Lecture 9 - Hybridization
1/44
Earn XP
Description and Tags
These flashcards cover key concepts from Southern blotting and microarray techniques, based on the lecture notes.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
45 Terms
What is Southern blotting used for?
To study purified DNA digested with restriction enzyme(s), which requires high-quality DNA; also used to identify genes, mutations, or DNA fingerprinting.
What are the basics of Southern blot?
Digested DNA fragments are separated by gel electrophoresis
The DNA fragments are transferred out of the gel onto a solid membrane (matrix)
The membrane is exposed to a DNA probe labeled with a radioactive, fluorescent or chemical tag
The tag allows any DNA fragments containing complementary sequences with the DNA probe to be visualized (detected).
What is the steps in the Southern blotting process?
Cut DNA with restriction enzymes.
Separate the DNA fragments by gel electrophoresis.
Transfer DNA to membrane (blotting).
Add labeled DNA probe.
Detect the target fragments.
What are the applications of Southern blot?
Detection of various DNA abnormalities such as deletions/insertions
Point mutations/polymorphisms
Structural rearrangements
Southern blot application examples:
In human tumors, changes in the number of copies or the structure of cancer-causing genes are frequent
Can be used to investigate whether a gene is amplified, deleted, or structurally rearranged in cancer cells as compared to normal cells
Although it is quite labor-intensive, it is particularly useful for detecting large deletions in tumor genomes
How is Southern blot used in Fragile X diagnosis?
Used to detect CGG repeat expansions in the FMR1 gene
Identifies methylation status and full mutations (>200 CGG repeats)
