MCAT Lab Techniques
1/59
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
60 Terms
gel electrophoresis (types, purpose, anode cathode charge)
Native-PAGE
SDS-PAGE
Reducing SDS-PAGE
Isoelectric focusing
Purpose: Separation of proteins, DNA, or RNA based on size and/or charge
anode: +
cathode: -
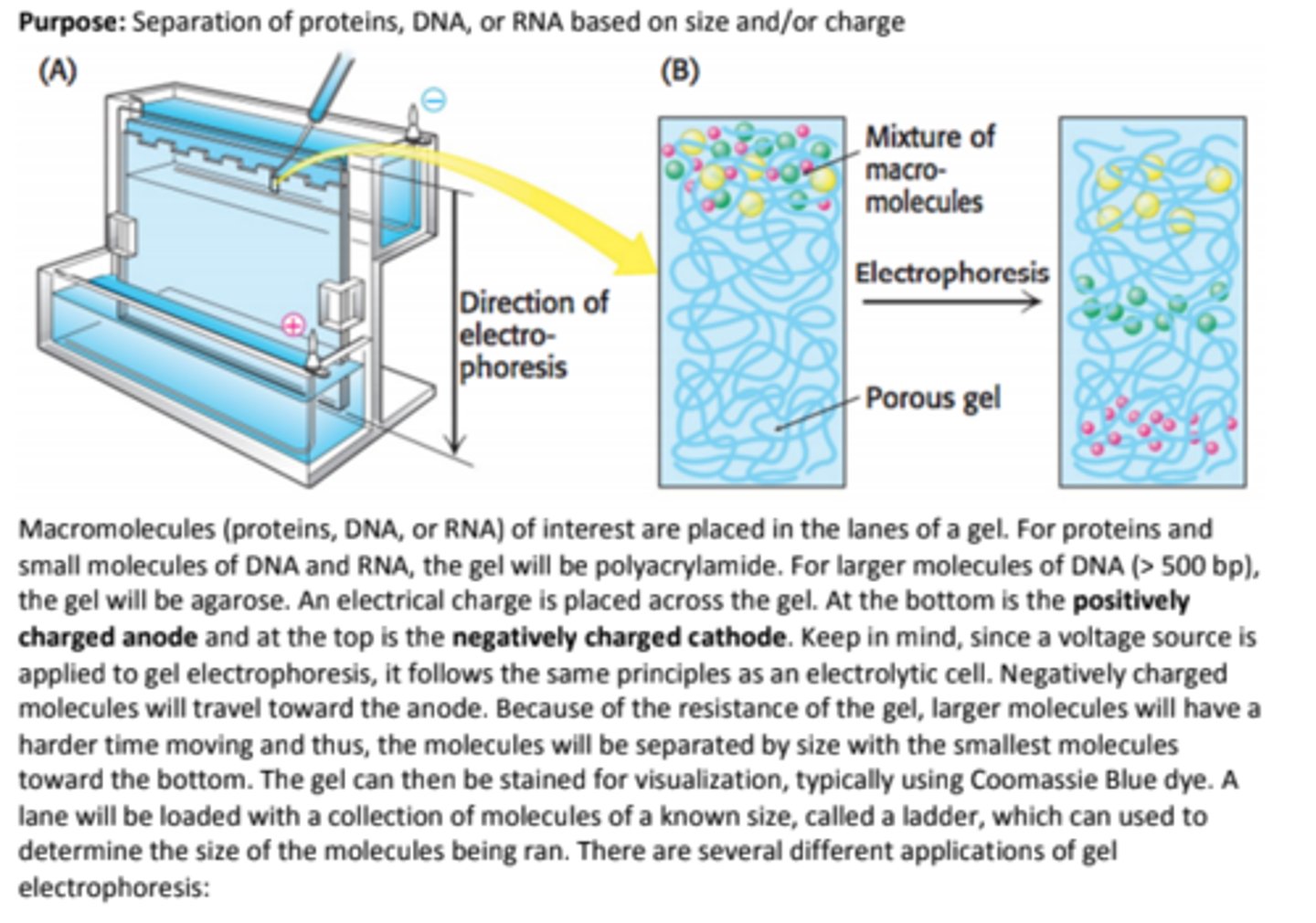
Native-PAGE
Native-PAGE is a polyacrylamide gel electrophoresis method for proteins that occurs under non-denaturing conditions.
This method will separate proteins by SIZE while retaining their structure.
Reducing SDS-PAGE
Reducing SDS-PAGE is exactly like SDS-PAGE, but with the addition of a reducing agent, like β-mercaptoethanol, which will separate proteins by size AND reduce disulfide bridges and result in a completely denatured protein.
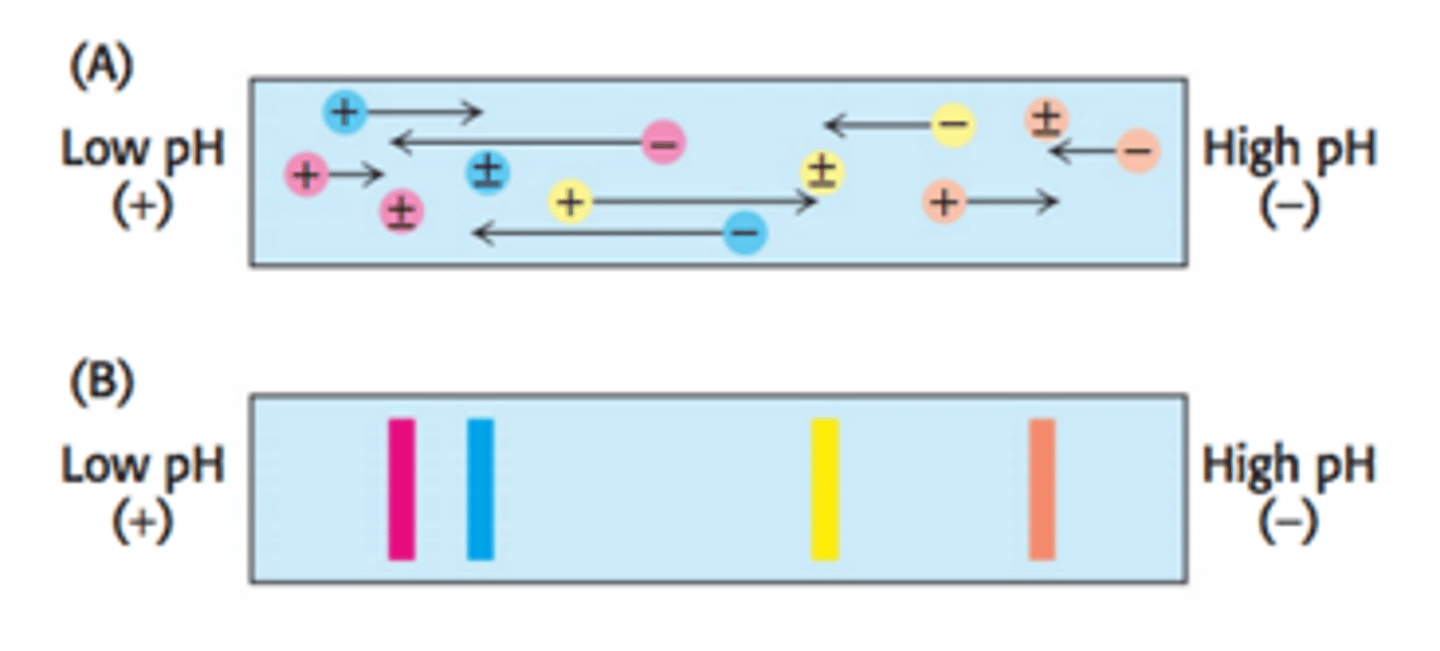
Isoelectric focusing
Isoelectric focusing is a gel electrophoresis method that separates proteins on the basis of their relative contents of acidic and basic residues. A polyacrylamide gel with a pH gradient (low pH on one side, high pH on the other) is used.
When proteins migrate through the pH gradient gel, they will travel toward the anode until the area of the gel with the pH that matches their isoelectric point (pI).
When a protein is at its pI, it has a net charge of zero and will not be attracted to the positively charged anode so it will not move.
