Day 2 (molecular analysis)
1/48
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
49 Terms

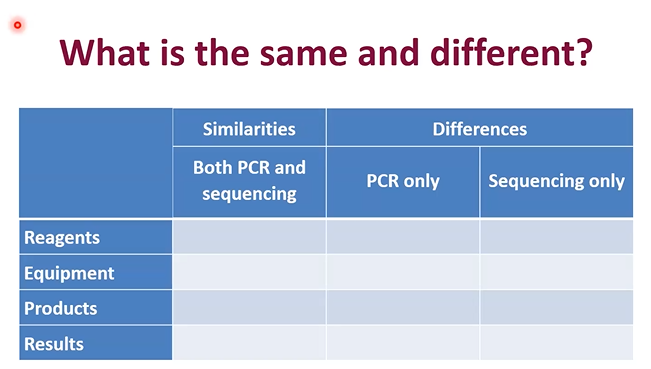
What is southern blot used for
Use this when you want to detect a specific DNA sequence in a DNA sample after cutting the DNA and separating the fragments on a gel.
What is FISH used for
Use this when you want to see whether a specific DNA sequence is present and where it is located in chromosomes, cells, or tissues.
What is DNA library used for
Use this when you want to store and screen many DNA fragments from an organism to find a gene or sequence of interest later.
What is PCR used for
Use this when you want to make many copies of one specific DNA region so it can be analyzed more easily.
What is a genomic library
When you take the genome of an organism, cut it up into a bunch of pieces, and insert them into a bunch of plasmids, and finally insert them into E.coli cells.
When should you use Sangar sequencing