Differentiates normal, premutation, and full mutation alleles
Essential for diagnosing Fragile X Syndrome and assessing carrier status
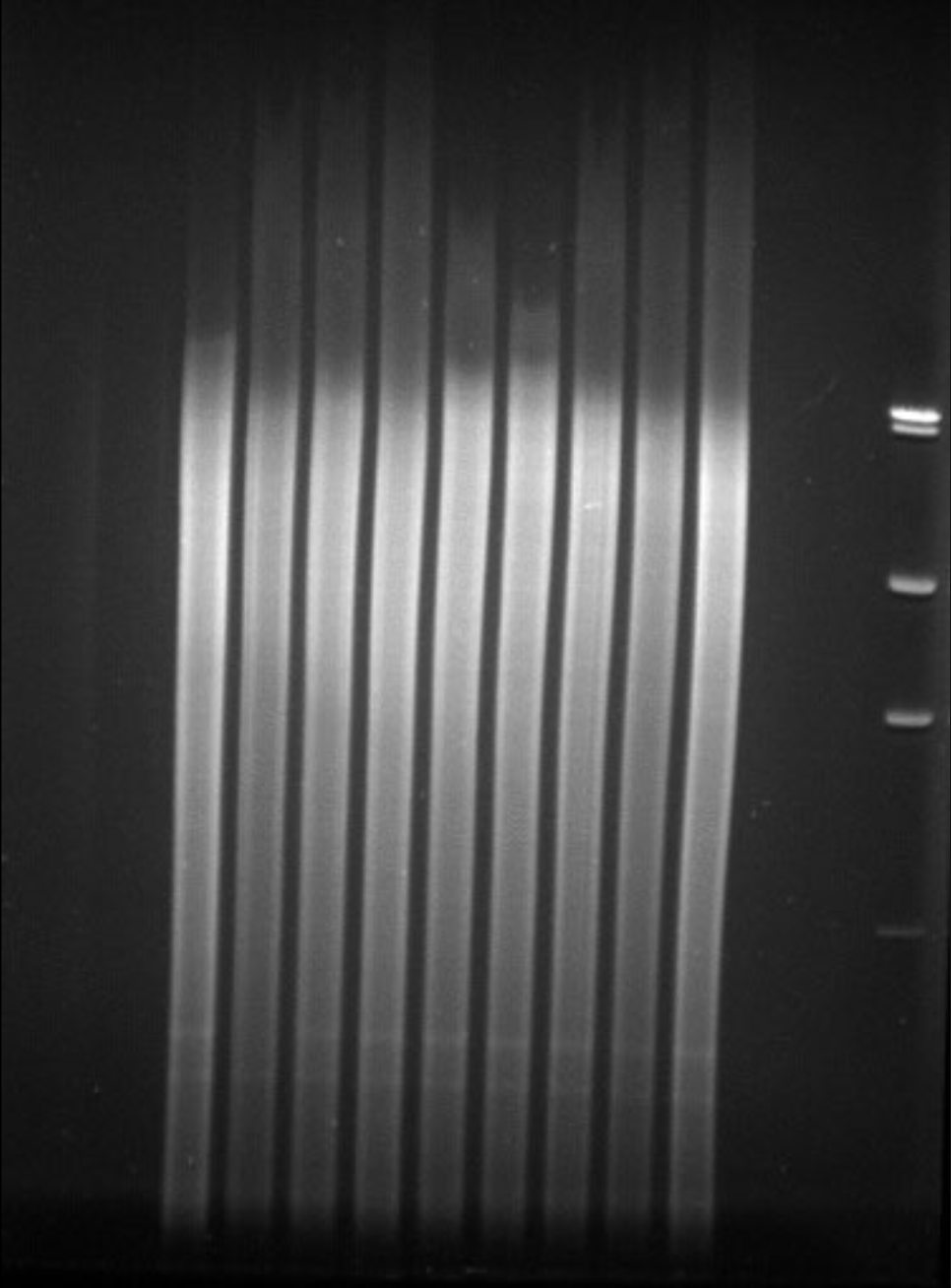
Southern blot pre-transfer gel photo
Represents endonuclease-digested genomic DNA of high quality and concentration, which has been electrophoresed through an agarose gel. After ethidium bromide staining and exposure to UV light, the DNA in each lane and the ladder/marker are bright and clearly seen.

What are the pre-transfer steps in Southern blot?
Completed prior to capillary transfer of DNA to the membrane (blot)
Prepares the DNA to move out of the gel
Reduces bulkiness
Gel is bathed in acid, then base
Depurination
Breaks bonds with purines using a weak acid (HCl)
Promotes higher efficiency transfer of HMW DNA
Denature
Produces single-stranded DNA using a strong base (NaOH)
What occurs during the transfer step in Southern blot?
Agarose gel is flimsy, difficult to work with, and only a temporary matrix used for separation of DNA
DNA is transferred to the membrane via capillary or vacuum methods
ssDNA strongly binds to positively charged nitrocellulose membranes
Irreversible hydrophobic and electrostatic interactions occur
What are the two methods of transfer we discussed in class?
Capillary transfer method and vacuum transfer method
What is the capillary transfer method?
High salt transfer buffer is added to the reservoir/tray
The membrane is placed directly on the gel
Trapped bubbles can interfere with transfer
Buffer will move by capillary action from the reservoir to the dry material on the top of the gel
DNA will be carried along
Usually transfers overnight
What is the vacuum transfer method?