Western Blotting (Protein)
Purpose: Detection of a specific protein in a sample
1) proteins from sample loaded into SDS-PAGE gel, separated based on size
2) proteins from gel transferred to polymer sheet
3) proteins exposed to radiolabeled antibody specific to protein of interest (sometimes 2 abs are used- one for protein of interest, another radiolabeled for visualization that binds to 1st ab)
4) polymer sheet is viewed using autoradiography used to view polymer sheet. Protein bound to radiolabeled ab = visible
Southern (DNA) and Northern (RNA) Blotting
Purpose: Detection of a specific DNA (Southern blot) or RNA (Northern blot) sequence in a sample
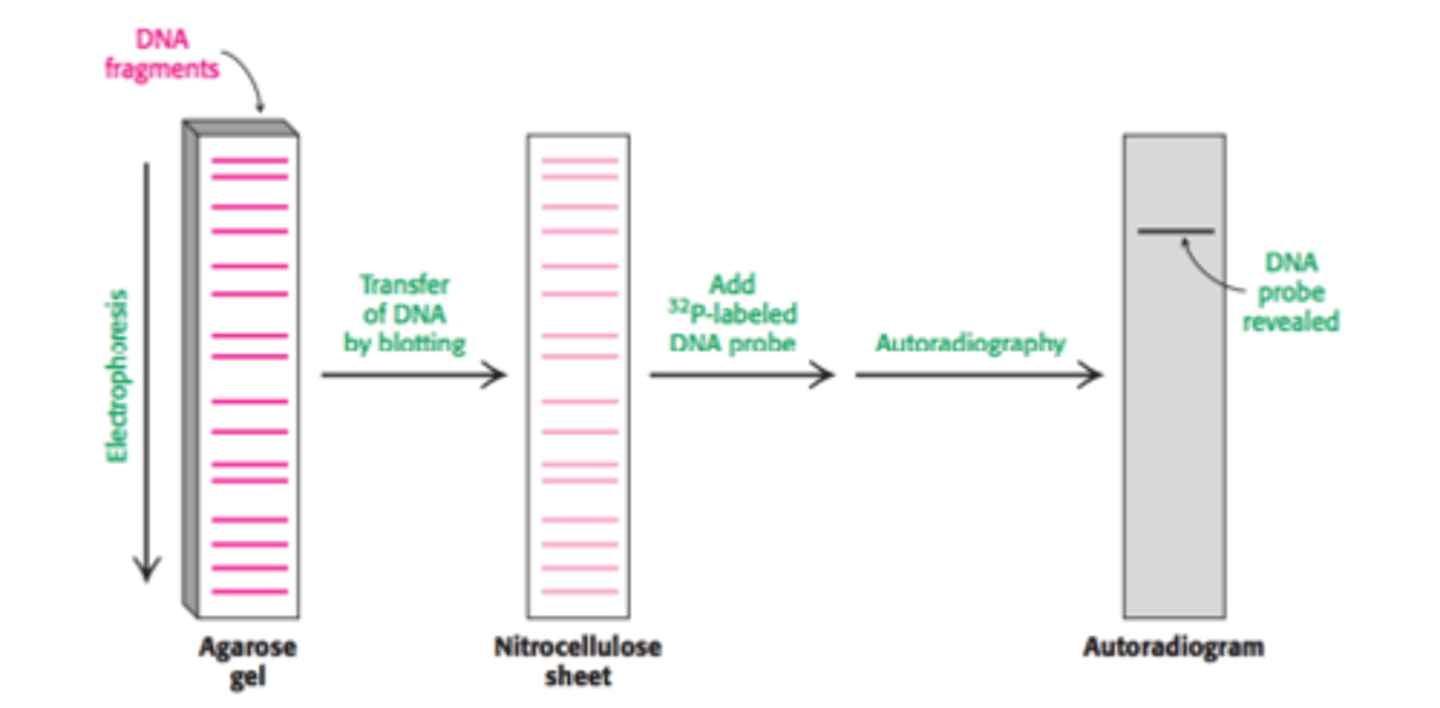
Step 1: The DNA strand of interest is exposed to restriction enzymes that cut the DNA strand into smaller fragments
Step 2: The newly cleaved strands of DNA are denatured using a solution of NaOH to create ssDNA strands
Step 3: The single stranded cleaved strands of DNA undergo gel electrophoresis, separating them by size. Smaller fragments will be found at the bottom of the gel. Polyacrylamide is used if the stands are less than 500 base pairs. Agarose is used if the strands are over 500 base pairs.
Step 4: The DNA from the gel is transferred to a sheet of nitrocellulose paper and then exposed to a 32P
radiolabeled DNA probe that is complementary to our DNA of interest.
Step 5: The nitrocellulose paper is then viewed using autoradiography to identify the strand of interest.
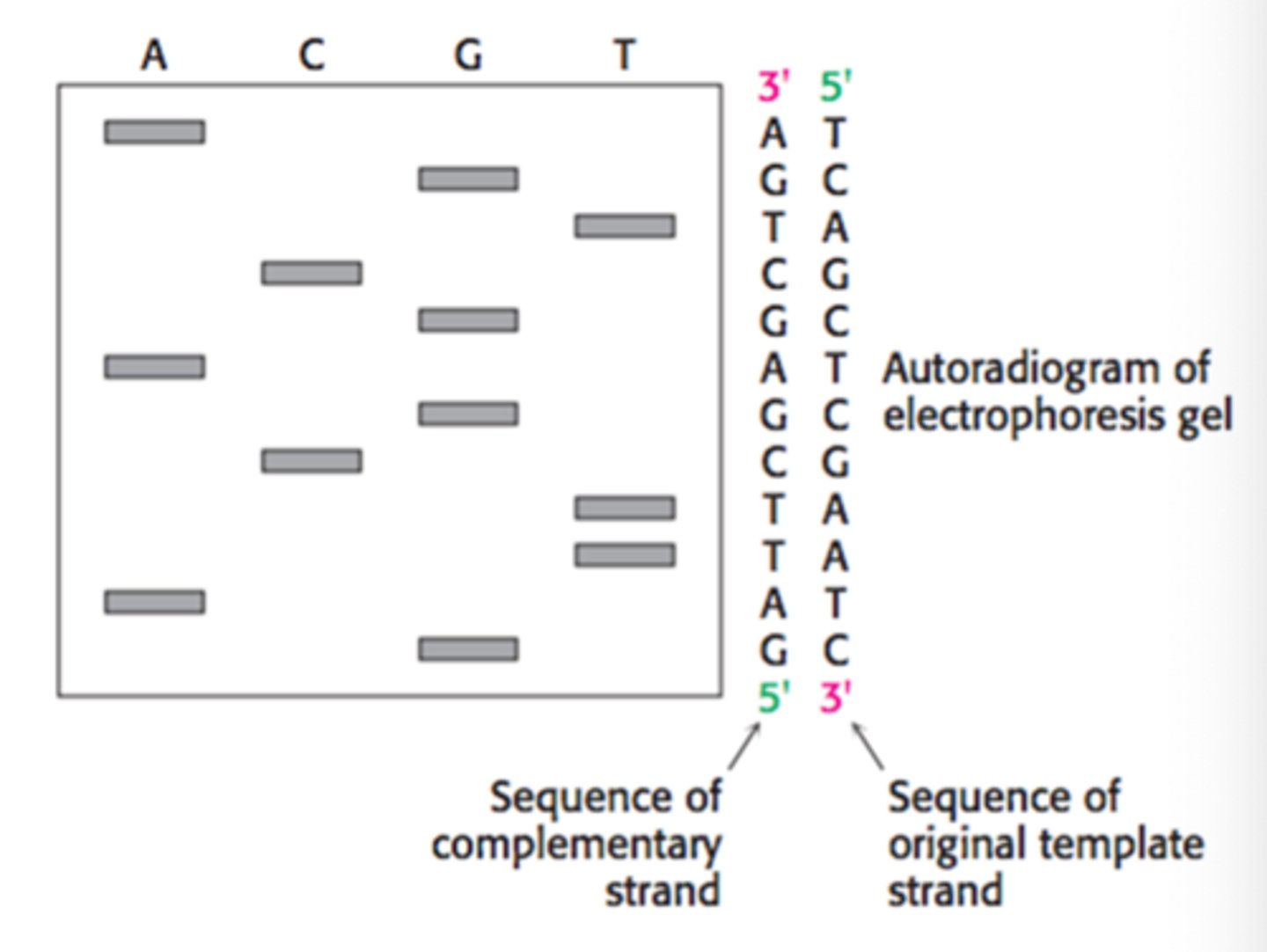
DNA Sequencing (Sanger Dideoxynucleotide Sequencing)
Purpose: Used to determine the sequence of nucleotides in a strand of DNA
Chromatography
Purpose: The separation of two or more molecules from a mixture using molecular interactions
- there is a stationary and a mobile phase with these
Traditionally, there are two components to chromatography:
- stationary phase, which is typically polar
- mobile phase, which is typically non-polar.
