Manuscript 2
1/23
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
what are the modifications in the oncolytic virus used in this paper
uses Ad5,
expressing E1A under TERT promoter (TERT usually off except in cancer cells) → transcriptional targeting
also has deletion in E1B to make it specific
add Ad11 fiber → transductional targeting
E3 region for potency → delete immunomodulatory genes
what is autophagy?
lysosome-mediated degradation of damaged proteins, foreign organisms and organelles
describe the process of autophagy
proteins and organelles to be removed have substrates that are recognized by autophagy receptors (eg. p62)
phagophore is formed
lipids added (eg. LC3-I) to membrane, tagging it to fuse with lysosome
forms autophagosome
autolysosome → fuses with lysosome with degradative enzymes
what does autophagy have to do with adenovirus?
autophagy is increased by adenovirus and seemed to increase adenovirus replication and lysis. Autophagosomes promote Ad replication:
increase amino acids via recycling
increase virus protein synthesis
provide platform for replication & assembly
what effect does autophagy have on cancer?
autophagy promoted immunogenic cell death
release of DAMPs (eg. ATP, F-actin, Calreticulin, HMGB)
autophagy helps package ATP into vesicles and secrete it into the extracellular space → tells cells around that cell is going through massive restructuring
describe the rationale behind the method the authors used in this paper
necause autophagy seems to increase oncolytic adenovirus, and promote immunogenic cell death, the authors will further augment autophagy using pyranose oxidase (P2O)
how does pyranose oxidase work?
pyranose oxidase is made by lots of fungi and catalyzes glucose, releasing H2O2 (hydrogen peroxide), which stresses the cell and activates autophagy
enzymatic P2O reaction and ROS production
H2O2 as stress signal → H2O2 is an ROS that induces stress to activate autophagy
what are the two methods one could use to deliver P2O and adenovirus to the same tumour cells
genetically encode P2O in recombinant Adenovirus → add P2O to genome
Co-enveloped Adenovirus and P2O in lipid particle → co-package
what are OMVs?
Bacterial Outer Membrane Vesicles → G- bacteria like E. coli naturally make OMVs for a variety of reasons:
sequester phage, antibiotics
gene transfer
deliver toxins
deliver enzymes
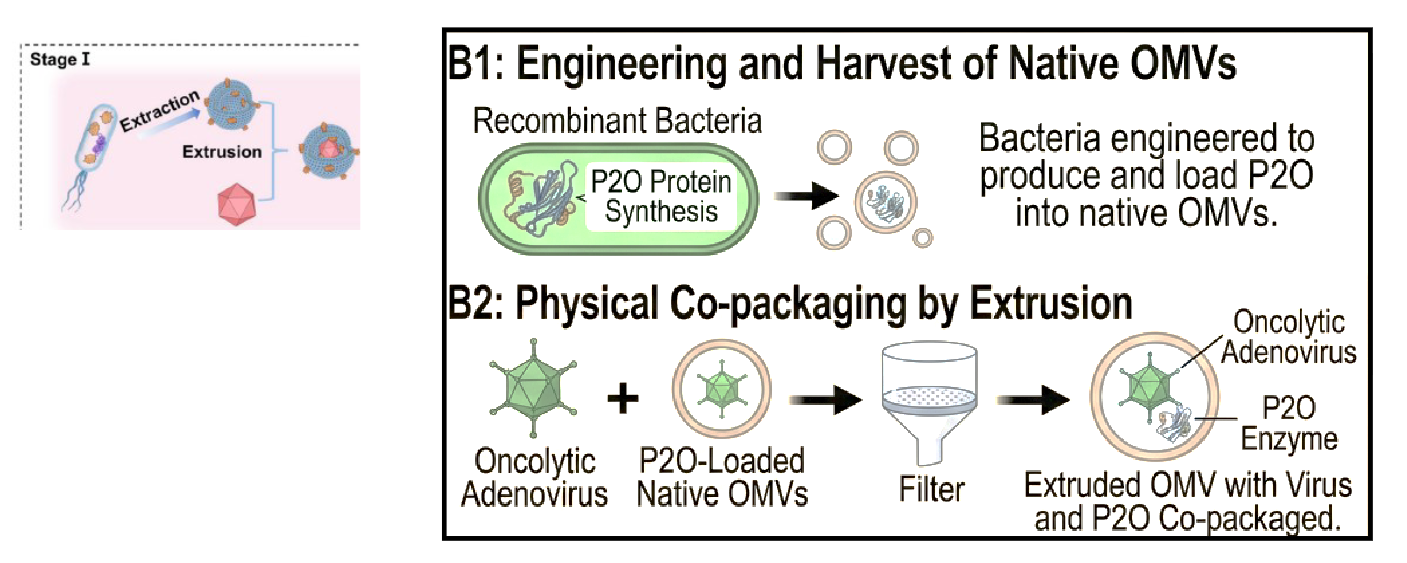
describe the rationale behind the treatment the authors produced?
