Genetics Practice Tests
1/19
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
20 Terms
Briefly describe the structure of lac operon and comment on the following diagram including the concepts about repressed transcription, basal and high level of transcription in situations keeping in view consumption of glucose and lactose in a bacterial cell.
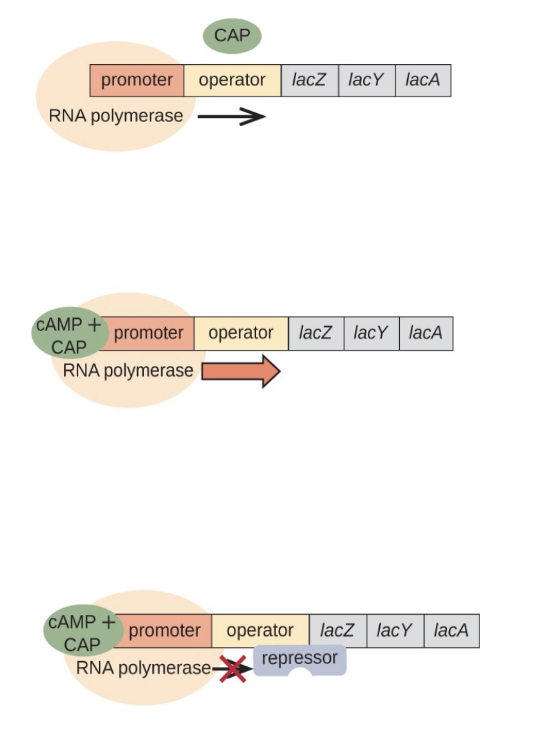
lac operon is a multi-gene sequence controlled by a singular promoter
codes for enzymes and proteins that aid in brining lactose into the cell (lac y → lac permease) and breaking down lactose (lac z → b galactosidase)
repressed when there is no lactose to break down
allolactose is allosteric regulator
CAP is not bound and the repressor is not bound
leads to basal levels of transcription
occurs when lactose and glucose are both present
cell preferentially uses glucose
CAP promotes the binding of RNA pol. but requires cAMP to bind
repressor binds to allolactose, blocking its ability to bind to the operator
CAP is bound and the repressor is not bound
leads to high levels of transcription
occurs when lactose, but not glucose, is present
cAMP is produced so CAP binds
allolactose prevents the repressor from binding, allowing RNA pol. to transcribe
CAP and repressor are bound
leads to repressed transcription
occurs when neither lactose nor glucose are present
cAMP is produced so CAP binds
there is no allolactose present so the repressor is free to bind to the operator
blocks RNA pol. from transcribing
Describe the purpose and principle of Electrophoretic Mobility Shift Assay?
studies the interactions between proteins and DNA
three samples
labelled probe (DNA)
labelled probe + protein of interest
labelled probe + protein + specific antibody
the samples are run on the gel and if the protein and DNA interact, they weigh more and don’t travel as far through the gel
(A) Dr. Song found a 15-nucleotide sequence located 300 bp upstream of gene X in cardiomyocytes. She suspects the sequence to be an enhancer and did EMSA with known cardiomyocyte specific transcription factors A and B. Observe the following radiographs of EMSA. Was it possible for her to validate the 15 bp sequence as ‘actual’ enhancer (Yes/No)? Comment on the gel images in detail keeping in view sequence-specific binding of specific transcription factors on the putative enhancer.