Use this when you want to determine the exact base sequence of a relatively short DNA fragment.
When should you use Genetic testing
Use this when you want to check for mutations, inherited conditions, or genetic differences in an individual.
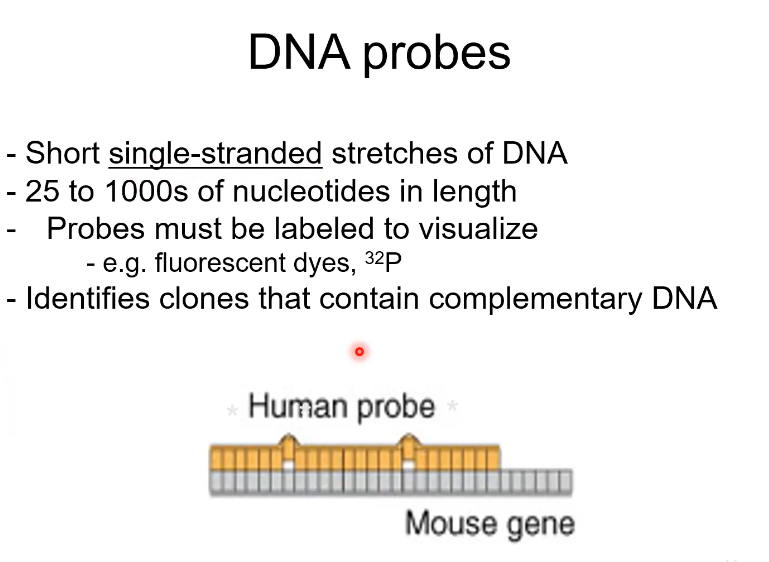
What is a DNA probe
A short DNA strand
25-1000s of bases long
They are labelled (like dyes) so you can isolate them
It can Identify clones that have complementary DNA
Like how drug testing is done on rats first instead of humans. We use a human probe
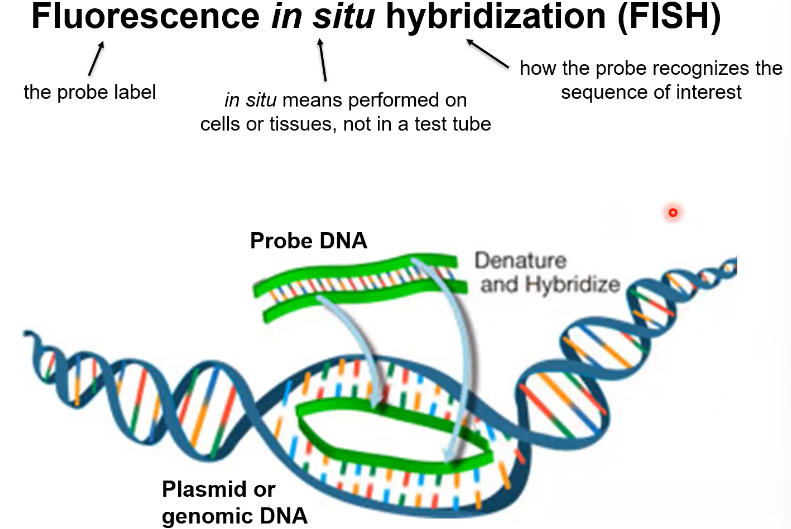
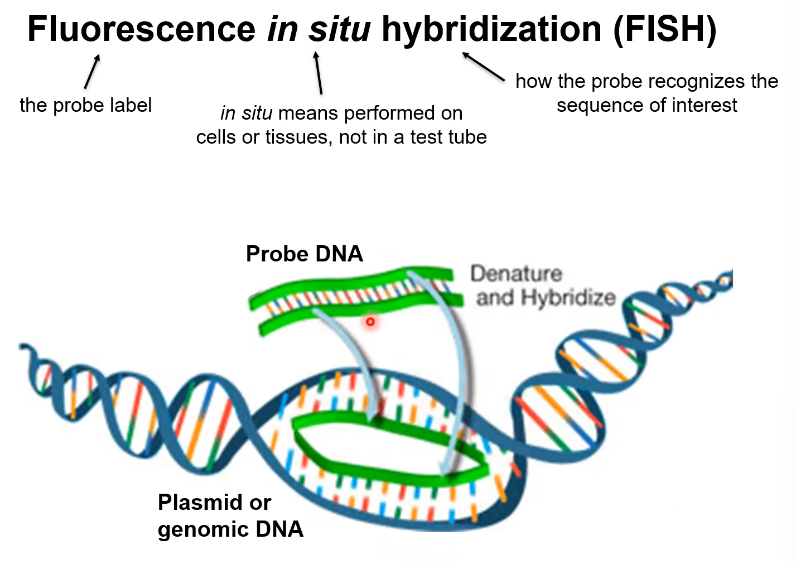
What does FISH stand for
Fluorescence in situ hybridization
What is FISH
It is a method to find a DNA sequence that was labelled with a probe that has attached to complementary DNA
How does probe DNA need to be added to a genome (general)
You need to sperate a section of the DNA and then add the Probe
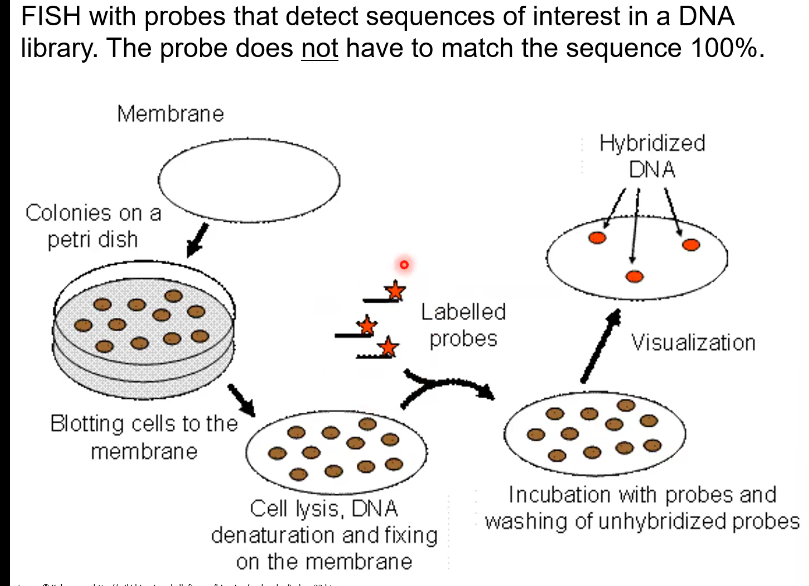