Polar molecules are separated from a mixture by staying with the stationary phase, while non-polar molecules stay with the mobile phase.
Liquid Chromatography
In liquid chromatography, silica is traditionally used as the stationary phase while toluene or another non-polar liquid is used as the mobile phase.
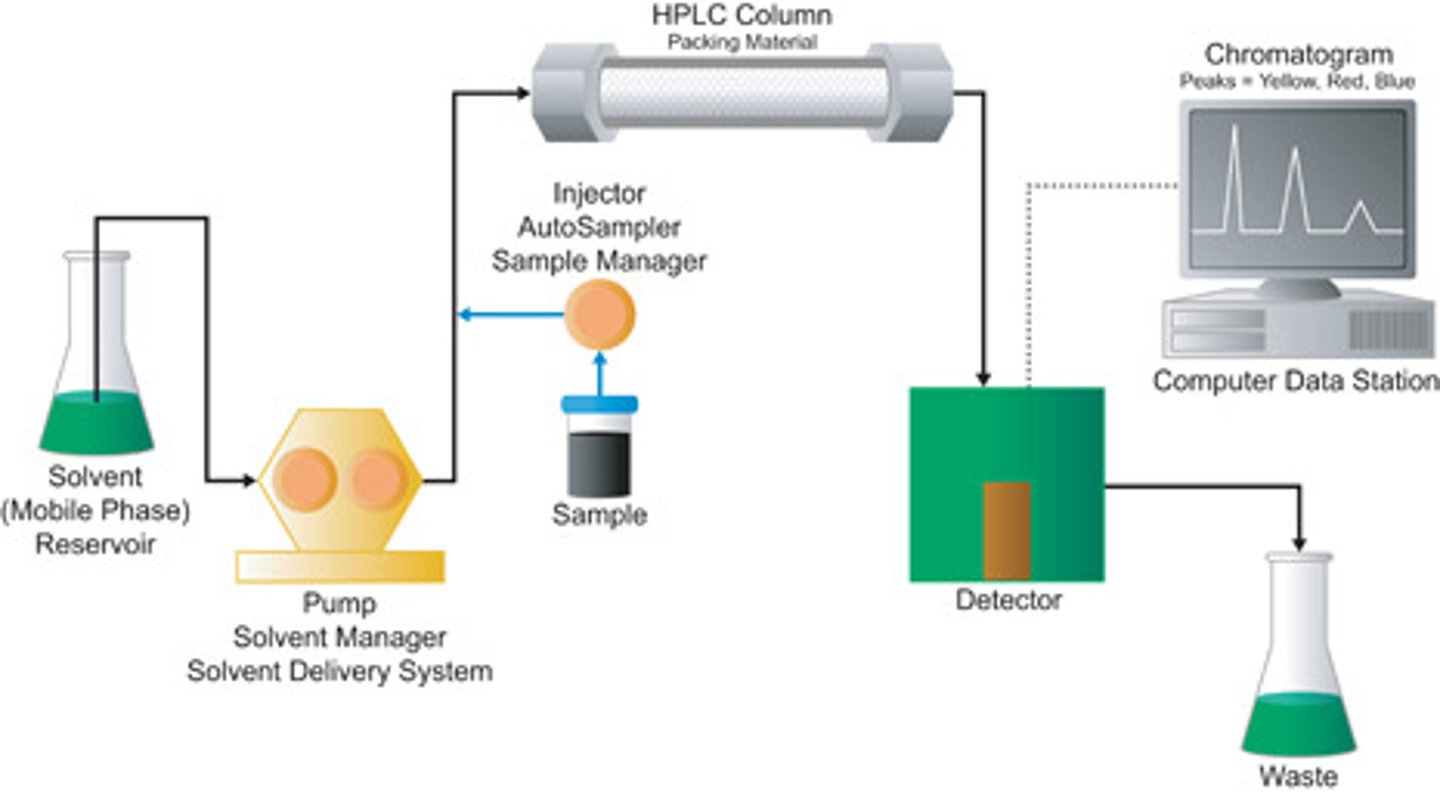
High-Performance Liquid Chromatography (HPLC)
HPLC is a type of liquid chromatography that utilizes high pressures to pass the solvent phase through a more finely-ground stationary phase, which increases the interactions between the molecules and the stationary phase, giving HPLC a higher resolving power. Molecules can then be determined based on their absorbance and elution time as seen on the right.
In normal phase HPLC, stationary phase is polar, mobile phase is nonpolar (so non-polar elutes faster)
Gas Chromatography
Gas chromatography (also known as gas-liquid chromatography) is used to separate and analyze molecules that can be vaporized.
The mobile phase is an inert or unreactive gas, such as helium or nitrogen, while the stationary phase is a thin layer of liquid or polymer that surrounds the walls of a tube.
The stationary phase allows more polar molecules to elute slower, giving them a higher retention time. The molecule with lowest boiling point comes out of the column first
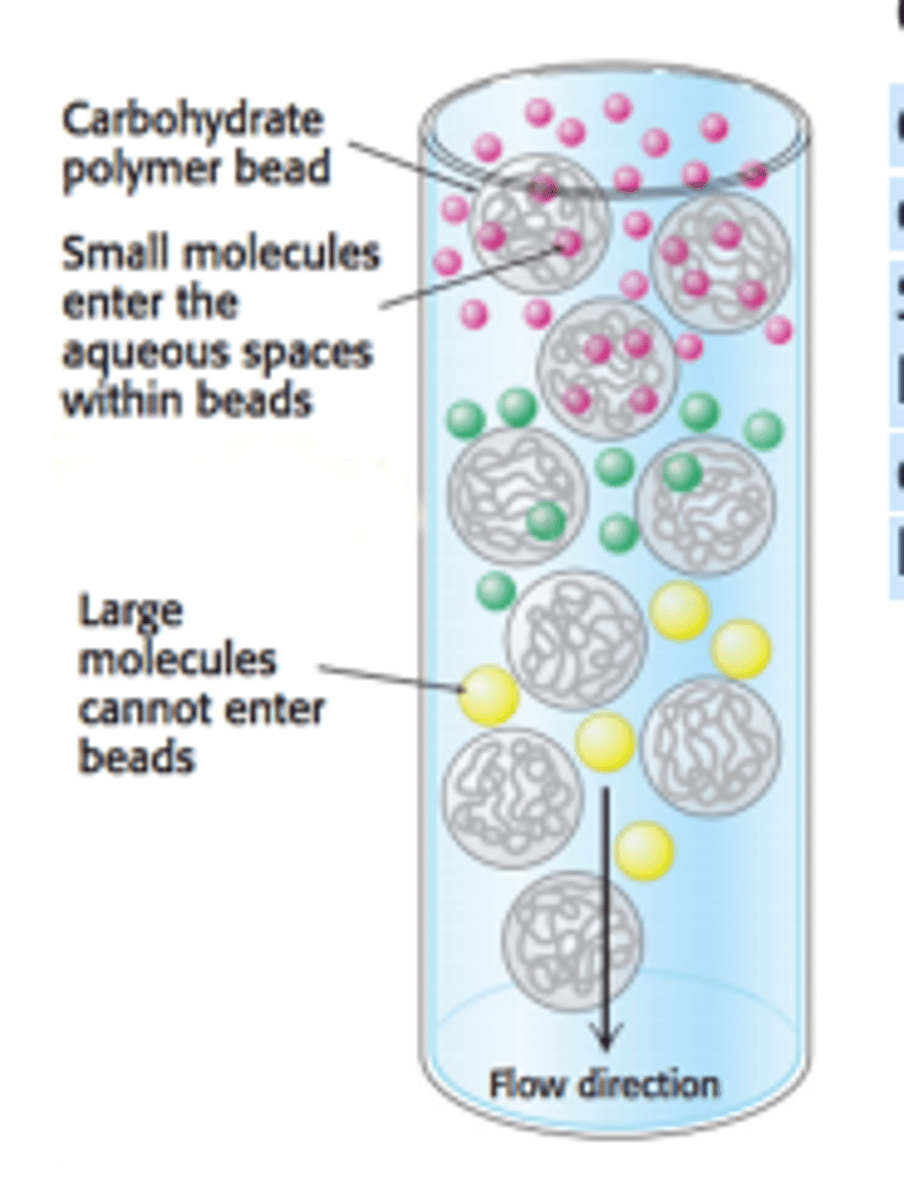
Gel-Filtration (Size Exclusion) Chromatography
Gel-filtration chromatography (also known as size-exclusion chromatography) is used to separate molecules by size rather than polarity.
