BCH LE 6 gene 1-3
1/259
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
260 Terms
Centromeres are located in the?
Middle of the chromosome
(?) is an adeninne-thymine (A-T) rich region.
Centromere
Telomeres are located?
At the end of the chromosome
(?) are Thymine-Guanine rich region.
Telomeres
What becomes shortened every DNA replication and division?
Telomere
What can be uses to determine how old one’s body is based on?
Length of Telomeres
Shorter telomeres =?
aging and age-related diseases
What is responsible for the coiling and packaging of DNA?
Histones
(*) Histone + DNA wrapped around it is called? =
Nucleosome!
What is chromatin composed of? (4)
Double stranded DNA
Histone
Non-histone
RNA
(?) proteins are responsible for
DNA replication and repair
RNA synthesis
Transport
Protein Translation
Non-histone proteins
Different areas of a Chromatin can either be Transcriptionally active or inactive.
what do you call:
(1) areas that are active?
(2) areas that are inactive?
Euchromatin = active
Heterochromatin = not-active
What is the basic unit of the Chromatin?
Nucleosome
Levels of Organization:
Nucleosome
chromosome
chromatin
arrange from smallest to largest
Nucleosome → Chromatin → Chromosome
Histones come in (?) parts or (?) in structure
8 parts or octameric in structure
What are the 4 main units in the histone?
H3 + H4
forms the tetramer (4 units)
Two H2A + H2B dimer
joins and completes the Octamer
Histones are (acidic/basic?)
why?
Histones are basic!
DNA is acidic
opposites attract = DNA wraps tightly around histones
H1 is known as
linker histone
not part of the core octamer
can easily be removed
this makes chomatin to be more soluble and easily worked with
What are Post -Transnational Modifications (PTMs)?
chemical groups added to histone tails after the protein has been made
tells the cell to OPEN or CLOSE the DNA
Two types:
Acetylation
Methylation
What does Acetylation do to DNA?
add acetyl group
relaxes grip between histoned and DNA
Makes DNA accessible for replication and transcription
Acetylation of
Histones H3 and H4 →
Core histone →
Phosphrylation of H1 →
ADP-ribosylation of histones →
Methylation of Histones
Monoubiquitylation of histones→
Sumoylation of histones →
Acetylation of
Histones H3 and H4 → activate or inactivate Gene transcription
Core histone → chromosomal assembly of DNA replication
w/o acetylation = dna remains tightly bound around histones and cannot separate.
Phosphrylation of H1 → Condensation of chromosomes during replication cycle
ADP-ribosylation of histones → DNA repair
Methylation of Histones → Activation or repression of Gene transcription
Monoubiquitylation of histones→ Gene Activation or repression + Gene Silencing
Sumoylation of histones → Transcription repression!
What does Methylation do to DNA?
Add methyl group
(activate) OR (inactivate) gene transcription
depends on WHERE the methyl group is placed
****Phosphorylation at H1 is associated with?
Condensing chromosomes during the Replication cycle
ADP-ribosylation: Linked to
ADP-ribosylation: Linked to DNA repair.
(?) is a versatile tag involved in activation, repression, and complete gene silencing (heterochromatin)
Monoubiquitylation
? is almost always associated with transcription repression (turning genes off).
Sumoylation: associated with transcription repression (turning genes off).
Sumo wrestler sits on u so u die (turn off)
According to the hierarchical assembly described in your notes, which specific units combine to form the final histone octamer?

A
***Which post-translational modification is explicitly described as essential for DNA replication by allowing the DNA to separate from the histones?


Methylation of histones is described in your lecture notes as having what effect on gene transcription?

C
Which part of the histone protein is primarily responsible for its 'basic' nature?


DNA is a double stranded polymer of nucleotides that have STRANDS that run in?
Opposite directions
5’ → 3’
3’ → 5’
Nucleotides are linked by (?) bond.
PHOSPHO DI ESTER bond
Each nucleotide is made up of? (3)
BASE (CGAT)
deOxyribose
Phosphate
Based on the rules of DNA synthesis described in the notes, where does the phosphate group of an incoming nucleotide always attach?


