EXAM 3
1/137
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
138 Terms
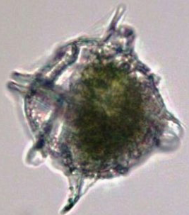
Pyrodinium bahamense
microbial eukaryote (domain: eukaryota)
produces saxitoxin
bioluminescent !! (in water)
phylum: Dinoflagellata
class: dinophyceae
order: Gonyaulacales
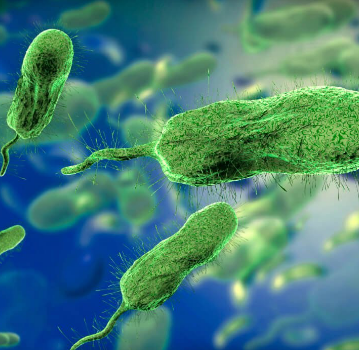
Vibrio vulnificus
gram negative
motile
rod, bacillus
present in marine environments
LPS (Lipopolysaccharides) — endotoxin, multiple exotoxins that cause the diseases
3 types of infection:
gastroenteritis (oysters)
severe wound infections (swimming in contaminated water)
sepsis (bloodstream; oysters/shellfish)
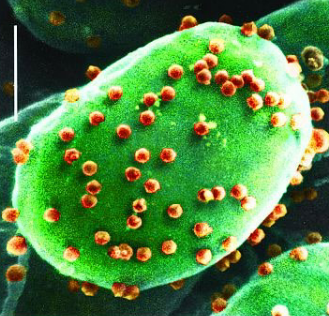
ATCV-1 Virus
Acanthocystis turfacea chlorella virus 1
giant dsDNA virus
type of Chlorovirus → infects green algae, common in fresh water like lakes and ponds
virus associated with reduced cognitive functioning
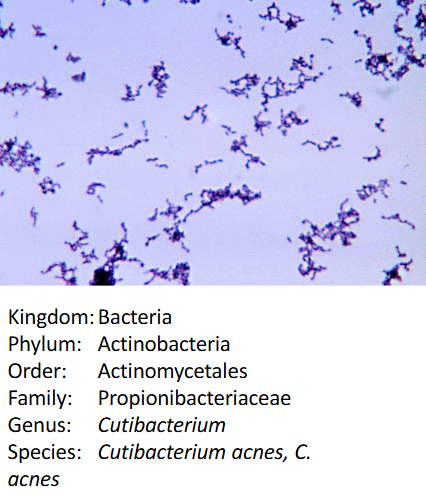
Cutibacterium acnes
slow-growing
typically aerotolerant anaerobic
Gram Positive
rod shaped
part of normal skin flora also found deep in hair follicles and pores → can cause acne
use sebum, cellular debris, and metabolic byproducts form surrounding skin tissue as primary sources of energy and nutrients
Saxitoxin
a highly potent natural neurotoxin
200 micrograms lethal for average-weight human
primarily known as the causative agent of paralytic shellfish poisoning (PSP) and pufferfish poisoning
mid-2000s FL saxitoxin outbreak from pufferfish caused by Pyrodinium bahamense
sxtA4
gene that appears to be specific for saxitoxin synthesis = proxy for toxin production
phylogenetic analysis indicates it is lost in nontoxic strains and species
Microbial Competition
success in environment based on ability to multiple and withstand adverse conditions
other factors in success → chemical warfare & streptomyces (many microbial compounds)
Microbial Communities
microbial mats
biofilm (distinct layers)
Biofilm Layers
cyanobacteria (top) → green photosynthetic pigments
purple sulfur bacteria → use light-harvesting pigments of dif. wavelengths than cyanos
sulfate-reducers (bottom)
Sulfate reducers
bottom layer of microbial mats
iron-reacting hydrogen sulfide
sulfate as terminal e- acceptor
Desulfovibrio oxyclinae
Desulfovibrio halophytica
Desulfonema & Desulfococcus species
Methods for Microbial Ecology
metagenomics (taxonomy & genetic/functional capabilities)
FISH (fluorescent in situ-hybridization)
DNA microarrays
Metagenomics
Who’s there and what are they capable of?