Smaller molecules can enter the porous gel beads, allowing them to elute later, while larger molecules that do not fit will elute faster.
The gel beads can be viewed as the stationary phase, while the solution in the column can be viewed as the mobile phase.
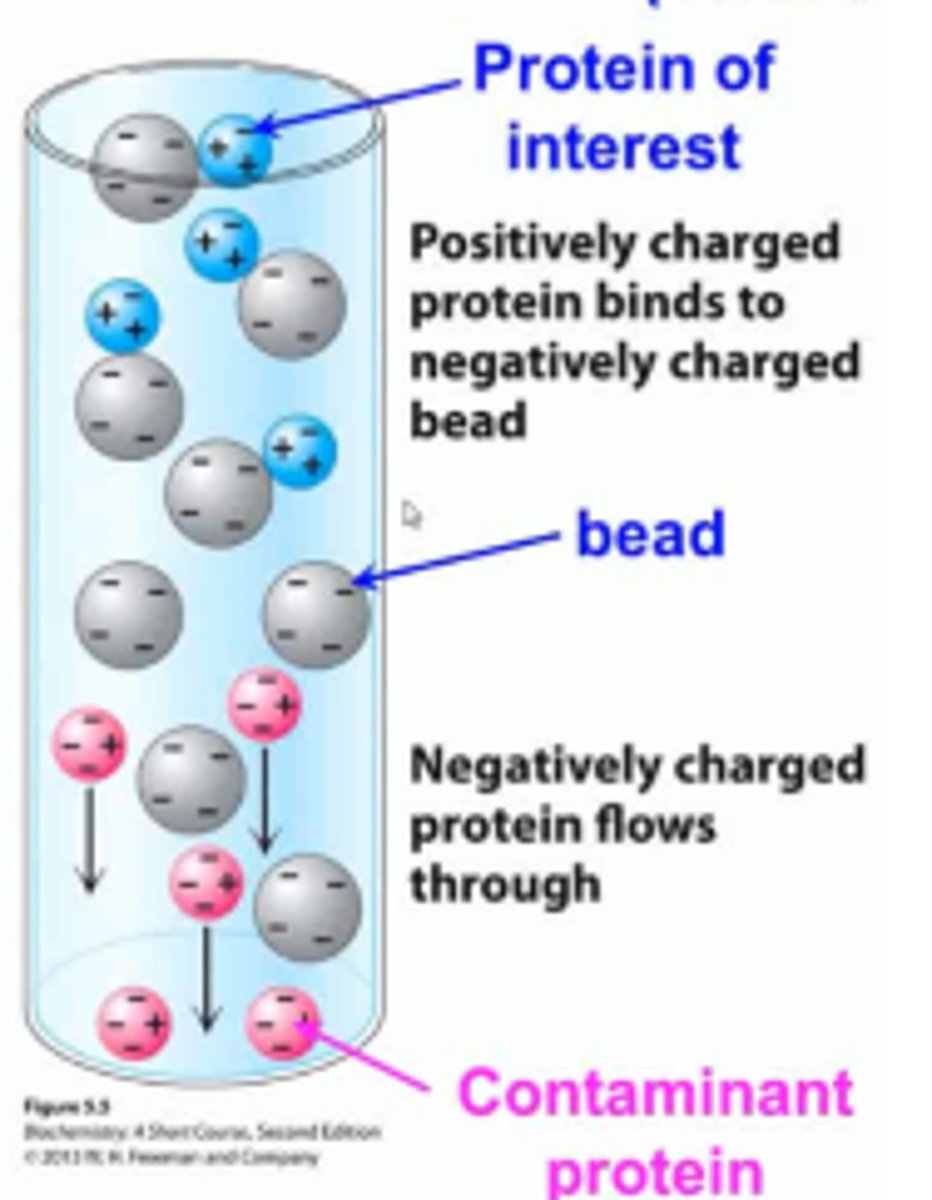
Ion-Exchange Chromatography
Ion-exchange chromatography will separate proteins by their net charge. The column is filled with charged beads, either positive or negative.
In cation-exchange, negatively-charged beads are used which attract positively charged proteins and negatively-charged proteins will elute first.
In anion-exchange, positively-charged beads are used which attract negatively-charged proteins and positively-charged proteins will elute first.
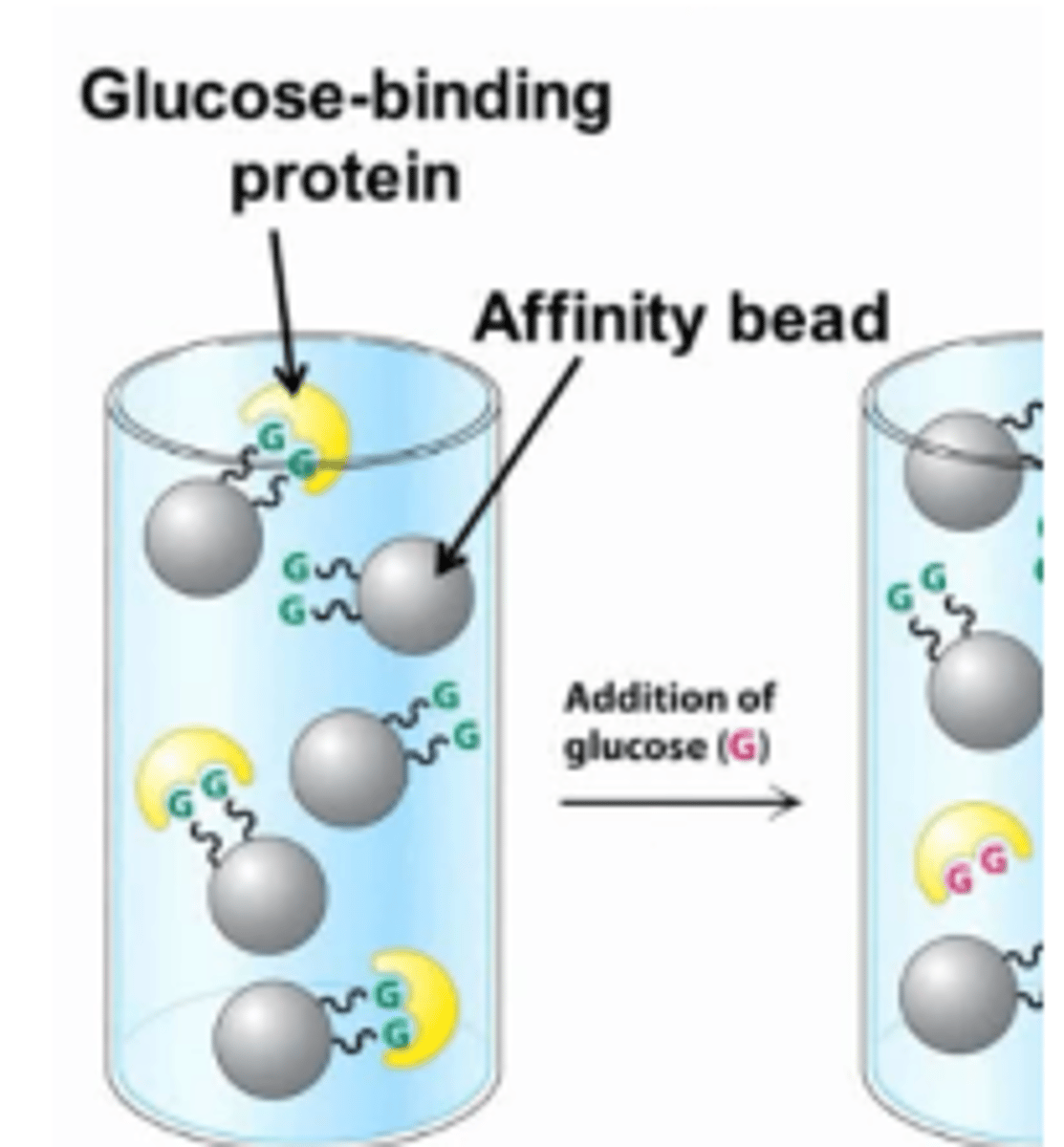
Affinity Chromatography
Affinity chromatography will separate proteins based on their affinity for a specific ligand.
