miRNA and Cancer
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
Biogenesis and function of canonical miRNAs of animals: Step 1
Transcription of a primary miRNA transcript (pri-miRNA) by RNA pol II
transcript typically has a 5’cap
3’polyA tail may or may not be present, depending on timing of the next processing step
depends on what machinery reaches the 3’end first (microprocessor Drosha, or polyadenylation)
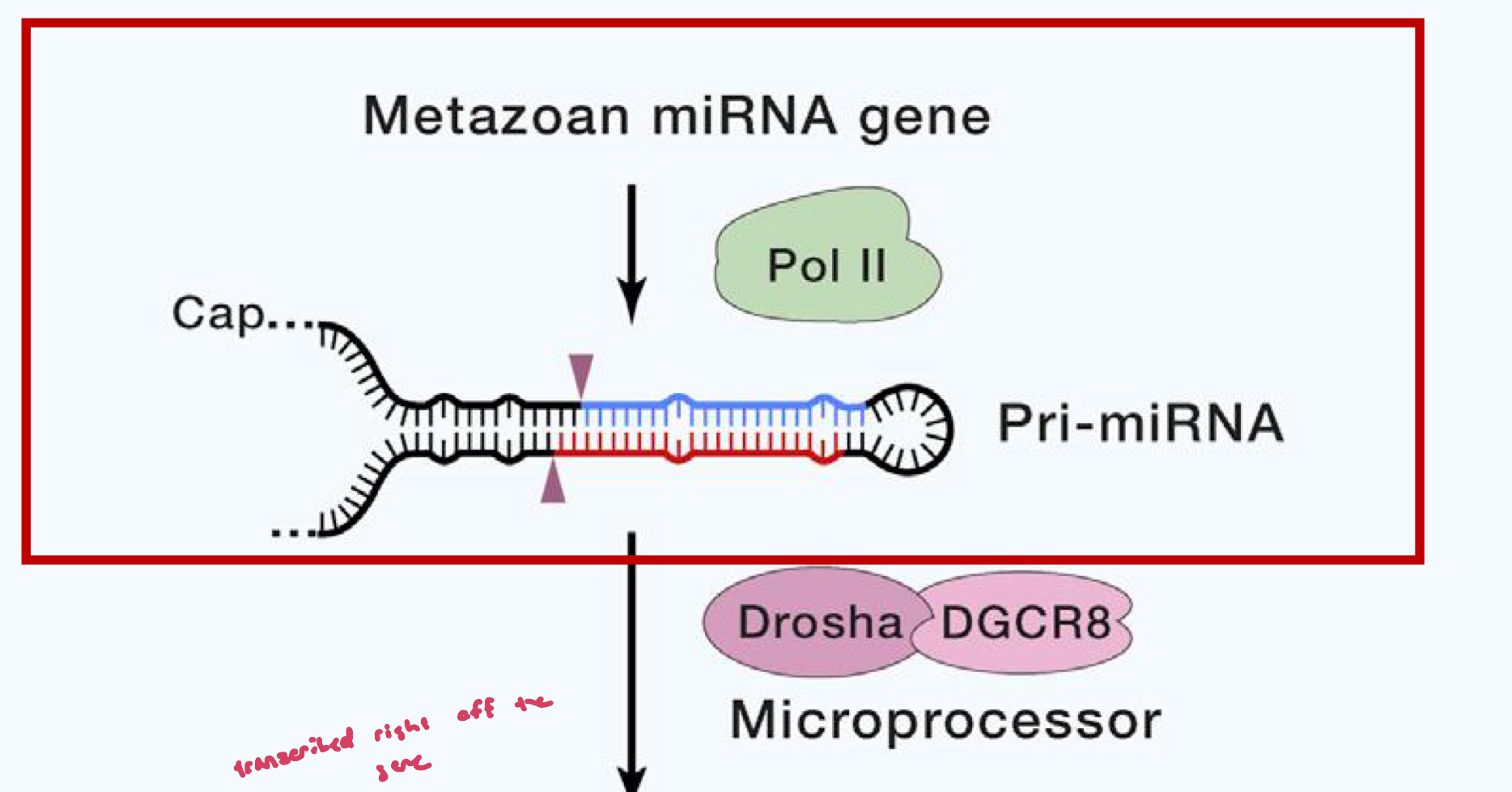
Biogenesis and function of canonical miRNAs of animals: Step 2
Cleavage by the microprocessor complex (Drosha +DGCR8) to form the Pre-miRNA hairpin
Drosha is an RNAse III enzyme, these act on dsRNA
DGCR8 is an essential cofactor, it recognizes the pri-miRNA
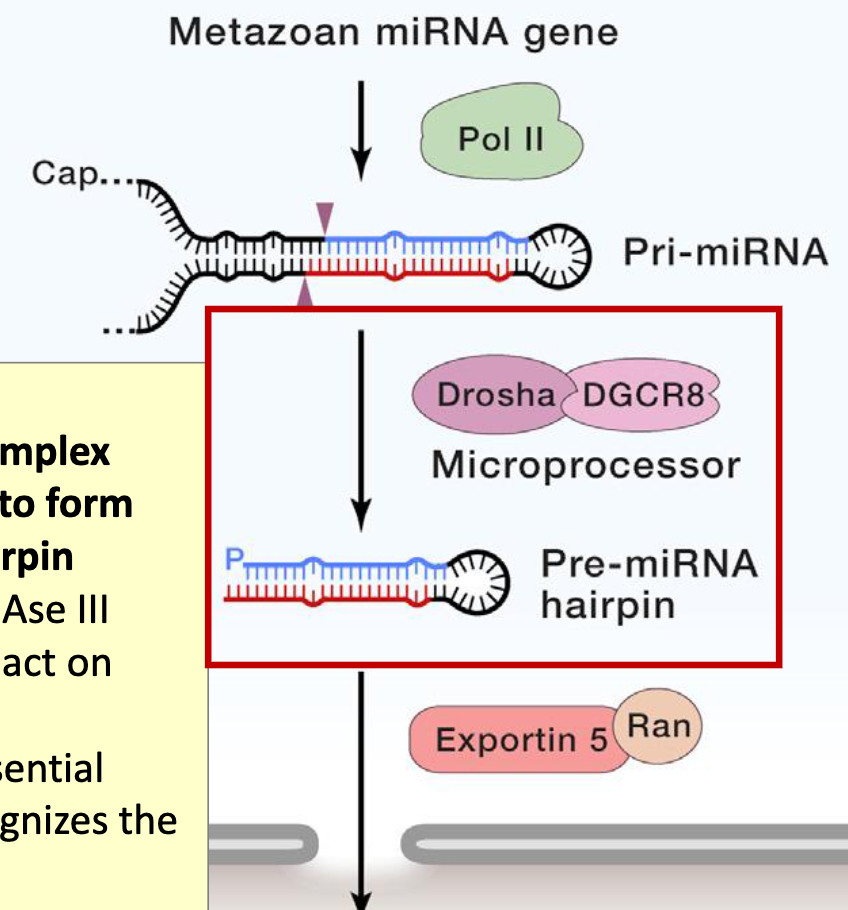
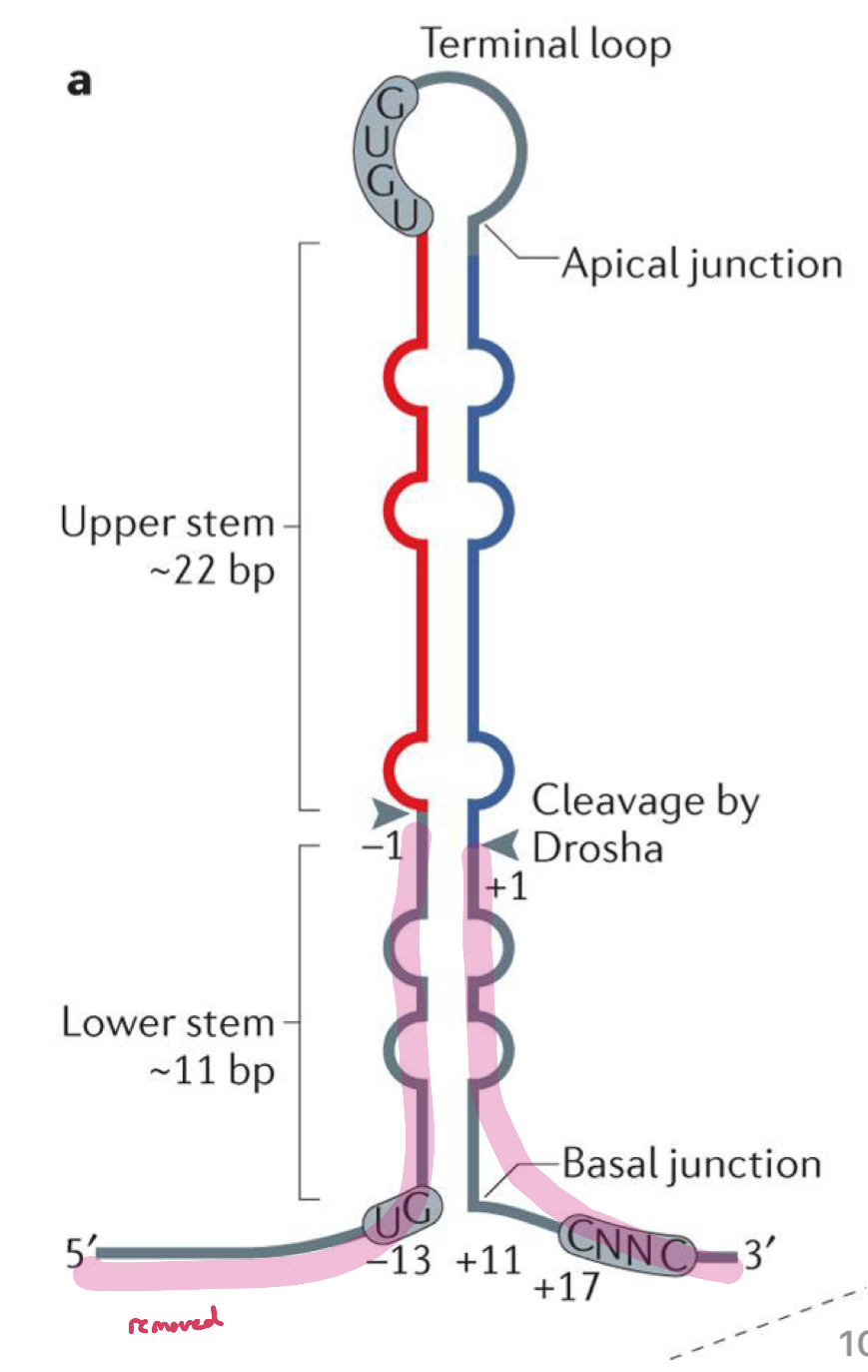
Drosha Cleavage
nuclear enzyme (a class II RNase III) that acts as the primary "scissors" in microRNA (miRNA) biogenesis
pri-miRNA (long transcript) → pre-miRNA (hairpin ~70nt)
pri-miRNA has a hairpin structure with the basal junction, which is a major reference point for Drosha cleavage. The other motifs also play a role in placement of the cut sites