Which of the following nitrogenous bases is classified as a Purine?
Guanine
Purines → Adenine and Guanine
purine = double ringed bases in dna and rna
Give 2 examples of PyriMidines.
Cytosine
Thymine
At which position on the sugar molecule is the nitrogenous base attached?
C1
What structural change occurs to the sugar molecule to allow the phosphate group to attach?
What type of DNA is found in
Mitochondrial DNA
Prokaryotic DNA
Eukaryotic DNA
Mitochondrial DNA → circular
Prokaryotic DNA → circular
Eukaryotic DNA → Linear*
***How does the genetic code differ from the standard code?
UGA is read as →
AGA & AGG is read as →
UGA is read as → Trp or Tryptophan
AGA & AGG is read as → STOP
Codon | Standard Meaning | Mitochondrial Meaning |
UGA | STOP | Trp (Tryptophan) |
AGA / AGG | Arg (Arginine) | STOP |
How does Mitochondrial DNA (mtDNA) differ from Nuclear DNA (eukaryotic cells)?
Circular
more compact
16,569 base pairs vs 3 billion in Nuclear DNA
2 stranded (1 heavy, 1 light)
**Compared to nuclear DNA, which of the following best describes the physical structure of human mitochondrial DNA (mtDNA)?

A
How many base pairs (bp) are contained within a single molecule of human mtDNA?
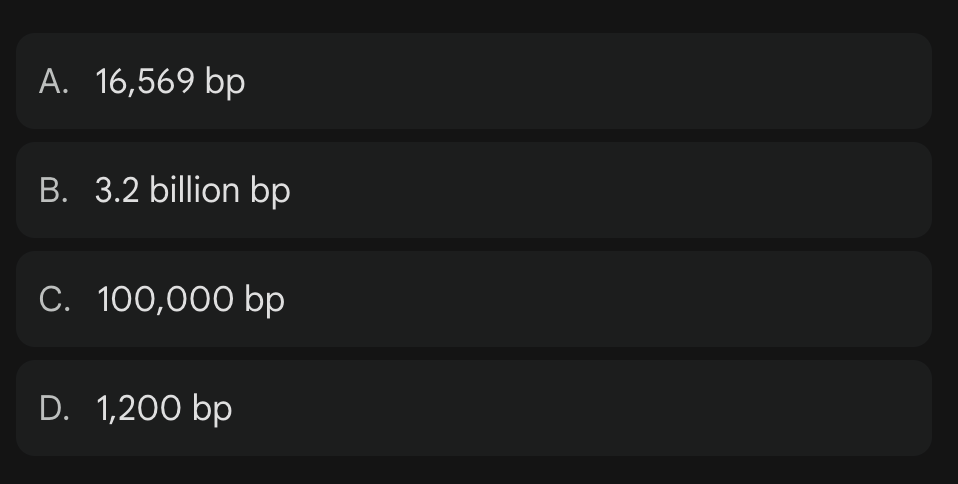
A
mitochondrial DNA bp < Nuclear/Eukaryotic DNA bp
Mitochondrial DNA encodes 13 protein subunits. What is the primary function of these specific proteins?


***In the unique genetic code of the mitochondria, what does the codon UGA represent?
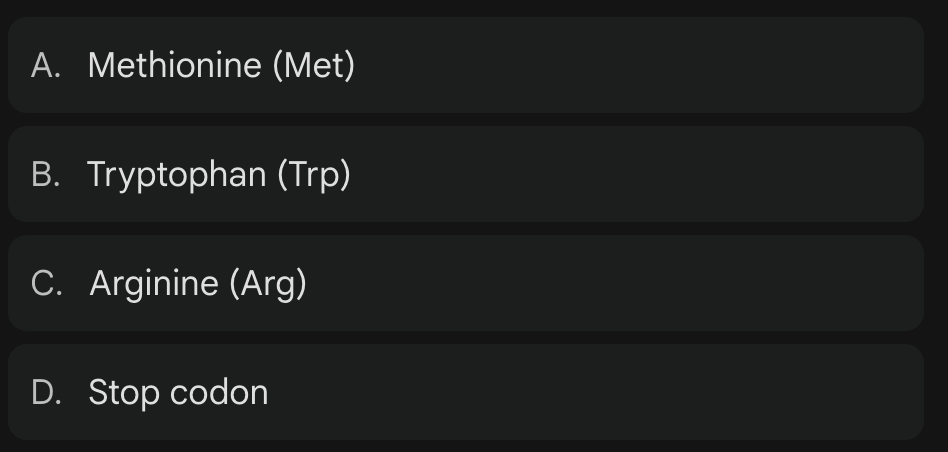
UGA = Tryptophan
Which codons are read as STOP signals in human mitochondria, even though they code for Arginine in the standard code?