will use OMVs from P2O-expressing bacteria - and extrude with Ad - to make them a ‘single package’
engineer and harvet native OMVs → recombinant bacteria engineered to produce and load P2O into native OMVs
physical co-packaging by extrusion → oncolytic Ad + P2O-loaded native OMVs → squeeze through filter, when come back together, Ad and P2O Co-packaged in OMV
how did the authors shield the OVMs from the immune system?
OMVs have foreign proteins that could lead to their clearance
So, to shield the OMVs, they were placed in a solution rich in calcium and phosphate ions.
The proteins and lipids on the OMV surface attract these ions, causing them to crystallize directly onto the vesicle.
Since calcium phosphate (CaP) is chemically similar to hydroxyapatite (the primary mineral in human bone and teeth), they are not recognized as foreign by the immune system
CaP is pH sensitive and degrades in the low pH endosome/lysosome
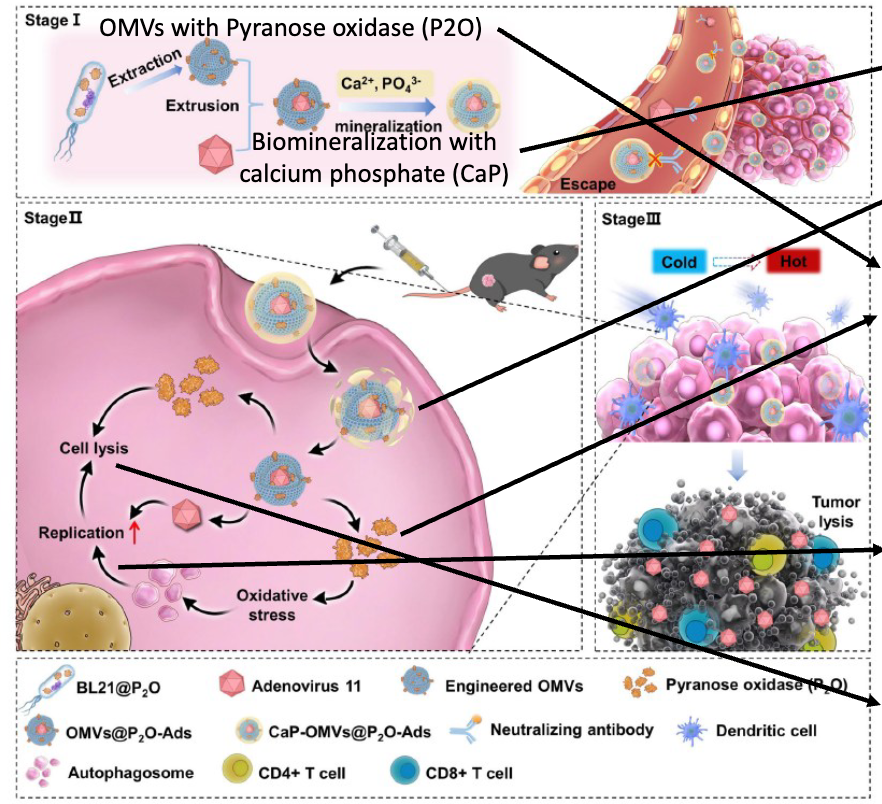
summarize the steps of how this oncolytic would work
Camouflaged ! CaP shell mimics natural mammal substance - protects from immune clearance – for example by hiding foreign epitopes
CaP is pH sensitive and degrades intracellularly
P2O converts glucose to H202 → ROS = reactive oxygen species, promotes autophagy
Autophagosomes promote Adenovirus replication: Increase AA etc via recycling, Increase virus protein synthesis, Provide platform for replication & Assembly
Autophagy promotes immunogenic cell death → Release of DAMPS & Tumor associated antigens
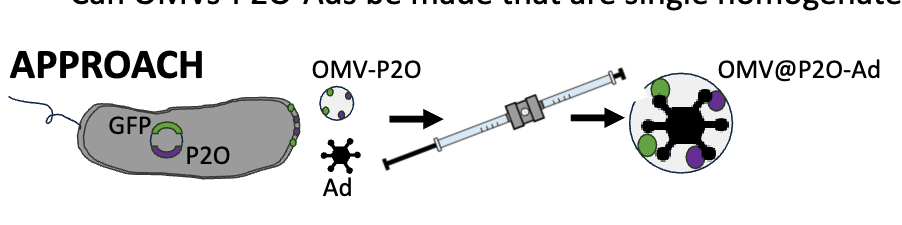
how did they test if OMVs-P2O-Ads be mde that are single homogenates with As inside OMV?