at least 3 of these motifs appear in 79% of human miRNAs (do not need to contain all 4 motifs for proper cleavage)
Drosha cuts the RNA about 11 NT up from the basal junction, and about 22 NT down from the apical junction
Biogenesis and function of canonical miRNAs of animals: Step 3
Export of the pre-miRNA hairpin to the cytosol
transport complex is formed by Exportin 5 (EXP5) plus Ran (a GTP-binding protein)
nucleic acids don’t have export signals, so it hitchhikes uses EXP5/Ran complex
GTP is hydrolyzed upon transport of the pre-miRNA thru the nuclear pore. This causes release of the pre-miRNA from the transport complex into the cytosol (complex becomes inactive)
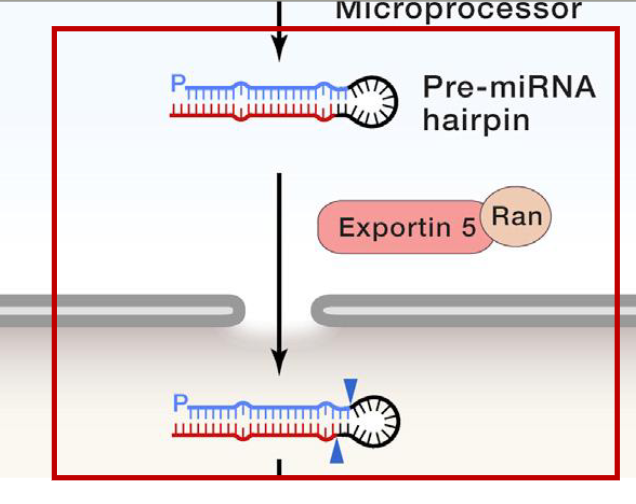
Biogenesis and function of canonical miRNAs of animals: Step 4
Dicer cleaves the loop off the pre-miRNA, leaving a miRNA duplex
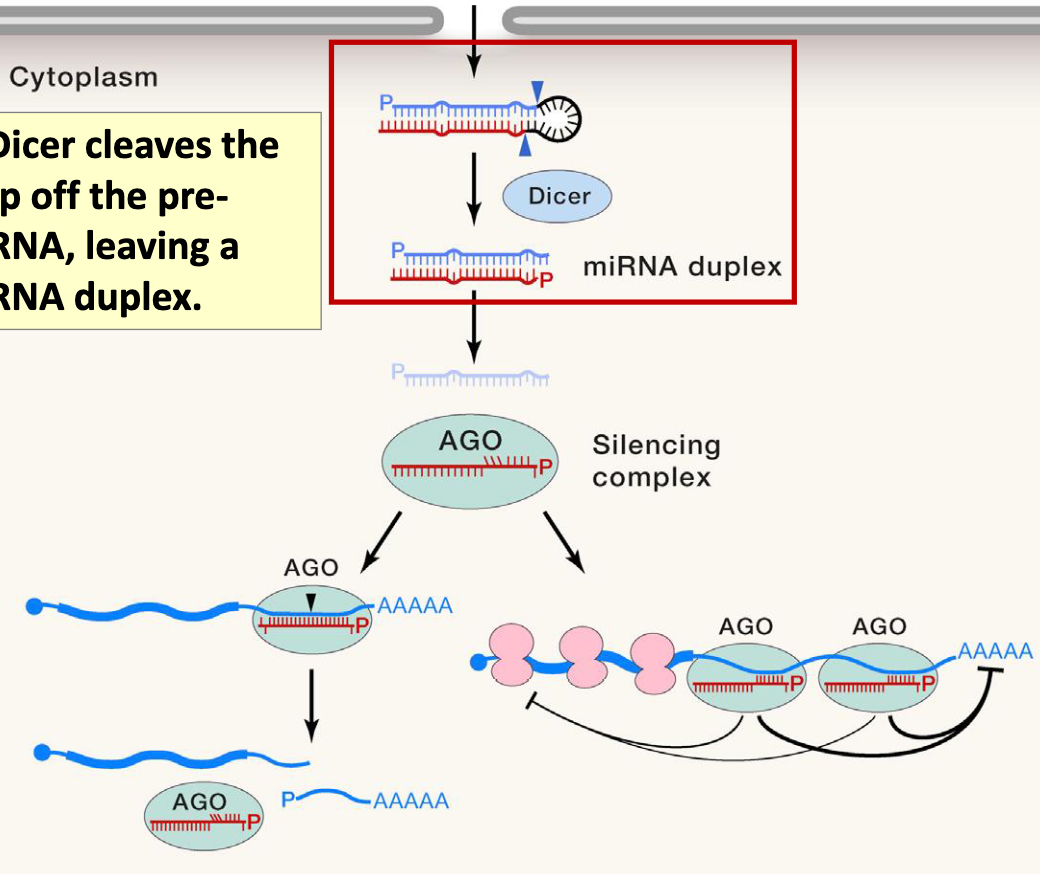
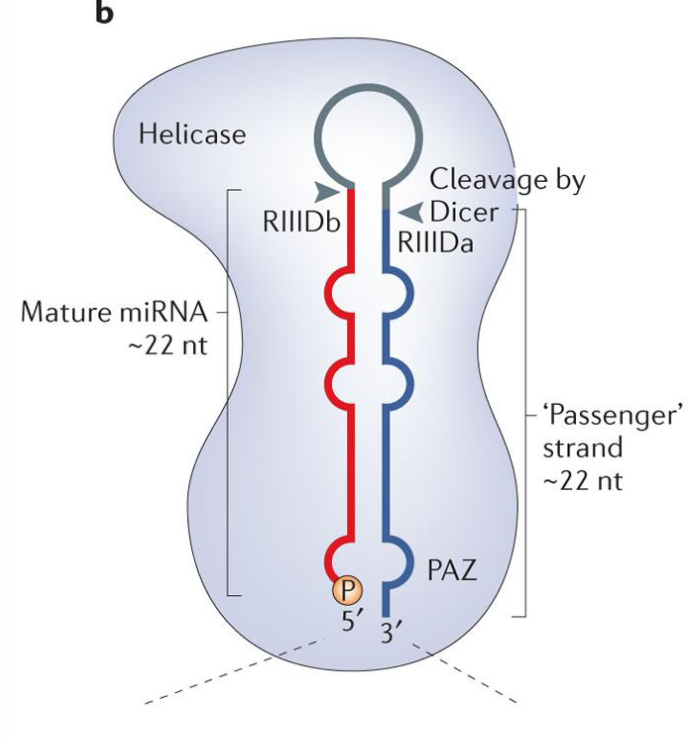
Dicer
Dicer binds the pre-miRNA in a particular orientation, based on the 3’ overhang left by Drosha