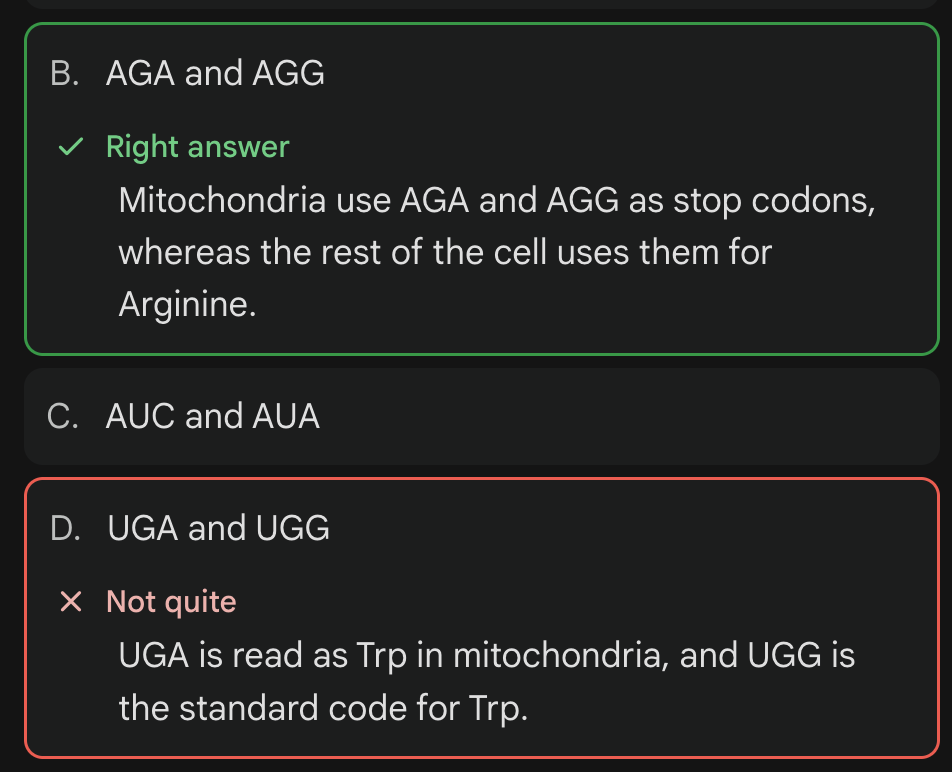
just remember GAGA…
The mutation rate of mtDNA is significantly higher than that of nuclear DNA. What is the estimated magnitude of this difference?





A

C

C
What are the 3 steps of Central Dogma? **
Replication
Transcription
Translation
*4. Reverse Transcription
When does Reverse transcription occur
Viruses
virus uses its RNA sequence to synthesize a complementary DNA
What determines ur genotype?
DNA
What determines Phenotype?
DNA
Environmental Factors
e.g. UV rays
DNA replication occurs from which prime to which prime?
REPLICATION: 5→3
*It is Bidirectional!
Begins at A-T rich regions (easier to melt)
Does DNA replication occur Continously?
No.
Semi-discontinuous
T of F? Eukaryotes only have a single origin of replication while Prokaryokes have multiple origins of replication.
False.
Eukaryotes - multiple origins
Prokaryotes - single origin of replication
T or F? In Eukaryotic DNA replication, BOTH DNA parents strands serve as the template
True
****What is the Initiator protein for Prokaryotes?
dnaA
****What is the Initiator protein for Eukaryotes?
ORC or Origin Recognition Complex
What are the 3 main phases of DNA replication?
Initiation
Unwinding
Primer formation
Elongation
formation of 4 daughter strands
addition of nucleotides
proofreading (removal of mismatched bases)
Termination
What helps add nucleotide sequences to pair with DNA?
DNA polymerase
What Protein is an ATP-driven, processive unwinding of DNA?
Helicase
What helps relieve the ‘tortional strain’ caused by Helicase?
Topoisomerase
What initiates the synthesis of RNA primers?
DNA primase
What prevents the premature reannealing of ssDNA strands to form dsDNA in prokaryotes?
***SSB (single strand binding proteins)
What seals the single strand nick between nascent chain and okazaki fragments on the LAGGING strand?
DNA ligase
Steps of Eukaryotic DNA replication
Locating: Proteins find the "start" buttons on the DNA.
Unpacking: The cell clears away protein spools (nucleosomes) and unzips the double helix (dsDNA) into two single strands (DNA FORK formation). *this process NEEDS ATP
Priming: A short "starter" (RNA primer (ssDNA)) is laid down as a Template.
Copying: DNA polymerase builds the new strands by matching partners to the original DNA.
Gluing: All the separate copied segments are stitched together into a solid chain.
Rewrapping: The new DNA is wound back onto its spools to keep it organized and safe.
Why is an RNA primer needed for DNA polymerase to begin synthesis?
Primer provides the free 3’OH group for DNA polymerase to begin elongation
What is the very last step in the eukaryotic DNA replication process?
Reconstitution of Chromatin Structure
Which of these best describes the state of the DNA template during Step 2?
It is a Single stranded ssDNA
DNA is unwound in step 2 to provide the ssDNA
**The addition of Nucleotides ALWAYS goes in what direction?
5’ to 3’

