BIOS 301: L5 Polysaccharides + Glycolysis Prepetory step
1/27
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
28 Terms
Preparatory phase of glycolysis – overall purpose
Spending energy, not producing ATP
Prepares glucose for cleavage into two 3-carbon molecules
Steps 1–5 (glucose → glyceraldehyde-3-phosphate)
Step 1 – Phosphorylation of glucose
Enzyme: Hexokinase (I–IV)
Reaction: Glucose + ATP → Glucose-6-phosphate (G6P) + ADP
Thermodynamics: ΔG°' = -16.7 kJ/mol → irreversible
Regulation:
Hexokinase I (muscle/brain) → high affinity, inhibited by G6P
Hexokinase IV/glucokinase (liver) → low affinity, regulated by [glucose]/[F6P]
Purpose: Traps glucose in cell and primes it for glycolysis
![<ul><li><p><strong>Enzyme:</strong> Hexokinase (I–IV)</p></li><li><p><strong>Reaction:</strong> Glucose + ATP → <strong>Glucose-6-phosphate (G6P) + ADP</strong></p></li><li><p><strong>Thermodynamics:</strong> ΔG°' = -16.7 kJ/mol → irreversible</p></li><li><p><strong>Regulation:</strong></p><ul><li><p>Hexokinase I (muscle/brain) → high affinity, inhibited by G6P</p></li><li><p>Hexokinase IV/glucokinase (liver) → low affinity, regulated by [glucose]/[F6P]</p></li></ul></li><li><p><strong>Purpose:</strong> Traps glucose in cell and primes it for glycolysis</p></li></ul><p></p>](https://assets.knowt.com/user-attachments/13a3dba7-ed1f-4914-a590-6108f8e66b42.png)
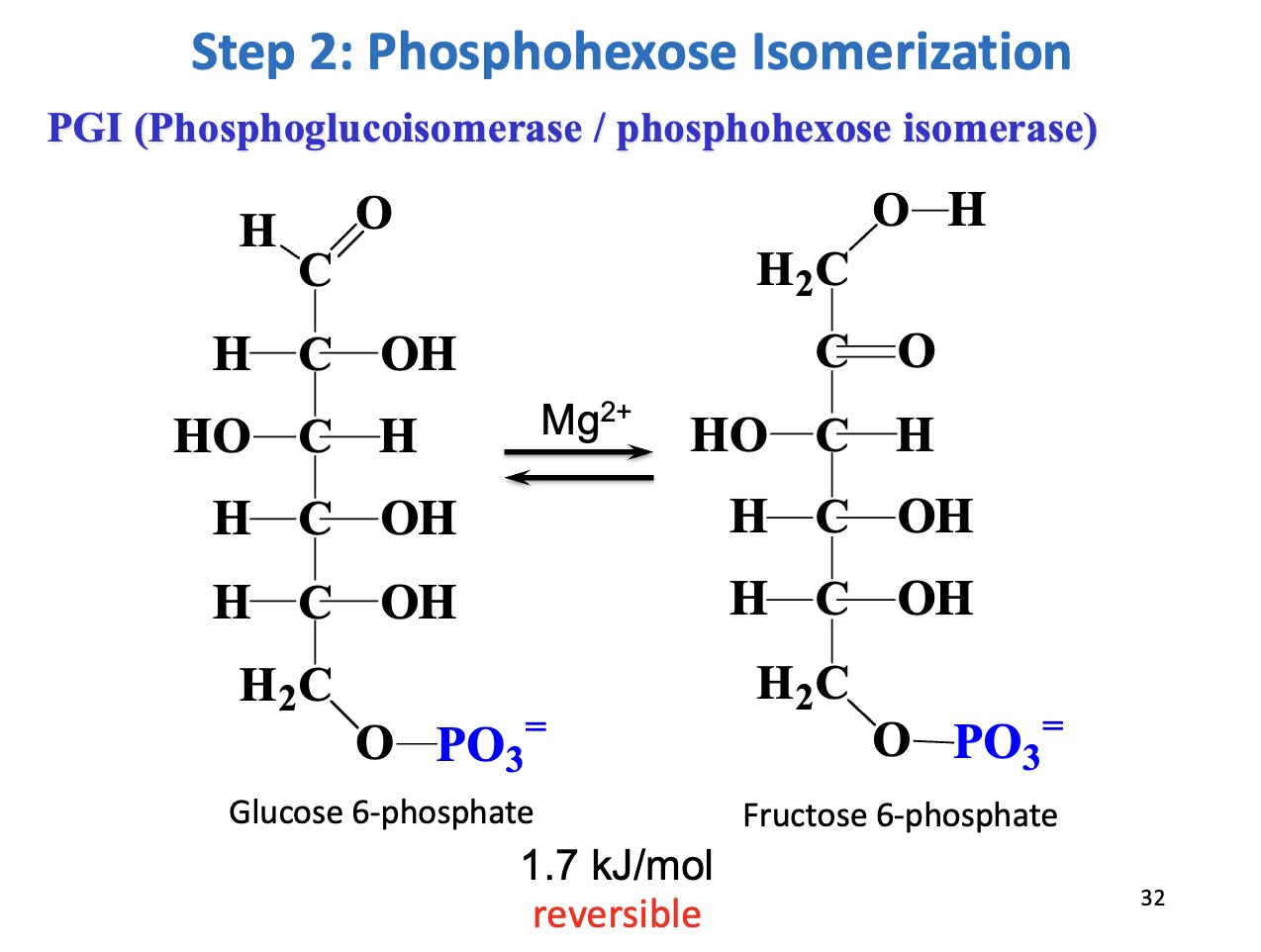
Step 2 – Phosphohexose isomerization
Enzyme: Phosphoglucoisomerase (PGI)
Reaction: G6P ↔ Fructose-6-phosphate (F6P)
Mechanism: Aldose ↔ ketose via enediol intermediate
Thermodynamics: ΔG = +1.7 kJ/mol → reversible
Purpose: C1 of F6P is phosphorylatable by PFK-1 & allows symmetrical cleavage by aldolase
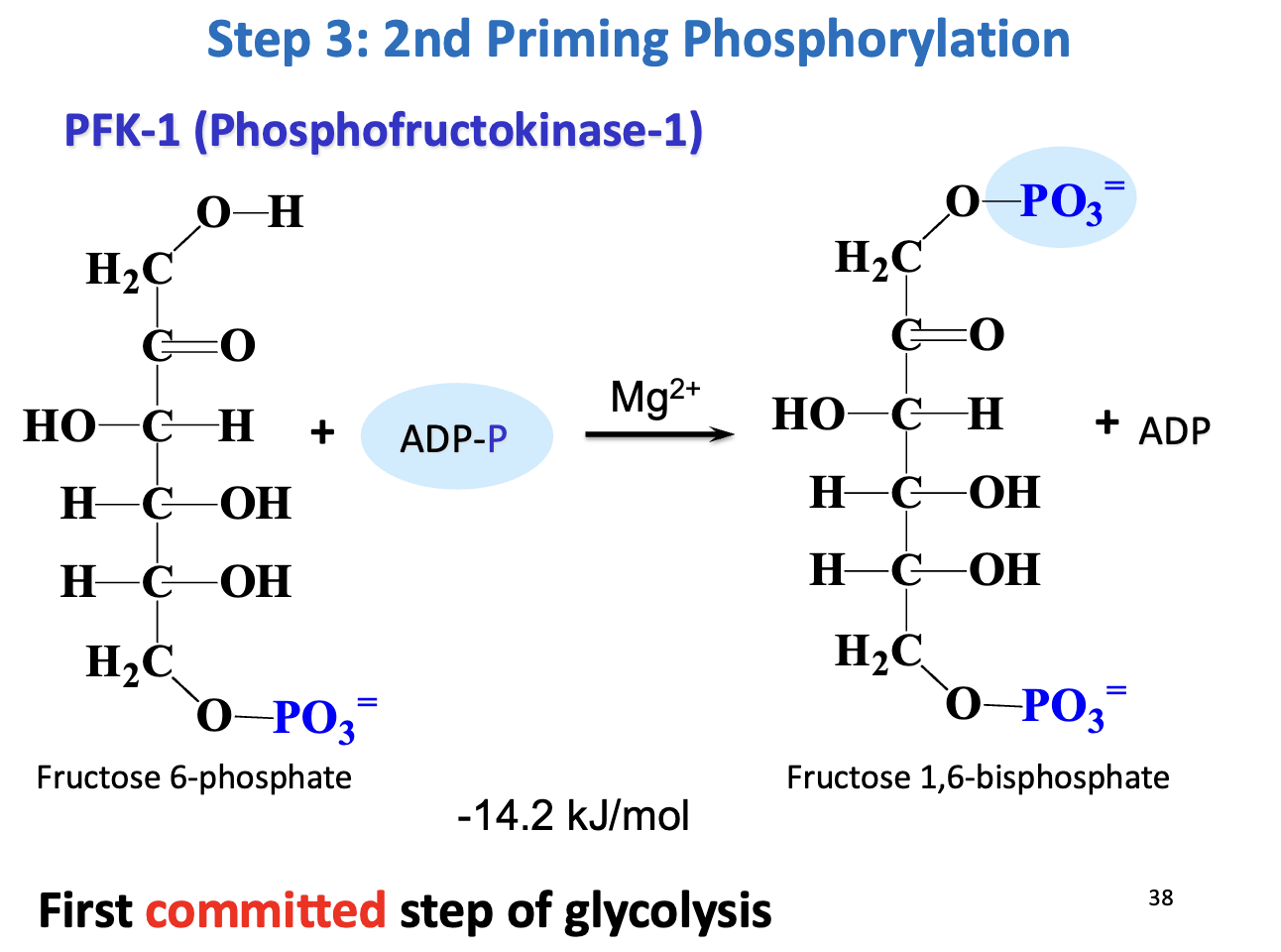
Step 3 – Second priming phosphorylation
Enzyme: Phosphofructokinase-1 (PFK-1)
Reaction: F6P + ATP → Fructose 1,6-bisphosphate (F1,6BP) + ADP
Thermodynamics: ΔG°' = -14.2 kJ/mol → irreversible
Significance: First committed step of glycolysis → commits glucose to energy production
Regulation: Activated by ADP, F2,6BP, inhibited by ATP
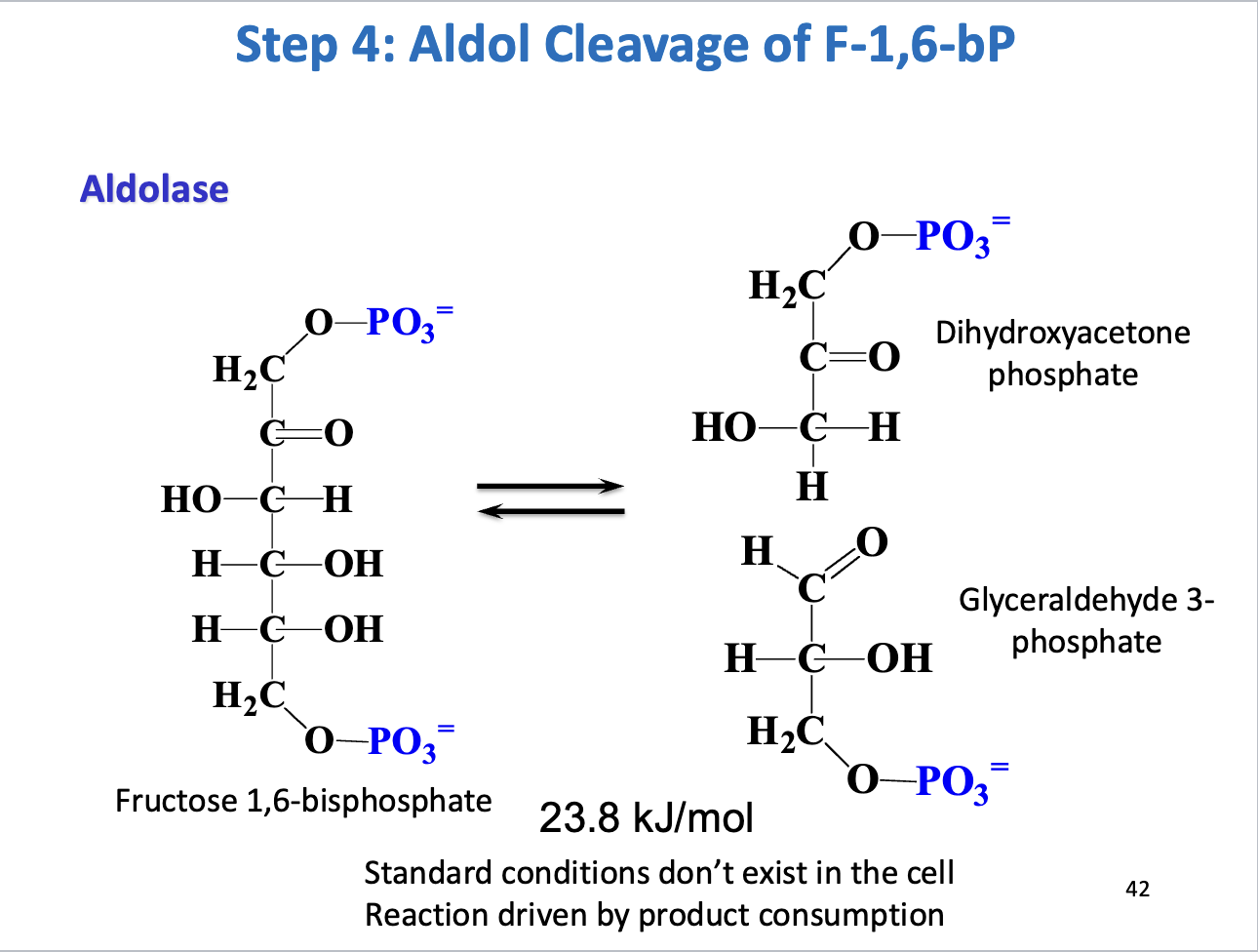
Step 4 – Aldol cleavage
Enzyme: Aldolase
Reaction: F1,6BP ↔ Dihydroxyacetone phosphate (DHAP) + Glyceraldehyde-3-phosphate (GAP)
Thermodynamics: ΔG°' = +23.8 kJ/mol → reversible, driven forward by product consumption
Purpose: Cleaves 6-carbon sugar into two 3-carbon molecules
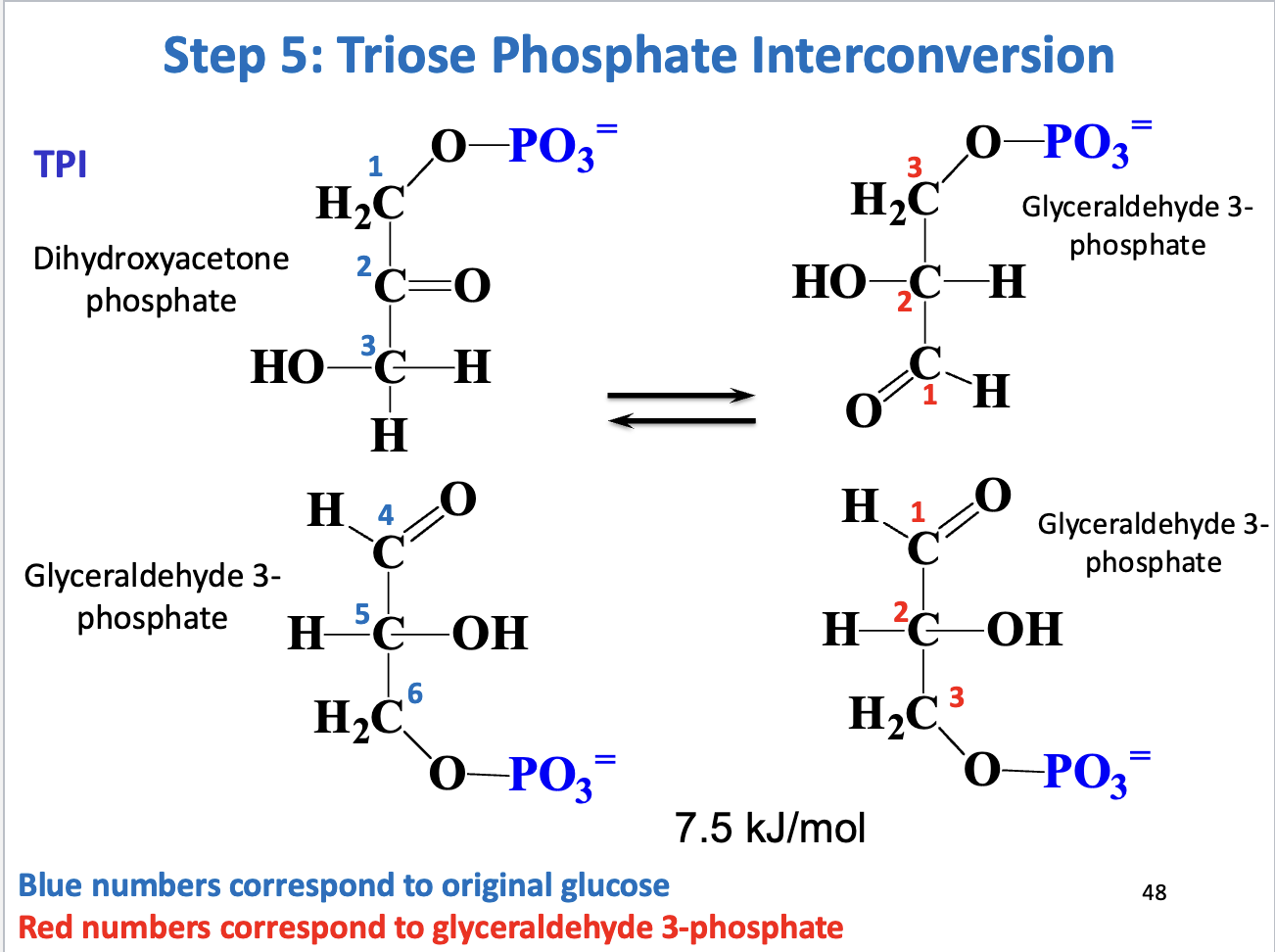
Step 5 – Triose phosphate interconversion
Enzyme: Triose phosphate isomerase (TPI)
Reaction: DHAP ↔ GAP
Significance: Only GAP continues in glycolysis
Thermodynamics: Reversible, driven forward by low GAP concentration
Purpose: Completes preparatory phase
Energy investment in preparatory phase
ATP used: 2 ATP per glucose
Step 1: Hexokinase → 1 ATP
Step 3: PFK-1 → 1 ATP
No ATP generated yet
Prepares glucose for payoff phase
Hexokinase I vs Hexokinase IV – affinity & regulation
Hexokinase I (muscle/brain)
Low Km (~0.1 mM) → high affinity, always near saturation
Inhibited by G6P (product feedback)
Hexokinase IV / Glucokinase (liver)
High Km (~10 mM) → low affinity, acts only at high blood glucose
Cooperative enzyme → activity increases sharply after meals
Graph interpretation:
X-axis: [glucose], Y-axis: enzyme activity
Hexokinase I curve is steep at low [glucose], plateaus quickly → saturates easily