Uses suction to move the DNA from the gel to the membrane in a recirculating buffer bath
Transfers DNA more rapidly than capillary transfer, usually 2-3 hours
No issues with trapped air bubbles causing discontinuous transfer
Equipment can be expensive and needs maintenance
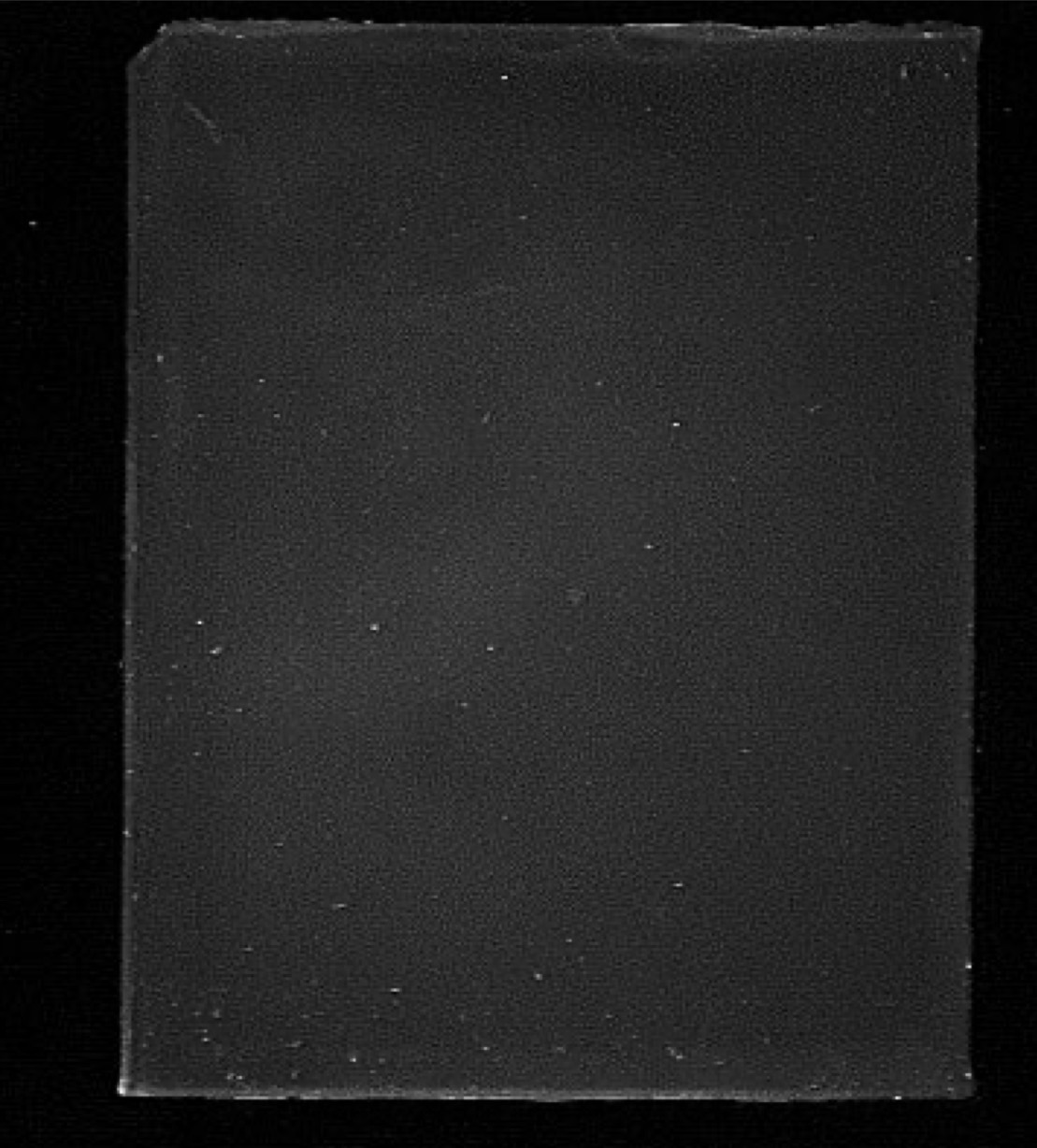
Southern blot post-transfer gel photo
Shows no DNA in the agarose gel following transfer to the membrane, after ethidium bromide staining and exposure to UVA’s light; represents a successful transfer with high efficiency. All DNA is now on the membrane.

How is DNA fixed to the membrane?
After transfer, the empty agarose gel is discarded
DNA is then permanently fixed (immobilized) to the membrane with covalent attachment to prevent loss during washes
Baking at 80° C
Cross-linking DNA with UV light
What are the steps for pre-hybridization in Southern blot?
