BIO417 Midterm
1/161
Earn XP
Description and Tags
Lectures 1-5
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
162 Terms
Genomics
Focuses on the structure function, evolution, mapping, and editing of genomes
Studies all its individual genes including interactions of those genes w/ each other, and also takes external signals (environment) into account
Genome biology
Studies quantification, packaging, storage, proof reading, and replication of genetic information
Genome size is measured by
Number of bases or C-value
C-value
Amount of DNA in a haploid nucleus (picograms)
The C-value paradox
Nuclear genome size varies strongly among species; there is no apparent relationship to the number of genes encoded in the genome that reflects the complexity of the organism
The phosphate groups and sugar backbone of the DNA are…
Polar and hydrophilic
The DNA bases are…
Hydrophobic; bases interact w/ each other to minimize interaction w/ the outside
Bases can’t mispair because…
It changes the space they take up (more or less) so they don’t fit into the backbone
Average twist angle for a single row
36º
Number of bp for a full twist
10
3 families of helicies
A-DNA, B-DNA, Z-DNA
Right handed helicies
A-DNA and B-DNA
Z-DNA
DNA-RNA hybrid structure; left handed
B-DNA
Average DNA conformation
Nuclosome
Basic unit of chromatin; made up of 147 bp of DNA, 2 H2A, 2 H2B, 2 H3, 2 H4, and a H1 linker histone
DNA-histone interaction
Small, basic, proteins attracted to negatively charged DNA; non-covalent
Histones
Small basic proteins (100-220 residues; 11-15 kDa, H1 ~22 kDa)
Conserved histone fold
Structured motif (more so than DNA sequence) conserved
N-terminal tail = variety of mods
Number of contact points between DNA and histones
12 overall (8 histones → 4 dimers x 3 contact points w/ DNA)
Histone dimers interact w/..
The minor grove
Total dimer interaction
147 bp / 10 bp for a full turn = 14.7 minor groves
Centromere function
Segment of chromosomal DNA that provides attachment sites for the kinetochore
Centromere structure
CEN region → minimal region that supports chromosomal segregation
Rich in repeats
Humans: 𝛂-satelite DNA; 171-bp
Heterochromatic
Binds platform proteins (kinetochore) which binds microtubles
CENP-A
Centromere-specific H3 variant; not sequence specific
Kinetochore and microtubule interaction
Lose until there’s force on the microtubule, then it gets really tight
CENP-B