co-express GFP & P2O on same plasmid → perform extrusion on OMV-P2O-GFP with Ad → should have OMV-P2O-Ad
use confocal laser microscopy to look fro DNA (Ad), and GFP (OMVs)
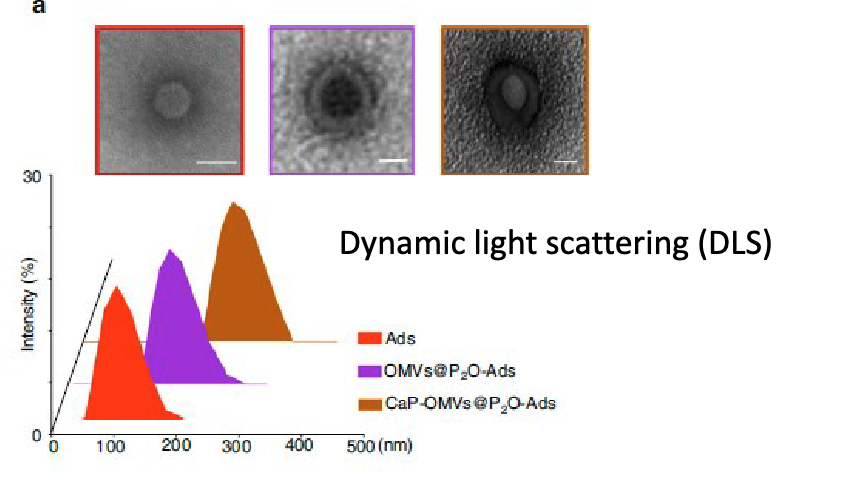
use TEM
dynamic light scattering to see size
describe how they checked if OMVs-P2O-Ads resulted in increased H2O2?
take Tc1 lung cancer cells + OMV-P2O-Ads or OMV-Ads → various assays to meaure H2O2/ROS, LCI/LCII, virus replication, and cell lysis
to measure ROS, used flow cytometry & DCFH-DA:
1. DCFHA-DA starts as non-fluorescent and is taken up by cells. Cellular esterases will convert it to non-fluorescent DCFH, which trap the probe inside the ce
2. ROX interact with DCFH to generate DCF, which is highly fluorescent → more ROS = more fluorescence
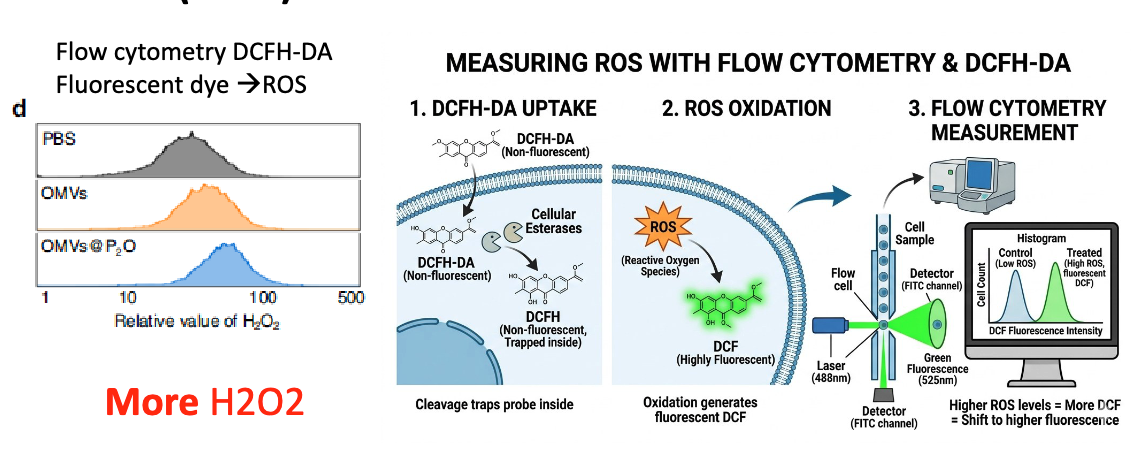
describe how they checked if OMVs-P2O-Ads resulted in increased H2O2?
