BB 492: Midterm 2
1/65
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
66 Terms
What is a Holliday Junction? How is it formed
Four-Way Crossover Point between double stranded DNA arms
two of the four strands exchange information here
Formation steps:
Initiation (Nicking): The process begins with single-stranded nicks at corresponding locations on two homologous DNA duplexes.
Strand Invasion: The broken strands leave their original complementary strand and pair with the opposite duplex, a process often guided by specific proteins.
Ligation and Junction Formation: The invading strands are ligated, creating a cross-shaped structure that links the two duplexes.
Branch Migration: This junction can move up or down the DNA sequence, increasing the region of heteroduplex DNA.
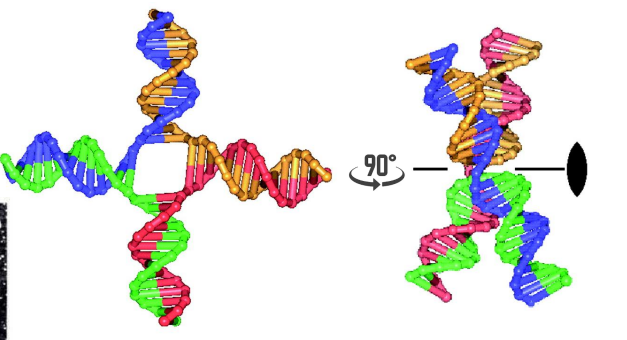
Nonrecombinant/ Patch Recombinant duplex
AB and ab
cleave the strands that crossed over resulting in exchange of a pair of homologous single stranded segments (non-recombinant but heteroduplex regions)
left side of image
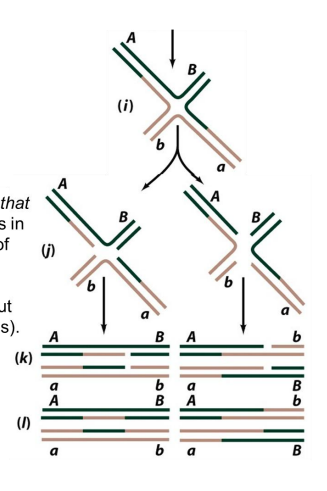
splice recombinant heteroduplex
Ab and aB
when you cleave the strands that didn’t crossover yet → recombinant heteroduplex after sealing the nick
right side of image
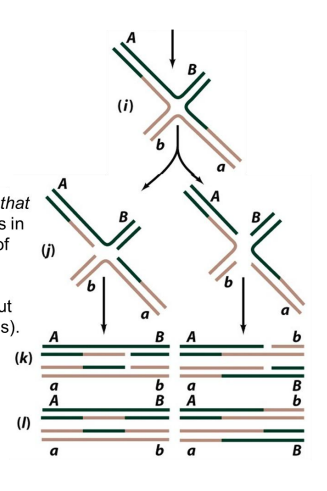
How is a Holliday junction separated?
Through the process of resolution:
specialized endonuclease enzymes (resolvases) catalyze symmetrical nicks in 2/4 DNA strands
cleavage occurs at horizontal or vertical axis
Resolution generates two double-stranded DNA products that are either patched (non-crossover) or spliced (crossover)
Steps for Using Homolgous Recombination to Repair DNA Damage
“Tailoring the ends of the DNA breaks to create a 3’ overhand of single-stranded DNA
Holliday Junction(s) formation and strand invasion
Branch migration (and patch DNA polymerization)
Resolving the Holliday Junctions
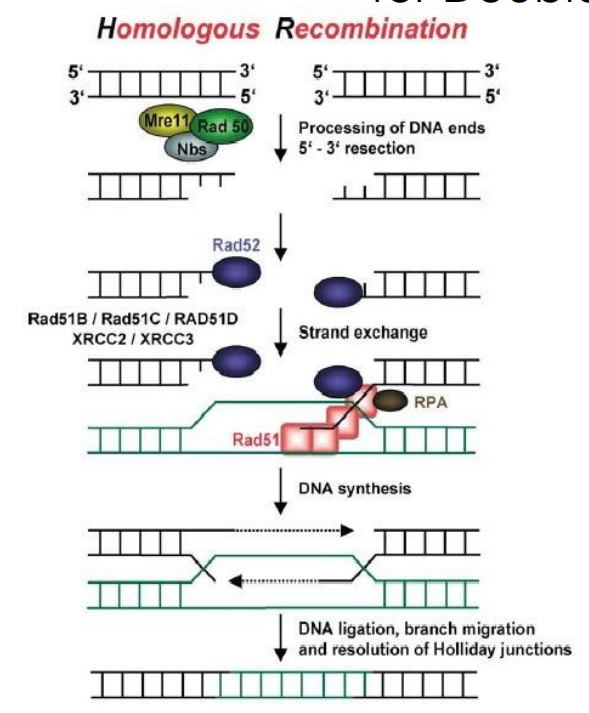
What sort of damaging agents may cause DNA double strand breaks?