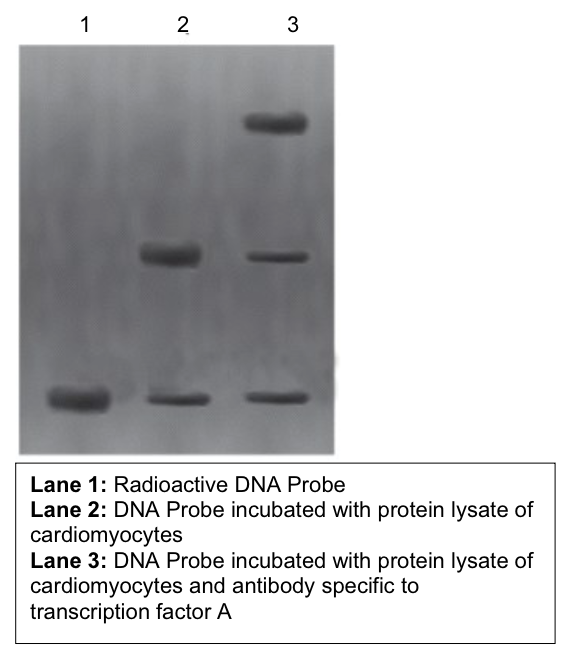
no we cannot determine that the 15 nt sequence is an enhancer
the gel does not show whether the gene has increased expression
only shows if the sequence interacts with specific protein
does appear that the sequence is specific to transcription factor A
top line in lane 3 represents the interaction between the DNA sequence, the sequence-specific protein, and the TFA-specific antibody
weighs more than the probe and the protein + probe
(B) Dr. Song found a 15-nucleotide sequence located 300 bp upstream of gene X in cardiomyocytes. She suspects the sequence to be an enhancer and did EMSA with known cardiomyocyte specific transcription factors A and B. Observe the following radiographs of EMSA. Was it possible for her to validate the 15 bp sequence as ‘actual’ enhancer (Yes/No)? Comment on the gel images in detail keeping in view sequence-specific binding of specific transcription factors on the putative enhancer.
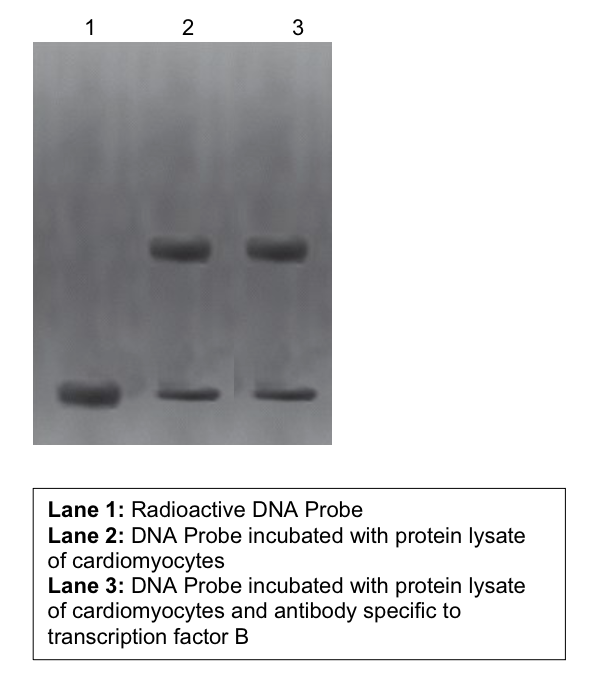
TRB does not bind to the specific sequence
if it did, there would be a third line in lane 3 where the protein + probe + antibody bound
the sequence is binding to a protein, but that protein is not TFB as it did not bind to the protein specific antibody
Fibroblast growth factor gene is regulated by GLI3 (a specific transcription factor).
Make a simple drawing of the FGF gene showing the arrangement, from 5ʹ®3ʹ, introns, exons (assume there are three exons), the promoter and enhancers (in all possible positions). On your drawing indicate all the sites where GLI3 can potentially bind, where RNA polymerase II and the general transcription factors would bind and label the start of transcription as +1.
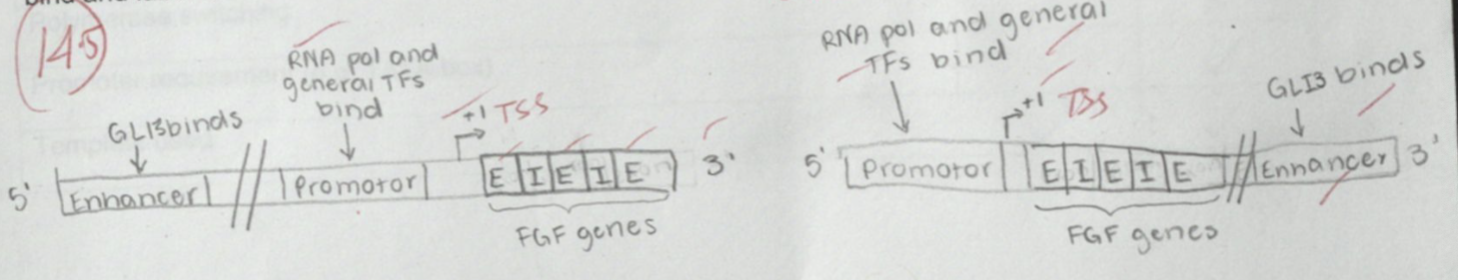
enhancers can also be found in introns
Carboxy Terminal Domain (CTD) of RNA polymerase II contains seven amino acid long conserved sequence repeated for fifty times. Which of the amino acid should be phosphorylated as a “go” signal for RNA pol II to leave promoter and start synthesizing mRNA.
serine 5
Enlist two main functions of TFIIH
helicase (unwinds supercoiled DNA)
CDK-dependent kinase (phosphorylates CTD)
What are the two main components of TFIID?