What are the steps of FISH
A membrane is pressed onto the original plate with the bacteria on it (like the velvet fabric experiment)
Cell lysis (the cell is broken open)
DNA denaturation (The DNA is split into single strands)
DNA fixing (when the DNA is attached to the cell membrane)
Then labelled probes are added and attach to some of the DNA
Then the cells are washed (Get rid of the cells that didn’t take in the probes)
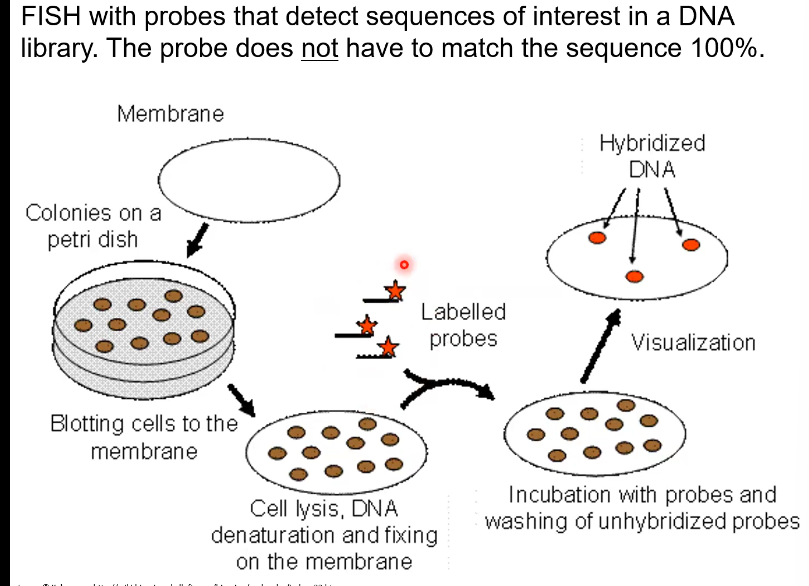
Can you replicate the cells that show a positive result from the FISH method
No, because they are dead after lysis (The cell is open). But you can locate the Identical cells on the original plate because you place the agar plate onto it, so they’re position is the same.
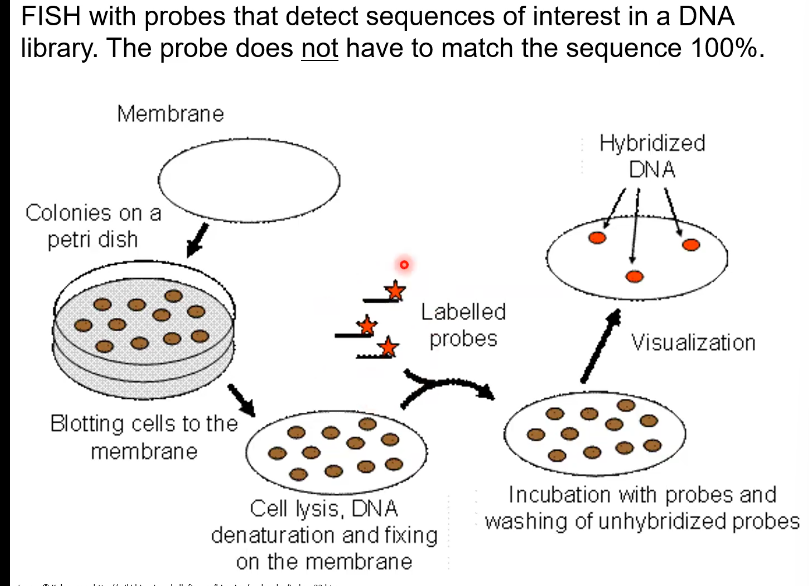
How can DNA probes be made
Chemical synthesis or PCR
Do probes have to be identical to the cell they’re going onto
No, they can have about 20% difference between them
