M4L1 - Transport to Peroxisome
1/22
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
Peroxisome Characteristics
Small-oval shaped organelle
Bound by a single membrane
Responsible for oxidative/synthetic functions
doesn’t have its own genetic info
Reproduces by binary fission
Peroxisome TEM
Contains dense regions with protein aggregates
Protein: catalase enzyme (anti-oxidant protein)
Function in Other Cells of Peroxisome
Animal Cells:
Peroxisome responsible for cholesterol synthesis
In nerve cells, it synthesizes plasmalogen
In liver cells, it is the site of toxin oxidation (alcohol)
Plant Cells:
Requires for conversino of fatty acid to carbs (important in seeds germination)
All Cells:
Responsible for catalysis of fatty acids
Major Function of Peroxisome
Breakdown of very long fatty acids through beta-oxidation
Animals: Long fatty acids are oxidized to medium ones
Goes to mitochondria for further processing
They can then be used as energy source
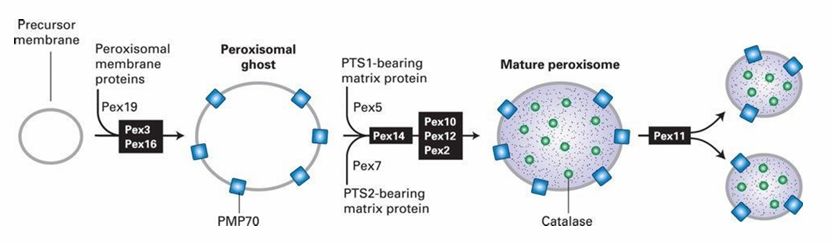
Fatty Acid Oxidation in Peroxisome
Done through beta-oxidation
Fatty acids yield the most ATP on E/gram basis compared to other macromolecules
Oxidation of fatty acids in peroxisome helps release metabolic energy
This process creates hydrogen peroxide byproduct (harmful)
Catalase Enzyme:
Abundant in peroxisomes
Converts H2O2 into O2 and H2O molecules (not harmful)
Catalase can be used as a marker for peroxisome visualization in cells bc it’s abundant
Catalase Enzyme (Peroxisome and Beta Oxidation)
Abundant in peroxisomes
Converts harmful H2O2 byproduct into O2 and H2O molecules (not harmful)
Catalase can be used as a marker for peroxisome visualization in cells bc it’s abundant
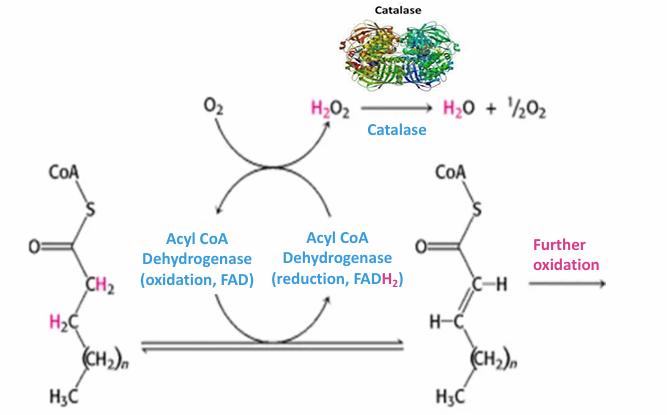
Fluorescence microscopy images of catalase in cell
Uses catalase primary antibody (immunofluorescence)
This is recognized by a red-flourescent secondary antibody
Each dot represents a peroxisome
Not high resolution, but you can identify there functional peroxisomes exist in the cell
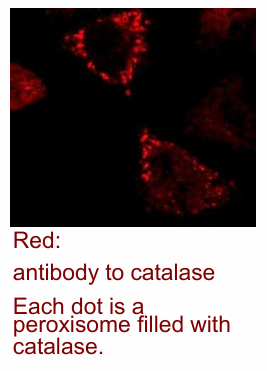
fluorescence image of a single cell (DNA/actin microfilament/catalase)
DNA stain (viswualize nucleas in red)
Actin microfilament network (shown in blue)
Catalase enzyme labelled with antibody (green)
Each bright feen stain represents individual peroxisomes
Fluorescence can look at living cells while TEM cannot
Peroxisomes aren’t static
They move around, change shape, undergo fission/fusion
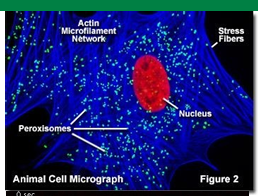
Peroxisome Biogenesis
Peroxisomal proteins are synthesized in the cytosol then transported to the peroxisome