Beads that are bound to a specific ligand will be used and proteins with a high affinity for that ligand will bind to the beads, allowing proteins with a low affinity to elute first.
The high affinity proteins are then eluted by increasing the concentration of the free ligand in the column, which competes for the active site of the bound proteins.
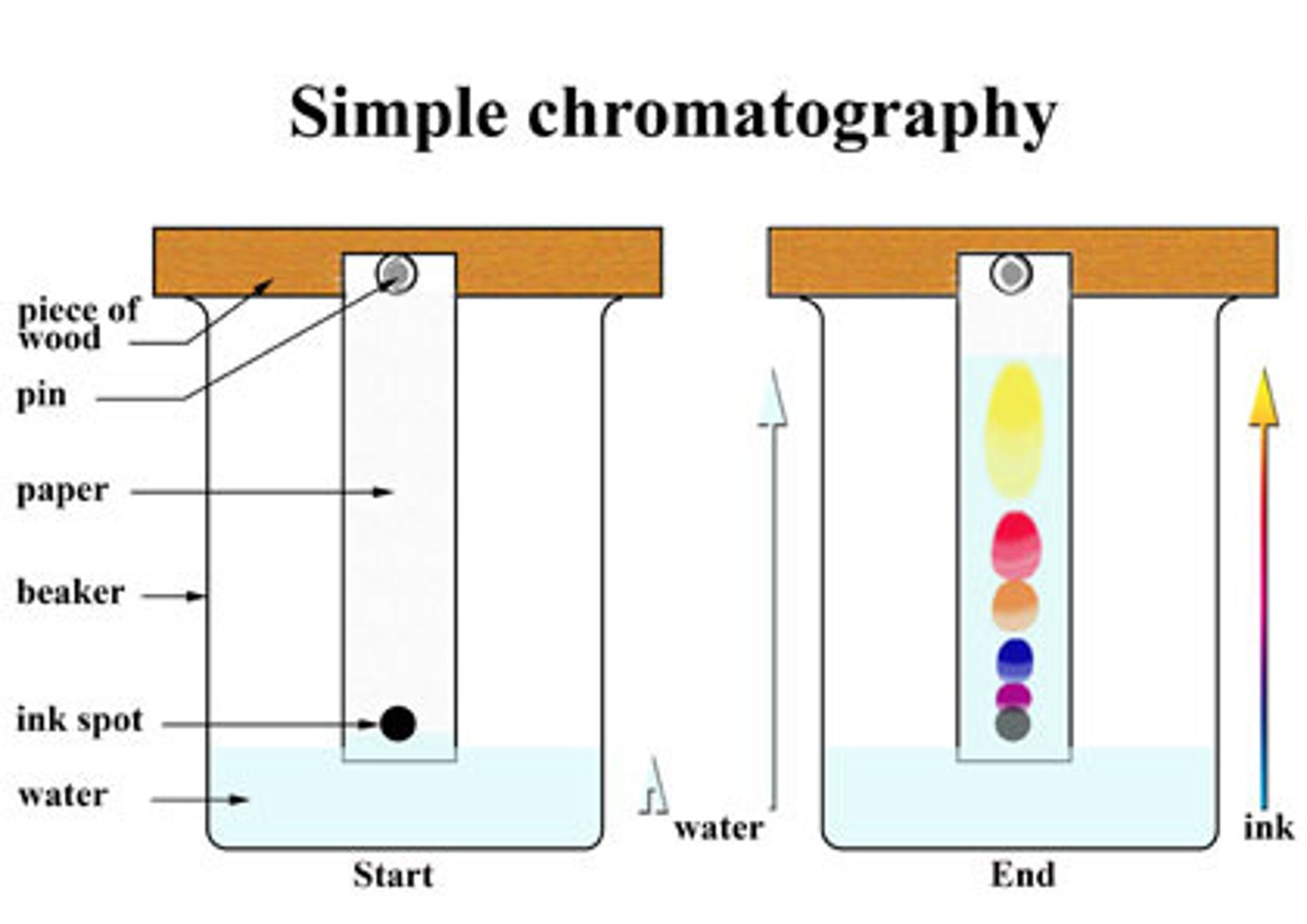
Thin-Layer Chromatography
Thin-layer chromatography consists of a small sheet of medium that is coated in an adsorbent material, such as silica gel.
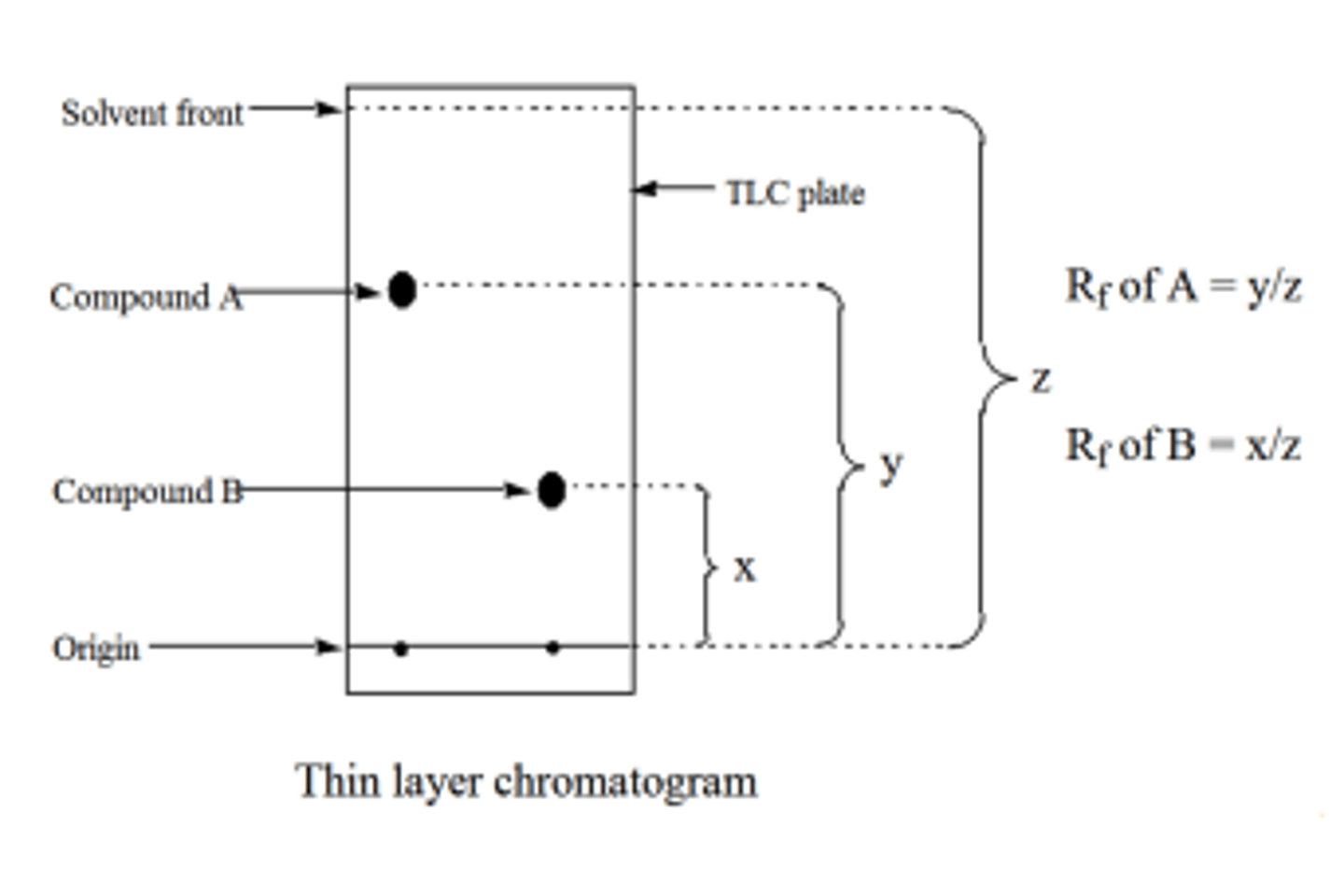
The polar silica is the stationary phase. The molecules of interest are added to the bottom of the sheet and the sheet is placed in a non-polar liquid, such as heptane, until it reaches the origin. The mobile phase then travels up the plate using capillary action, allowing the molecules to move with it if they are relatively non-polar. The spots are then visualized using UV light.