Membrane must be prepared before hybridization with a probe
Membrane is incubated in pre-hybridization (pre-hyb) buffer, which includes an aqueous mixture of detergent and blocking reagents (DNA and proteins)
Critical step to help prevent non-specific hybridization
During Southern Blot, what does incubating the membrane in pre-hyb buffer achieve?
Helps block non-specific binding
What pre-hyb buffer blocking reagents are used in Southern Blot, and what do they block?
Herring or salmon sperm DNA is most often used
Blocks DNA to prevents non-specific binding of nucleic acid probe
Milk protein and Denhardt’s solution (albumin and Ficoll)
Blocks protein to reduce background interference
Southern Blot Hybridization
Annealing/binding of ssDNA or RNA probe with ssDNA or RNA target
Formation of complementary base pairs
Goal is to optimize specific target:probe binding while minimizing non-specific binding
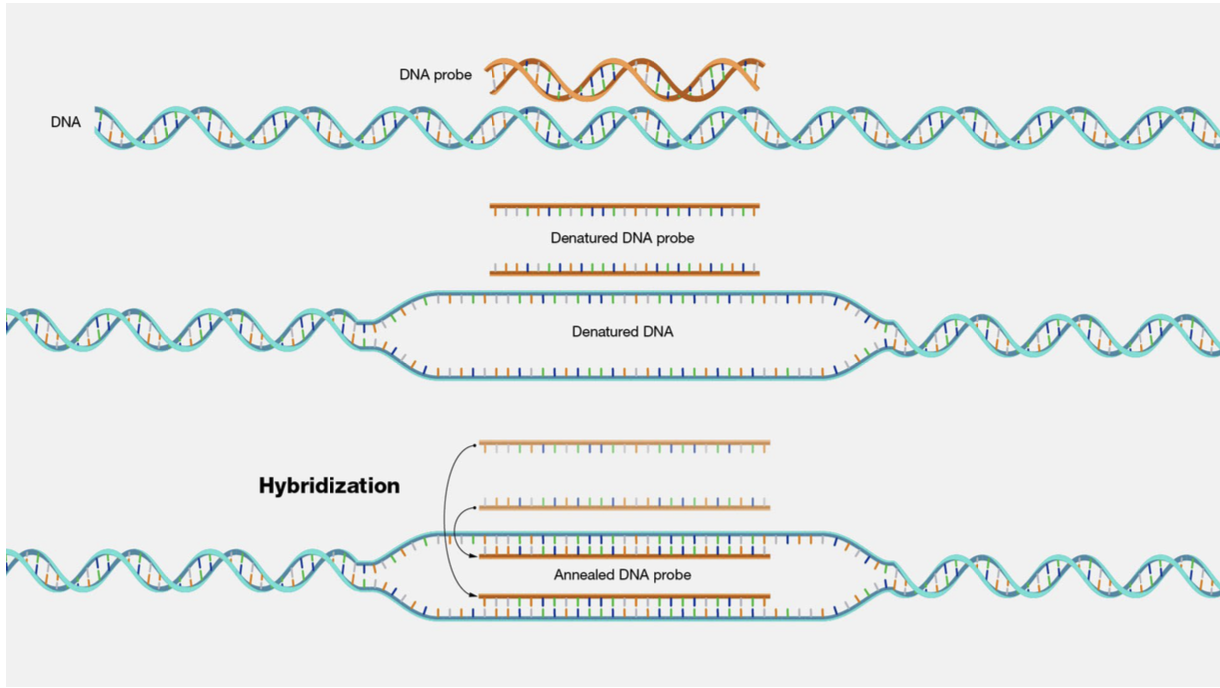
Why must probes be carefully designed for optimal hybridization?
To ensure specificity of the results
What techniques are used to produce probes?
PCR, cloning, or rapid synthesis; synthetic probes are immediately ready upon production
Probes in Southern Blot
Probes are labeled to attach a detector molecule
Large (long) probes are most often used in S blot
Sequences are rare or unique in the genome
Prevents/reduces cross-hybridization
Produces low background with clear results
Vectors used in cloning
Plasmid – up to 10 kb insert
BAC – 100-150 kb insert
YAC – up to 1000 kb insert
What are the three types of probe labeling mentioned in lecture?