Peroxisomal membrane proteins (PMP70) attached to precursor membranes creates ‘peroxisomal ghost’
They’re used to transport peroxisomal matrix proteins such as catalase inside
More proteins synethesized = more growth of peroxisome
New peroxisomes can form by fission
This lets cells make more peroxisomes and give them to new cells
Luciferase: Peroxisomal Protein Transport
Enzyme that allows the bioluminescence in fireflies
This enzyme is found in the peroxisome of cells in firefly abdomens
Experiment:
Inserted luciferase genes into mammals
Found that after expression, it went straight to peroxisomes
Showed mechanism for sending it to the peroxisome is same for both fireflies and mammals
Signals and transport proteins involved are the same
5 Rules for Protein Transport
A signal sequence on the transported protein
A receptor for that signal sequence on the target organelle/membrane
A translocation channel to help get the protein across
Required energy (ATP) at some step in the process
A way of targeting a protein to specific locations within the organelle
Rule 1 for Protein Transport: Peroxisome Protein Ex
PTS1: Peroxisomal-Transport Sequence 1 (aka SKL)
Found in many species’ peroxisomal proteins
Tri-peptide made of 3 amino acids
serine
lysine
leucine
Found at C-terminus of translated peroxisomal protein
N-terminus translated first
C-terminus translated last
Because the signal sequence isn’t available until the entire protein is finished, the protein is sent to the peroxisome after translation is complete.
This is called post-translational transport
It folds in the cytosol, leaving the C-terminal sequence visible
Facilitates transport
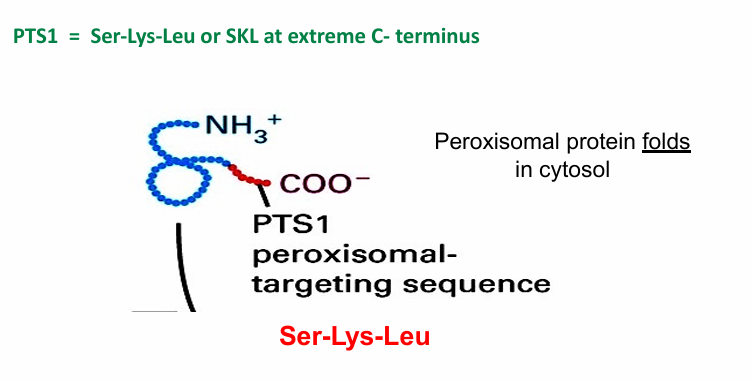
PTS1: Necessary for Protein Import?
Basis:
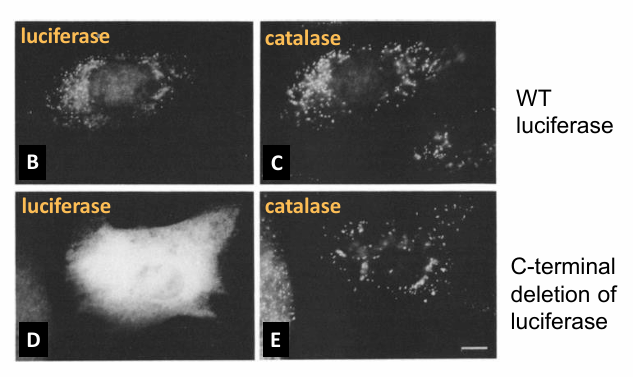
Wild-type Luciferase
Signal Sequence is present so protein is visible within bioluminescent peroxisomes in this single cell
C-terminus Deletion
Hypothesis: If PTS1 is necessary for protein transport into peroxisome, then luciferse with the deletion will no longer go
Results:
Image B: Luciferase in spots in the cell
Image C: Same cell stained with antibody to catalase
Colocalization of the spots confirms luciferase is in the same location as catalase
Suggests successful transport to peroxisome
Image D: Modified luciferase protein (no signal)
Bioluminescence is shown throughout the cytosol
Image E: Same cell as D but stained with antibody to catalase
Not in the same areas only
Conclusion: PTS1 is necessary for transport
Parallel Further Experiment
Is each of the amino acids in the sequence necessary for transport?