The relative distances traveled between the molecules is represented by the Rf value, which is measured as the ratio of the distance the molecule traveled from the origin to the distance the solvent front traveled from the origin.
Distillation
Simple Distillation
Simple distillation is used to separate two molecules from a solution when their boiling points differ by 25 C or greater.
Fractional Distillation
Fractional distillation is used to separate two molecules from a solution when their boiling points differ by less than 25 C.
Polymerase Chain Reaction (PCR)
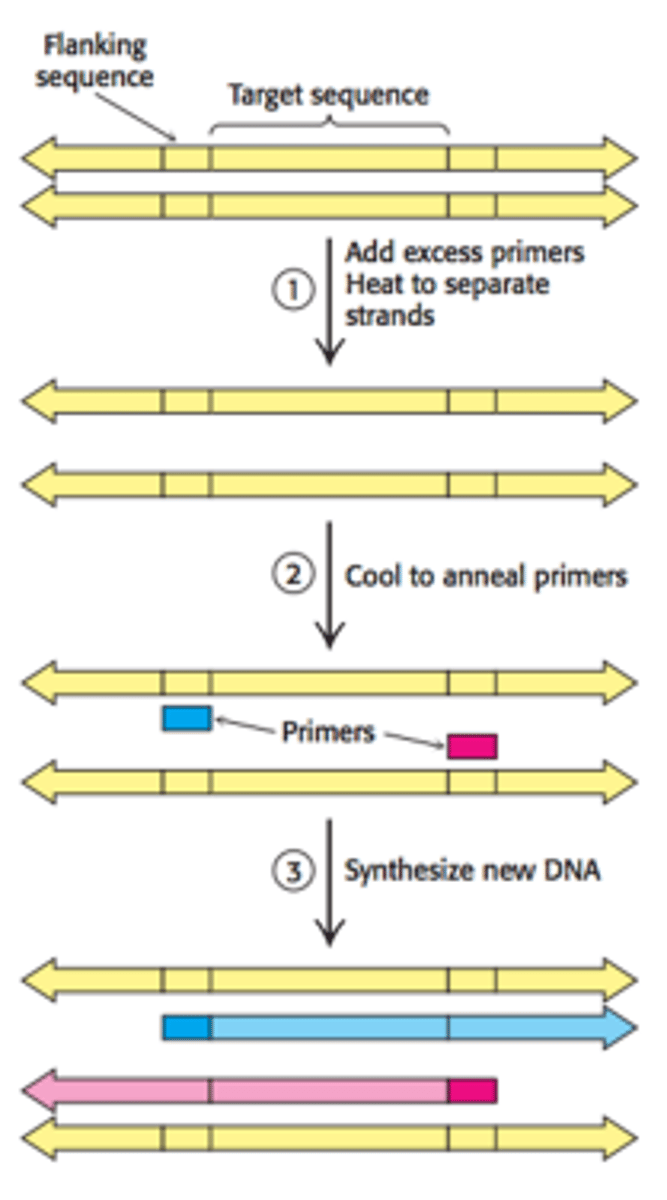
Purpose: Used to amplify a small quantity of DNA by several orders of magnitude
Step 1: DNA strands and complementary DNA primers are heated to 95 C for 15 seconds to separate the strands.
Step 2: The solution is abruptly cooled to 54 C to allow the
primers to anneal to each ssDNA.
Step 3: The solution is heated to 72 C and new complementary
strands are synthesized using Taq DNA polymerase.
Step 4: The cycle is repeated until the desired quantity of DNA is synthesized.
H-NMR Spectroscopy
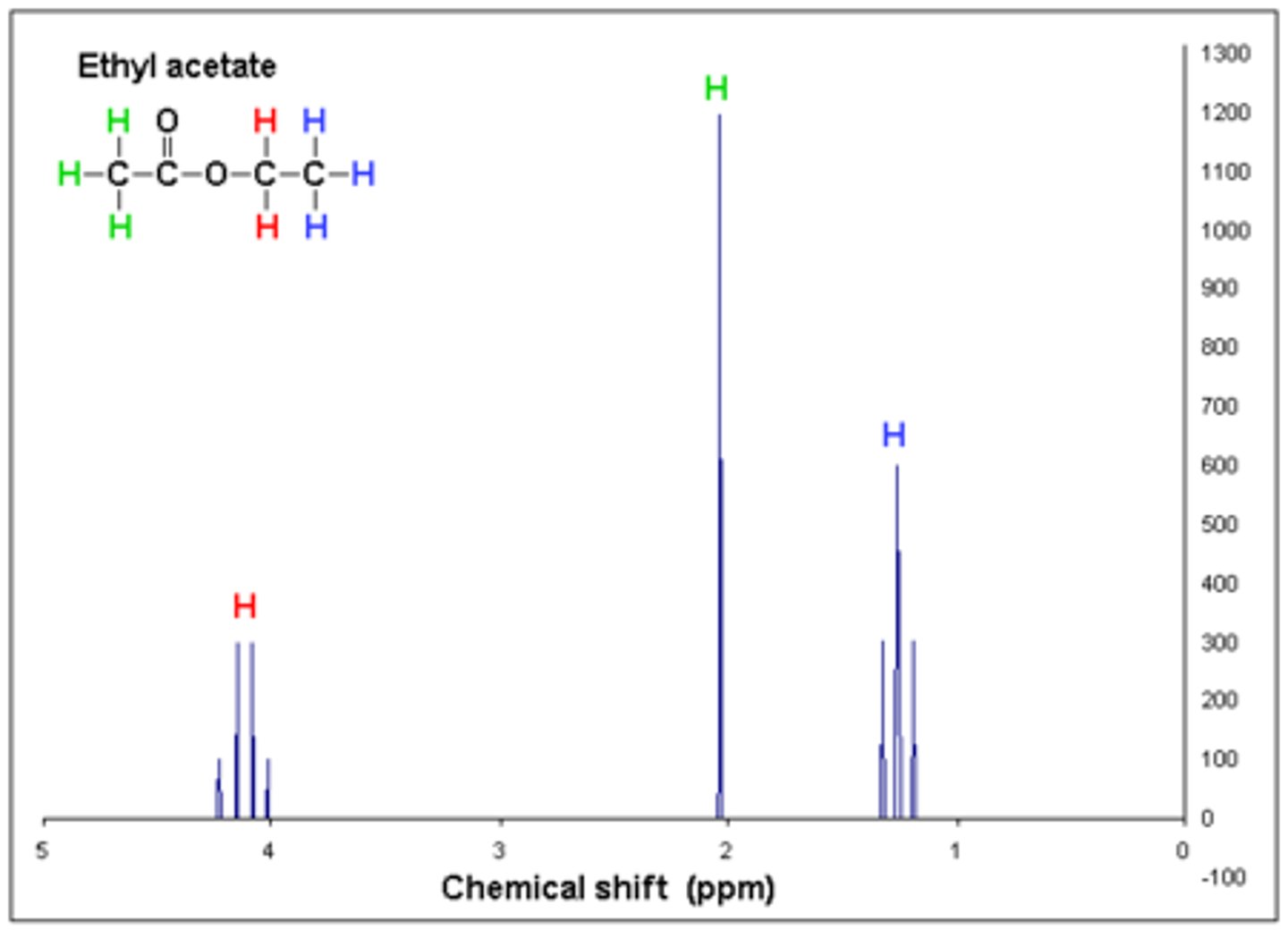
Chemical shift: The chemical shift on the x-axis (δppm) represents the amount of deshielding of electrons that is caused by an adjacent heteroatom or pi bond.