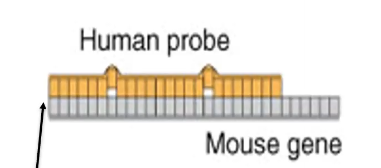
What is the probe base sequence designed after
It is designed after the specific target DNA sequence you are targeting
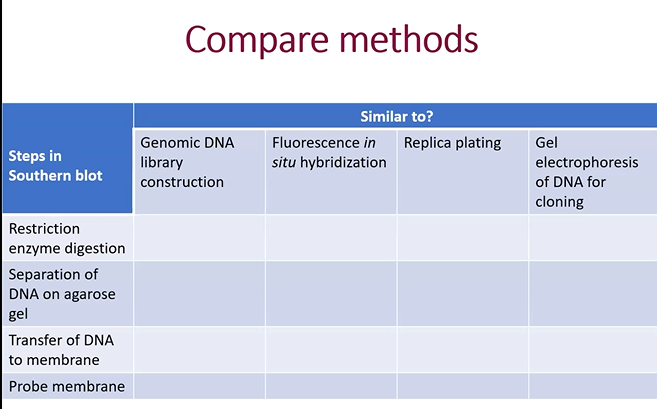
What is southern Blot analysis
It is when you break up a genome with restriction enzymes and put them on an electrophoresis gel and see where the DNA ends up. (Bigger DNA closer to negative side, smaller DNA closer to positive side).
You can then observe them in a fancy machine, or a simple large contraption.
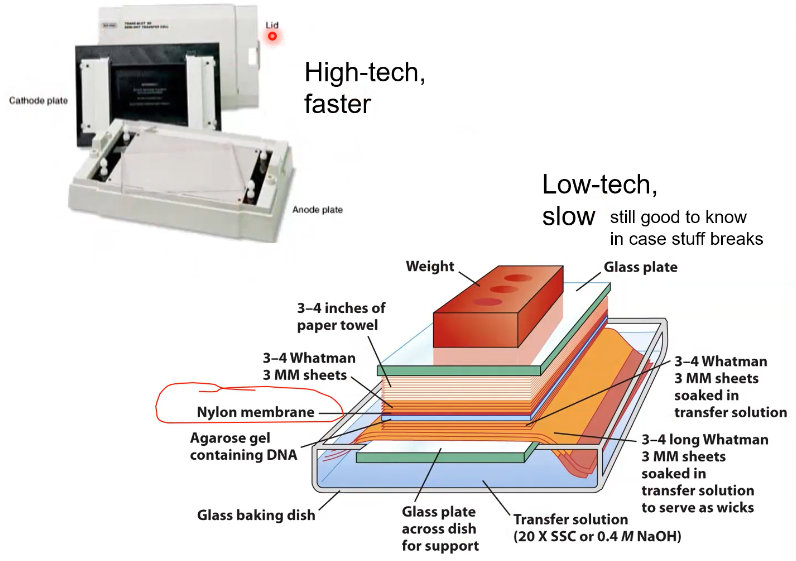
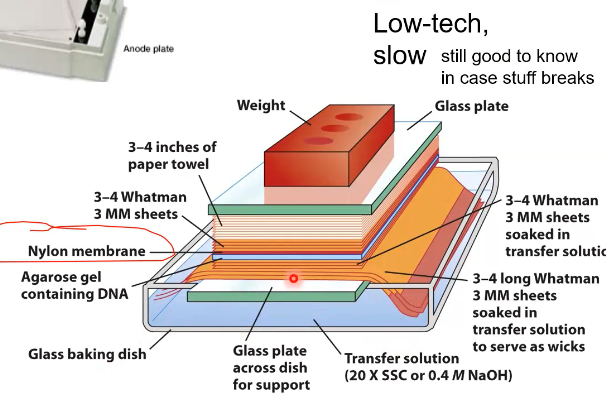
What is the goal of this large contraption
To get the DNA from inside of the electrophoresis gel to the surface of the Agar plate. (why the weight is needed to push down the contraption)

What is the goal of this high-tech machine
To move the DNA from the electrophoresis gel, to the agar plate. This is done by electrical current pulling the DNA up out of the gel.