Hexokinase IV curve is sigmoidal, rises slowly at low [glucose], accelerates at high [glucose] → acts as “glucose sensor”
![<ul><li><p><strong>Hexokinase I (muscle/brain)</strong></p><ul><li><p><strong>Low Km (~0.1 mM)</strong> → high affinity, <strong>always near saturation</strong></p></li><li><p><strong>Inhibited by G6P</strong> (product feedback)</p></li></ul></li><li><p><strong>Hexokinase IV / Glucokinase (liver)</strong></p><ul><li><p><strong>High Km (~10 mM)</strong> → low affinity, <strong>acts only at high blood glucose</strong></p></li><li><p><strong>Cooperative enzyme</strong> → activity increases sharply after meals</p></li></ul></li><li><p><strong>Graph interpretation:</strong></p><ul><li><p>X-axis: [glucose], Y-axis: enzyme activity</p></li><li><p>Hexokinase I curve is <strong>steep at low [glucose]</strong>, plateaus quickly → saturates easily</p></li><li><p>Hexokinase IV curve is <strong>sigmoidal</strong>, rises slowly at low [glucose], accelerates at high [glucose] → acts as “glucose sensor”</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/953ffb1d-a576-4dd7-85d8-722fb13f79b4.png)
Regulation of Hexokinase IV (Glucokinase – Liver)
Function: Phosphorylates glucose → glucose-6-phosphate (G6P) only at high blood glucose (after meals)
Low affinity: Km ~10 mM → inactive at normal glucose (~5 mM)
Cooperative enzyme: Activity increases sharply with rising glucose
Regulation by subcellular localization:
Stored in nucleus when glucose is low or F6P builds up → inactive
Released into cytoplasm when [glucose] rises → active
Helps liver prevent excess G6P production when not needed
Feedback by metabolites:
F6P buildup → sequesters enzyme in nucleus
High glucose → enzyme released to cytoplasm
Purpose: Acts as a glucose sensor, matches liver glucose phosphorylation to nutrient status
![<ul><li><p><strong>Function:</strong> Phosphorylates glucose → glucose-6-phosphate (G6P) only at <strong>high blood glucose</strong> (after meals)</p></li><li><p><strong>Low affinity:</strong> Km ~10 mM → inactive at normal glucose (~5 mM)</p></li><li><p><strong>Cooperative enzyme:</strong> Activity increases sharply with rising glucose</p></li><li><p><strong>Regulation by subcellular localization:</strong></p><ul><li><p>Stored in <strong>nucleus</strong> when glucose is low or F6P builds up → inactive</p></li><li><p><strong>Released into cytoplasm</strong> when [glucose] rises → active</p></li><li><p>Helps liver <strong>prevent excess G6P production</strong> when not needed</p></li></ul></li><li><p><strong>Feedback by metabolites:</strong></p><ul><li><p>F6P buildup → sequesters enzyme in nucleus</p></li><li><p>High glucose → enzyme released to cytoplasm</p></li></ul></li><li><p><strong>Purpose:</strong> Acts as a <strong>glucose sensor</strong>, matches liver glucose phosphorylation to nutrient status</p></li></ul><p></p>](https://assets.knowt.com/user-attachments/6e4e4a20-1fed-469e-b26a-12b2dba23318.png)
Metabolic “Crossroads” of Glucose-6-Phosphate (G6P)
1. Entry & Trapping:
Glucose transporters:
Muscle/fat → GLUT4
Liver → GLUT2
Glucose → G6P via Hexokinase I (muscle/brain) or Glucokinase/Hexokinase IV (liver)
Costs 1 ATP
Hexokinase I: high affinity, always grabs glucose → energy for “selfish” organs
Glucokinase: low affinity, only active at high [glucose] → liver buffers excess
