Quiz 4 Content
1/67
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
68 Terms
Main differences between DNA and RNA
2’ -OH
Uracil instead of Thymine
RNA can fold up to look like a helix
single stranded vs. double stranded
monocistronic
codes for only 1 polypeptide
in eukaryotes and bacteria
polycistronic
codes for 2+ different polypeptides
in bacteria and archaea
transcriptome
the sum of all the RNA molecules produced in a cell under a given set of conditions
mRNA
encodes the amino aicd sequences of polypeptides
tRNA
read the mRNA and transfer the appropriate AA to a growing polypeptide chain during protein synthesis
rRNA
constituents of ribosomes, the cellular machines that synthesize proteins
ncRNAs
have a variety of catalytic structural, and regulatory functions
RNA polymerase
catalyzes transcription
requires a DNA template
add ribonucleotide units to the 3’ -OH end
has 2 Mg2+ groups just like DNA polymerase where a pyrophosphate leaves
overall RNA polymerase rxn
(NMP)n + NTP → (NMP)n+1 + PPi
template strand
DNA strand that serves as template or RNA synthesis
Nontemplate strand
aka coding strand
DNA strand that is identical in base sequence to the transcribed RNA, with U in RNA in place of T in DNA
promoter sequence
directs RNA polymerase to a specific chromosomal location
-35 region → - 10 region (TATAAT)
sigma factor
recognizes the promoter sequence
sigma70 RNA polymerase
binds between -70 — +30
RNA polymerase + supercoils
a transcription bubble forms when the DNA duplex unwinds
RNA polymerase generates positive supercoils ahead and negative supercoils behind
DNA can twist but cannot swivel, besides when topoisomerase is used
RNA can exite through an RNA exit
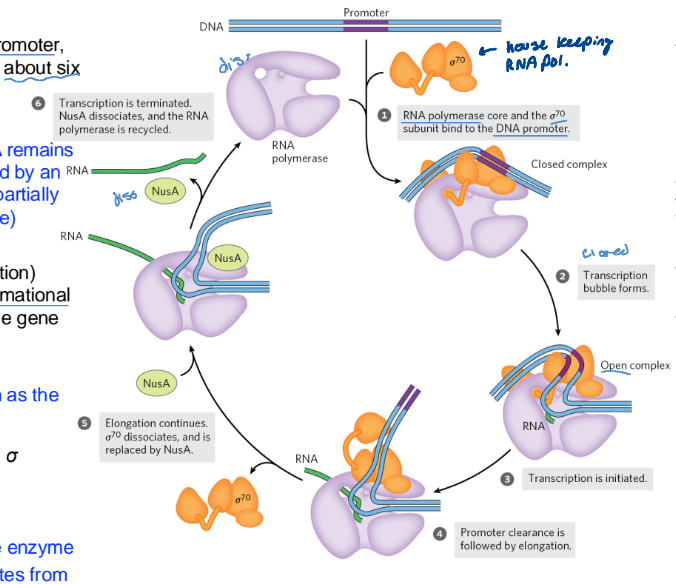
Initiation + Elongation by E. coli RNA Polymerase (6 steps)
RNA polymerase binds to the promoter, directed by a sigma factor
Closed complex (bound DNA remails double-stranded) forms, followed by an open complex (bound DNA is partially unwound near the -10 seq.)