once bound, it cleaves the hairpin at a fixed distance from the 3’end (typically 21-25 NT, depending on the species and type of Dicer protein)
Dicer may also be bound w/ cofactors, these are other dsRNA-binding proteins
eg. TRBP= TAR RNA binding protein, a protein that modulates Dicer function in humans
Biogenesis and function of canonical miRNAs of animals: Step 5
miRNA duplex is then loaded onto the AGO (Argonaute) protein to form the RNA-induced silencing complex (RISC)
assembly involves loading and unwinding the miRNA duplex
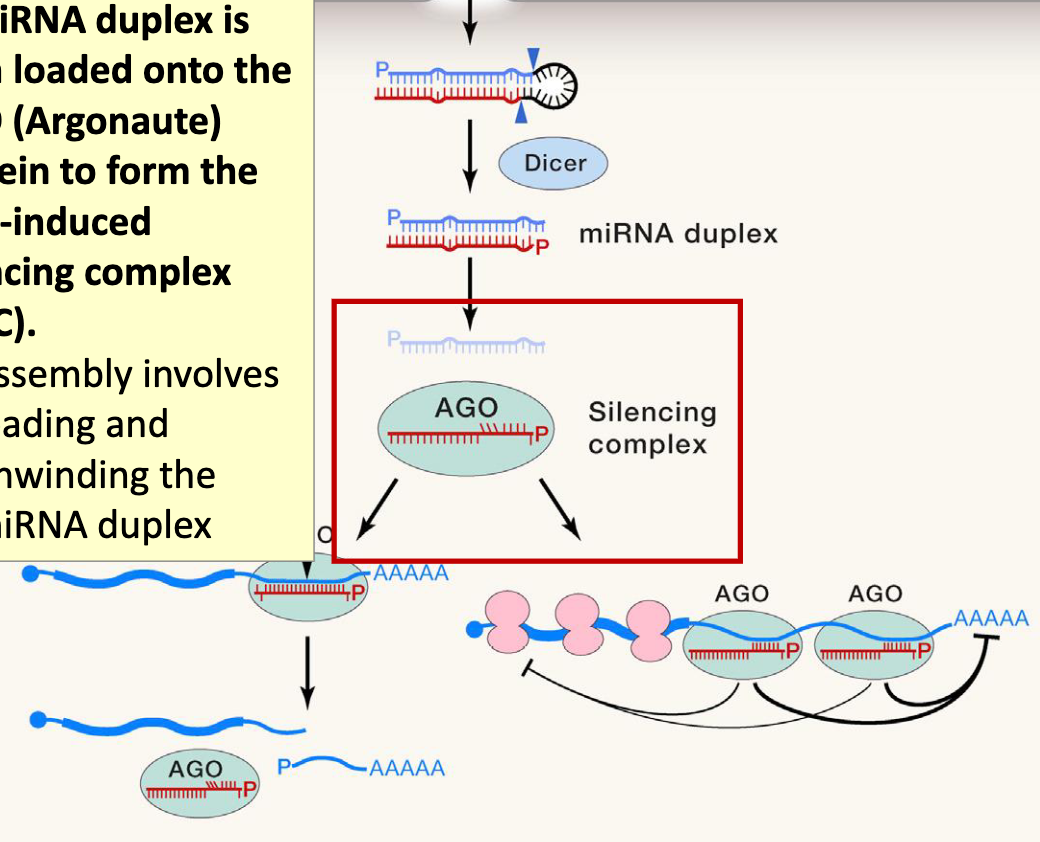
RNA-induced Silencing Complex
multi-protein ribonucleoprotein complex that uses a small guide RNA—microRNA (miRNA) or small interfering RNA (siRNA)—to target and silence specific messenger RNAs (mRNAs)
core of this complex is an Argonaute (AGO) protein, which binds the guide RNA and acts as an endonuclease (or "slicer") to degrade target mRNA, thereby reducing gene expression
the loading complex binds the dsRNA and brings it to the Ago
one of the RNA strands is degraded the remaining RNA strand becomes the guide RNA