2. Four Major Fates of G6P:
Storage as Glycogen: liver (whole body) & muscle (local)
Glycolysis:
Muscle/brain → ATP for energy
Liver → building blocks for fatty acid synthesis
Glucose Secretion (Liver only): via Glucose-6-phosphatase → maintains blood glucose (~90 mg/dL)
Structural Polysaccharides: for ECM & glycoproteins
3. Key Comparisons:
Feature | Brain/Muscle | Liver |
|---|---|---|
Enzyme | Hexokinase I (high affinity) | Glucokinase (low affinity) |
Primary Goal | ATP for survival/movement | Blood sugar regulation |
Can release glucose? | No | Yes (via G6Pase) |
Takeaway: G6P is a decision point → metabolism follows the body’s needs: energy, storage, blood sugar, or structural components.
![<p><strong>1. Entry & Trapping:</strong></p><ul><li><p><strong>Glucose transporters:</strong></p><ul><li><p>Muscle/fat → <strong>GLUT4</strong></p></li><li><p>Liver → <strong>GLUT2</strong></p></li></ul></li><li><p>Glucose → <strong>G6P</strong> via <strong>Hexokinase I (muscle/brain)</strong> or <strong>Glucokinase/Hexokinase IV (liver)</strong></p></li><li><p>Costs <strong>1 ATP</strong></p></li><li><p><strong>Hexokinase I:</strong> high affinity, always grabs glucose → energy for “selfish” organs</p></li><li><p><strong>Glucokinase:</strong> low affinity, only active at high [glucose] → liver buffers excess</p></li></ul><p><strong>2. Four Major Fates of G6P:</strong></p><ol><li><p><strong>Storage as Glycogen:</strong> liver (whole body) & muscle (local)</p></li><li><p><strong>Glycolysis:</strong></p><ul><li><p>Muscle/brain → ATP for energy</p></li><li><p>Liver → building blocks for fatty acid synthesis</p></li></ul></li><li><p><strong>Glucose Secretion (Liver only):</strong> via <strong>Glucose-6-phosphatase</strong> → maintains blood glucose (~90 mg/dL)</p></li><li><p><strong>Structural Polysaccharides:</strong> for ECM & glycoproteins</p></li></ol><p><strong>3. Key Comparisons:</strong></p><table style="min-width: 75px;"><colgroup><col style="min-width: 25px;"><col style="min-width: 25px;"><col style="min-width: 25px;"></colgroup><tbody><tr><th colspan="1" rowspan="1"><p>Feature</p></th><th colspan="1" rowspan="1"><p>Brain/Muscle</p></th><th colspan="1" rowspan="1"><p>Liver</p></th></tr><tr><td colspan="1" rowspan="1"><p>Enzyme</p></td><td colspan="1" rowspan="1"><p>Hexokinase I (high affinity)</p></td><td colspan="1" rowspan="1"><p>Glucokinase (low affinity)</p></td></tr><tr><td colspan="1" rowspan="1"><p>Primary Goal</p></td><td colspan="1" rowspan="1"><p>ATP for survival/movement</p></td><td colspan="1" rowspan="1"><p>Blood sugar regulation</p></td></tr><tr><td colspan="1" rowspan="1"><p>Can release glucose?</p></td><td colspan="1" rowspan="1"><p>No</p></td><td colspan="1" rowspan="1"><p>Yes (via G6Pase)</p></td></tr></tbody></table><p><strong>Takeaway:</strong> G6P is a <strong>decision point</strong> → metabolism follows the body’s needs: energy, storage, blood sugar, or structural components.</p>](https://assets.knowt.com/user-attachments/c4debda9-a686-401e-8947-b4274660bd4a.png)
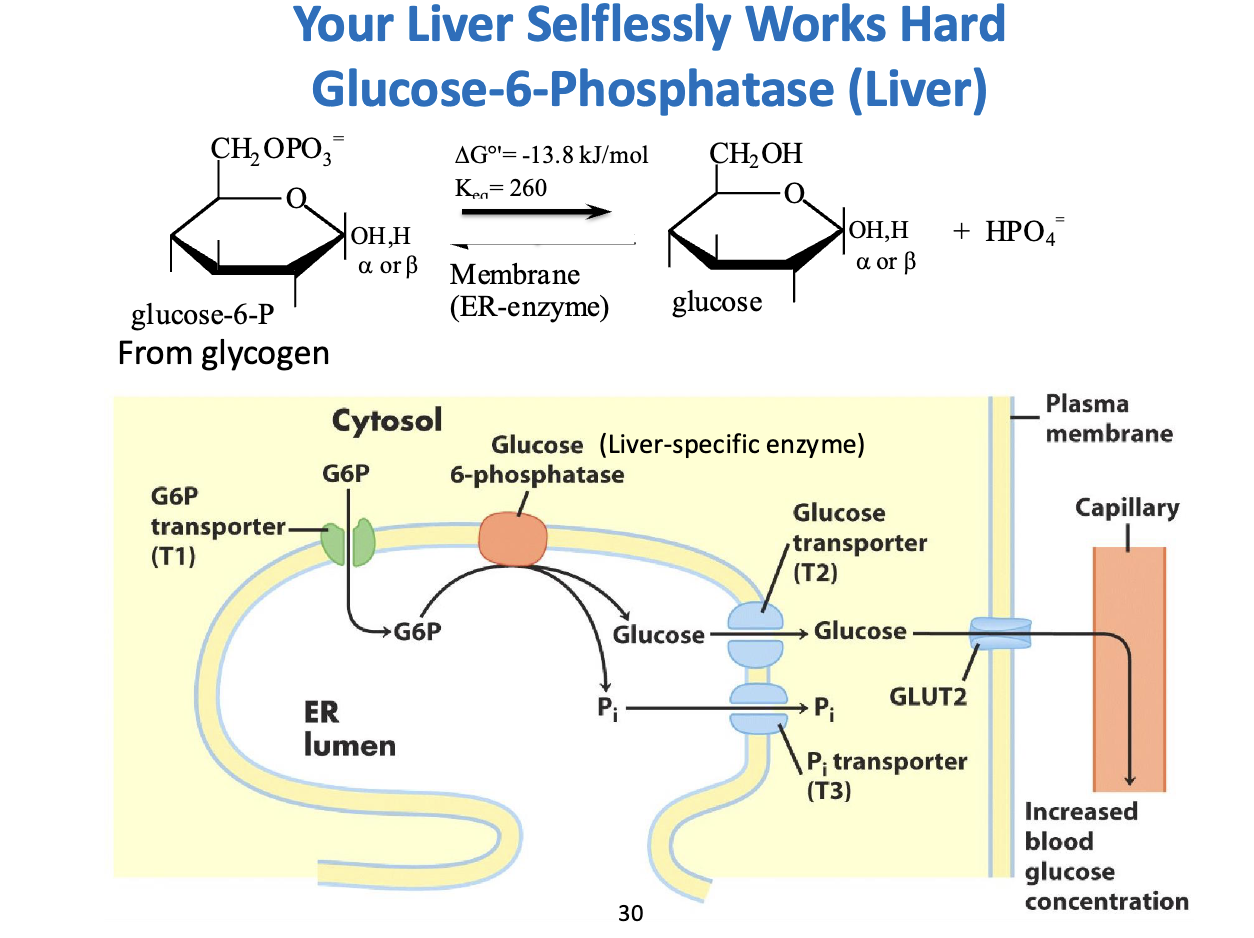
How the Liver “Selflessly” Exports Glucose – Glucose-6-Phosphatase
1. The Chemistry:
Reaction: G6P + H₂O → Glucose + Pi
ΔG°' = -13.8 kJ/mol → spontaneous, exergonic
Purpose: G6P is charged → cannot cross membranes. Dephosphorylation makes free glucose that can exit the cell.