Initiation causes RNA polymerase conformational change that moves it towards the gene
promoter clearance follows by elongation
NusA protein competes with the sigma subunit to bind the polymerase. Leads to sigma 70 dissociating
Transcription is terminated
NusA dissociates from the enzyme
RNA polymerase dissociates from the DNA
RNA polymerase is recycled
σ70 RpoD
housekeeping sigma factor
σ32 aka RpoH
stress-induced promotor from heat shock
has different -10 — -35 region
σ54 RpoN
specific for the promoters that control nitrgen assimilation
Importance of RNA polymerase orientation
orientation is dictated by the promoter sequence
anti-sense strand = template strand or nontranscribed strand
sense strand = transcribed strand
three main things that control protein activity
modification/ligand binding
subcellular localization
synthesis and degradation
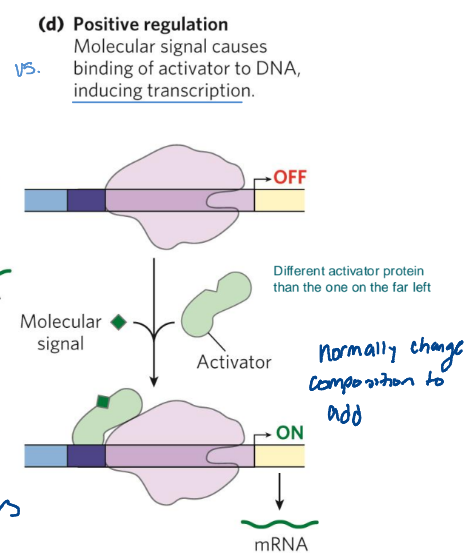
Positive induction
molecular signal causes binding of activator to DNA, inducing transcription
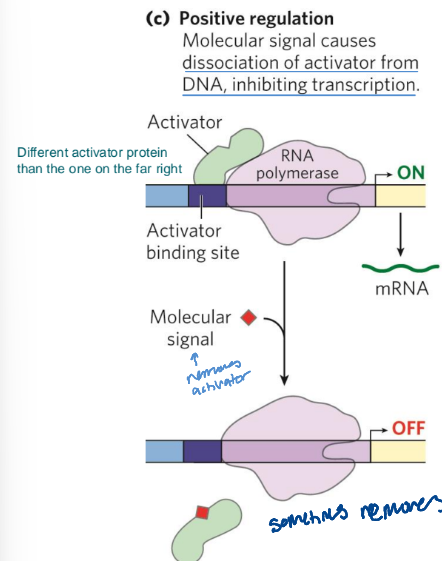
positive repression
molecular signal causes dissociation of activator from DNA, inhibiting transcription
negative induction
Molecular signal causes dissociation of repressor from DNA, inducing transcription
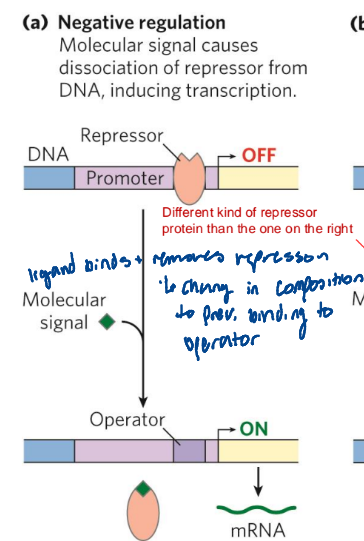
negative repression
molecular signal causes binding of repressor to DNA, inhibiting transcription
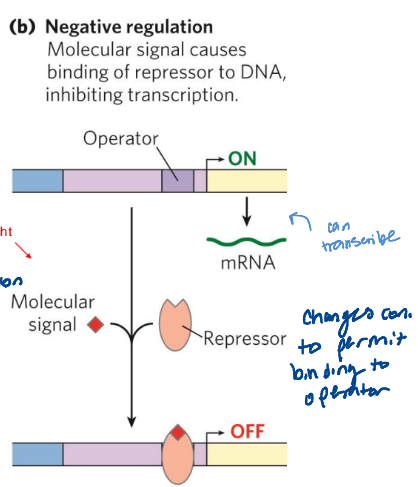
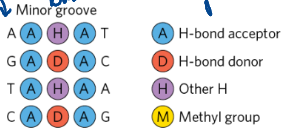
Functional groups - major groove binding specificity
make a code protein that can recognize w/ specificity