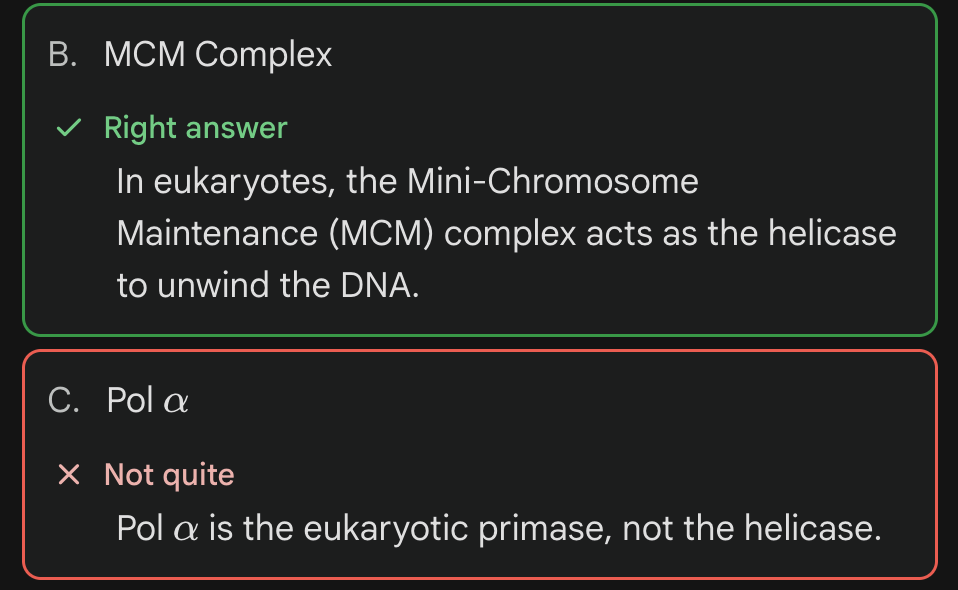
(?) is a Type of DNA helicase that is specifically found in the Eukaryotes.
MCM complex

**LOOKING and IDENTIFYING A-T rich regions

What is the primary function of Single-Strand DNA Binding Proteins (SSBs/RPA)?
Prevent premature reannealing of DNA strands
***Which eukaryotic DNA polymerase is specialized for leading strand synthesis?
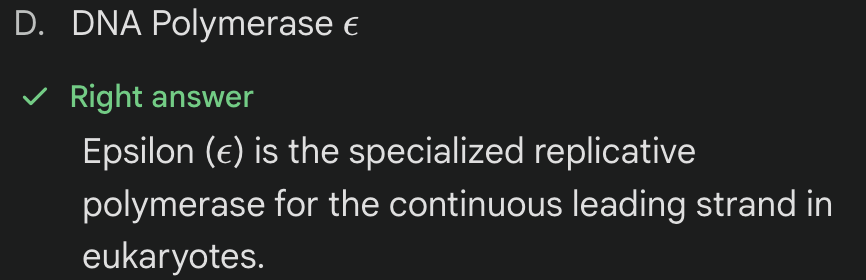
** DNA polymerase ∂ = lagging strand
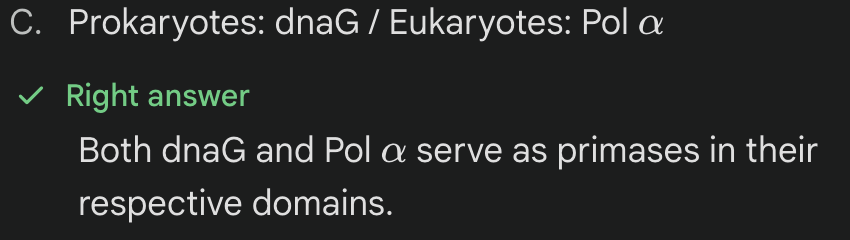
The protein 'dnaG' in prokaryotes is equivalent to which eukaryotic protein?
dnaG & Pol a
function as Primase
Which prokaryotic enzyme acts as the 'Main replicative polymerase' for both strands?
DNA polymerase III (pol III)
BOTH leading and lagging strands
SSBs - (prokaryotes or eukaryotes?)
RPAs - (prokaryotes or eukaryotes?)
SSBs - (prokaryotes)
RPAs - (eukaryotes)
How often does DNA Helicase unwind the DNA according to the table?
Once every 10 nucleotide pairs
In eukaryotes, which enzyme is primarily responsible for lagging strand synthesis?

