A3.1 Diversity of organisms
1/41
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
42 Terms
Define taxons
The category organisms are grouped in
Linnaeus’ hierarchy system
Kingdom
Phylum
Class
Order
Family
Genus
All species in the same genus have a common ancestor
Species
(Mnemonic: Keep Ponds Clean Or Frogs Get Sick)
What are species (Linnaeus definition)
Groups of organisms with similar traits
Classification by morphology and its limitations
Categorisation based on the physical appearance of an organism, which has been traditionally used
Limitation: Variations in characteristics need not be physical appearance (e.g. variations in mating calls)
Reading binomial nomenclature
significance of genus name
Scientific names follow the format of → Genus species
When writing scientific names, they are to be underlined, e.g.: Genus species
After writing a binomial once in text, it can be abbreviated to the initial letter of the genus name with the full species name, e.g. H. sapiens or H. sapiens
species in the same genus have similar traits
Define species (biological species concept)
species are groups of organisms that can potentially interbreed to produce fertile offspring (i.e. can produce more offspring)
who proposed biological species concept
ernst mayr
why must offspring be fertile to be considered species
it is possible for organisms of different species to breed to produce infertile hybrid offspring.
Limitations of the biological species concept
Cannot be applied to organisms that reproduce asexually (and therefore do not breed), e.g. bacteria (binary fission)
Hybrids from parents of closely related but separate species are usually, but not always, infertile
Some species have DNA from multiple species, making classification into a distinct species difficult
Impossible to apply to extinct species, as it is not possible to tell whether members of a population can interbreed from skeletal remains
In addition, in organisms like bacteria there is horizontal gene transfer where plasmids (genetic material) are transferred without mating, and can take place between different species of bacteria
When a bacterium has a mix of genes from its own species that it received from previous generations, mixed with genes from a donor species during its lifetime, it poses a challenge to the idea of sharing a common ancestry with the other members of its species.
If horizontal gene transfer happened several times in previous generations and again during its lifespan, the genetic material inside the bacterium would be a mosaic of genes from various sources.
Gene transfer can occur from bacteria to archaea, viruses to eukaryotes, and bacteria to eukaryotes
Parthenogenesis in certain insects such as stick insects, which produce young without mating
Vegetative propagation in plants, e.g. strawberries and potatoes, which produces new plants that are identical copies of the parent plant
Other ways to define species, aside from morphology and biological species concept
Ecological Niche
Single celled microbes are an example of the limitations of morphology in defining species
The environmental conditions and what these microbes consume can be used to classify microbes into different species
Genetics
DNA sequences that do not match known samples can indicate the finding of a new species
Lineage
This applies mostly to extinct species, and is used on any bodily remains
Is not the same as genetic classification
E.g. fossils of an extinct snail with a shell similar to that of a modern similarities can be assigned a species name from the same part of the evolutionary tree as existing species
Secretory products
Based on the molecules produced by an organism, which is especially useful in classifying microscopic organisms without easily observable features
E.g. microbes can be classified based on whether they produce carbon dioxide, methane, or hydrogen
What is speciation
Speciation is the process by which a population is separated into two groups that can no longer reproduce together, involving the gradual accumulation of changes in traits
Process of speciation
Speciation may occur abruptly, but most instances of speciation involves gradual change of the allele frequencies within a population over multiple generations
Hence, when one population is evolving from another population, there can be difficulties to distinguish whether the two populations are regarded as the same or different species
One part of the population evolves one way and the other, living with different selection pressures and producing different sets of mutations, evolves in a different way.
The two populations become different enough over time that they can no longer interbreed to produce a fertile offspring.
As a result, a new species has branched off from the previous one, resulting in two species that have a common ancestor.
Sometimes when two populations are in the midst of speciation, the decision whether they are of the same or different species can be arbitrary, and decided during scientific conventions
Case studies for speciation being arbitrary
Case study: African Cichlid Fishes
There are over 200 species of African Cichlid fishes in Lake Victoria, which all appear to have evolved from a single species
Each species has evolved in its own niche and split off from the others, with various species specializing in eating algae, plankton, or snails
However, the split in species would have taken many generations. Even during those generations, the population that started to split off would still interbreed with some success with the original population, though the success rate would diminish as the populations became more different
It is difficult to decide when a speciation occurs, therefore making it arbitrary and subjective
Case Study: Woolly Mammoth
The woolly mammoth appears to share many similar characteristics with Asian elephants, which is why it was originally classified in the same genus.
