BIMM 100 Exam 1
1/110
Earn XP
Description and Tags
covers LE 1-6, week 2-3 DI notes & slides, exam 1 review notes & slides (incl. practice problems), problem sets 1-3 (3rd is the optional problem set, but likely include flashcards on it)
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
111 Terms
def. gene
2 conditions for it to be the molecular basis of a gene
a unit of hereditary trait that shapes a specific trait of a cell or organism
(hereditary traits - traits passed through generations)
_
DNA
should be transmitted from cell-to-cell (or parent-to-child)
should generate a trait
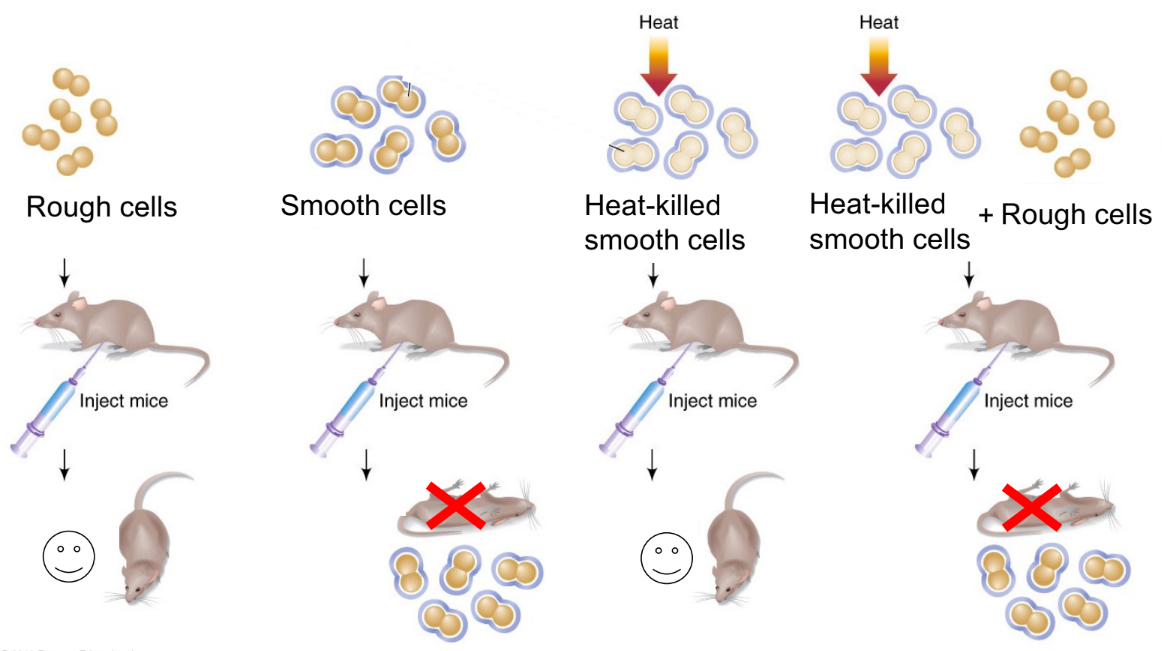
what did Griffith’s experiment discover? (1)
_
describe rough vs. smooth bacteria (← 2 each)
(& what happened to mice infected with each?)
_
what happens to:
mice w/ smooth cell bacteria introduced to heat (←explain w/ 2)
mice w/ smooth cell bacteria introduced to heat + rough cells (← explain w/ ~3, where 1 of them def. diff. b/w genes and a specific term)
_
are dead bacteria virulent or non-virulent?
_
genes shape __, while chromosomes do what? (1)
discovered that a trait, like bacterial virulence (cause death/disease), can be transferred from cell-to-cell
_
rough bacteria is non-virulent
do NOT have polysaccharide “cloak”
are destroyed by the mouse immune system
SO mice infected with R cells survive
smooth bacteria is virulent
have polysaccharide “cloak”
sneak past the mouse immune system
SO mice infected with S cells die
__
heat-killed smooth cells → mice survive
b/c heat-killed (dead) smooth cells are non-virulent
heat kills bacteria
dead bacteria are non-virulent
dead smooth cells + live rough cells → mice die
b/c live smooth cells are recovered from the dead mouse
dead cells can transfer their traits to live cells
^ virulence can be transferred from cell-to-cell
SO something else is transmitted from S cells → R cells in order to change the virulence of R cells, which are CHROMOSOMES
genes shape traits, while chromosomes carry genetic info & have both nucleic acids AND proteins
def. chromosomes
describe (2)
___
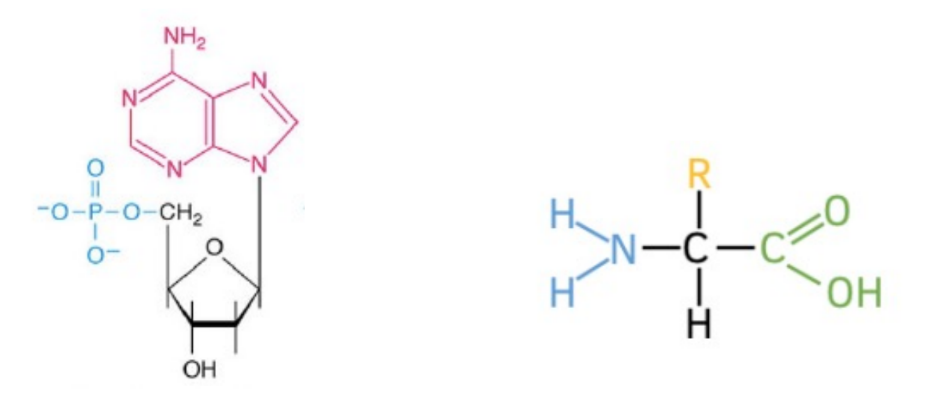
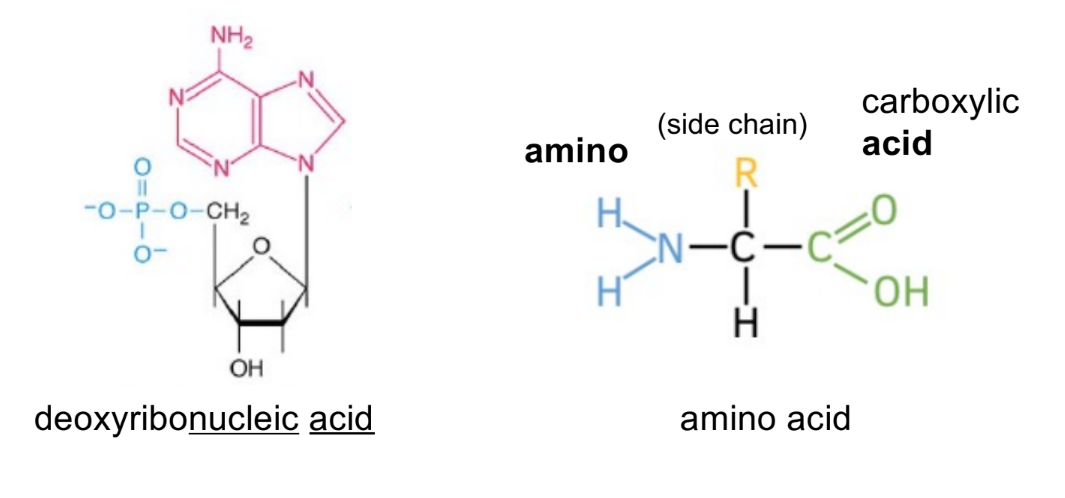
label which is nucleic acid vs. amino acid
_
identify 5 parts of an amino acid
#of common amino acids
_
identify 3 parts of nucleic acid
name the 5 nucleotides (which 2 belong to what specifically)
are carriers of heritable traits/genes & are made of nucleic acids AND proteins
when a cell divides, chromosomes are transmitted to both daughter cells
inheritance of specific traits correlates to inheritance of chromosomes
__
amino acid has central carbon (alpha carbon), attached to:
amino group (NH2)
side chain (R)
carboxylic acid group (COOH)
H
__
nucleic acid has multiple:
phosphate group
sugar ring (is deoxyribose sugar for DNA)
Nitrogenous base
nucleotides (have one of each of the above)
adenine, cytosine, guanine, thymine (DNA), uracil (RNA)
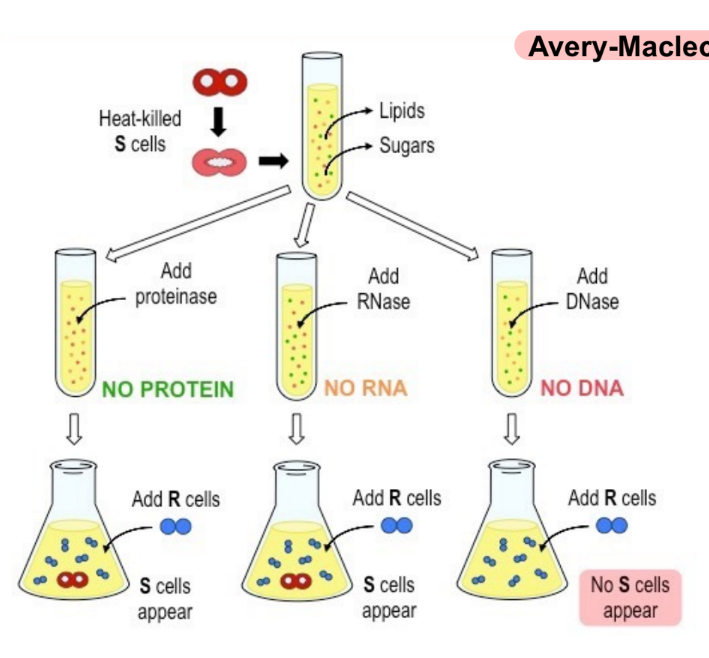
review: what molecule is needed to transmit S cells → R cells in order to change the virulence of R cells
conclusion of Avery-Macleod-McCarty experiment (1)
_
what happens to R cells if you:
break open/lyse S cells, eliminate protein w/ proteinase, add R cells to S-cell lysate that lacks protein
lyse S cells, eliminate DNA w/ DNase, add R cells to S-cell lysate that lacks DNA
chromosome, that contain nucleic acids (RNA, DNA) & proteins
which led to hypothesis that nucleic acid or protein was the genetic molecule
_
conclusion:
absence of DNA (the genetic material) will block rough/non-virulent cells from transforming into smooth/virulent cells (whereas, smooth cell DNA transforms rough cells into smooth cells)
(aka smooth cell traits are transmitted to rough cells by DNA)
__
w/ proteinase
S-cell lysate can still cause R/non-virulent cells to transform into S/virulent cells B/C the lysate doesn’t have protein BUT does have nucleic acid
→ protein is not the genetic material
w/ DNase
R/non-virulent cells do NOT transform into S/virulent cells B/C the lysate does not contain DNA, even though it does contain RNA and proteins
→ DNA is the genetic material
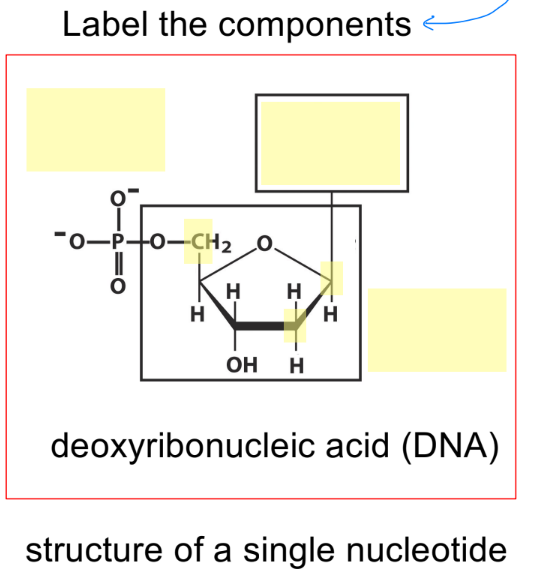
include C1-5
_
DNA is 2 strands of __ in a helix
DNA is 2 strands of nucleotides in a helix
(picture should be deoxyribose sugar ring)
(remember that a nucleotide is made up of 1 phosphate group, 1 sugar ring, 1 nitrogenous base)
(a nucleic acid is made of multiple nucleotides)
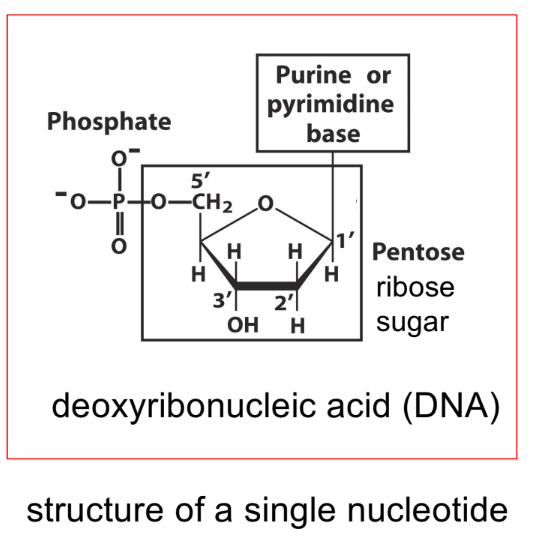
diff. b/w results & conclusion
results = data/ / observations
conclusion = interpretation
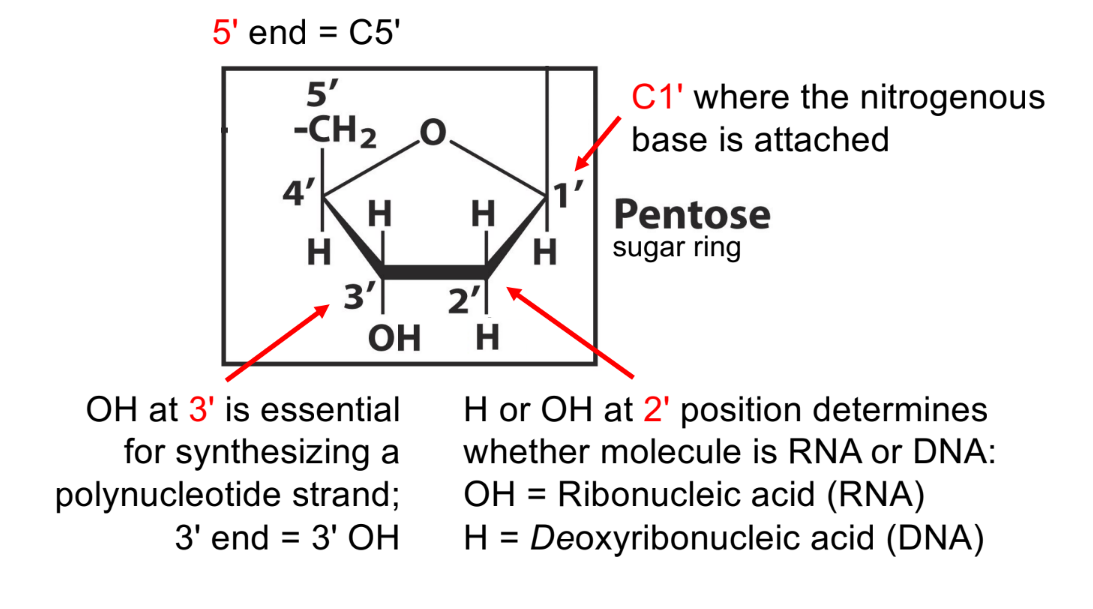
state what is attached at positions C1-3 and C5 in a pentose sugar ring
for 1 of them, state its importance
C1’ — where nitrogenous base attaches (purine or pyrimidine base)
C2’ — determines if molecule is RNA or DNA (w/ hydroxyl group)
2’ OH means RNA (think RUH “ROH”)
not 2’ OH, but 2’ H, means DNA
C3’ — where OH/hydroxyl group attaches, to form a polynucleotide strand (3’ OH)
C5’ — where phosphate group attaches
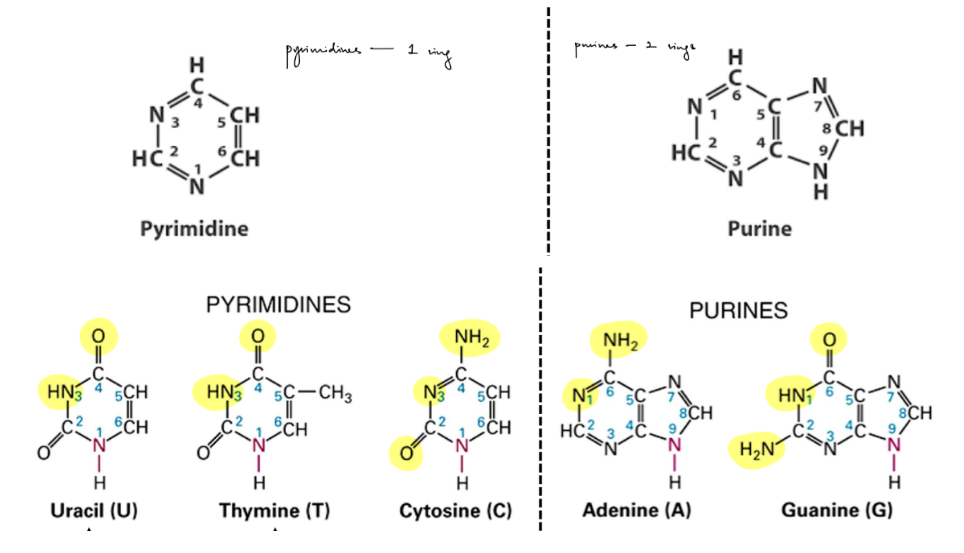
diff b/w purine vs. pyrimidine
which of the 5 bases are a part of each?
___
draw all the 5 bases (might help to think about the basic structure of purines & pyrimidines)
& highlight which parts are hydrogen bond donors & acceptors
^ which 2 bases have 3 total donors/acceptors vs. 2
(MB know #s w/in a ring)
^^ can also refer to drawings in midterm notebook for better visualization
purine - 2 rings
A, G
pyrimidine - 1 ring
C, T, U
__
C-G have 3 H bonds
A-T have 3 H bonds
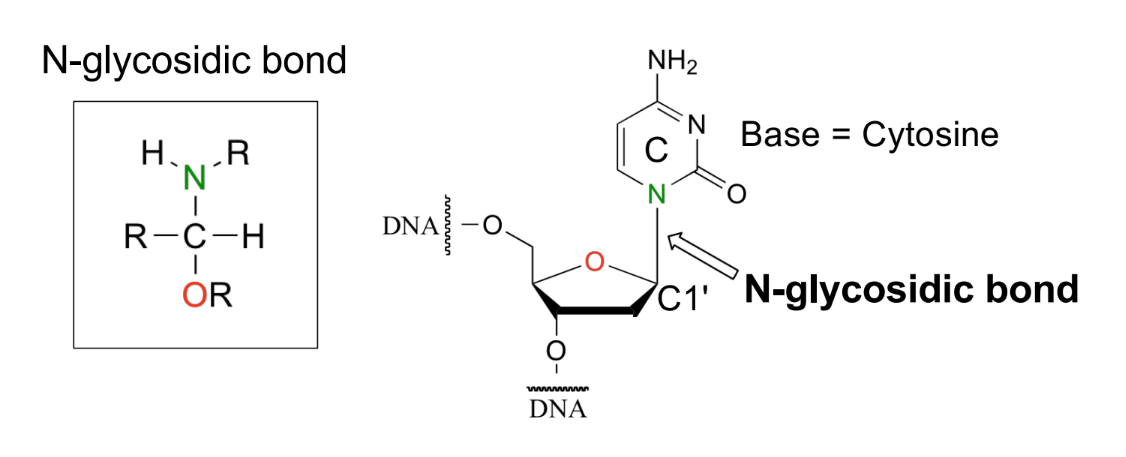
def. glycosidic bond
enzyme that cleaves it
bond b/w C1’ of sugar ring & N-base
cleaved by glycosylase
nucleoside vs. nucleotide (1 each)
name 4 nucleosides
t/f: different #s of phosphates can compose a nucleotide (meaning?)
nucleoside has attached OH (hydroxyl) group
adenosine, thymidine, guanidine, cytidine
nucleotide has attached phosphate group
(^ both have sugar ring & N-base)
__
true
dAMP (deoxy””-5’-monophosphate), dADP (bi-), dATP (tri-)
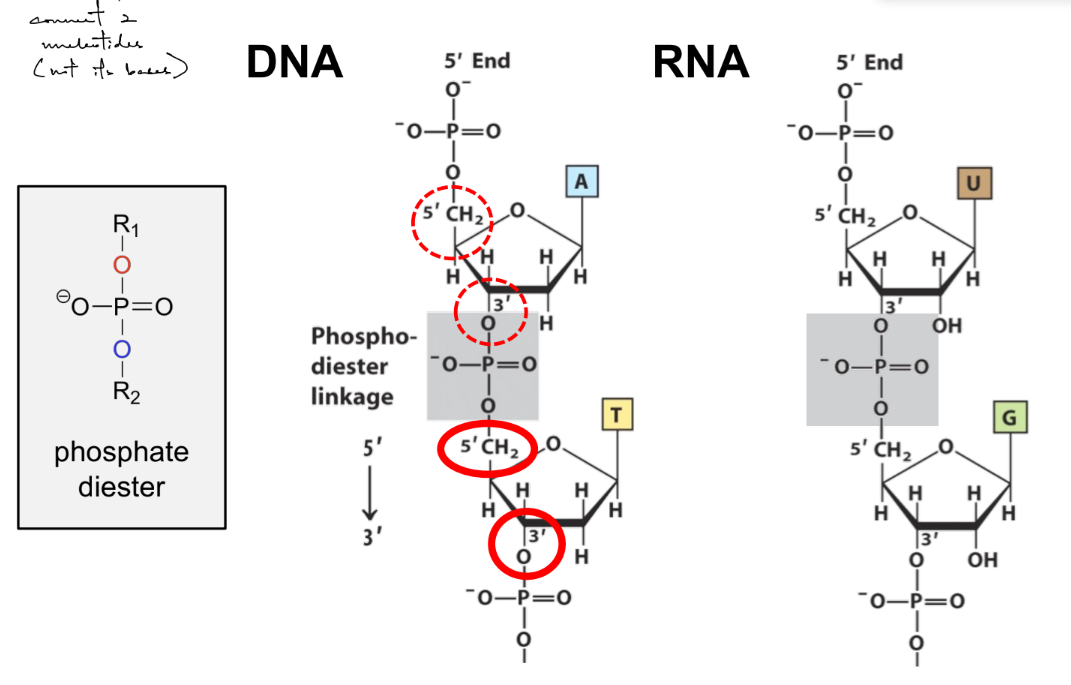
name 4 things that make RNA diff. from DNA
RNA
ribose sugar, where 2’ OH (think RUH “ROH”)
has uracil, not thymine
usually single stranded
can function as an enzyme: “ribozyme”
def. phosphodiester bond
what enzymes forms it (2) ← & name b/w what 2 things the bond forms between
what enzymes cleave it (2)
this bond forms (1)
bond b/w 3’ OH & 5’ of 2 nucleotides (specifically b/w 2 sugar rings)
(where 3’ OH becomes 3’ O that’s part of phosphate, which binds to the 5’ of another ring)
3” OH can be either with :
5’ phosphate w/in the strand — DNA ligase
5’ phosphate of incoming dNTP — DNA polymerase
__
made from DNA ligase & DNA polymerase
cleaved from exonuclease & endonuclease
__
phosphodiester bond forms the sugar-phosphate backbone w/in the SAME strand
describe 4 protein structures
bond of secondary? structures (can only name 1)?
