HLSC322 exam 4 - can't believe there's more!
1/16
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
17 Terms
Polycistronic vs Monocistronic mRNA
encoding several proteins from same gene vs. encoding only one protein from one gene, occurs in prokaryotes and eukaryotes respectively
Cis-Regulatory Element (CRE) vs. Trans-Regulatory Element (TRE)
both regulate transcription, though former is noncoding DNA sequence located on the same chromosome as the promoter, while latter is diffusible molecule encoded by gene on different chromosome
Levels of DNA condensation
Nucleosome, 30 nm fiber, 300 nm loops, 250 nm fiber, chromatid
Histone modifications
Histone methylation may either open or close chromatin / activate or repress transcription depending on location. Performed by histone methyltransferases
Histone acetylation generally opens chromatin and activates transcription (reduces + charge on histone tails, weakening affinity for DNA). Performed by histone acetyltransferases
Modifications are passed on to daughter cells ft. maintenance methylases / acetylases
Topologically Associating Domains (TADs)
foundational 3D structural units of chromatin, typically 200 kb to 5 Mb in size, where DNA interacts frequently within themselves but rarely with outside regions (important for gene regulation, i.e. correct enhancer-promoter interaction)
Chromatin remodeling proteins
multi-subunit protein complexes that expose DNA at promoter / CRE by removing / repositioning nucleosomes
Pioneer factors
specialized transcription factors that can bind to closed chromatin / nucleosomal DNA and induce local decompaction so other factors can begin assembling at promoter
Coregulators
proteins that interact with transcription factors to activate (coactivators) or repress (corepressors) gene expression by remodeling chromatin.
Histone methyltransferases
Histone demethylases
Histone acetyltransferases (HATs)
Histone deacetylases (HDACs)
ChIP-seq
Chromatin Immunoprecipitation sequencing, a technique to determine which DNA sequences are bound to a certain protein
Cross-link DNA and bound proteins, incubate in antibody specific to protein of interest, pull proteins of interest out, separate out DNA and sequence
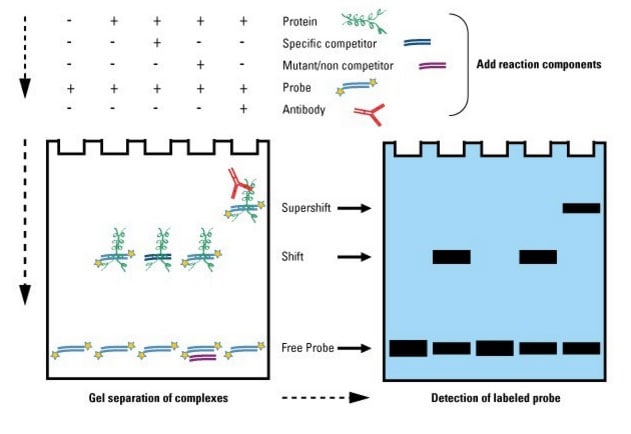
Gel shift assay
anotha technique to define DNA-binding sites on a protein.
A DNA sequence (probe) thought to contain the binding site is labeled (eg. with radioactive phosphorus isotope 32P)
Running the gel:
Lane 1: probe only
check for purity of DNA and obtain baseline migration speed to compare to protein-bound complex
Lane 2: probe + protein suspected to bind
if binds, should migrate slower than 1 = “shift”
Lane 3: probe + protein + competing DNA
should see no shifted band, indicating competitor used up protein binding sites → binding site was correct
Lane 4: probe + protein + noncompeting DNA
check that binding is actually at particular sequence and not just binding to any random DNA
Lane 5: probe + protein + antibody
confirm identity of protein. If binds, should migrate slowest = “supershift”
Reporter gene
molecular tool used to study gene regulation by encoding easily detectable proteins—eg. fluorescent proteins (GFP), enzymes (lacZ/𝛽-galactosidase), or luminescent proteins (luciferase)—that produce measurable signals.