A: H-bond acceptor
D: H-bond donor
H: Other H
M: Methyl group
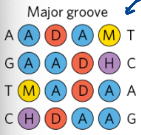
Functional groups - minor groove binding specificity
3 in the minor groove vs. the 4 in the major
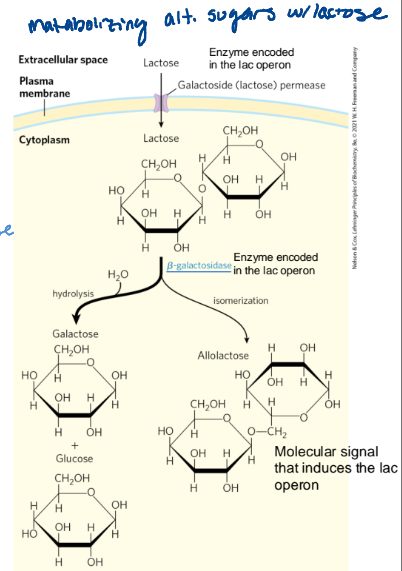
Lac operon in E. coli
Lactose is used as an energy source. Uses beta-galactosidase to break it down (encoded by the lac operon)
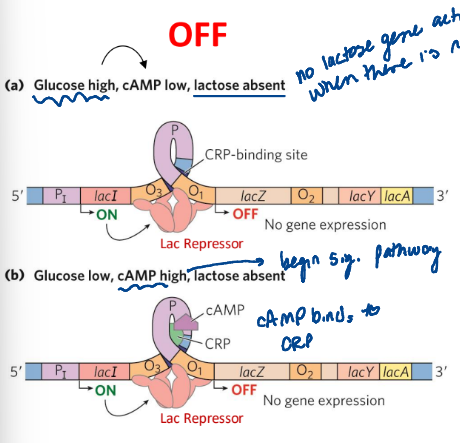
negative regulation by Lac repressor
a. glucose high, cAMP low, lactose absent
b. glycose low, cAMP high, lactose absent
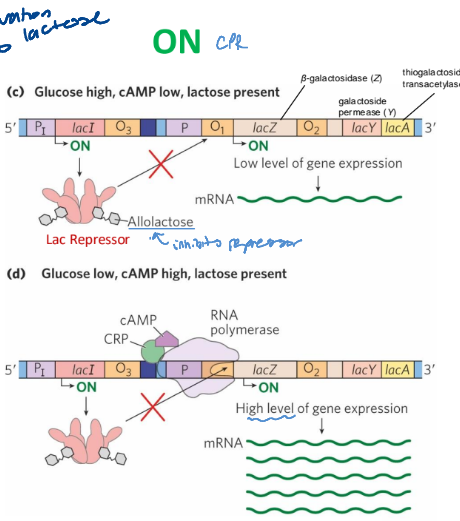
positive regulation by CPR
a. glucose high, cAMP low, lactose present
b. Glucose low, cAMP high, lactose present
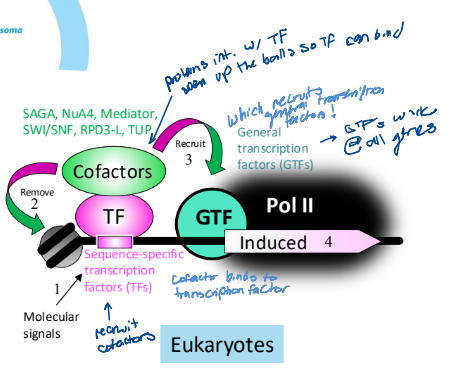
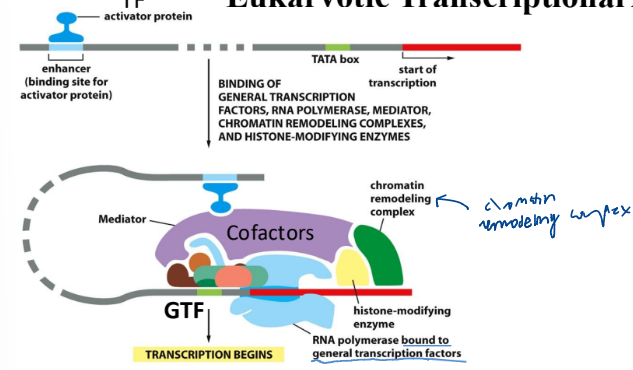
Cofactors role in regulation in Eukaryotes
interact with the transcription factor to remove the nucleosome
recruits GTFs
Eukaryotic Homeodomain
eukaryotic transcriptional regulators
play a special role during development
Zinc Finger
Zn2+ is coordinated to 4 Cys and/or His
feels the major groove out to help with binding
on its own it is weak, but many zinc fingers are substantially enhancing binding
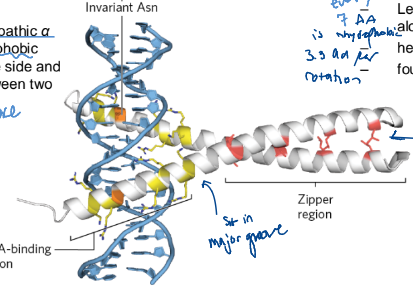
Leucine Zippers
an amphipathic alpha helix with a series of hydrophobic amino acid residues on 1 side and a hydrophobic surface between 2 polypeptides of a dimer
Leu @ every 7th position
about 1 every 2 turns (3.3 residues per rotation)
RNA polymerase I (eukaryotes)
rRNA
consitiuents of ribosomes, the cellular machines that synthesize proteins
