M3L3 - Biomembranes and Cell Architecture
1/20
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
21 Terms
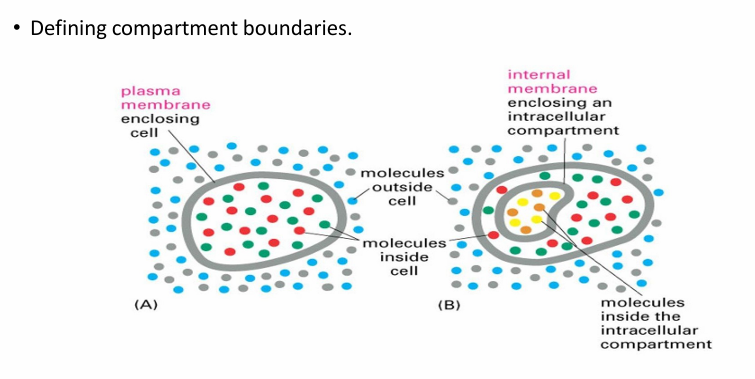
Biomembranes Importance
They define boundaries in a cell by seperating interior and exterior
Creatures a unique internal env
They also define internal microenvs by surrounding organelles
Biomembranes Characteristics / Functions
Selectively permeable: Few molecules can freely travel across
They contain proteins for cell signaling (receptors) and adhesion to other cells or the environment.
Membranes are flexible and dynamic, letting them change shape for processes like movement and cell division.
Biomembrane Terminology: Cells and Organelles
Cells
Exoplasmic Face (Faces outside of cell)
Cytosolic Face (Faces inside the cell)
Organelles
Cytosolic Face (Faces the cytosol of cell)
Lumenal Face (Faces inside the organelle)
Intermembrane space
Space between 2 membranes inside organelle
Mitochondria has this
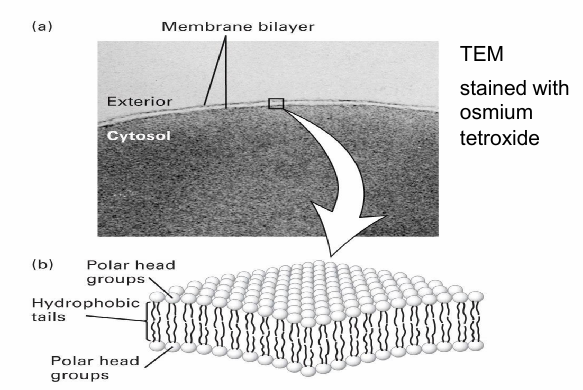
Bilayer Structure of Biomembranes
Visualized using TEM
2 thin parallel lines seen at cell surface indicates bilayer structure
Row of polar groups facing inside/outside
Hydrophobic tail (np) and polar headgroup (hydrophilic)
Hydrophobic core as tails face eachother in (aq) cell env
Phospholipids
Amphipathic molecule (hydrophilic/phobic)
Spontaneously arrange in (aq) solution to form a micelle
bubble-like structure
Produced when single sheet of phopholipids assemble
Hydrophilic wall and hydrophobic core
At higher [phospholipids], they spontaneously assemble to form bilayer
![<ul><li><p>Amphipathic molecule (hydrophilic/phobic) </p></li><li><p>Spontaneously arrange in (aq) solution to form a micelle</p><ul><li><p>bubble-like structure</p></li><li><p>Produced when single sheet of phopholipids assemble </p></li><li><p>Hydrophilic wall and hydrophobic core </p></li></ul></li><li><p>At higher [phospholipids], they spontaneously assemble to form bilayer </p></li></ul><p></p>](https://knowt-user-attachments.s3.amazonaws.com/32bea08b-d456-4194-9582-91c73d52ebec.png)
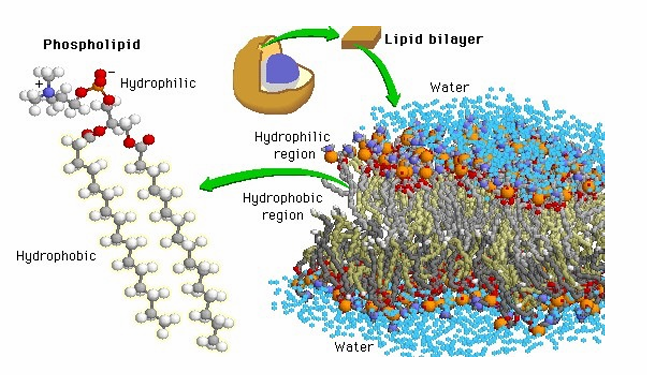
Chemical Makeup of Phospholipid
A diglyceride has two fatty acids (long hydrocarbon chains with a carboxyl group) linked to glycerol.