Exogenous Agents (e.g. ionizing radiation, chemotherapeutic drugs, UV light) and endogenous factors (e.g. reactive oxygen species, replication stress)
What proteins in prokaryotes are important in the repair process by homologous recombination?
Recombinase proteins (Rec’s) identify broken ends, remove bases at the damage site, and create a platform for strand invasion
Rec BCD Complex: initiates recombination by editing DNA to create a 3’ single-stranded overhang
RecA: Binds ssDNA & forms a nucleoprotein filament that allows DNA strand invasion and recombination
What does the RecBCD complex do?
Initiates Recombination!
The complex binds double stranded DNA, but DNA leaves the complex as singled stranded (one is a single-stranded 3' overhang coated with repair proteins)
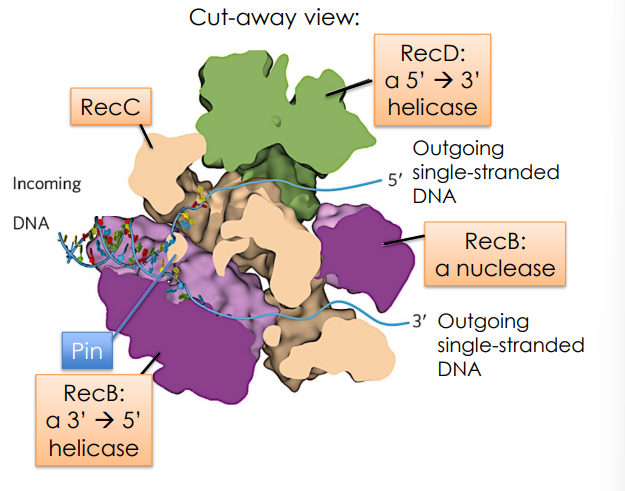
RecC
has a protrusion called “the pin” which facilitates strand separation
part of RecBCD complex
RecD
a 5’→3’ helicase
part of RecBCD complex
RecB
a 3’→5’ helicase
a nuclease
will degrade both strands but cleaves the 5’ end strand more than the other
part of RecBCD complex
How is the 3’ overhand created
RecBCD unwinds and degraded DNA UNTIL
encounters “Chi” (5’-GCTGGTGG-3’)
RecC binds tightly on Chi sequence, stopping degradation on 3’ strand. (Note: nuclease activity continues on the 5’ strand)
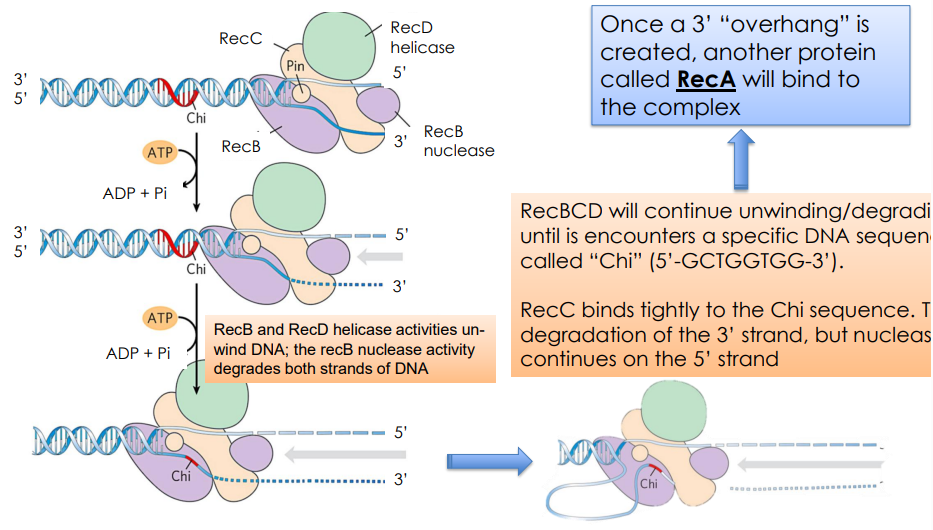
What does RecA do?
Binds ssDNA & forms a nucleoprotein filament that allows DNA strand invasion and recombination
What must be created at the double-stranded break before repair can occur?
3′ single-stranded DNA (ssDNA) overhangs
What happens after the overhang is made?