2. Compartmentalization in the ER:
Step 1 (T1): G6P transported from cytosol → ER lumen
Step 2: Glucose-6-phosphatase in ER removes phosphate → glucose + Pi
Step 3 (T2 & T3): Glucose exits ER via T2, Pi exits via T3
Step 4: Free glucose leaves cell via GLUT2 → enters bloodstream
3. Why “Selfless”?
Muscle lacks G6Pase → must use G6P for own energy
Liver can release glucose to feed other organs (brain, muscles) during fasting
ER acts as a secure room → prevents accidental glucose loss; release is regulated (e.g., by glucagon)
Takeaway: The liver has a specialized ER-based system to convert stored G6P into blood glucose, supporting other tissues when needed.
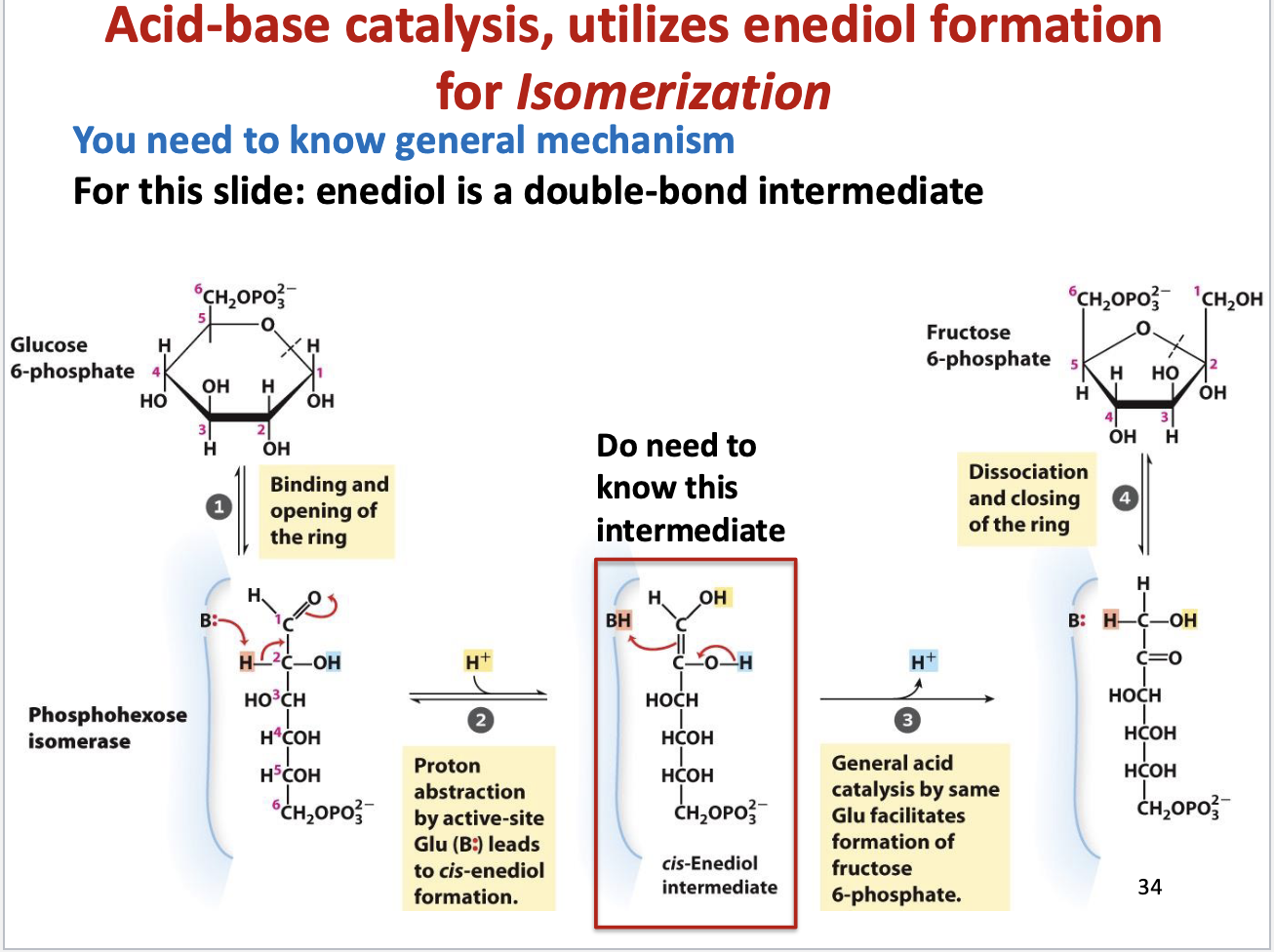
Step 2 of Glycolysis – G6P → F6P (Phosphohexose Isomerization)
Enzyme: Phosphoglucose/Phosphohexose Isomerase
Mechanism:
Ring Opening:
Enzyme binds G6P and opens the 6-membered pyranose ring → linear chain.
Proton Abstraction & Enediol Formation:
Glutamate (B:) abstracts proton from C2 → forms cis-Enediol intermediate (double bond between C1-C2, two -OH groups)
Key intermediate – must know for exams
General Acid Catalysis:
Proton added back “upside down”
C1 (aldehyde → alcohol)
C2 (alcohol → ketone)
Converts aldose (glucose) → ketose (fructose)
Ring Closing:
F6P forms a 5-membered furanose ring
Product leaves enzyme
Why Important:
Moves carbonyl from C1 → C2 → primes for Step 4 (Aldolase cleavage)
Easier symmetrical cleavage of 6-carbon sugar
Key Exam Points:
Intermediate: cis-Enediol
Catalysis type: General acid-base
Active site residue: Glutamate (Glu) acts as base
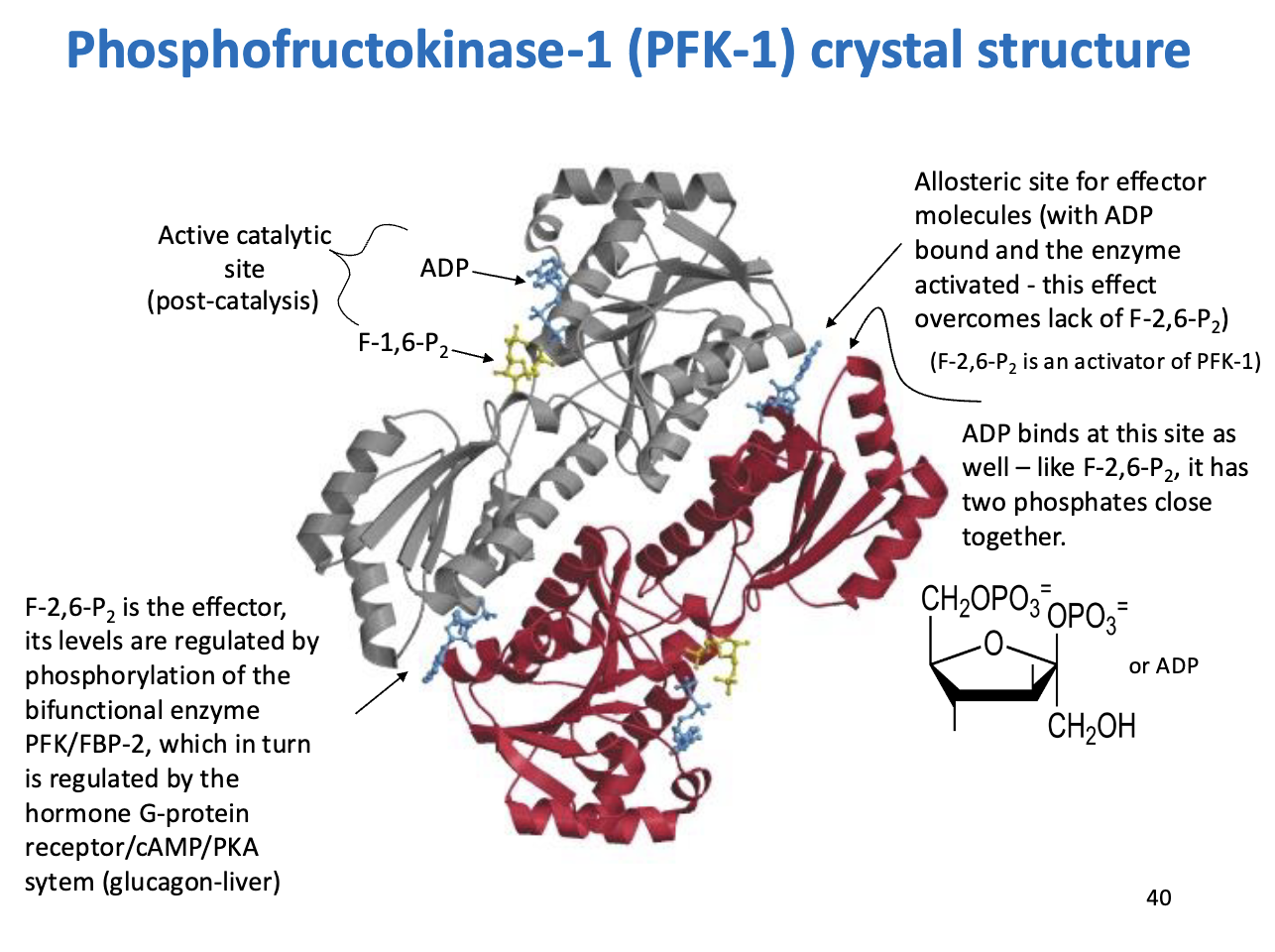
Phosphofructokinase-1 (PFK-1) – Regulation and Effectors
Enzyme: PFK-1 – catalyzes F6P → F1,6-bisphosphate (first committed step of glycolysis)
Regulatory Highlights:
