6.6 Gene Expression and Cell Specialization
1/12
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
13 Terms
Eukaryotic Gene Regulation
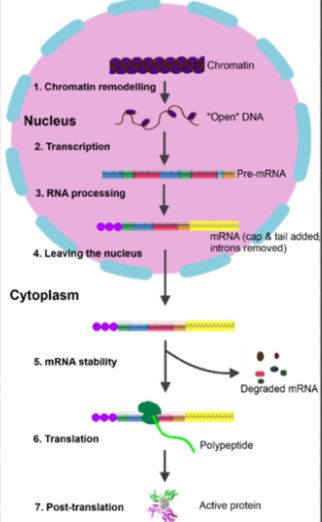
gene regulation in eukaryotes can occur at several different stages:
chromatin remodeling
transcription (most genes regulated at this level)
RNA processing
RNA stability
Translation
Post-translation
Chromatin Remodeling
if DNA is tightly wound it is less accessible for transcription
how can it be modified?
histone acetylation loosens (accessible)
DNA methylation condenses (inaccessible)
epigenetic inheritance
Histone Acetylation
adds acetyl groups to histones, which loosens the DNA (accessible)
DNA Methylation
adds methyl groups to DNA, which causes the chromatin to condense (inaccessible)
Epigenetic Inheritance
inheritance that involves changes in how a gene is expressed without any changes in that gene’s nucleotide sequence (ex. methylation)
can be passed on during cell division
modifications can be reversed, unlike mutations
explains why one identical twin may express a disease while the other does not
Transcription