After overhang is made, RecBCD recruits RecA
two RecA proteins bind, displacing SSBs
when ssDNA is made, SSB rushes in to coat it
HOWEVER SSB get in the way of the repair process
RecBCD makes sure two RecA proteins can bind to the ssDNA, so more RecA can follow
RecA associates with the phosphodiester backbone of the 3’ DNA single strand
NOTE: ATP must bind to RecA in order to make it competent to bind DNA
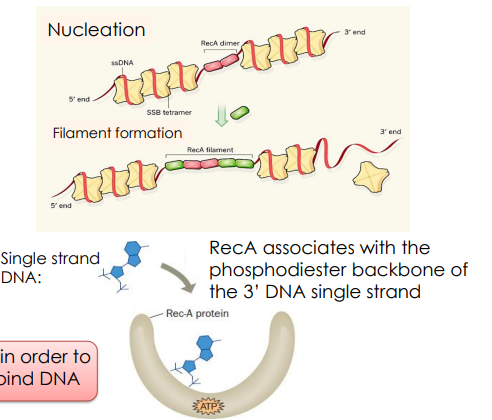
Describe how RecA creates a filament with DNA.
Nucleation: small number of RecA monomers bind to a segment of ssDNA
Polymerization: after nucleated, more RecA monomers are rapidly added
it does so without regard to base sequence— RecA is a generalist loader—it will "zip up" onto any available single-stranded DNA regardless of what genetic information that DNA is carrying.
DNA Stretching: Ultimately, RecA forms a nucleoprotein filament with extended dimensions: 18 nucleotides/ filament turn
What does RecA need to make it competent to bind DNA?
ATP must bind to RecA in order to make it competent to bind DNA
What type of binding sequeunces does RecA have?
1 sequence recognizes single stranded DNA
1 sequence recognizes double stranded DNA
How many strands of DNA can a RecA filament accommodate at one time?
three!
RecA single strand DNA polymer looks for a double stranded DNA segment with similar homology to parts of 3’ DNA overhang
dsDNA binds to the second site
duplex DNA is scanned by RecA filamet until homologous segment is found
ssDNA filaments begin to base pair w/ complementary strand from duplex DNA
What process is catalyzed by the RecA filament? Describe this process and what is formed?
strand invasion
RecA forms a filament on ssDNA
homologous dsDNA segment incorporates into the complex
invasion of the single strrand begins at 5’ end
rotation of complex spools homologous DNA out of dsDNA, spools in ssDNA
Strand exchange and continued branch migration require ATP hydrolysis
if ssDNA and dsDNA are linear molecule, a new ssDNA is released
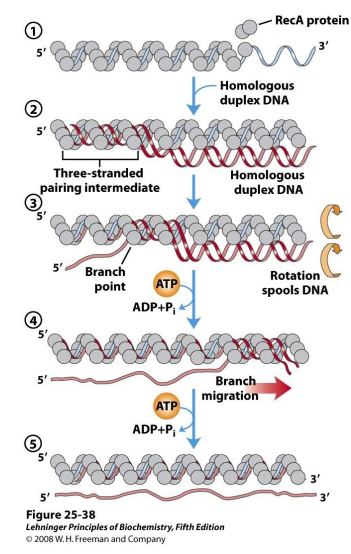
What does strand exchange look like?
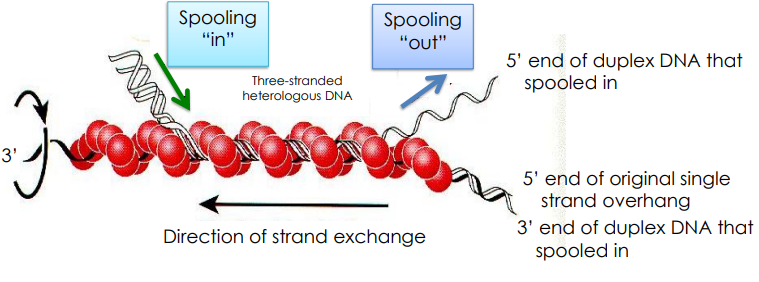
How do we fix strand breaks using Holliday Junctions?
RecA does Holliday Junction nucleation using its spooling mechanism to do strand exchange
Once a Hollidar Junction if formed, RecA polymers dissociate
are templates are made yay! Now, DNA polymerases/DNA ligases fill the gaps in the DNA strands
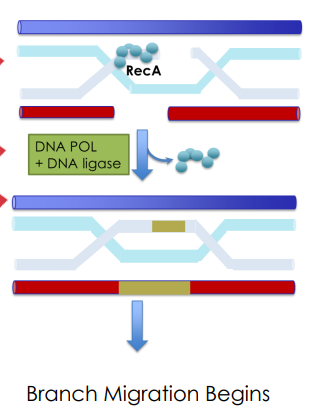
What two enzymes drive branch migration during DNA repair by homologous recombination?