Allosteric Regulation by F-2,6-bisphosphate (F-2,6-P2):
Activator of PFK-1
Levels controlled by PFK/FBP-2 bifunctional enzyme
PFK/FBP-2 regulated via glucagon → G-protein/cAMP/PKA signaling in liver
Allosteric Site for ADP:
ADP can also bind here
Mimics effect of F-2,6-P2 (two phosphates close together)
Helps activate PFK-1 when F-2,6-P2 is low
Active Catalytic Site:
Where F6P binds and is phosphorylated
Takeaway:
PFK-1 integrates energy signals:
High F-2,6-P2 → glycolysis ON
High ATP → glycolysis OFF (not in this slide but important)
ADP can override low F-2,6-P2 to ensure energy production
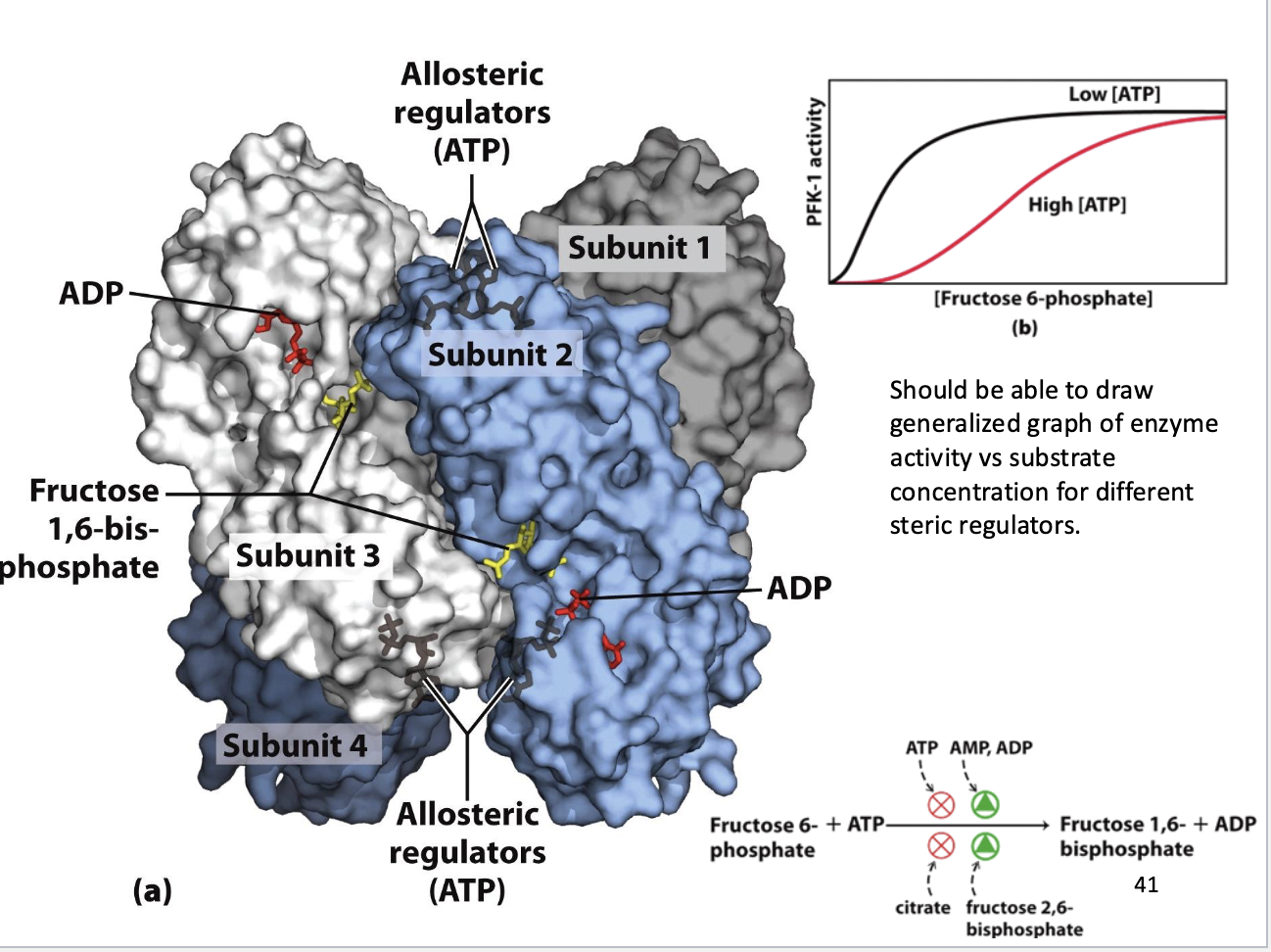
Phosphofructokinase-1 (PFK-1) – Control Valve of Glycolysis
1. Structure:
Tetramer (4 subunits)
Active sites: F6P + ATP binding → catalysis
Allosteric sites: regulators bind → “on/off switches”
2. Regulators:
Red Lights (Inhibitors):
ATP: high energy → binds allosterically → slows enzyme
Citrate: Citric Acid Cycle backed up → inhibits glycolysis
Green Lights (Activators):
AMP/ADP: low energy → activate enzyme
Fructose 2,6-bisphosphate (F-2,6-P2): most potent activator
3. Sigmoidal Kinetics:
Low ATP: hyperbolic curve → enzyme efficient at low substrate
High ATP: S-shaped (sigmoidal) curve → decreased substrate affinity, higher $K_m$
Reason: ATP allosterically decreases PFK-1 affinity for F6P to prevent excess glycolysis
4. Biological Takeaway:
PFK-1 = metabolic “logic gate”
High energy → glycolysis slows, saves glucose
Low energy → glycolysis accelerates, makes ATP
5. Exam Tip:
Sigmoidal vs hyperbolic curve often tested
F-2,6-P2 = strongest activator; remember it’s not the product
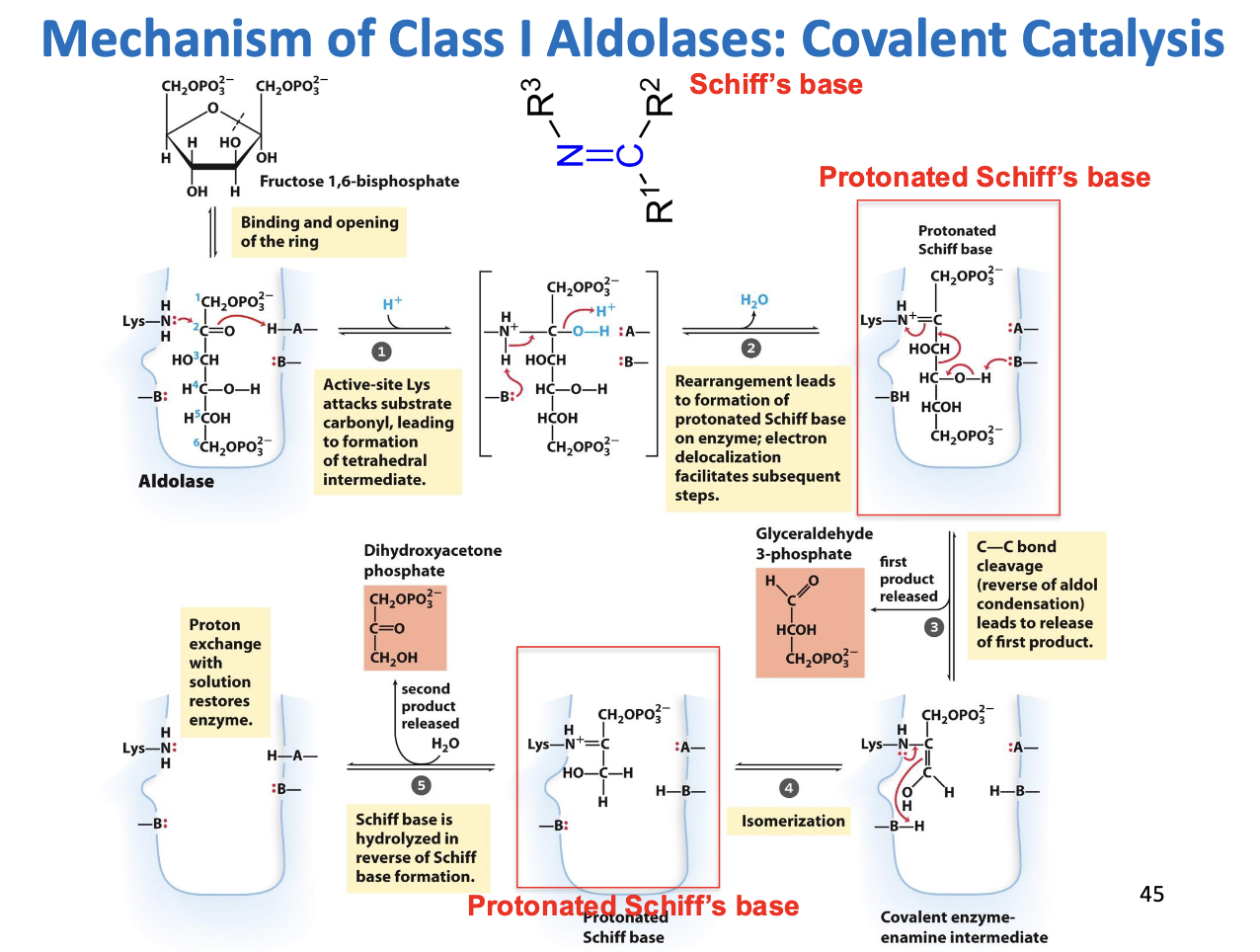
Mechanism of Class I Aldolases: Covalent Catalysis
Mechanism Step by Step 1. Ring Opening
F1,6BP is normally a ring (pyranose form).
