Genetics Exam 1 (Rutgers Soliman)
1/151
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
152 Terms
Nucleic Acid
This must be able to contain complex information, replicate, encode the phenotype, be different in phenotypes, and must be complex so it can be carried from one organism to another
DNA
Found in the nucleus and can be found in mitochondria, viral, bacterial, etc. and can exist as double stranded
RNA
Found in the nucleus as a messenger, ribosomal, transfer, anti, etc. and have an Uracil base with single and double stranded
Rough Strain
The nonvirulent form that is unable to invade the immune system due to having no outer capsule
Smooth Strain
The virulent, wild type form that can invade the immune system with an outer capsule
die
When the mice were injected smooth strain bacteria, did they survive or die?
survive
When the mice were injected rough strain bacteria, did they survive or die?
survive
When the mice were injected heated smooth strain bacteria, did they survive or die?
die
When the mice were injected heated smooth strain mixed with rough strain bacteria, did they survive or die?
Transformation
What did Fred Griffith conclude about his results?
Avery, Macleod, and McCarthy
The experiment was meant to prove that DNA was the transforming principle through enzymes like proteases, RNAses and DNAses
Hershey & Chase
This experiment used bacteriophages to examine traces of DNA through replication of those viruses with radioactive sulfur or phosphorous (proved that DNA was genetic material continuously replicating)
Rosalind Franklin
Did her research on DNA structure and gathered an X-ray diffraction image of DNA which concluded that DNA is helical, contains 10 nucleotides and have a diameter of 2 nm
Watson & Crick
Discovered more points about DNA structure and concluded that DNA is double stranded & runs antiparallel and have complementary pairing
Grooves
Meant to make the DNA more visible for protein bidning
Nucleoside Analogs
Can be used to stop cancer cell from dividing through with no OH groups on the end
C-G
This pairing has 3 bonds
A-T
This pairing has 2 bonds
Conservative
When the original strand served as a template for a new molecule of DNA
Dispersive
When both DNA molecules break down like fragments and serve as a new template
Semi-conservative
The original strand unwinds and used as a template to generate a new strand
DNA was semiconservative! As when the 2 different samples of Nitrogen was added, they ended up making a half/half of one another.
In the Meselon & Stahl Experiment, what did they conclude after growing the radioactive Nitrogen?
Theta Replication
A bidirectional step process that takes place in circular DNA like Ecoli
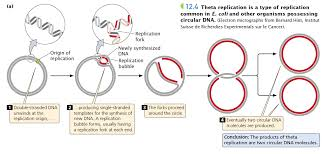
1) Double strand will begin to unwind produce single strands which can function as a template for the new strand at the replication bubble (reminder that the plasmid is melted first)
1st step in Theta Replication
Unwinding is continuous and the bubble gets larger (location where there is unwinding is replication fork)
2nd step in Theta Replication
3) DNA synthesis would occur in the 5’ to 3’ direction
3rd step in Theta Replication
Rolling Circle Replication
Takes place in typically viruses and Ecoli where there would be a single break unidirectionally
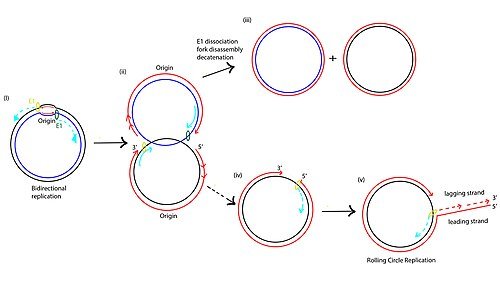
A single strand break in 3’ OH & 5’ P group
1st step in Rolling Circle Replication
2) A new strand is added to the 3’ OH break using the inner strand as a template
2nd step in Rolling Circle Replication
The original strand will be displaced and roll off the entire plasmid (and continues until it has several copies) as the linear fragment can be used as another template!