A phosphate group attaches to the third –OH of glycerol, forming a phospholipid, which often has additional charged groups on the phosphate.
The fatty acid tails are hydrophobic (water-insoluble), while the phosphate head is hydrophilic (water-soluble).
Because of this, phospholipids spontaneously form bilayers in water — their hydrophobic cores face inward and hydrophilic surfaces face outward, creating the lowest free-energy configuration.
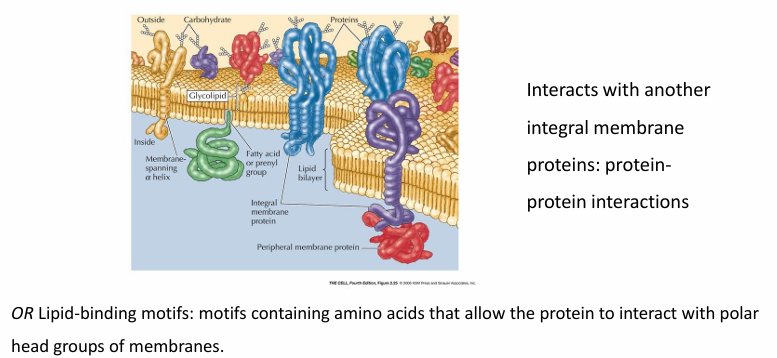
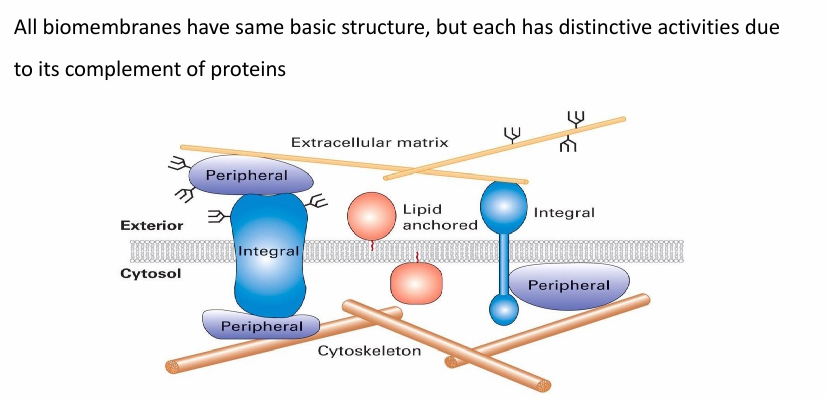
Proteins associated with bilayer
Integral membrane proteins (embedded in hydrophobic core)
Lipid-anchored and peripheral membrane proteins
Associated with one surface of bilayer
The proteins associated determine the functinos of a membrane
Some membranes are dense w proteins
Inner mitochondrial membrane with 76% protein composition
Some have very few proteins
Myeline membrane with 18% protein composition
Why is it good that the membrane is dynamic and fluid?