Aldolase binds the sugar and opens the ring to the linear form.
This is necessary because the C-C bond cleavage happens in the linear molecule.
2. Formation of Schiff Base (C=N)
Key player: Active-site Lysine (Lys).
Lysine has an amine (-NH2) group that is nucleophilic.
It attacks the carbonyl carbon at C2 of F1,6BP.
Water is eliminated, forming a Schiff base (C=N), also called an imine.
Why does this happen?
The Schiff base acts as an electron sink.
When the C3-C4 bond breaks, electrons flow toward the nitrogen, stabilizing the negative charge.
Without this, the bond would be very hard to break.
⚡ Think of it as the enzyme “holding the electrons” so the sugar can be chopped safely.
3. C-C Bond Cleavage
Now the C3-C4 bond is activated by the electron sink (Schiff base).
The bond breaks, releasing the first product: GAP.
The remaining three carbons (DHAP portion) are still attached to Lys as a covalent enamine intermediate.
4. Rearrangement of the Enamine
The remaining 3-carbon piece is still bound to the enzyme.
It rearranges internally to form a Schiff base with proper orientation for hydrolysis.
5. Hydrolysis – Release of DHAP
Water attacks the Schiff base, breaking the C=N bond.
Lys is regenerated (freed) for another reaction.
DHAP, the ketose product, is released.
Key Concepts
Schiff Base (C=N):
Covalent bond between Lys nitrogen and sugar carbonyl.
Stabilizes electrons during bond cleavage → acts as electron sink.
Covalent Catalysis:
The enzyme physically “glues” to the substrate temporarily.
Allows unusual chemistry like breaking a strong C-C bond.
Why the Mechanism Exists:
F1,6BP is symmetrical. By forming a Schiff base, aldolase can selectively break the bond between C3-C4.
Makes the reaction controlled and reversible.
Products:
GAP (aldose) released first
DHAP (ketose) released second
How to Remember It
Think of the Lys as a hook: it grabs the sugar and pulls electrons, so the “chop” can happen.
Schiff base = electron stabilizer = makes impossible bond breaking possible.
Class I Aldolase uses covalent catalysis; Class II (bacteria/fungi) uses metal ions instead.
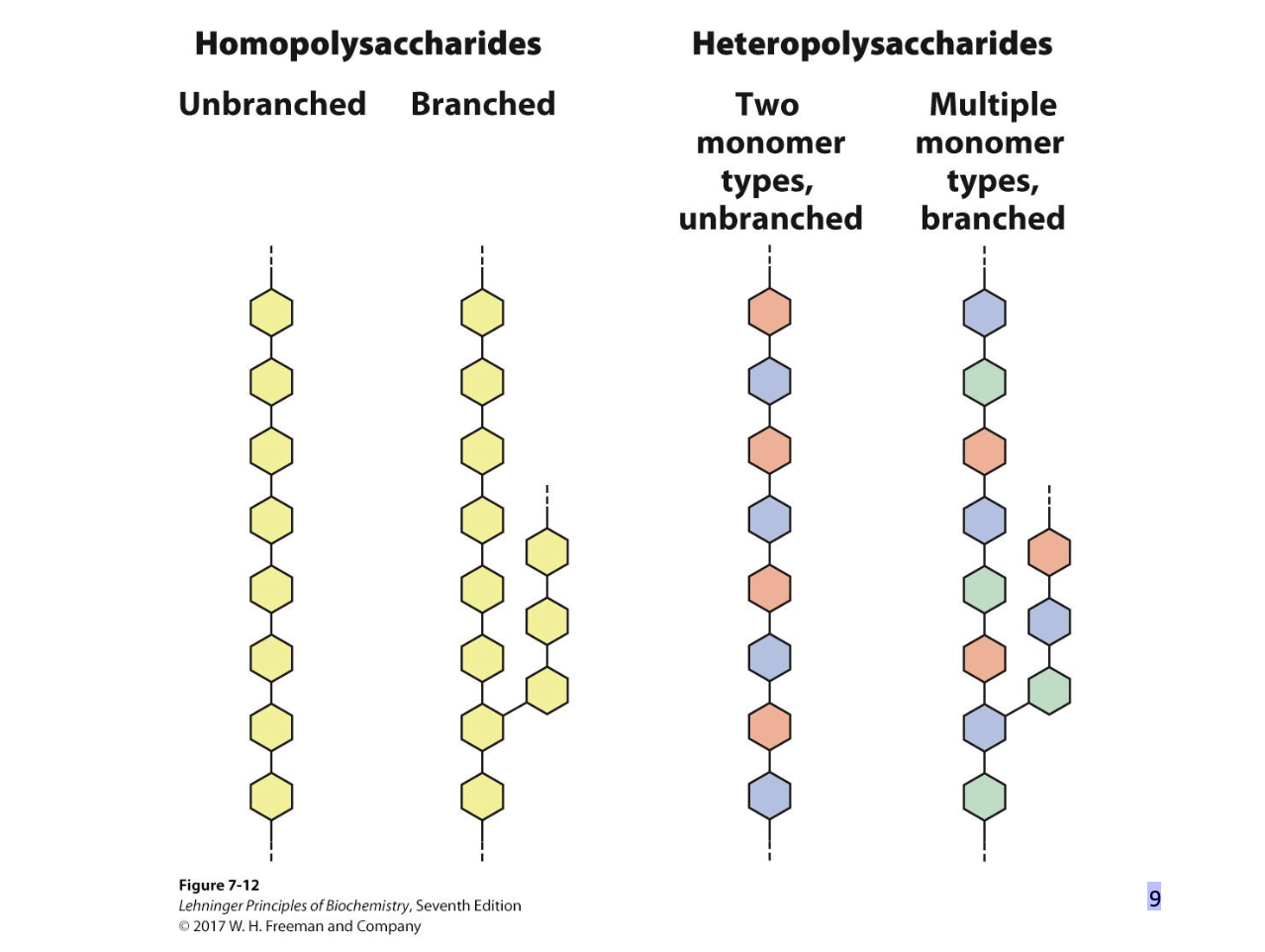
Polysaccharides
Natural carbohydrates are usually found as polymers.
• These polysaccharides can be:
– homopolysaccharides (one monomer unit)
– heteropolysaccharides (multiple monomer units)
– linear (one type of glycosidic bond)
– branched (multiple types of glycosidic bonds)
• Polysaccharides do not have a defined molecular weight.
– No template
– Are in flux (constantly being degraded and rebuilt)
3 examples of Homopolymers of Glucose
Glycogen – mainly (
1 → 4) bonds; branching with (
1 → 6) every
8-12 residues. Storage polysaccharide in animals. MW – n*106. Water
insoluble. Highly branched molecule
Starch – amylose (linear (
1 → 4) bonds) + amylopectin (branching
with (
1 → 6) every 24-30 residues). Storage polysaccharide in
plants. MW –up to 2*108. Water insoluble. Moderately branched.
Cellulose – linear (
1 → 4) chains. Hydrogen bonds between
adjacent monomers and chains. The most abundant polysaccharide
in nature. Water insoluble. Cannot be digested by humans. No
branching.