Solve
A, because Gel electrophoresis for DNA cloning doesn’t use Restriction enzymes to break the DNA into a bunch of parts
What does PCR stand for
Polymerase Chain Reaction
What is PCR
It is a method for making a bunch of copies of a DNA sequence
It can be observed using electrophoresis
It is used in research and diagnostics
What else is PCR used for
It is used for paternity testing, finger prints, HIV testing, etc
What does PCR use for it’s primer
It uses DNA instead of RNA
What are dNTP’s
They are free nucleotide bases floating around
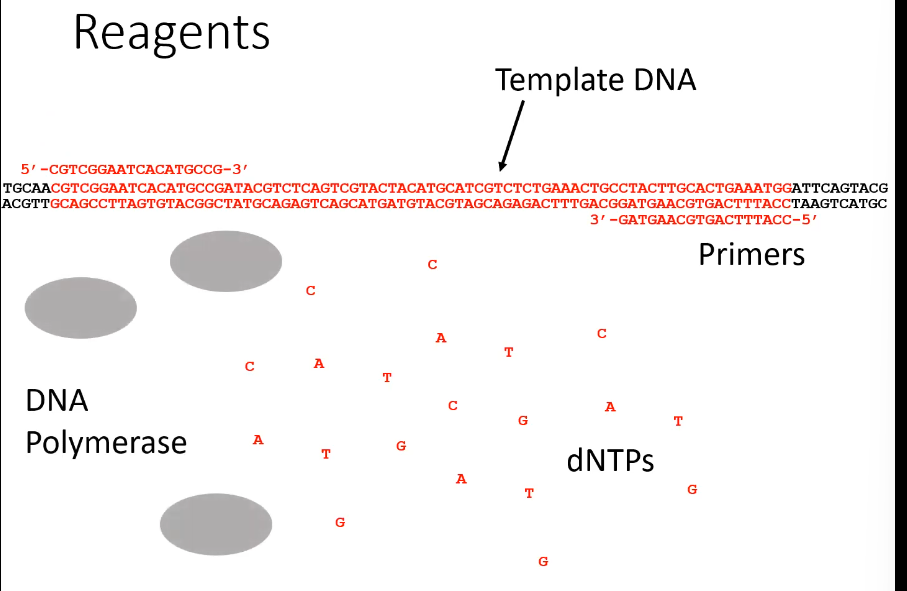
What components are needed for PCR
The target DNA, DNA primers, DNA polymerase, free nucleotide bases, and a buffer solution
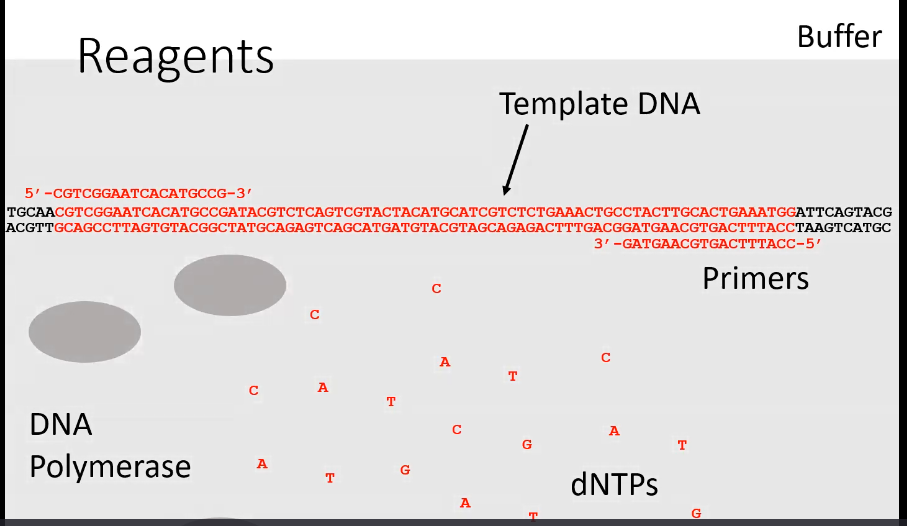

What is the general process of PCR
Heating and cooling of the components
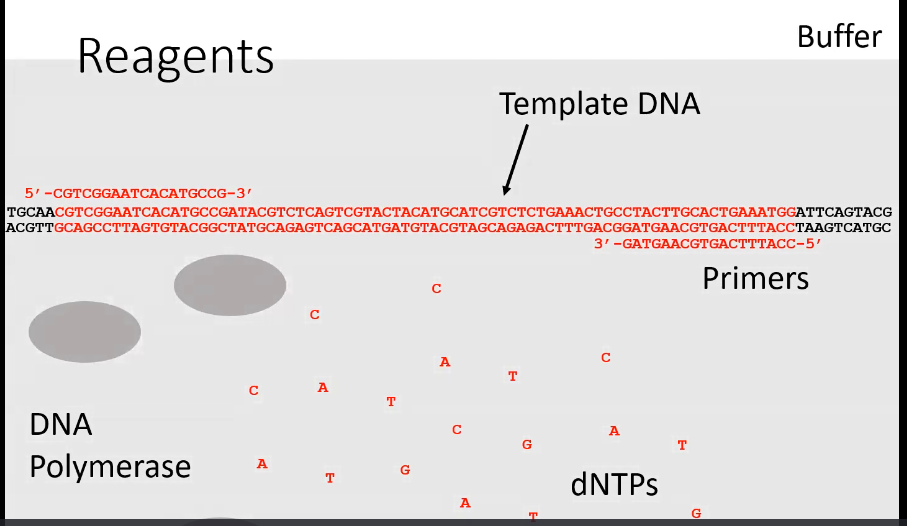

What are the steps of PCR
Denature (95-98 degrees Celsius)
Separating the strands of DNA
Anneal (a range of temperatures)
Allows the DNA primers to pair with the separate strands of DNA
Extend (68-72 degrees Celsius)
Allows the DNA polymerase to extend the DNA primers using the free nucleotide bases
This cycle repeats several times
What does step 1 of PCR look like and Name
Denature
Strand of DNA has separated into single strands