RNA polymerase II (eukaryotes)
mRNA
encode the amino aicd sequence of polypeptides
ncRNA
variety of catalytic structures and regulatory functions
RNA polymerase III (eukaryotes)
tRNA
reads the mRNA and transfer the appropriate AA to a growing polypeptide chain during protein synthesis
5s rRNA
ncRNA
Chloroplast + Mitochondria’s DNA polymerase
have their own RNA polymerase
evidence to support the endosymbiotic theory
all RNA polymerases have a common ancestor!
TATA binding protein (TBP)
recognize eukaryotic promoters
the first component to bind to the preinitiation complex (PIC) at the TATA box of a typical Pol II promoter
used in Pol I II and III as well as archaea!
similar to the role of sigma factors in bacteria
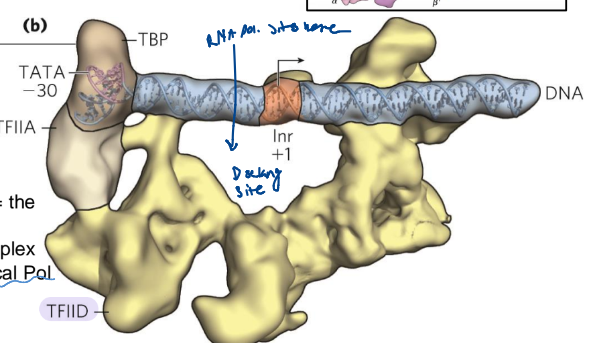
TFIID Role
positions the TBP and Pol II on the promoter
binds the promoter DNA where the TBP subunits are anchored in the DNA minor groove
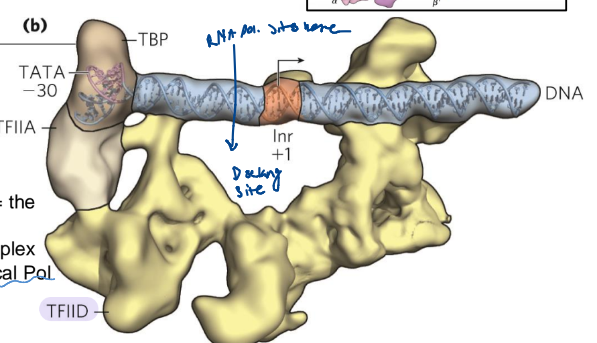
TATA box promoter element
typically located at -30
TATA(A/T)A(A/T)(A/G) sequence
recognized by the TATA binding protein (TBP)
not very common
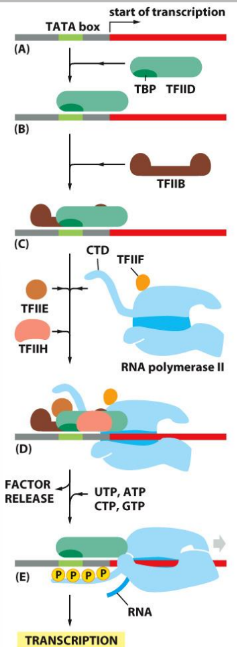
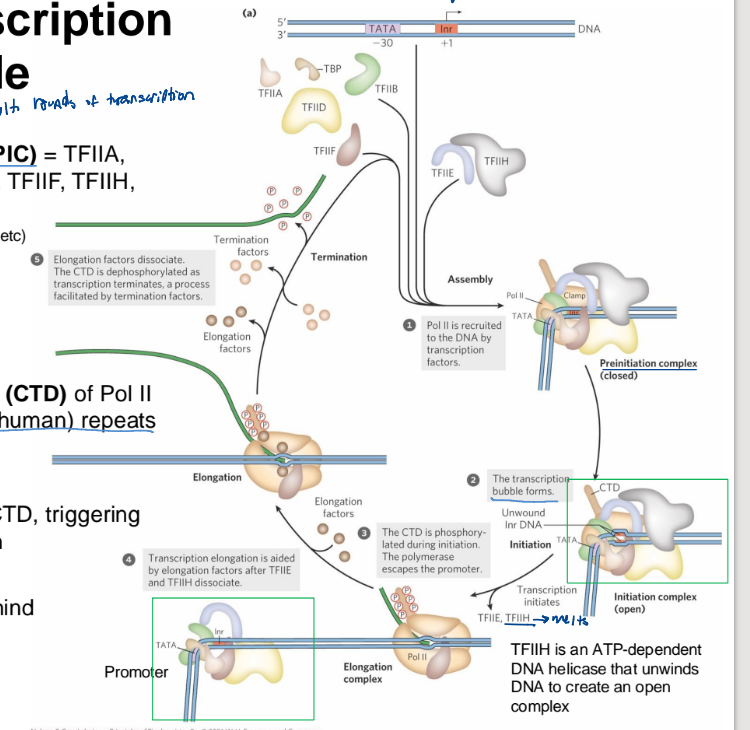
Assembly of eukaryotic transcription RNA polymerase II initiation complexes (w/ general transcription factors!)
TFIID recognizes the promoter sequence (with help form TFIIA)
TFIIB, TFIIE, TFIIF, and TFIIH bring Pol II to the promoter, and initiate DNA melting
TFIIH uses ATP energy to ram the DNA into polymerase which melts the site of trancription initiation
TFIIH also phosphorylates RNA polymerase