RuvA and RuvB
Describe RuvA
DNA-binding protein that binds Holliday junctions with high affinity
RuvA is a tetramer w/ four channels that are lined with basic AA residues that will bind to our Holliday Junction
it has a central pin of eight acidic AA which serve to repel the negatively charged DNA backbone, which causes the DNA to not go to the center of the holiday junction
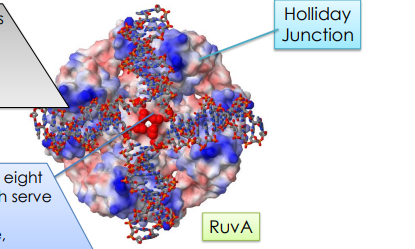
RuvB
An ATP-dependent helicase spools DNA through RuvA unit
Describe RuvA/RuvB Complex Formation
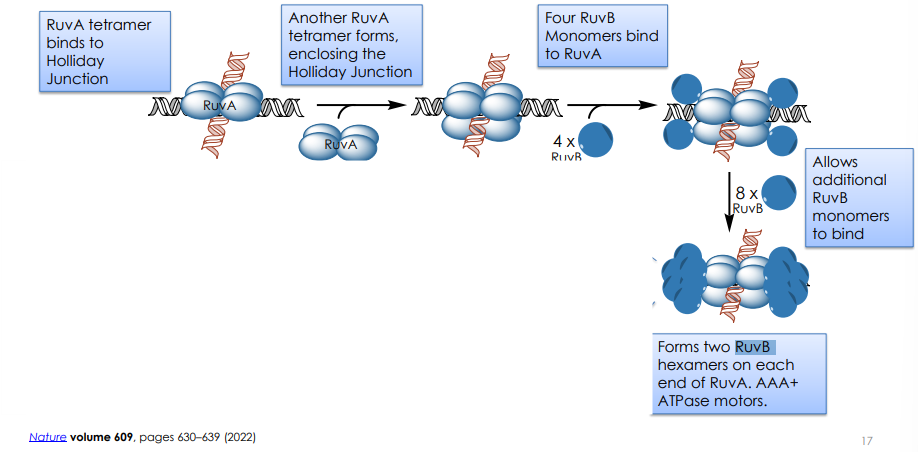
Which protein provides the motor for branch migration?
RuvB: RuvB is a hexameric ATPase (a type of molecular motor) that forms rings around the DNA arms flanking the RuvA tetramer. It uses the energy from ATP hydrolysis to pull the DNA through the junction, effectively "ratcheting" the branch point along the DNA.
Which protein resolves the Holliday junctions during repair? What does it do and what cofactordoes it require?
RuvC
Cleaves Holliday Junctions when homology between the strands end
Mg2+ dependent
RuvC mechanism
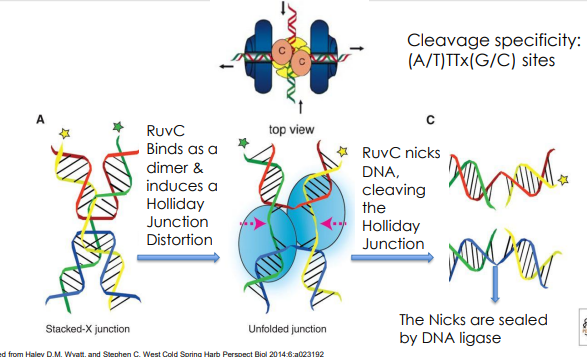
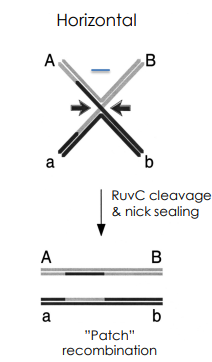
What is a patch recombinant?
RuvC-Mediated Horizontal Strand Cleavage
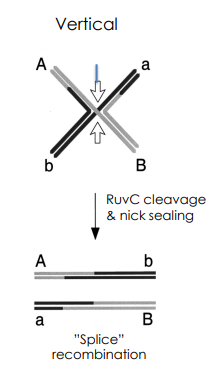
What is a splice recombinant?