Allows biomembranes to
fuse with one another
deform without tearing
change in shape to accompany any cell movement (ex cell division)
Proteins and phospholipids can move laterally through the membrane
Proteins can cluster in membrane areas called microdomains
Helps perform specific cellular activity
They can disperse after cell activity is complete
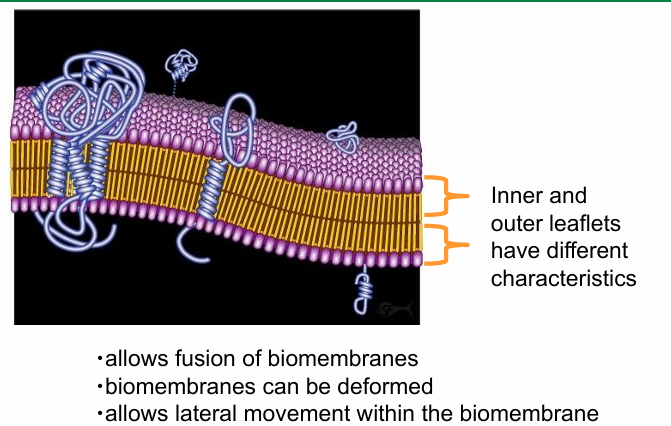
Fluid Mosaic Model
Proposed by SJ singer and G nicolson
Describes structural features of biomembranes
Defined as fluid bc
membrane components more laterally or sidewars throughout
Not solid, more fluid
Defined as mosaic bc
made of many different kinds of macromolecules
Fluidity is seen in both the outer and inner leaflets of membrane bilayer
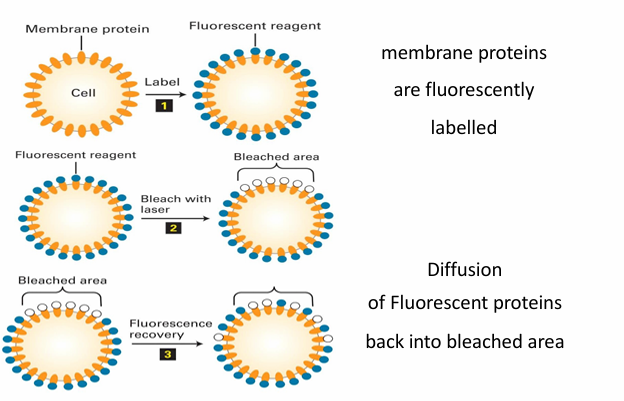
FRAP in determining membrane fluidity
Microscopy labelling technique: Fluorescence recovery after photobleaching
Allows to track and measyre the fluidity of proteins in membrane
Mechanism
Protein can be tagged by adding an antibody or fusion to GFP
The fluorescent molecule can be damaged by exposure to too much light
They then become bleached and no longer fluoresce
Experiment
At start, the proteins are evenly placed in membrane so fluoresce evenly
A small patch is then bleached by a laser
Depending on membrane fluididt, the patch can be recovered
The molecules dont regain ability to fluoresce but they just moved to disperse
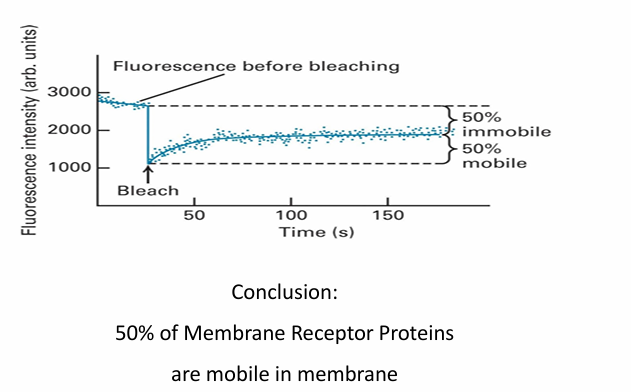
FRAP: Calculating Fluidity
y-axis: Fluorescence in arbitrary units
x-axis: Time
Initial fluorescence patch is 3000 units
Bleaching step reduces to 1000 units
Overtime, fluorescence inc as fluorescennt molecules move in while bleached ones move out
At 50s, the fluorescence of patch reaches 2000 units
about 50% of initial compared to when bleached
Suggests half proteins are mobile and able to move laterally but other half are immobile
Other experiments, proteins may be less mobile or not at all (less fluid membrane)
If very fluid, it can go back up to 100%
Regulating Membrane Fluidity: Lipid Composition
Hydrophobic fatty acid tails can be saturated (no C=C bonds)
Can pack together more closely, dec fluidity
They can be unsaturated (many C=C bonds)
Kinked so more fluid
Long fatty acid chains (18C)
Can pack together tightly to dec fluidity
Presence of cholesterol