___
name if single vs. double-stranded nucleic acid chains
primary - linear polypeptide chain of amino acids
secondary - folding of backbone w/ H bonds
alpha helix, etc.
tertiary - 3D folding of chain
quaternary - multiple chains into a protein
__
single - RNA
double - DNA
for discovering structure of DNA
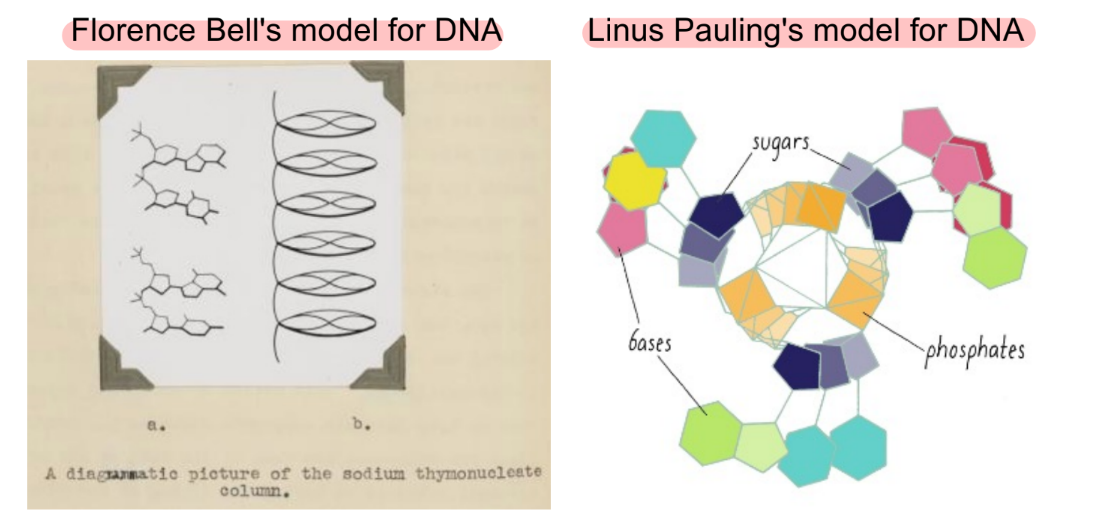
brief description of Florence Bell vs. Linus Pauling model for DNA ← both were incorrect models of DNA
what’s incorrect about Pauling’s model (2)
3 parts of correct DNA structure? ← by Franklin-Wilkins-Watson-Crick
direction of DNA movement
Bell — stacked nucleotides
Pauling — triple helix
wrong about:
3 strands of DNA in the helix (triple helix) — should be double-helix
bases being outside the helix — should have bases from opp. strands pair w/ e/o in the center of the helix
__
2 strands of DNA (double-helix)
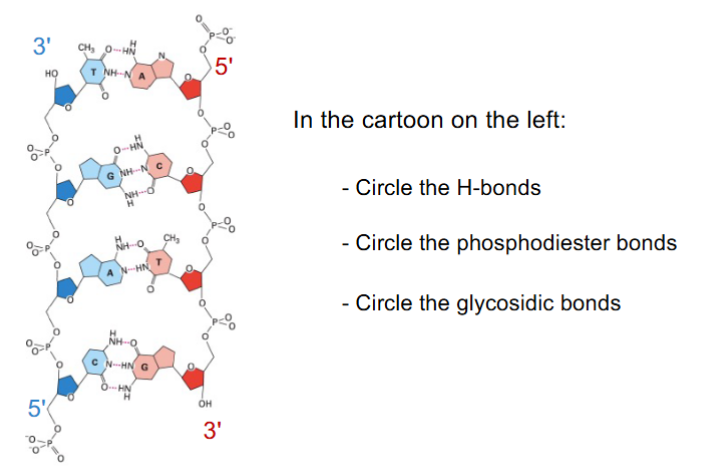
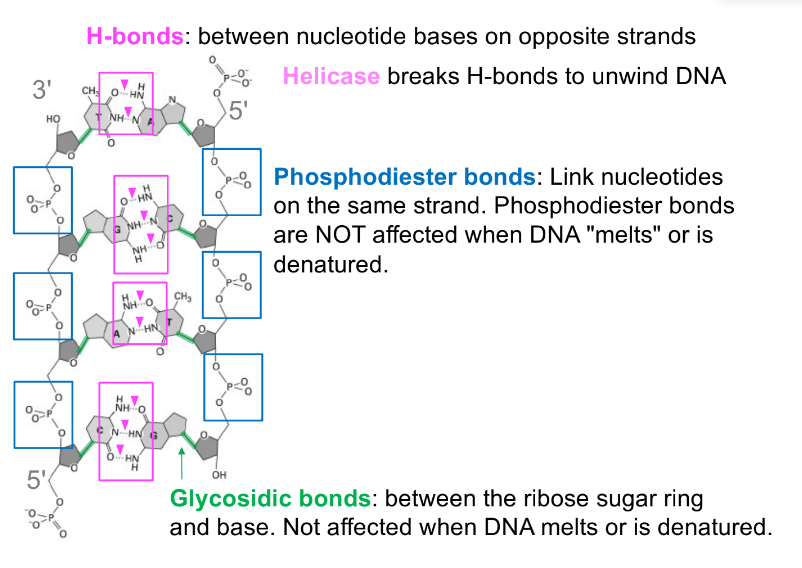
base pairs, where nucleotide bases on opposite strands form hydrogen bonds
the 2 strands are anti-parallel to e/o (5’→3’, 3’→5’ which refer to the C in the sugar ring)
DNA moves from 5’ → 3’ end
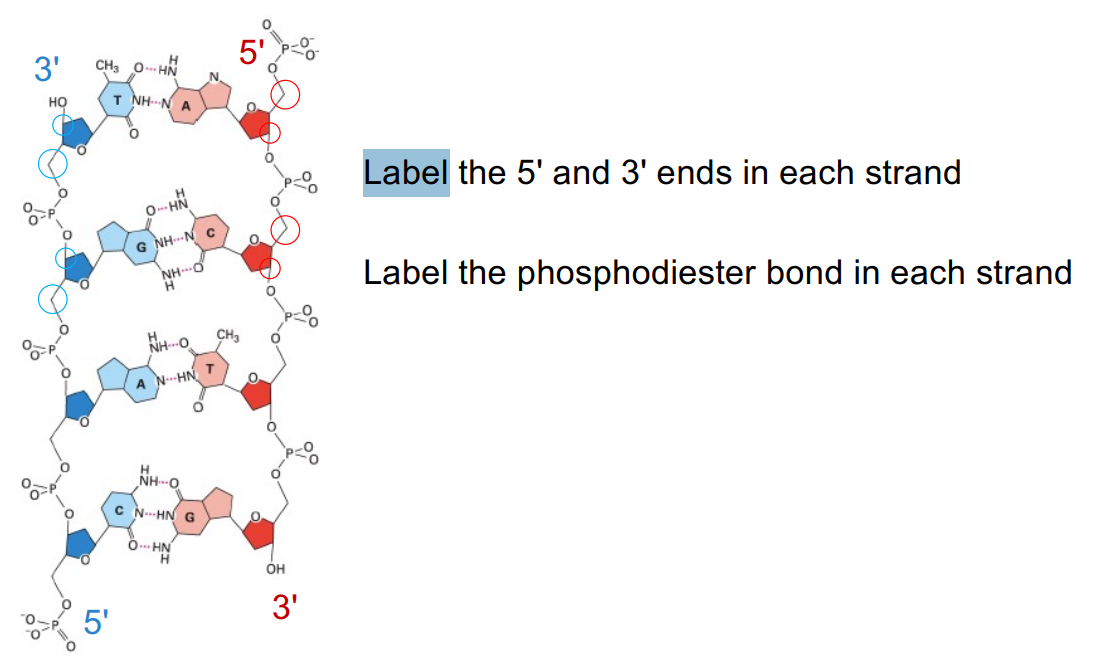
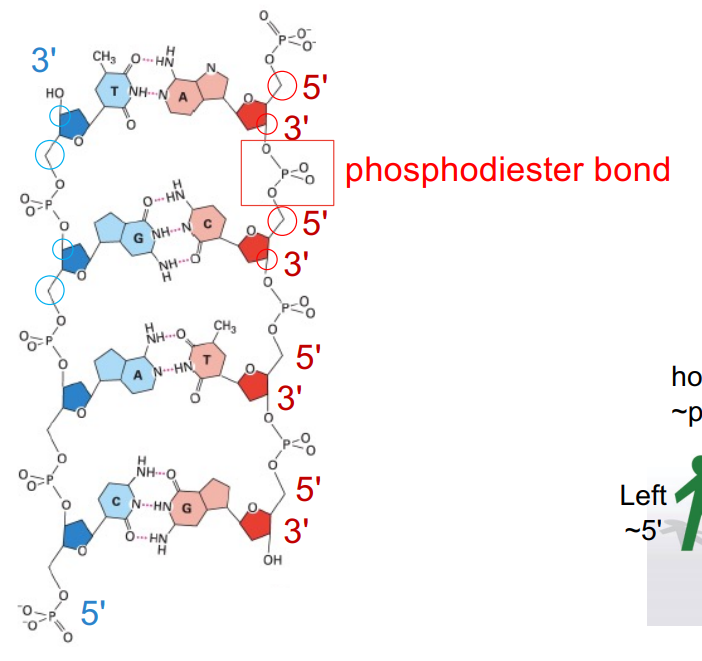
can RNA be antiparallel? explain
yes
even though it’s single-stranded, it can become a complex structure, where the single strand folds in on itself & the complementary bases can pair together ← anti-parallel
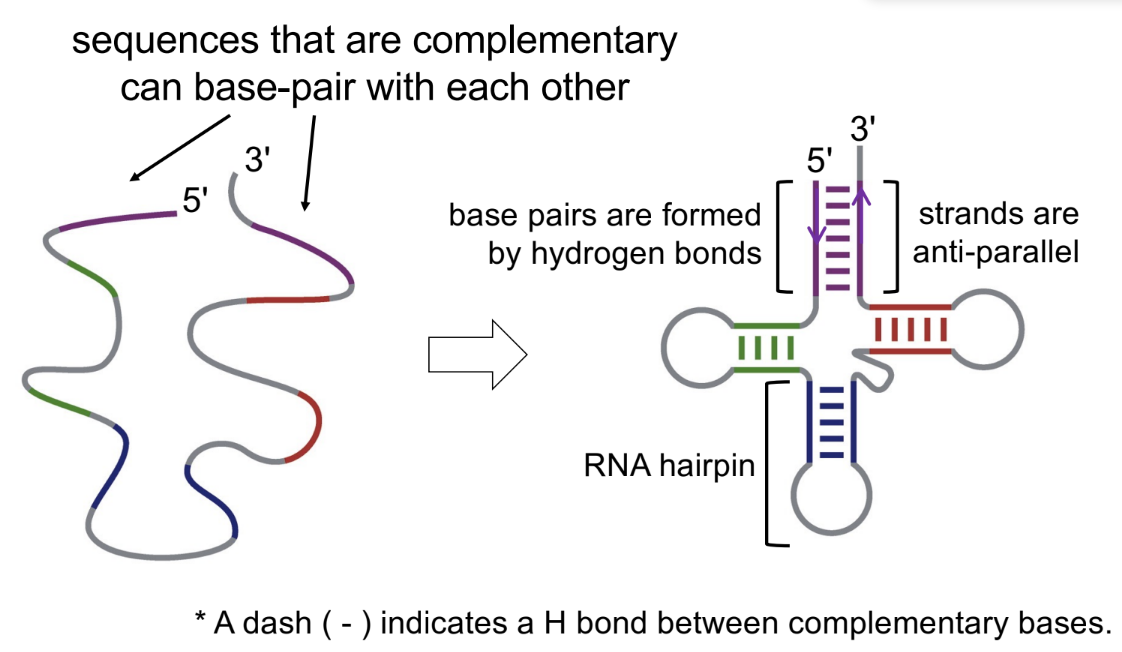
name the 2 types of intermolecular interactions present in DNA (& what are the interactions)
what about another one w/ what specific types of bonds?
__
(2) are important for stability of DNA
hydrogen bonds — base pairing (aka b/w bases of opposite strands)
Van der Waals forces — base stacking
(does NOT include any of the 3 bonds talked about earlier)
_
covalent bonds
glycosidic bonds
phosphodiester bonds
__
base pairing & base stacking for stability
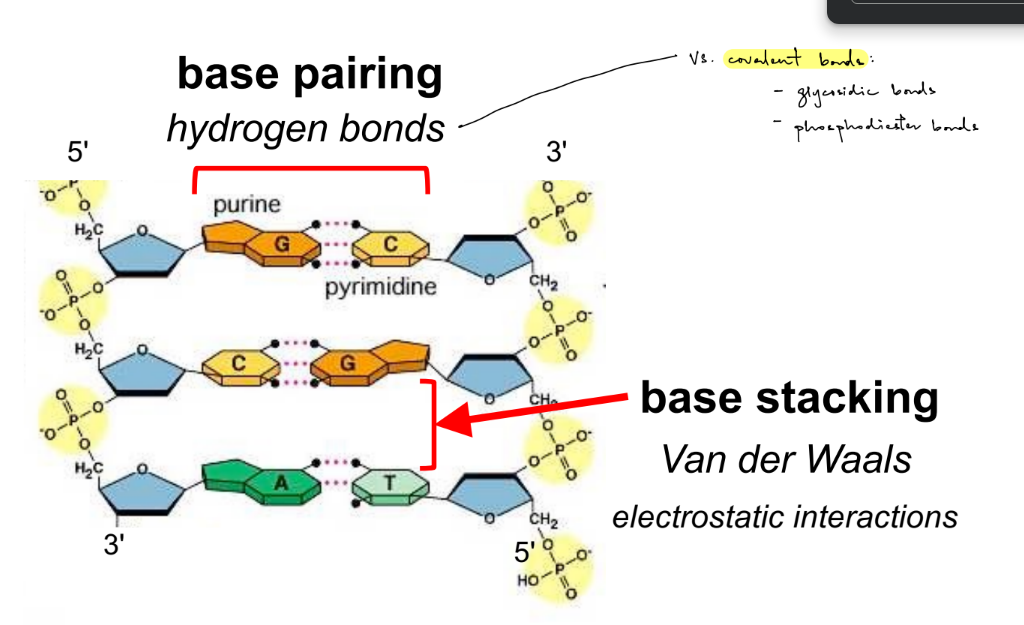
what 2 types of hydrogen bonds are for DNA (aka the molecules that base pair for DNA) vs. 1 type of H bond for proteins
DNA
O — H(N)H
N — H(N)
proteins
O — H(N) ← for alpha helix, beta sheet
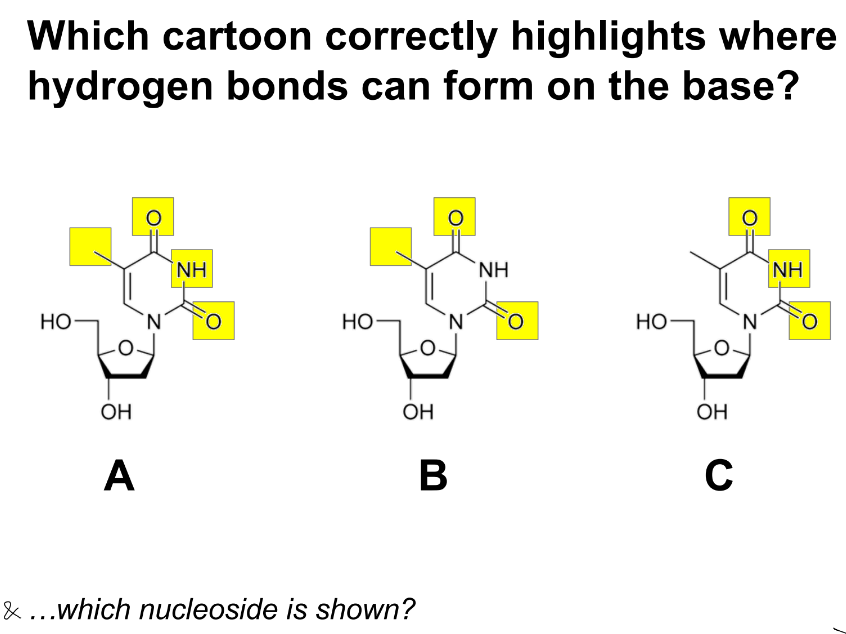
relation b/w H bonds & methyl group (CH3)
C
thymidine (“-side”)
__
hydrogen bonds aren’t usually formed w/ methyl groups (CH3)
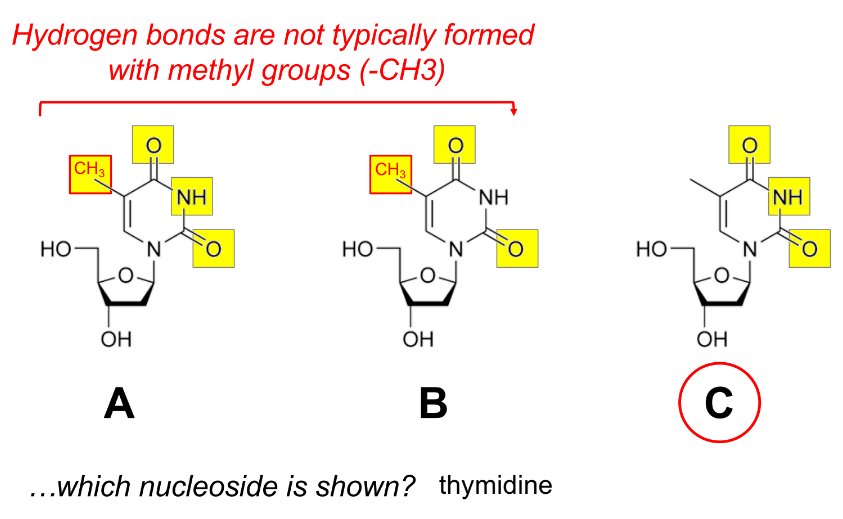
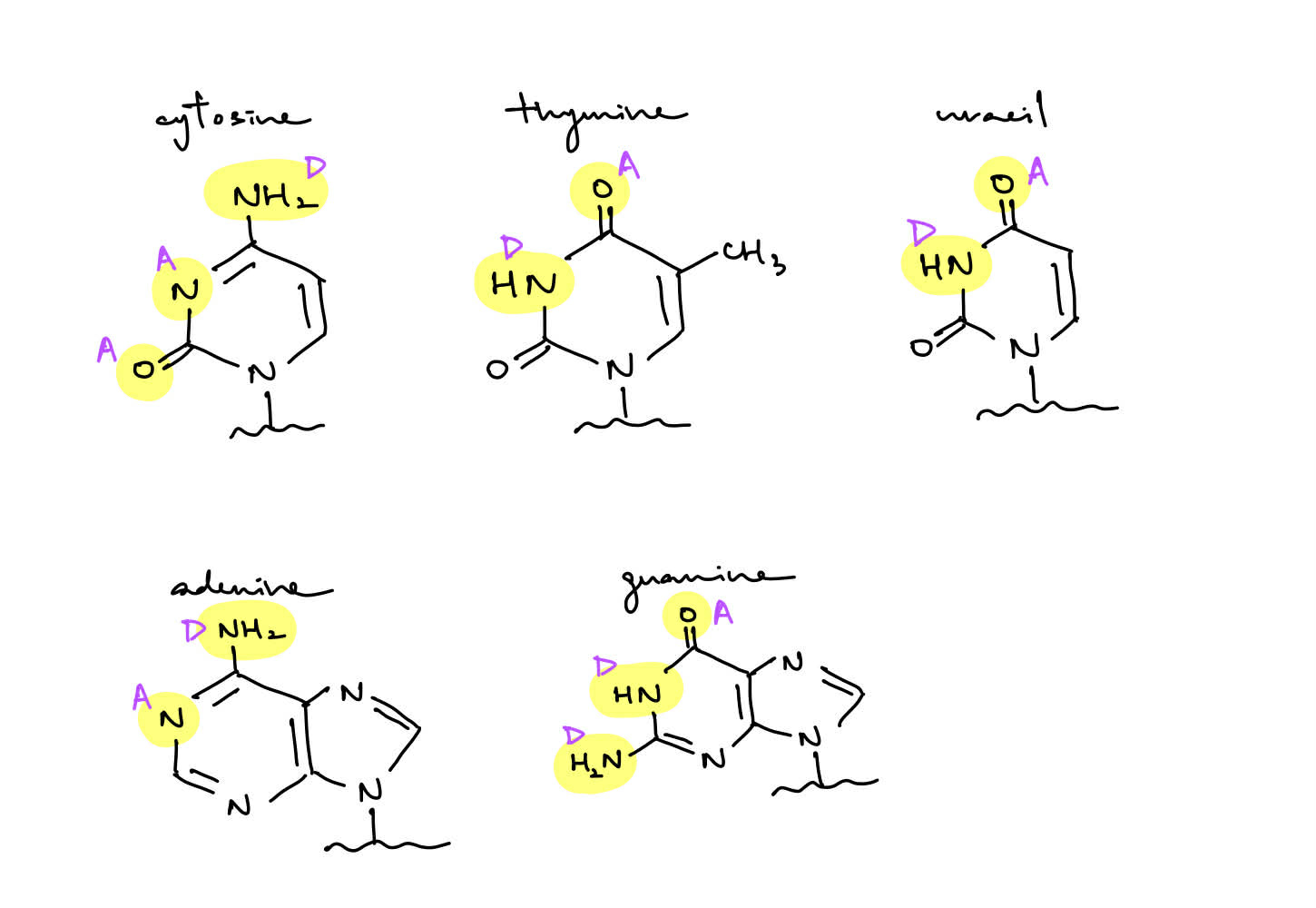
review(ish)
locate hydrogen bond acceptors vs. donors for 5 bases (probably draw it out)
__
t/f: molecules have additional H-bond donors and acceptors that aren’t highlighted by arrows
t/f: base pairs form due to alignment of hydrogen bond donors and acceptors
true
(i.e. the other H in NH2)
_
true
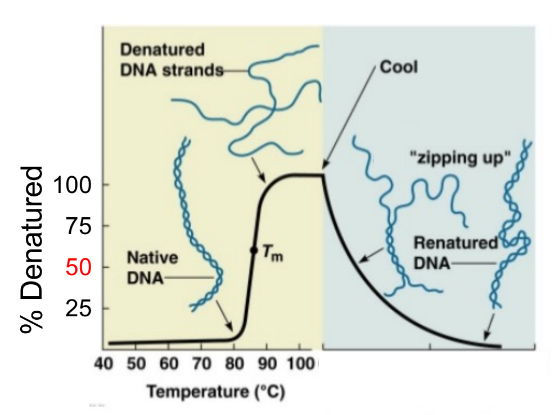
the secondary structure of DNA and RNA, specifically (1), is sensitive to (2 ← specify a bit)
__
def. Tm
what happens as temp increases
relate this to Tm (melting temperature) (2) ← & explain each
what happens as temp decreases
H bonds of secondary structure is sensitive to temperature & pH
high temp & extreme pH (very high or very low pH), disrupt the secondary structure
_
Tm (melting temp)
temperature where the helix is half double-stranded & half single-stranded (aka temp when 50% denatured)
_
as temp increase, the strands of DNA helix will denature/separate
stable DNA helix = high Tm
^ stable helix needs more heat to denature, so high Tm
(denatures 50% at higher temp, so more stable)
unstable DNA helix = low Tm
^ unstable helix needs less heat to denature, so low Tm
(denature 50% at lower temp, so more unstable)
as temp decreases, DNA strands w/ complementary sequences will anneal/zip up
t/f: DNA sequence also affects DNA stability & Tm of DNA (in addition to temp and pH)
explain
true
A/T and C/G have/form different #s of H bonds (2 vs. 3) and base-stacking interactions
where C/G pair have a higher Tm
meaning C/G is more stable at higher temp than A/T which is more unstable at higher temp / more stable at low temp)