method of DNA extraction
overall numerous short sequencing reads that are software-analyzed
method of microbial ecology
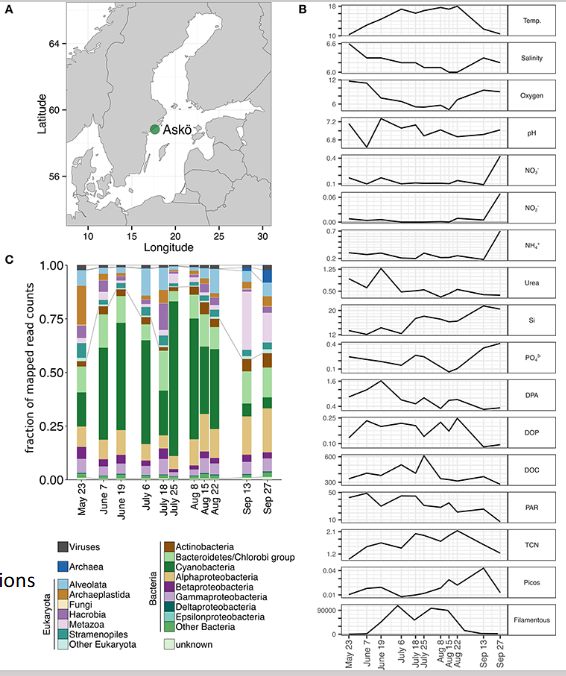
Metatranscriptomics
What are they doing?
RNA sequences
nucleic extraction method
short half-life
presence of Rnases
lots of rRNA
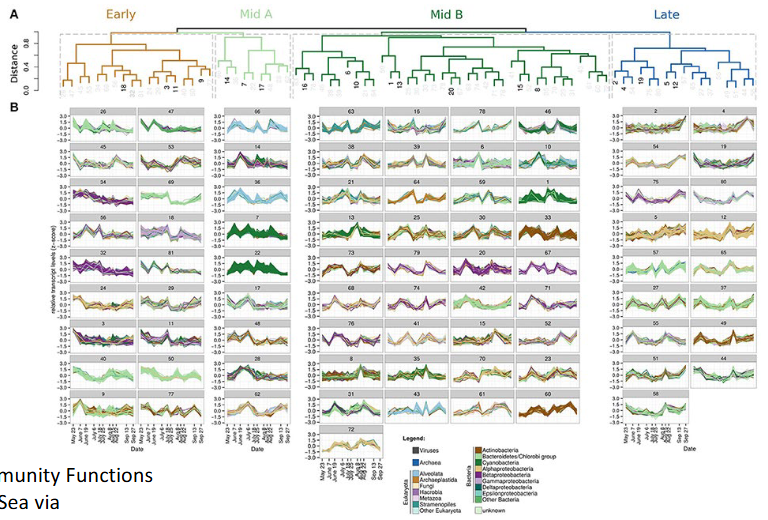
Taxonomic profile comparison at genus rank
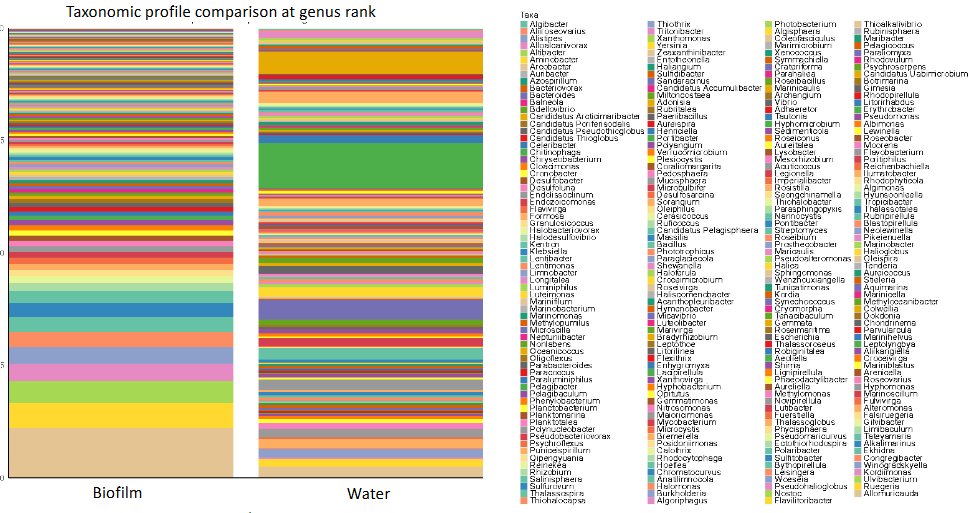
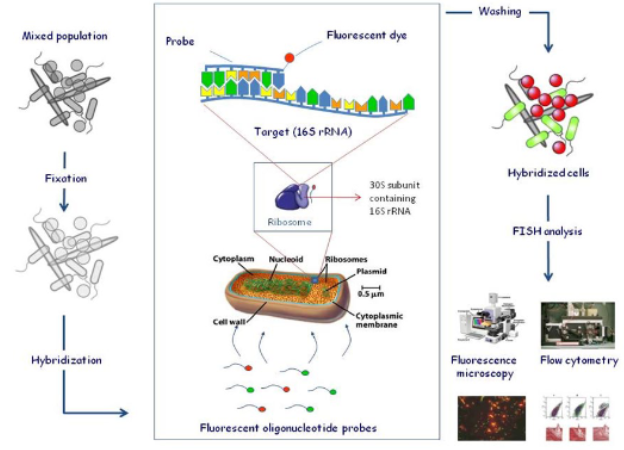
FISH
fluorescent in-situ hybridization
method for microbial ecology
can be used to distinguish subsets of prokaryotes that contain a specific nucleotide sequence

Which of the following is false?
a) culture techniques are an accurate way of determining which members in a microbial community are most common
Oligotrophic
nutrient poor
Eutrophic
nutrient rich
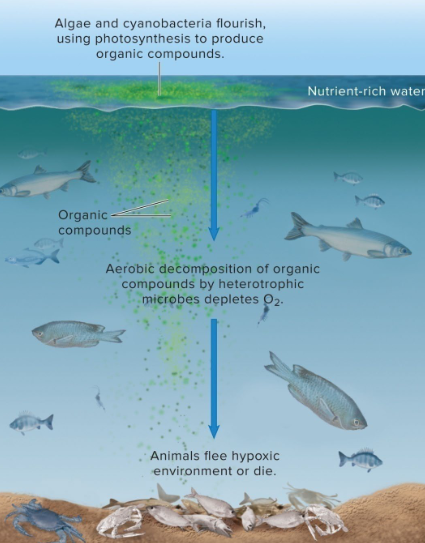
Eutrophication
excessive enrichment of water bodies with nutrients—primarily nitrogen and phosphorus—leading to dense algal blooms, severe oxygen depletion (hypoxia), and ecological degradation
Aquatic Habitats: Marine