dec fluidity to maintain integrity
without it, membrane would be too fluid and permeable
At high [cholesterol], it helps seperate phospholipids so the fatty acid chains don’t come together and crystalize
Good for hibernating animals
Thus, it prevents extermes in mebrane fluidity (either too fluid or too gel-like)
![<ul><li><p>Hydrophobic fatty acid tails can be saturated (no C=C bonds) </p><ul><li><p>Can pack together more closely, dec fluidity </p></li></ul></li><li><p>They can be unsaturated (many C=C bonds) </p><ul><li><p>Kinked so more fluid </p></li></ul></li><li><p>Long fatty acid chains (18C) </p><ul><li><p>Can pack together tightly to dec fluidity </p></li></ul></li><li><p>Presence of cholesterol </p><ul><li><p>dec fluidity to maintain integrity </p></li><li><p>without it, membrane would be too fluid and permeable </p></li><li><p>At high [cholesterol], it helps seperate phospholipids so the fatty acid chains don’t come together and crystalize </p></li><li><p>Good for hibernating animals </p></li><li><p>Thus, it prevents extermes in mebrane fluidity (either too fluid or too gel-like) </p></li></ul></li></ul><p></p>](https://knowt-user-attachments.s3.amazonaws.com/28bee6e7-335f-4ec9-bbf3-9420515ce6ea.png)
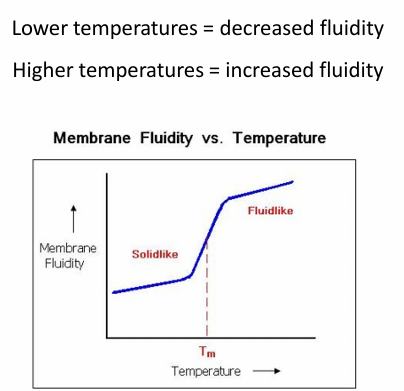
Regulating Membrane Fluidity: Temperature
Low temp = dec fluidity
high temp = more fluid
Cells respond to temp by altering membrane makeup
High temp = add cholesterol to dec fluidity
Ex. Bacteria
Respond to low temp by cleaving fatty acid chains from 18C to 16C
OR
They activate desaturase enzyme to introduce C=C
Ex. Cold-tolerant plants
Have greater % of unsaturated fatty acid chains in phospholipids to prep for lower temp
Ex. Cold-blooded animals
incorporate more cholesterol into membrane in response to cold
Lipid Rafts
Demonstrates difference in fluidity within subregions of membrane
They’re a little taller than the rest of membrane due to longer fatty acid chain
There’s higher [cholesterol] in this region
These factors make the membrane in that area less fluid
The phospholipids/proteins are less mobile within this region
The raft as a unit is mobile within the surroundings
It also doesn’t float on top of the membrane, but has 2 leaflets so it’s a part of it
Movement between leaflets
Difficult to move proteins or phospholipids from one leaflet to another