End-labeling, nick translation, and random prime labeling
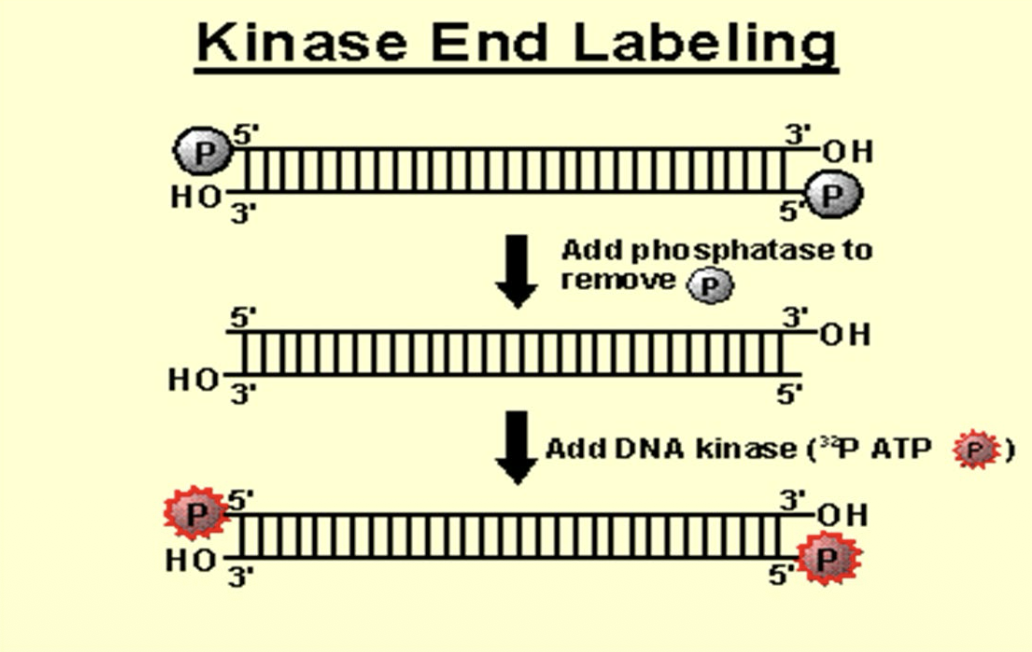
What is end labeling?
Labeling of 5´ ends of small, linear fragment (PCR amplicon)
Only one marked residue per probe molecule is formed, so activity of the label is lower than in other methods (less intense signal)
Uses phosphatase and kinase enzymes
Phosphodiester bonds are formed
DNA is not denatured
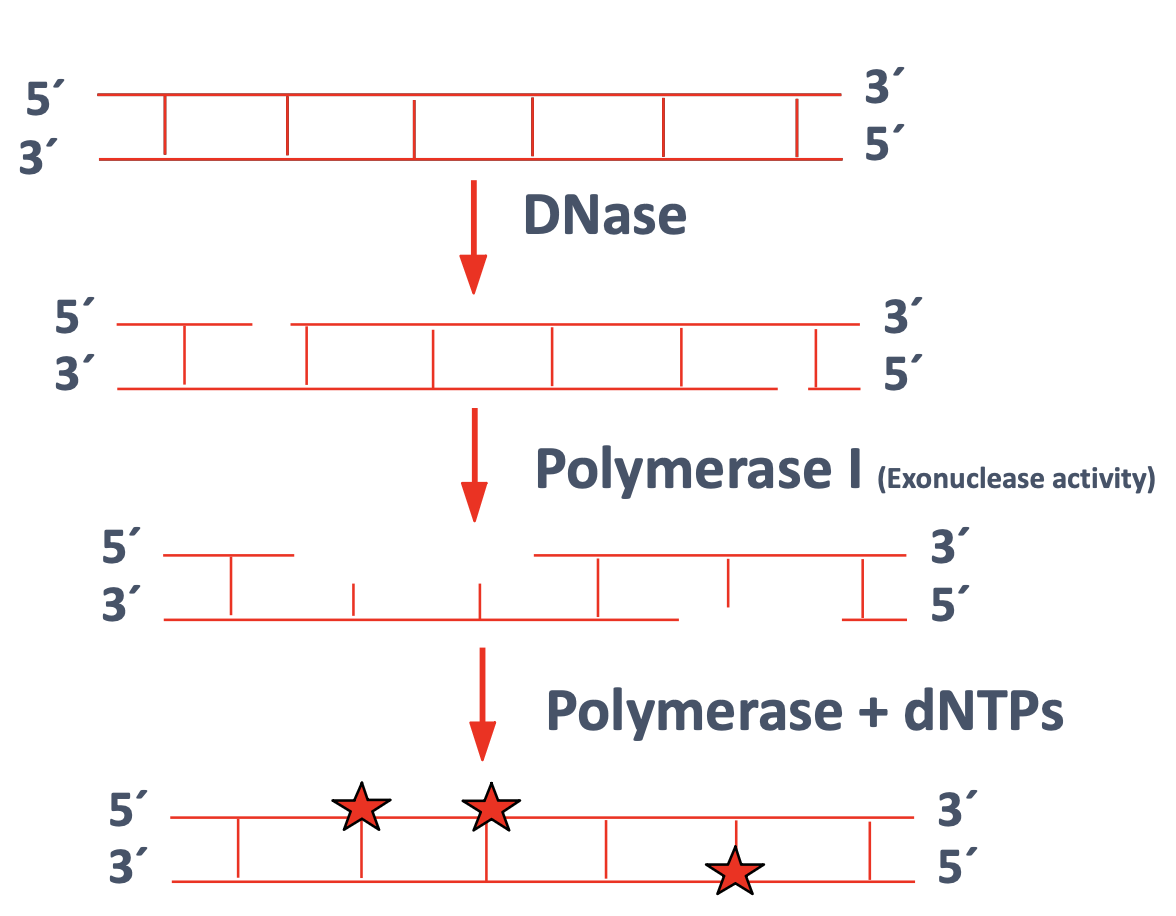