RuvC-Mediated Vertical Strand Cleavage
Model for repair of DNA double-strand breaks
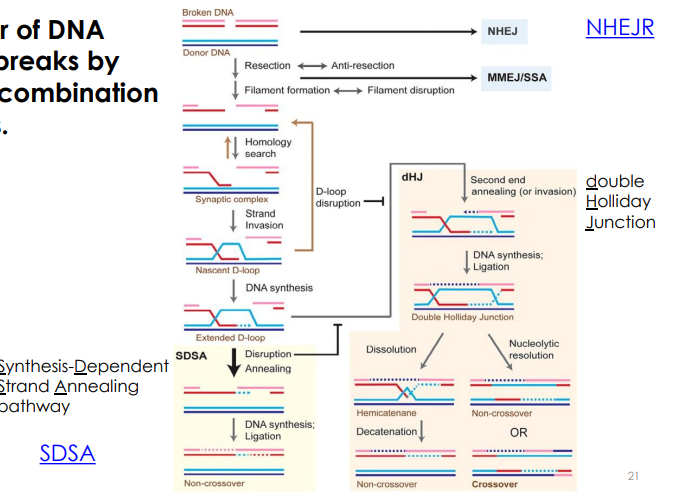
Eukaryotic Homolog of RecA
Rad 51
Eukaryotic Homolog of RuvAB
Rad54
Eukaryotic Homolog of RuvC
Resolvase A
What processes does genetic recombination affect?
specilized forms of DNA repair
specialized DNA replication situations
genetic diversity
regulation for expression of certain genes
chromosome segregation in eukaryotes
immune adaptability
emryogenesis
Three classifications of genetic recombination
Homologous
site specific
DNA transposition
Homologous Genetic Recombination
Involves genetic exchange between any two DNA molecules
Must share extended regions of near identical sequences
Does site-specific recombination require ATP or DNA synthesis?
No ATP needed: The energy required for cleaving and rejoining the DNA strands is stored in a covalent protein-DNA intermediate.
No DNA synthesis: This process involves the direct cutting and pasting of existing DNA strands without generating new DNA.
High specificity: It occurs at specific, short DNA sequences recognized by specialized enzymes called recombinases.
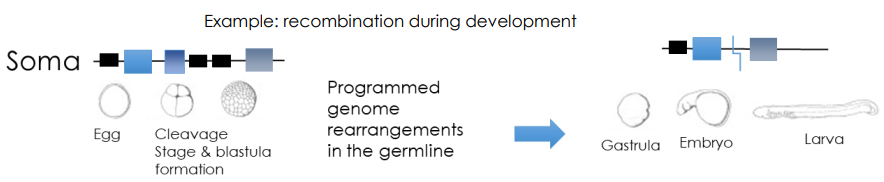
What are programmed genomic rearrangements? What purpose do they possible serve?
programmed genomic rearrangments that reorganize an organism’s DNA that occur at different stages of devlopment
Purpose:
massive diversity from limited code
permanent gene silencing
immune evasion
streamlining the genome
organism's DNA is not a static instruction manual, but a dynamic, editable script
Programmed DNA Elimination