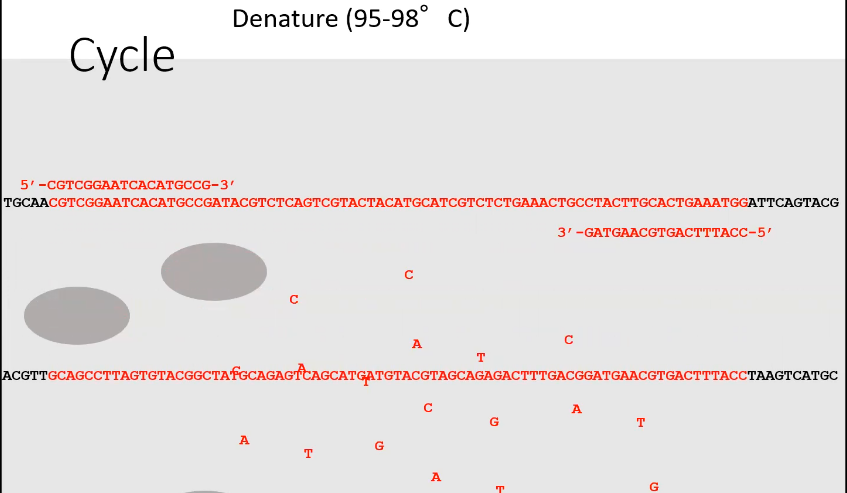
What does step 2 of PCR look like and called
Anneal
DNA primers have paired with the DNA section that is complementary to it
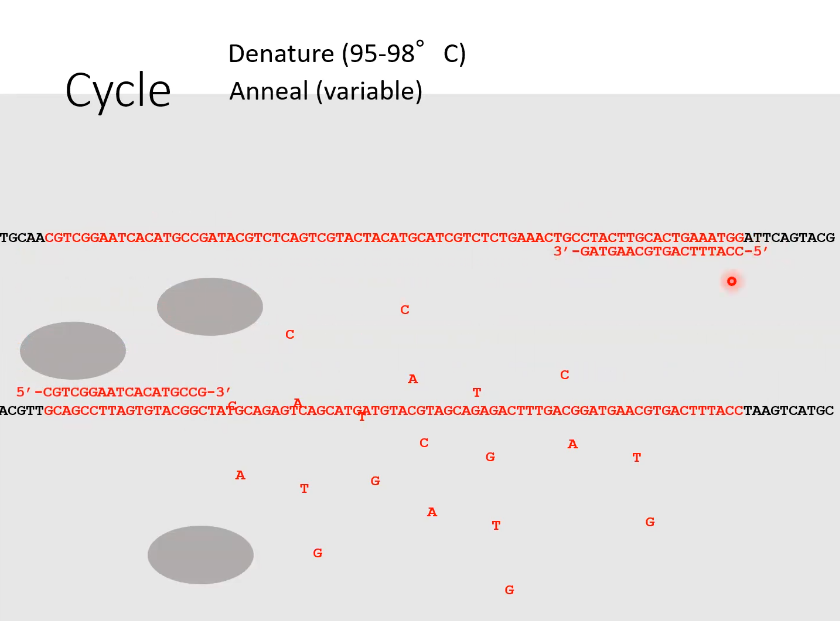
What is the 3rd step of PCR and it’s name
Extend
Allows the DNA polymerase to extend the DNA primers using the free nucleotide bases

How does the DNA polymerase know when to stop copying DNA
It doesn’t, the way the target section of DNA gets multiplied is by the DNA primers switching sides of the DNA to start replicating on for each replication.
This means that after the 2nd replication, only the target section of DNA will be replicated.

How fast does PCR replicate the target section of DNA
Very fast, since the number doubles every replication (2, 4, 8, 16, 32…)
What is the sanger method used for
Smaller DNA sequences, similar to PCR
What end does DNA polymerase build off of
the 3’ end
What componenets does the Sanger method have
Same as PCR, but with ddNTP’s
What is pretty much the only 2 differences between PCR and the Sanger method
PCR has the DNA polymerase go as far as they can, the Sanger method uses ddNTP’s to tell them when to stop replicating.
PCR also has 2 primers while Sanger only has 1.
What are the steps of the Sanger method