- 0 - 5 ppm → Alkane region
- 3 - 5 ppm → Alkane with a heteroatom region
- 5 - 7 ppm → Alkene region
- 6 - 8 ppm → Aromatic region
- 9 - 10 ppm → Aldehyde region
- 10 - 13 ppm → Carboxylic acid region
Integration: The integration of the peak determines the number of equivalent hydrogens a signal
represents.
Neighbors: The number of peaks determines the number of neighboring hydrogens that are ≤ 3 bonds away. The number of peaks equals the number of neighbors + 1.
- Singlet → No neighboring hydrogens
- Doublet → One neighboring hydrogen
- Triplet → Two neighboring hydrogens
- Quartet → Three neighboring hydrogens
- Quintet → Four neighboring hydrogens
- Sextet → Five neighboring hydrogens
- Septet → Six neighboring hydrogens
- Multiplet → Seven or more neighboring hydrogens
IR spectroscopy
IR spectroscopy is a method that is used to identify certain functional groups within a molecule. Only molecules that have a dipole moment will show absorbance. The x-axis is reported in wave numbers (1/cm) and the y-axis is reported in % absorbance.
Important regions:
- 1700 - 1750 → Carbonyls (sharp peak)
- 1720 - 1740 → Aldehydes
- 1700 - 1725 → Ketones
- 1735 - 1750 → Esters
- 1700 - 1725 → Carboxylic acids
- 3200 - 3600 → OH groups (broad peak)
- 3300 - 3400 → Amines
The number of peaks are relative to the number of hydrogens on the amine (e.g., 1 degree amines will have two peaks, 2 degree amines will have one peak)
UV-Via Spectroscopy
As the number of conjugated pi bonds increase, the
energy gap between the highest occupied molecular
orbital (HOMO) and lowest unoccupied molecular
orbital (LUMO) decreases, which means that light of a
lower energy is absorbed.
This results in light with a longer wavelength to be emitted (as seen on the right). If a molecule absorbs green light, we see it as red.
mass spectrometry
Purpose: Used to determine the molecular weight of a compound and aid in determining the molecular structure.
In mass spectrometry, the sample is vaporized and subjected to ionizing conditions. The charged molecule collides with an electron, resulting in the ejection of an electron from the molecule, making it a radical. The charged radical can undergo fragmentation or being detected.
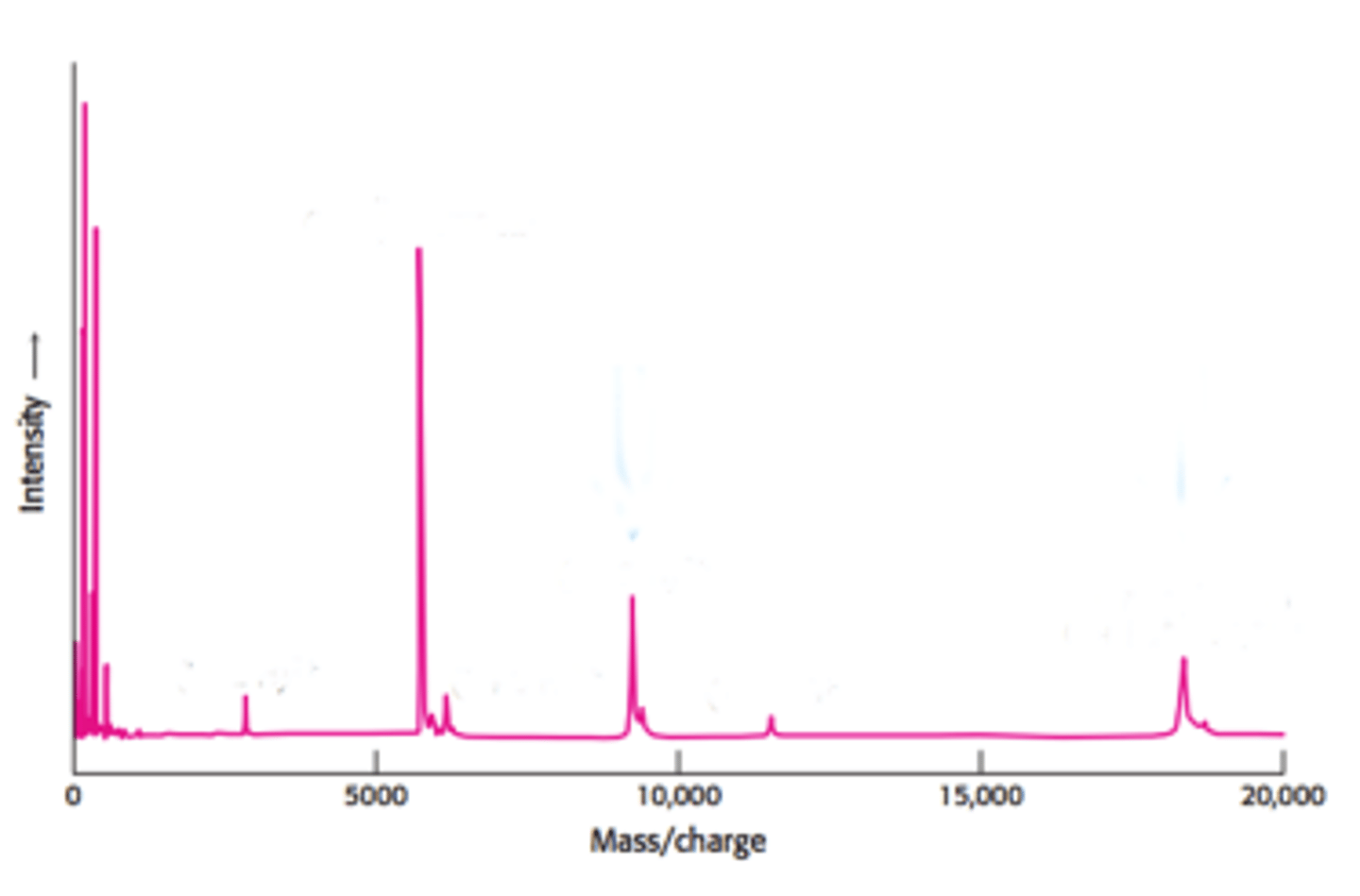
The x-axis represents the mass/charge ratio (m/z), which
essentially just means the molecules mass using only the lowest isotopes of the atoms involved (e.g.,
12C, 1H, 35Cl). The y-axis represents the intensity, or relative abundance of the molecule, usually given as
a percentage.
Base peak: The tallest peak. This does not always represent the actual intact molecule, as it may
sometimes be a fragment of the molecule that is found in higher abundance.
Molecular ion peak (M): The peak that represents the molecule. The m/z value of this peak represents
the molecular weight of the molecule.
M+1 peak: The relative abundance of 13C in the molecule. Found in a relative abundance of 1.1%. So if
there is a M+1 with a m/z value of 4.4, it means that there are 4 carbons present (4.4/1.1 = 4).
M+2 peak: The relative abundance of either 37Cl or 81Br in the molecule. 37Cl will be found in a 3:1 ratio
relative to the M peak (e.g., if the M peak has a relative abundance of 90%, if the M+2 peak is at 30%, it
means there is chlorine present in the molecule). 81Br will be found in a 1:1 ratio relative to the M peak (e.g., if the M peak has a relative abundance of 90%, if the M+2 peak is also at 90%, it means there is bromine present in the molecule).