Phopholipid bc the hydrophilic head would have to go through the hydrophobic core
Thus proteins and phospholipds are places in correct orientation during synthesis
Enzyme flippases are capable of helping the flipping of phospholipids if needed
It’s possible but requires a lot of energy as well as the enzyme
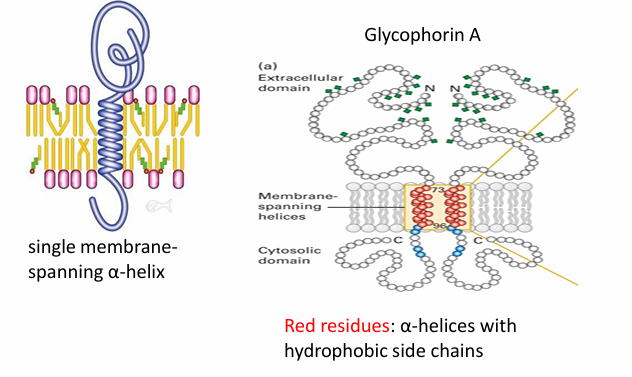
Integral Proteins: Single Pass
Leaves domains on both exterior and interior surfaces of membrane
Ex. glycophorin A found in human RBC
It’s a homodimer
(red) amino acids make up the hydrophobic alpha helix spanning the hydrophobic core
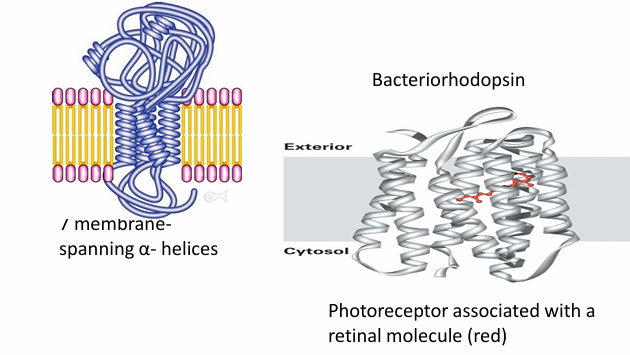
Integral Proteins: Multi Pass
Pass through membranes many different time
Ex. Bacteriorhodopsin
Uses 7 membrane spanning domain with 7 alpha helices
They interact to form a transmembrane domain
Ion channels have this structure
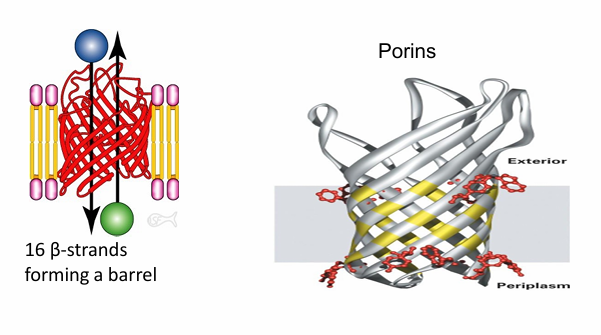
Integral Proteins: Beta Barrel
Ex. Formed from 16 beta strands
Exterior is hydrophobic so it can interact with hydrophobic membrane
The interior is hydrophilic
Forms hydrophilic pre through the hydrophobic membrance
Ex. porins found in bacteria cells, chloroplasts and mitochondria
Integral Proteins: Collection of Alpha Helices
Shown in different colours in image
The exterior is hydrophobic while the interior is hydrophilic
This is an aquaporin that creates hydrophilic channel for water movement across cells during osmosis
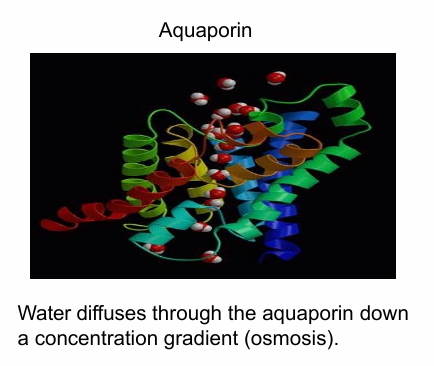

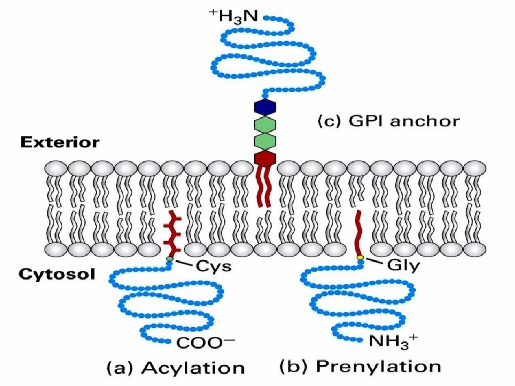
Lipid-Anchored Proteins
Associated with one leaflet by covalently attached lipid modifications
During synthesis, they’re modified by acetylation
14C or 16C long acyl chains
Attaches lipid anchor to the N-terminus
And also prenylation
15C OR 20C long unsaturated chains
Adds lipis anchor to C-terminus
Some lipid-anchored proteins have a structure called GPI anchor that forms hydrophobic anchor
enables the association of a protein to membrane
Peripheral Protein
Peripheral proteins interact with membrane-embedded or anchored proteins, attaching indirectly to the membrane.
Lipid-binding motifs let them bind to the polar head groups on the membrane surface.
They often reversibly attach or detach through reversible modifications like phosphorylation or allosteric structural changes.