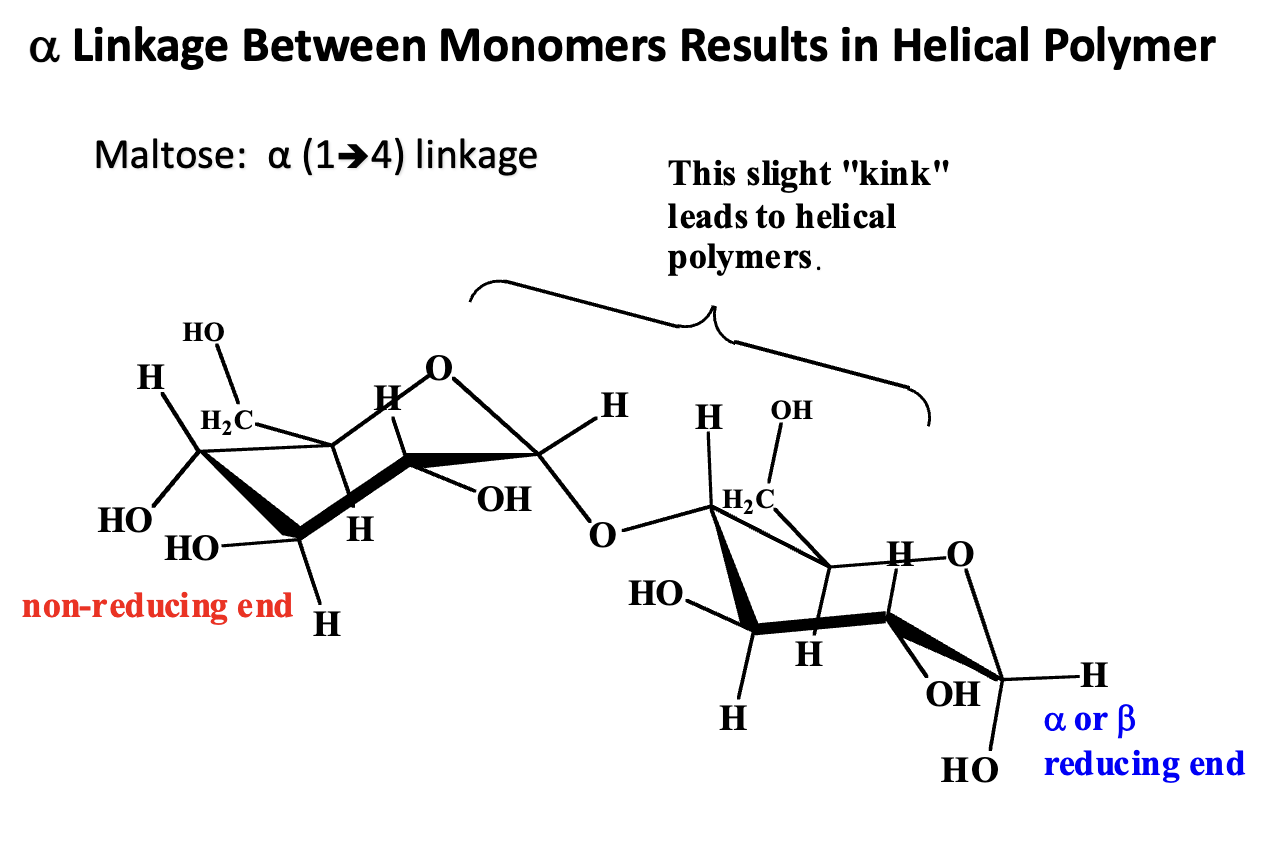
What is the structural consequence of an α-glycosidic linkage between monosaccharide monomers?
α-Glycosidic Linkage:
A bond between the anomeric carbon (C1) of one glucose and a hydroxyl group (usually C4) of another glucose.
In α-linkages, the OH on C1 is “down” (opposite side of CH2OH at C5 in glucose).
Structural Consequence:
The α-linkage forces the glucose units to bend slightly relative to each other.
Repeating α-linkages produce a helical polymer rather than a straight chain.
Example:
α(1→4) linkage in starch (amylose) forms a right-handed helix.
The helical structure is compact, ideal for energy storage.
Contrast with β-Linkages:
β(1→4) linkages (as in cellulose) produce straight, rigid chains that can form hydrogen-bonded sheets, not helices.
Takeaway:
α-linkages → helix → storage polysaccharides (starch, glycogen)
β-linkages → straight → structural polysaccharides (cellulose)
Starch Detection
Starch forms long helices that bind polyiodine.
Charge transfer of electrons from starch to iodine allows absorption of yellow-red light resulting in a blue color.
Starch Structure: Forms long helices.
Iodine Binding: Iodine fits inside the helix.
Electron Transfer: Electrons move between starch and iodine.
Color Change: This absorbs yellow-red light, so we see blue.
Takeaway:
Blue color = starch present (used in iodine tests).
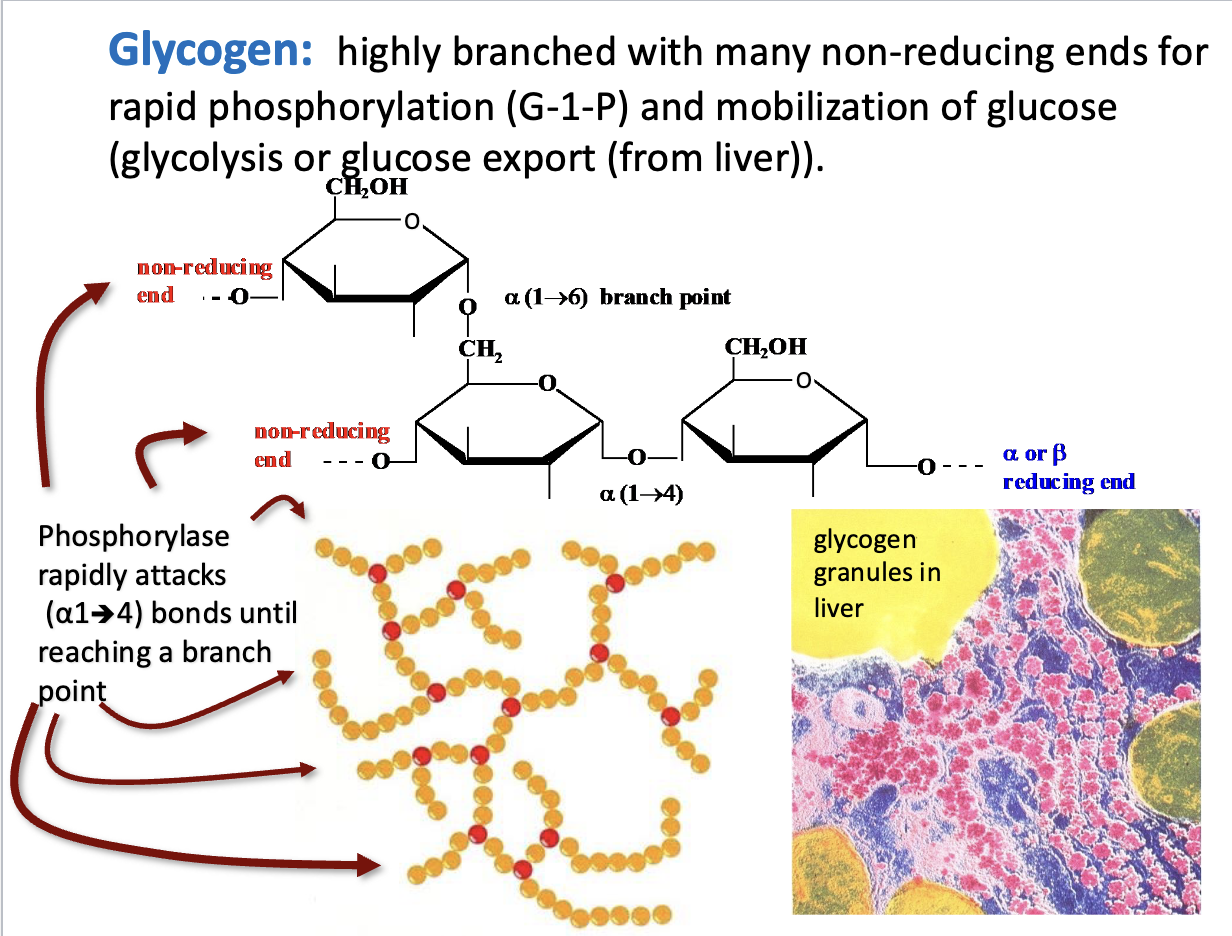
Glycogen
Highly branched structure: Many α(1→6) branches.
Non-reducing ends: Sites where glycogen phosphorylase can rapidly remove glucose as glucose-1-phosphate (G-1-P).
Function: Allows fast glucose mobilization for glycolysis or export (liver).
Mechanism: Phosphorylase works on α(1→4) bonds until it hits a branch.
Takeaway: Branching = faster energy release.
