Exam 3 - Do Not Share
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
the sequence of one strand of DNA is 5’ - TCGATC - 3’. The sequence of the complementary strand would be…
5’ - GATCGA - 3’
suggestion: write out the answer with corresponding ends and nucleotides before looking at answer choices
RNA differs from DNA in all EXCEPT which of the following ways?
Reword: In what way does RNA and DNA relate?
polynucleotides form a sugar-phosphate backbone
Nucleotides that have not yet been incorporated into a DNA molecule (free nucleotides) occur in the cell in which form?
deoxynucleoside triphosphate (dNTP)
Has three phosphates
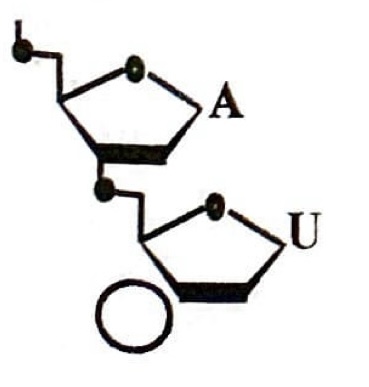
Refer to the figure of the dinucleotide; is it DNA or RNA? Also, is the circle closest to the 5’ or 3’ end?
RNA
3’ end
A pyrimidine is paired with what?
A purine in DNA or RNA.
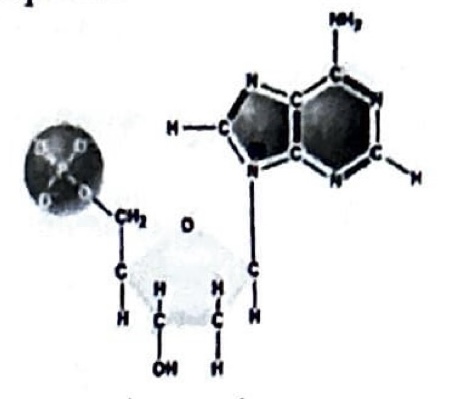
In a double stranded DNA molecule, this nucleotide would be paired with a:
Pyrimidine
Which is true of the structure of a double stranded DNA molecule?
Phosphodiester bonds between sugar-phosphate groups are on the outside of the DNA molecule
Mutations that arise from agents of normal cellular process are called:
spontaneous mutations
Oblique mutations are defined as…
mutations that occur due to external factors or environmental influences, rather than from errors during DNA replication.
What is the difference between oblique and induced mutations?
Oblique mutations arise from external factors or environmental influences, while induced mutations are specifically caused by known mutagens or agents, leading to alterations in the DNA sequence.
Examples of induced mutations?
Chemical agents, radiation, viral infections.
Mismatch, deamination, and deprivation are what type of mutations?
Spontaneous mutations
Which of the following mutation categories is most likely to be the consequence of UV radiation exposure (non-spontaneous)?
thymine-thymine dimer (induced mutation)
Which of the following is most true with respect to a tautomeric shift?
Tautomers do not follow normal rules of complementation resulting in replication errors
Which of the following is NOT true of spontaneous mutation rates?
The spontaneous mutation rate cannot be measured in complex animals such as mammals
How do TC-NER pathways differ from GG-NER pathways
In TC-NER pathways RNA polymerase stalling at a damaging DNA region signals the start of the NER pathway, and in GG-NER the XPC/Rad protein complex detects the damaged DNA
Homologous recombination is used by the cell to repair specific types of DNA damage because:
the homologous chromosome can act as a template to repair the mutated chromosome
Individuals with Xeroderma Pigments accumulate mutations caused by UV induced DNA damage at a much more rapid rate than individuals without the disease. One common form of the disease is caused by a mutation in a gene that produces a non-functional XPC protein. Why does this result in a rapid accumulation of mutations?
Without a functioning XPC, UV induced mutations cannot be detected and the repair process never starts
Which of the following are produced by the process of transcription?
mRNA
tRNA
rRna
sncRNA
All of these are produced during transcription, which is the synthesis of RNA from a DNA template.
mRNA
tRNA
rRna
sncRNA
In the genome, which of the following is a general function of the promoter sequence?
mRNA
tRNA
rRna
sncRNA
mRNA
tRNA
rRna
sncRNA
Which of the following is a general function of the promoter?
guides RNA polymerase to start transcribing immediately downstream (+1 site)
In a double-stranded section of genomic DNA that functions as a gene, both DNA strands function as the template for transcription. True or false?
False, only one strand serves as the template.
Which of the following most accurately summarizes the structure and functions of the RNA polymerase core enzyme?
multiple subunits, DNA unwinding, 5’ to 3’ polymerization
If a mutation occurred that eliminated the start codon from the template DNA molecule, what would be the consequence for the process of transcription?
This mutation would not have a consequence for transcription
if a mutation occurred that caused the elimination of a terminator sequence, what would be the consequence for the process of transcription?
A longer RNA molecule
RNA polymerase moves along the template strand of DNA in the _______ direction, and adds nucleotides to the __________ end of the growing transcript.
3’ to 5’
3’
What would be the result if the sigma subunit of bacterial RNA polymerase did Not function during transcription?
