CMMB 343 Final
1/105
Earn XP
Description and Tags
CMMB 343 Final UofC
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
106 Terms
PDL23: GENETICS PART 2
Prokaryote Translation: Steps (3)
Translation begins when the ribosome binds mRNA at the RBS (Shine-Dalgarno sequence) and initiates translation at a start codon (AUG)
During elongation, tRNAs deliver amino acids based on complementary codon-anticodon interactions
Translation ends at a stop codon (UAA, UAG, UGA)
Prokaryote Translation: mRNA Differences (2)
Prokaryotic mRNA is often polycistronic, meaning it contains multiple coding regions with multiple RBS sites (allowing simultaneous translation of proteins from a single transcript)
Additionally, prokaryotic mRNA does not undergo post-transcriptional processing and therefore lacks: 5’ cap, poly-A tail, and introns
Prokaryote Translation: Mechanistic Differences (5)
Prokaryotes use 70S ribosomes (30S small subunit. and 50S large subunit) while eukaryotes have 80S ribosomes
Translation initiation begins with incorporation of formyl-methionine (fMet) as the first amino acid rather than Met
Codon bias differs among species
Not all ORFs in polycistronic mRNA are in the same reading frame (allows diverse protein production from single transcript)
Certain prokaryotes can reinterpret stop codons to encode uncommon amino acids such as selenocysteine (Sec) and pyrrolysine (Pyl)
Prokaryote: Coupled Transcription/Translation (3)
In prokaryotes, transcription and translation are coupled processes due to the absence of a nucleus; therefore, does not have to be exported before translation begins
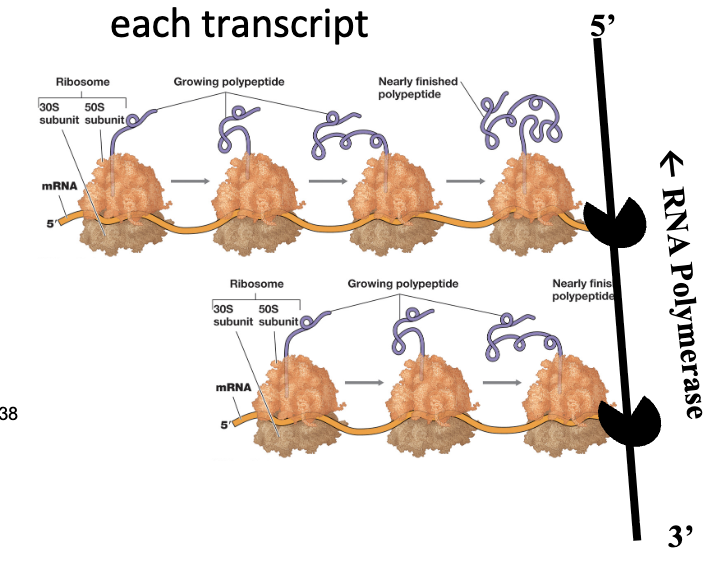
As RNA polymerase synthesizes mRNA in the 5’ to 3’ direction, ribosomes can bind to the emerging 5’ end and initiate translation before transcription is complete
Additionally, multiple ribosomes can attach to a single mRNA molecule, forming polysomes, enabling simultaneous production of many copies of a protein from one transcript
Protein Processing: Post-Translational Modifications (2)
Molecular chaperones play a role in assisting proper protein folding, facilitating the assembly of multi-subunit complexes, and aiding in the incorporation of necessary cofactors
Proteins also contain specific targeting signals that direct them to their correct cellular locations
Translational Regulation: sRNAs (3)
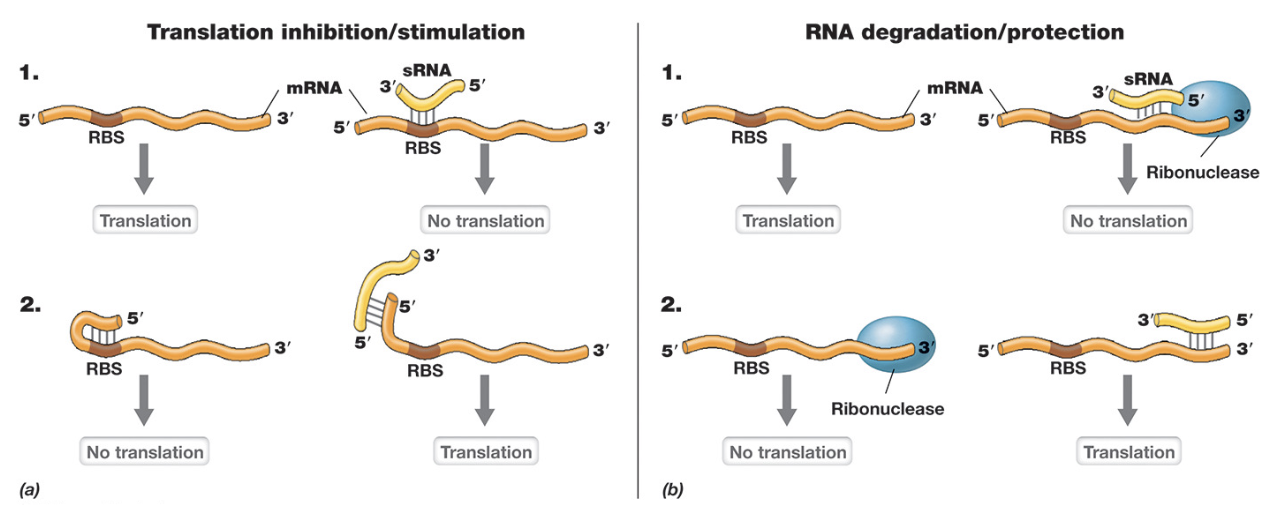
sRNAs are short single-stranded RNA molecules that regulate gene expression by base pairing with target mRNAs (altering translation efficiency)
Binding near the RBS can block ribosomes across and inhibit translation. (3’ end), while in other cases it can expose the RBS to promote translation (5’ end)
sRNAs also influence mRNA stability by either recruiting ribonucleases to promote degradation or protecting the mRNA from degradation (increasing lifespan)
Translational Regulation: Riboswitches (4)
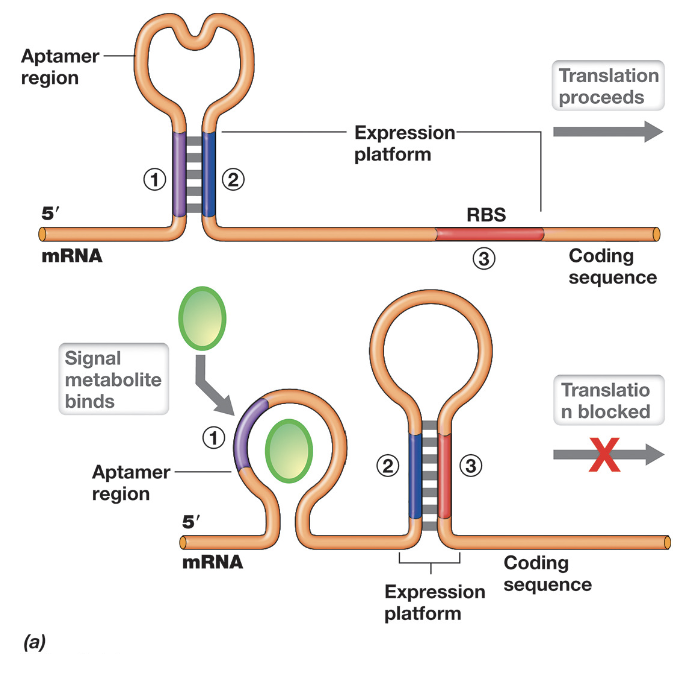
Riboswitches are regulatory elements located within the 5’ UTR of mRNA that control translation through structural changes in response to metabolite binding
They contain an aptamer region that can form alternative secondary structures (stem-loops) depending on whether a specific metabolite is present
In the absence of the metabolite (e.g. low serine), the mRNA folds in a way that exposes the RBS, allowing translation of enzymes involved in metabolite synthesis
When the metabolite is abundant, it binds to the aptamer region, inducing a conformational change that sequesters the RBS within a stem-loop structure, blocking ribosome access and inhibiting translation
Transcriptional Regulation: Attenuation (2)
This is unique to prokaryotes and only possible because of the coupling of transcription & translation
During transcription, the 5’ leader region of mRNA can form one of two alternative step-loop structures, and the structure formed is determined by how quickly the leader peptide is translated
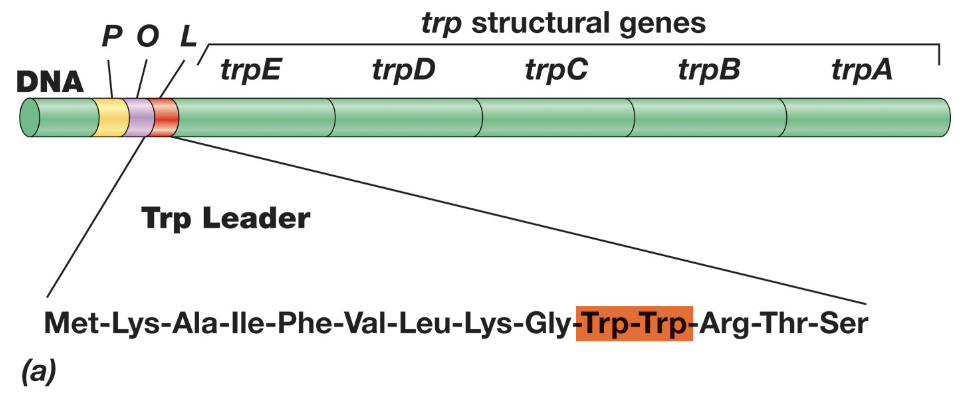
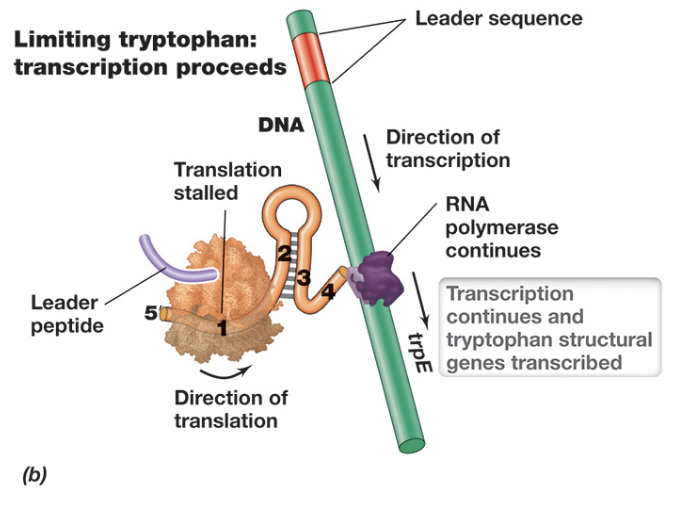
Transcriptional Regulation: Attenuation - When Tryptophan Levels Are Low (3)
When tryptophan levels are low, attenuation allows continued transcription of the Trp operon
Due to insufficient charged tRNATrp, the ribosome stalls at the tryptophan codons within the leader peptide, preventing it from progressing past region 1
This stalling allows regions 2 and 3 of the mRNA to base pair and form an anti-terminator hairpin, continuing transcription and translation
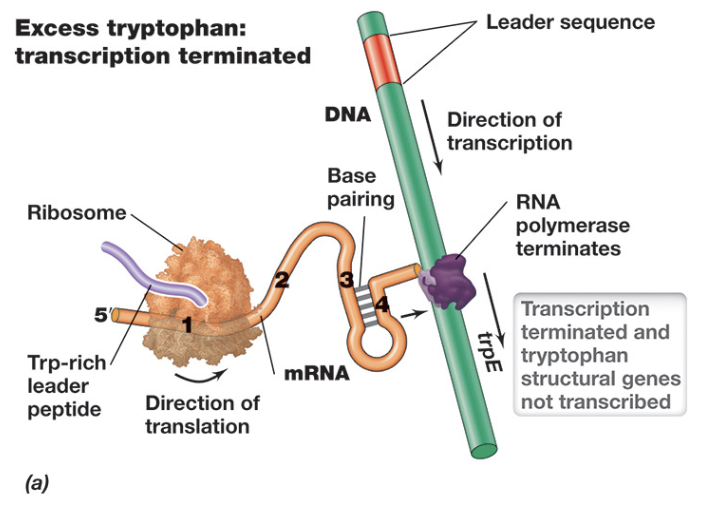
Transcriptional Regulation: Attenuation - When Tryptophan Levels Are High (3)
When tryptophan levels are high, attenuation leads to termination of transcription in the Trp operon
Because charged tRNATrp is readily available, the ribosome rapidly translates the leader peptide and does not stall, covering regions 1 and 2 of the mRNA
This prevents formation of the anti-terminator structure, and instead allows regions 3 and 4 to base pair, forming a terminator hairpin (terminating transcription)
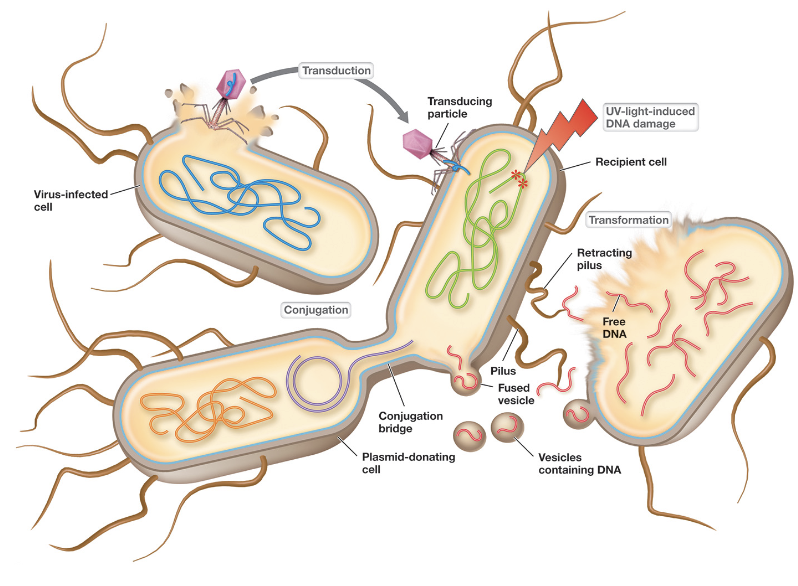
Horizontal Gene Transfer Can Occur Through 3 Mechanisms:
Transformation
Conjugation
Transduction
DNA Transferred By HGT Can Have Three Possible Fates:
Degraded by endo/exonucleases
Exist and replicate autonomously (plasmid)
Recombine with the recipient chromosome
Recombination Can Occur Through Two Mechanisms:
RecA-mediated homologous recombination