What is nick translation?
Relies on the action of DNase and DNA Polymerase I enzymes
Under limiting conditions, DNase produces nicks
Physical nicking through vortexing is another option sometimes used
Polymerase I acts as an exonuclease to remove nucleotides at the nick sites as well as a polymerase to replace nucleotides, some of which are labeled
DNA is not denatured
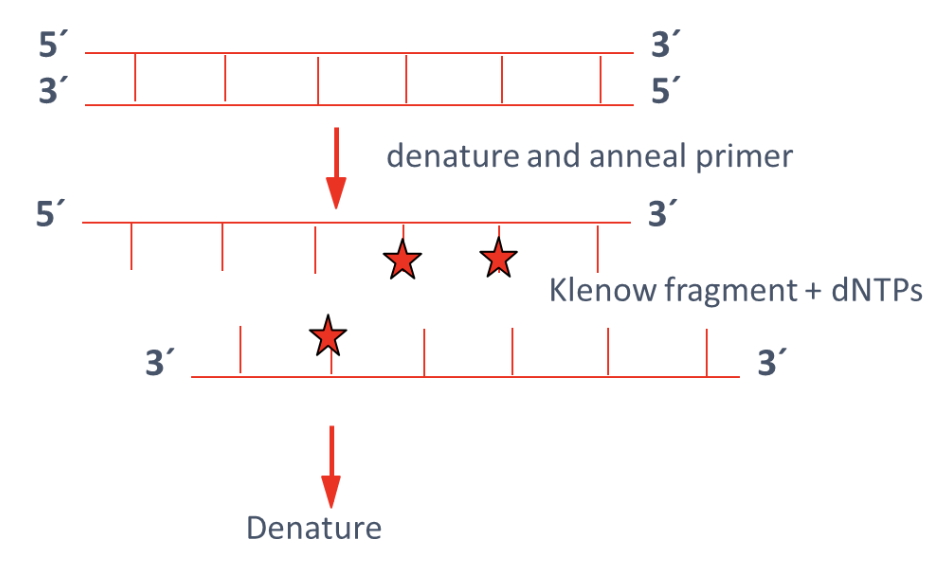
What is random prime labeling?