Which of these is the correct pairing for the function of initiating RNA primer synthesis?



Does Topoisomerase use ATP?
no…
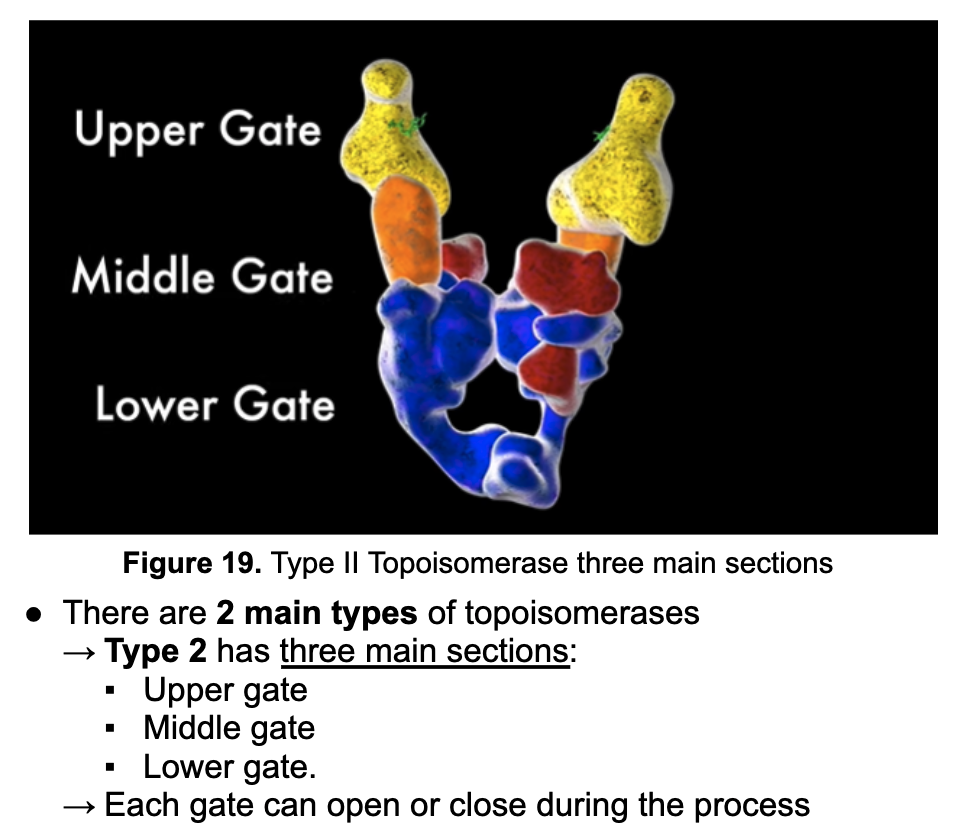
*Which segment of Topoisomerase breaks apart one segmanet of DNA apart?
Middle Segment*
What does Topoisomerase do to help mitigate the tension caused by Helicase?
Cuts: It makes a temporary break in the DNA backbone.
Untwists: It lets the DNA swivel to release the pressure.
Re-seals: It glues the backbone back together.
***What proteins carry out elongation in
Prokaryotes
Eukaryotes
Elongation
Prokaryotes → DNA pol III and I
remember prokaryotes = Numbers
Eukaryotes → DNA pol Delta & Epsilon
remember eukaryotes = greek alphabet
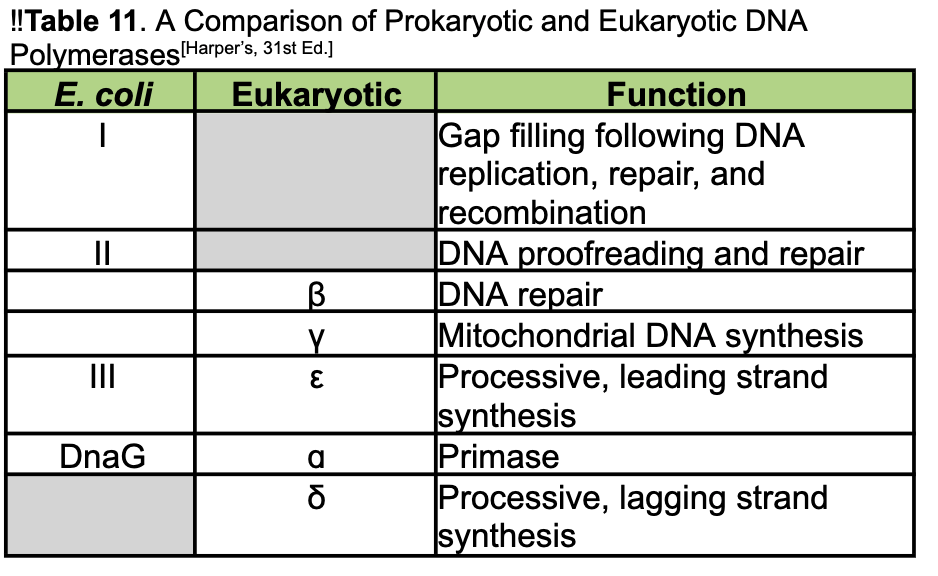
also note:
Beta - DNA repair
Gama - Mitochondrial DNA synthesis
III = epsilon = leading strand synthesis
Delta - lagging strand synthesis
What is the absolute requirement for DNA polymerization?
pre-existing 3’ OH (provided by primase)
Which strand is synthesized continuously in the same direction that the DNA unzips?
Leading
5’ to 3’
What are Okazaki fragments?
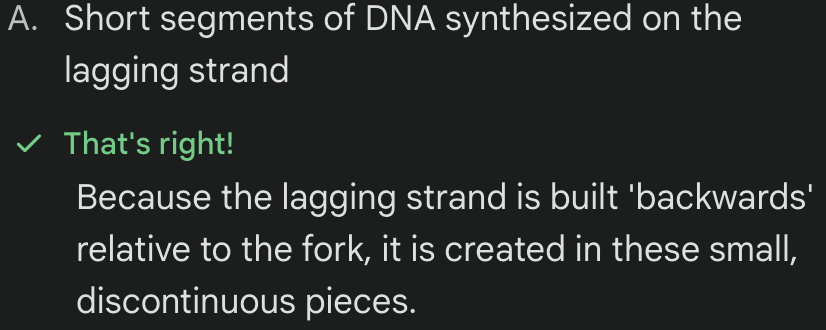
****Why does the lagging strand require more RNA primers THAN the leading strand?