Site-specific (non-homologous) recombination
Horizontal Gene Transfer: Transformation (3)
Cells take up “naked” (free) DNA from the environment, often originating from lysed cells that released fragmented dsDNA
Only certain bacteria (termed competent cells) are capable of this process: typically take up ssDNA (examples include Streptococcus, Bacillus, Vibrio)
Competence can be artificially induced in labs by methods that increase membrane permeability such as heat shock, cold shock, or electroporation
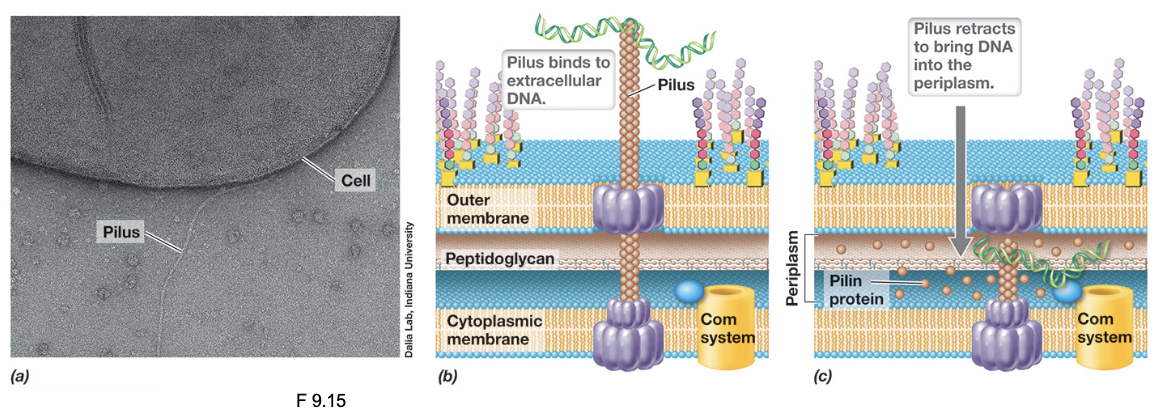
Transformation & Type IV Pili (2)
Extracellular dsDNA binds to the tip of an extended pilus, then retracting to pull DNA toward the cell surface
In Gram-negative bacteria, this process brings DNA into the periplasm; in Gram-positive bacteria DNA is transported toward the cytoplasmic membrane
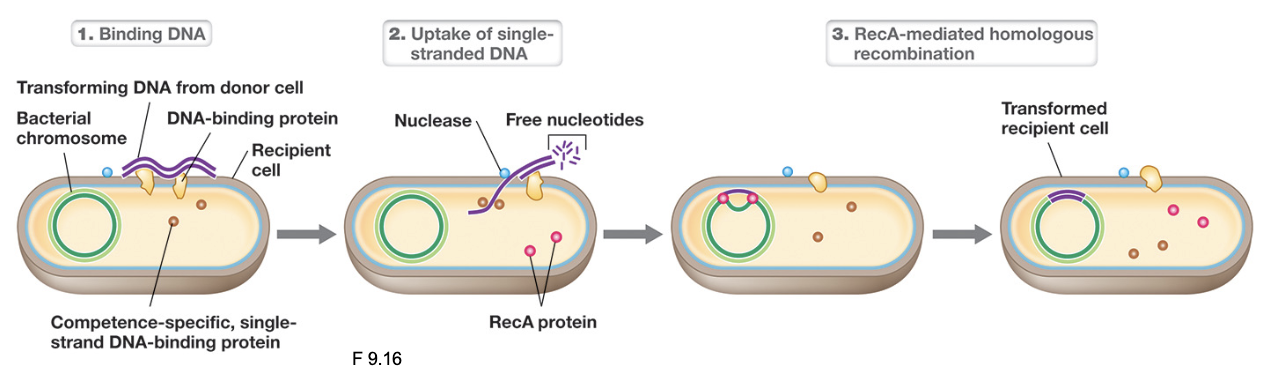
Transformation & The Competence System (3)
Once at the membrane, a competence-associated transport system (such as Rec2/Com proteins) binds extracellular dsDNA
As the DNA is transported across the membrane, one strand is degraded by nucleases while the other strand is imported into the cytosol as ssDNA (strand retained is random)
Once inside, ssDNA may be degraded, but if sufficient sequence homology exists with the host chromosome, RecA mediates homologous recombination
Horizontal Gene Transfer: Conjugation (2)
Conjugation is the main mechanism of HGT involving direct transfer of plasmid DNA from a donor cell (F+) to a recipient cell (F-) through cell-cell contact mediated by a conjugative pilus
Conjugative plasmids carry the genes necessary for this transfer; not all cells are conjugative, but non-conjugative plasmids can move between cells during conjugation
F Plasmid: Components (3)
Contains a tra (transfer) region which encodes the F pilus (conjugative pili) and Type IV secretion system
The plasmid contains an origin of replication (oriV) that allows it to replicate independently within the cell, and an origin of transfer (oriT) where DNA transfer is initiated
The F plasmid also includes insertion sequences (IS) and transposons (Tn) that can facilitate genetic rearrangements
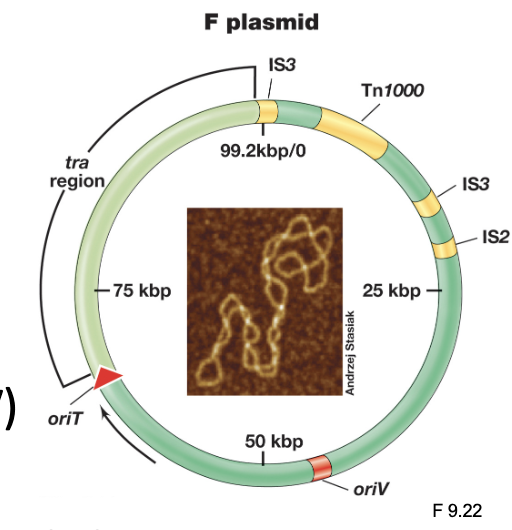
F Plasmid: Only the genes in the ___ region are needed for conjugation; the rest are a bonus
Tra
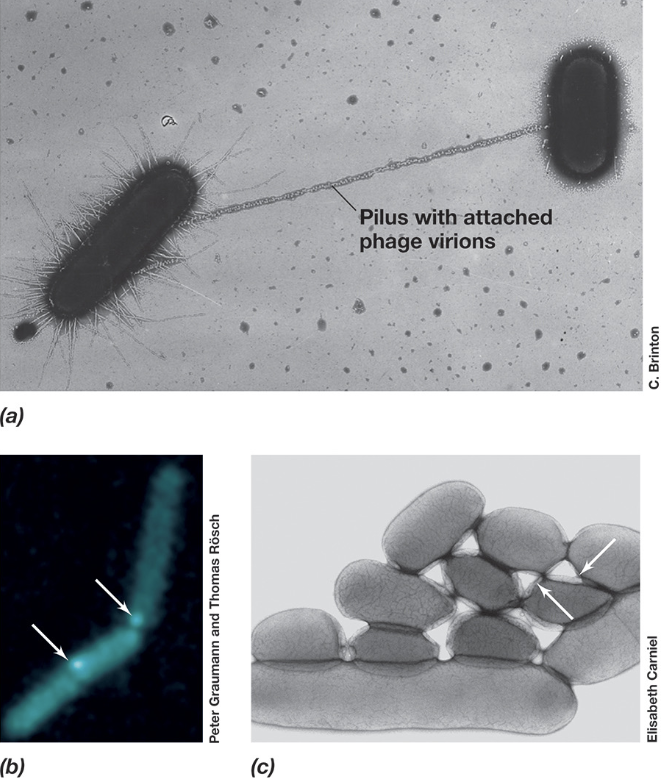
Bacterial Conjugation: Mating Pair Formation (3)
During mating pair formation, the F pilus produced by an F+ donor cell extends outward to locate and bind a F- recipient cell via surface receptors
After contact, the pilus retracts to bring the cells into close proximity, allowing the formation of a stable conjugation junction
DNA transfer then occurs through a Type IV secretion system, creating a channel between the two cells through which plasmid DNA is transferred
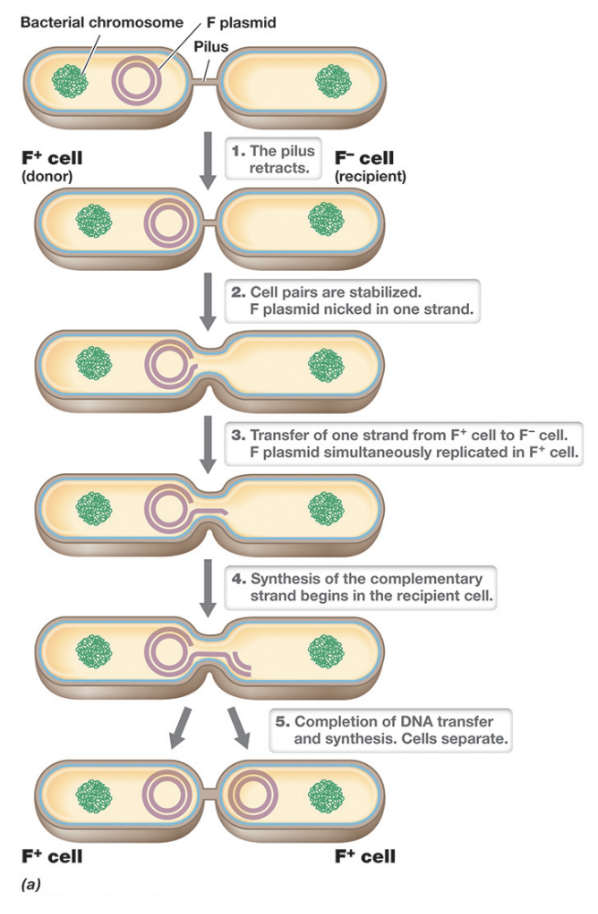
Rolling Circle Replication (4)
Mechanism by which conjugative plasmids are transferred during bacterial conjugation, and is induced by mating pair formation through activation of the tra operon
The process begins when one strand of the plasmid DNA is nicked at the oriT, the nicked strand is then progressively displaced (“rolled off”) as DNA polymerase simultaneously synthesizes a complementary strand in the donor cell to replace it
The displaced ssDNA is transferred into the recipient cell through the conjugation machinery where it circularizes, and a complementary strand is synthesized
The recipient cell is converted into a donor (F+)
Hfr Cells (2)
Hfr (high frequency recombination) cells arise when the F plasmid, which is an episome, integrates into the bacterial chromosome via homologous recombination at shared IS elements
Once integrated, the F plasmid genes remain functional and initiate conjugation
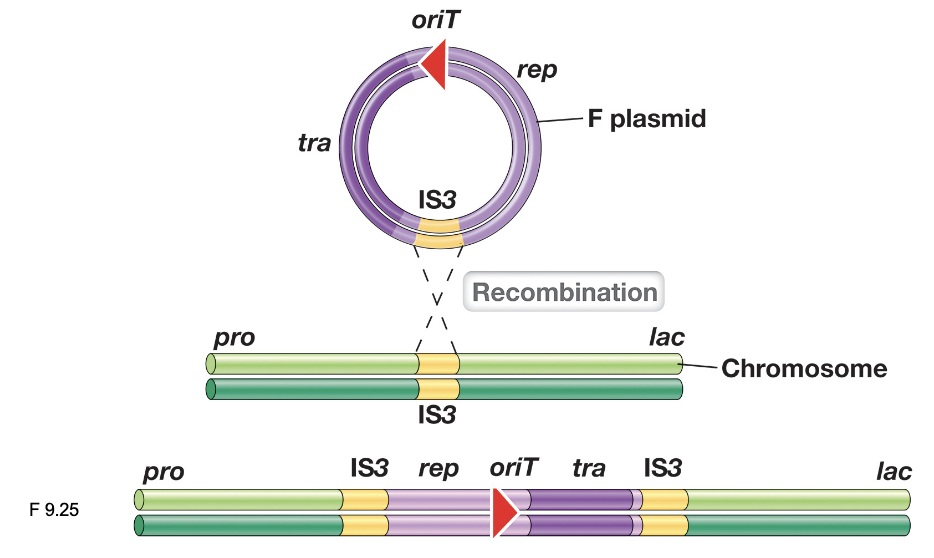
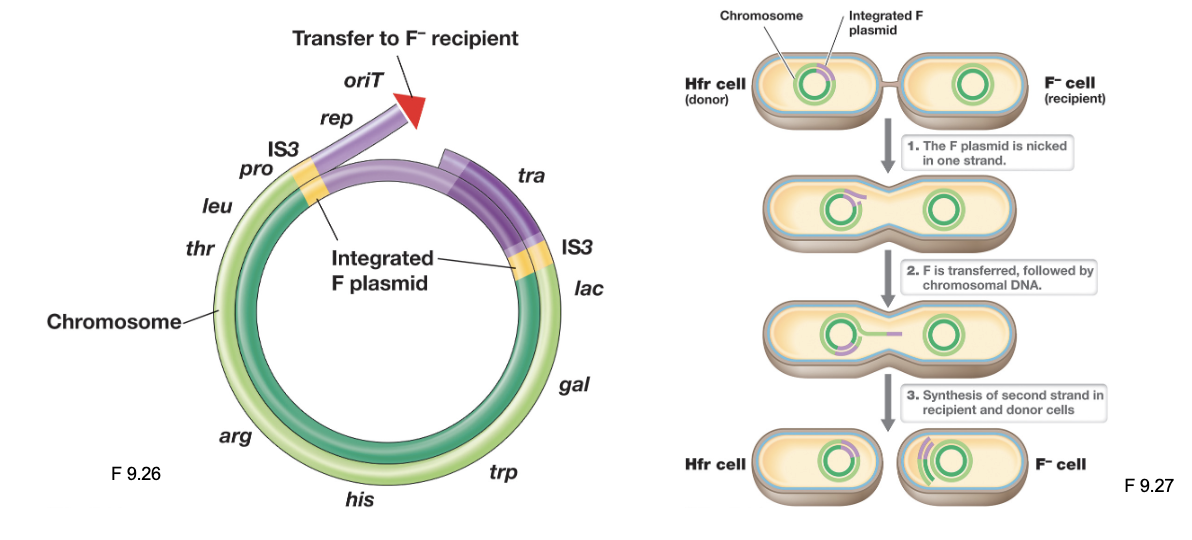
Hfr Cells: Conjugation (3)
Once a mating pair forms, DNA transfer occurs through rolling circle replication, moving chromosomal DNA into the recipient cell
It is unlikely the entire donor chromosome transfers during recombination; as a result, the donor cell remains an Hfr cell, and the recipient remains an F-
The linear stretch of DNA transferred may be degraded, or the donor DNA can recombine with the recipient chromosome “at high frequencies”
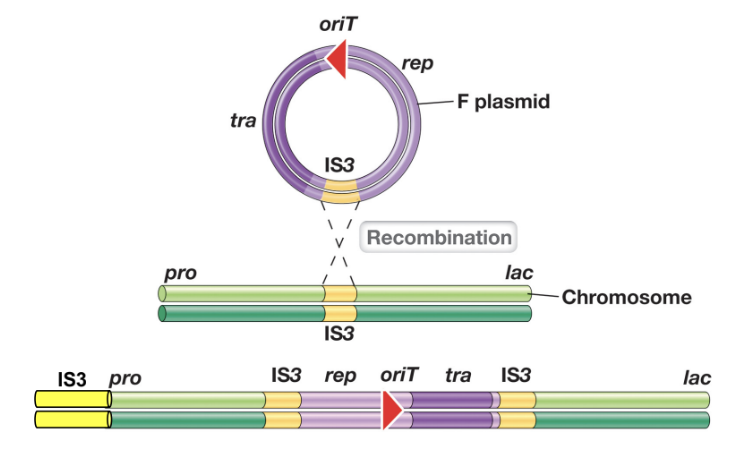
F’ Plasmids (3)
F’ (F prime) plasmids arise when a chromosomally integrated F plasmid is excise incorrectly, carrying adjacent bacterial chromosomal genes
During conjugation, the F’ plasmid is transferred to a F- recipient, allowing the recipient to obtain donor chromosomal genes
The recipient may contain two copies of certain chromosomal genes (partial diploidy)
Transposition (3)
Transposons are transposable genetic elements found in prokaryotic chromosomes, plasmids, and some viral genomes
Transposable elements have the ability to “jump” within the the same DNA molecule, or between DNA molecules
These transposable elements when inserted can potentially disrupt genes
Horizontal Gene Transfer: Transduction (3)
Transduction is the transfer of bacterial chromosome DNA from a donor cell to a recipient cell, mediated by a bacteriophage (phage vector)
Phages can accidentally package fragments of host DNA and deliver them to another bacterium upon infection
Transduction is medically important because it facilitates the spread of virulence factors and antibiotic resistance genes (e.g. GI tract); and in biotechnology (e.g. moving genes from one organism to another)