Certain organisms completely destroy a large percentage of their genome in cells destined to become body tissues (soma), while keeping it intact in reproductive cells (germline)
type of programmed genomic rearrangment

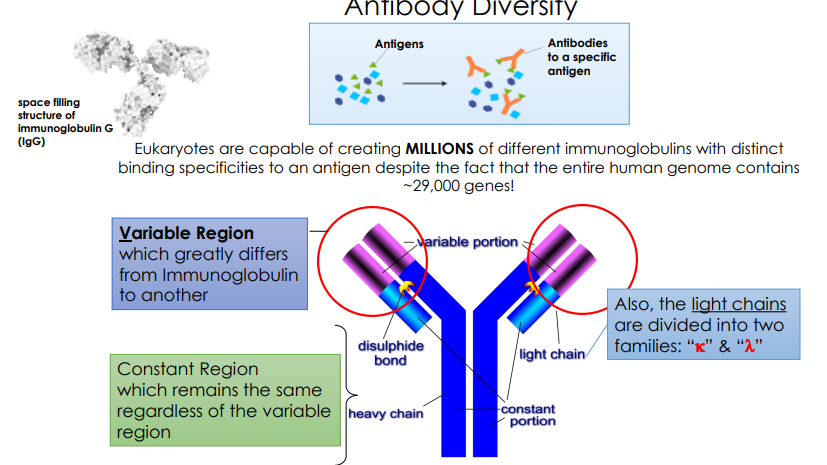
Antibody Diversity
type of programmed genomic rearrangment
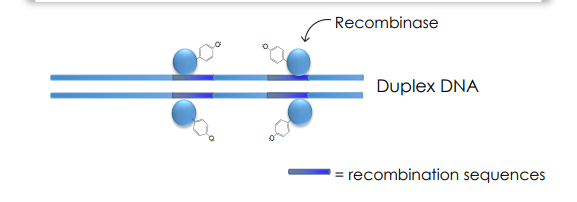
Recombinases Mechanism of Action
Recombinases use tyrosine or serine to act as a nucleophile to attack the phosphodiester backbone of DNA
A total of four recombinases will bind independently to the recombination sites and sequentially act to break DNA for recombination
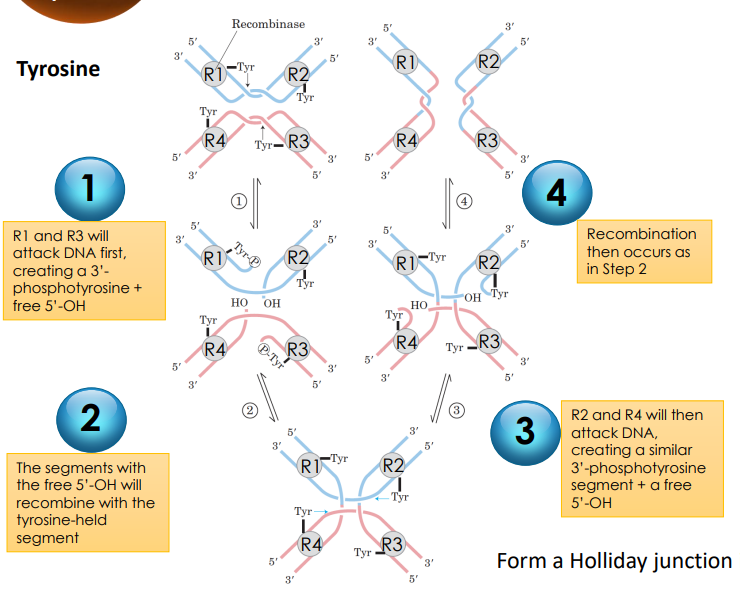
What are two types of site-specific recombinases?
Tyrosine recombinases and Serine recombinases
The Mechanism of Site-Specific Recombination for Tyrosine Serine Recombinases
Describe how programmed gene rearrangements are able to generate such incredible antibody diversity.
The body creates nearly infinite diversity by mixing and matching small segments to make the variable region tips, scrambling the connections, pairing random light and heavy chains, and then snapping them onto a standard constant region.
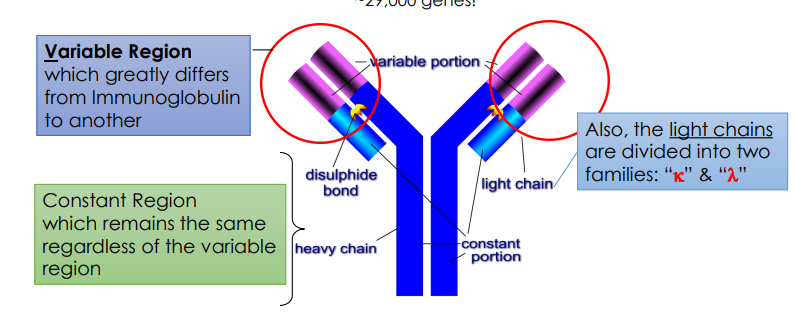
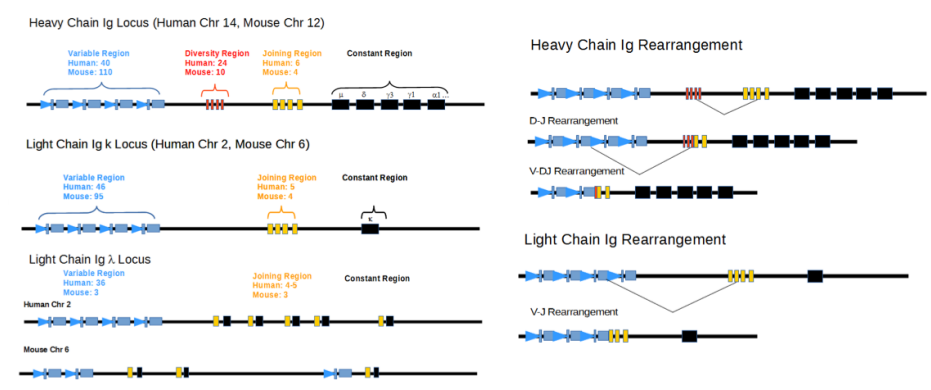
Which genes are responsible for heavy and light chain rearrangements?
RAG1 and RAG2 (Recombination-Activating Genes):
code for a specialized protein complex that acts as a targeted molecular scissors.
They scan the DNA, recognize specific signals flanking the variable regions, and physically slice the double-stranded DNA.
They are only active in developing lymphocytes.