Protein in humans that recognizes centromere-specific repeats and turns sequences into centromere formation (accessory; not neccessarily required b/c the process isn’t sequence specific)
Telomere function
Region of specific repetitive DNA sequences at the end of a linear eukaryotic chromosome that protects chromosome ends and shortens slightly as cells divide
Telomere structure
Short tandem repeat
T1-4A0-1G1-8
Human: TTAGGG
Tetrahymena: TTTGGGG
Arabidopsis: TTTAGGG
Can form G-quadruplexes
Length in humans: 5-20 kb
Shelterin complex
T-loops
Shelterin complex
Cover and compact repeats on the telomere; protect form repair
T-loops
Protruding single strand end, loops back into telomeric repeat
Hayflick limit
After a certain number of cell divisions, the telomere shortens until the cell stops dividing (apoptosis)
Telomerase
Attracted to telomeric repeats and synthesizes additional repeats; DNA pol unable to replicate the 3’ ends of DNA strand
Key features of the chromosome
Centromere
Telomeres
Many origins of replication
p and q arm (p = shorter; q = longer)
Genes (interspaced)
Kinetochore protein contact points
Repetitive sequences
Pericentromeric region
Metacentric chromosome
Centromere in middle (ex Chr1)
Submetacentric chromosome
Centromere towards the middle (ex Chr 4)
Acrocentric chromosome
Centromere towards the tip (ex Chr 14)
Telocentric chromosome
Centromere at the tip
Cytogenetic stains
Allow to further characterize chromosomes
Giemsa → G-bands
Quinacrine → Q-bands
Reverse giemsa → R-bands
Telomere visualization → T-bands
Silver nitrate staining → NOR (nucleolar organization region; code for rRNA)
Giemsa stain
Typical stain
Specific for the phosphate groups of DNA in AT-rich regions (heterochromatic regions = dark)
Mix of methylene blue, eosin, and Azure B (readymixed powder)
With cell arrest in metaphase or prometaphase
Used to generate karyogram
Karyotype
General appearance of the full set of chromosomes of an individual
Can be obtained and represented in image (micrograph) of an individual’s metaphase chromosomes
Karyotype sorted by
Size (assigned a number based on size, largest #1)
Chromosome type (autosomes first, sex chr last)
Centromere position
Short (p) arm on top
Further divided into groups based on size and centromere position
Ideogram for G-banding
Standardized map of a chromosome set
Centromeres = dark grey
Used as a reference point
Can visualize missing/extra chromosomes or pieces of chromosomes