TATA binding protein (TBP)
TATA binding associated factors (TAFs)
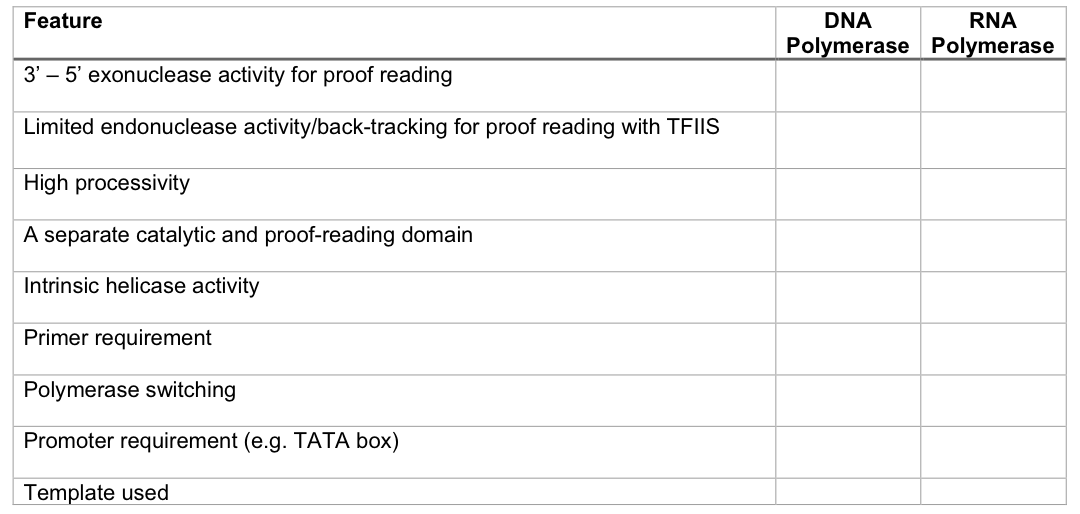
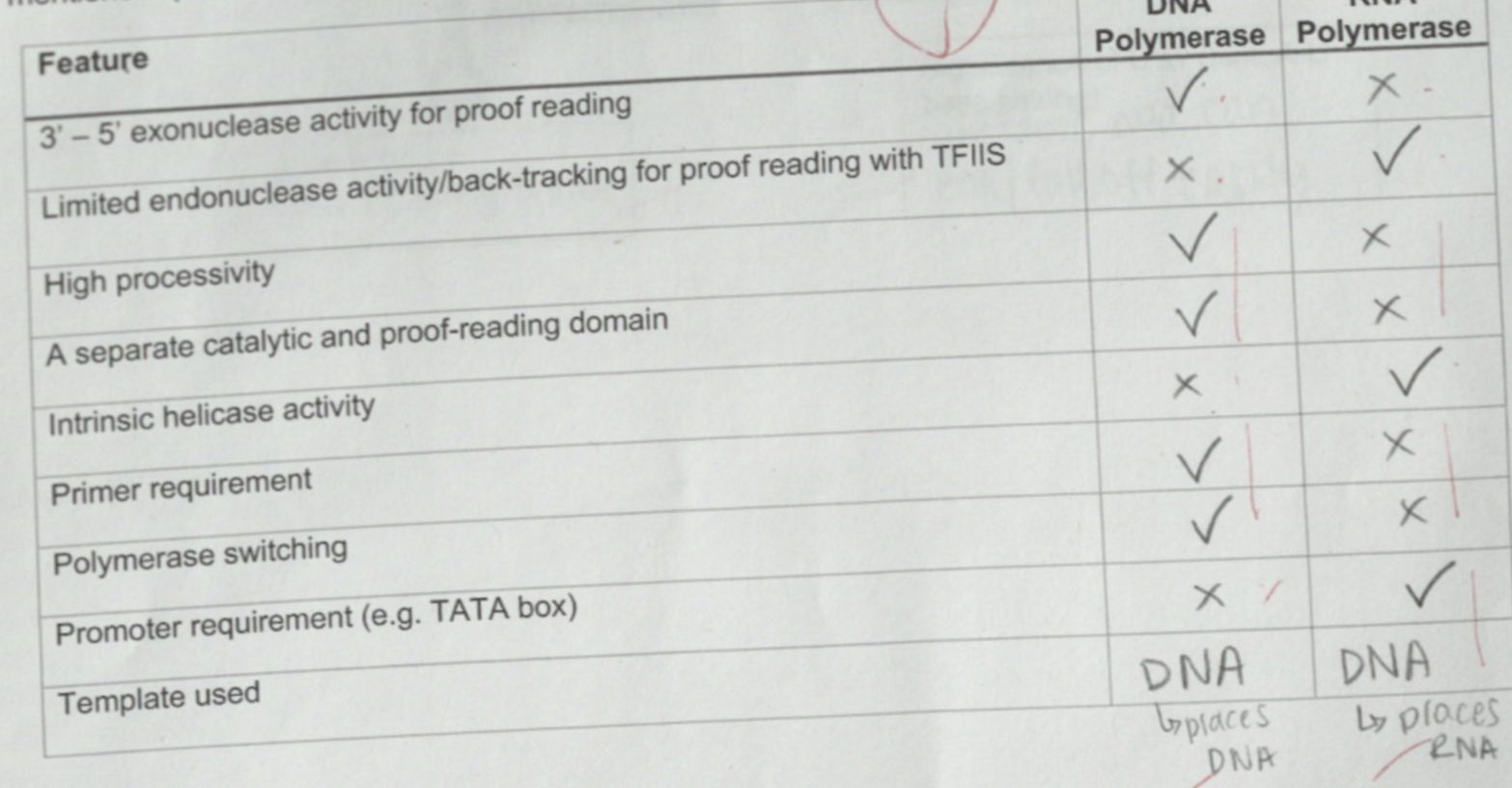
Fill the following table using ✓ and x to show presence and absence respectively of the features mentioned.
Observe the following riboswitch. Answer in the boxes provided.
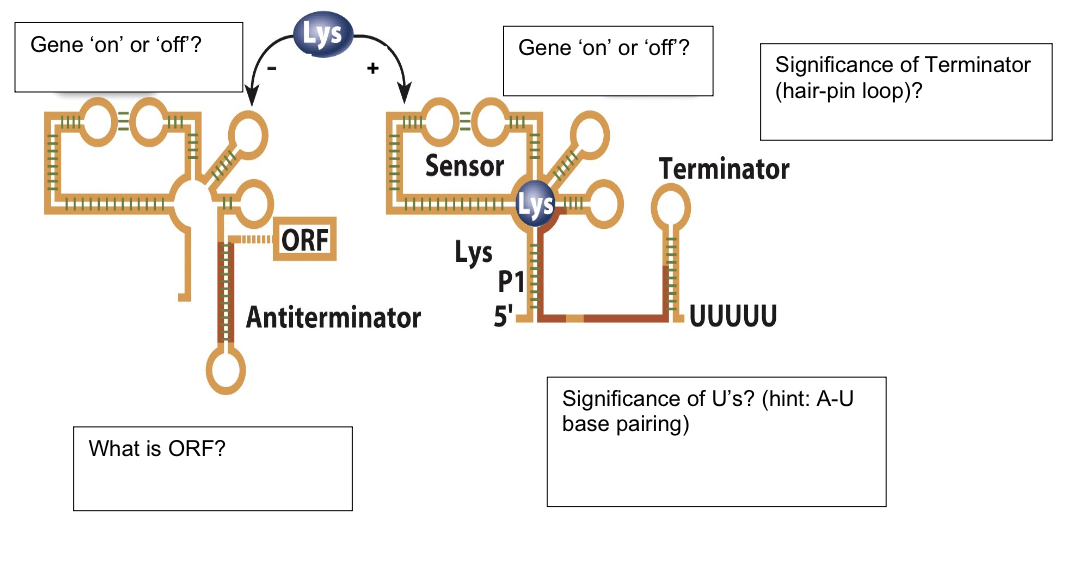
first gene is ON
ORF
open reading frame
second gene is OFF
significance of Us
A-U base pairing is unstable, so RNA Pol falls off, terminating transcription
significance of terminator (hair-pin loop)
allows for rho-independent termination because it is an obstacle for RNA polymerase
Fill in the empty boxes with relevant information. Briefly comment on the diagram.