take Tc1 lung cancer cells + OMV-P2O-Ads or OMV-Ads → various assays to measure H2O2/ROS, LCI/LCII, virus replication, and cell lysis
performed westen blot looking at ratio of LC3-II/LC3-I → more LC3II = more autophagy
also used confocal microscopy → used ethidium bromide to dye DNA, and MDC dye to stain autophagosome (selectively accumulates via lipid and ion interactions)
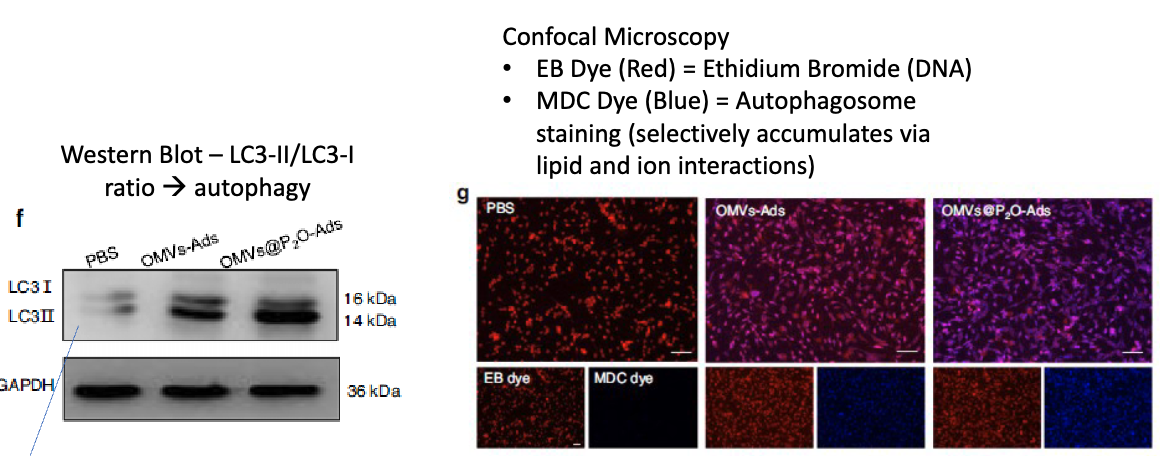
describe how they checked if OMVs-P2O-Ads resulted in increased Ad replication?
take Tc1 lung cancer cells + OMV-P2O-Ads or OMV-Ads → various assays to meaure H2O2/ROS, LCI/LCII, virus replication, and cell lysis
checked for [Ad DNA] by qPCR → replication
found more adenovirus rep with P2O-Ad
As control, also had group with P2O-Ad + 3MA
![<p>take Tc1 lung cancer cells + OMV-P2O-Ads or OMV-Ads → various assays to meaure H2O2/ROS, LCI/LCII, virus replication, and cell lysis </p><ul><li><p>checked for [Ad DNA] by qPCR → replication </p></li><li><p>found more adenovirus rep with P2O-Ad</p></li><li><p>As control, also had group with P2O-Ad + 3MA</p></li></ul><p></p>](https://assets.knowt.com/user-attachments/461306b8-1ad1-4478-a1c0-8bf71ad1b653.png)
describe how they checked if OMVs-P2O-Ads resulted in increased cell lysis?