Holocentric chromosomes
One size-restricted centromere per chromosome
In diverse eukaryotic linages including algae, some nematodes, insects, spiders, plants (sedges)
Convergent evolution
Diffuse kinetochores/several distinct microtubule binding sites
Rules for karyotyping
22 autosomes by size
Type (sex chromosomes at end)
p arm oriented to the top
Chr classified into 7 groups (A-G) by length and centromere position
Ideograms for G-banding of human metaphase chromosomes
>400 bands (n)
Centromeres = dark grey

Myrmecia pilosula
Primitive ants
2n = 2-32
Model for chromosome set evolution
Diploscapter pachys
Single pair of chromosomes
Relative of C. elegans
Truncated meiosis (absence of functional meiosis I)
Only asexual reproduction
Lack of telomeric repeats and genes for telomere protection
Polyommatus atlantica
2N = 458
10x more chromosomes than most butterflies
Highest number of chromosomes in non-polyploid organisms (smalles chr size possible)
Result of rapid fragmentation of autosomes
Centromeric region across whole chromosome
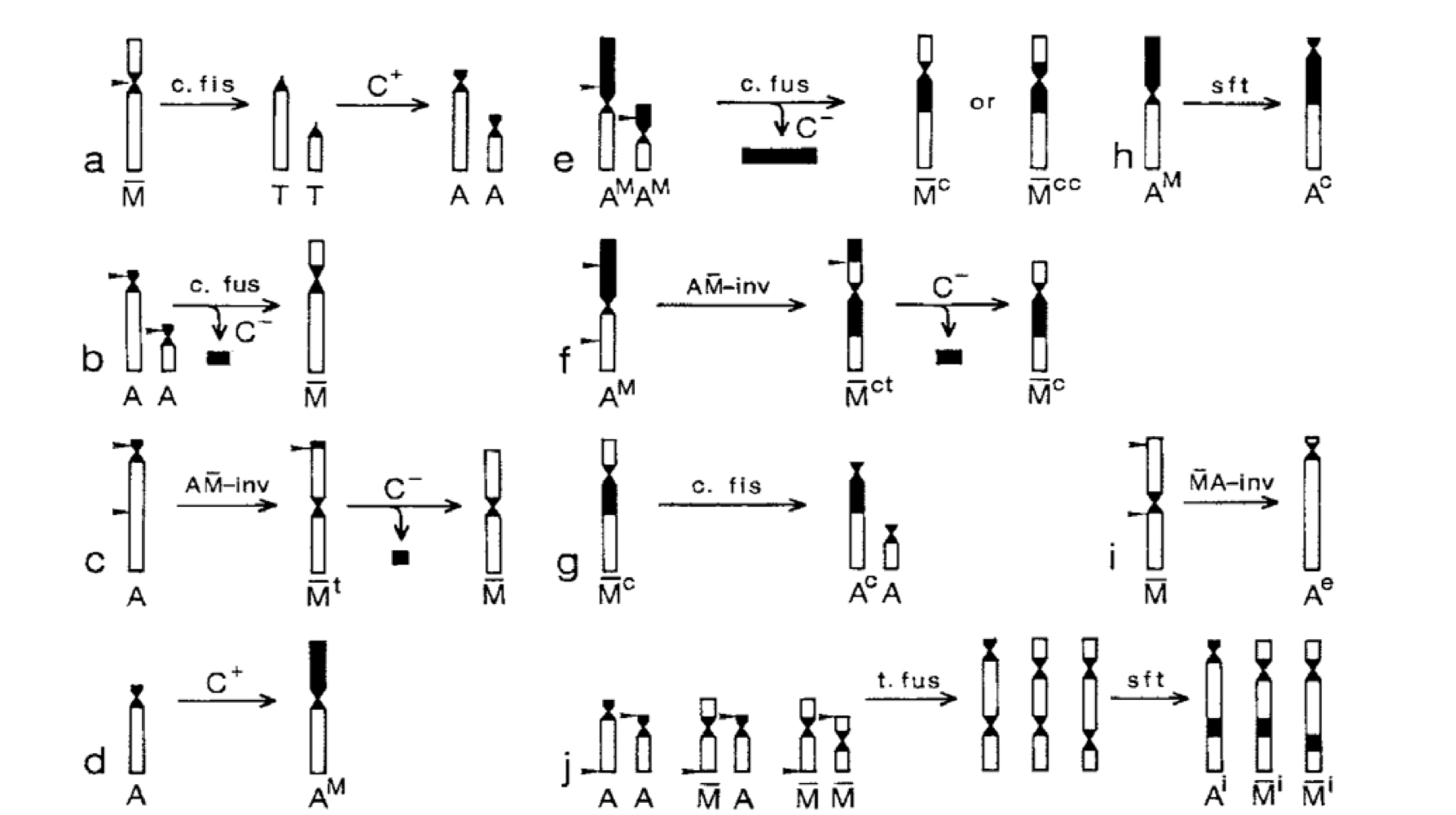
Chromosome aberrations
Substantial change to the structure of a chromosome; aka chromosome mutations
Types of chromosomal aberrations
Deletions, inversions, insertions, duplications, exchanges, fusions (translocations)
Deletions and duplications
Alter amount of genes
Inversions and translocations
Rearrangements of genes
Deletions
Chromosomal breaks
Recombination at incorrect places
Often detrimental
Phenotypic consequences often correlate with the size
Cri-du-chat syndrome characteristics
High pitched cry
Intellectual disability
Delayed development
Small head size
Low birth weight and weak muscle tone
1:20,000 - 50,000 newborns
Cri-du-chat syndrome chromosomal defect
Section of chromosome 5 missing
5p- / 5p15.2 / 46,XX,del(5)(p15.2)
Cri-du-chat syndrome molecular mechanism
CTNND2 gene affected: delta catenin, neural migration
Others (SEMA5A, hTERT)
Subfunctionalization
Domains are separated into different genes, allowing for more precise regulation
Neofunctionalization
Second copy takes on different function (evolution)
Duplications can result in…
Subfunctionalization, neofunctionalization, or degeneration/gene loss
Duplication and gene families example
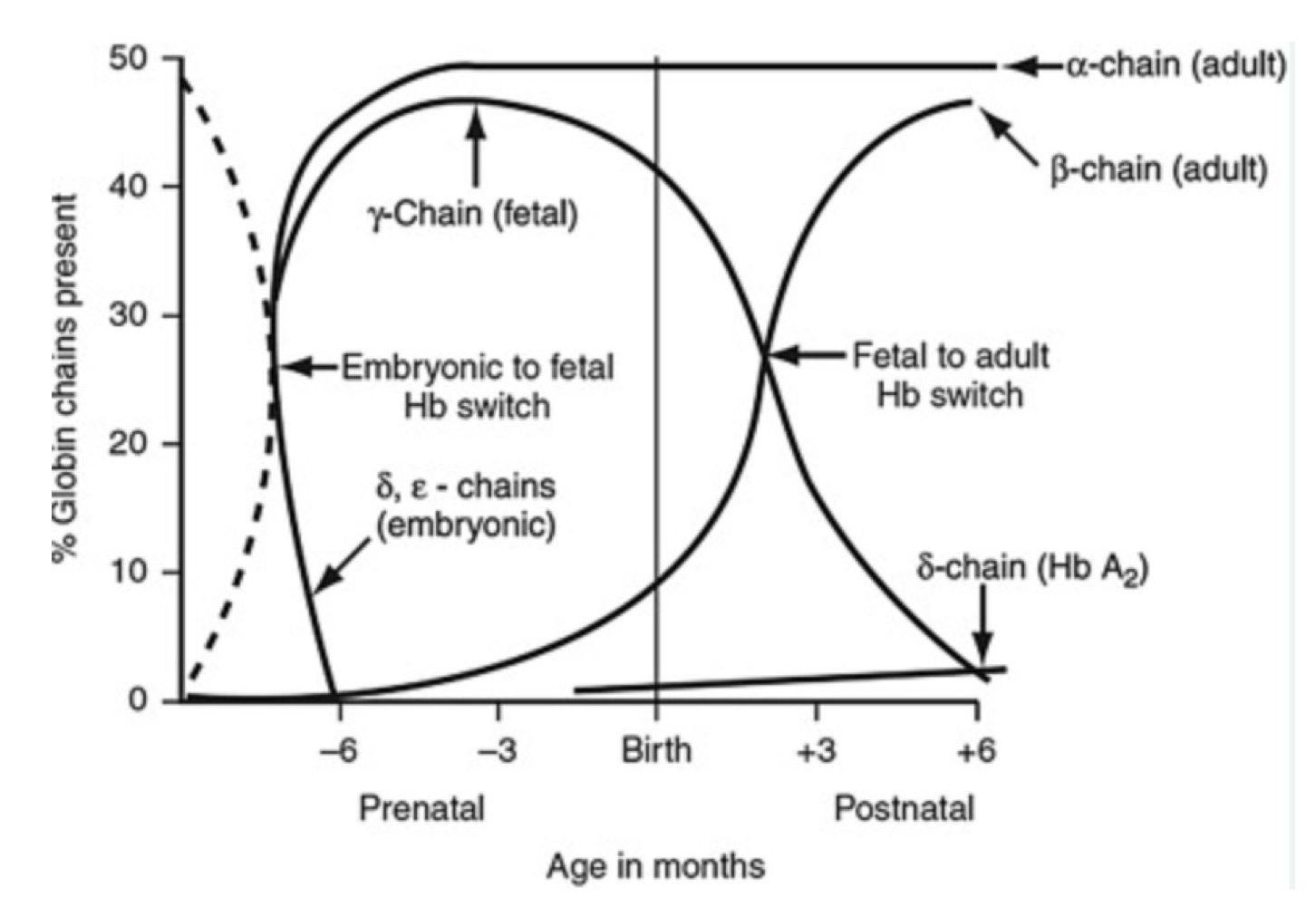
Evolution of the globin gene family in humans
Hemoglobin β chain genes
Has control region that regulates entire gene cluster