However, because there are no living wooly mammoths left, it is not possible to know whether they can breed with Asian elephants to produce fertile offspring
The scientific name of the mammoth has since been changed to Mammuthus primigmius without resolving this question, making the decision relatively arbitrary from the perspective of the biological species concept </aside>
significance of chromosome numbers
The number of chromosomes is a characteristic of a species
The number of chromosomes does not necessarily correlate with the apparent complexity of an animal or a plant (e.g. a dog has more chromosomes than humans and chimpanzees.)
diploid cells have an even number of chromosomes.
How many chromosomes do humans and chimpanzees have
Chimpanzees and humans are closely related, although chimpanzees have 48 chromosomes and humans 46.
It is thought that one pair of chromosomes from the ancestor of chimpanzees and humans fused during evolution, leaving humans with 46 chromosomes.
what is a karyogram, which phase of cell division is taken
A karyogram shows the chromosomes of an organism in homologous pairs of decreasing length
Chromosomes are the most obvious in metaphase, giving the clearest view.
karyotype vs karyogram
Karyotype is a property of a cell – the number and type of chromosomes present in the nucleus.
Karyogram is a photograph or diagram showing the chromosomes.
evaluate the hypothesis that chromosome 2 in humans arose from fusion of chromosomes 12 and 13 with a shared primate ancestor
Modern humans have 46 chromosomes.
Gorillas and chimpanzees are the species most closely related to humans.
However, when we prepare a karyogram of the contents of their nuclei, both gorillas and chimpanzees have 48 chromosomes instead of 46.
Two possible hypotheses can be formulated:
1.a complete chromosome disappeared
2.two chromosomes from an earlier common ancestor fused to become a single chromosome.
It is unlikely that an entire chromosome was deleted and disappeared, because removing hundreds of genes in that way would cause a major threat to the viability of the species.
To test the second hypothesis, we can look for evidence, and can start by examining the two characteristics that help identify a chromosome: its shape (position of the centromere) and its banding patterns.
Shape of chromosome
One shape a chromosome can have is the metacentric shape, with the centromere close to the centre.
Chromosomes can also have an acrocentric shape, meaning the centromere is at one end, making one arm of the chromosome much shorter and the other much longer.
All primates have both types.
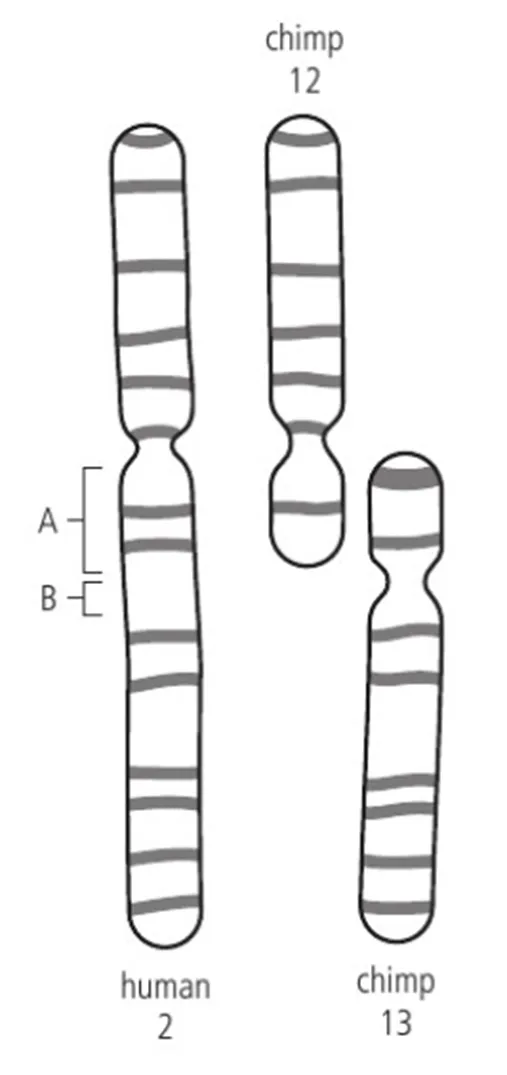
One hypothesis is that chromosome 2 in humans arose from the fusion of chromosomes 12 and 13 in a shared ancestor.