DNA polymerase connects the 3’OH to the next nucleotide at the 5’ phosphate (3’OH is important)
Then Chain termination is used. This creates a bunch of fragments of DNA.
Then we use fluorescent labelling to tell which base caused the termination. (Each ddNTP has a different colour. Like ddATP, ddCTP, ddGTP, ddTTP.)
This results in a bunch of different length strands.

What is dNTP
A normal DNA nucleotide, that has a 3’ OH (dNTP)
What is a ddNTP
A DNA nucleotide that does not have a 3’ OH (ddNTP), just an H. This causes the DNA polymerase to be unable to add and Nucleotide bases to it.
This is called Chain termination sequencing
What is chain termination sequencing
When a DNA polymerase adds a ddNTP (a nucleiotide 3’OH it needs for DNA polymerase to extend the DNA strand. So the replication stops.
What is capillary electrophoresis
When you add the DNA strands produced from the Sanger method to a vertical tube and allow the different ddNTP strands to fall to different heights.
Then smaller strands fall to the bottom, bigger ones stay at the top. A detector at the bottom scans which ones pass through first.
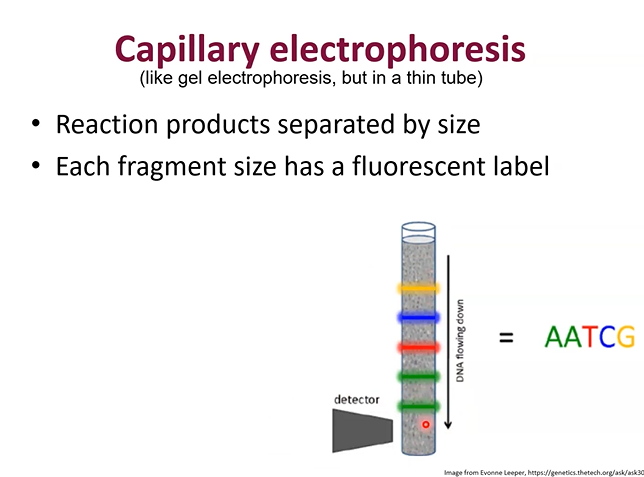
How do you use the results from the electrophoresis
You upload them into 2 files, 1 will be a graph, 1 will be just values.