Genes begin w/ embryonic globin chain and the order corresponds with development in the order of use
Locus as chromatin hub
Shared locus control region
Folding and gene activation depending on developmental need
Inversion
Chromosome segment is flipped in opposite orientation
Classified according to location relative to centromere
Often has no phenotypic consequence (genetic info still there)
Variation in size; rather common
Pericentric inversion
Contains centromere
Paracentric inversion
Doesn’t contain centromere
Examples of inversions with a phenotype
Hemophilia (type A, X linked inversion) → factor VIII, blood clotting protein
Crossing over with paracentric inversions
Creates a normal chr, an inverted viable chr, a dicentric chr, and an acentric chr
Acentric fragment is lost and dicentric fragment randomly breaks, creating two non viable deletion products
Crossing over with pericentric inversion
Creates a normal chr, an inverted viable chr, and two duplication/deletion chrs that are nonviable
Translocations
Cross over and non-homologous recombination
Balanced/reciprocal are often without phenotype
Can produce abnormal gametes (semisterility)
Philadelphia chromosomes
Chronic myelogenous leukemia (CML)
Translocation creates new fusion gene on chr 22
Kinase that becomes more active
Cells proliferate without being regulated by cytokines
Ring chromosomes
Fusion of broken ends
Maintenance doesn’t work, so the cell tries to connect to something so its not completely unprotected
Usual ring chromosomes
14, 10, 13, 4, 20
Aneuploidy
Unbalanced chromosome set; number of chromosomes is not a multiple of a set (trisomy and monosomy)
Colchicine
Inhibits microtubules polymerization (binds tubulin subunits)
Inhibits assembly of the mitotic spindle
Can be used for generation of altered chromosome sets (polyploids, aneuploids)
Used in karyotyping
Alkaloid from autumn crocus
Datura stramonium (jimsonweed)
Early model for studies on chromosome sets
Plant of nightshade family
Produces tropane (nitrous, bicyclic) alkaloids, psychoactive compounds
n=12 chromosomes; diploid 2n=24
Systematic study of trisomies in jimsonweed
Identified all 12 of the possible 12 forms (complete set)
Cocklebur
2n+Chr6
Narrow leaves
Weak
Inclined to droop
Number of monosomies and tetrasomies in jimsonweed
Only 1 of 12 forms identified
Number of ploidies (1n, 2n, 3n, 4n) in jimsonweed
All 4 of 4 possible forms identified