Not all ___, and not all ___
Phages are transducing
Prokaryotes are transducible
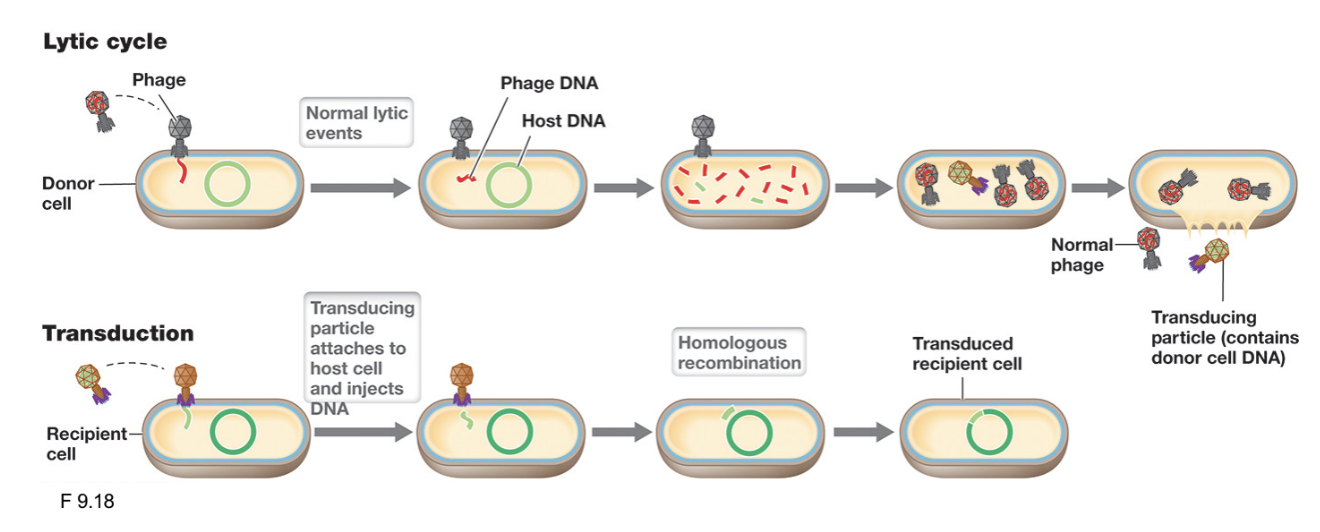
Generalized Transduction (3)
Occurs during the lytic cycle of virulent bacteriophages when host bacterial DNA is accidentally packaged into phage capsids instead of viral DNA
Following release, these defective phage (transducing particles) can still infect new host cells; the transferred DNA may be degraded or undergo homologous recombination
Successful transduction events are rare because progeny phages are virulent, and recombination is not guaranteed
PDL24: MICROBIAL CELL-CELL COMMUNICATION
Protein Secretion (3)
Protein secretion in microbes places proteins either within the cell membrane or exports them outside the cell
This process is mediated by translocase systems, which transport proteins across membranes using energy derived from ATP, GTP, or PMF
Two universal translocation systems are the Sec pathway (general secretion system), and the Tat pathway (twin-arginine translocase)
Protein Secretion: Sec Translocase System (2)
Facilitates the co-translation of membrane-associated proteins into the cytoplasmic membrane via the SRP (signal recognition particle), using GTP as an energy source
It also exports unfolded extracellular proteins post-translationally out of the cytosol through a SecA-mediated process that uses ATP to drive translocation (proteins fold after crossing membrane)
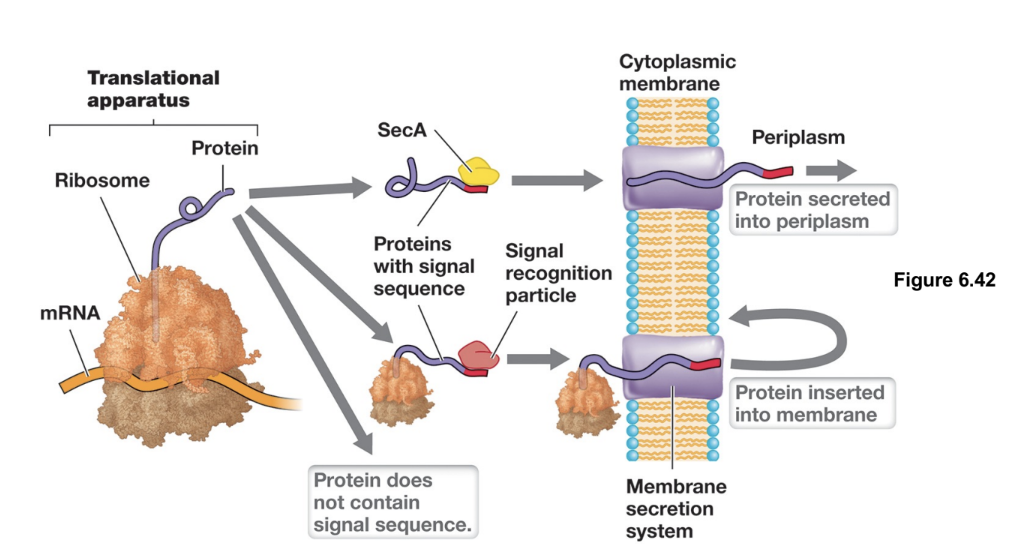
Protein Secretion: Twin-Arginine Translocase
Transports folded extracellular proteins out of the cytosol, recognizing substrates by a twin-arginine (RR) signal sequence in its leader peptide, and uses PMF to power translocation
Gram-Negative Secretion Systems (2)
Two-step translocases (e.g. Type II and V) first transport proteins across the inner membrane into the periplasm, and then across the outer membrane
One-step translocases (e.g. Type I, III, IV, VI) form continuous channels that span both membranes, allowing proteins to be transported from the cytoplasm to the extracellular space or recipient cell in a single step
T2SS: Type II Secretion Systems (4)
In the first step, proteins are transported across the inner membrane into the periplasm via the Sec or Tat systems
In the second step, these proteins are folded and transferred across the outer membrane through a secretion pore
This process is driven by ATP-dependent pseudopilin extension, where pilin-like proteins assemble and push the substrate protein out of the cell
The T2SS is commonly used for secretion of AB toxins
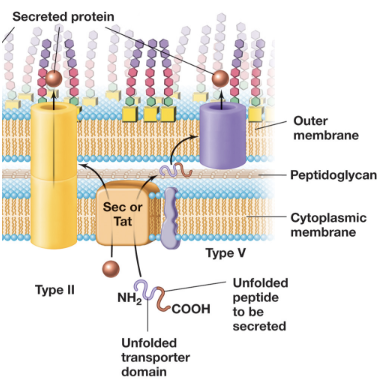
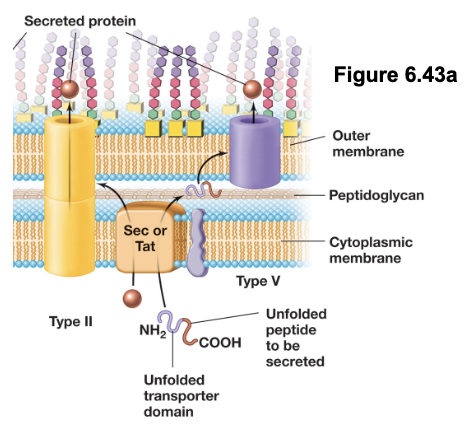
T5SS: Type V Secretion Systems (4)
In the first step, the Sec system transports the autotransporter protein in an unfolded state across the inner membrane into the periplasm
The autotransporter contains two domains: a β-barrel transporter domain and a passenger domain
The transporter domain inserts into the outer membrane and forms a pore, allowing the passenger domain to pass through to the cell surface or extracellular environment (folds into functional form); an autoproteolysis separates the two domains
T5SS is used to secrete virulence factors such as exoenzymes (e.g. IgA protease), adhesins (e.g. pertactin), cytotoxins (e.g. vacuolating cytotoxin) (all contribute to bacterial pathogenicity)
T1SS: Type I Secretion Systems (3)
Consists of an inner membrane ABC transporter, a periplasmic membrane fusion protein, and an outer membrane channel that forms a continuous conduit across both membranes
ATP hydrolysis by the ABC transporter provides the energy required to drive protein export in a single step
T1SS is commonly used to secrete molecules such as bacteriocins, biofilm-associated proteins (e.g. LapA), and RTX toxins

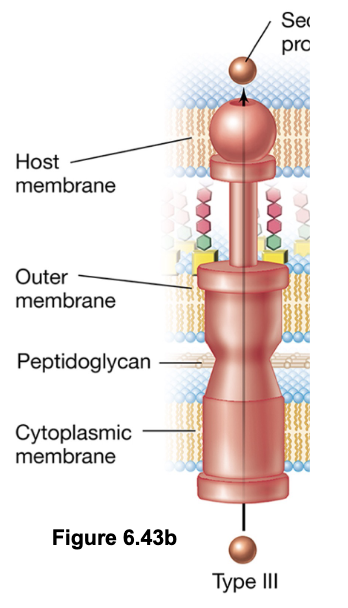
T3SS: Type III Secretion Systems - “Injectisomes” (5)
Deliver proteins directly from the bacterial cytosol into the cytosol of a a target cell (can be either prokaryotic or eukaryotic)
Structurally related to the bacterial flagellar apparatus, this system forms a needle-like complex that spans both membranes
Once contact is established, the tip of the injectisome forms a translocation pore in the host membrane, allowing effector proteins to be injected directly into the host cell
Protein export is powered primarily by the PMF, although ATP is used to prepare and load substrates
T3SS is commonly used by pathogenic bacteria to deliver cytotoxins, and by symbiotic bacteria (e.g. rhizobia) to deliver signaling molecules (e.g. Nod factors) into host cells
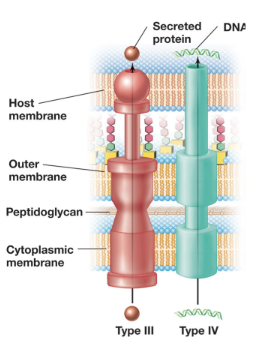
T4SS: Type IV Secretion Systems (4)
Deliver proteins directly from the bacterial cytosol into the cytosol of a a target cell (can be either prokaryotic or eukaryotic)
This is the most common type of secretion system and is powered by ATP hydrolysis
T4SS plays a role in bacterial conjugation, mediating HGT (transferring F and Ti plasmids)
It can also function as a protein secretion system, delivering virulence factors (e.g. pertussis toxin) either into host cells or the extracellular space
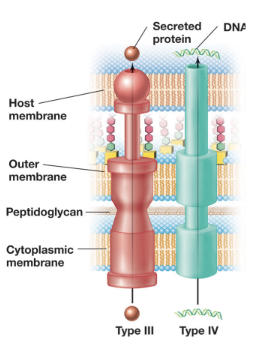
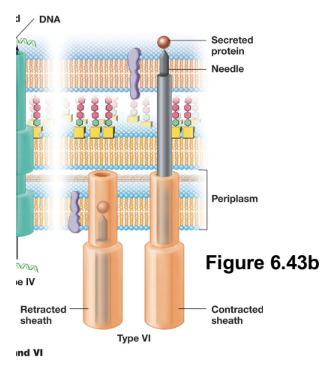
T6SS: Type VI Secretion Systems (4)
Deliver proteins directly from the bacterial cytosol into the cytosol of a a target cell (can be either prokaryotic or eukaryotic)