the actual process of loading a dsRNA complex and activating the RISC is the same for miRNA and siRNA
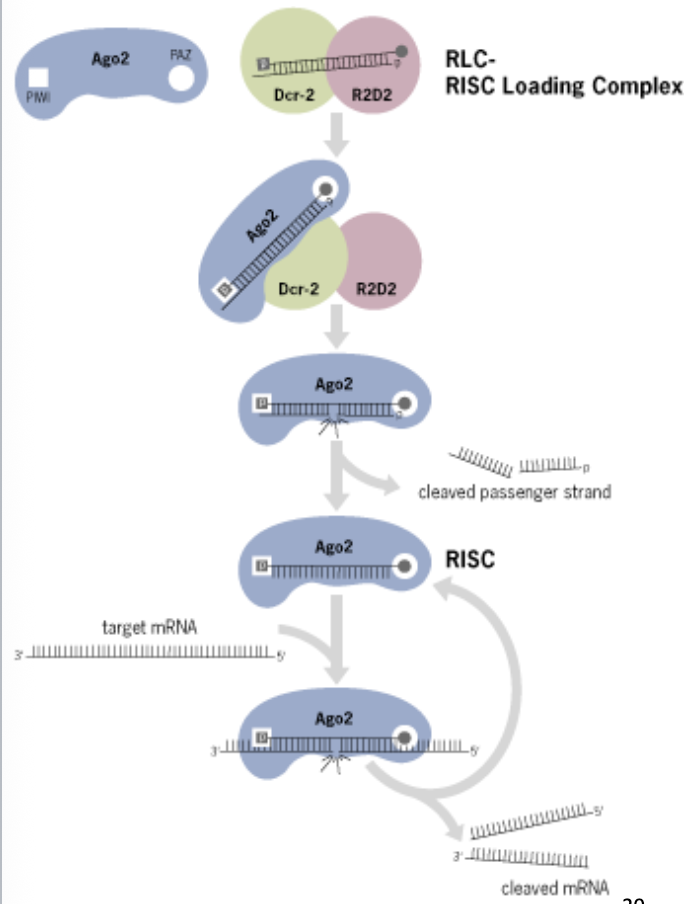
Biogenesis and function of canonical miRNAs of animals: Step 6
RNA-induced silencing complex (RISC) uses the guide RNA to bind to the target mRNA
the complex then silences the mRNA
if the gudie RNA is a perfect match, the target mRNA is degraded (siRNA pathway)
if the guide RNA has some mismatch, translation of the target mRNA is suppressed (miRNA pathway)
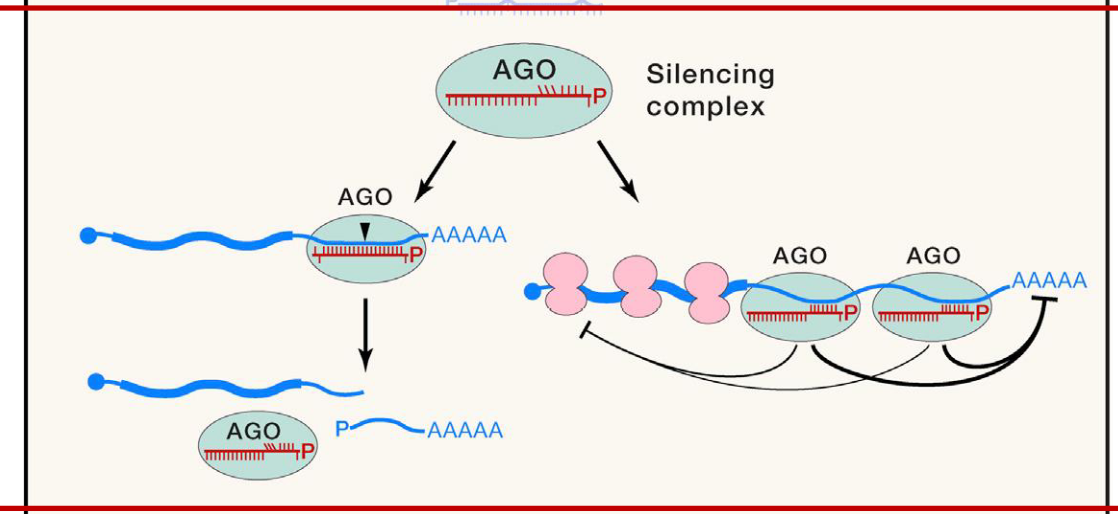
Human Ago Proteins
4 different Ago proteins (Ago1 → Ago 4)
only Ago2 is capable of cleaving perfectly matches RNAs
all 4 Ago proteins interact w/ translational machinery and suppress translation of mRNAs. The mRNAs eventually decay
there is no strict sorting of dsRNA types, they all work with miRNA and siRNA duplexes
Contrast w/ Drosophila Ago proteins
Drosophila have 2 diff Ago proteins (Ago1 and Ago2)
Ago1 preferentially loads miRNA duplexes, whereas Ago2 preferentially loads siRNA duplexes
sorting is also guided by slight differences in the 5’NT (is it a U or a C?)