CTD - carboxy-terminal domain
aka repeat central
The largest subunit of RNA polymerase II has a repeating AA sequence at the carboxy-terminal end
for human RNA polymerase II, a sequence of 7 AA is repeated 52 times
thought to help tether Pol II to the general transcription factors

TFIIH
helicase-like activity
starts initiation by phosphorylating the CTD of RNA polymerase II, promoting melting
because it phosphorylates, it is called a kinase
TFIIH is also a kinase
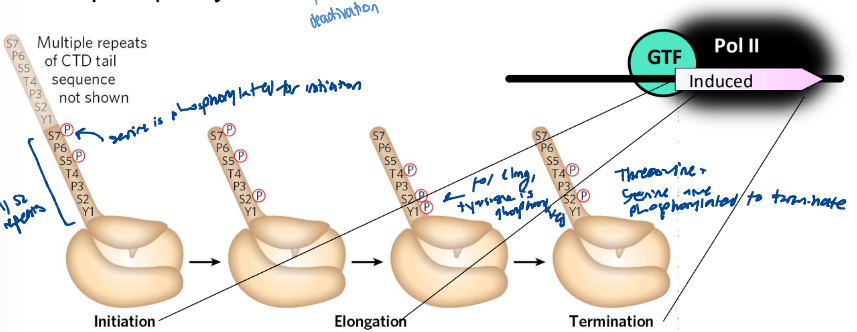
Phosphorylation of the CTD in RNA Pol II
the phosphorylation state changes during trancription
when Pol II dissociates, the gene is dephosphorylated
Eukaryotic Transcriptional Activates and Enhancers
can be very far upstream
bind to the cofactors to help enhance gene expression
Pol II transcription cycle
Pre-Initiation Complex (PIC): TFIIA, TFIIB, TFIID (TBP), TFIIE, TFIIF, TFIIH, and Pol II
the CTD - carboxyl-terminal domain of Pol II consists of 26 yeasts to 52 human repeats of -YSPTSPS- AAs
TFIIH phosphorylates the CTD to trigger Pol II to initiate transcription
Pol II is recruited to the DNA by the GTP, TFIID finds the promoter site and TFIIH directly binds it.
transcription bubble forms (w/ TFIIH which phosphorylates the CTD to melt it further")
CTD is further phosphorylated during initiation
Transcription elongation is aided by elongation factors after TFIIE and TFIIH dissociate
Elongation factors dissociate. The CTD is dephosphorylated as transcription terminates a process facilitated by termination factors.