oceans cover 70%+ of the Earth
oligotrophic = nutrient poor
eutrophic = nutrient rich
eutrophication can lead to hypoxic conditions (too little O2)
heterotrophs in lower layers → consumption of organic compounds + dissolved O2
Dead Zones
Eutrophication in coastal areas
Gulf of Mexico Dead Zone
Harmful Algal Blooms
many difference species and toxins
Dinoflagellates (microbial eukaryotes) & cyanobacteria
Pyrodinium bahamense
Lake Stratification — Freshwater Systems
the separation of lakes into three distinct horizontal layers based on water density differences caused by temperature variations
Epilimnion
the top-most, warmest, and least dense layer of water in a thermally stratified lake
Hypolimnion
the dense, cold, and dark bottom layer of a thermally stratified lake, located below the thermocline
Metalimnion
the middle layer of a thermally stratified lake, located between the warm surface water (epilimnion) and cold deep water (hypolimnion)
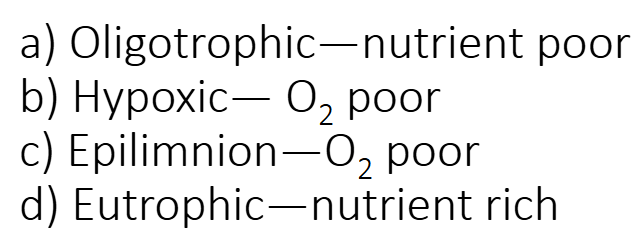
Which of the following pairs that relate to aquatic environments does not match?