Structurally, it resembles an inverted bacteriophage tail, and functions as a T4 phage-like injection system powered by ATP
Upon contact with a target cell, a contractile sheath undergoes rapid conformational change, driving a rigid inner spike through the target cell membrane and creating a pore for exoproteins
T6SS is primarily involved in interbacterial competition (“microbial warfare”)
Prokaryotic Cells: Two Mechanisms of Signal Regulated Gene Expression
Signals can act directly as effector molecules
Inducers bind to transcription factors and turn transcription on
Signals can act indirectly by binding to cell surface receptors
Initiate intracellular signalling pathways to alter gene expression

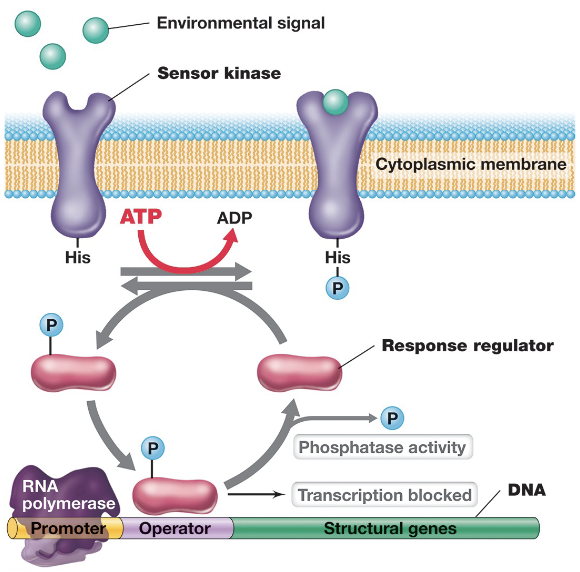
Two-Component Regulatory Systems (4)
This system is used by prokaryotic cells to respond to external signals that cannot directly enter the cell, and consists of two main components: a sensor kinase, and a response regulator
The sensor kinase is a transmembrane histidine kinase that detects an environmental signal, and upon activation, autophosphorylates using ATP on a histidine residue
The phosphoryl group is then transferred to the response regulator which is a transcription factor that becomes active when phosphorylated; alters gene expression accordingly
A phosphatase activity (either intrinsic or separate) can remove the phosphate group from the response regulator, resetting the system and terminating the response
Quorom Sensing (4)
Quorum sensing is a density-dependent mechanism of cellular communication in prokaryotes that enables coordinated population-level responses
Bacterial cells produce extracellular peptide or non-peptide signaling molecules called autoinducers that accumulate as cell density increases
As population grows and a threshold concentration (quorum) is reached, autoinducers bind to regulatory proteins and trigger synchronized gene expression across the entire population (“all-or-nothing” manner)
Regulates biofilm formation, sporulation, competence, bioluminescence, and production of virulence factors
Gram-Negative Autoinducers (2)
Gram-negative bacteria use quorum sensing systems based on species-specific autoinducers called acyl homoserine lactones (AHLs)
AHLs are produced constitutively by AHL synthase enzymes and freely diffuse across the cell membrane into the surrounding environment (small, lipid-soluble properties)
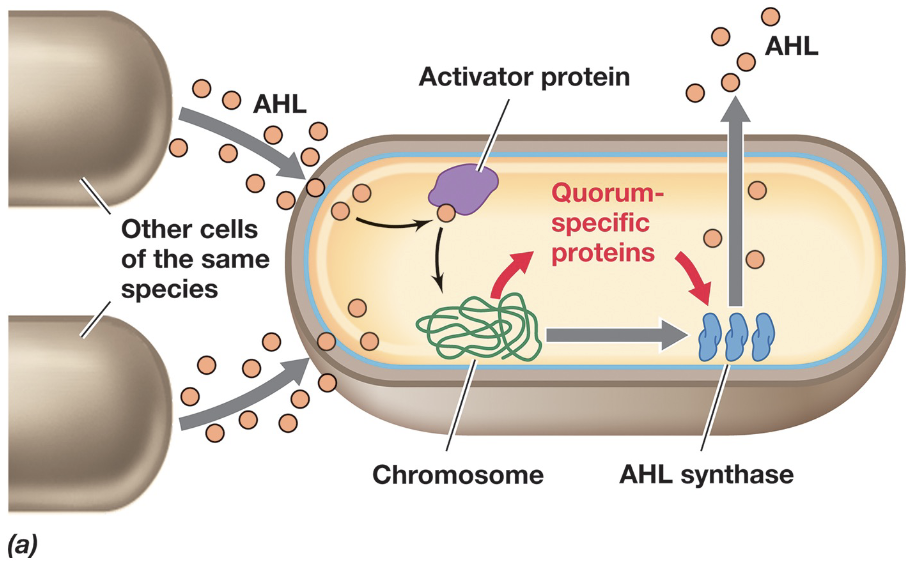
*Quorum Sensing & AHLs: Information Blurb (4)
At low cell density AHLs diffuse away from the cell and remain at low concentrations (preventing activation of gene expression); As cell density increases, AHLs accumulate in environment and diffuse back into cell
A rise in intracellular AHL reaches a threshold, after which AHL binds to an intracellular activator protein, forming an active complex that induces transcription of quorum-sensing genes (e.g. AHL synthase)
This creates a positive feedback loop that rapidly amplifies AHL production and results in coordinated expression of quorum-specific proteins across the population
If cell density decreases, AHL concentrations drop, the activator complex dissociates, and gene expression is reduced
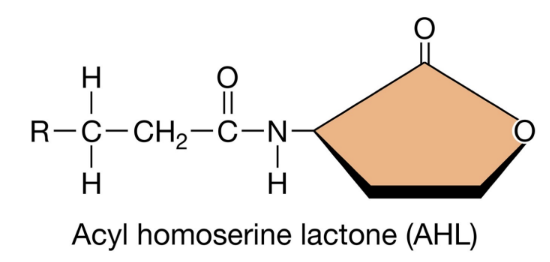
AHLs: Structural Variations (3)
AHLs share a common core structure consisting of a homoserine lactone ring linked to an acyl side chain
The acyl side chain can vary in length from 4-18 carbons, and from modifications to the R group (-OH, =O, -H)
Structural differences determine the specificity of signalling as only bacteria with compatible AHL receptors can recognize and respond to a given AHL molecule
Gram-Positive Autoinducers (2)
Gram-positive bacteria (and some archaea) use quorum sensing systems based on species-specific oligopeptide autoinducers called autoinducing peptides (AIPs)
Unlike AHLs, AIPs are not lipid-soluble, and therefore cannot diffuse freely across the membrane; instead synthesized inside the cell as precursor peptides and actively exported into extracellular environment via ABC transporters
*Quorum Sensing & AIPs: Information Blurb (3)
At low cell density, AIP concentrations remain low and no response is triggered
As cell density increases, AIPs accumulate outside the cell, and upon reaching threshold bind to membrane-bound sensor kinases (2CRS) as part of a two-component regulatory system
This process results in coordinated upregulation (and sometimes downregulation) of quorum-sensing genes across the population
Bacterial Bioluminescence (3)
Example of quorum sensing in which some marine bacteria such as Vibrio, Aliivibrio, and Photobacterium produce light via bioluminescence
Bioluminescent bacteria can colonize the light organs of marine creates such as squid and flashlight fish
In this mutualistic interaction, bacteria provide light that aids in host defense or predation, while the host supplies nutrients and protected environment
Quorum-Sensing Regulated Bioluminescence: Symbiosis Between Aliivibrio fischeri and Euprymna scolopes (Bobtail Squid) (5)
The squid’s light organ is sterile at birth, but is rapidly colonized by A. fischeri from the surrounding environment
Ciliated ducts and a mucous layer help trap bacteria, while antimicrobial compounds and nitric oxide eliminate non-specific species
Chemotactic sugars then attract A. fischeri into the light organ, where its presence stimulates maturation