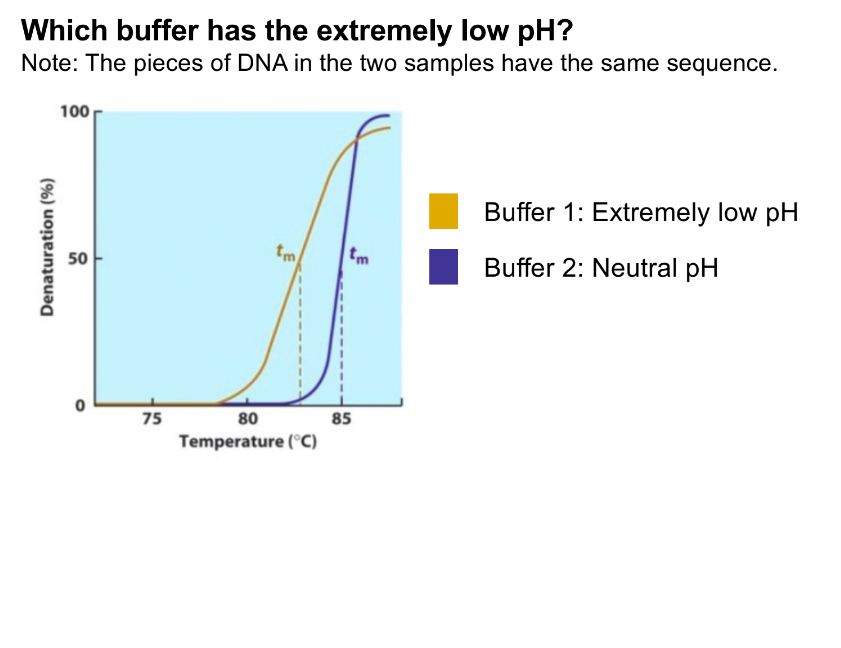
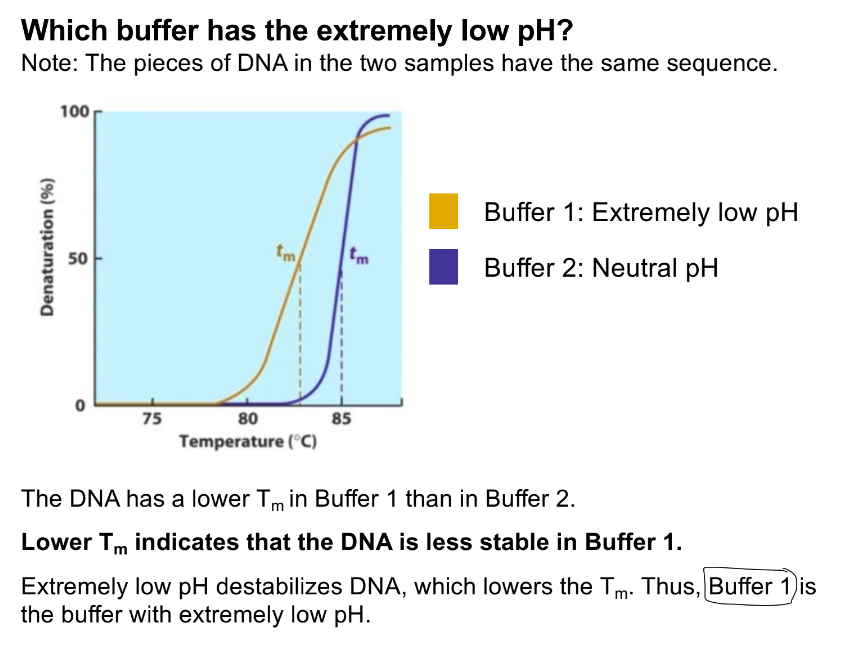
low pH is unstable
unstable helix is low Tm (needs less heat to denature)
buffer 1 has the lower Tm
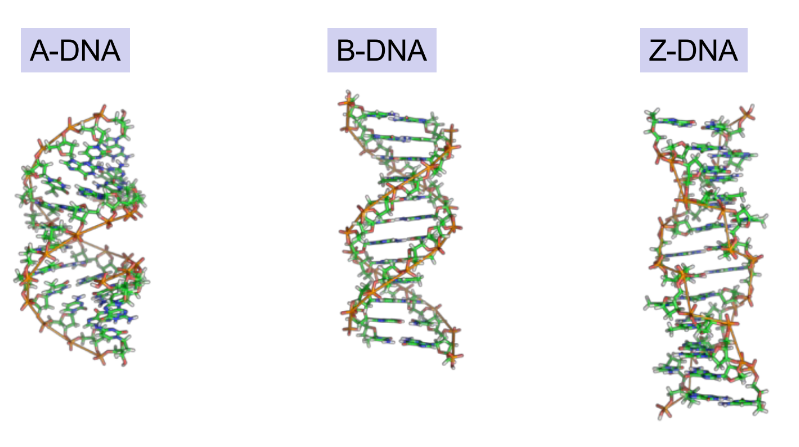
3 conformations/forms of DNA in tertiary structure
which one allows nucleic acid bases & proteins to interact
__
function of quaternary structure
A-, B-, Z-DNA (have diff. configurations)
_
B-DNA conformation allows proteins to bind to specific DNA sequences
this is essential for gene expression
____
quaternary structure of DNA helps in packing genomic DNA in cells
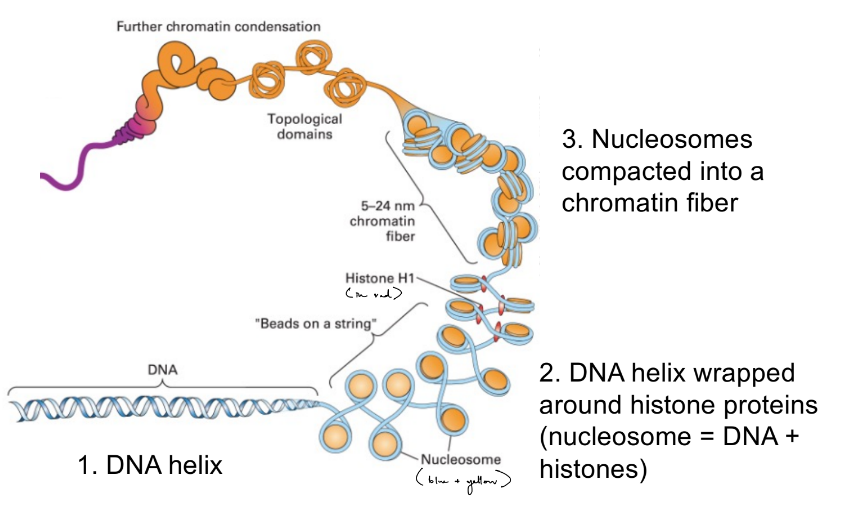
DNA is stored as __ in EUkaryotic cells
describe
what about how DNA is packaged in PROkaryotic cells?
DNA is stored as chromatin in eu- cells
DNA helix
DNA helix + histones (are proteins) = nucleosomes
nucleosomes are compacted into chromatin fiber
__
pro- genomic DNA is supercoiled
review:
state central dogma w/ processes (which process allows genetic info to be transmitted from parent cell to daughter cells via cell division)
genes are encoded in __
DNA → RNA → proteins
replication (of DNA), transcription (DNA ←→ RNA), translation (into protein)
DNA replication allows genetic info to be transmitted from parent cell to daughter cells
__
genes are encoded in DNA
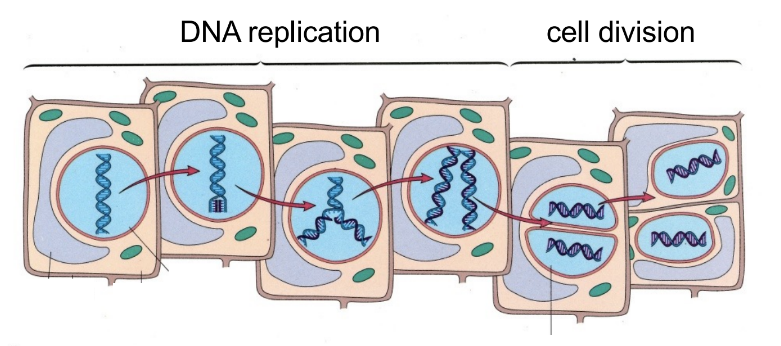
discovery from Meselson-Stahl experiment (1)
what serves as template to make daughter strand?
DNA replication is semi-conservative
after replication, EACH helix has 1 strand of template parent DNA & 1 strand of newly synthesized daughter DNA
parent strand is the template for daughter strand
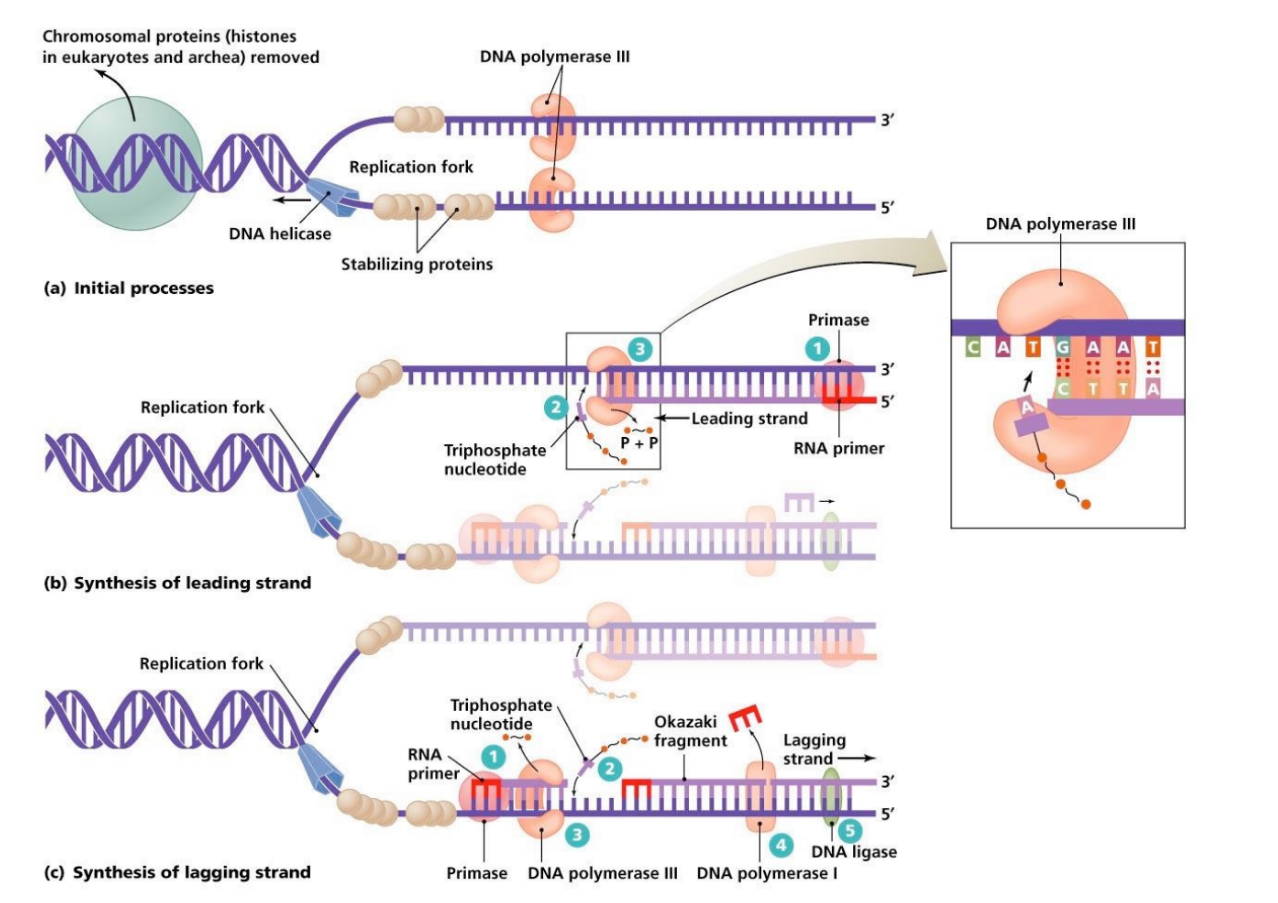
state key steps & enzymes involved in DNA replication (8 steps)
helicase denatures/unwinds DNA
topoisomerase relieves helix strain that occurs when unwinding DNA
RNA primase (aka primase) creates a short RNA primer to start DNA synthesis
DNA polymerase (DNA pol III) adds nucleotides via phosphoryl-transfer rxn
DNA pol “proofreads” via exonuclease activity
DNA pol (DNA pol I) removes RNA primer & replaces it w/ DNA
DNA ligase (aka ligase) joins together the DNA fragments by forming a phosphodiester bond
telomerase acts at the end of chromosomes to maintain their/the chromosome’s length
each human cell has ~__ feet of DNA, which takes ~__ hours for the human cell to replicate its genome b/c during mitosis, the cell copies the entire length of DNA before it divides
where does genome replication start (1)
describe (1)
in pro- vs. eukaryotes
~6 ft of DNA, ~8 hours
__
DNA replication starts at an origin of replication (“ori”)
at an ori, 2 strands of DNA helix are denatured
pro- have single ori
eu- have multiple ori
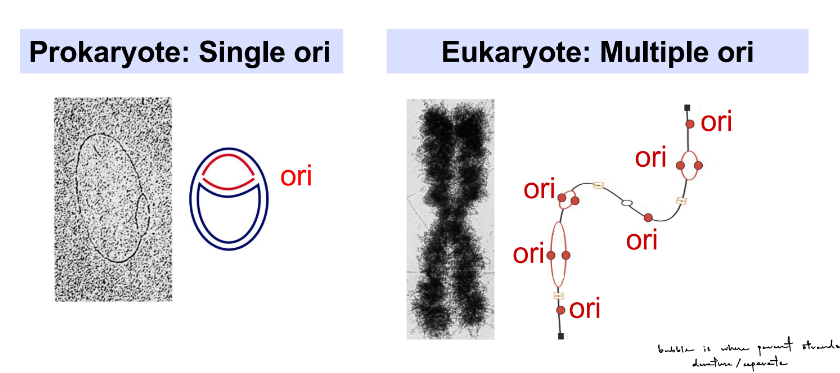
why do eu- have multiple ori?
multiple ori allows for FAST genome replication
(would take ~1 year for human cell to divide if human genome only had 1 origin of replication)
(OVERALL, not only regarding multiple ori, you want DNA replication to have a balance b/w being both fast & accurate)
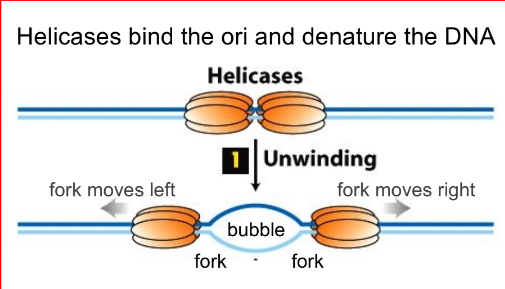
function of helicase (~2), which causes __ that looks like __
direction of movement
forms (1 w/ 1)
helicases bind the ori & denature/unwind DNA by breaking the H-bonds, which causes tension that looks like supercoils
(locally denature DNA at the ori)
move in opposite directions to unwind the helix & separate the strands
forms the “replication bubble”, where there are 2 “replication forks” that move in opposite directions away from the ori
function of topoisomerase (1)
function of SSB (is NOT an enzyme, but a protein)
as helicase unwinds DNA, topoisomerase relieves torsional stress that was caused by strand separation (prevent supercoils from forming)
__
SSB (single-strand binding protein ← not an enzyme)
stabilizes single-stranded DNA (works together w/ topoisomerase in DNA replication)
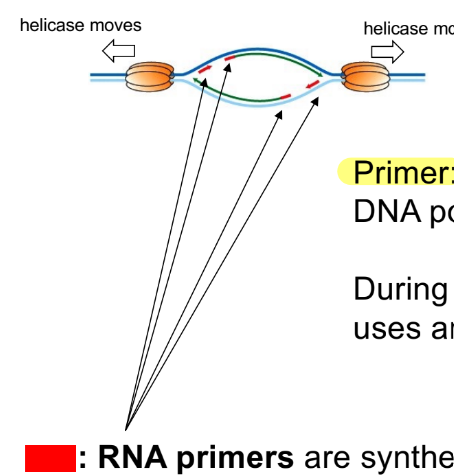
function of primase (← which is a __ __)
def. (RNA) primers
why is it needed?