In terms of shape, these two acrocentric non-human chromosomes, when placed end to end, have a similar length to the human chromosome, although some parts overlap.
The position of the centromere in human chromosome 2 lines up with the chimpanzee chromosome 12 but not with chromosome 13.
However, in the zone marked B on the human chromosome, there is satellite DNA (which consists of short repeating sequences of DNA).
This zone corresponds to the position of the centromere in non-human chromosome 13, given creditability to the hypothesis.
Banding of chromosome
In terms of banding patterns, the long arm of chimpanzee chromosome 12 matches that of the short arm of human chromosome 2, and the long arm of chimpanzee chromosome 13 matches the banding patterns of the long arm of human chromosome 2.
Presence of telomeric DNA
Besides shape and banding patterns, other evidence to support the idea of fusion is the presence of telomeric DNA in the centre of human chromosome 2.
The telomeres are caps at the tips of chromosomes that contain repeating sequences of DNA and provide protection, the same way that bumpers protect cars and aglets protect the ends of shoelaces.
Such repeating telomeric DNA is not supposed to be in the centre of chromosomes, only at the tips.
And yet, at position A in the human chromosome 2 shown in Figure 3, telomeric DNA is present at the position where the two chromosomes would have fused.
It is very important to understand that this evidence does not say we descended from chimpanzees.
The fusion of the chromosomes would have happened after the speciation split of a common ancestor that led to the evolution of chimpanzees on one branch of the tree of life and the evolution of humans on another branch.
What is the genome
how do species affect differences in genome
all the genetic information stored in DNA that is present in an individual, which includes the summation of all the genes present in the individual
organisms in same species share most of their genome but variations such as single nucleotide polymorphisms give some diversity
what are single nucleotide polymorphisms
differences in single nucleotide base sequences in genes
functionality of single nucleotide polymorphisms
Only about 5% of SNPs are functional, meaning that they actually produce a difference in a person’s body.
Most are neutral, meaning that they will not affect a person’s phenotype (physical expression of a gene e.g blood type).
significance of the human genome project
1.identification of all human genes.
2.discover protein structures and functions.
3.find evidence for evolutionary relationships.
4.find mutations e.g. base substitutions or single nucleotide polymorphisms.
5.find genes causing diseases.
6.develop new drugs based on base sequences (new gene therapies)
7.tailor medication to individual genetic variation (pharmacogenomics)
promote international co-operation
what is the genetic code
the correspondence between each of the 64 codons and the amino acids into which they are translated.
what variation exists between eukaryotic genomes
Overall size - determined by total amount of DNA
Sequences of the bases - variation in sequences are larger between species than within species
Genes can vary in length
what is junk dna
it is important to note that the genomes of organisms are not completely made up of regions that encode for genes
Many of such regions are known as “junk DNA”, and do not code for any genes, or have any regulatory functions, and “junk DNA” may consist of repeated sequences
significance of mitochondrial DNA in comparing genetic diversity
All eukaryotes have mitochondria, and the way mitochondrial DNA (present only in the egg, not in the sperm cell) is passed down from mother to offspring, means there is not the shuffling and mixing that we see in chromosomal DNA.
It is estimated that, within a species, roughly 1 in 1,000 of the genetic code letters is different between individuals' mitochondrial DNA.
To see differences between individuals within a species, or to see differences between species, it is possible to look up the amino acid sequences for a particular gene in a database and match them to see if there are amino acids missing, added or modified.
Kbp and Mbp unit
kilo base pairs (thousand base pairs)
mega base pairs (million base pairs)
why is amount of DNA not a reliable indicator of complexity
The proportion of DNA that acts functional gene varies.
The amount of gene duplication varies between organisms.
what is phylogenetics
compare whole genome sequences to see how organisms are related to each other
why is whole genome sequencing more feasible now
increased speed
decreased cost
current uses and future uses of genome sequencing
current use: research into evolutionary relationships
future use: personalized medicine, sometimes called precision medicine: information about a person's genetic makeup can be applied to an individual when prescribing treatments.