likewise, C. elegans Ago proteins distinguish b/w miRNA and siRNA
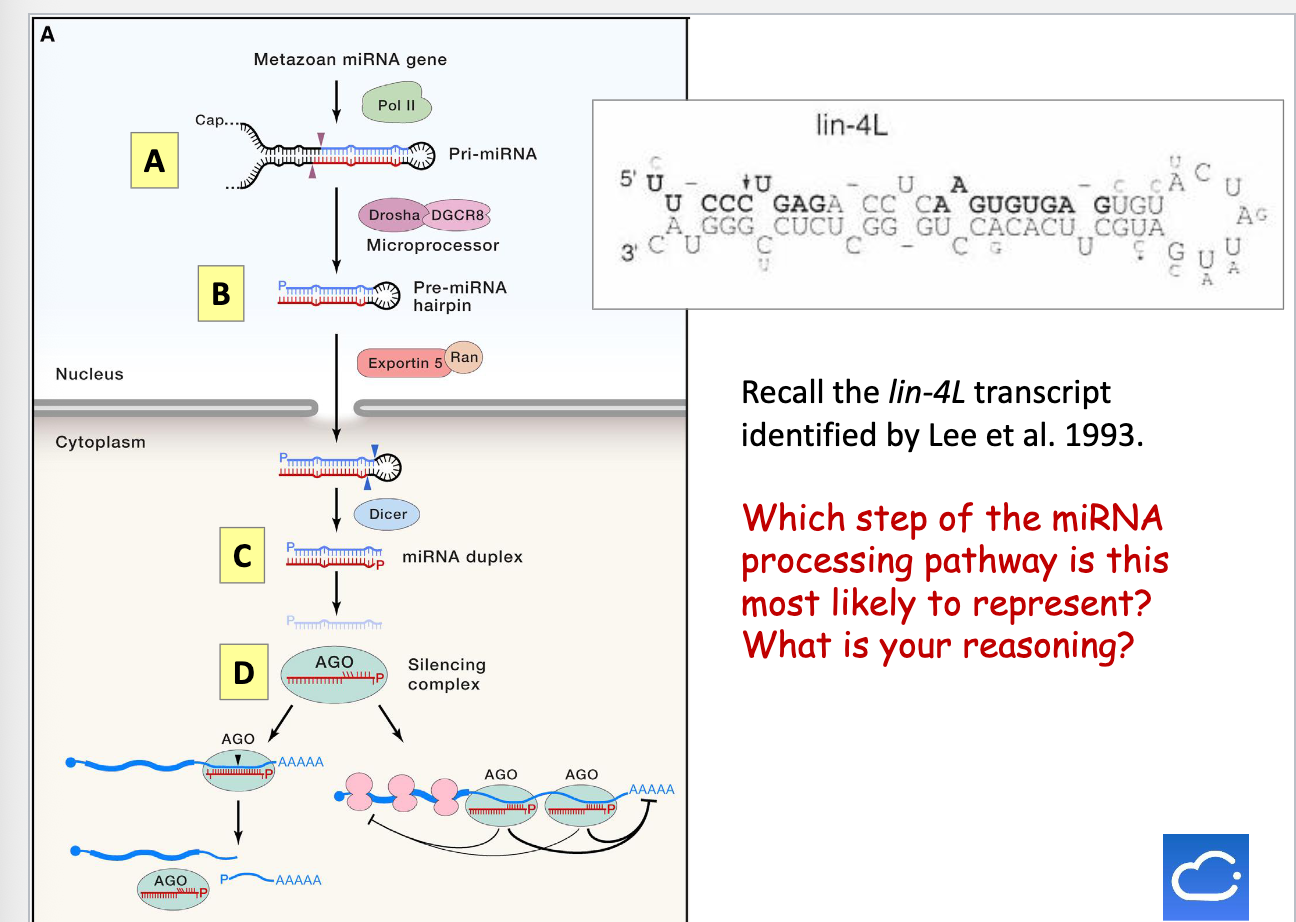
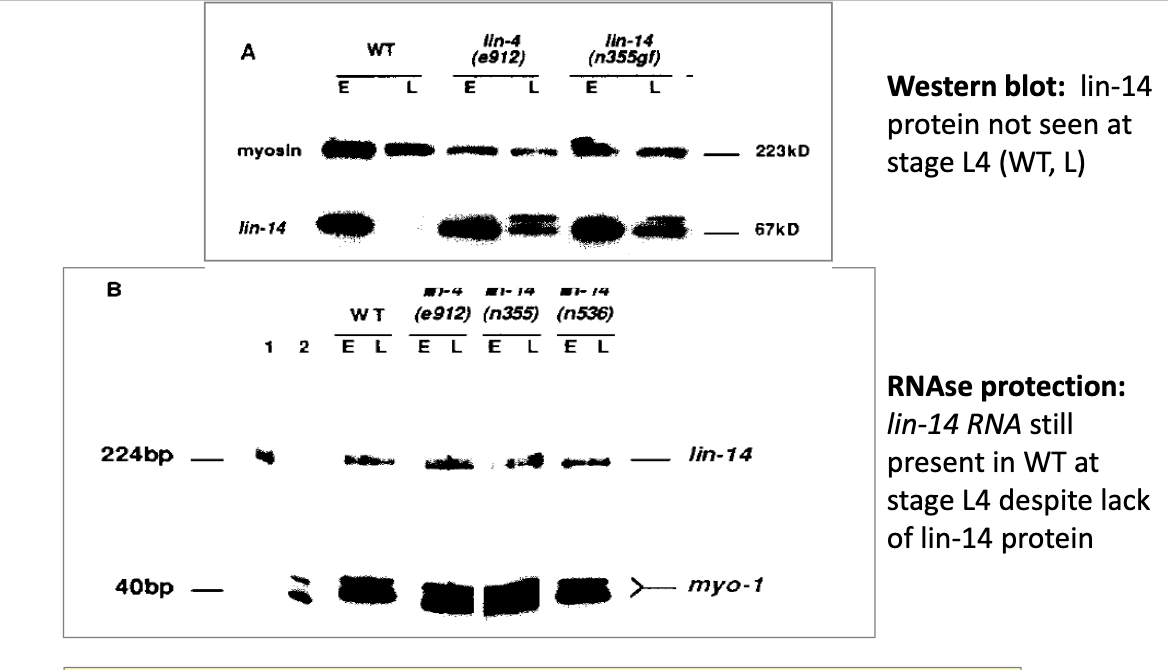
Recall the lin-4L transcript identified. Which step of the miRNA processing pathway is this most likely to represent?
Step 2: Cleavage by the microprocessor complex (Drosha +DGCR8) to form the Pre-miRNA hairpin
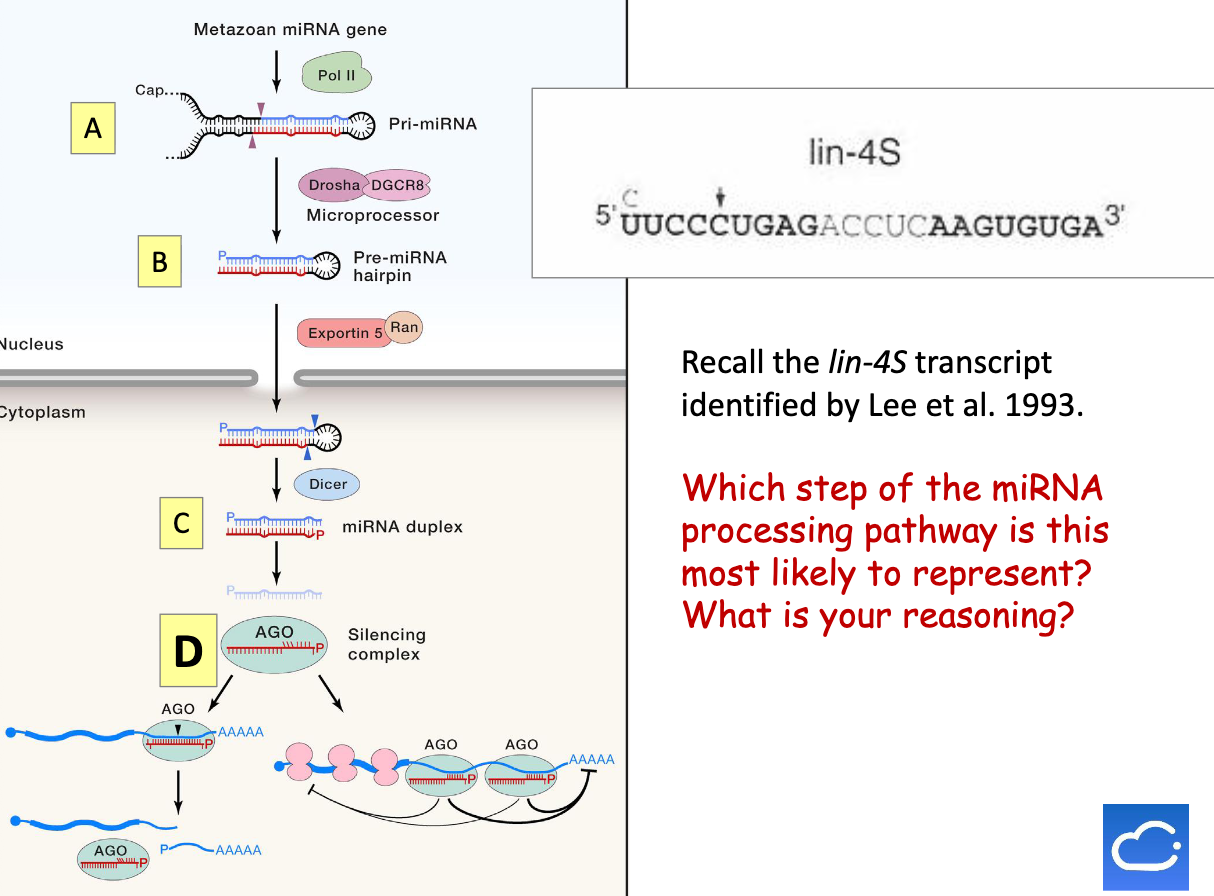
Recall the lin-4S transcript identified. Which step of the miRNA processing pathway is this most likely to represent?