RNA primase (an RNA polymerase)
makes RNA primers based on DNA template
primers - short piece of DNA or RNA (DNA/RNA primer) that DNA polymerase needs to make DNA
BUT we use RNA primers in genome replication
__
is needed b/c DNA polymerase (next step) CANNOT bind DNA and initiate DNA synthesis w/o a primer
replication forks move in what direction?
move in same direction as helicase moves aka same direction as DNA separating
state function of DNA polymerase, where it adds nucleotides (1 ← long)
daughter strands are made __
__
DNA pol. can only add nucleotides to the (1), so if it is the very beginning of genome replication w/o some synthesized part, what is needed?
explain
is the primer part of the immediate product (not talking abt final product)?
adds the complementary nucleotide to the template base
by forming phosphodiester bond b/w 3’ OH & 5’-phosphate / alpha-phosphate of incoming dNTP (polymerization, via phosphoryl transfer rxn)
^^^ (daughter strands are made simultaneously)
^^^ dNTP as in dATP, dGTP, dCTP, dTTP
__
DNA pol. can ONLY add nucleotides to the 3’ OH at the 3’ end of a primer (at the primer’s 3’ end)
if starting new DNA strand (5’ → 3’), then need primase to form a primer, so that primase can add the first nucleotide to initiate a strand
primer is part of immediate product, located at the 5’ end of the synthesized DNA strand
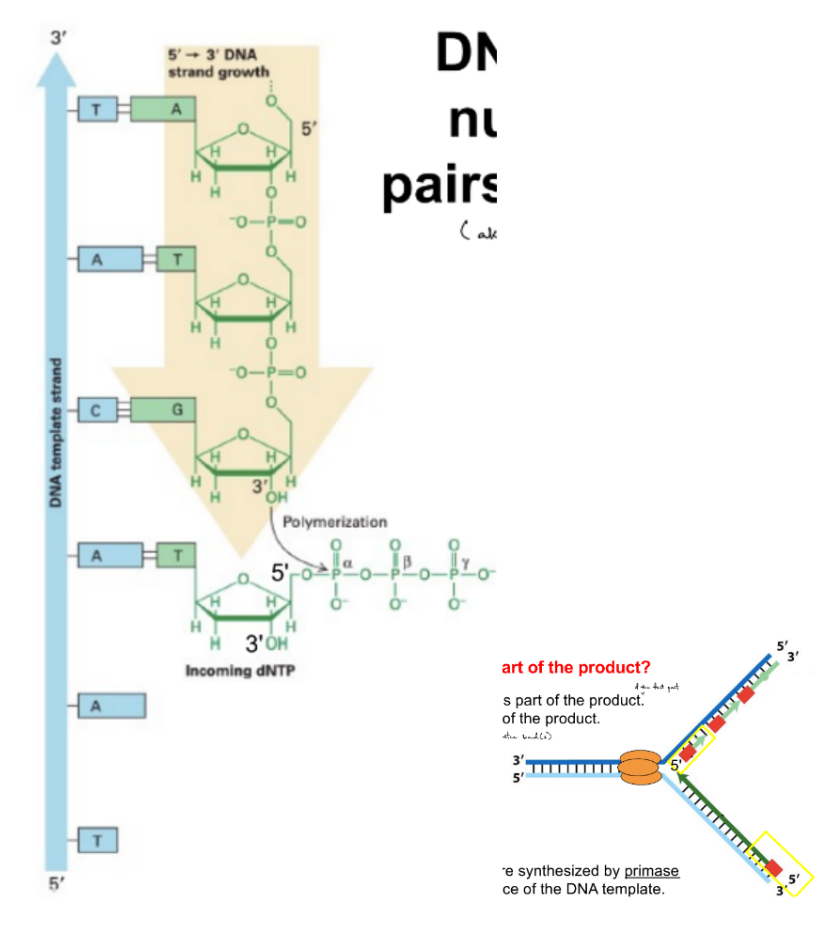
t/f: DNA polymerase can’t initiate synthesis, but (RNA) primase can
true
DNA polymerase can’t initiate synthesis, so it needs RNA primer
def. nucleoside analogs (2)
function ← how?
give example & describe for (viruses):
HIV infections
Covid-19
describe for cancer
compounds that are structurally similar to nucleosides & that are used to combat DNA & RNA viruses and cancer
block replication of viral genomes
nucleoside analog is incorporated into the strand during DNA (or RNA) synthesis, but the analog stops further synthesis which prevents viral replication
__
ex:
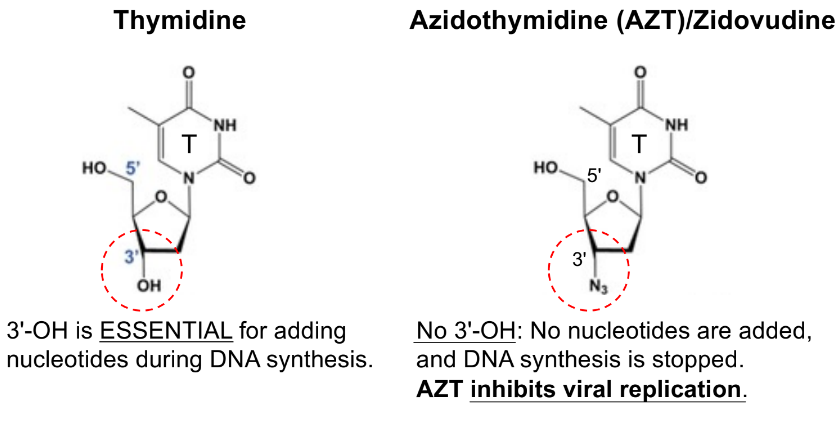
AZT treats/combats HIV infections, where AZT is an analog of the nucleoside thymidine
b/c thymidine has 3’ OH for adding nucleotides during DNA synthesis, but AZT does NOT have 3’ OH so no nucleotides added → DNA synthesis stops
AZT inhibits viral replication
ex:
Remdesivir combats Covid-19, where Remdesivir is an adenosine analog
__
slow mitosis & proliferation of cancerous cells
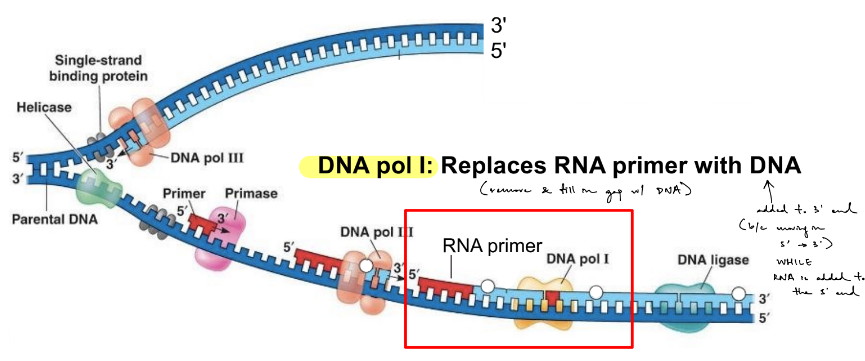
how are RNA primers removed & replaced? (& by what?)
DNA polymerase 1 (DNA pol I) removes RNA primers & fills in the gap w/ DNA
(aka simultaneously removes the primer from the 5’ end of newly synthesized strand & replaces it w/ DNA)
(DNA pol binds at the 3’ end of the RNA primer & moves in the direction of 5’ → 3’)
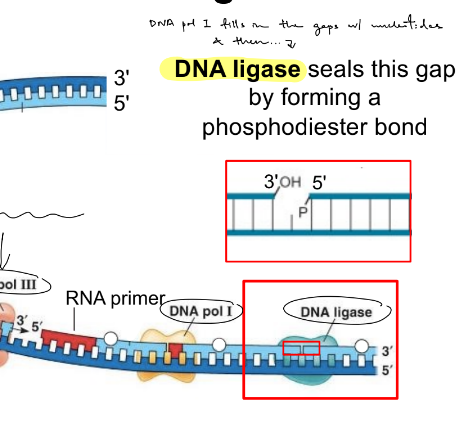
function of DNA ligase
seals the gap by forming a phosphodiester bond
(the gap is b/w where DNA pol filled in the gap & part of synthesized strand)
(aka when the primer is replaced by DNA, a new piece of DNA is made BUT that piece is not attached to the neighboring DNA, which creates a gap b/w the 3’ & 5’ ends of neighboring pieces)
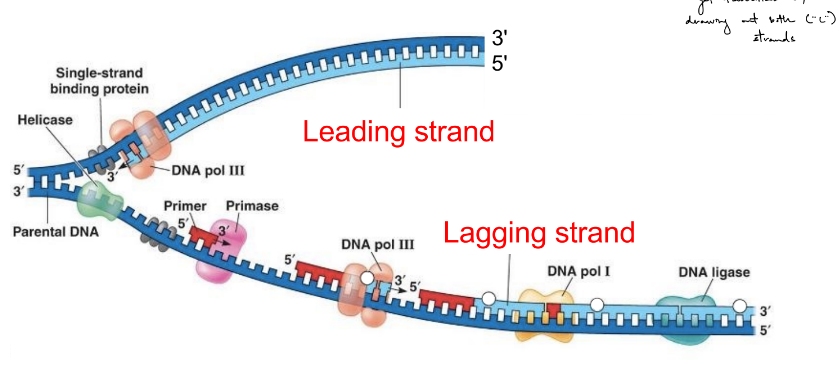
name these 2 synthesized daughter strands that: (& def. ← 2 each)
during genome replication, the 2 synthesized daughter strands (that are anti-parallel) in the 5’ → 3’ direction (b/c only direction DNA pol. moves in)
^ these 2 daughter stranded are synthesized __
leading strand (aka leading daughter strand)
is made in 1 piece, using only 1 (RNA) primer
made in the same direction that the replication fork moves
lagging strand (aka lagging daughter strand)
is made in pieces (Okazaki fragments), using multiple (RNA) primers
made in the opposite direction of the replication fork
__
simultaneously
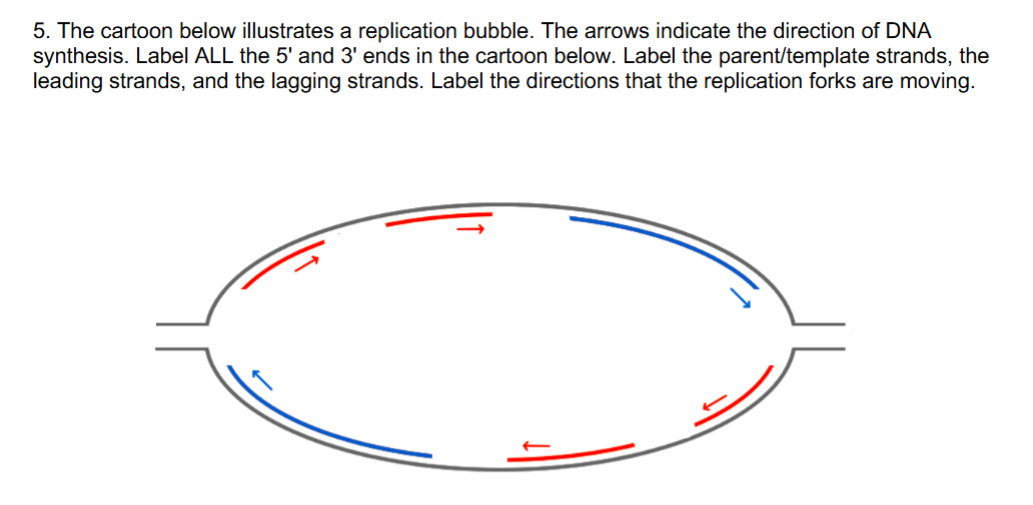
also, which part would be the chromosome end (aka __)
telomere (aka chromosome end) is the end where the 2 parent strands haven’t unwound/separated/denatured yet (aka where parent strands are still together/intact)
for the problem in pic:
you can identify the 5’ and 3’ end from the direction of the synthesized DNA daughter strands that move in 5’ → 3’ direction, SO the ends of parent strands would the opposite of the synthesized strands (b/c anti-parallel)
you can identify the leading vs. lagging strand by splitting the diagram in half vertically
the blue move in same direction as fork
the red move in opp. direction from fork
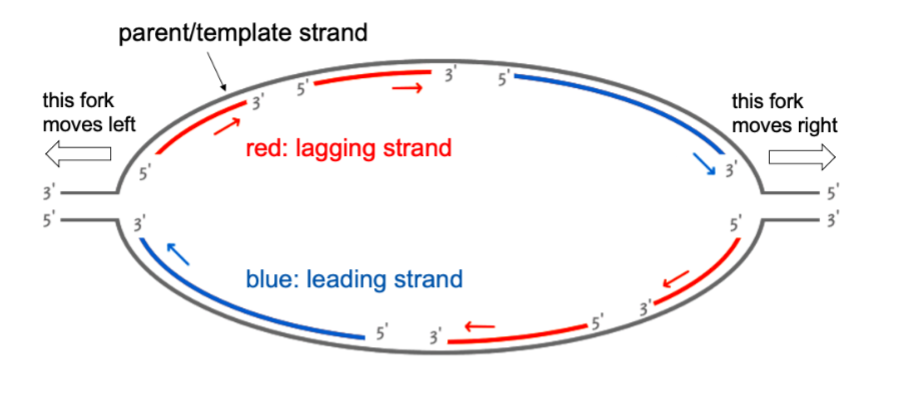
at the ends of linear chromosomes (aka at telomeres), what happens if nothing after ligase happens & you continue DNA replication?
there is a gap (b/w 3’ and 5’ of neighboring ends), but (the leading strand has no gap)
for lagging strand synthesis:
primase adds an RNA primer at 5’ end of chromosome
when primer is removed, there’s a gap at the end of the new chromosome
ligase joins the 2 Okazaki fragments w/ phosphodiester bonds
there is a gap at the end of the telomere WHERE chromosomes get shorter each time IF you don’t fill in the gap and just continue genome/DNA replication
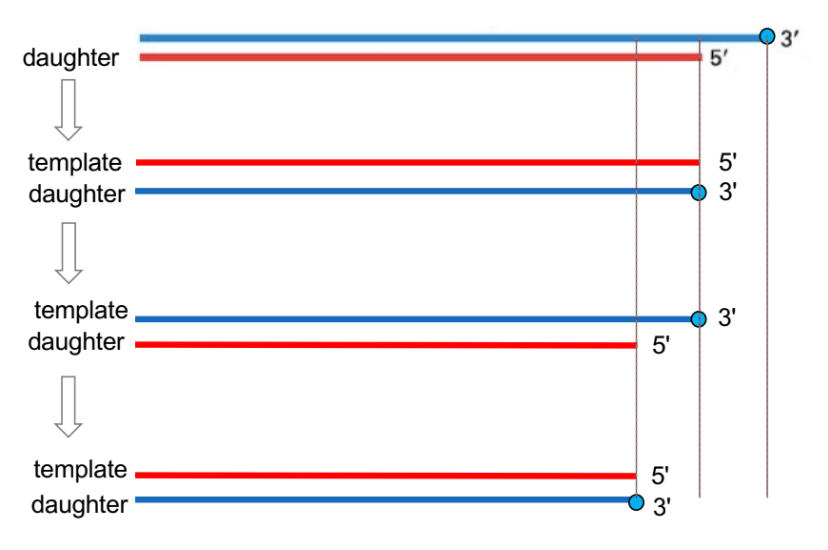
function of telomere (1), which is a __ polymerase
telomerase is a __ __ (← term relates to transcription)
_
after telomerase performs its role, what happens to the lagging daughter strand?
telomerase (a DNA polymerase)
extends the 3’ end of the parent strand, using a RNA template, to maintain chromosome length
telomerase is a reverse transcriptase
(b/c makes DNA based on an RNA template, where central dogma is DNA → RNA → protein)
_
then, the lagging daughter strand extends via primase + DNA pol + ligase using this newly synthesized parent strand as the template
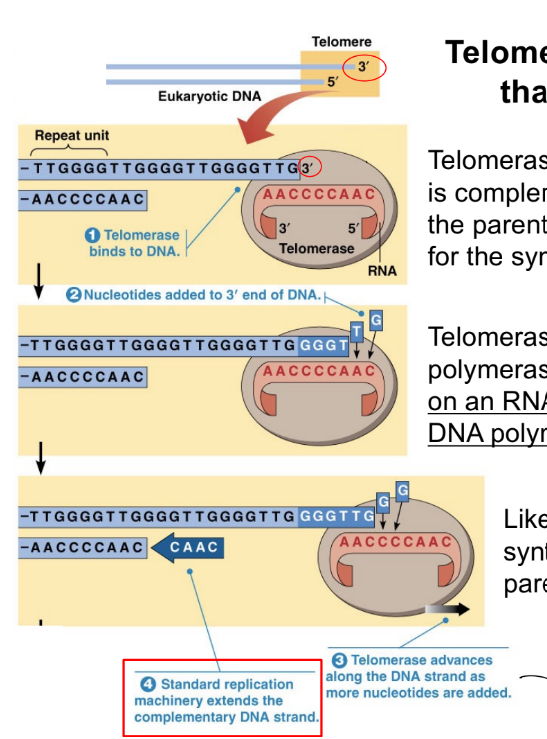
multiple ori allows for fast genome replication, but this can lead to (1)
you want …
fast replication can lead to mistakes in DNA replication, like mismatches
instead, you want a balance b/w replicating a genome quickly & accurately (i.e. making only 2-3 mistakes)
def. mutation
can be caused by (1) or (1)
can be either (3)
ANY change in DNA sequence of a cell
caused by mistakes during cell division (replication) OR exposure to DNA-damaging agents in the environment
can be harmful, beneficial, or have no effect (neutral)
just state the 2 mechanisms of repairing mismatches during DNA replication (1 of them also has a specific term)
which is immediate vs. not
exonuclease activity by DNA polymerase (“proofreading”)
immediate / during replication
mismatch repair
not immediate / after replication is done
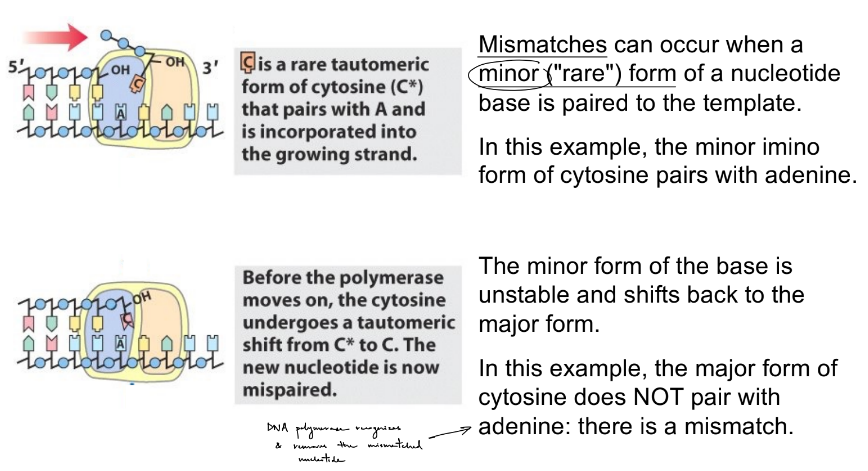
t/f: different forms of nucleotide bases can result in mismatches (“errors”) during DNA synthesis
explain generally
then, explain w/ ex. of cytosine
how do these spontaneous ““ shifts in the base occur? they can cause more __ nucleotides
true
nucleotide bases can exist in 2 forms called tautomers (aka shifting of protons), where 1 form is more stable than the other: predominant, major vs. minor form
the minor and major forms have different patterns of H bond donors and acceptors
__
cytosine has a major & minor form
the minor form of C can pair w/ A (adenine), but the major form cannot & only binds to G (guanine)
this C/A can lead to a change in the DNA sequence
_
these spontaneous proton shifts in the base can cause more mismatched nucleotides
occur by:
minor form is paired to the template (w/ the complementary nucleotide base)
before DNA polymerase continues to add nucleotides after this, the minor base changes back to the major form B/C the minor form is unstable
this would be a spontaneous mismatch

what are the 2 enzyme activities of DNA pol? what bond is affected?
specific term for 1 of them
(^^ this question is not completely referring to how to repair mismatch)
polymerization, where DNA pol adds nucleotides via polymerase domain (Pol)
exonuclease activity, where DNA pol removes mismatched nucleotides via exonuclease domain (Exo)
“proofreading”
^ both affect phosphodiester bonds (make vs. break, respectively)
(can refer to past flashcard for pic of polymerization, via phosphoryl-transfer rxn)
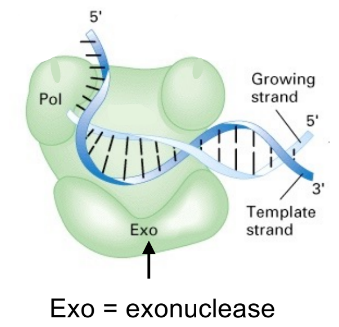
DNA pol adds/removes nucleotides to/from the __ end
3’ end
(i.e. DNA pol adds nucleotide at the 3’ OH of prior neighboring sequence, in the direction of 5’ to 3’)
DNA polymerase is what type of exonuclease & what type of polymerase?
function of both
DNA polymerase I is what type of exonuclease & what type of polymerase?
function of both
DNA pol
5’-to-3’ polymerase (adds DNA)
start at 3’ end
3’-to-5’ exonuclease / 3’ exonuclease (“proofreading” — remove incorrectly added bases, like mismatch and insertion/deletion)
starts acting at the 3’ end
DNA pol I
5’-to-3’ polymerase (adds DNA)
start at 3’ end
5’-to-3’ exonuclease / 5’ exonuclease (remove RNA primer)
starts acting at the 5’ end
(the green and purple ovals are the Exo / exonuclease domain)
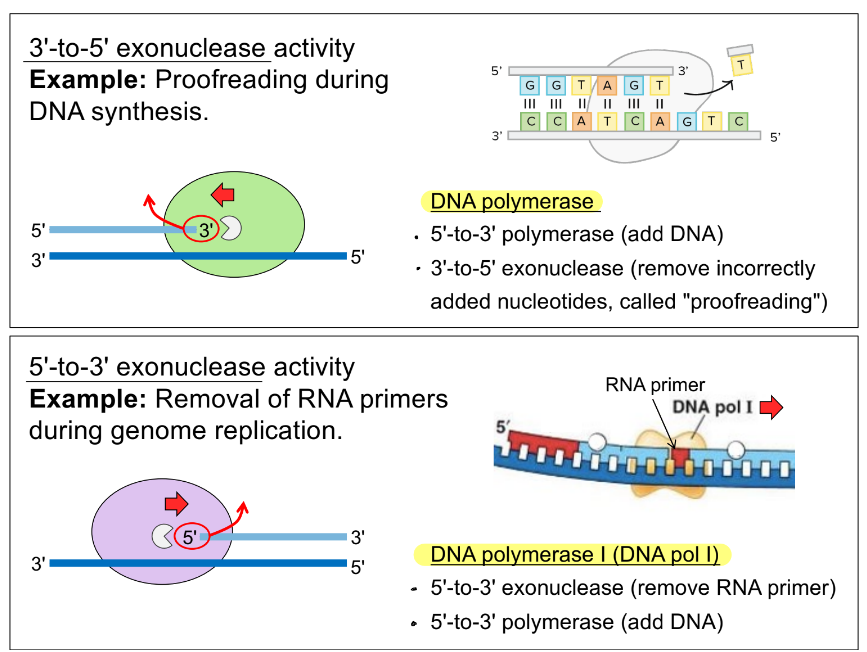
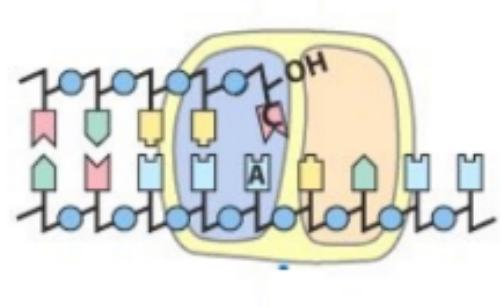
how would DNA pol recognize & remove mismatched nucleotides, based on the picture
DNA pol recognize mismatched base b/c 3’ OH of cytosine is not positioned properly
DNA pol then uses its 3’-to-5’ exonuclease activity to remove the mismatched base
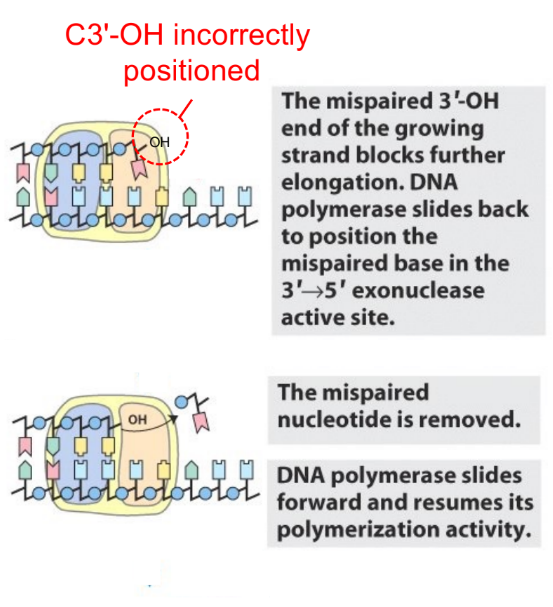
exonuclease activity by DNA pol starts when there’s a spontaneous tautomeric shift that stops continuation of DNA synthesis,
so why doesn’t mismatch repair stop DNA synthesis when there’s a mismatch?