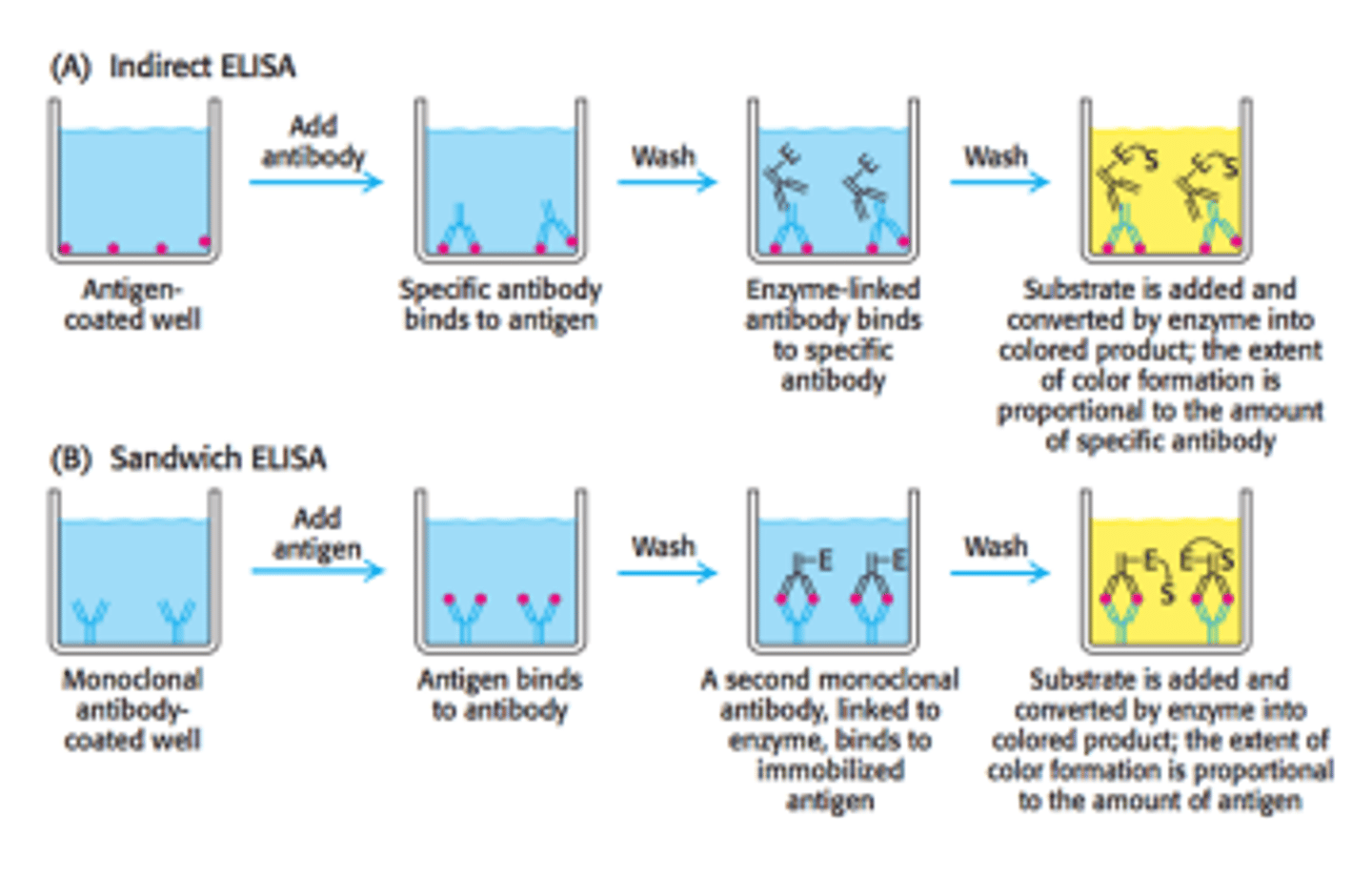
Enzyme-Linked Immunosorbent Assay (ELISA)
Purpose: Used to identify the concentration of a molecule of interest in a given sample
ELISA is an assay that uses primary antibodies that are specific to a molecule of interest and secondary antibodies that are specific to primary antibodies and are conjugated with a fluorophore, so their presence can be measured via spectrophotometry. There are two different methods of ELISAs that are used.
Gram Staining
Purpose: Used to differentiate bacteria into two groups, gram-positive or gram-negative, based on the
content of their cell wall
- Gram-positive bacteria: Appears purple on the slide. Contains a thick peptidoglycan layer that takes up
the stain.
- Gram-negative bacteria: Appears pink on the slide. Contains a thin peptidoglycan layer sandwiched
between two lipid bilayers that take up the safranin stain.
Tollen's Test
Tollen's reagent tests for the presence of an aldehyde and can distinguish between aldoses and ketoses.
Ketones do not react unless they are α-hydroxy-ketones. - A positive Tollen's test is characterized by the
precipitation of elemental silver.
Tollen's reagent consists of [Ag(NH3)2]NO3.
Benedict's Test
Benedict's reagent tests for the presence of an aldehyde. Ketones do not react unless they are α-
hydroxy-ketones. A positive Benedict's test is characterized by a change in color from clear blue to brick-red with the formation of a precipitate.
Benedict's reagent consists of a mixture of sodium carbonate, sodium citrate, and copper (II) sulfate
pentahydrate.
1 SDS for every ___ amino acids
binds 1 SDS for every 2 amino acids
SDS-PAGE gel type
polyacrylamide gel
SDS-PAGE only interrupts ____ bonds
covalent
_____ sized proteins are found towards the bottom of the gel in SDS-PAGE
small proteins at the bottom
reducing SDS-PAGE reduces ________ resulting in a completely __________ protein
reducing SDS-PAGE reduces disulfide bridges resulting in a completely denatured protein
chromatography purpose
separation of 2+ molecules from a mixture
liquid chromatography stationary/mobile phases
stationary: silica
mobile: toluene or another non-polar liquid
high-performance liquid chromatography
uses high pressure to pass the solvent through a more finely-ground stationary phase
gas chromatography
used to separate and analyze molecules that can be vaporized
gas chromatography mobile/stationary phase
mobile: an inert or unreactive gas (i.e. helium or nitrogen)
stationary: thin layer of liquid or polymer that surrounds the walls of a tube
gas chromatography higher retention time
polar molecules elute slower giving them a higher retention time
gel-filtration chromatography
used to separate molecules by size rather than polarity
what elutes faster in gel-filtration chromatography
larger molecules because they do not enter the porous gel beads
gel-filtration chromatography mobile/stationary phase
mobile: solution in the column
stationary: gel beads
cation-exchange chromatography
(-) charged beads/resins are used which attract (+) charged proteins
(-) charged proteins will elute first
ion exchange chromatography
molecules separated based on net surface charge
-stationary phase is made of either negatively or positively charged beads (attract & bind compounds that have opposite charge)
-salt is added to elute proteins stuck to column
Anion exchange chromatography
(+) charged beads/resins are used which attract (-) charged proteins
(+) charged proteins will elute first
affinity chromatography
separates proteins based on their affinity for a specific ligand
proteins with a ____ affinity will elute first in affinity chromatography
proteins with a low affinity will elute first in affinity chromatography
in TLC, _____ molecules move farther
non-polar
TLC stationary/mobile phase
stationary: polar silica gel
mobile: use capillary action to travel
simple distillation
separates 2+ molecules from a solution when their boiling points differ by 25C or more and the BPs are under 150 C
Fractional distillation
separates 2+ molecules from a solution when their boiling points differ by less than 25C
vacuum distillation
separates 2 molecules from a solution when their boiling points are high (>150 C) and risk changing chemically
1700-1750 sharp peak IR
carbonyls
3200-3600 broad peak IR
OH groups
3300-3400 IR
amines
primary: 2 peaks
secondary: 1 peak
.... sized molecules will move more in gel electrophoresis
smaller molecules
in gel electrophoresis negatively charged molecules travel towards the ... while positively charge molecules travel towards the ...
- charged molecules go to anode
+ charged molecules go to cathode
(opposite charges = attraction)
location of cathode/anode in gel electrophoresis
cathode (-): top
anode (+): bottom
gel electrophoresis follows the same principles as ....
electrolytic cell b/c a voltage source is applied (non-spontaneous, requires a battery, anode is + and negatively charged species are attracted to it)
gel for large molecules in gel electrophoresis
agarose
gel for small molecules in gel electrophoresis
polyacrylamide
purpose of gel electrophoresis
separation of proteins, DNA, or RNA based on size and/or charge
What is the underlying goal behind extractions, distillations, and chromatographies?
to separate two or more compounds by their characteristics