By attaching these genes to a target promoter, researchers can measure promoter activity, visualize protein localization, or monitor cellular signaling pathways in real time
Basal transcription factors
bind to core promoter (tho not necessarily directly to DNA itself) and recruit RNA polymerase
TATA box
conserved DNA sequence (usually TATAAA) in eukaryotes, usually 25-35 bp upstream of transcription start site. Binding site for TATA-binding protein (TBP) which recruits other TFs and eventually RNAP
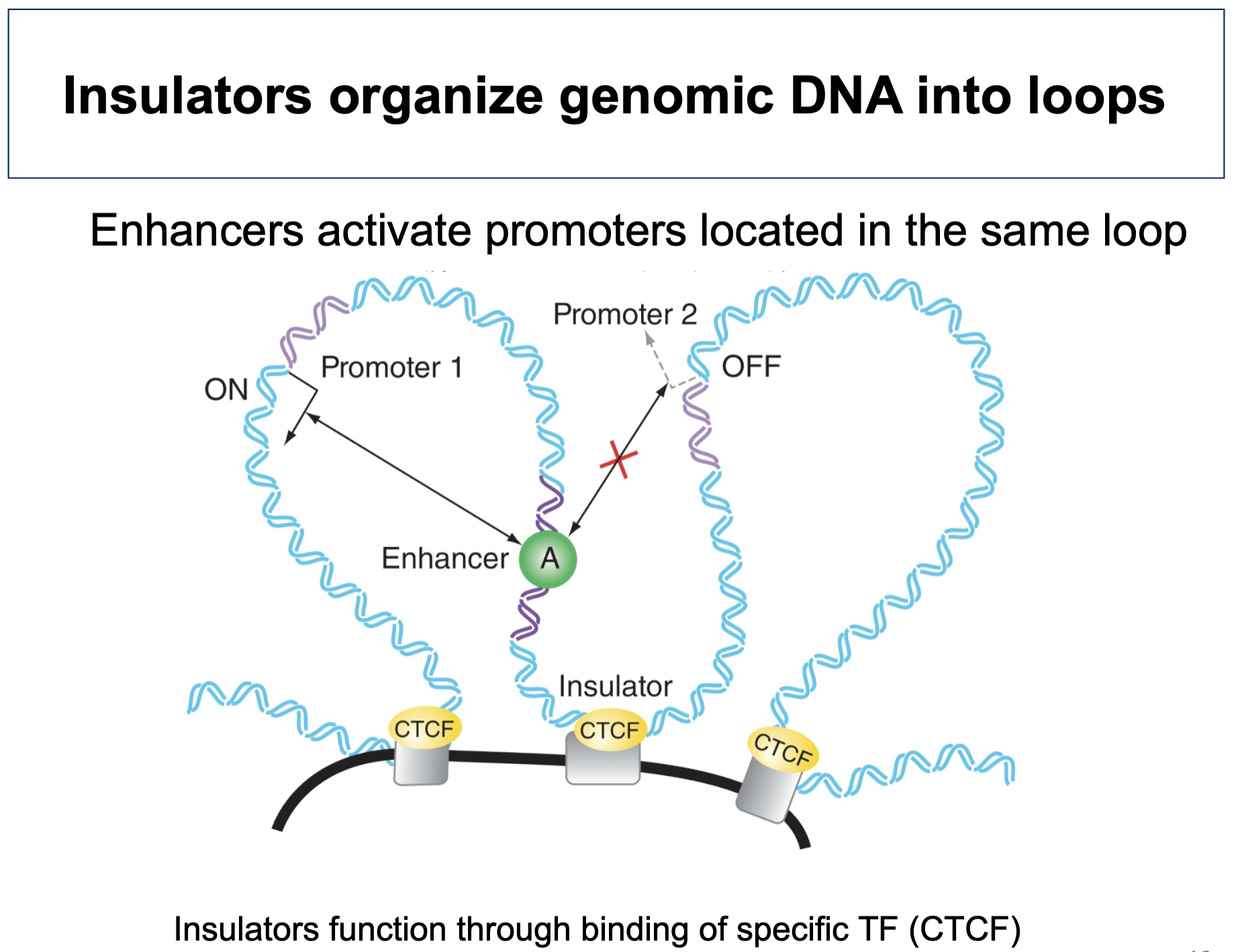
Insulators
sequences between enhancer and promoter that prevent the 2 from interacting, often involving CTCF binding and formation of chromatin loops as physical barriers
CCCTC-binding Factor (CTCF)
binds DNA ft. zinc fingers
establishes TADs
binds insulator sequences → DNA loops wherein enhancers can only activate promoters in same loop
Jun & Fos
Transcription factors important for cell growth, proliferation and differentiation (expressed more highly in response to growth factors and/or stress).
Jun-Fos heterodimers are more stable / have higher DNA binding affinity than Jun-Jun homodimers, so they are more common in most cells.
Jun-Fos bends DNA towards major groove while Jun-Jun bends towards minor groove.
Myc & Max
Transcription factors with the same DNA-binding sequence and dimerization domain, but only Myc contains trans-activation domain
Myc-Max heterodimers have higher binding affinity than Max-Max homodimers
Myc will displace one of the Max and form the heterodimer → activate cell proliferation genes