5) miRNA duplex is then loaded onto the AGO (Argonaute) protein to form the RNA-induced silencing complex (RISC)
How would you predict the lin-4 miRNA to regulate expression of lin-14
Repress the translation of lin-14 mRNA
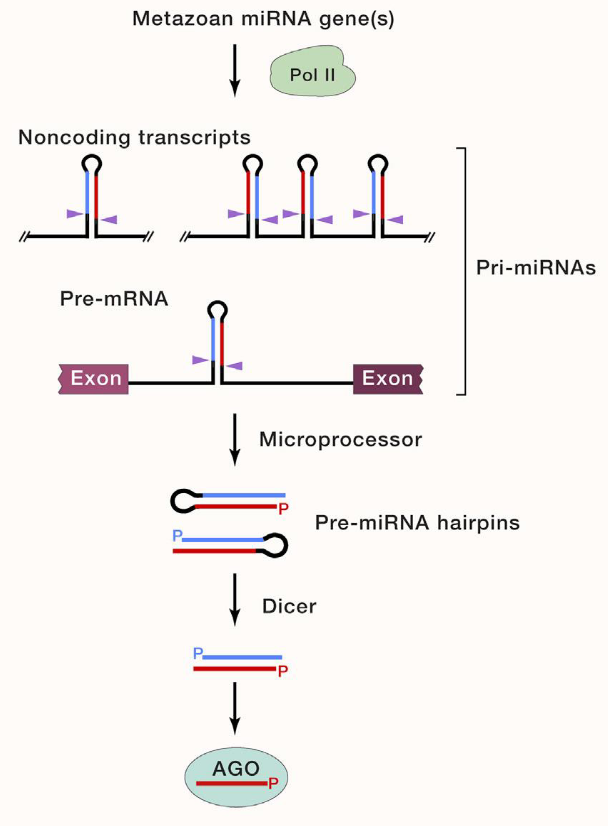
Typical miRNAs
transcribed by RNA pol II
are transcribed from noncoding sequences outside genes or within introns
are sometimes derived from exon sequences
are sometimes derived from transcripts that form multiple hairpins (and multiple miRNAs)
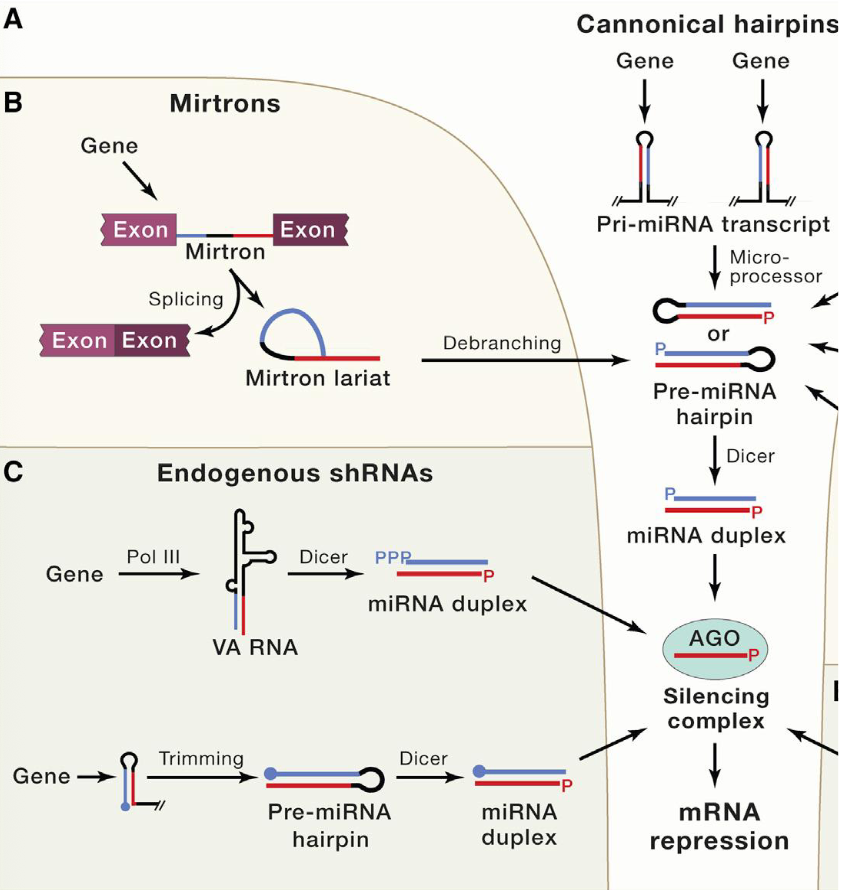
Can miRNAs enter the hairpin processing and silencing complex thru other sources
Yes?