Tested by single amino acid sub in PTS1 sequence
PTS1: Sufficient for Protein Import? (DHFR)
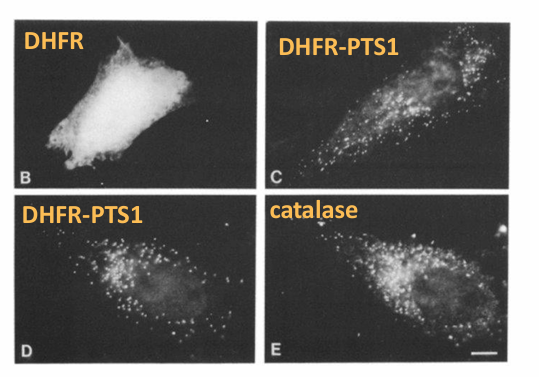
Basis:
Check is PTS1 sequence can for any protein into the peroxisome
Ex. DHFR
Cytosolic protein
Wildtype: Normal without sequence
DHFR plus C-terminal luciferase
Hypothesis: If PTS1 sequence is sufficient and added to DHFR, then it will be transported to the peroxisome
Result:
Image B: Wildtype DHFR localizes in cytosol
Image C: DHFR-PTSL is found in punctate pattern in cell
Image D + E: Co-localization of DHFR-PTS1 and catalase
Indicates the' DHFR-PTS1 has moved to the peroxisomes
Conclusion: Yes it is sufficient
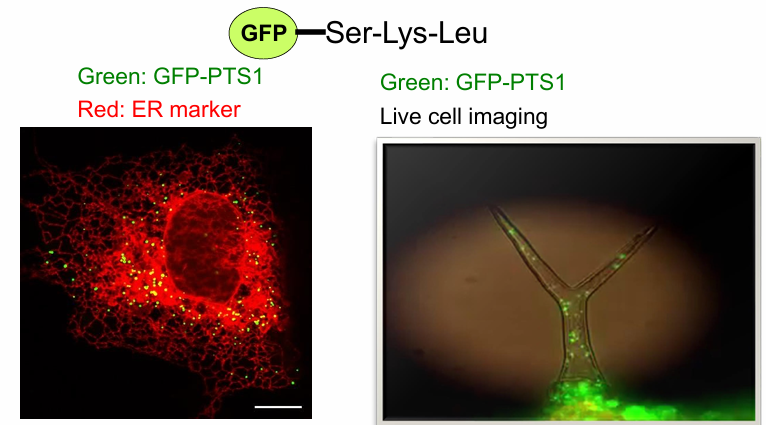
PTS1: Sufficient for Protein Import? (GFP)
GFP: Green fluorescent protein found in jellyfish
It’s a cytosolic protein
When modified with PTS1 sequence, it moves to the peroxisome as well
This is also shown in the tip of Arabidopsis plant
GFP-PTS1 found in peroxisomes as they move within the root tip cells
Conclusion: Yes, sufficient once again!
Rule 2: Signal Receptor for PTS1?