3rd step in Rolling Circle Replication
Intitation
When a DNAa protein will bind to the origin of replication to cause a conformational change to twist and unwind the region
Helicase
Will undo hydrogen bonds
Single Strand Binding Proteins
Will bind to the DNA to stabilize DNA strand once unwound
DNA gyrase (topoisomerase)
Will prevent supercoiling and upstream torsional strain to build up during unwinding
Topoisomerase I
Create single stranded breaks in DNA
Topoisomerase II
Creates double strand breaks in DNA as it passes through double helix and reseals it
Elongation
The addition of new nucleotides on an old template to make a new strand
Primers
Needed for a 3’ OH groups so nucleotides can be added → generated by primase so 10 bps of RNA can be generated
DNA Polymerase III
Replicates DNA using RNA primer (5’ to 3’ for replication and 3’ to 5’ for exonuclease activity)
DNA Polymerase I
Removes the RNA primer and replaces it with DNA (works on lagging strand) with 5’ to 3’ for replication & exonuclease activity to remove primers and replace & 3’ to 5’ for exonuclease activity to backup and check errors
DNA Ligase
Will put together nicks once polymerase I removes and replaces primer
Termination
Two ways that this process can work:
1) DNA can termination once 2 forks meet
2) Some needs the protein tus to bind to the ter sequence so the protein can block replication
Leading Strand
Reads from 3’ to 5 but synthesized from 5’ to 3’ direction since it is continuous and starts from replication bubble
Lagging strand
Reads from 3’ to 5 but synthesized from 5’ to 3’ direction since it is discontinuous and starts from replication fork
Licensing Factor
Like a signal to tell DNA polymerase that DNA replication can intitate in euykaryotes (geminin will bind and remove licensing factors)
DNA polymerase Alpha
Has primase activity in DNA for eukaryotes
DNA polymerase Eplison
After primers are added on, will act on the leading strand
DNA polymerase Delta
After primers are added on, will act on the lagging strand
Trans Lesion Polymerase
This polymerase can bypass error if necessary by adding an incorrect base
Nucleosomes
When histone proteins come together with the DNA wrapped around it (8 histone proteins)
Telomeres
These typically shorten which is part of the normal aging process, but kept when needed for critical genes (if shortened too quickly, this can cause cancer)
Tus Protein
Will bind to this if termination needs to occur (not all DNA needs)
Only one strand (template strand)
How many strands are copied in transcription?
No
Is the promoter transcribed into mRNA?
The promoter needs to be found, with the consensus sequence known as the TATA (bacteria & eukaryotes) or TTGACA in the bacteria
1st step of transcription
pre-mRNA (or mRNA in prokaryotes) will be transcribed by RNA polymerase (RNA polymerase II in eukaryotes) from 5’ to 3’
2nd step of transcription
Rho Independent & Dependent
What are two ways in which transcription can be terminated?
1) RNA polymerase stops at terminator RNA
1st step of Rho Dependent
Rho will attach to a sequence known as Rho ultilization site
2nd step of Rho Dependent
Rho will move to the 3’ end and unwind DNA
3rd step of Rho Dependent
Rho independent terminators
There could be an inverted repeated sequence that causes the formation of secondary structure
Eukaryotic transcription
This type of transcription that has more complexity and uniqueness for genes depending on level of expression as nucleosomes are gripping tightly to DNA
Acetyl Group
Groups that will bind to histones to loosen DNA
RNA polymerase I
Synthesize rRNA
RNA Polymerase II
synthesize mRNA, snRNA, miRNA
RNA Polymerase III
Synthesizes tRNAs, small rRNAs
Use a protein to bind to downstream sequence
How does Poly I terminate?
Works for 100s nucleotides past its stop (Rat1 or xrn1)
How does Poly II terminate?
Transcribing a long strand of Uracils
How does Poly III terminate?
Splicing
This is when a consensus sequence can help form the spliceosome that is responsible for the removal intron
Polyadenylation
Poly-A-Tail (5-20 Adenine) is added to the end thanks to a signal
Guide mRNA
These bind to the mRNA to do last minute editing
AUG
Start codon
UAA, UAG, UGA
Stop codons
Degeneracy & Wobble
Different codons coding for the same amino acid as the first 2 nucleotides pair normally according to watson and crick
Poly-A-Tail, Cap binding proteins (poly A bind to cap binding proteins)
Example of initiation factors in translation
Shine Delgarno sequence
UAAGGAGGU is the what sequence
small subunit in ribosome
What will bind first in the mRNA?
Elongation
Will continue from the A site and elongate from there (only AUG binds to P-site)
Termination
When release factors bind to the ribosome from the A site as the polypeptide is hydrolyzed from the P site
Cis Acting Factors
Factors that will affect transcription but do not move around the cell
Trans Acting Factors
Factors that will affect transcription but move around the cell
Nonsense Mediated mRNA Decay
Will handle premature stops when the ribosome reaches the stop codon as the release factor will bind and junction proteins will sense that there is something wrong with the mRNA and degrade it.
Nonstop Mutation
This mutation can be stopped by having an empty A site in the end, and signaling an enzyme to degrade the mRNA
Enhancers
Can promote direction in transcribing for the lac operon.
Silencers
Have negative control over expression
DNA binding proteins
Will bind to DNA binding domains and recognize them.
Helix Loop Helix
Has the secondary structure that can be crystalized to find the DNA binding domain
Zinc Finger Domain
The DNA binding protein can find this sequence and bind to it so it can carry on positive and negative regulation
Leucine Zippers
Will wrap around the DNA and can control positive and negative regulation
Regulatory Gene (aka Lac I)
Will make the repressor or activator
Promoter
This is where RNA polymerase binds to before it begins transcription
Operator
Will have the repressor bind to it
Lac Z
Can make beta galactosidase that will break down to glucose and galactose
beta galactosidase
break down to glucose and galactose (converts lactose to allolactose)
Lac Y
Encodes permease that permits entrance of lactose into the cell
Permease
Permitting entrance of the cell
Lac A
Forms transacetylase
Glucose
This can help regulate the operon because the operon (lac) prefers this over lactose
CAP (when cAMP binds to CAP protein)
Will bind to a site that will increase transcription in the operon