DNA is denatured and then annealed to random primers in a cycling procedure
Modified Klenow fragment of DNA Polymerase I lacks exonuclease activity
Klenow fragment extends primers while incorporating labeled nucleotides
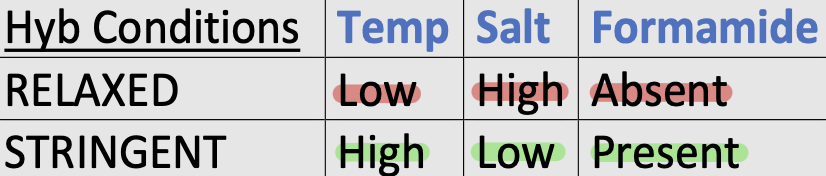
What is (hybridization) stringency?
The hybridization conditions that control the specificity of binding between two single-strand sequences; usually probe and immobilized target sequence
What can exist at conditions of high stringency?
Only duplexes with near perfect sequence complementarity
What happens if stringency conditions are too low?
The probe will bind unrelated target sequences which is non-specific hybridization; analogous to non-specific priming (annealing) in PCR
What is formamide?
A chemical denaturant often used in S blot hybridization buffers
Stringent vs. relaxed hybridization conditions
Under stringent conditions, only perfect complementary probes will hybridize to specific target
High stringency reduces non-specific hybridization
Radioactive (isotopic) label detection
Photographic exposure to x-ray film
Non-radioactive label detection
Chemiluminescence, photographic exposure to x-ray film
Colorimetric reactions, direct visualization with visible light
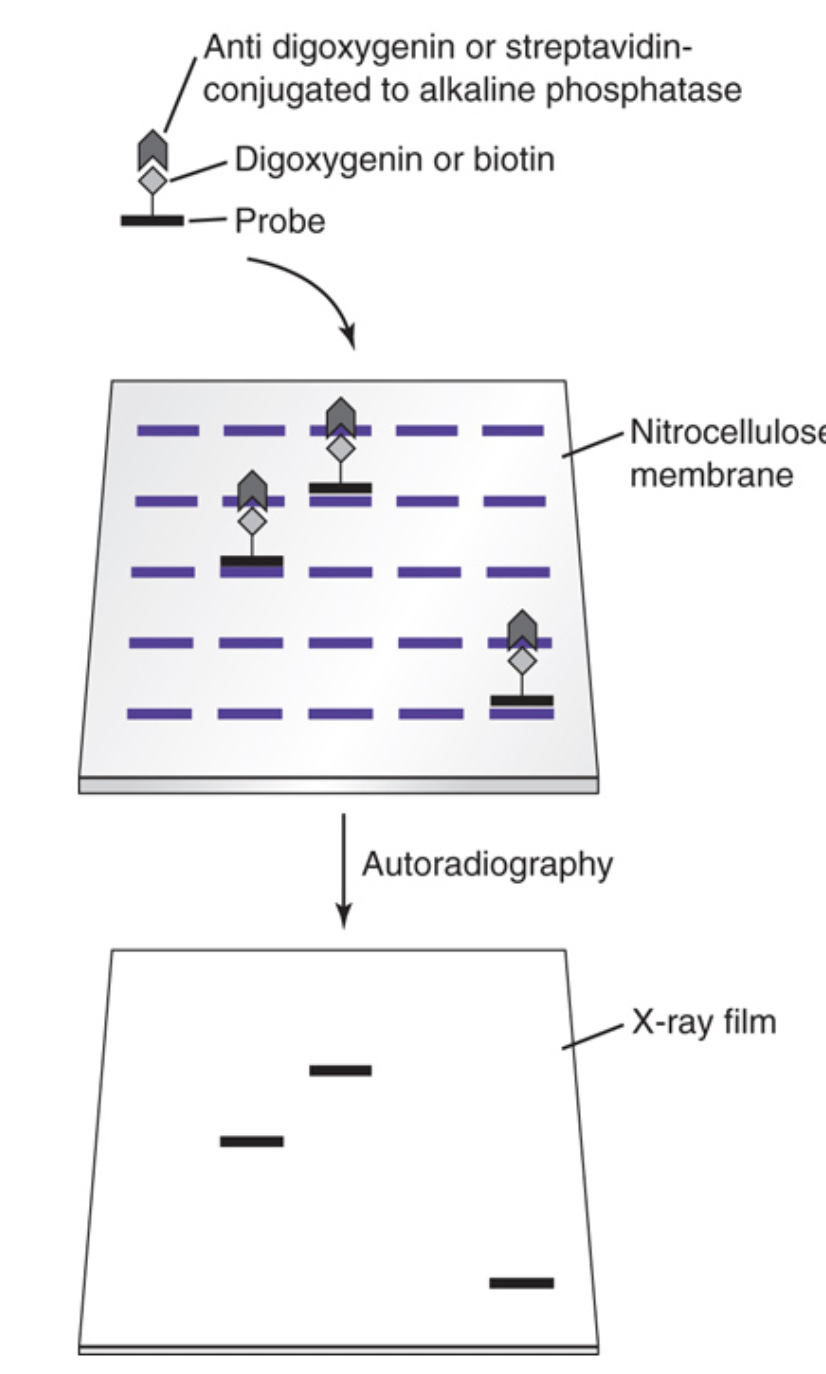
Anti-digoxigenin antibody or streptavidin conjugated to alkaline phosphatase (AP conjugate) is added to the reaction mix to bind the digoxigenin- or biotin-labeled probe:target complex
Steps of indirect nonradioactive label detection
The probe is covalently attached to digoxigenin or biotin
After hybridization, the probe is bound by antibodies to digoxigenin or streptavidin conjugated to alkaline phosphatase (AP)
This complex is exposed to color- or light-producing substrates of AP, producing color on the membrane or light detected with autoradiography film
Signal-to-noise ratio