Compare + Contrast promoters across the 3 domains of life
bacteria:
-35 to TAAT box at -10
Sigma factors are very common
DNA lacks nucleosomes
Eukaryotes:
TATA box at -30 while initiation begins at +1
TATA boxes are uncommon
Archaea:
Have regulatory factors that are Bacterial-like
have initiation factors and an RNA polymerase that is Eukaryotic-like
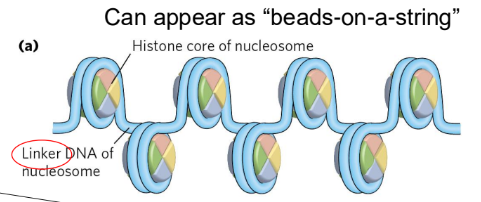
Nucleosomes
fundamental organizational units of chromatin
contain 8 core histone molecules: 2 of H2A, H2B, H3, H4
150 bp are bound to nucleosomes
left-handed coil
Histones
Small, Basic Proteins
make up a nucleosome
H2A, H2B, and H3 are in an evolutionarily related gene family
H1 binds to linkers and is repressive
binds to the nucleosome exterior near the linkers
H2A
includes H2A.Z, H2A.X, macroH2A and more
in eukaryotes
H2A.Z is in the first nucleosome of a Pol II transcription unit
phosphorylated H2A.X exists primarily at sites of DNA breaks (help in DNA repair)
macroH2A is involved in transcriptional repression
H2B
also includes H2B.1
is in eukaryotes
H3
also includes H3.2, and H3.3
in both archaea and eukaryotes
H3.3 is enriched near transcriptionally active promoters
H4
no other family members
in archaea and eukaryotes
Histone Amino-Terminal Tails (Epigenetics!)
intrinsically disordered
where most of the histone modification occurs
amino-terminal tails likely interact with one another
four main types:
acetylation (ac)
methylation (me)
ubiquitylation (ub)
phosphorylation (ph)
acetylation (ac)
K
H3K9ac, H3K27ac, H4K18ac
function is opening chromatin for transcription
methylation (me)
K and R
H3K4me1: enhancer mark
H3K4me3: first few nucleosomes of a transcribed gene
H3K36me3: marks nucleosomes in genes downstream of H3K4me3
H3K9me3: repressive mark of heterochromatin
H3K27me3: dynamic repressive mark
Ubiquitylation (ub)
K
H2BK123ub: helps return nucleosomes to gene bodies during transcription
