Chapter 16b - Eukaryotic Regulation
1/22
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
In prokaryotes, expression of
many genes is __ by default,
until something turns it __
on, off
In Eukaryotes, expression of
most genes is __ by default,
until something turns it __
off, on
Each of our cells carry the exact same DNA with the exact same genomic sequence.
So how do we get such different properties?
Genomic Regulation
Eukaryotes have a variety of ways to regulate.
At the structural level (has to do with tightly wound around histones our linear DNA molecules are)
At the transcription level (combinatorial action of many different transcriptions factors and sometimes ncRNA)
At the translation level (has to do with the pace at which mRNA molecules degrade and how translation initiates)
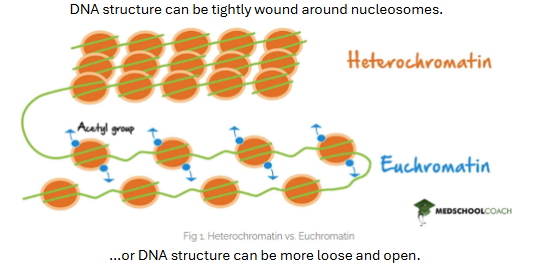
If DNA is in heterochromatin state, expression of
genes in that area is __ . But if it is in the euchromatin state, expression in that area is __
repressed, possible
Eukaryotic cells shift
different regions of DNA
between euchromatin and
heterochromatin as a
__ strategy
regulatory
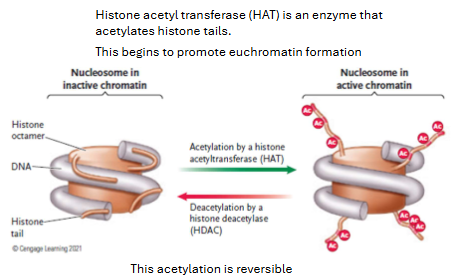
Acetylation makes nucleosomes bind DNA less ___
tightly
Epigenetics = the study of how gene expression is
changed by chemical modifications to chromatin that
__ gene sequence
do not impact
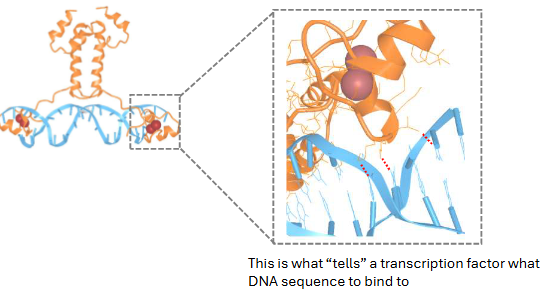
Regulation at Transcription Level:
Transcription factors participate in __ intermolecular _ with DNA. This is x
weak, interaction, non-covalent
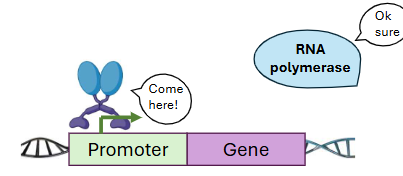
Some transcription factors __ gene expression. They do this by recruiting _ to the promoter region.
active, RNA Polymerase
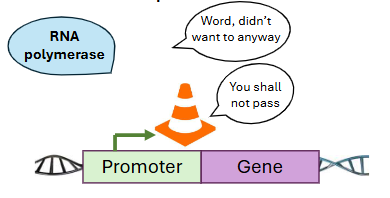
Other transcription factors act like traffic cones, __ gene expression.
silencing
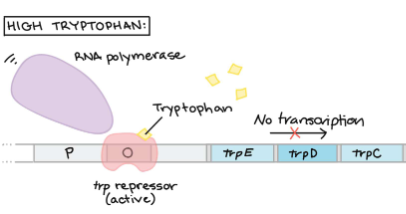
Tryptophan Operon (TRP) is a transcription factor that __ transaction.
represses
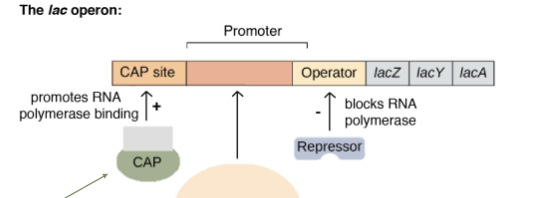
CAP is a transcription factor that activates when there isn’t any glucose around. It is an __ transcription factor.
activating
There are approximately ~___ different transcription
factors in the human body that we know of
1,600
Eukaryotes do not have __
Each Eukaryotic gene has its own promoter and
polyadenylation signal.
operons
Eukaryotes employ ____ of
transcription factors to regulate how __
transcription __
complex combinations, often, initiates
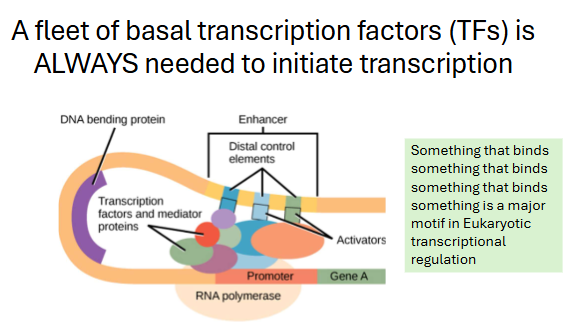
Each colorful circle is a ___ transcription factor (TF)
These have nothing to do with regulation, they must be
__ every time RNAPII initiates transcription.
general, recruited
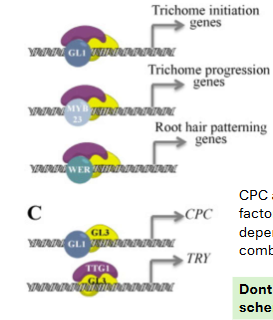
Don’t forget. Similar combinatorial schemes to combinational TF-based regulation also exist for gene ___.
repression
Hormone cascades coordinate regulation of related
genes via TFs. This will assist in turning on all __ genes throughout the _.
related, geonome
Long noncoding RNA (lncRNA) does not carry gene sequence, and carries out diverse regulation. Here, long RNA means RNA greater than __ bases in length. They can act like transcription factors. They can bind to _ themselves and prevent or promote translation.
200, mRNA
Translation Level:
Manipulation of UTRs, caps, and tails of
mRNA all present opportunities for post-
transcriptional regulation
3’ UTR and polyA tail largely influence mRNA half-life. Often, longer poly(A) tails = more stable mRNA molecule. The lengths and motifs of sequence contained in 3’ UTRs can either increase or decrease mRNA half-life.
Rates of initiation is another post-transactional regulatory strategy