In which direction is a new DNA strand ALWAYS synthesized?
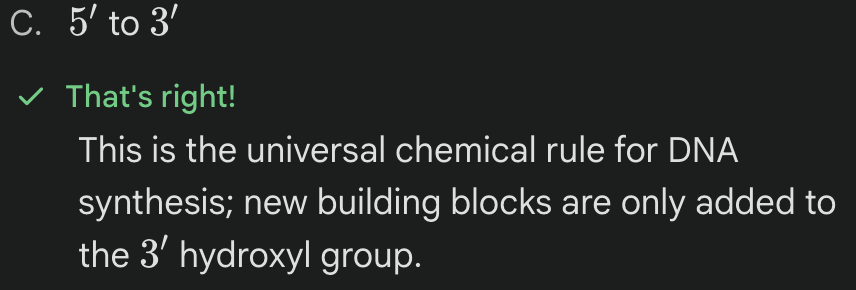
What is the main reason for the 'gaps' mentioned in the lagging strand notes?

How does DNA polymerase know where to start building a new Okazaki fragment?

*What degrades Telomeres?
Exonucleases
What is a specialized reverse transcriptase enzyme that maintains chromosomal stability by extending telomeres?
Telomerase
TeRT = catalytic reverse transcriptase protein subunit
TeRC = RNA compenent that serves as the Internal Template for DNA synthesis
Telomerase recognizes and binds to the (?)
Telomerase recognizes and binds to the 3’ overhang of the telomere.