Epilimnion — O2 poor

Adding high levels of nutrients to a lake or inshore area would have all of the following effects in that environment except…
increased levels of dissolved O2
Soil Microbiome
great diversity
Rhizosphere = important component of soil microbiome → plant roots (secret organic compounds)
Streptomyces (soil bacteria) = antimicrobials, geosmins (soil smell)
Bacillus = endospores for survival
microbes in soil are good source of bioremediation potentials

Which of the following is NOT a matching pair?
soil — minimal biodiversity
Biogeochemical Cycles
carbon cycle
nitrogen cycle
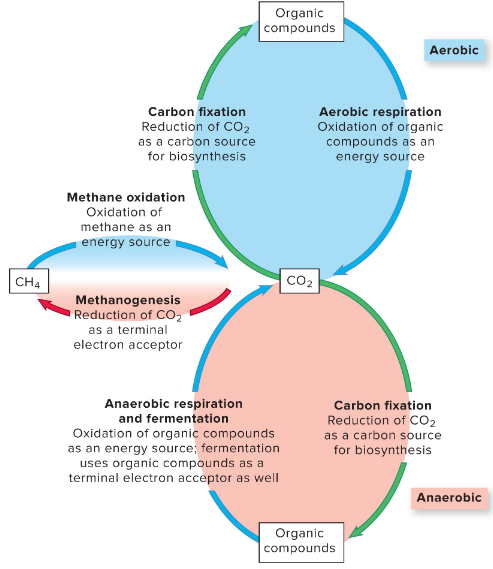
Carbon Cycle
carbon fixation — autotrophs convert CO2 into an organic form
respiration & fermentation — CO2 usually produced via activities of heterotrophs
methanogenesis & methane oxidation
methanogens — archaea oxidize hydrogen gas, CO2 as terminal e- acceptor, make methane
methyltrophs — methane as energy source
the carbon cycle revolves around carbon dioxide !!
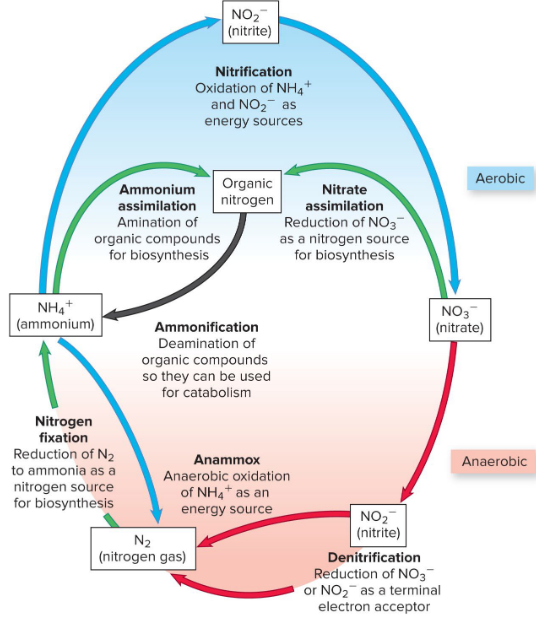
Nitrogen Cycle
nitrogen fixation → nitrogenase, diazotroph (azotobacter, clostridium)
ammonification
nitrification
denitrification
anammox
Food Poisoning
when a pathogen or its toxin is consumed
Exotoxin
highly potent, protein-based toxins secreted by living bacteria into their environment
Foodborne Intoxication
illness that results form consuming exotoxin
Staphylococcus aureus
Clostridium botulinum
Staphylococcus aureus
many toxins
gastroenteritis — enterotoxin = protein exotoxin
strain detection: Qpcr, MLST, PFGE
Clostridium botulinum
anaerobic
gram positive
lab isolation — in tryptose sulfite cycloserine (TSC) in anaerobic (<2%) oxygen
endospores
neurotoxin produced by vegetative cells = botulism
8 neurotoxins = A-G; A, B, E, F, H cause human illness
Toxin binds to nerve endings, so nerves cannot signal to heart
Qpcr for toxin genes A, B, E, F

Which of the following organisms cause foodborne intoxication?