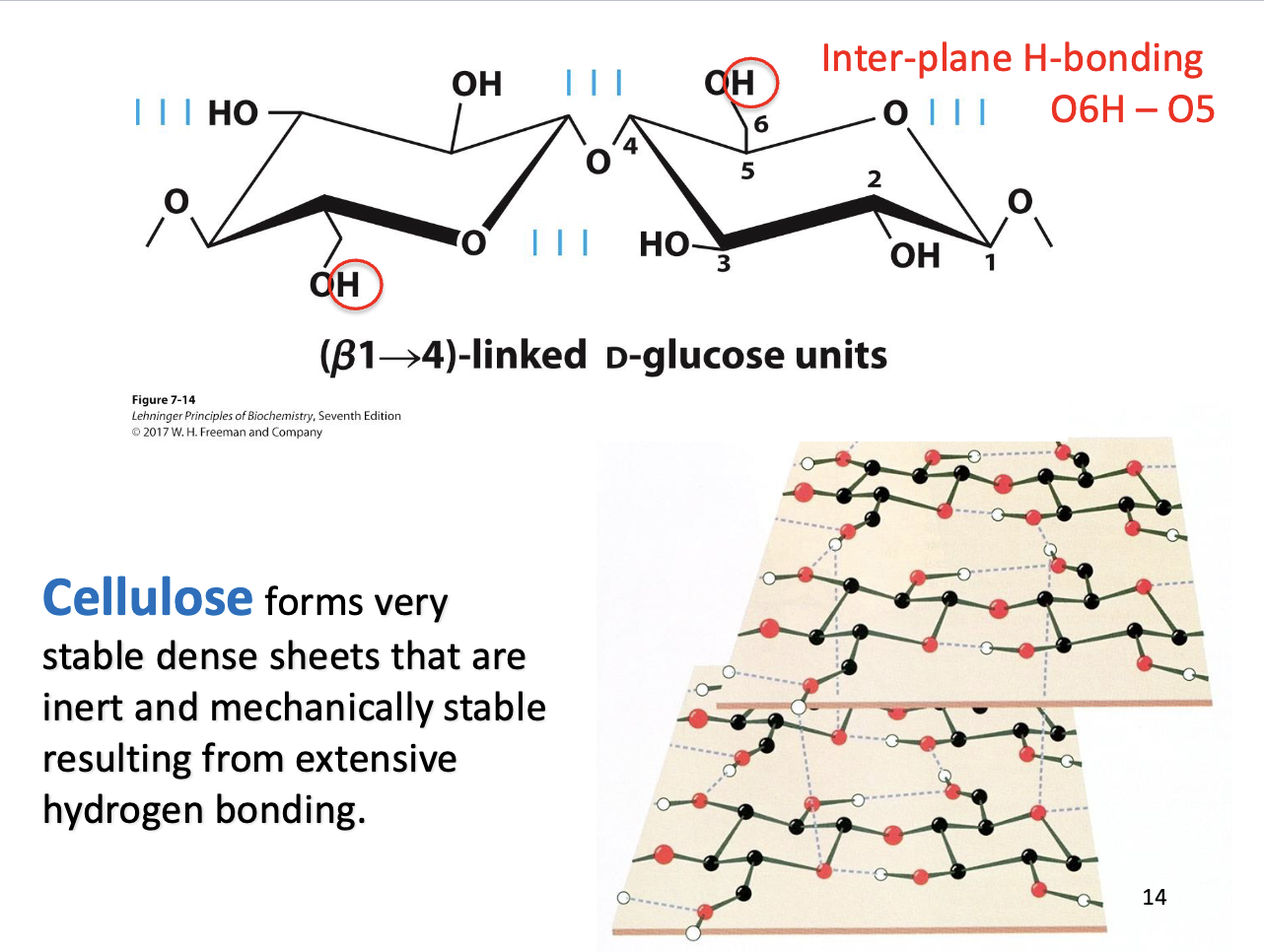
Cellulose
Structure: Forms dense sheets.
Stability: Extensive hydrogen bonding between chains.
Result: Mechanically strong and chemically inert (resists breakdown).
Takeaway: Cellulose’s hydrogen-bonded sheets give plants structural support.
What are the major pathways of utilization of Glucose
Synthesis of structural polymers = Extracellular matrix and cell wall polysaccharides
Oxidation via pentose phosphate pathway = Ribose 5-phosphate
Storage = Glycogen, starch, sucrose
Oxidation via glycolysis = Pyruvate
Central Importance of Glucose
• Glucose is an excellent fuel.
– yields good amount of energy upon oxidation
• −2,840 kJ/mol glucose
– can be efficiently stored in the polymeric form
– Many organisms and tissues can meet their energy
needs on glucose alone.
• Glucose is a versatile biochemical precursor.
– Many organisms can use glucose (or metabolic
derivatives of glucose) to generate:
• all the amino acids (only some in humans)
• membrane lipids
• nucleotides in DNA and RNA
• cofactors needed for the metabolism
How do different organs share and use fuel in the body?
Liver – The Central Hub
Stores glycogen, makes new glucose (gluconeogenesis), converts excess sugar to fat.
Manages blood glucose for the whole body.
Brain – The Picky Consumer
Uses only glucose or ketone bodies.
Burns fuel completely to $CO_2$ and $H_2O$.
Heart – Steady Burner
Mainly uses fatty acids, but can also use glucose.
Muscle – Flexible Burner
Uses glucose and fatty acids.
Stores glycogen for quick energy.
Produces lactate under anaerobic stress (Cori Cycle).
Adipose Tissue – The Storage Unit
Stores energy as triacylglycerides.
Releases fatty acids during fasting or exercise for other organs to use.
Cori Cycle (Stress Loop):
Lactate from muscles → blood → liver → converted back to glucose.
Key Takeaway: Fuel is shared and managed; everything ultimately ends as CO2 and H2O.
What are the three stages of glucose/fuel metabolism and their main outcomes?
Stage 1: Acetyl-CoA Production
Fuels: Carbs, fats, proteins → Acetyl-CoA
Pathways: Glycolysis → Pyruvate → Pyruvate Dehydrogenase Complex → Acetyl-CoA + $CO_2$
ATP Yield: 2 ATP (fast but inefficient)
Stage 2: Acetyl-CoA Oxidation (Citric Acid/Krebs Cycle)
Goal: Harvest electrons, not ATP directly
Output: NADH, FADH₂, and more $CO_2$
Stage 3: Electron Transfer & Oxidative Phosphorylation
Process: NADH & FADH₂ electrons → Electron Transport Chain → O₂ → H₂O
ATP Yield: ~30 ATP (protons drive ATP synthase)
Key Takeaway:
Total ATP from glucose: ~32
Without mitochondria/O₂: Only 2 ATP → lose ~94% of energy
Glycolysis: Features
Sequence of enzyme-catalyzed reactions by which
glucose is converted into pyruvate
• Pyruvate can be further aerobically oxidized (TCA cycle).
• Pyruvate can be used as a precursor in biosynthesis.
Some of the free energy is captured by the synthesis of ATP and NADH.
• Research of glycolysis played a large role in the development of modern biochemistry.
What key reaction in glycolysis produces a direct net gain of 2 ATP per glucose?
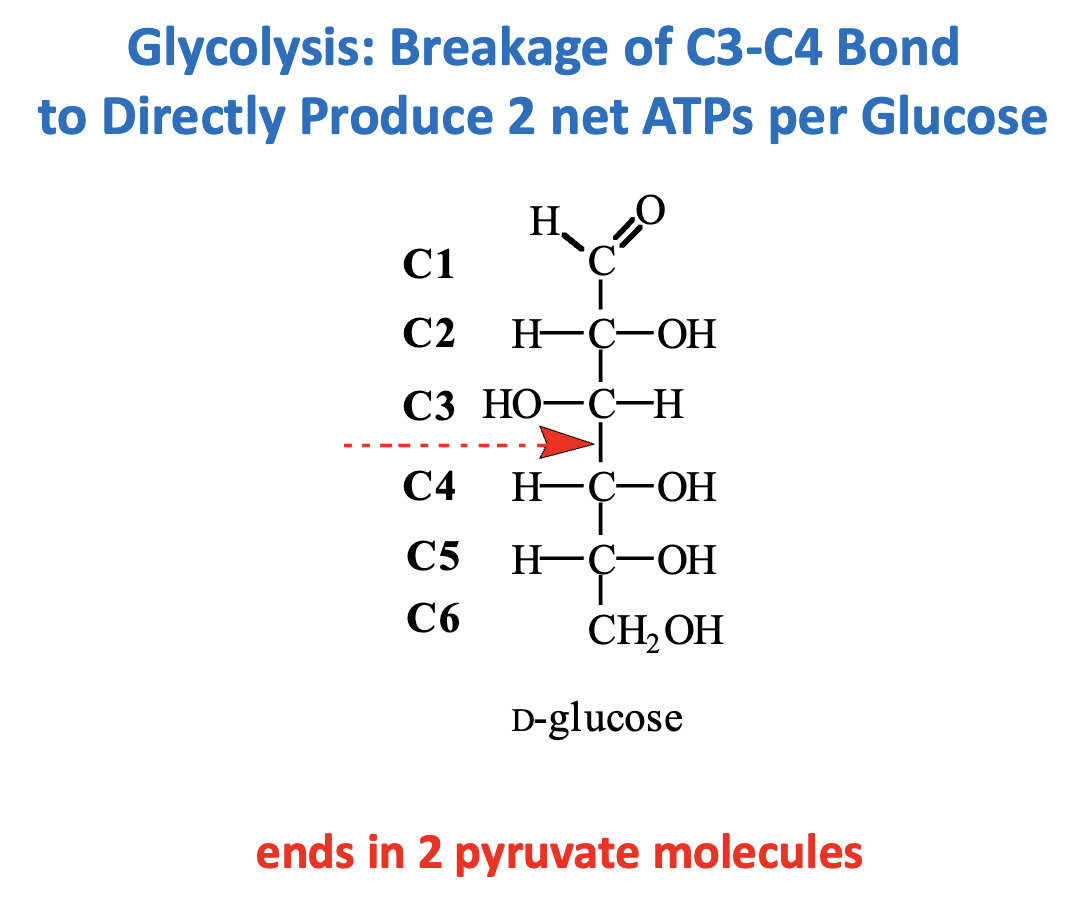
Reaction: Cleavage of Fructose 1,6-bisphosphate (F1,6BP) → Glyceraldehyde 3-phosphate (GAP) + Dihydroxyacetone phosphate (DHAP)
Step: C3–C4 bond is broken (Step 4, Aldolase reaction)
Outcome: Prepares two 3-carbon molecules for subsequent energy-producing steps
Direct ATP: 2 net ATP per glucose (from later substrate-level phosphorylation in glycolysis)
Key Point: This bond cleavage is essential for generating two “units” that each yield ATP, doubling the energy output from a single glucose.
What are the main categories of chemical reactivity in metabolism?
1. Cleavage & Formation of C–C Bonds
Making or breaking carbon-carbon bonds (e.g., Aldolase in glycolysis).
2. Internal Rearrangements & Eliminations
Changing molecule structure without adding/removing atoms (e.g., isomerization like G6P → F6P).
3. Group Transfers
Moving functional groups between molecules (e.g., H⁺, CH₃⁺, PO₄³⁻).
4. Oxidation–Reduction (Electron Transfers)
Moving electrons from one molecule to another (e.g., dehydrogenases, reductases).
Key Point: Almost all metabolic reactions can be categorized into one of these four types.