Once established, bacteria lose their flagella and proliferate to high densities; quorum-sensing activates bioluminescence allowing the squid to avoid predators and assist in hunting at night
Every morning, the squid expels 90% of the bacterial population, resetting the system so that quorum sensing and light production is re-established daily
Quorum-Sensing Regulated Bioluminescence: Biochemistry - Aliivibrio fischeri (3)
Bacterial bioluminescence is catalyzed by the enzyme luciferase, which produces light through an oxidation reaction requiring luciferin and oxygen
In A. fischeriI, luciferin consists of two components: a long-chain aldehyde (RCHO) and reduced flavin mononucleotide (FMNH2)
In the presence of oxygen, luciferase catalyzes the reaction: RCHO + FMNH2 + O2 → RCOOH + FMN + light, where oxidation of aldehyde and FMNH2 releases energy in the form of blue-green light

Quorum-Sensing Regulated Bioluminescence: Physiology - Euprymna scolopes (Bobtail Squid) (3)
In the bobtail squid, the host directs oxygen into the light organ at night to drive the reaction
Emitted light is used for counter-illumination, allowing the squid to mask itself (and shadow) from predators at night
The squid can regulate light intensity using structures such as its ink sac, and an iris-like mechanism
Quorum Sensing in Aliivibrio fischeri (3)
LuxI is an AHL synthase that is constitutively expressed at low (basal) levels and produces the autoinducer 3OC6-HSL, an AI-1 type signalling molecule
Once quorum is reached, 3OC6-HSL binds to LuxR (transcriptional activator), forming the 3OC6-HSL-LuxR complex
This complex binds to promoter regions of the lux operon, activating transcription of genes required for bioluminescence, and LuxI (positive feedback loop)
Quorum Sensing in Aliivibrio fischeri: The Lux Operon (4)
LuxI is the first gene of the operon (*effects covered earlier)
The genes LuxC, LuxD, and LuxE produce a multicomponent fatty acid reductase that synthesizes a long-chain fatty aldehyde (RCHO)
The genes LuxA and LuxB encode the α and β subunits of luciferase enzyme
The gene LuxG encodes an FMN reductase that generates FMNH2
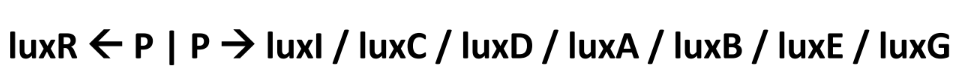
Symbiosis Between Staphylococcus aureus and Homo sapiens (3)
S. aureus is a Gram-positive bacterium that inhabits human skin and nasal passages
It is an opportunistic pathogen capable of causing disease when it breaches innate human defenses
It can cause a broad spectrum of infections, particularly pyogenic (pus-forming) infections

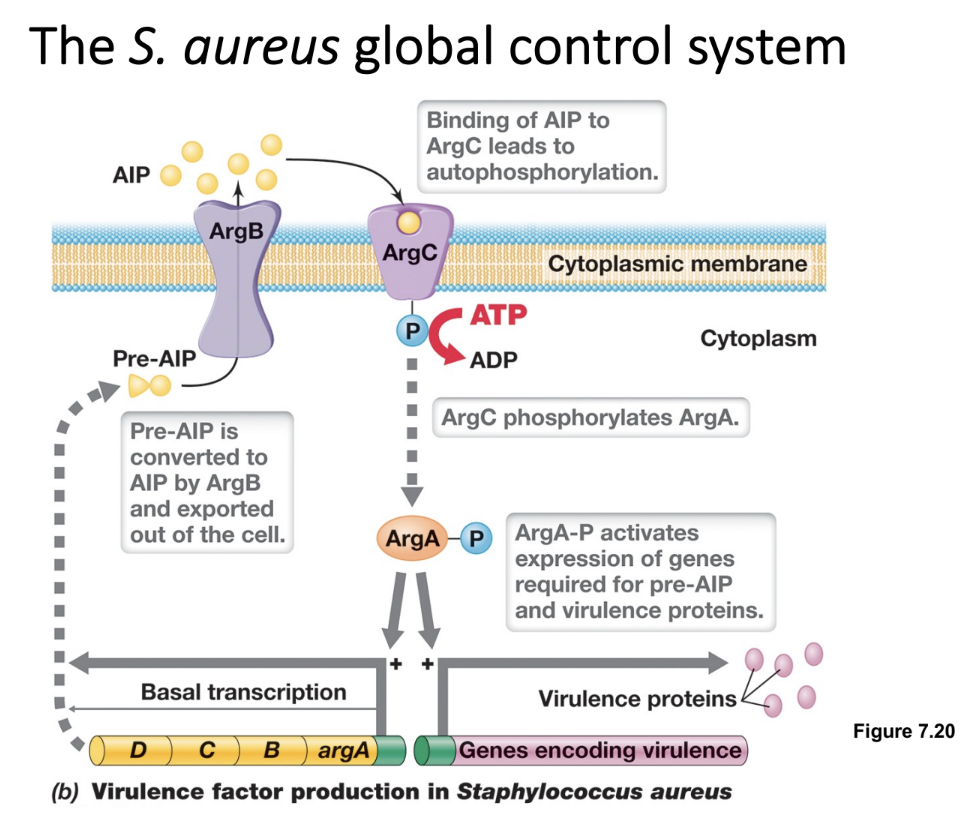
Quorum Sensing in Staphylococcus aureus: The Agr System (3)
AgrD synthesizes the precursor of an AIP, which is exported by the cell membrane transporter AgrB
At quorum, AIP binds to AgrC, a membrane-bound histidine kinase that autophosphorylates and transfers the phosphate to AgrA (a response regulator)
Phosphorylated AgrA-P acts as a transcriptional activator binding to promoter regions of multiple operons (regulons), leading to upregulation of virulence factors
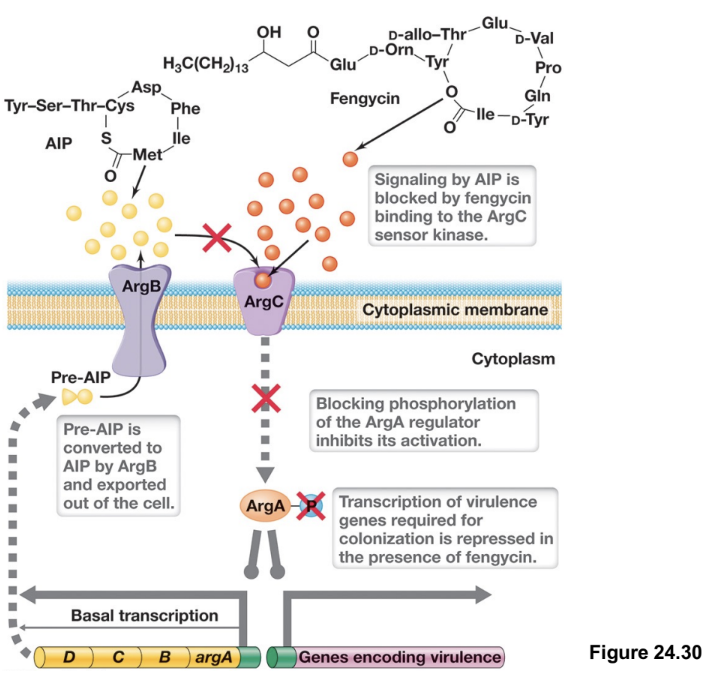
Microbial Warfare By GI Microbiota: Inhibition of S. aureus Global Control System (2)
In the gut, S. aureus can grow and cause disease, but commensal bacteria such as Bacillus species produce antimicrobial compounds such as fengycin that interfere with this process
Fengycin disrupts signalling by blocking AIP from binding to AgrC, preventing activation of the two-component system
PDL25:MICROBIAL INFECTIONS & PATHOGENESIS
The Germ Theory of Disease (3)
The germ theory of disease emerged shortly after the discovery of microbes, replacing the miasma theory of infection
A key figure in establishing this was Robert Koch, who was the first to definitively link specific bacteria to specific diseases (studied anthrax and tuberculosis)
Formulated Koch’s postulates for determining the microbial cause of infection
Koch’s Postulates: Four Steps
The suspected pathogen must be consistently associated with diseased individuals, and not healthy ones
The organism must be isolated from the diseased hist and grown in a pure culture
The cultured organism should reproduce the same disease when introduced into a healthy, susceptible host
The same organism must be re-isolated from the newly diseased host and shown to be identical to the original pathogen
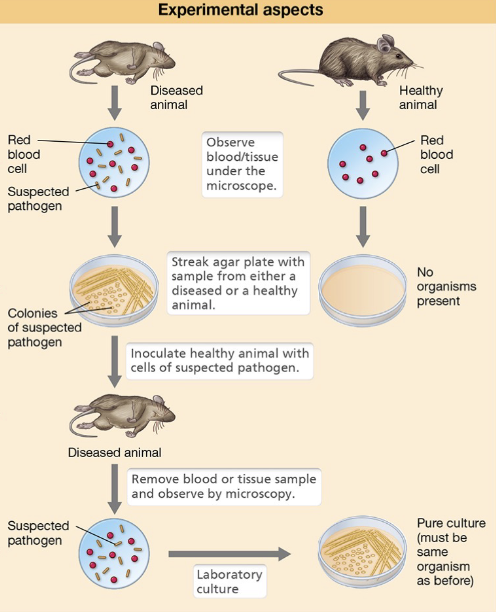
Koch’s Postulates: 3 Limitations
Many pathogens are found in asymptomatic carriers
Additionally, some diseases are caused by toxins rather than the organism itself
Many pathogens are unculturable
Many pathogens lack a suitable animal model
May only infect humans (ethical constraints)
The modern approach to establishing causation uses ___
Genomic-based approaches
Koch’s Postulates: Helicobacter pylori & Gastritis (4)
Australian scientists Barry Marshall and Robin Warren investigated gastritis and ulcers which at the time were believed to result from excess stomach acid, anxiety, and spicy foods, and was treated with bland diets and lifelong antacid use
They isolated Helicobacter pylori from tissue despite the belief that microbes could not survive in acidic stomach environments
However, they lacked an appropriate animal model as attempts to infect other animals did not reproduce the disease
Barry Marshall drank a broth of Helicobacter pylori and proved that the bacteria caused gastritis and ulcers
Helicobacter pylori: Physiology & Infection (3)