Staphylococcus aureus
Virus
requires host cells
most only visible with electron microscopy
prokaryotic viruses (bacteriophages) vs eukaryotic viruses
~10-800 nm in size
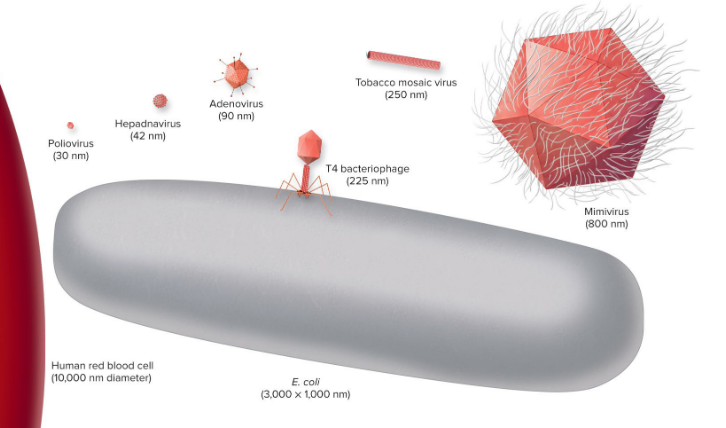
Virus structures
Virion
Nucleocapsid
Capsomeres
Enveloped vs non-enveloped/naked viruses
tail fibers (phages) and spikes (animal) for specific receptors
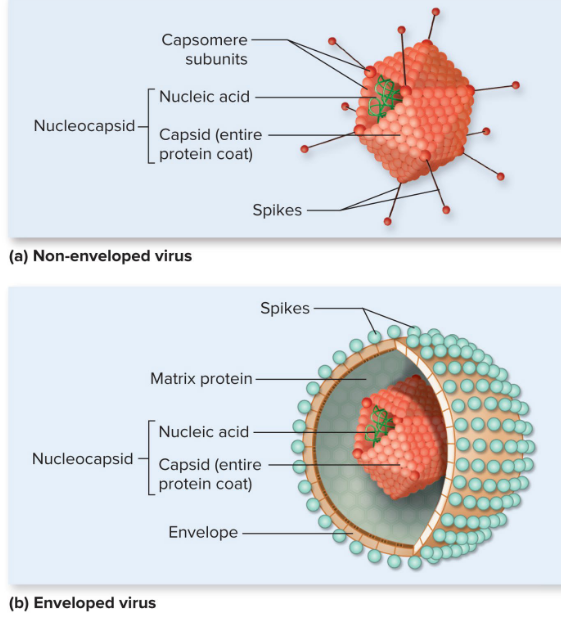
Virion
complete extracellular viral particle
capsid = protection + infection
DNA/RNA — linear or circular, DS or SS, type of nucleic acid used in classification
Nucleocapsid
nucleic acid + capsid
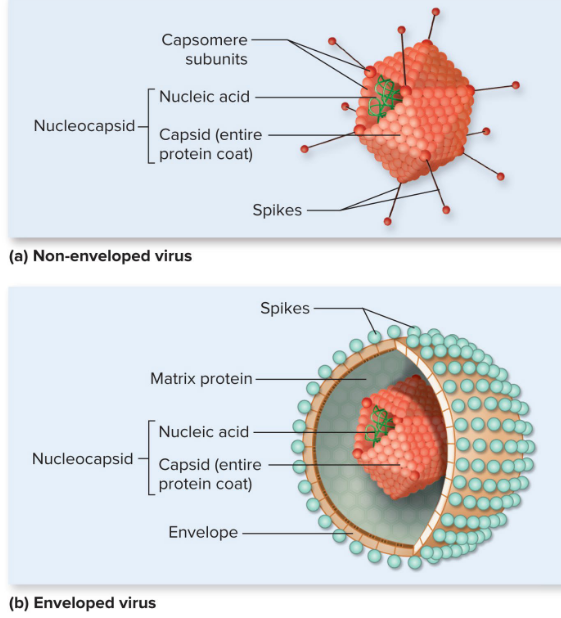
Capsomeres
protein subunits
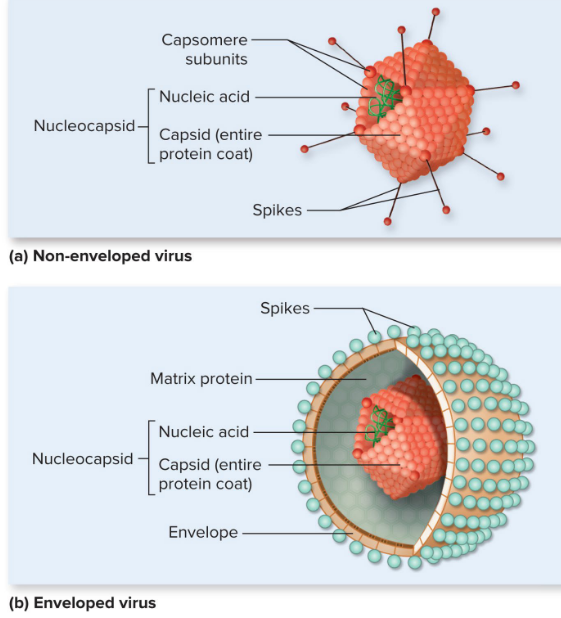
Enveloped vs Naked Viruses
lipid bilayer is unique to enveloped viruses
phage = naked and non-enveloped
enveloped viruses are more susceptible to disinfectants