Each peak qin the graph is a band from the electrophoresis passing the scanner.
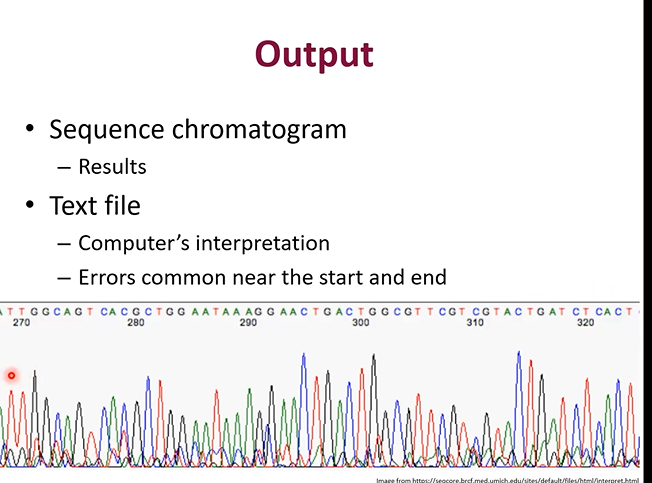
How should you interpret the Electrophoresis graph
the peaks are the bands passing the scanner
The peaks should be evenly spaced apart
The peaks should be well above the baseline noise, or else it should not be considered a peak
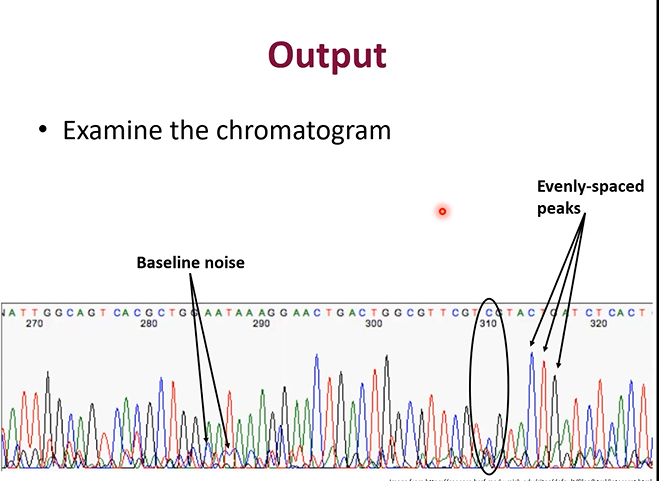
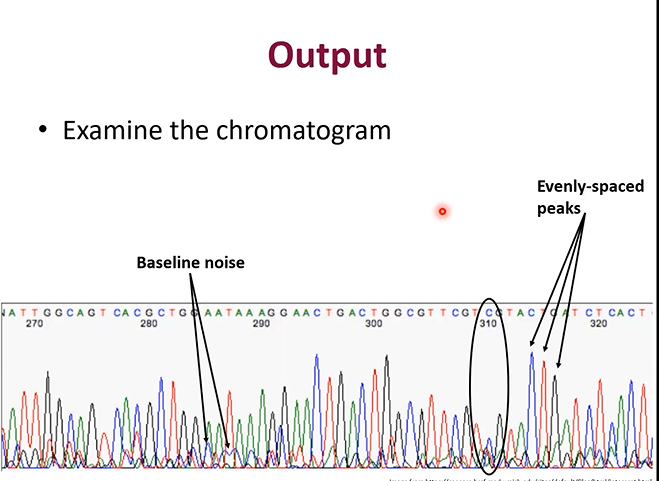
Why are the peaks different height in the electrophoresis graph
Because just by chance, there may be more of a certain ddNTP than others.