Exons (the actual segments of the DNA that carry the code to make proteins) make up 1.5% of the human genome
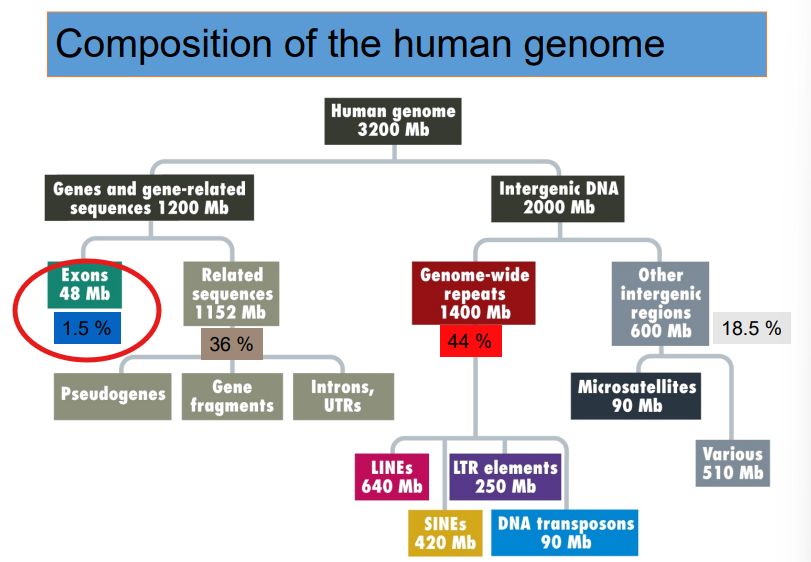
What percentage of the genome is made up of LINES and SINES?
~33%
NOTE: they are retrotransposons (a class of "jumping genes")
What is a “jumping gene”?
a Transposons!!: specific sequences of DNA that can physically move or duplicate themselves from one location in the genome to another.
Transposition
Recombination that allows the movement of transposable element
not based on DNA sequence homology
Enzymes catalyzing transposition recognize specific short sequences
Involves DNA synthesis
Involves duplication of the target site • Often the transposable element is replicated
Transposable elements can restructure a host chromosome
Transposons code for enzymes that insert it into the recipient DNA
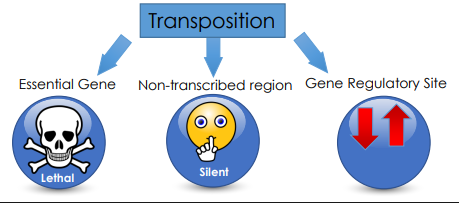
What are the potential effects of transposition on a gene?
if inserted on exon → often, lethal/loss of function (bcs of Frameshift Mutations or Premature Stop Codons); however sometimes we may get exonization
Insertion into a Nontranscribed Region → silent/ no immediate effect
insertion into a promoter → Altered Gene Expression
What are hallmarks of transpositions?**
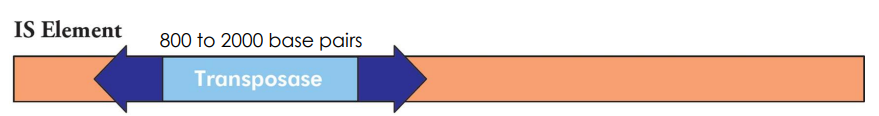
What is an IS element?
Insertion Sequence (IS) for transposition + transposase gene (encodes of the enzyme that promote transposition)
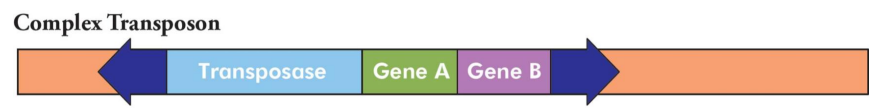
What is a complex transposon?
Insertion Sequence (IS) for transposition + transposase gene(s) + extra genes not directly involved in transposition
What is a composite transposon?
Consists of two identical or nearly identical IS-like modules (blue) flanking a central region carrying various genes. The IS-like modules may have either direct or inverted relative orientations.

Insertion sequences mechanistically
The IS Element (The Guest): This is the actual mobile piece of DNA. It contains only the genes necessary for its own movement (the transposase enzyme) flanked by terminal inverted repeats. It does not contain the target sequence.
The Target Sequence (The Host): This is the specific site on the host's bacterial chromosome where the IS element decides to land.
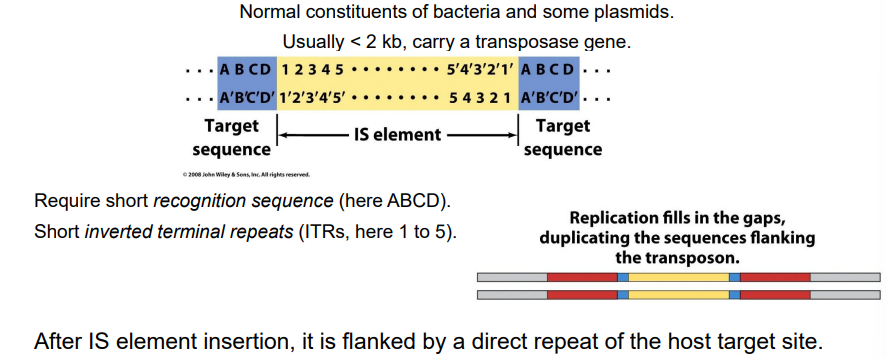
What happens to the host target site after transposition?
short, direct repeating sequences of the host's own DNA are created, flanking the left and right ends of the newly arrived transposon
What is a medical problem that can occur because of transposition?