DNA synthesis continues b/c the mismatched nucleotide was not recognized
(mismatch repair occurs after DNA synthesis is done b/c there’s no proofreading)
while exonucl. activity by DNA pol is proofreading that occurs during DNA synthesis
exonuclease activity/proofreading occurs during DNA synthesis
mismatch repair (MMR) occurs after end of DNA synthesis
BER and NER occurs after the end of DNA replication
name & def. 2 mismatch repair enzymes
1 of them can act __ or __
_
review(ish):
another name for each (& function of each):
3’-to-5’ exonuclease
5’-to-3’ exonuclease
endonuclease — cleaves phosphodiester bonds w/in a strand
can act EITHER upstream (at 5’ end) OR downstream (at 3’ end) of the mismatch (cleaving ONE phosphodiester bond in mismatch repair)
exonuclease — cleaves phosphodiester bonds at the end of a DNA strand
EITHER upstream OR downstream
(^^ both cleave phosphodiester bonds, just at different positions in a DNA strand)
__
3’-to-5’ exonuclease (aka 3’ exonuclease) (aka 3’-5’ ““)
proofreading / remove mismatch
5’-to-3’ exonuclease (aka 5’ exonuclease) (aka 5’-3’ ““)
remove RNA primer
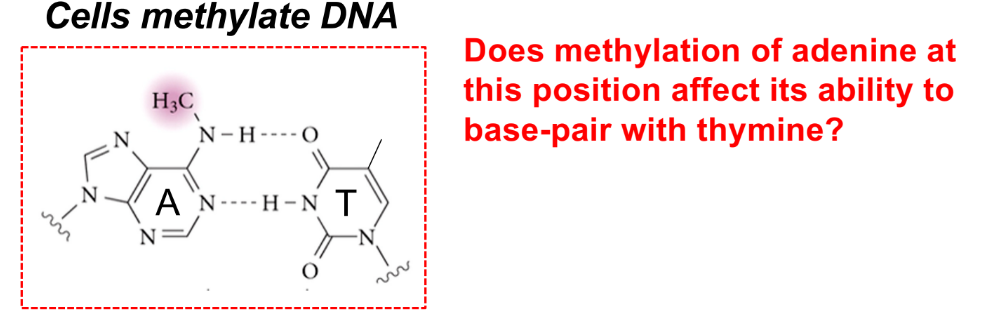
which DNA strand do endonucleases cleave?
how do they know?
why is it like this?
mismatch repair machinery cleaves the non-methylated DNA
the mismatch repair machinery can identify the parent strand b/c it is methylated (CH3), while the daughter strand is not methylated
over time, genomic DNA accumulates methylation (parent strand is old, while daughter strand is new)
__
picture:
No b/c methylation at this position doesn’t disrupt the hydrogen bonds that form b/w adenine & thymine (A-T base pair is intact)
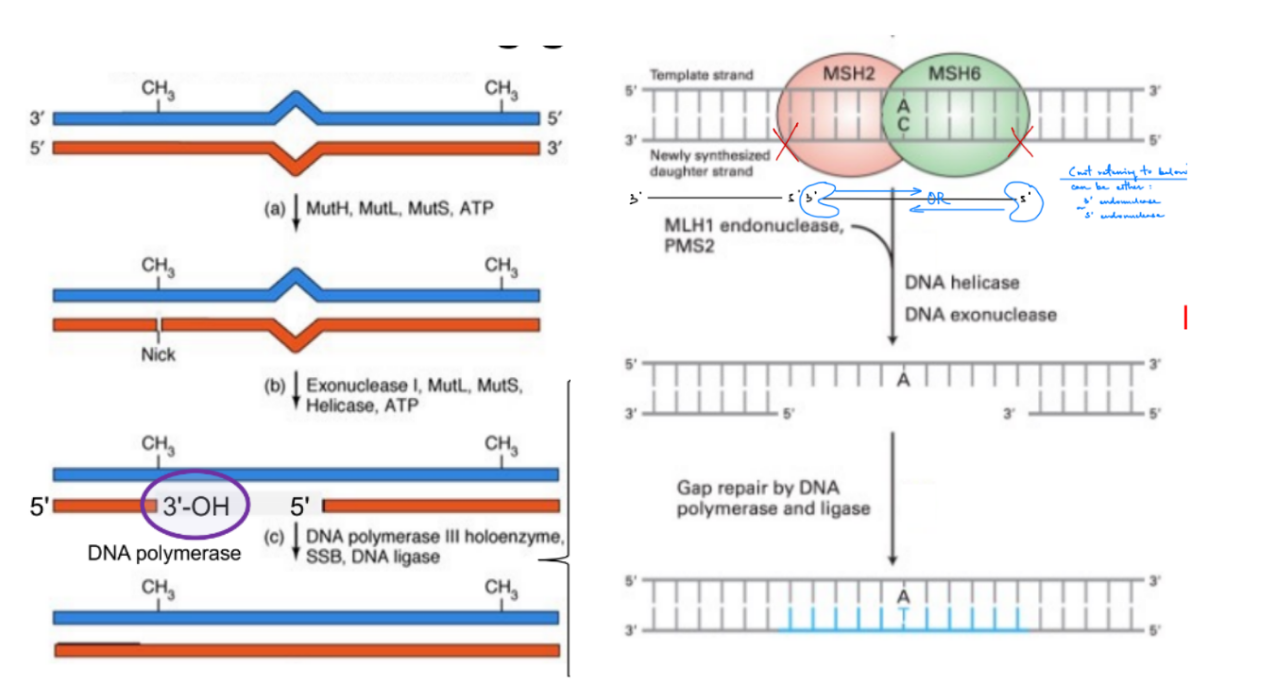
how do they both work together in mismatch repair:
endonuclease, exonuclease, mismatch repair machinery, helicase, ligase, DNA pol (← is stated out of order)
^^ aka steps of mismatch repair (5)
_
& include which specific exonuclease is used depending on relative positions of two specific things (← 2)
repair machinery/proteins recognize the mismatch & identify the unmethylated daughter strand
endonuc. makes a “nick”/cut ONE phosphodiester bond near the mismatch
(nicks forms “near” aka EITHER upstream OR downstream of the mismatch, but not “at” the mismatch)
helicase unwinds the DNA for exonuclease to access the DNA
exonuc. removes nucleotides past the mismatch (aka including and after the mismatch), EITHER - OR -:
5’ (5’ to 3’) exonuclease IF “nick” is 5’ / upstream from the mismatch
so moving from nick to mismatch is downstream (5’→3’)
3’ (3’ to 5’) exonuclease IF “nick” is 3’ / downstream from the mismatch
so moving from nick to mismatch is upstream (3’→5’)
DNA pol synthesizes new NDA
ligase joins the 3’ OH & 5’ phosphate of neighboring nucleotide w/in the same strand (fills in the gap) in newly synthesized daughter strand
^^ endonuclease can cleave either upstream (5’ → 3’) or downstream (3’ → 5’)
name 2 of various ways that double-stranded breaks (DSBs) occur
_
name 2 ways that double-stranded DNA breaks (DSBs) can be repaired
which is error-prone vs. precise repair? why?
exposure to environmental mutagens
deliberate breaks caused during gene editing (aka gene editing machinery) (will go over in later unit)
__
double-stranded DNA breaks can be repaired via:
end-joining (EJ)
error-prone (b/c mutations are introduced a lot at the break)
homology-directed repair (HDR), such as homologous recombination
precise repair (b/c homologous sequences act as the template for precise repair of the break)
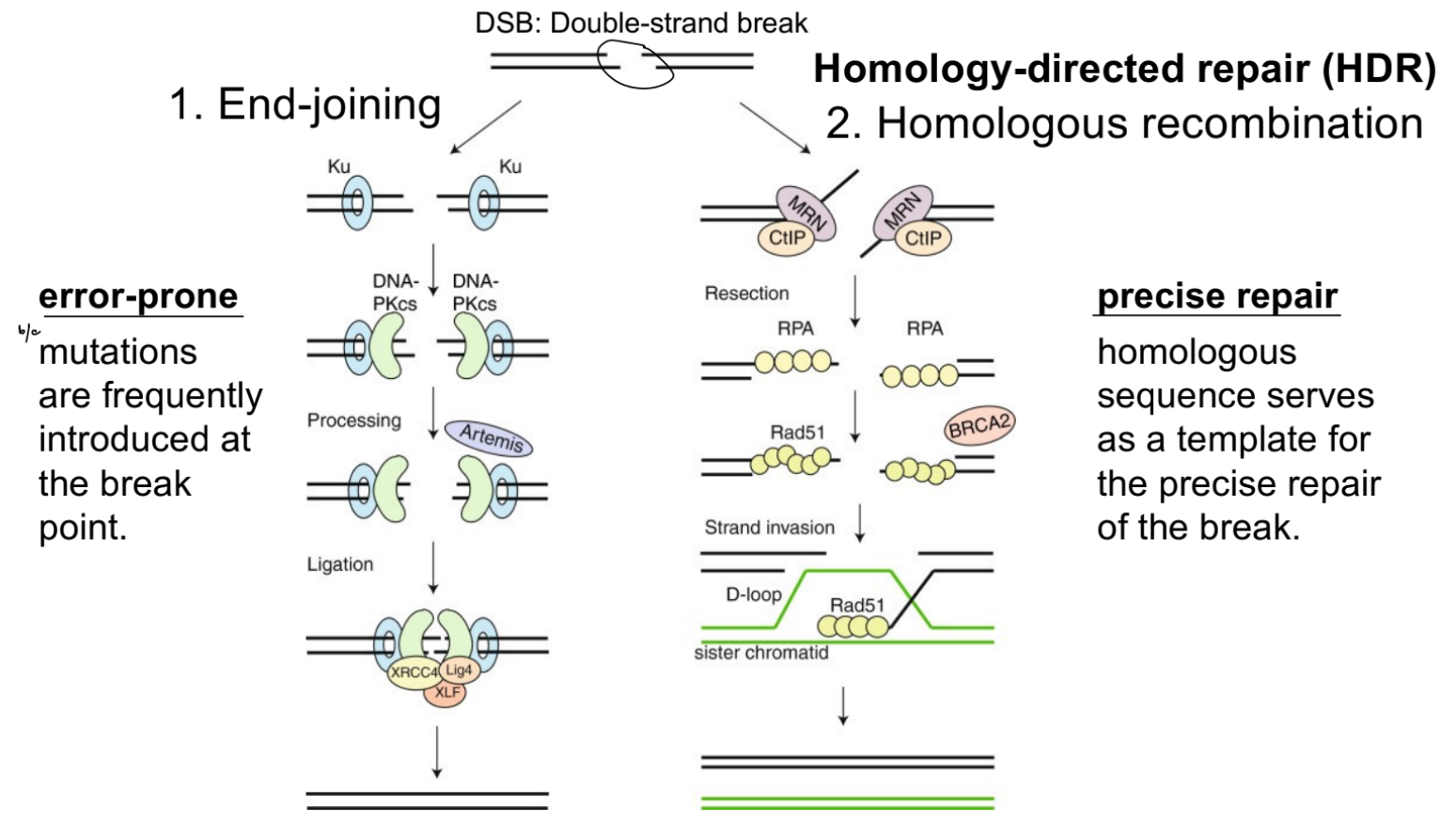
briefly state what happens in end-joining vs. homologous recombination (1 each)
& again, explain why it’s error-prone or precise repair
end-joining
broken ends are joined together
^^ during EJ, nucleotides might be added or removed around the break → changing the DNA sequence
__
homologous recombination
new DNA is made to repair the break
^^ a similar/homologous sequence acts as the template sequence for repairing the break

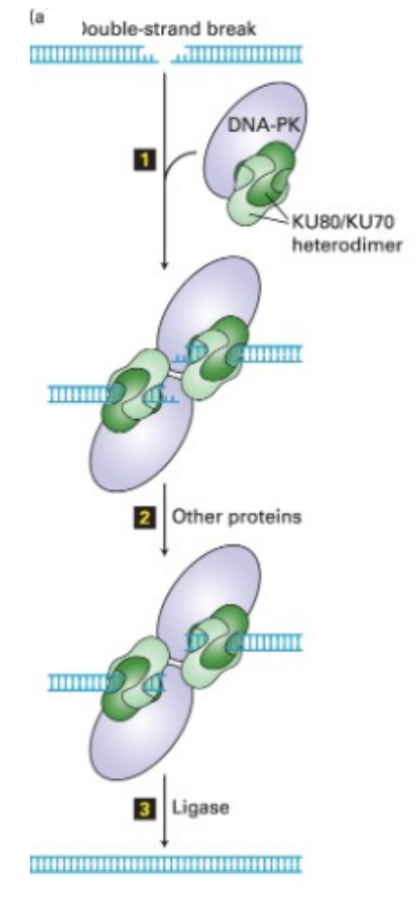
explain the process of end-joining (for double-stranded break repair) (3)
a protein complex (Ku + DNA-PK) binds to the broken ends & bring the ends together
broken ends are modified (in varied ways) that change the DNA sequence
(aka usually have mutations at the site of the break)
DNA ligase makes sure that the broken DNA is re-sealed
__
^^ the modifications are variable (i.e. exonuclease often removes a variable # of nucleotides from the ends) → SO variations in end-repair
briefly describe the 2 DNA helices involved in homologous recombination
__
where can a cell find a homologous sequence to repair the DSB? (2 w/ explanation)
a DNA helix w/ 2 broken strands
a DNA helix w/ a homologous sequence to the double-stranded broken helix
__
find homo- sequence via:
sister chromatid — human cells are diploid, meaning they have 2 copies of each chromosome (1 from each parent) SO chromosome pairs are homologous chromosomes
exogenous DNA — DNA w/ homologous sequence can be injected into cells during gene editing

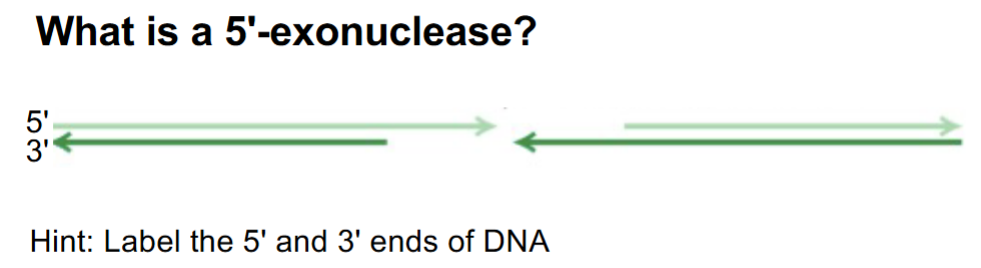
answer the Q
& what does it result in/create? (1)
draw the 3’/5’ ends & the 5’ exonuclease with arrows of direction its moving in
5’ exonuclease — removes nucleotides from the 5’ end of one strand
SO that the 3’ end of the opposite strand is single-stranded → 3’ overhang
(red shows 3’ overhangs)
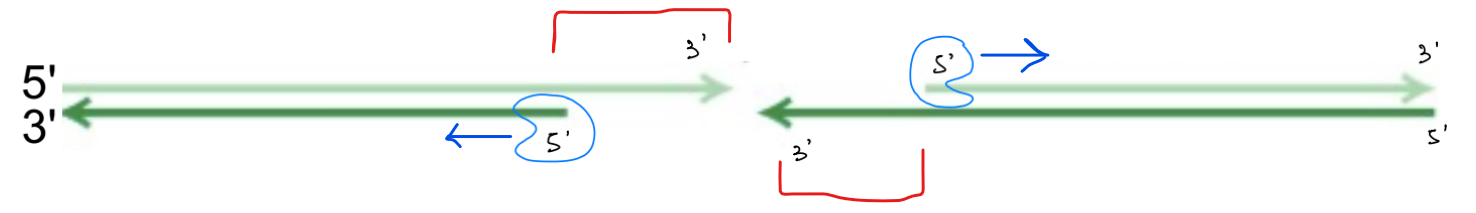
explain the process/steps of homologous recombination (for double-stranded break repair) (6)
in the helix w/ DSB, 5’ exonuclease cleaves the nucleotides at the 5’ end of one strand → make 3’ single-stranded ends on the opposite strand aka 3’ overhang
3’ overhang invades the helix w/ the homologous sequence
DNA pol. extends the 3’ end of this invading strand UNTIL the displaced strand (of homo- sequence) base pairs w/ the OTHER 3’ overhang (to cover the break)
(3’ overhang acts a primer for DNA pol., except it is part of the DNA sequence)
DNA pol. extends this other 3’ end (to cover the break)
DNA ligase seals the two gaps → to form Holliday structures (“X”)
the 2 Holliday structures are resolved → get repaired DNA
(in 2 ways: cut → non-crossover & cut → crossover) ← don’t have to know
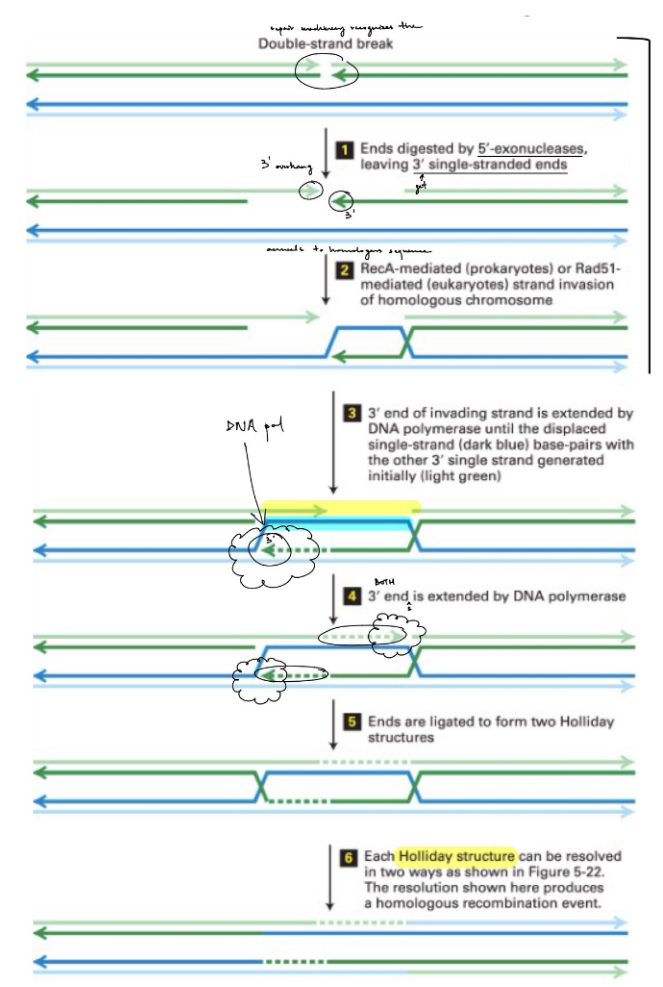
why is it important to form the 3’ overhangs (1) & explain
the 3’ overhang brings in DNA polymerase
b/c the 3’ overhang has a free 3’ OH, that DNA pol can add nucleotides to (b/c DNA pol starts adding nucleotides at the 3’ end)
^ this is also why you don’t need RNA primase in homologous recombination
review:
which specific exonuclease is used depending on relative positions of nick & mismatch (← 2)
__
review:
which removes single mismatch vs. large nucleotide sequence that can contain multiple mismatches: exonuclease activity via DNA pol OR mismatch repair
5’ (5’ to 3’) exonuclease IF “nick” is 5’ / upstream from the mismatch
so moving from nick to mismatch is downstream (5’→3’)
3’ (3’ to 5’) exonuclease IF “nick” is 3’ / downstream from the mismatch
so moving from nick to mismatch is upstream (3’→5’)
^^ just depends on relative position of nick and mismatch
can act EITHER upstream or downstream of the nick, but not in both directions at the same time
__
removes single mismatch — exonuclease activity via DNA poly
(b/c acts once it recognize a mismatch during replication)
large nucleotide sequence that can contain multiple mismatches — mismatch repair
(b/c acts after replication is over, so can possibly have multiple mismatches that are removed by the exonuclease)
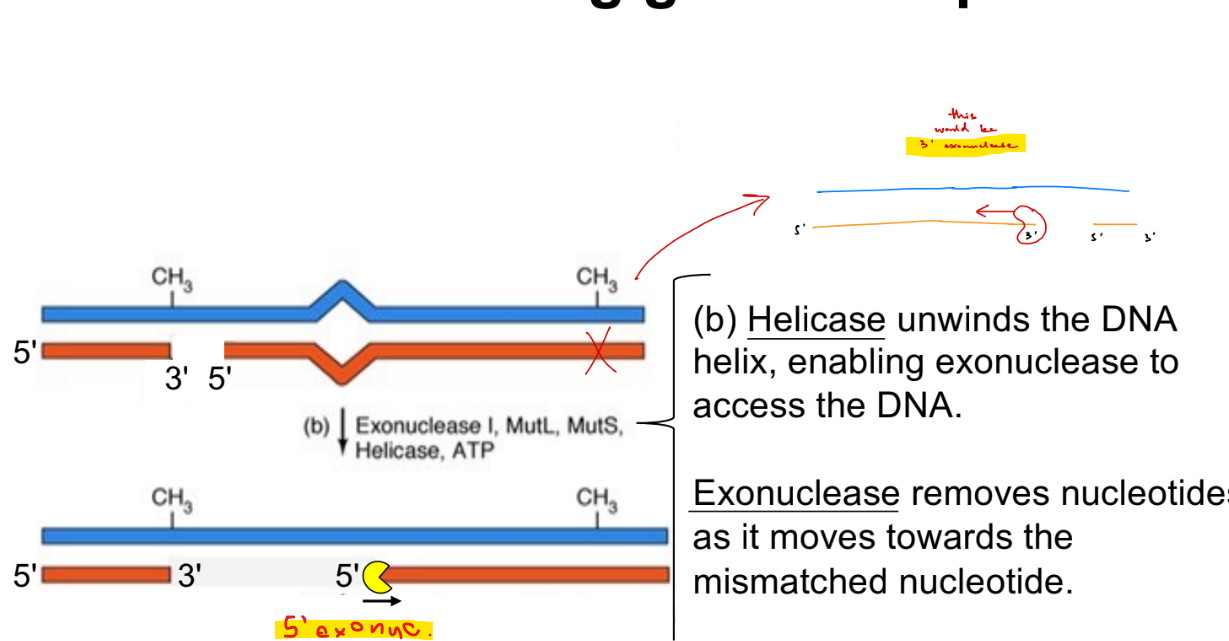
what 2 enzymes catalyze the formation of phosphodiester bonds?