take Tc1 lung cancer cells + OMV-P2O-Ads or OMV-Ads → various assays to meaure H2O2/ROS, LCI/LCII, virus replication, and cell lysis
used confocal microscopy:
Calcein (green) = living cells → intracellular esterase converts to fluorescent mol
Propidium Iodide (PI) = dead cells → enters permeabilized cells and binds nuclei
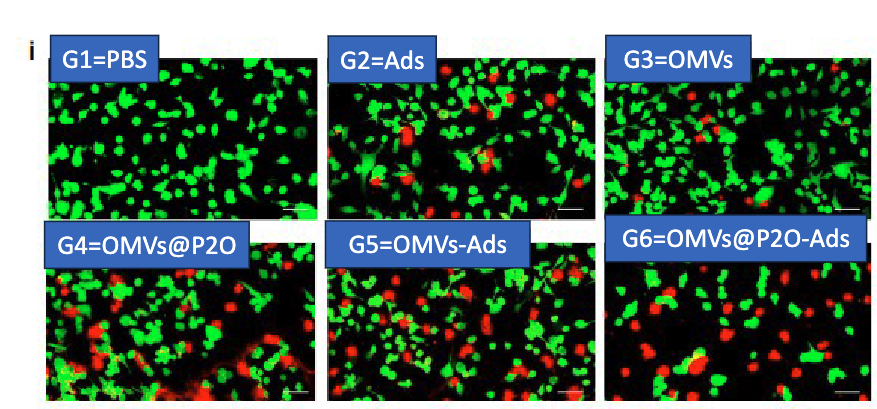
describe how they tested if OMV-P2O-Ads exhibit increased oncolytic activity in vivo?
inoculated mice with TC-1 lung cancer cells 7 days prior to treatment → gave intratumoral treatment every 3 days for 4 treatments → at 18 days collect tumor for analysis
looked at tumor volume
did ELISA on mouse serum and looked for TNFa, IL-6, and IFNg
found OMVs-P2O-Ads had best increase in cytokines
how did they exxamine what genes alter expression in the tumor after OMVs=-P2O-Ads
transcriptomic analysis: Illumina RNAseq
competing tech: nanopore sequencing → PORE_forming proteins from bacteria + tethering
found increased immune response genes
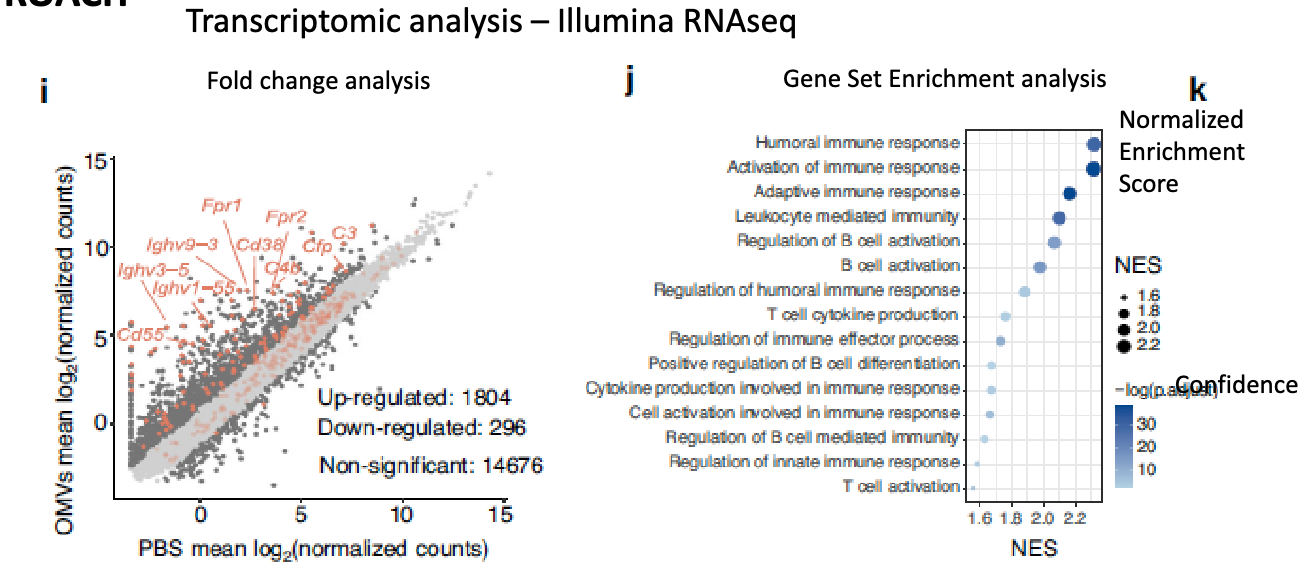
how did they look at whether the OMVs-P2O-Ads could be further calcium biomineralized?