H2AK119ub: repressive mark
Phosphorylation (ph)
S and T
H3S10ph: is involved in chromatin condensation
Chromosome territories
subnuclear region that constrains the entire structure of each chromosome
little or no intermingling of DNA in different territories
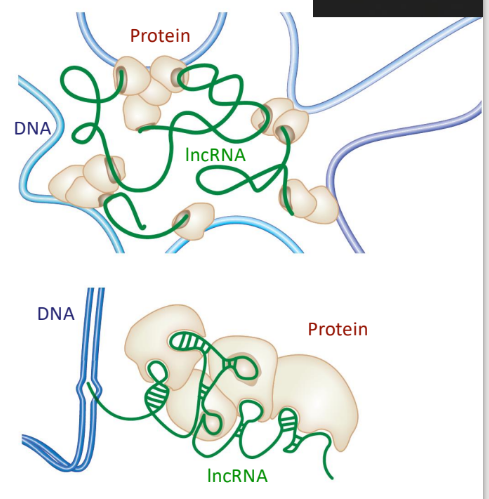
Long noncoding RNAs (lncRNAs)
play a functional role in defining the chromosome structure
many provide a scaffold for proteins
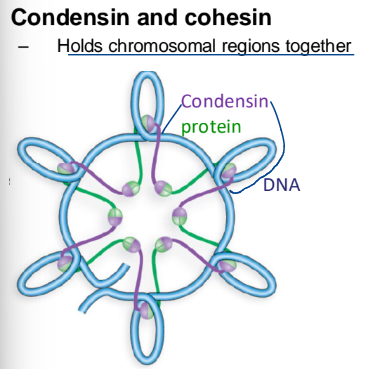
Condensin and cohesion
holds chromosomal regions together
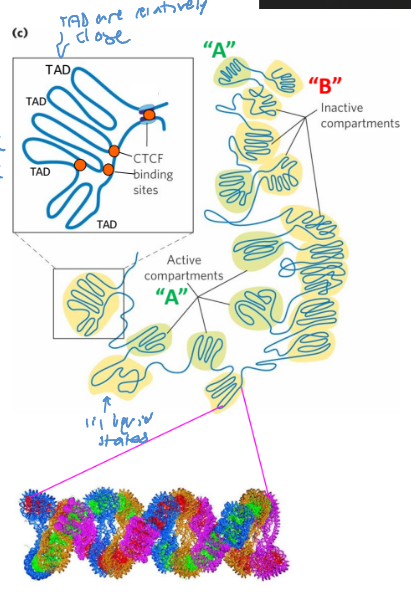
Active + inactive compartments
Active (“A”) - compartments withh reduced chromatin condensation (active)
Inactive (“B”) - compartments (heterochromatin) are highly condensed
deactivating and repressive
H3K27 methylation could occur here
H3K9 methylation for permanent storage
Topologically associated domains (TADs)
large segments of DNA are organized in loops
binding of CTCF, cohension, and topoisomerase II to bordering sites brings the DNA into these loops
The TADs are relatively close
Epigenome
regulates information flow
backregulation
includes nucleosomes with their modifications
Acetylation, methylation, phosphorylation, ubiquitylation
Is a stable type where it is passed down through generations through inheritance.
Hereditary material
instructs its own replication
stably passed on to its progeny
through readers and writers the instructs are passed to daughter nucleosomes
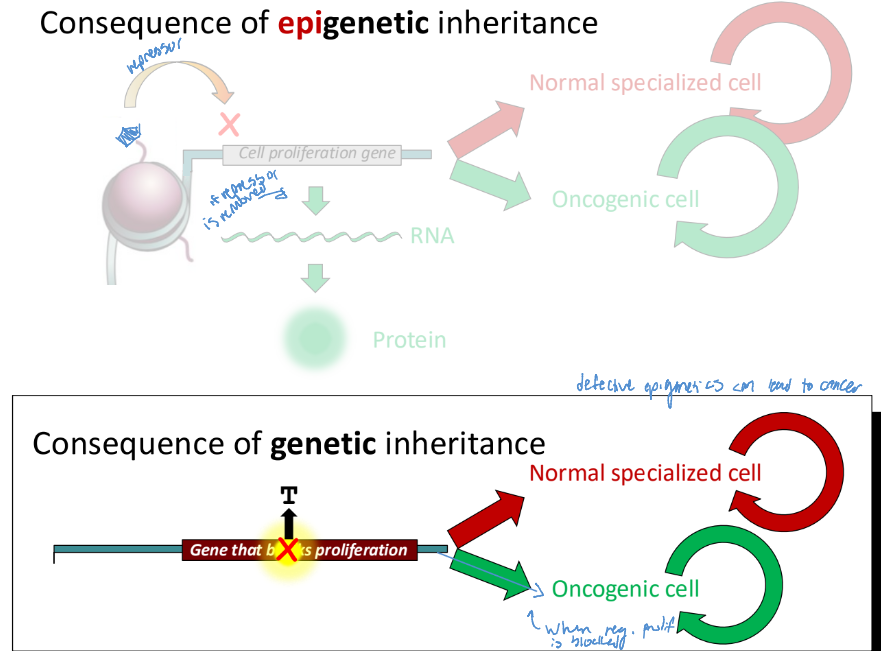
Consequence of epigenetic inheritance vs. genetic inheritance (FIX)
a loss of a nucleosome edit, such as methylation, can lead to an oncogenic cell
Defective epigenetics can lead to cancer
When regulation is blocked, an oncogenic cell is made (which can become cancerous)
however, the epigenome isi dymanic and there can be dynamic changes w/o inheritance that are stable