Hemophilia A
How Transposition Causes Hemophilia A
The Target: The gene responsible for producing Factor VIII (a critical blood-clotting protein).
The Event: An active LINE-1 (L1) element jumps and inserts itself directly into an exon of the Factor VIII gene.
The Result: The insertion disrupts the genetic code. The cell can no longer create the functional clotting protein, leading to the severe bleeding disorder known as Hemophilia A
Describe the process of “cut and paste” transposition which is also known as direct transposition.
1. Tn5 transposase monomers bind to “outside end” inverted repeat sequences (green).
2. Formation of transposition complex. DNA bends and two transposases interact.
3. After dimerization, each transposase activates a water molecule for nucleophilic attack of the opposite strand of DNA.
A hairpin is formed at each end, which excises the transposon. Hairpin is hydrolyzed. = CUT
Transposase dimer and transposon DNA (“synaptic complex”) find target site in genome. Another nucleophilic attack by the transposon’s 3’-OH group results in insertion. = PASTE
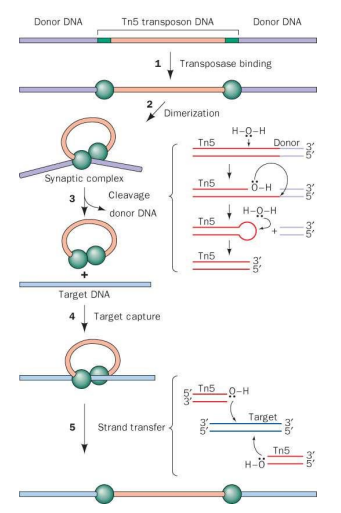
What provides the nucleophile for attack of the DNA strand during excision in “cut and paste” transposition?
During excision a water molecule acts as the nucleophile
What provides the nucleophile for the attack on the target DNA?
During Attack on the Target DNA (Strand Transfer), The exposed 3’-hydroxyl group (3’-OH) at the ends of the excised transposon.
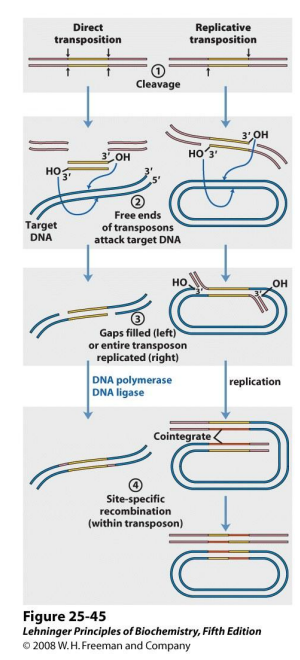
Direct vs. Replicative Transposition
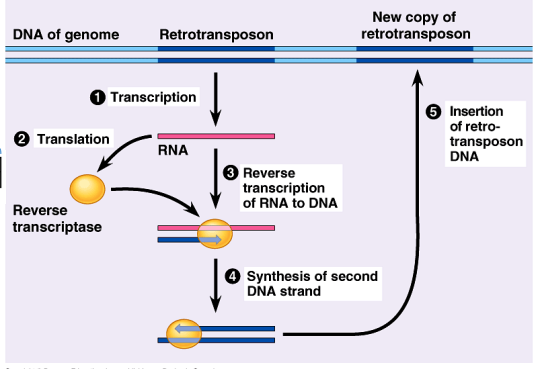
Describe the basic steps of retrotransposition
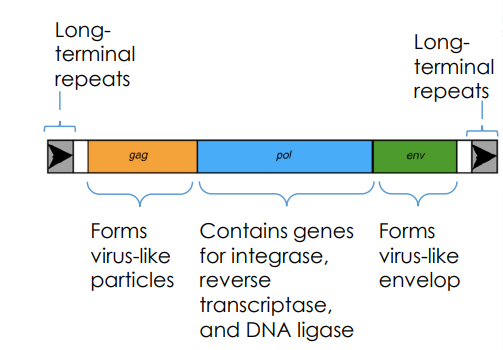
Long-Terminal Repeat (LTR) Retrotransposons
contain genes that act like retroviruses which allow for reverse transcription of an RNA to DNA, and also for integration of the transposon into the genome
NOTE: LINES and SINES use retrotransposition!!
What type of eukaryotic transposon increases in number?
replicative