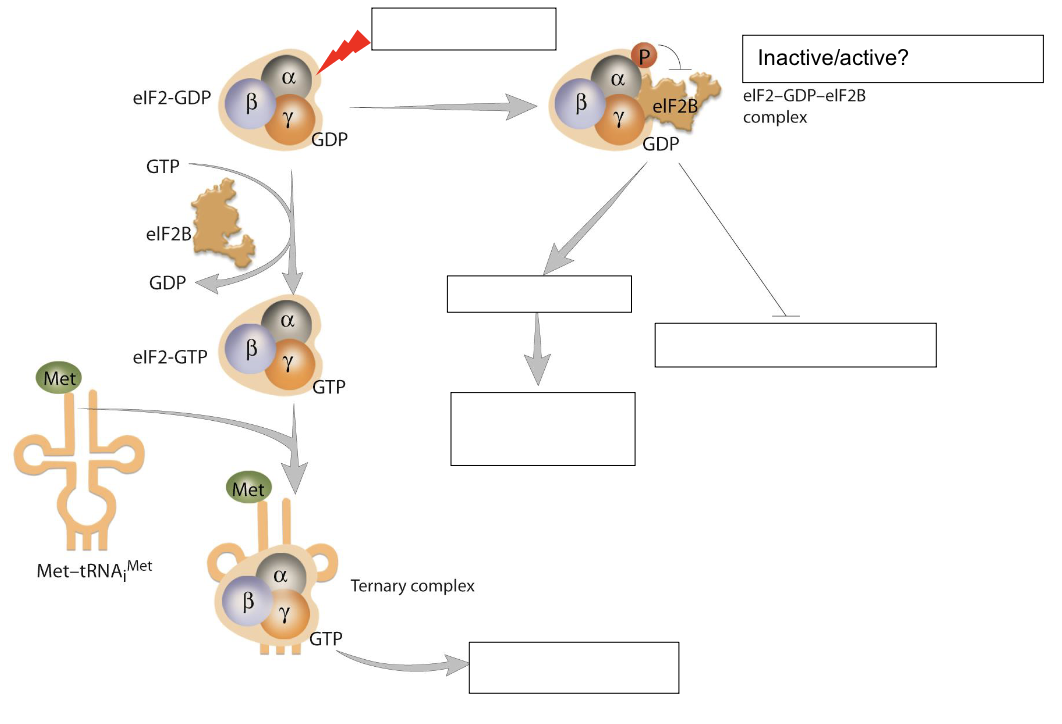
Boxes
DNA damage
Inactive
selective translation → activates stress response protein synthesis
global translation (inhibited)
binds to 40S subunit and mRNA
eIF2 is an initiation factor active when bound to GPT
when DNA damage occurs, phosphorylation of a-subunit blocks the activity of eIF2B, so it cannot perform the guanosine nucleoside exchange necessary for eIF2 to form the ternary complex
inactive complex prevents translation initiation of most proteins, but does allow for selective translation of stress-response proteins to repair the damage
Comment on the Gene Targeting Knock-out Models shedding light on neomycin, neor, ganciclovir and tk and fill the empty boxes regarding information about cell survival in the presence/absence of neomycin and ganciclovir
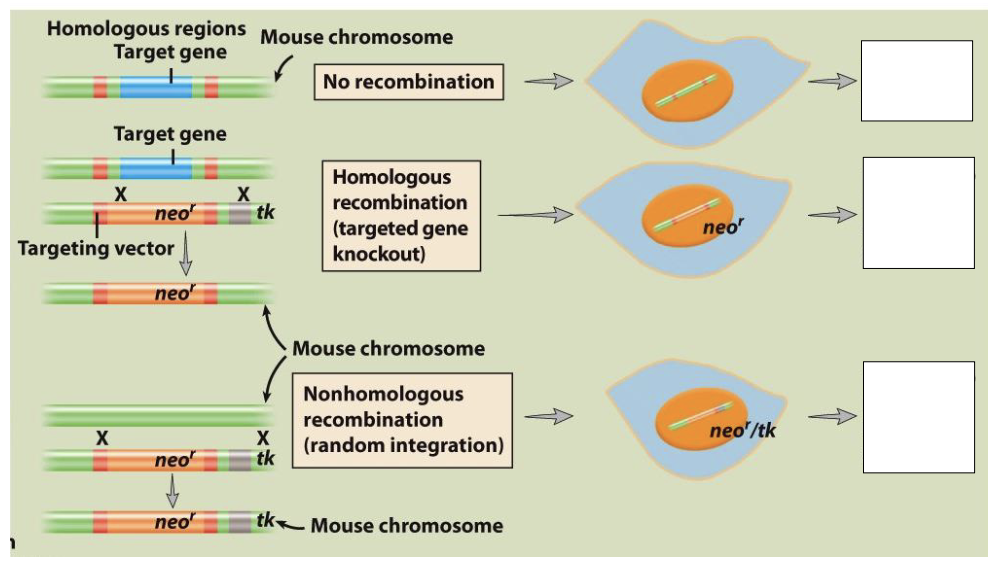
Boxes
killed by neomycin
resistant to neomycin and ganciclovir
resistant to neomycin, killed by ganciclovir
Scenario A
no recombination of neo r gene occurs
mouse is not resistant to neomycin so it dies
Scenario B
homologous recombination occurs
target gene replaced by neo r gene giving the mouse resistance of neomycin
no tk integrated so the mouse is resistant to ganciclovir
Scenario C
non-homologous recombination occurs
mouse gains neo r and tk genes
resistant to neomycin but not to ganciclovir
Describe back splicing. What is its advantage? Give three examples
back splicing occurs when the wrong splice sites are targeted so the wrong introns and exons are removed or not removed
advantages
stability
closed loop makes it less prone to degradation
translational control with small RNA
whilhey are made in small amounts, they can exist in high amounts in cells that need stable RNAs and suffers heavily under RNA degradation that hinders the synthesis of proteins
protein diversity