Pex5: Cytosolic protein
Recognizes PTS1
Binds to the PTS1 sequence at C-terminus of target protein
Pex14: Peroxisomal Membrane protein
Pex5 associates with Pex14
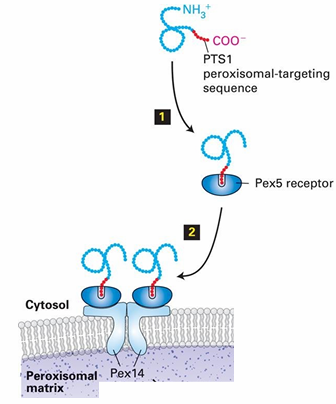
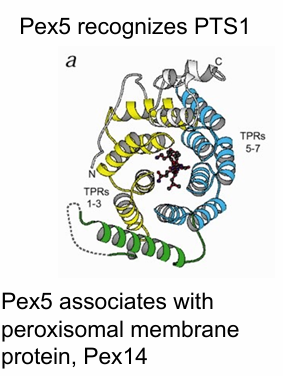
Pex5 binding mechanism to PTS1 Signal
Contains 7 TPR motifs
Each motif has 2 a-helices connected by a loop
The 7 TRP motifs form a PTS1 binding pocket
Inside pocket: PTS1 tripeptide
Rule 3: Translocation channel on peroxisome?
Pex14
Important for Pex5 recognition
Forms translocon that transports Pex5 with target protein to the inside (peroxisomal matrix)
When inside, Pex5 dissociates from the protein
It is then recycled back out to the cytosol
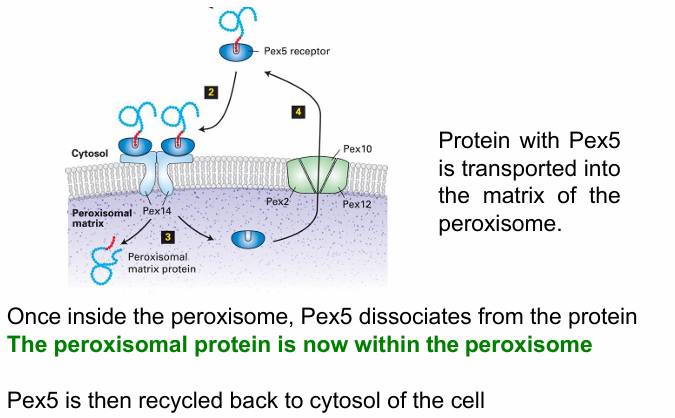
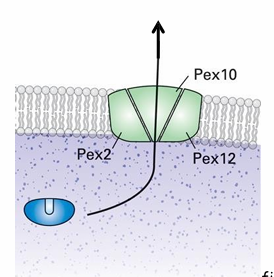
Rule 4: Energy Requirement for Peroxisomal Protein Transport?
Yes during ubiquitinylation
Pex5 is ubiquitinylated and deubiquitinylated to release the protein in the matrix and be brought back to cytosol
Pex2 and Pex10 are ubiquitin ligases
Form complex with Pex12
Makes another transllocon to export Pex5
ubiquitinylation requires ATP hydrolysis
Identification of Peroxisomal Transport Pathway Components
Done by genetic screens to identify mutations preventing normal transport
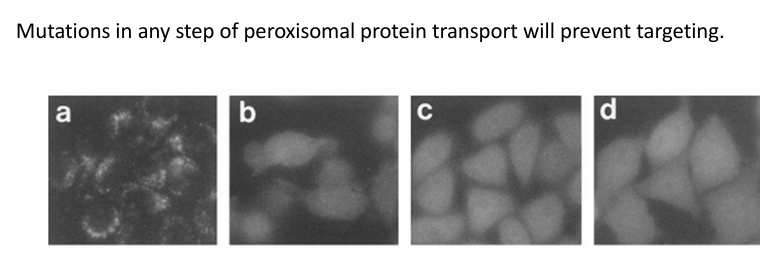
Figure a: GFP-PTS1 is localized to peroxisomes
Random mutations to genome were made, one per cell line
Each cell line was tested for its ability to transport GFP-PTS1 to peroxisome
If unsuccessful, the cell line carried a mutation in a gene coding for a protein in the pathway
The genes were named in order of identification
Pex1/Pex2/Pex3/etc
Figures b-d: Examples of cell lines where GFP-PTS1 remained in cytosol
Carry mutations in the pex genes
Rule 5: Protein Transport to Different Interior Compartments (Pex12 Mutation)