don’t memorize the figure, but know that there are other RNA transcripts that can be cleaved to generate pre-miRNA hairpins or RNA duplexes, that can enter the miRNA pathway and be processed for loading onto Ago
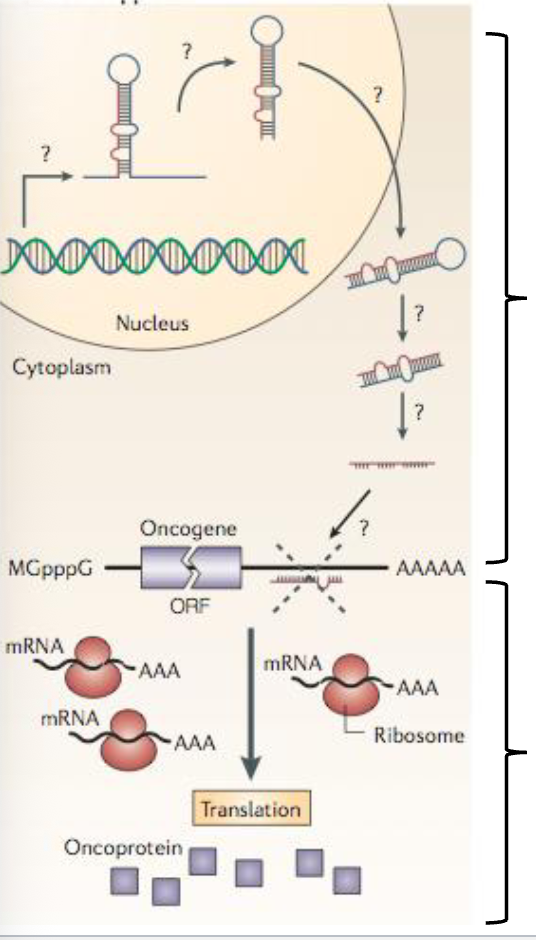
Recall: Tumor Suppressors + Proto-Oncogenes
Tumor suppressors: their normal function is typically to suppress cell proliferation and induce cell death. Loss of function can result in cancer
Proto-oncogene: their normal function is to promote proliferation and to protect against cell death. Gain of function can result in cancer
miRNAs can also act as tumor suppressors or proto-oncogenes
You identify a miRNA that suppresses cell proliferation. Which genes are more likely to have binding sites for this miRNA within their mRNA transcripts?
A. Map kinase: a protein that phosphorylates and activates other kinases in a cell signal pathway.
B. p21: A protein that binds and inhibits Cdk/cyclin complexes.
C. Wee1: A kinase that adds an inhibitory
A) Map kinases act in signal transduction pathways to promote cell proliferation
You identify a miRNA that suppresses p21, a protein that binds and inhibits Cdk/cyclin complexes. What would happen if this miRNA had a gain of function mutation?
A. Increased cell proliferation
B. Decreased cell proliferation
A
miRNA as tumor suppressor
normal role is to suppress translation of a proto-oncogene
a loss of function mutation could result in elevated levels of oncogene expression, and formation of tumor formation
miRNA as proto-oncogene
normal role is to regulate the translation of a tumor suppressor gene
a gof mutation could promote cancer by abnormal blocking expression of the tumor suppressor
Are miRNAs associated w/ human cancers?
Yes, several!
e.g. let-7 family members
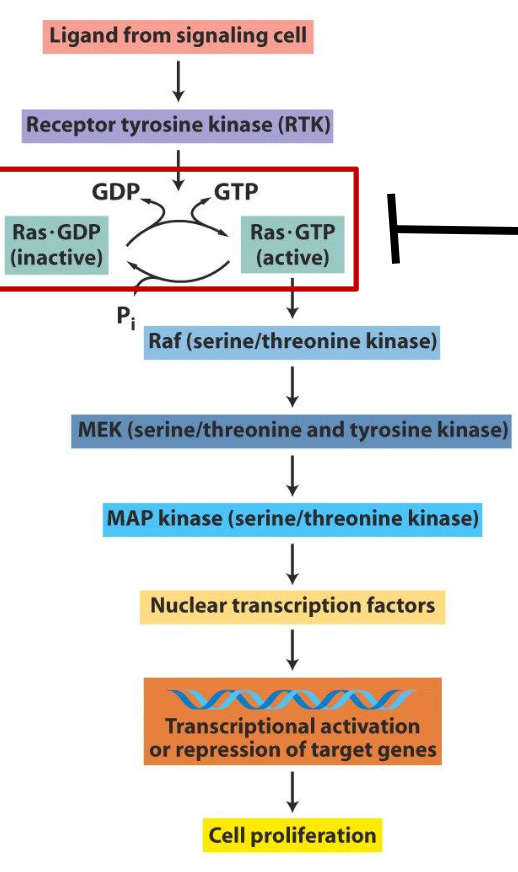
Let-7 on Cancer: Ras
recall the role of Ras in receptor tyrosine kinase signaling (RTK) pathways; when RTK is activated, adaptor proteins help load Ras w/ GTP to activate Ras
active Ras then triggers downstream pathways that promote cell growth and division