The bacteria uses its helical shape and polar flagella to penetrate the thick mucous lining of the stomach epithelium
Epithelial cells secrete carbonate to reduce acidity, and H. pylori enhances its survival by producing urease which generates ammonia to neutralize gastric acid
The bacterium produces toxins (VacA) that damages epithelial cells to access nutrients; eventually causes ulcers (can be resolved with antibiotics)

Define Infection, Disease, Pathogen, Opportunistic Pathogen, Pathogenicity, Pathogenesis, Virulence, and Virulence Factors (8)
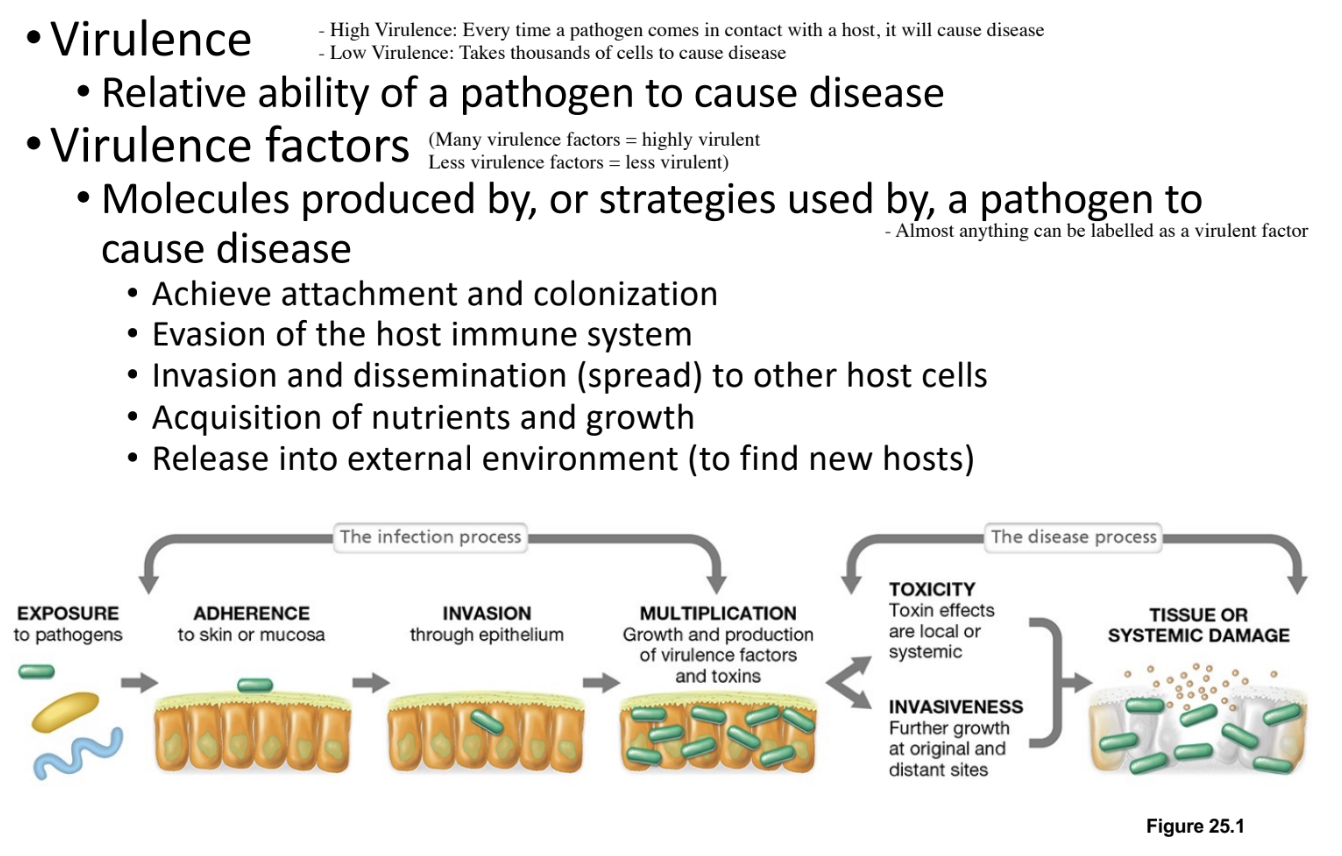
Infection: Portals of Entry - Definition & Examples (2)
Infection requires a portal of entry as all pathogens must gain access to the host through specific routes
Common portals include the skin (breaches such as wounds or bites), gut (need to survive acidic environment - enter using food), and mucous membranes of the nose, mouth, eyes, etc.,
Infection: Portals of Entry - Mucosal Surfaces (3)
Mucosal surfaces are coated with mucus composed of mucin (glycoproteins and polysaccharides) which trap microbes
Mucin also contains immune components such as secretory IgA (sIgA), and defensins
Although most microbes cannot penetrate this barrier, some possess virulence factors such as flagella and mucin-degrading enzymes
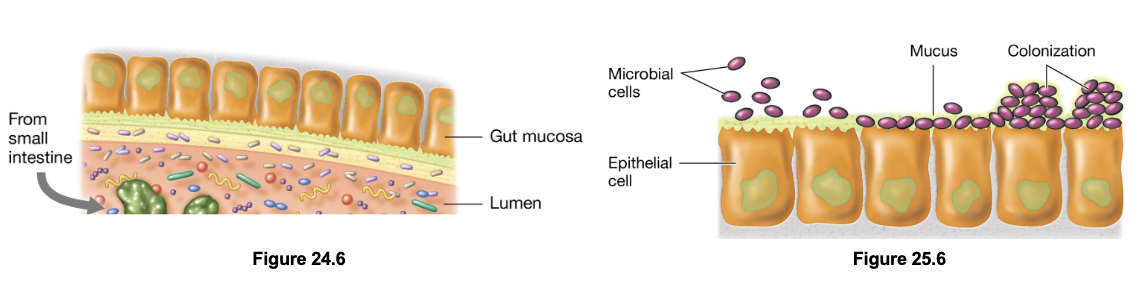
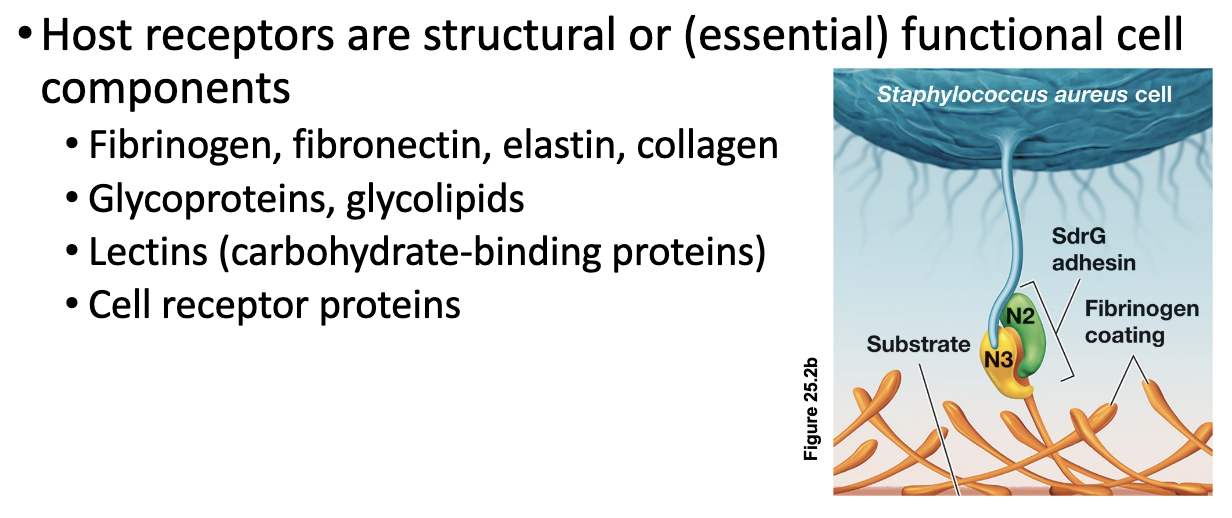
Adherence Molecules (4)
Pathogens must attach to host cells to avoid being removed by physical defenses such as coughing, sneezing, or fluid flow
Microbial adhesins, which are extracellular glycoproteins or lipoproteins, mediate this attachment by binding to specific host cell receptors
Fimbrial adhesins facilitate initial attachment, while afimbrial adhesins promote close, intimate contact with host cell
Once attachment is established, the pathogen is less likely to be flushed out and can proceed with colonization

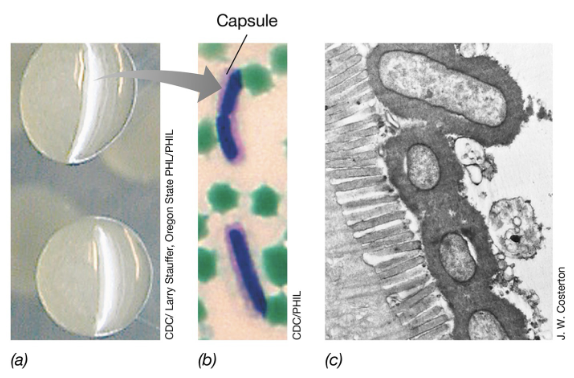
Adherence Structures: Capsules (3)
Capsules are composed of an extracellular polysaccharide or polypeptide coat that surrounds bacterial cells to facilitate attachment
Capsules also provide protection against host immune defenses including phagocytosis mediated by antibodies, and lysis by antimicrobial peptides (e.g. defensins)
Examples of encapsulated pathogens include Bacillus anthracis and Streptococcus pneumoniae
Adherence Secretion Systems (2)
Used by pathogens to inject proteins/toxins directly into host cells; major types include T3SS, T6SS, and sometimes T4SS
Examples of Gram-negative pathogens that use these systems includes E. coli, Shigella, Yersinia, and Salmonella)
Adherence Secretion Systems: E. coli O157:H7 (3)
Pathogenic E. coli O157:H7 uses T3SS to inject Tir (translocated intimin receptor) protein into host cells
Initially, the bacterium uses fimbriae to approach the host cell, after which Tir is delivered across both the bacterial and host membranes into intestinal epithelial cells
Tir becomes embedded in the host cell membrane and functions as a receptor for the E. coli surface adhesin intimin (allowing adherence between bacterium and host cell)
Induced Pathogenesis (3)
Mechanism used by intracellular pathogens to enter host cells after adhesion where the pathogen induces the host cell to trigger its own uptake
This process is mediated by invasin proteins that activate host cell signalling pathways, leading to reorganization of the actin cytoskeleton and the formation of pseudopod-like structures to engulf the bacterium
Once internalized within a phagosome, pathogens employ different strategies to survive, either escaping the phagosome before it fuses with a lysosome, or inhibiting fusion altogether
Induced Pathogenesis: Listeria monocytogenes (2)
Uses invasins called internalins to induce endocytosis
Then releases listeriolysin O to disrupt the phagosomal membrane and escape into the cytoplasm
Induced Pathogenesis: Salmonella (2)
Salmonella uses a T3SS to deliver invasins that induce uptake
A second T3SS inserts proteins into the phagosome membrane, preventing lysosome fusion, and allowing intracellular replication
3 Ways Pathogens Evade The Immune System
Immunoglobulin Proteases
Antigenic Variation
Hide in Plain Sight
Immune System Evasion by Pathogens: Immunoglobulin Proteases (2)
Immunoglobulin proteases degrade Ig proteins to prevent opsonization
IgA proteases are particularly important in mucosal environments where secretory IgA (sIgA) is abundant
Immune System Evasion by Pathogens: Antigenic Variation (2)
Antigenic variation can alter the structure of surface components (e.g. flagella, pili/fimbriae, etc.,)
Antigenic variation can also alter cell envelope components like teichoic acids, LPS, and peptidoglycan to evade defensins
Immune System Evasion by Pathogens: Hide in Plain Sight (2)
Pathogens reduce or stop the expression of recognizable structures like flagella or pili
They incorporate host-derived polysaccharides into their capsules, or coat their surface with host proteins
Nutrient Acquisition by Pathogens: Degradative Enzymes (2)
Degradative enzymes such as lipases, proteases, and glycosidases break down host tissues
Phospholipases disrupt host cell membranes to release intracellular nutrients
Nutrient Acquisition by Pathogens: Eliciting Immune Response
Limited inflammation damages host cells and indirectly releases nutrients
Nutrient Acquisition by Pathogens: Iron Acquisition (3)