Define alternative splicing? What is its significance? Describe the mechanism by using illustration highlighting Exon Splice-site Silencer (ESS), Exon Splice-site Enhancer (ESE), Intron Splice-site Silencer (ISS), Intron Splice-site Enhancer (ISE), branch point, activator, repressor protein, snRNPs
one or more gene (pre-mRNA) can be spliced different ways to create multiple mRNA transcripts
allows for multiple proteins to be synthesized by the same gene
exon splice-site silencers and exon splice-site silencers repress the silencing at intron and exon splice sites
ESS and ISS bind to repressors
exon and intron splice-site enhancers activate splicing at intron and exon splice sites
ESE and ISE bind to activator proteins
branch point is where the 5’SS end of the intron binds after the first transesterification
forms an intron lariat loop which will be degraded after the second transesterification
bulged adenosine
snRNPs are the ribonucleoproteins that identify the splice sites and perform the catalysis of the introns
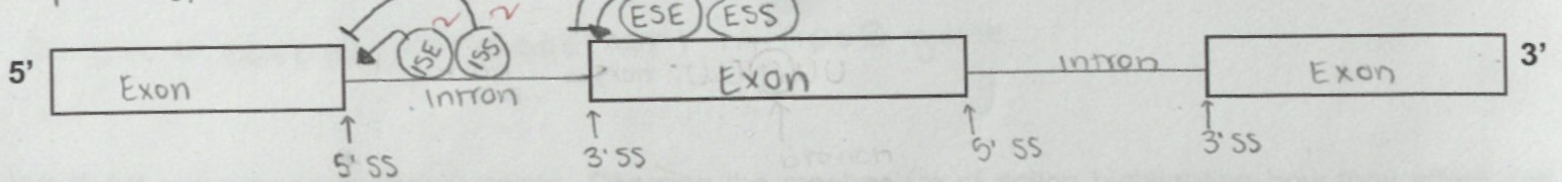
Descibe two fates of mRNA targeted by miRNA
translationally repressed
synthesizing minimal to no proteins
degraded
nucleotides may be cut which destabilizes the mRNA
Describe biogenesis of miRNA in terms of steps/intermediates in the nucleus and cytosol
miRNA is endogenous
encoded for in the genome and translated into pri-mRNA
DROSHA cleaves the pri-mRNA to make smaller transcripts called pre-MRNA
pre-MRNA transported from the nucleus to the cytosol
in cytosol, it binds to RISC
has an argonaut subunit with slicer activity
separates the double stranded pre-miRNA into two single-stranded fragments
one is degraded and the other is mature miRNA and is used as guide RNA
Ribosomes are ribozymes have A, P, and E sites. Which part of eukaryotic ribosomes provide peptidyl transferase activity?