Incidence of aneuploidy in humans
About 35% of spontaneous abortions but only 0.3% of live births
Most common aneuploidys in humans
Trisomy of chromosome 13, 18, or 21 (smaller chrs; only trisomies that can be tolerated)
Sex chromosome aneuploidies are…
Often nonlethal
How to detect aneuplodies?
Karyogram or FISH
Monoploid individuals
Drone bees
Polyploidy is common in…
Plants
Polyploidy is common in plants and crops b/c…
Plants larger in size
Better stress resistance
Larger flowers
Higher yields
Better adaptability
Polyploidy in wheat
Hexaploid; result of a hybridization between a domesticated tetraploid
Autopolyploids
Multiple genomes of a single species
Allopolyploids
Multiple genomes derived from 2 or more species
Polytene chromosomes
Model for genetic experiments and analyses
Found in Drosophila and other dipteran flies; salivary gland (larvae); interphase nuclei
Endomitosis
Specialized cells undergo repeated round of DNA replication without cell division; found in interphase nuclei
Mutations
Any change in base-pair sequence of DNA
Source of genetic variation
Raw material for natural selection
Occur spontaneously
Can be induced by external factors (e.g. chemicals, radiation)
Can be detected and repaired: proofreading, correction of replication errors, BER (base excision repair), homologous recombination repair
Mutation classification is based on
Molecular change
Phenotypic effects
Location
Mode of generation
Frameshift
Deletion or insertion of any number of bases, except multiples of 3
Substitution
Missense (difference amino acid), nonsense (stop), silent (same amino acid), transition, or transversion
Transition
Pyrimidine replaces pyrimidine, or purine replaces purine
Transversion
Purine replaces pyrimidine or v.v.
Sickle cell anemia cause
Missense mutation
Causes of duplications and deletions
Abnormal events during recombination (non-allelic homologous recombination)
Cross over at misaligned sites
Depurination
Loss of a base in intact double helix (mostly A or G)
Most common type of naturally occurring chemical change
Release of purine base, leaving apurinic site
1:10,000 purines
Recognized by repair, but if missed → mutation
Deamination
Amino group (NH2) → keto group (O)
Changes cytosine to uracil and 5-methylcytosine to thymine (keto)
U can be recognized by repair, it not, mutation during replication
Guanine damination
Changes to xanthine (still pairs with C)