RNA polymerase would fail to initiate transcription
The process where general transcription factors aggregate to form the pre-initiation complex (PIC) is necessary for:
RNA polymerase binding
Regulatory sequences in bacterial genomes that form hairpin loops are important for:
transcription termination
Function(s) of the 5’ - cap in eukaryotic mRNA includes which of the following
ribosomal binding
transport
protection
Alternative splicing allows for which of the following to occur in eukaryotes?
a single gene to encode more than one protein
For any organism, genes must be expressed in the correct place at the correct time, but the amount (level) of expression doesn’t matter. true or false?
False, as the level of expression is crucial for normal biological function and development.
Which of the following has the capacity to increase transcription rates?
mutation in silencers
activators
the cell’s ability to respond to physical or chemical signals through the expression of appropriate gene products is facilitated by:
transcription factors
You have identified a mutation in humans that decreases transcription levels of a gene 5000 nucleotides away from the mutation site. What regulatory sequence has likely been mutated in this instance?
enhancer sequence
In eukaryotic genes, the 3’ polyadenylation sequence (AAUAAA) functions to:
signal other molecules to cut the RNA from RNA polymerase to terminate transcription
Some introns contain functional non-coding RNA or cis-regulatory sequences. True or false?
True; some introns can regulate gene expression and produce non-coding RNAs.
Which of the following explains why a larger number of proteins are produced by the human genome than is predicted by the number of genes humans possess?
different exons can be included or excluded during post-transcriptional processing
RNA editing is important for tRNA function because the cell insert nucleotides other than A, G, C, or U. True or False?
True; RNA editing allows for the modification of tRNA nucleotides, which is essential for their proper function in protein synthesis.
in eukaryotic genomes, cis-regulatory sequences are typically:
a greater proportion to genes
The common form of the disease Beta-thalassemia is caused by a mutation in the TATA box of the beta-hemoglobin gene. This mutation results in a disease phenotype because:
this mutation is a regulatory mutation of the promoter resulting in under-production of the beta-hemoglobin protein
What is the most likely consequence of a mutation in an intronic splice recognition site?
A non-functional gene
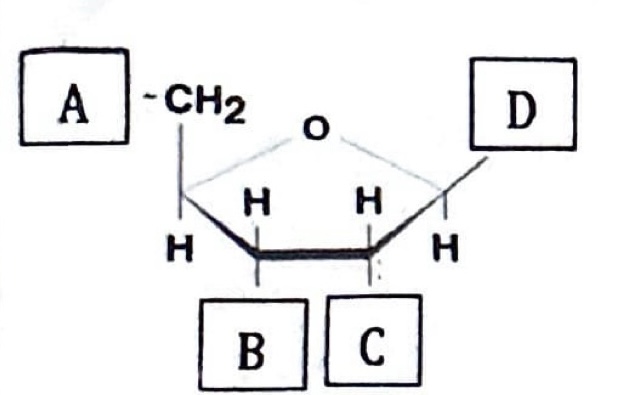
Below is a figure of a nucleotide with regions/structures indicated by letters (A-D). Match the appropriate region/structure. (answers may be used more than once)
Phosophate group —> A
Where hydrogen bonds will form with a complementary base —> D
Used to designate the 3’ end —> B
Match the DNA repair mechanism to the category of mutations they recognize:
Double stranded DNA Repair —>
Recombination
Match the DNA repair mechanism to the category of mutations they recognize:
Double stranded DNA containing a pairing between a normal thymine and a normal guanine —>
Mismatch repair
Match the DNA repair mechanism to the category of mutations they recognize:
Destabilizing/bulky DNA damage such as pyrimidine dimers —>
nucleotide excision repair
Match the DNA repair mechanism to the category of mutations they recognize:
Deamination or deprivation —>
Base excision repair
Match the sequence with the molecule that recognizes it in the context of regulating expression in eukaryotes. (Answers may be used once, more than once, or not at all, one answer per question).
TATA-binding protein (TBP) →
Core promoter
Match the sequence with the molecule that recognizes it in the context of regulating expression in eukaryotes. (Answers may be used once, more than once, or not at all, one answer per question).
Activator —>
Enhancer
Match the sequence with the molecule that recognizes it in the context of regulating expression in eukaryotes. (Answers may be used once, more than once, or not at all, one answer per question).
Repressor —>
Silencer
Match the sequence with the molecule that recognizes it in the context of regulating expression in eukaryotes. (Answers may be used once, more than once, or not at all, one answer per question).
snRNPs —>
GU or AG exon-intron boundary sequence
Use these symbols and the underlined representation of the eukaryotic transcript molecule below to answer the following questions.
E = exons
I = intron
UTR = untranslated region
5’ - UTR E1 I1 E2 I2 E3 I3 E4 UTR - 3’
Which components of the transcript molecule will also be found in immature mRNA?
5’ - UTR E1 I1 E2 I2 E3 I3 E4UTR - 3’
Use these symbols and the underlined representation of the eukaryotic transcript molecule below to answer the following questions.
E = exons
I = intron
UTR = untranslated region
5’ - UTR E1 I1 E2 I2 E3 I3 E4 UTR - 3’
Which components of the transcript molecule will also be found in mRNA after post-transcriptional processing?
5' - UTR E1 E2 E3 E4 UTR- 3'