Pathogens can secrete siderophores (iron-chelating molecules) that bind iron with high-affinity
They can also use receptors to capture iron from host-binding proteins such as lactoferrin, transferrin, ferritin, and heme; aided by exoenzymes that degrade these complexes
Some pathogens exhibit iron abstinence by avoiding reliance on iron altogether (e.g. Borrelia burgdorferi which uses manganese as an enzyme cofactor - lyme disease)
Genetics of Virulence Factors (3)
Chromosomal virulence factor genes are clustered together in regions known as pathogenicity islands; facilitates their spread by HGT as they are excised together
Many (and possibly most) virulence factor genes are located on plasmids, further facilitating HGT events
Some virulence genes are derived from non-bacterial sources and are found within prophage DNA (viral origin)
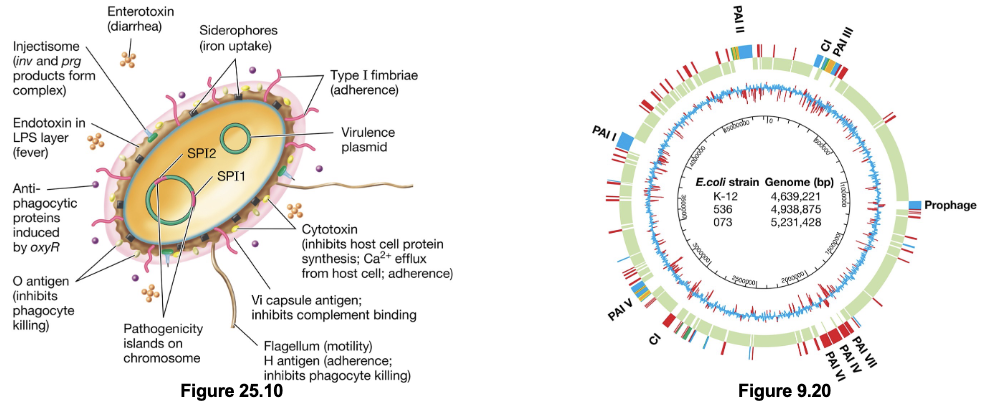
Virulence Enzymes & Toxins (2)
Virulence enzymes and toxins are key mechanisms by which pathogens cause disease in order to acquire nutrients for growth, replication, and spread
The tissue damage caused by pathogens reflects their toxicity
Virulence Enzymes & Toxins: Three Mechanisms of Toxicity
Enzymes that break down components of host cells/tissues
This includes exoenzymes that are secreted into the surroundings or delivered directly into host cells
Protein toxins that affect host cell/tissue function
Exotoxins are secreted into the surroundings or directly injected into host cells
Enterotoxins specifically act in the GI tract to impair normal function
Endotoxins; structural components of the bacterial cell wall rather than secreted products
Includes the lipid A portion of LPS in Gram-negative bacteria
Includes lipoteichoic acids in Gram-positive bacteria
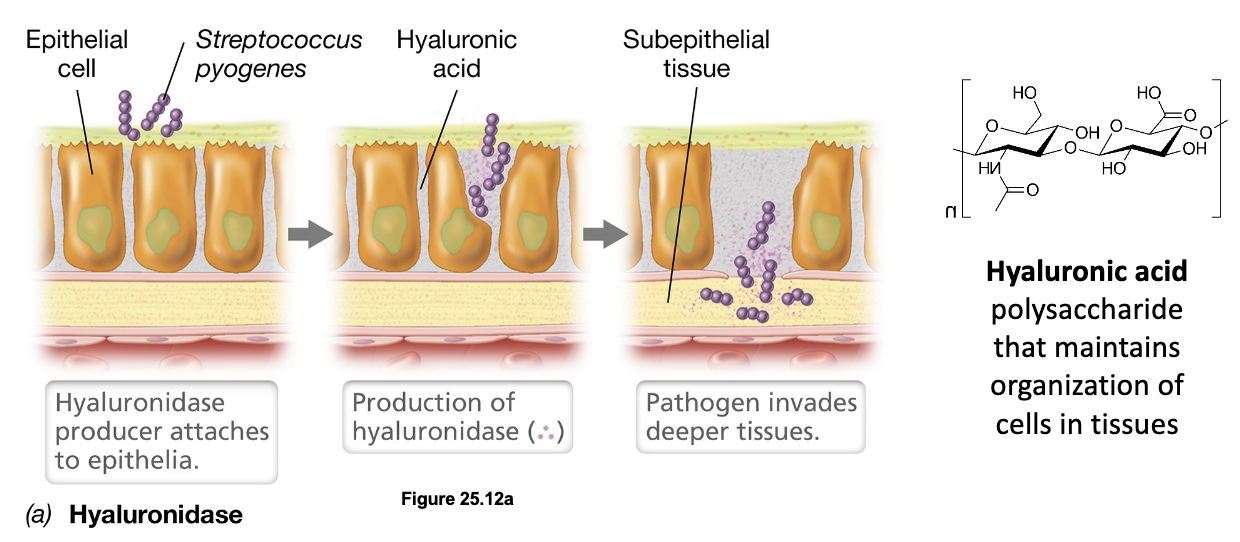
Tissue-Destroying Enzymes: Hyaluronidase (3)
Tissue-destroying enzymes are exoenzymes produced by invasive pathogens that degrade components of the extracellular matrix, allowing microbes to spread beneath the epithelium
For example, Streptococcus pyogenes produces hyaluronidase which breaks down hyaluronic acid (a polysaccharide maintaining structural organization of cells in tissues), opening pathways for deeper invasion
Pathogens produce a variety of other degradative enzymes including collagenases, elastases, proteases, lipases, and nucleases
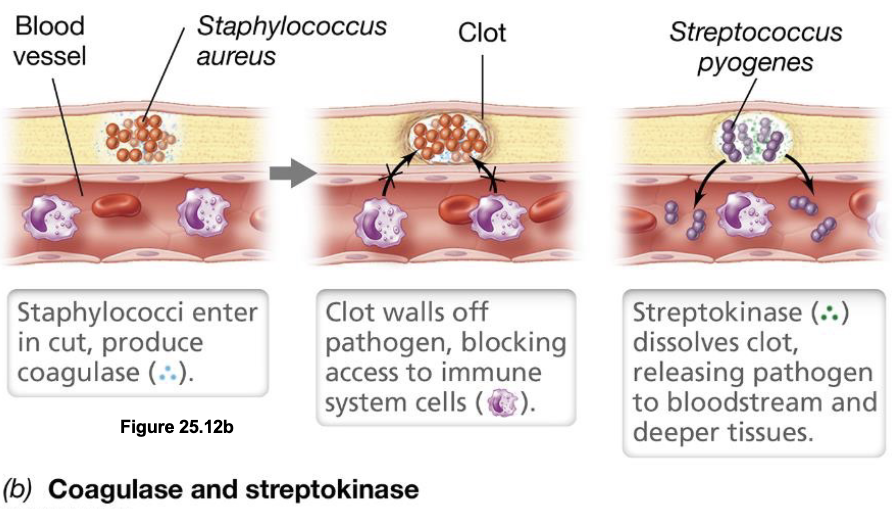
Clot-Forming & Dissolving Enzymes: Coagulase & Streptokinase (2)
Coagulase produced by bacteria such as Staphylococcus aureus induces clot formation around the infection site, walling off the pathogen to protect it from immune cells as it reproduces
Pathogens later switch strategies by producing clot-dissolving enzymes such as streptokinase (or staphylokinase), which break down the clot to release the bacteria into surrounding tissues (or bloodstream)
Pyogenic Infections (4)
Pyogenic infections are pus-forming infections caused by opportunistic pathogens such as Staph. aureus, Staph. epidermidis, and Strep. pyogenes that enter the body through skin cuts
Once inside, these bacteria use virulence factors like coagulase to induce fibrin clot formation, and produce toxins such as hemolysins and leukocidins to destroy RBCs and WBCs
This leads to the accumulation of dead cells, bacteria, and cellular debris which forms the pus
As infection progresses, pressure builds within the localized abscess until it may rupture, facilitating further spread of the pathogen
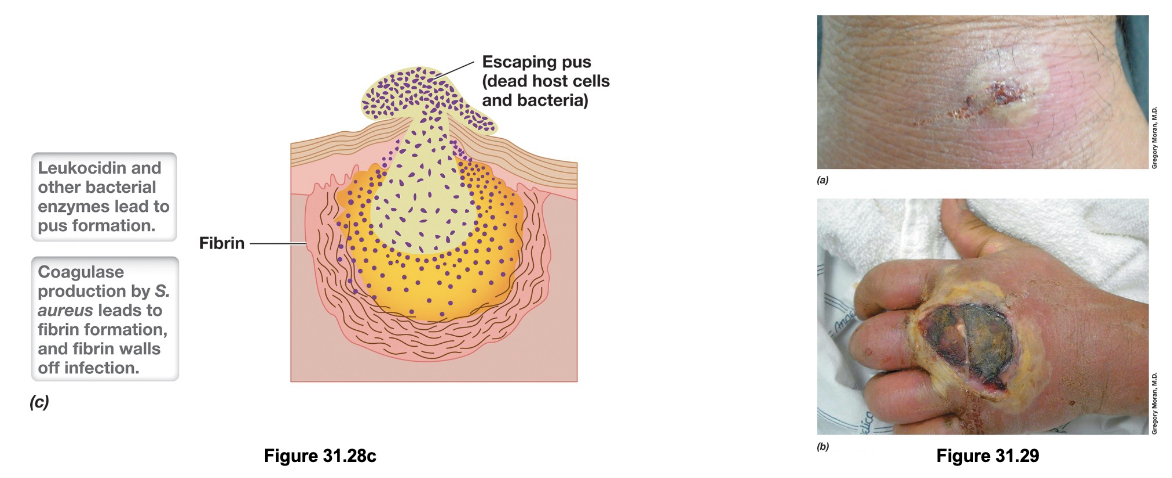
Staphylococcus aureus: α-Toxin (3)
Cell destruction in Staph. aureus skin infections is mediated by α-toxin (α-hemolysin), a pore-forming cytotoxin that disrupts membrane integrity
This leads to uncontrolled efflux of cytoplasmic contents and influx of extracellular molecules, leading to cell lysis; accumulation of dead cells in the area forms pus
These infections are treated with antibiotics and often require drainage to remove pus
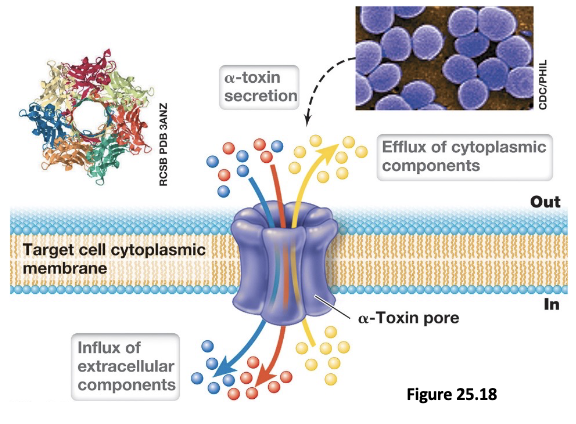
Pathogenic Invasion of Deep Tissue: Local & Distal Effects (2)
Pathogens use multiple adhesins to bind ECM proteins and enzymes such as hyaluronidase to degrade structural components and penetrate into subcutaneous tissues; leads to localized infections such as impetigo, scalded skin syndrome, and strep throat
As bacteria invade further and potentially enter the bloodstream, they can disseminate, resulting in septicemia; associated with severe disease outcomes such as scarlet fever, rheumatic fever, necrotizing fasciitis, and toxic shock syndrome (sepsis)
AB-Type Exotoxins (3)
AB-type exotoxins are two-component protein toxins composed of a binding (B) subunit and an active (A) subunit that work together to disrupt host cell function
The B-subunit binds to receptors on the surface to facilitate entry/translocation of the A-subunit into the cell
Once inside, the A-subunit exerts toxic effects through enzymatic activity, often targeting essential cellular processes such as transcription, translation, or intracellular signalling
Diphtheria (4)
Diphtheria is a respiratory disease caused by Corynebacterium diphtheriae that colonizes the throat and upper respiratory tract; typically transmitted through aerosolized droplets or direct contact with infected individuals
The disease is primarily driven by an AB-type exotoxin that damages host cells, leading to accumulation of fibrin, dead cells, and immune components; forms a pseudomembrane in the throat (along with swelling of the neck)