transmission electron microscopy
dynamic light scattering (DLS(
found the CaP mineralization produces larger homogenous particles
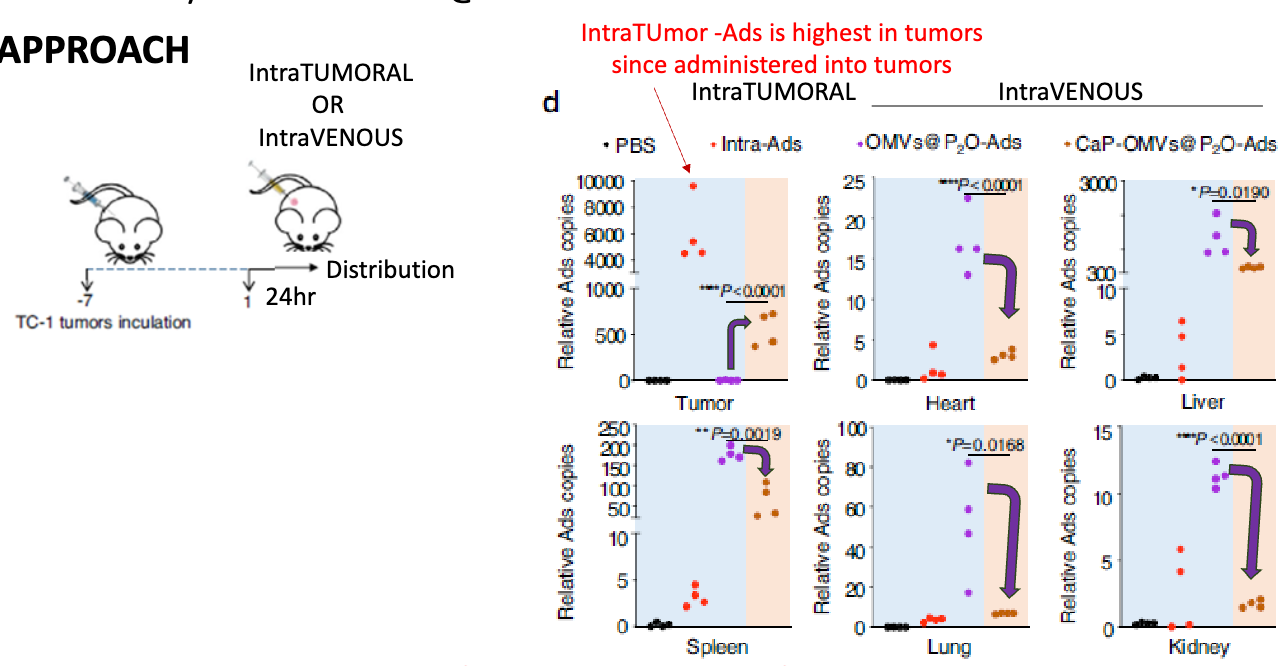
how did they look at how adding CaP to OMVs-P2O-Ads affects biodistribution
inoculated mice with TC-1 tumours for 7 days, then inoculated with either PBS, Intra-tumoral Ad, OMV-P2O-Ad, CaP-OMVs-P2O-Ad, examine biodistribution after 24h → looked ay relative Ad genome copy number in tumor, heart, liver, speen, lung, and kidney
found that adding CaP increases tumor-specific accumulation of IV OMV-Ads
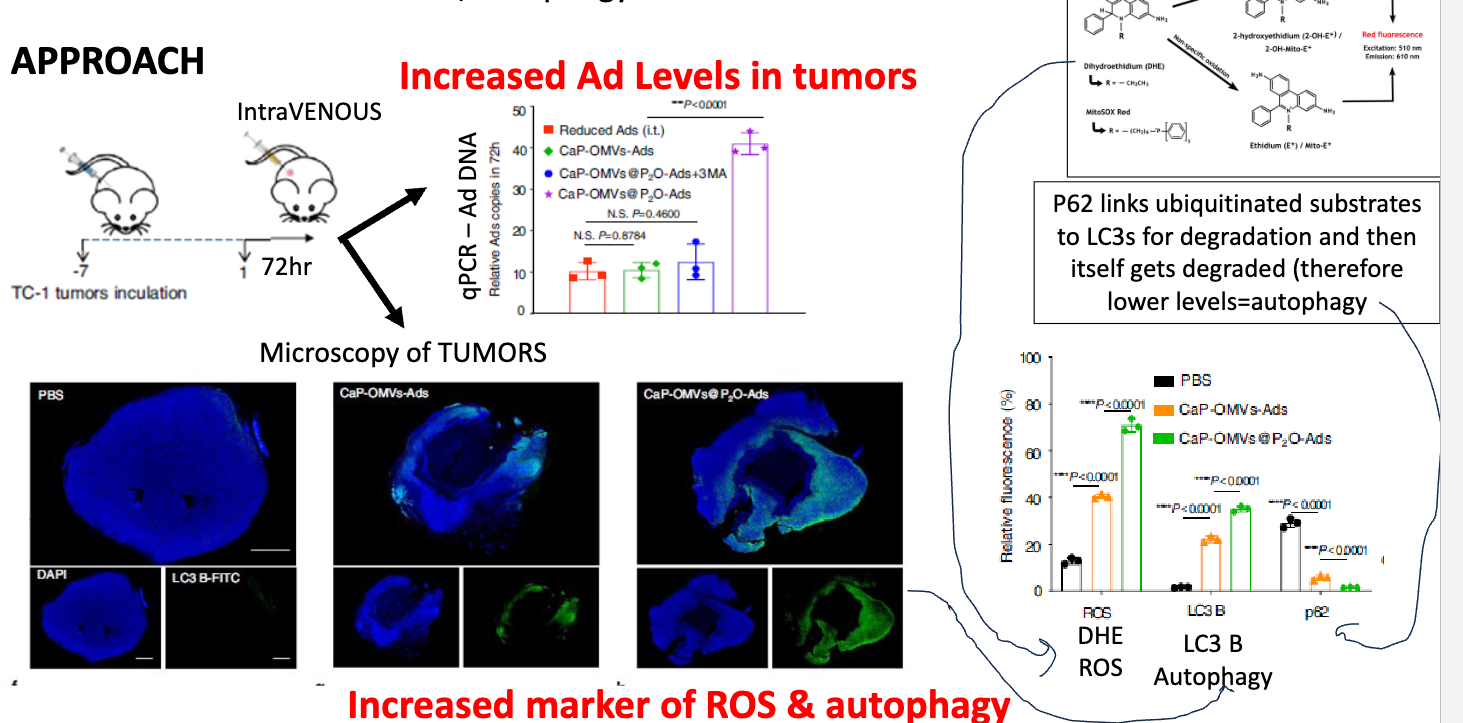
how did they test if ± CaP and ± P2O onto OMV-Ads affect ROS, Autophagy, and Ad levels?
inoculate with TC-1 tumors for 7 days → inject construct, perform analysis 72h later
using qPCR, detected higher levels of Ad-DNA when treated with Ca-OMVs-P2O-Ads
performed microscopy of tumoura → found increased markers of ROS and autophagy
p62 linked ubiquitinated substrates to LC3s for degradation and then itself gets degraded → therefore lower levels = autophagy → p63 used as measure of autophagy
how did they examine what the oncolytic activity of OMVs-P2O-Ads was in vivo on primary tumor?