once chromatin modifications allow the DNA to be more accessible, specific transcription factors can bind to control elements
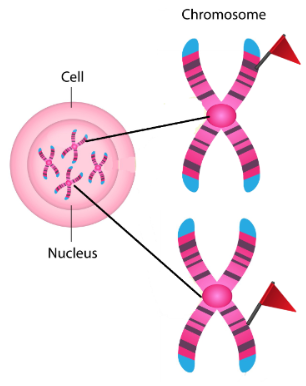
groups of genes with related functions may be regulated together if they share common control elements recognized by the same transcription factors (even if located on different chromosomes), which allows for coordinated regulation of related processes
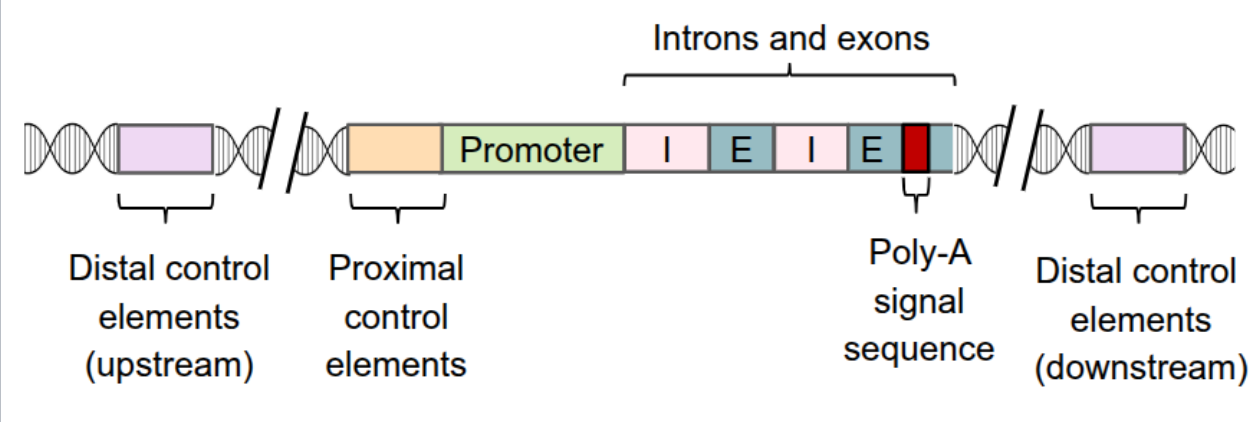
Control Elements
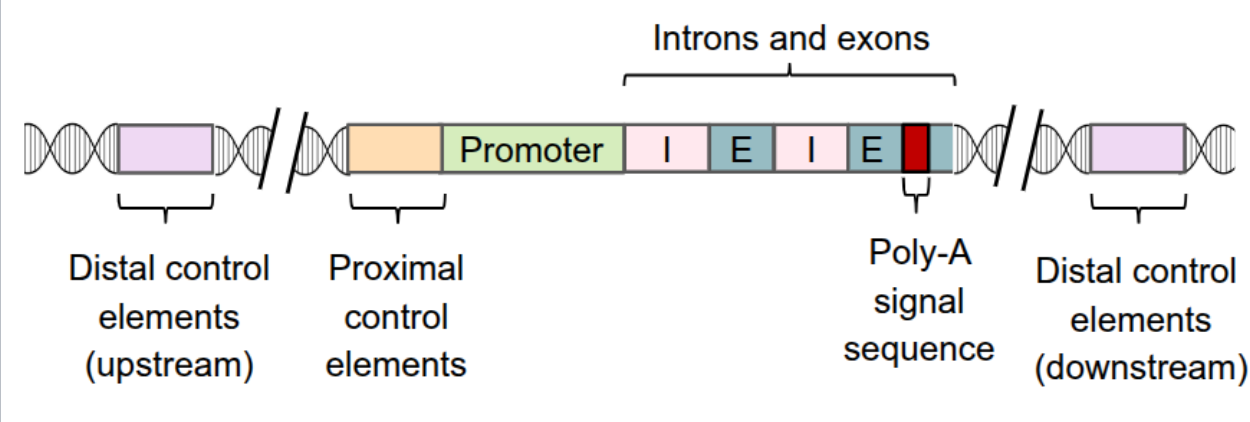
sections of noncoding DNA that can be located near (proximal) or far (distal) from the promoter
includes enhancers
the rate of gene expression can be regulated by binding of transcription factors
groups of genes with related functions may be regulated together if they share common control elements recognized by the same transcription factors (even if located on different chromosomes)
this allows for coordinated regulation of related processes
Enhancers
distal control elements can be grouped together as enhancers
can be either upstream or downstream of the transcription start site
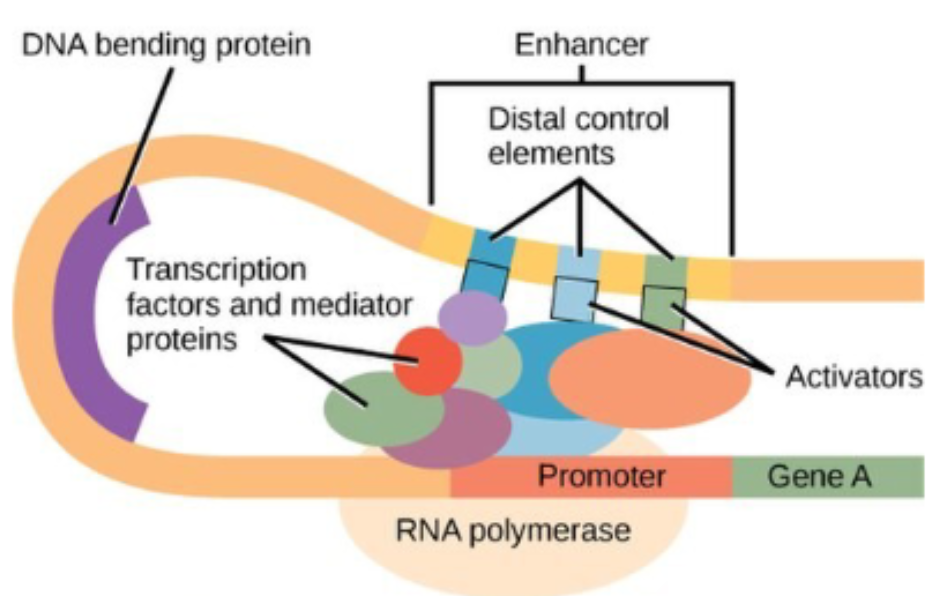
Activators
can bind to enhancers to increase the rate of transcription