This pseudomembrane can obstruct the airway and cause suffocation if untreated, while systemic spread can result in damage to the heart and liver
Diphtheria has been eliminated in developed countries due to widespread vaccination with the DTaP vaccine, and is treated with antibiotics
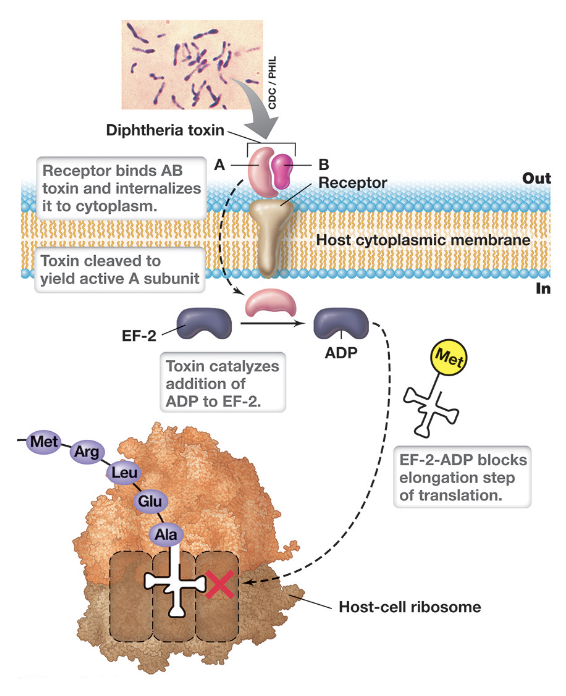
Diphtheria Toxin (DT) (3)
DT is an AB-type exotoxin encoded by a lysogenic bacteriophage (β phage) within Corynebacterium diphtheriae (best studied)
Once inside, the A-subunit exerts its toxic effect by ADP-ribosylating elongation factor-2 (EF-2)
This modification inactivates EF-2 and blocks the elongation step of translation (halting protein synthesis), leading to cell death
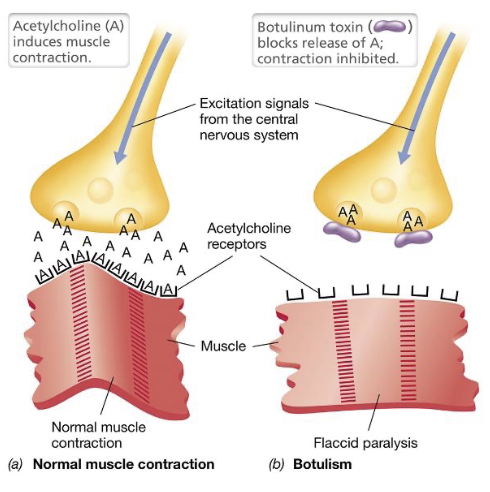
Botulism (4)
Botulism is a neuromuscular disease caused by Clostridium botulinum in which either ingestion of the preformed toxin, or bacterial colonization of the GI tract leads to toxin exposure
The toxin is absorbed through the stomach or small intestine into the bloodstream, and specifically targets motor neurons at neuromuscular junctions
At these junctions it blocks neurotransmitter release, preventing muscle contraction, and resulting in flaccid paralysis; paralysis becomes life-threatening if respiratory muscles are affected
Treatment includes antitoxin to neutralize circulating toxin and antibiotics, but once the toxin has bound to nerve terminals its effects are irreversible; if the body does not replace the neuron, damage is permanent; (severe cases may require mechanical ventilation)
Botulism Toxin (BoNT) (2)
Botulinum toxin (BoNT) is an AB-type neurotoxin produced by Clostridium botulinum, with its genes found on the chromosome, plasmids, or prophage elements (most potent toxin known)
The B-subunit binds specifically to receptors on motor neurons; once inside, A-subunit acts as an endoprotease that cleaves SNARE proteins required for synaptic vesicle fusion
As a result, synaptic vesicles cannot merge with the presynaptic membrane, preventing the release of acetylcholine into the synaptic cleft; without acetylcholine muscle fibers cannot be stimulated, leading to flaccid paralysis
Botulism Can Occur Through Three Routes:
Food Botulism
We eat C. botulinum all the time; it normally cannot compete with normal gut bacteria, unless bacteria produce toxin in food before we eat (e.g. unpasteurized milk, honey, juice, etc.,)
Infant Botulism
Babies lack a well-established gut microbiome, allowing C. botulinum spores to colonize the intestine
Wound Botulism
Spores enter a wound, wounds “heal”, organisms continue to grow in anaerobic conditions and produce toxin
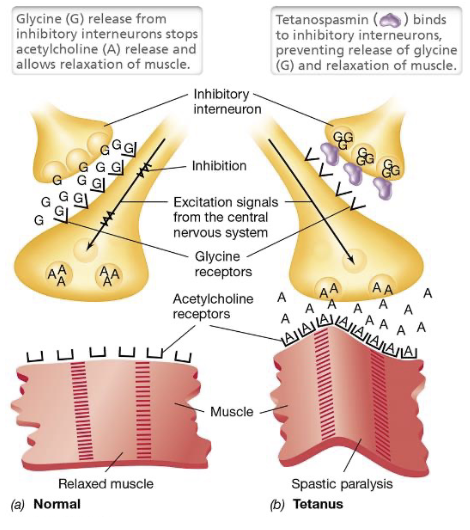
Tetanus (4)
Tetanus is a neuromuscular disease caused by Clostridium tetani that occurs when the bacterium colonizes a deep, anaerobic wound and produces tetanus toxin (tetanospasmin)
The toxin enters the bloodstream and targets inhibitory interneurons that normally regulate motor neuron activity
By blocking the release of inhibitory neurotransmitters, the toxin removes the signal that tells muscles to relax, resulting in sustained contractions - spastic paralysis
Tetanus is rare in developed countries due to DTaP vaccine; treatment includes antibiotics and antitoxin to neutralize unbound toxin (require neuronal regeneration or damage is permanent once bound)
Tetanus Toxin (TeNT) (4)
TeNT is an AB-type neurotoxin produced by Clostridium tetani that shares structural homology with botulinum toxin, but targets inhibitory neurons rather than motor neurons; genes located on plasmid
The B-subunit binds to receptors on inhibitory interneurons; once inside, A-subunit acts as an endoprotease that cleaves SNARE proteins (such as synaptobrevin)
This prevents synaptic vesicle fusion, and blocks the release of inhibitory neurotransmitters like glycine
Without inhibition, motor neurons become overactive and continuously release acetylcholine (causing spastic paralysis)
Pertussis (4)
Pertussis (whooping cough) is a respiratory disease caused by Bordetella pertussis, which colonizes the ciliated epithelial cells of the lower respiratory tract
The bacteria produces virulence factors including pertussis toxin (PT), an AB-type exotoxin secreted via T4SS, and tracheal cytotoxin (TCT), which damages and kills ciliated cells
Loss of these cells impairs mucociliary clearance, leading to accumulation of mucus and prolonged forceful coughing episodes, with the characteristic “whoop” occurring during inhalation after coughing fits
Protection against pertussis is provided through the DTaP vaccine
DTaP vaccine stands for ___
Diphtheria, Tetanus, and acellular Pertussis
Cholera (4)
Cholera is an intestinal disease caused by Vibrio cholerae that colonizes the small intestine following ingestion food or water contaminated with human feces (infects only humans)
The bacteria produce cholera toxin, an AB-type exotoxin that targets intestinal epithelial cells and disrupts ion transport, leading to massive secretion of water and electrolytes into the intestinal lumen
This results in profuse, watery “rice-water” diarrhea that can cause rapid and severe dehydration; rapid fluid loss can lead to hypovolemic shock and death if untreated
Management primarily involves aggressive rehydration therapy to replace lost fluids and electrolytes, along with antibiotics to reduce disease severity