difference b/w the 2 things that the bond forms between
DNA ligase
b/w 3’ OH and 5’ phosphate w/in a strand (that already exist, which is why they’re w/in the strand) of 2 different nucleotides w/in a strand
DNA polymerase
b/w 3’ OH & 5’ phosphate of incoming dNTP of 2 different nucleotides w/in a strand
t/f: primase (a type of RNA polymerase) can initiate DNA synthesis de novo
what does “de novo” mean?
true
“de novo” — from the beginning
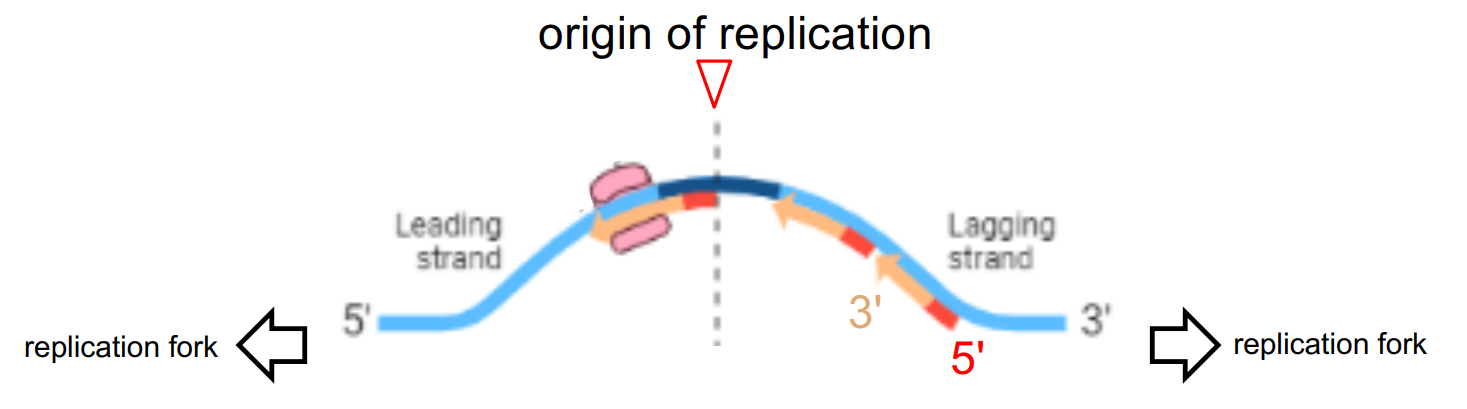
just notice that:
one side of ori is the synthesized leading strand, other side of ori is the synthesized lagging strand
DNA always synthesized from 5’ → 3’
leading or lagging strand is deter. by its synthesis relative to the direction the replication fork moves in
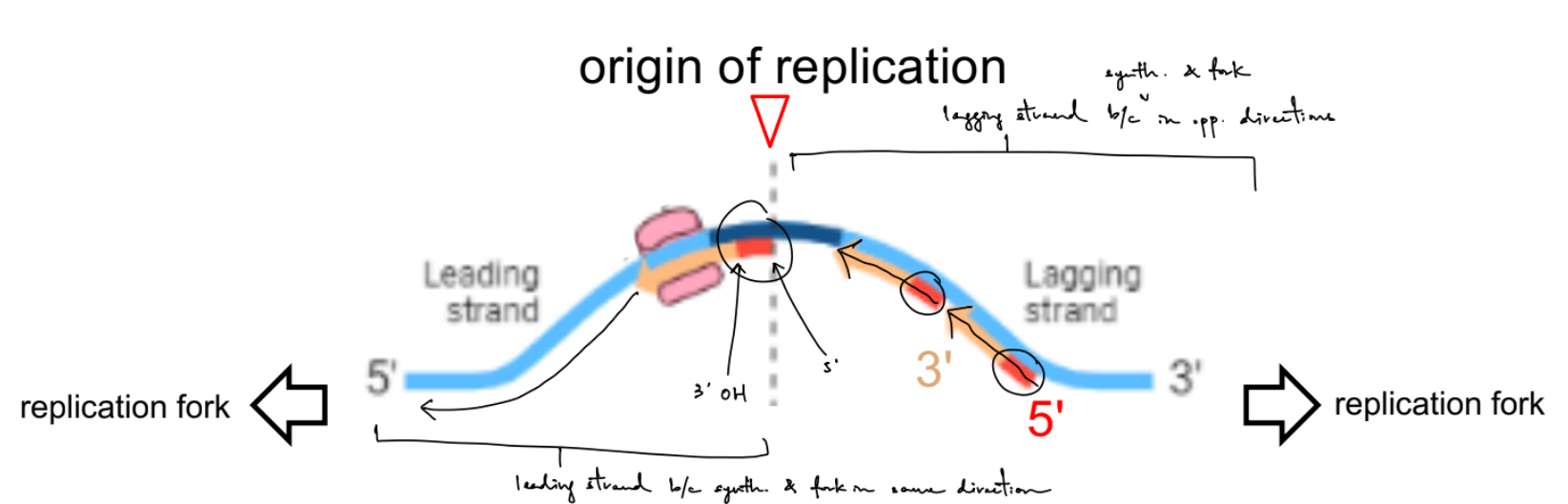
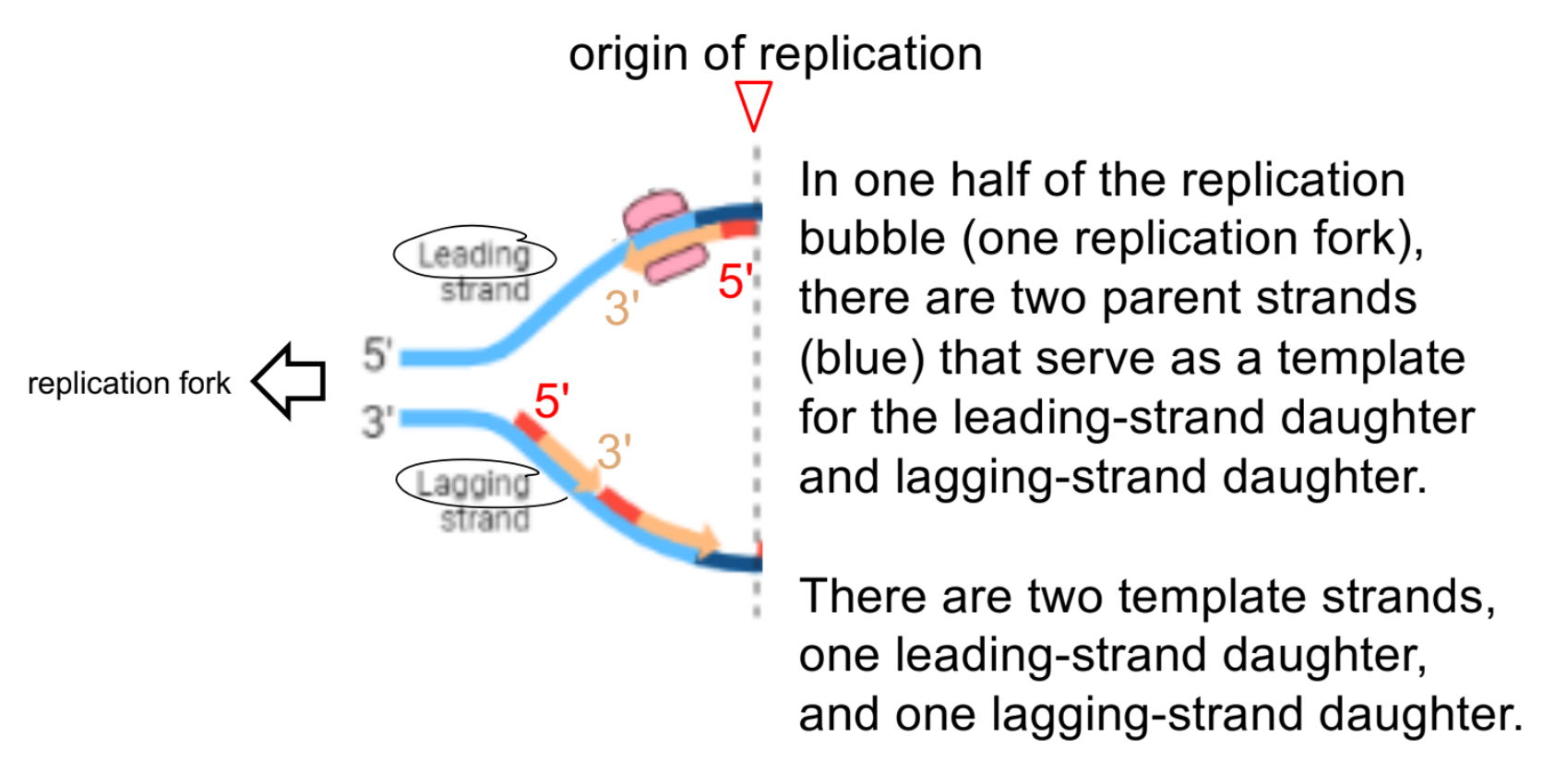
just notice that:
name & def. 3 types of mutagenic chemical modifications that affect the DNA base
(^ aka changes to nucleotide bases can cause mutations if not repaired via DNA repair mechanisms)
alkylation — addition of methyl or ethyl that disrupts a H-bond from forming (can’t form H bond aka disrupts base pairing)
(methylation w/ CH3 or ethylation w/ CH2CH3)
deamination — remove amine group & replace w/ carbonyl group
(replace “—NH2” w/ “=O”)
depurination — remove the entire purine base
is methylation of guanine O6 mutagenic? (may help to draw picture)
what happens to base pairing if methylated guanine O6?
when is methylation mutagenic?
_
diff. b/w methylation & alkylation (revew-ish)
yes b/c prevents it from being a H-bond acceptor
instead, O6-methylguanine pairs with thymine (G6-me-T, instead of G-C)
__
methylation is mutagenic if it occurs at a position that disrupts hydrogen bonds b/w standard base pairs
__
methylation — only adds a methyl group
alkylation — can add either a methyl or ethyl group
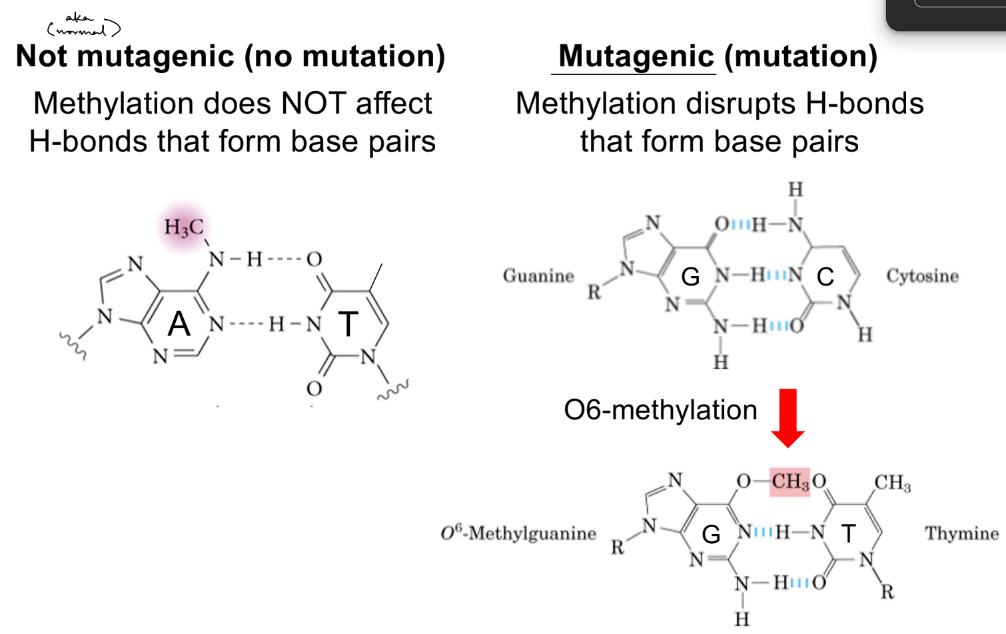
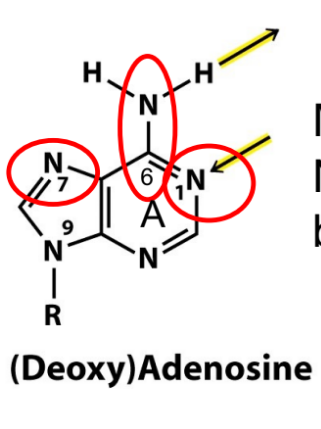
try to draw out adenosine w/o looking at picture & figure it out
for (deoxy)adenosine, is methylation at the circled positions N1, N6, and N7 mutagenic or not? explain
at N1: mutagenic
N1 is acceptor in a base pair w/ T (thymine)
at N6: not mutagenic
N6 is donor in a base pair w/ T, BUT N6 has 2 H’s (b/c NH2)
at N7: not mutagenic
even though N7 is a hydrogen bond acceptor, it is NOT involved in base pairing
aka N7 is not a H-bond acceptor in a standard base pair
(review that deamination is removing amine group & replacing it w/ carbonyl group)
t/f: deamination of a base can change the base’s identity
__
deamination occurs __, but can also be catalyzed by __
__
deamination of C will convert C into __
which makes the standard base pair of (1) into (1) mismatch, which can lead to mutations ← usually the most common mutation from this specific mismatch is called (1)
what may explain why DNA lacks uracil (U)?
true
__
deamination occurs spontaneously, but can also be catalyzed by enzymes
_
deamination of C: regularC (cytosine) → U (uracil)
standard base pair of C:G into U:G mismatch
U:G mismatch can cause mutations, most commonly the C>T mutation (aka C → T mutation)
(C>T mutation goes from standard base pair of C:G → T:A)
__
DNA may lack uracil b/c of spontaneous deamination of cytosine
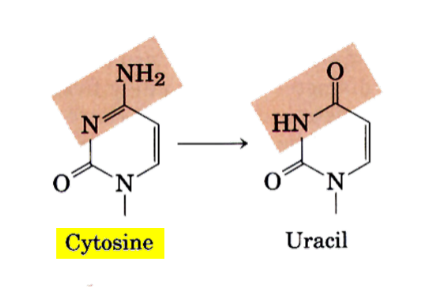
what base cannot be deaminated?
__
base-deamination changes these bases into what:
cytosine
5-methylcytosine
adenine
guanine
& know the structure of 5-methylcytosine (draw it)
thymine can’t be deaminated b/c thymine has no amine group to replace w/ carbonyl group
__
C → U
C5-me → thymine
A → hypoxanthine
G → xanthine
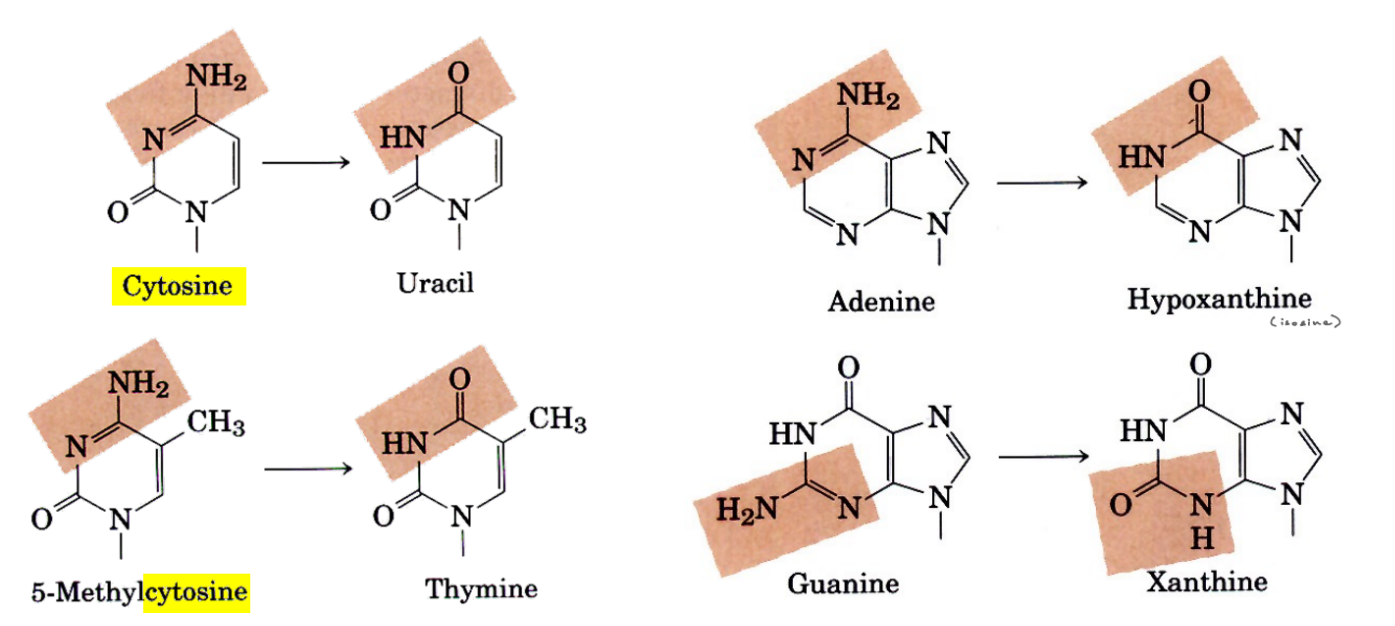
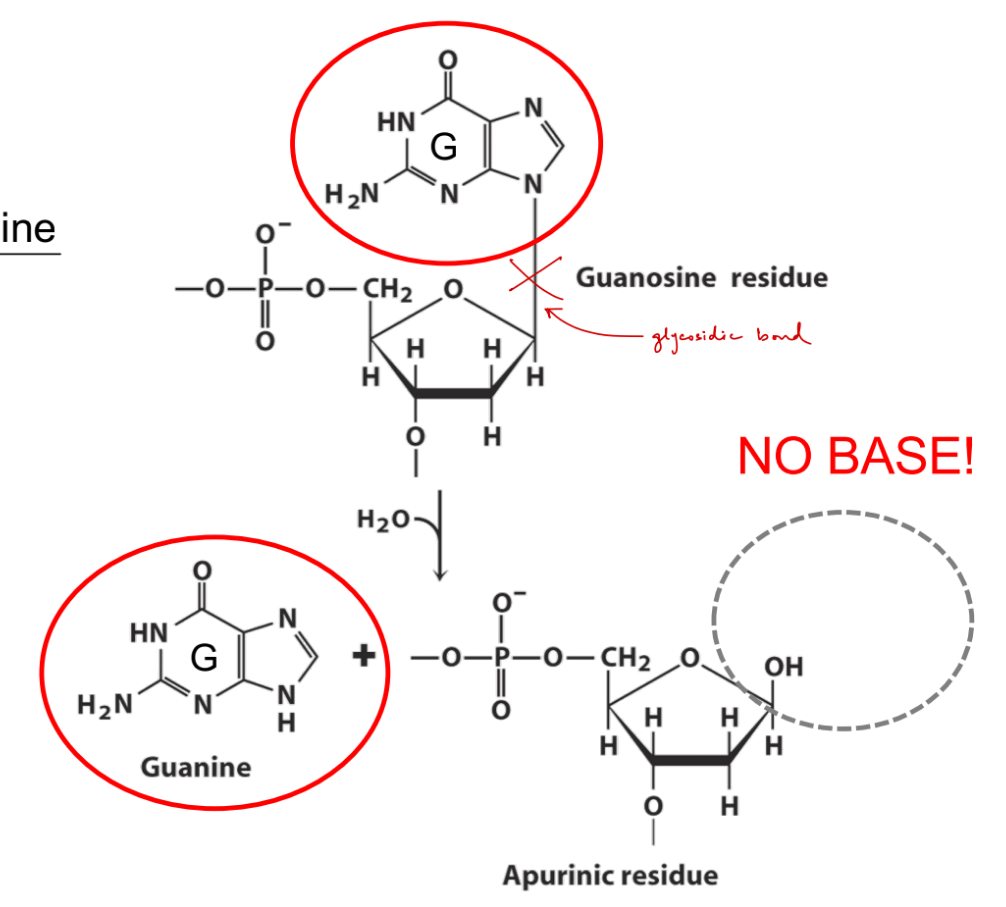
review:
def. of depurination
what bases are affected
def. apurinic
what bond is cleaved/removed?
remove entire purine base (adenine, guanine)
cleaves the glycosidic bond
_
apurinic — no purine base b/c it has been removed
(NOT apurinic if it has a pyrimidine base)
alkylation, deamination, depurination were all mutagenic CHEMICAL modifications, but
what is the mutagenic STRUCTURAL modification (1 ← name & def.)