23S rRNA (often near the P site)
Enlist two types of RNA editing mechanisms
U insertions/deletions
C → U
A → I
Enlist two tumor suppressor genes. Describe the mechanism of action highlighting how they affect cell cycles
pRB
represses formation of retinoblastoma tumors
controls the cell cycle by requiring a specific transcription factor to activate the expression of S-phase genes
controls the transition from G1 to S-phase (replication
CDK phosphorylates pRB which causes a conformational change, loosening EF2’s grip on pRB
more phosphorylation causes EF2 to fall of entirely
EF2 acts as a transcription factor and promotes the expression of s-phase genes
allows the cell to move from the G1 to S phase
p53
regulates the cell’s ability to move from the G1 to S phase
activated when DNA damage is recognized
activated p53 then acts as transcriptional repressor and inhibits the expression of CDF-21
stops the cell in the G1 phase
the cell must repair the damage to inactivate p53 to allow the transition into the S phase
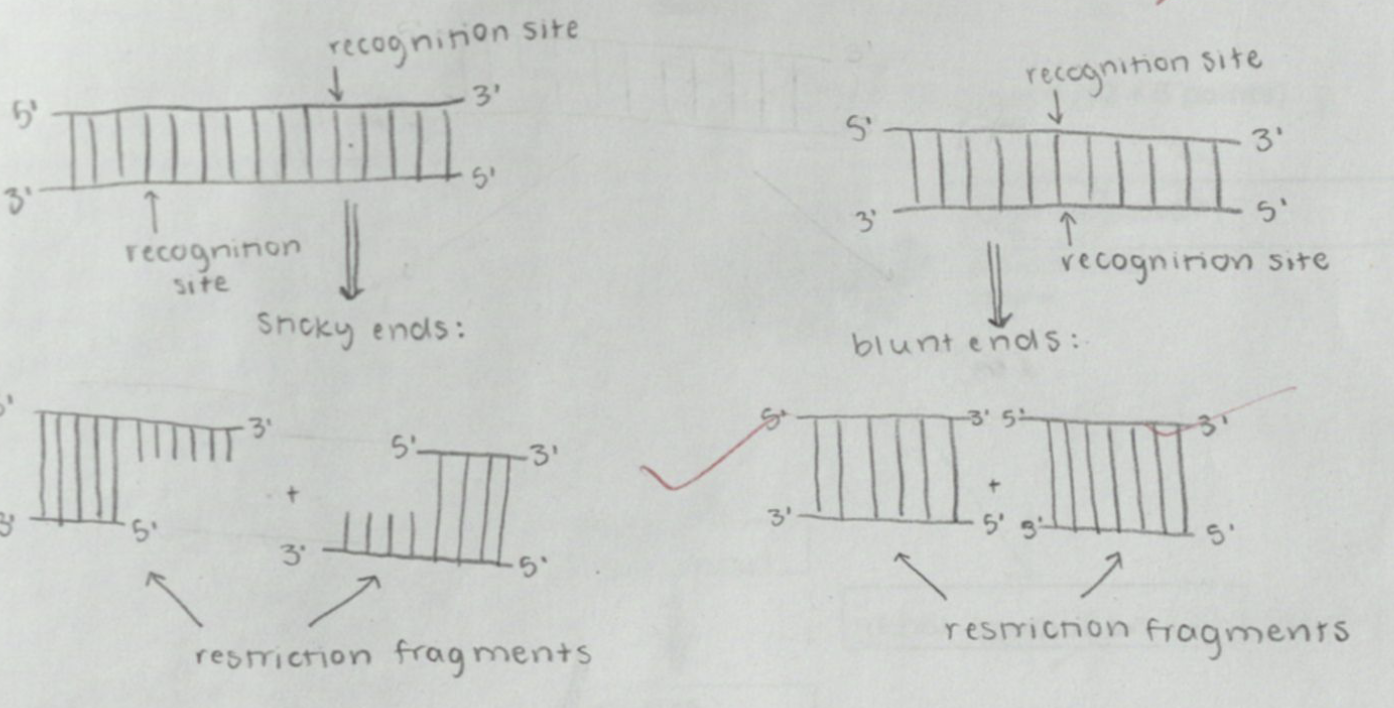
What are restriction endonucleases? Give any two examples. Illustrate sticky and blunt ends.
cut at restriction sites to form restriction fragments
BAMHI
Sma I