2 Places in peroxisome
Membrane (Ex. components of translocon)
Matrix (Ex. catalase)
They use different pathways to do so
Experiment:
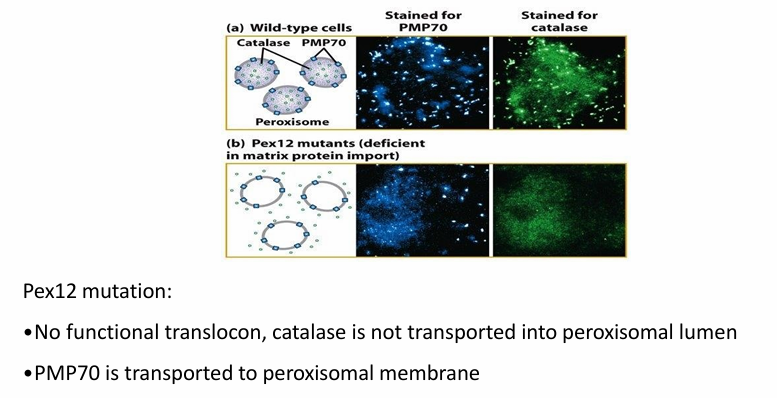
Images follow antibody to membrane protein PMP70 (blue) OR antibody to catalase (green)
Figure a: Wild-type PMP70 and catalase shown in punctate pattern
Figure b: Mutation in Pex12 gene
Catalase not transported into peroxisome
PMP70 successfully transported
Conclusion: There are distinct pathways of transport where Pex12 is only required for transport of proteins into matrix
Rule 5: Protein Transport to Different Interior Compartments (Pex3 Mutation)
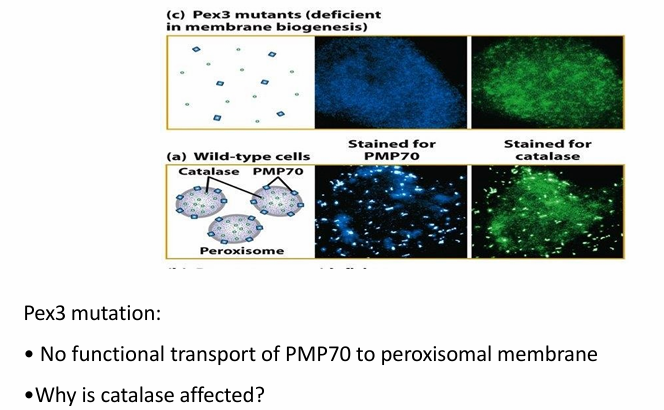
Mutation in Pex3 disrupts PMP70 transport
Figure c: Has mutation on Pex3
PMP70 is found within cytosol (blue)
Pex3
Needed to help form peroxisome membrane as well
Affects transport of both PMP70 and catalase
Proteins needed for catalase transport are peroxisomal membrane proteins as well (ex. Pex14)
If system to transport proteins into the membrane is faulty, the cell can’t transport proteins into the matrix as well
Zellweger’s syndrome
Caused by failure to transport proteins which results in nonfunctional peroxisomes
Autosomal Recessive disorder caused by mutations in genes encoding Pex2/3/5/10/12
Mutations in any of the genes lead to the same phenotype
Result
Cells accumulate long fatty acid chains
Impairs the normal function of many organ systems by disrupting
neuronal migration
nueronal positioning
brain development
Image:
Figure a: Human cells carrying mutation in Pex19
Impaired pathway as catalase is found in cytosol
Figure b: Rescued cells by addition of functional Pex19 gene
Catalase is now localized
There are no treatments to the condition though :(