2 affected bases
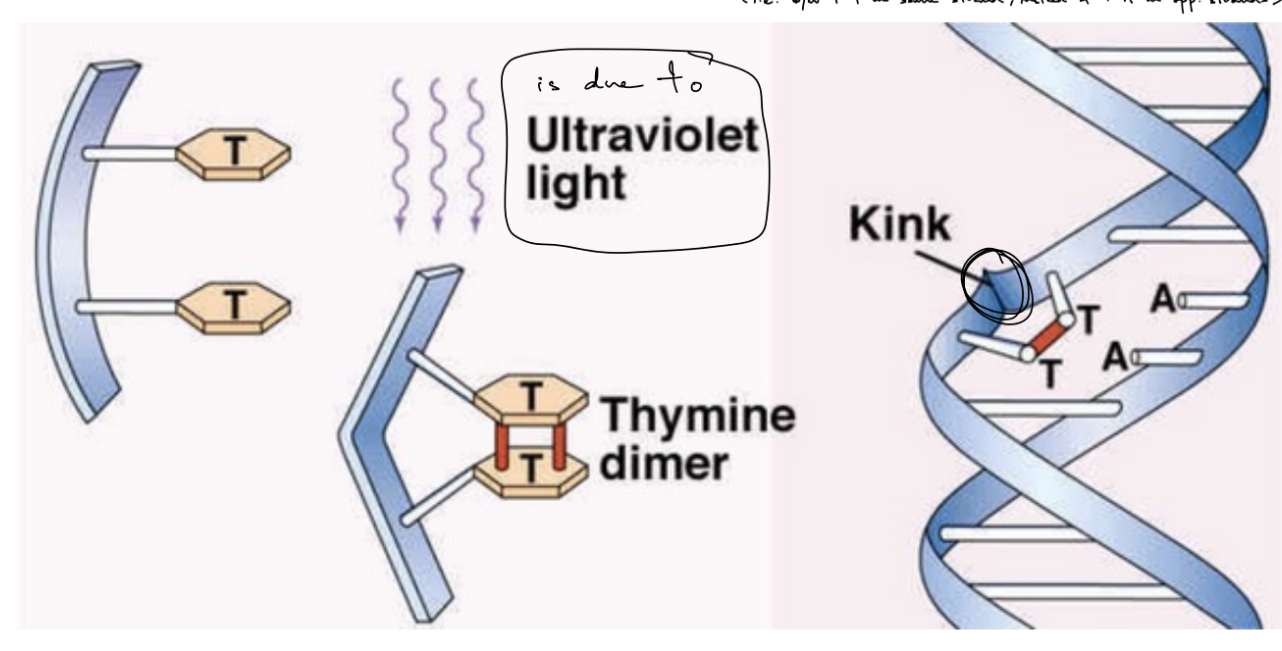
def. kink
usually forms b/w …
pyrimidine dimers
UV light causes a kink to form B/C of covalent bond that is made b/w pyrimidines on the same strand
kink — structural change to DNA helix
affects pyrimidine bases: thymine, cytosine
NOT usually uracil
SO like covalent bond b/w T-T on same strand
usually forms b/w T-T
just name 3 DNA repair mechanisms (that repair damaged base(s) that occur independently from replication)
base excision repair (BER)
nucleotide excision repair (NER)
direct repair
def. direct repair
is not a pathway for DNA repair, but is instead, a class of different enzymes that change/modify a base
(i.e. methyltransferase does demethylation/remove methyl group to revert the mutagenic methylation of O6 → into normal/unmethylated guanine)
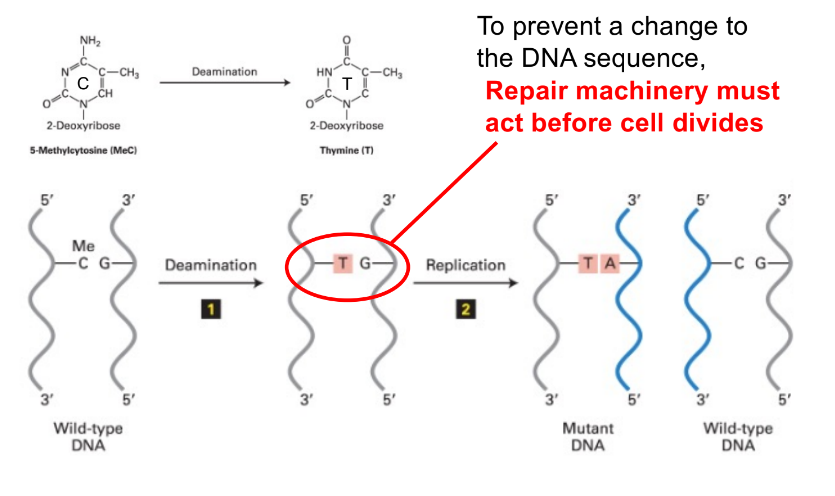
how do you prevent a mistake from becoming a mutation?
repair machinery must act before the cell divides
aka
DNA repair must happen before replication happens, or else the mistake will become a PERMANENT mutation (after replication, mismatch becomes a mutation)
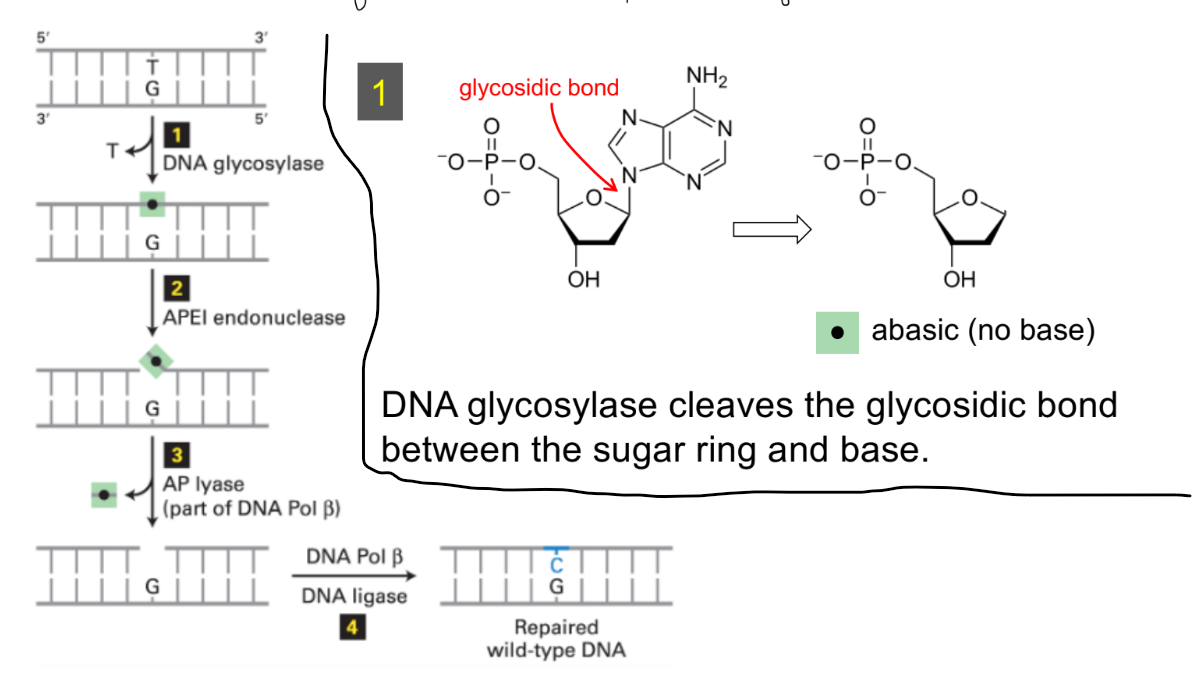
def. base excision repair (BER)
explain w/ example of: BER steps for deaminated 5-methylcytosine aka guanine (4)
repairs that change DNA at ONE or few nucleotides & do NOT cause a major distortion of DNA
(remove 1 or few damaged/mismatched bases before replication occurs)
__
BER w/ deaminated 5-methylcytosine aka guanine:
DNA glycosylase cleaves the glycosidic bond (b/w the sugar ring & base) —→ to make abasic nucleotide (aka nucleotide w/o base)
glycosylase is NOT specific in which strand’s mismatched base is changed (aka recognizes the mismatch/damage, but not the strand)
AP endonuclease cleaves the phosphodiester bonds to remove the abasic nucleotide w/in the strand → forms a gap
(AP endonuc. is either apurinic or apyrimidinic endonuclease)
DNA polymerase synthesizes new DNA to fill in gap
DNA ligase seals the remaining nick
is DNA glycosylase need to repair depurinated bases? why? what would happen as the first step?
no
b/c depurinated bases don’t have a base, meaning that there is no glycosidic bond (which usually forms b/w sugar ring + base)
so glycosylase (which cleaves glycosidic bonds) is not needed for depurinated bases
SO BER’s the first step would be AP endonuclease (skip the glycosylase step)
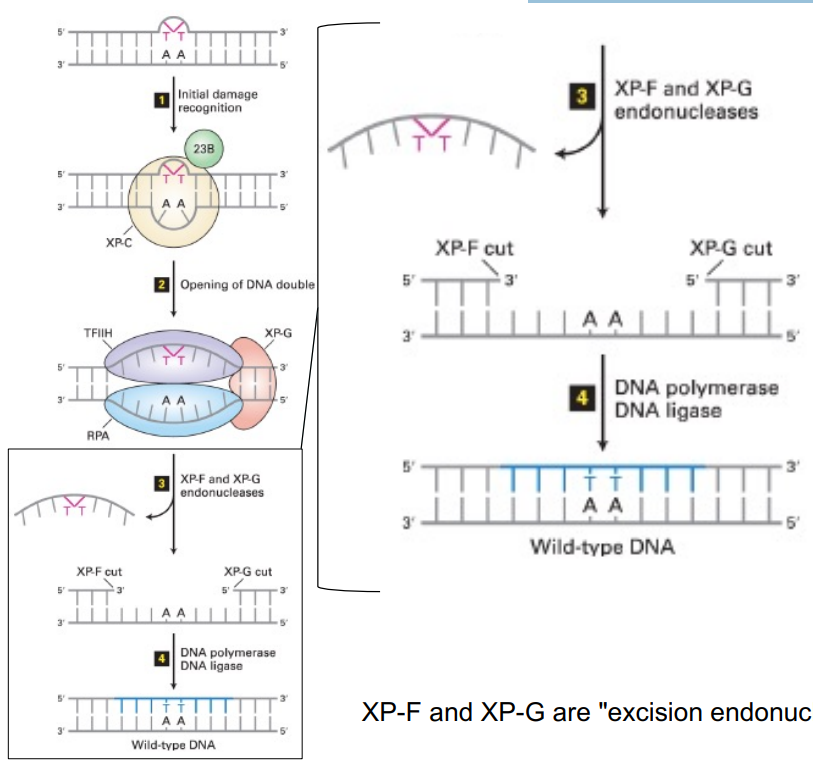
def. nucleotide excision repair (NER)
name example of NER
explain steps of NER using pyrimidine dimers
repairs that change DNA at multiple nucleotides & causes a major distortion of DNA
like pyrimidine dimers
__
NER using pyrimidine dimers:
recognize damage (aka pyrimidine dimers, like T-T)
helicase opens/unwinds the helix
endonucleases remove multiple nucleotides surrounding the damaged region (on the same strand)
specifically, excision endonucleases aka excinucleases, like XP-F and XP-G
DNA polymerase synthesizes new DNA to fill in the gap
DNA ligase seals the remaining nick
explain how (regular/unmethylated) C deamination in dividing cells → results in base-pair mismatch & in a C>T mutation (3 steps)
& state transition of standard & non-standard/mismatched base pairs @ each step
(unmethylated) C is deaminated into U
so goes from C:G → U:G
cell division/replication occurs (dividing cells), where U is though to be part of the parent strand, SO DNA pol. pairs it w/ A in newly synthesized daughter strand
aka U:G —(cell div./repl. using U strand as the parent template strand)→ U:A
after cell division is done, BER occurs to replace U w/ T (B/C it recognizes that uracil is not apart of DNA)
so U:A → T:A
(C:G → U:G → U:A → T:A ————— this is a C>T mutation)
^^^ b/c replication happened when mismatch was still present/not fixed, it caused a (permanent) mutation
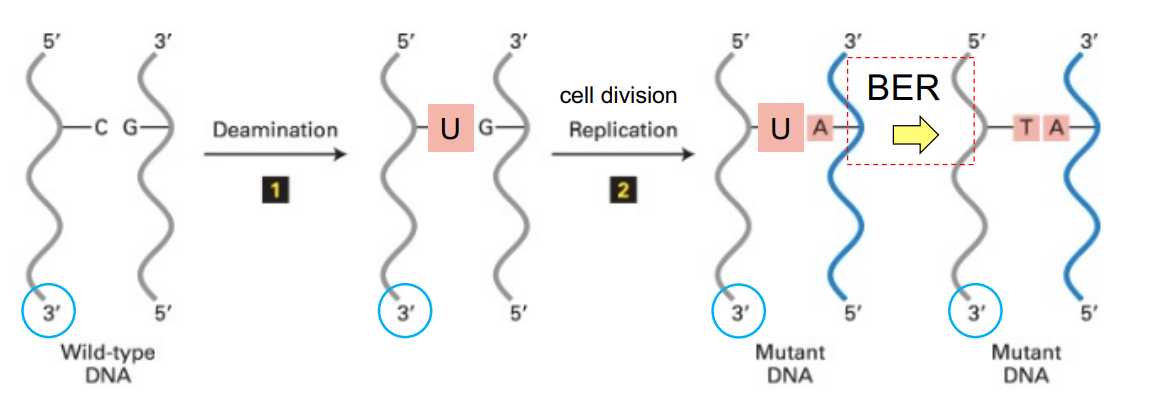
explain how 5-methycytosine deamination in dividing cells → results in mismatch & in mutation
deamination of C5-me, where C5-me:G → T:G
cell division/replication occurs (dividing cells), where makes 2 daughter strands: 1 mutant T:A & 1 wild-type C:G strand
mutant strand is T:A (b/c has diff. base pair from original starting, standard base pair)
wild-type strand is C:G (b/c has same base pair as original starting base pair)
(C>T mutation in mutant strand)
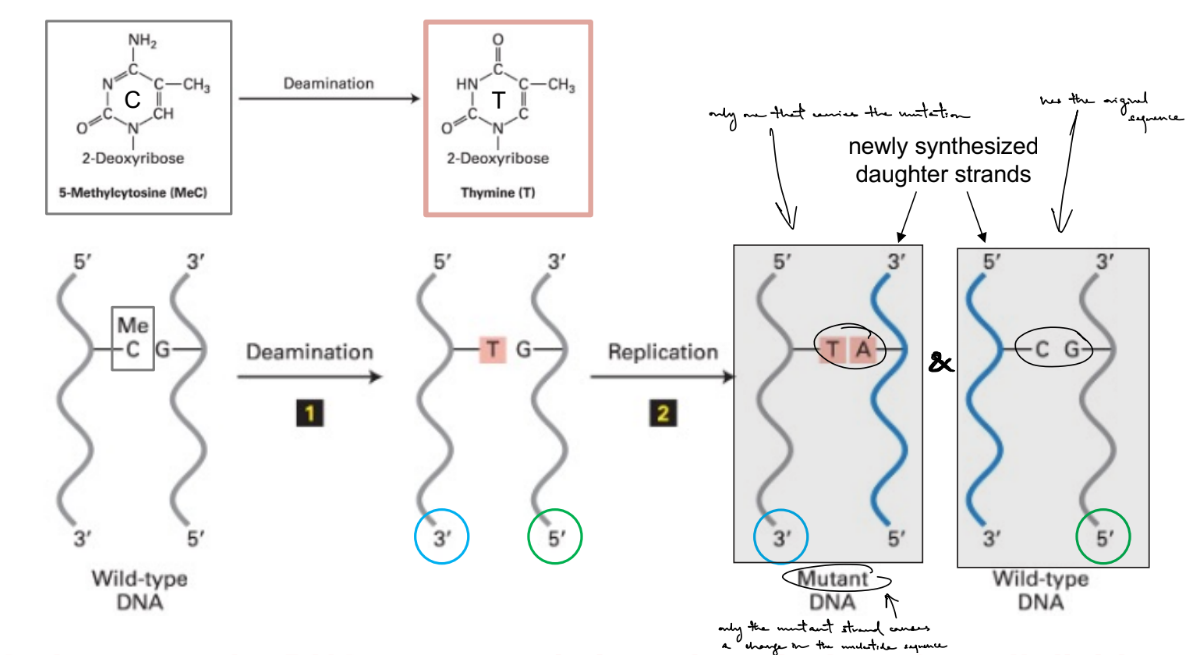
explain how 5-methycytosine deamination in NON-DIVIDING cells → results in mutation
specifically results in C>T mutation
b/c deamination of C5-me, where C5-me:G → T:G
(aka C5-me → T)
can a mutation (like a C>T mutation) occur from:
deamination of C in dividing cell
deamination of 5-methylcytosine in non-dividing cell
deamination of C in non-dividing cell
deamination of 5-methylcytosine in dividing cell
deamination of C in dividing cell
YES, b/c C:G (when deamination)→ U:G → U:A (when cell division/replication) → T:A (when BER of T)
deamination of 5-methylcytosine in non-dividing cell
likely YES
b/c C5-me:G (when deamination) → T:G
aka C turns into T, so is C>T mutation
deamination of C in non-dividing cell
likely NOT
b/c C:G (deamination) → U:G (BER) → original C:G
b/c BER recognizes that U is not part of DNA
this happens despite the glycosylase step of BER doing non-specific cleaving of a strand, BER will recognize that U is usually part of RNA, not DNA
deamination of 5-methylcytosine in dividing cell
YES, b/c C5-me:G (when deamination) → T:G (when replication) → get two daughter strands of T:A & C:G
mutant strand of C>T mutation is T:A (b/c causes a change in/is different from the starting standard base pair)
wild-type DNA strand is C:G (b/c is the same base pair as the original standard base pair)
^^^
if mismatch/mutation is NOT repaired before a cell divides, it will cause a (permanent) mutation in the parent strand & in synthesized daughter strand
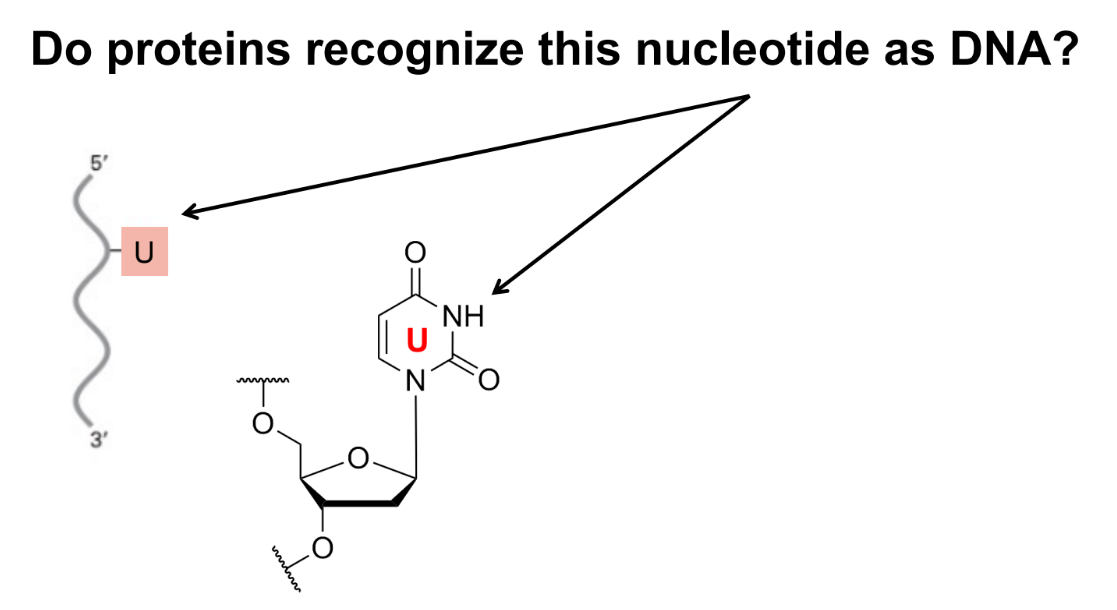
yes b/c there’s no 2’ OH (think RUH “ROH”), so that means it’s DNA
b/c DNA vs. RNA is deter. by the 2’ sugar ring, not the type of base (i.e. not U vs. T)
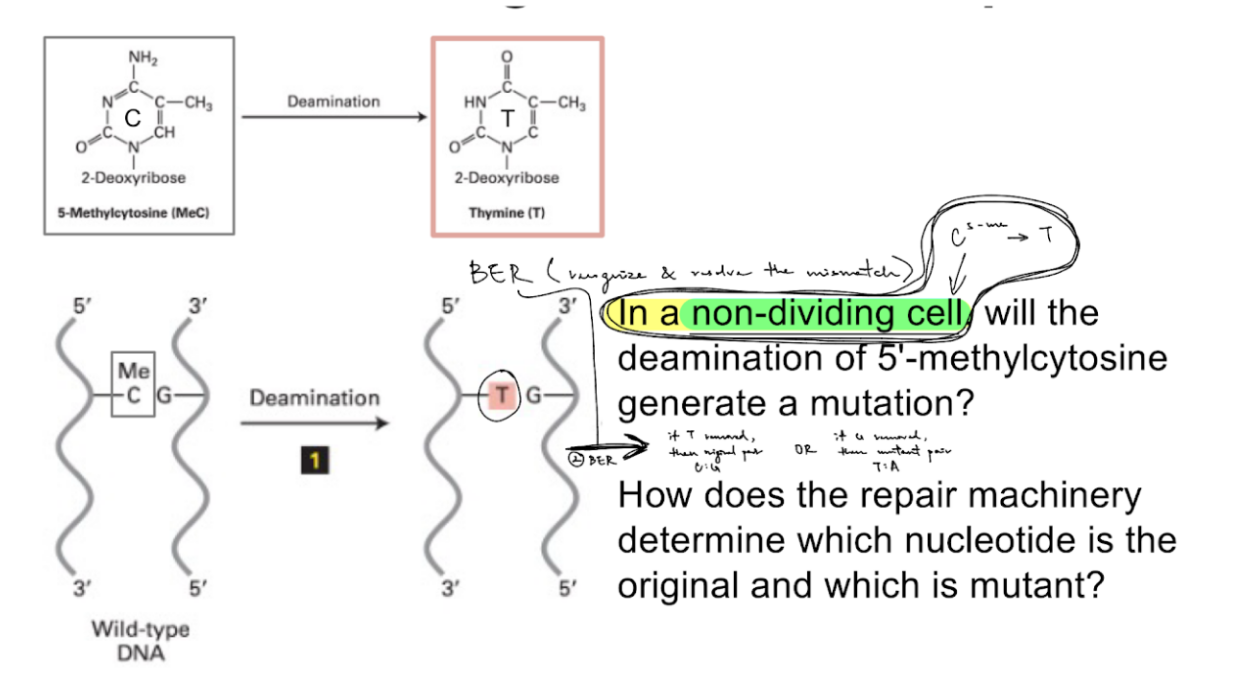
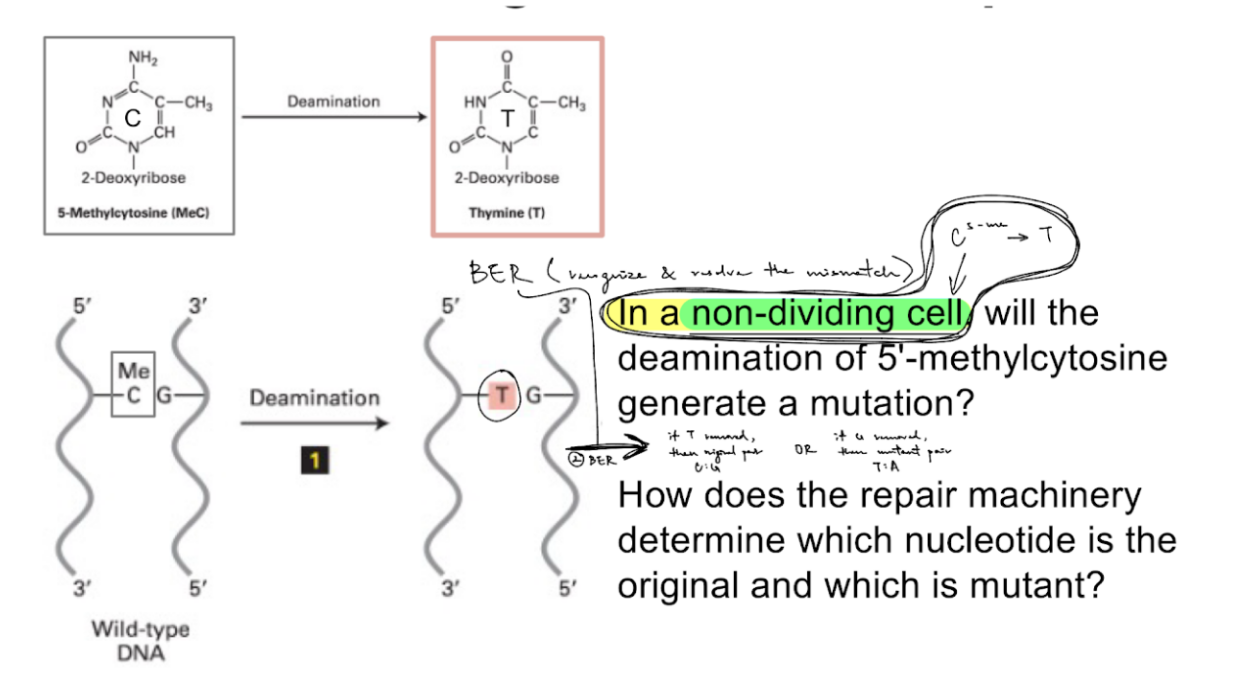
for 5-methylcytosine deamination in non-dividing cells (shown in picture),
how does the repair machinery deter. which nucleotide is the original vs. mutant?