A successful nucleic acid hybridization experiment shows a high signal-to-noise ratio
A specific, robust signal is generated
In some protocols, specificity of detection may be sacrificed to get a stronger signal
Controlled with stringency conditions
What is used to confirm results of a microarray experiment?
Quantitative PCR (qPCR)
What is FISH?
Fluorescence In Situ Hybridization, a molecular cytogenetic technique that uses fluorescent DNA probes to detect specific chromosomal locations.
What are common applications of Southern blotting?
Detection of DNA abnormalities, point mutations, deletions/insertions, and structural rearrangements in cancer.
What is the purpose of the blocking reagents in pre-hybridization?
To prevent non-specific binding during the hybridization process.
What is the difference between capillary and vacuum transfer methods in Southern blotting?
Capillary transfer is slower and uses capillary action, while vacuum transfer is faster, typically taking 2-3 hours, and uses suction.
What does a successful Southern blot analysis indicate about signal-to-noise ratio?
It indicates a high signal-to-noise ratio with robust, specific signals generated.
What fluorescent signals indicate in a microarray experiment when comparing tumor sample mRNA to normal control?
Green indicates more mRNA in tumor, red indicates more in normal control, yellow indicates the same expression level.
What are dual color break-apart probes used for in FISH analysis?
To detect translocations by binding to intact chromosomes flanking the translocation breakpoint.
What does the HER2 gene amplification indicate in some breast cancers?
It indicates the presence of amplified HER2, which can influence treatment decisions.
What is photobleaching?
Quenching or fading of the fluorescent signal occurs when FISH signals are exposed to light; involves photochemical destruction of the fluorophore molecules