Ras turns off when it hydrolyzes GTP → GDP
let-7 family members negatively regulate Ras
if there’s a loss of let-7 → no repression of Ras → increase in cell division
Let-7 in C.elegans
let-7 is required for cell fate determination and terminal differentiation of seam cells (specialized epithelial cells)
cells exit cell cycle after terminal differentiation
let-7 LOF mutation results in seam cells that keep cycling
Let-7 in Humans
let-7 family members have mapped to regions that are deleted in cancers - a tumor suppressor
overexpression of let-7 miRNA in a human lung cancer cell line inhibited its growth
Exp to Identify the Target Genes of the miRNA let-7
Strategy: computational search for C. elegans genes with sequence complementary to let-7
revealed that miRNA let-7 binds to the 3’UTR of let-60 and represses its expression
let-60 has complementary sites within its 3’UTR → human ortholog is RAS (a proto-oncogene)
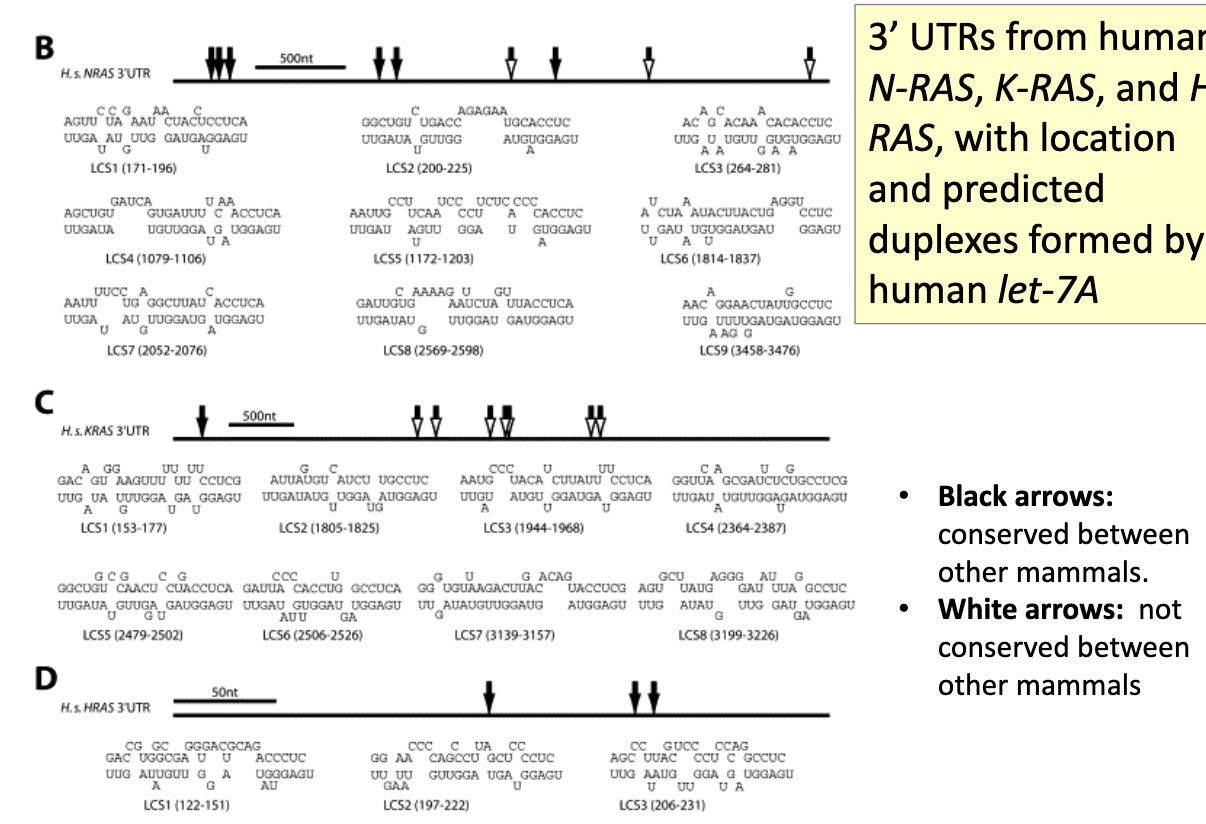
After showing let-7 targets let-60 (Ras) in worms, does this also happen in humans? Exp
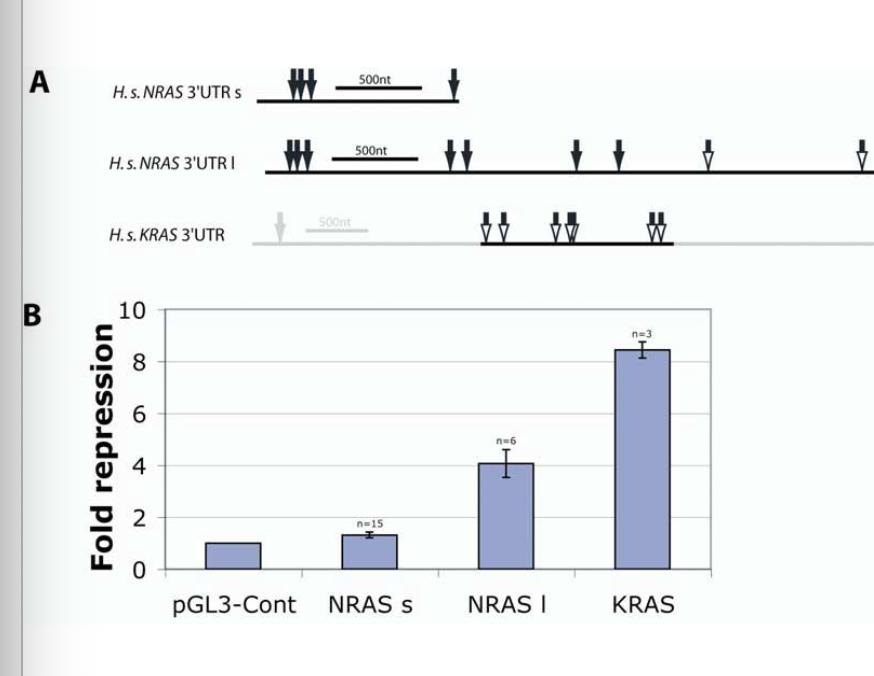
looked at 3’UTRs from human N-RAS, K-RAS and H-RAS, w/ location and predicted duplexes formed by human let-7A
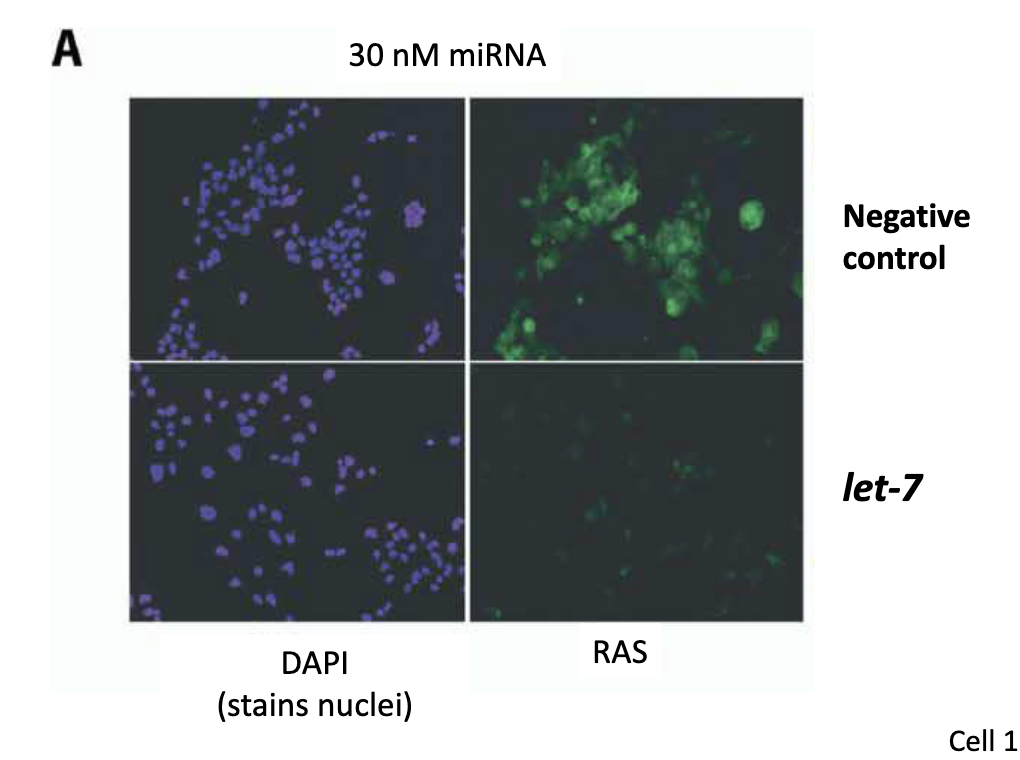
Test if let-7 suppresses translation of its target Data
Experiment:
human liver cell carcinoma cells transfected w/ a dsRNA let-7 precursor
human RAS protein detected w/ fluorescent antibodies
Describe the data:
let-7 represses Ras expression (same amount of nuclei, but little fluorescence)
Test if let-7 regulates translation of RAS through through the RAS 3’UTR
Experiment:
N-RAS (short and long forms) and K-RAS 3’UTR was fused to a luciferase reporter gene.
Reporter constructs were transfected into HeLa cells (human cell line), and luciferase activity was normalized to the control sample (luciferase vector lacking the RAS 3’UTR)
Interpretation of Results
let-7 suppresses expression of the luciferase reporter
The longer N-RAS 3’UTR is subject to greater regulation