Personalized medicine is better adapted for diseases that are dynamic (e.g. cancer) and require different treatments at different stages of the illness.
what are xenologs
When sequencing and matching genes, sometimes a sequence of DNA is found that has more in common with another species than the one it is found in.
Such genes, known as xenologs or jumping genes, travel in plasmids from one bacterium to another.
how do xenologs challenge biological species concept
Sometimes we find the same identical gene in several very different species of bacteria.
Usually when we look at similar DNA sequences, we think that the organisms shares a common ancestry. But sometimes, the host species are on different branches of the tree of life and do not share the same lineages, e.g. yeast cells (a fungus) containing bacterial DNA.
Such examples challenge the concept of species because it questions the idea organism classified in the same genus or species have a common ancestor.
case study of why cross breeding between closely related species is unlikely to produce fertile offspring if parent chromosome numbers are different
A female horse (64) and a male donkey (62) can mate and produce a mule (63).
A mule having 63 chromosomes makes it difficult for homologous pairs of chromosomes to match up during meiosis, and thus production of gametes can be difficult.
Because the offspring (the mule) are not fertile, no new species has been created.
Instead, a mule is called an interspecific hybrid.
Hybrids face several challenges to continue as a population.
The vast majority of hybrids are infertile.
Even if one generation of hybrids is produced, a second generation is highly unlikely.
This presents a genetic barrier between species.
limitations of dichotomous keys
Keys may use technical terms that only an expert would understand.
It is possible that there may not be a key feature for the type of organism to be identified.
Some features of organisms cannot be easily established in the field e.g. whether or not an animal has a placenta; whether an animal is endothermic or ectothermic (warm- or cold-blooded).
Some organisms significantly change their body shape during their lifetime (e.g. frogs have an aquatic tadpole juvenile form which is very different from the adult), which keys must take into account.
Many insects, e.g. show difference between males and females of the species, which can cause difficulties when identifying species.
cannot be applied to microbes which require microscopes
Unidentified plants that are not flowering or trees without leaves can be hard to identify
what is a DNA barcode
A DNA barcode is a short sequence of DNA (several hundred base pairs) inside an organism's cells that can be used to quickly identify the species.
what DNA is often used to identify animals or prokaryotes
mitochondrial DNA for animals
ribosomal DNA for prokaryotes
what is environmental DNA and why is it present
The DNA extracted from the water or soil is sequenced and barcodes are isolated for analysis.
Such DNA collected from the environment rather than from an organism is called environmental DNA or eDNA.
It is present in the environment because organisms release dead cells, produce faeces or die and start to decay.
bioindicator species and example
organisms are so sensitive to certain types of pollution that their presence in an ecosystem indicates a lack of pollution.
Conversely, their sudden disappearance from an ecosystem would suggest the appearance of a source of pollution.
e.g. Caddisfly larva live underwater in streams and can be used as bioindicators for water health. A large caddisfly population is reassuring, but a small or disappearing caddisfly population is a signal to investigators that they should start looking for potential contamination upstream.
how to perform and read results of ecological survey
To perform an ecological survey, one species at a time is identified and counted, and often (e.g. insects) this requires capturing them, which can disrupt the population.
With eDNA metabarcoding, a single sample can be sequenced for dozens or hundreds of species without having to capture individual organisms.
This is more time efficient for the specialists carrying out ecological surveys.
Over months and years, if the number of species in an ecosystem decreases, we know that the diversity of that habitat is unhealthy or declining.
In contrast, if the number of species is stable or increases over time, we know that biodiversity is stable or improving, suggesting that the ecosystem is healthy and flourishing.
advantages of DNA barcoding
This is a way to monitor the presence of species in an environmental without introducing the stress of the presence of human observers (or cameras) that may drive certain species away
The speed and ease of use of the method allows for constant monitoring of the species in the location with multiple samples taken at regular intervals
The method is sensitive to detect the presence of rare species, or species too small to observe easily
disadvantage of using eDNA to track species
Firstly, it only gives an indication or the presence or absence of a species, not the population size.
Secondly, the DNA does not indicate if it is from a living organism or a dead one.
Thirdly, certain chemical incompatibilities exist with the processing of soil samples because substances in the soil can interfere with the sequencing process, giving rise to inaccurate results.