even though enhancers are located far from the promoter, DNA is flexible and can bend
DNA’s flexibility and bending allows activators to interact with mediator proteins, which recruit general transcription factors and RNA polymerase to begin transcription
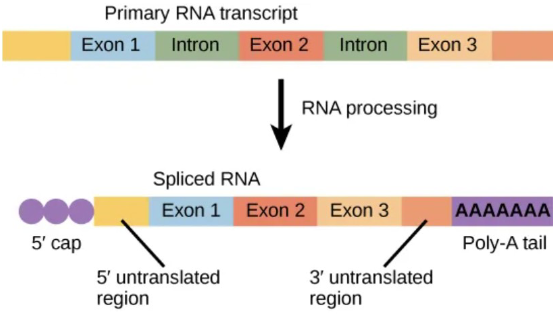
RNA Processing
adding poly-A tail and 5’ cap
alternative splicing of pre-mRNA
RNA Stability and Transport
RNA transport: controlling access to nuclear pores regulates mRNA transport to cytosol
RNA stability: lifespan of mRNA in cytosol affects how many protein molecules can be translated from it
microRNAs and small interfering RNAs can bind to mRNA and degrade it or block translation
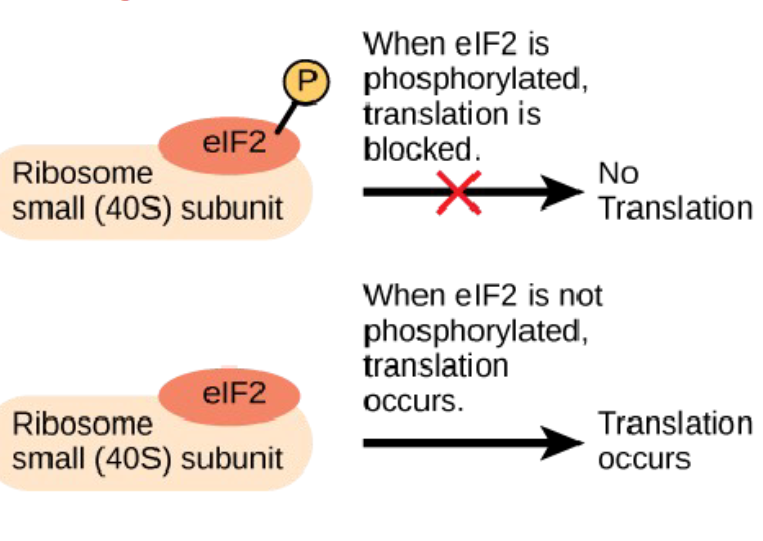
Translation
the initiation of translation can be blocked by regulatory proteins that bind to sequences in the 5’ cap or poly-A tail, which prevent the ribosome from binding
modification (activation/inactivation) of proteins that help ribosomes attach to mRNA
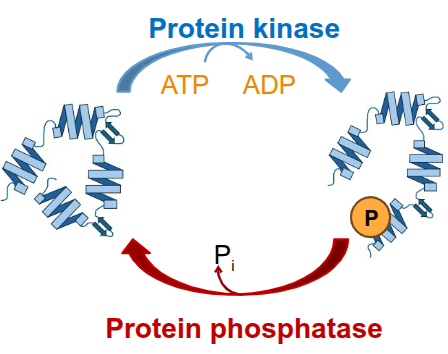
Post-Translation
protein modifications
ex. addition/removal of phosphate groups for activation/inactivation or chemical tags for degradation