it can’t
b/c the “mutant” deaminated 5-methylcytosine is T (thymine)
BER occurs as the next step after deamination (b/c non-dividing cells):
sometimes BER removes T → then restored to original base pair of C:G
sometimes BER removes G → then sequence will mutate/change into T:A
t/f: spontaneous deamination of cytosine changes our genome & results in C>T mutations
# of cytosines spontaneously deaminated per day
% of human genome that is made of cytosine
__
t/f: C>T mutations are highly represented in “mutational signatures” associated w/ cancer
_
t/f: malfunctioning DNA repair machinery leads to predisposition to disease
true
100-500 Cs deaminated / day
20% of human genome that is made of cytosine
__
true
b/c cancers are actively dividing cells
__
true
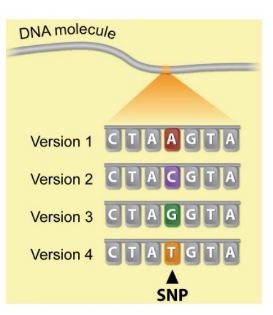
human genome has (#) base pairs
t/f: no two genomes have identical sequences b/c our genomes have many SNPs / single-nucleotide polymorphisms
t/f: a mutation may not even be detected b/c there may not be a “correct”/common nucleotide for that position in the genome
3.2 billion base pairs
_
true
true
incorrect base incorporation during DNA replication is fixed how?
__
chemical changes to DNA bases (alkylation, deamination, depurination) is fixed by (← state for each of the 3)
structural changes to DNA (pyrimidine dimers) is fixed by (1)
fixed by DNA polymerase’s exonuclease activity OR mismatch repair
__
chemical changes to DNA bases (alkylation, deamination, depurination) is fixed by:
alkylation — by BER or direct repair (methyltransferase)
deamination — by BER (need glycosylase)
depurination — by BER (don’t need glycosylase)
structural changes to DNA (pyrimidine dimers) is fixed by — NER or direct repair
early discoveries from mapping genes (~4)
first genetic map of chromosome (Sturtevant)
DNA is the genetic molecule (…McCarty experiment)
pieces of DNA “hop” around the genome, which disrupts genes & changes traits
structure of DNA solved
for study on location genes w/in a genome:
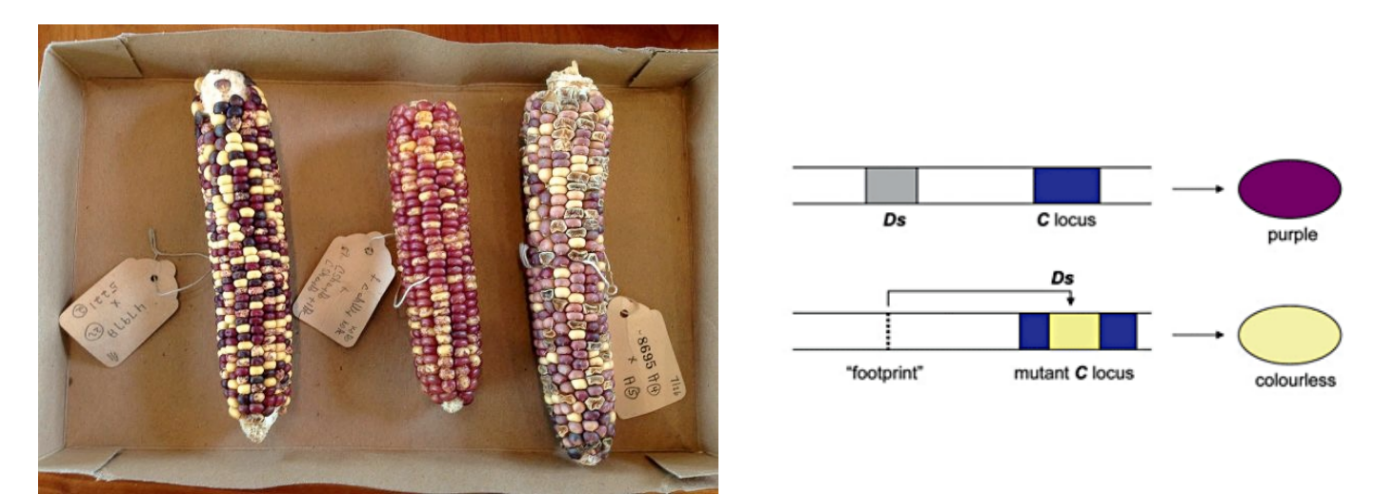
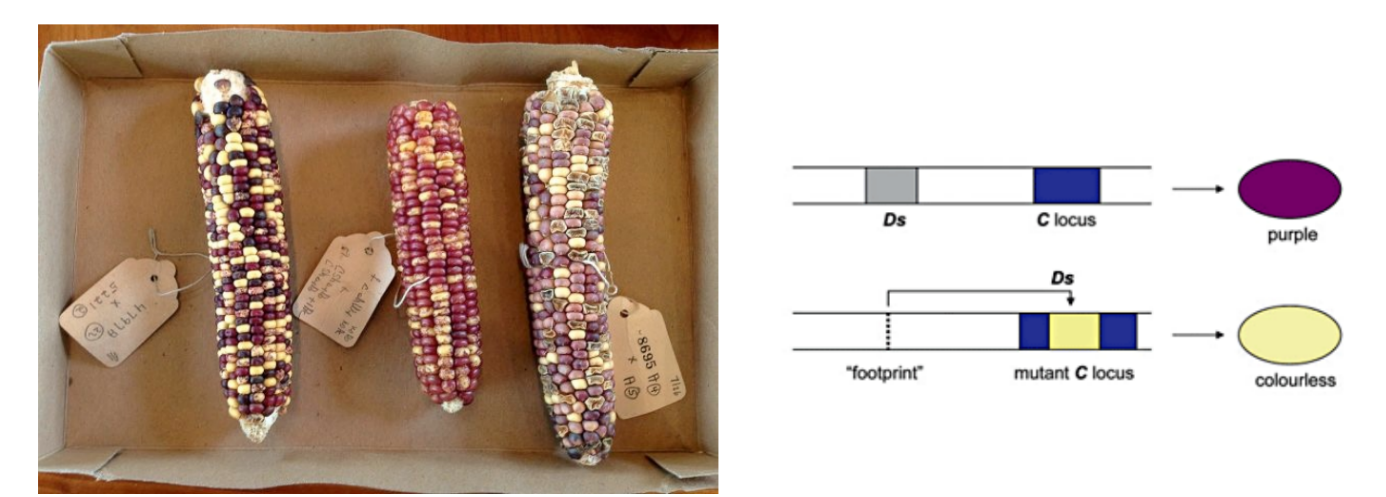
explain experiment w/ pigmentation of corn
kernel color (trait), corn/maize genome
genomic DNA (chromosomes) are treated w/ DNA stain to get patterns of light and dark bands → used to find parts of genome that contain genes correlating w/ a specific trait
when the genomic fragment Ds that jumps around is in a position → purple kernels
when genomic fragment Ds is in another position → colorless/yellow kernels
(the genomic fragment Ds is the transposon)

def. Mendelian inheritance
__
def. transposons
name 2 broad classes of transposons & briefly def.
t/f: more than half the human genome s made of transposable elements
genes are transmitted in a constant, predictable way from parent-to-child
__
transposons — pieces of genomic DNA that can “jump” around (aka can be pasted/inserted into different positions of a genome to affect the expressed traits)
DNA transposon — cut & paste
retrotransposon — copy & paste
__
true
describe how DNA transposons work (incl. enzymes involved) (2)
DNA transposons
transposase cut/excises DNA transposon out of the original position in genome/DNA sequence (& flanking DNA leaves) → make a DNA intermediate
transposase pastes DNA inter. into another part of the genome (the target DNA) → make a transposed mobile element, where original/same DNA transposon is in different position
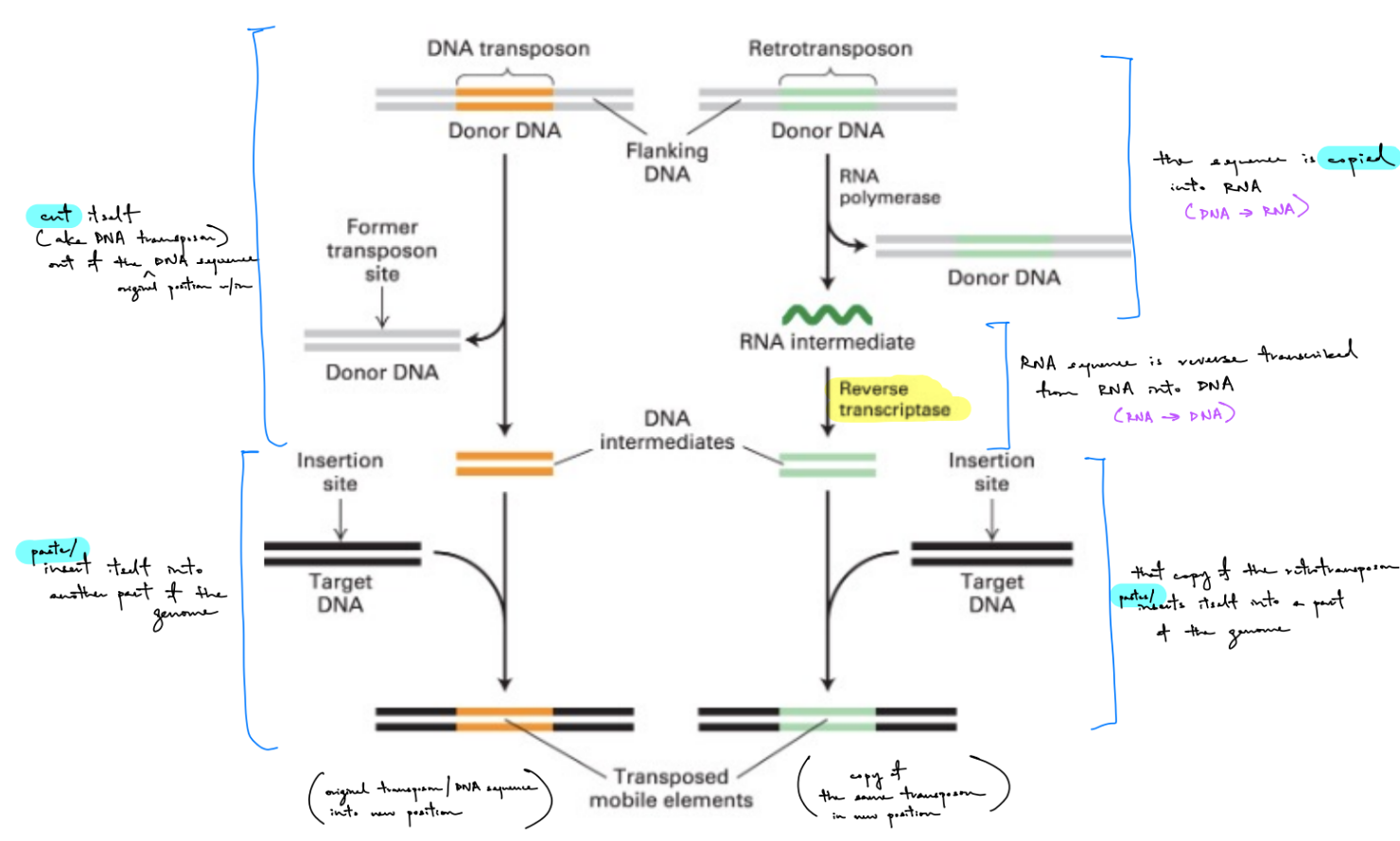
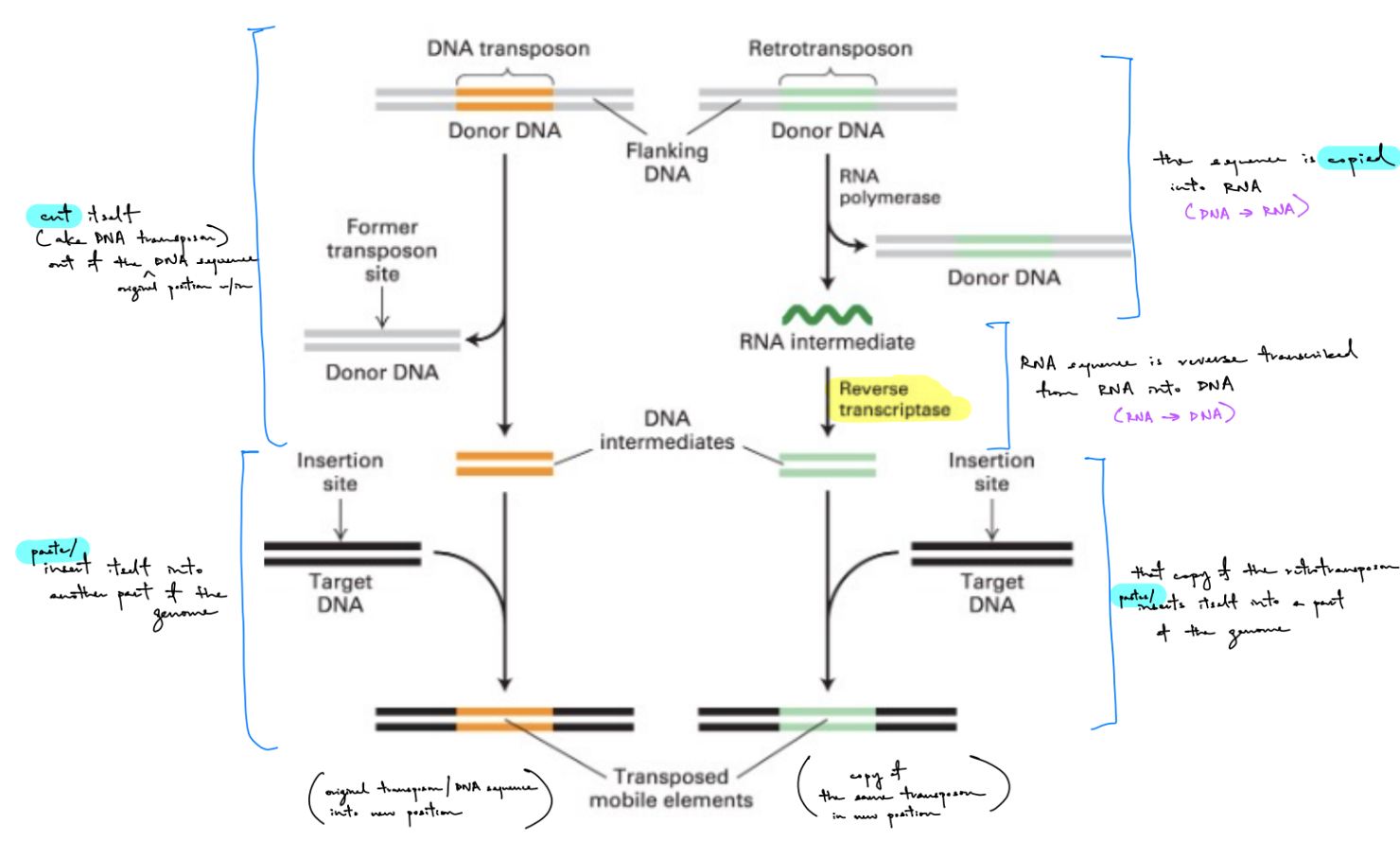
describe how retrotransposons work (incl. enzymes involved) (3)
__
how many copies of the retrotransposon do you have at the end?
_
def. reverse transcriptase (give an example)
retrotransposons
RNA polymerase copies retrotransposon into RNA (& original sequence w/ retrotransposon + flanking DNA leaves ← no excision of original retrotransposon) → make a RNA intermediate
via reverse transcriptase: RNA sequence is reverse transcribed from RNA into DNA → make a DNA intermediate
DNA inter. is pasted into another part of the genome (the target DNA) → make a transposed mobile element, where copy of retrotransposon is in different position
__
2 copies of retrotransposon: original (“donor DNA”) & newly synthesized transposon (“transposed mobile element”)
reverse transcriptase is a DNA polymerase that uses an RNA template
like telomerase
cont. kernel pigmentation study
the movement of a transposon in the maize genome causes variation in the (1)
t/f: insertion of transposon can have the effects of decreasing OR increasing gene expression (i.e. pigment). ALSO, the presence and absence of transposons can also affect the expression of the trait
_
t/f: transposons induce variations in traits, which can include intermediate expressions of the trait
_
t/f: transposons induce variation in traits that contribute to evolution
kernel pigmentation
b/c insertion of transposons in new positions w/in genome causes variations in expressed traits
(aka transposons induce variations in traits)
__
true
w/ inserted transposon → either decrease or increased expression of trait
w/ or w/o inserted transposon → either decrease or increased expression of trait
__
true
i.e. no transposon is black grapes, w/ transposon is white grapes, partially removed/piece of transposon is red grapes
_
true
i.e. rise in industry caused moths to evolve from peppered to black appearance

where are genes located in the genome (2 parts)
genes are present in consistent, predictable positions (fixed positions) w/in genome of an organism
BUT some pieces of DNA called transposons can jump in & out of genes (insertion + excision), which affects traits
t/f: gene structure is sim. in all eukaryotic genomes
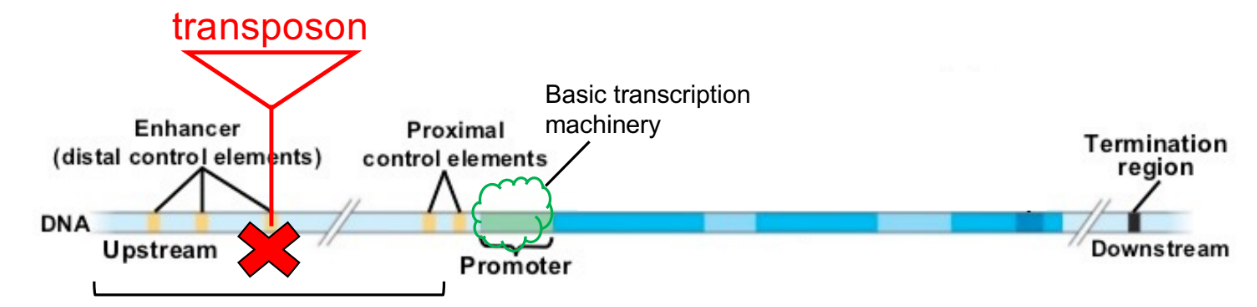
name & def. 3 common parts of a gene in eu-
what does 1 of them tell us (2)
true
__
enhancer
part of DNA sequence that binds proteins, like TFs, that stabilize or destabilize the RNA polymerase bound to the promoter
promoter
part of DNA sequence that binds RNA polymerase, SO when stably bound, starts RNA transcription
transcription unit
part of DNA sequence that encodes for RNA (aka DNA template for RNA synthesis / DNA → RNA)
where if RNA codes for → proteins → trait, then that transcription unit is part of the “blueprint”
but can also have RNA encoding for→ trait
__
enhancer tells us where a gene is expressed & the amount of RNA transcribed
name & def. two elements/parts of mRNAs / of transcription unit
exons — part of mRNA that is transcribed & kept in mature mRNA
introns — is removed from mRNA
what happens if transposon is inserted/pasted into an enhancer?
gene is expressed in different areas/different tissues (← pattern)
amount of RNA expression (from RNA transcription) & amount of protein changes
BUT
protein made is normal (just the amount of protein changes) b/c has no effect on transcription units, which code for RNA
^ basically functions of enhancers