beginning of exon and UTR
3’ splice site - cut between end of exon and UTR
Branch point - A nucleotide in between 5’ss and 3’ss
Describe the process of mRNA splicing
1. U1 (snRNP) binds to 5’ splice site via base pairing
2. U2 (snRNP) binds to branch point sequence via base-pairing, except for the A nucleotide, which then bulges out the A and its free 2’ OH
3. splisome folds up intron to enable cleave and removal of intron sequence
4. free 2’OH of A at branchpoint attacks phosphate group at 5’ exon, cleaving exon 1 off
5. exon 1 joing its now open 3’OH with ’ phosphate group of nucleotide at beginning of next exon at 3’ss
6. intron released, exons joined
How does alternative splicing affect DNA and mRNA?
DNA - not at all, alternative splicing happens to pre-mRNA
RNA - go different distances on RNA gel, extra exon doesn’t travel as far