inoculated mouse with TC-1 tumour for 7 days, then given injection of either PBS, IV-OMV-P2O-Ads, IV-CaP-OMVs-Ads, IT-Ad, IV-CaP-OMVs-P2O-Ad, IT-Ads (high dose) every 3 days for 3 doses → at day 15 mice were dissected, flow cytometry performed & tumor volume analyzed
found mix had advantage → another benefit, given IV not IT
flow siggested that TME is becoming less immunosuppressive → but doesn’t show it’s anti-tumour
of IV OMVs, CaP and especially CaP-P2O improve over PBS, but not notable advantage over IT-Ads
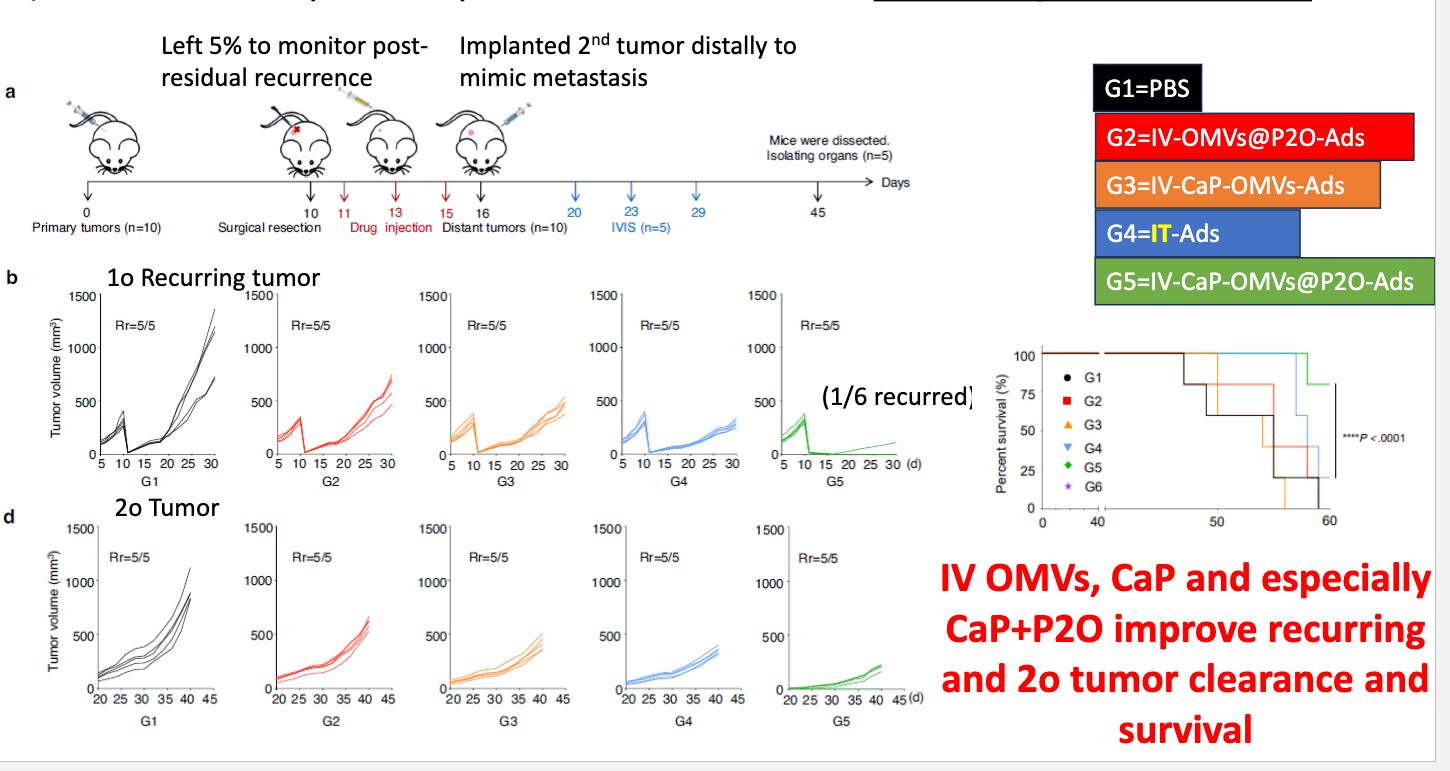
how did they determine the oncolytic activity of OMVs-P2O-Ads on recurring tumor and secondary site?
inoculated with tumor → surgically removed 95%, leaving 5% on day 10 → drug injection day 11, 13, 25 → implant 2nd tumor distally to mimic metastasis → IVIS day 20, 23, 29 → mice dissected day 45
found IV OMVs, CaP and especially Ca+P2O improve recurring and 2o tumor clearance and survival