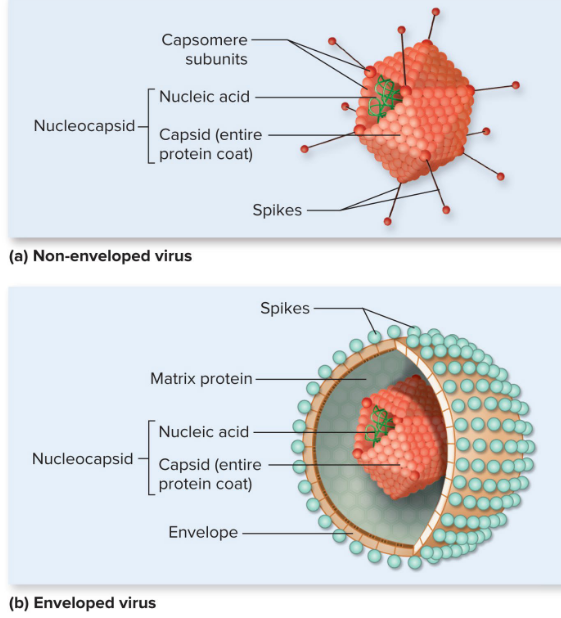
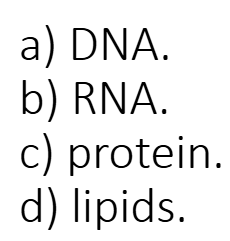
Capsids are composed of…
protein

The tail fibers on phages are associated with…
attachment
Shape of viruses
determined by capsid
icosahedral
helical
complex (most phages)
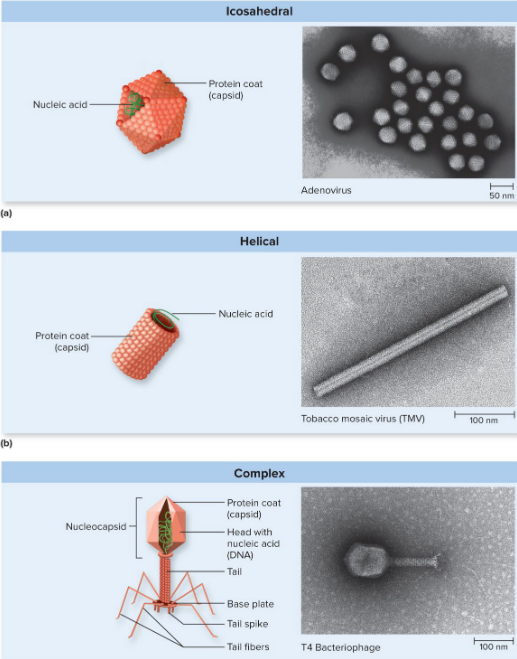
Virus Taxonomy
key characteristics for classification:
genome structure (nucleic acid, strandedness)
host
shape and disease
families: -viridae
geographic location
genera (many within a family): -virus
species: based on disease
no bacterial nomenclature
route of transmission rather than taxonomical relatedness
Zoonotic
disease transmitted from animal to human
Arboviruses
virus spread by arthropods, can infect many different species (Wes Nile; yellow fever)
Bacteriophages
viruses that infect bacteria
Phage Replication
similar mechanisms as for viruses that infect animal cells
productive infection
latent state
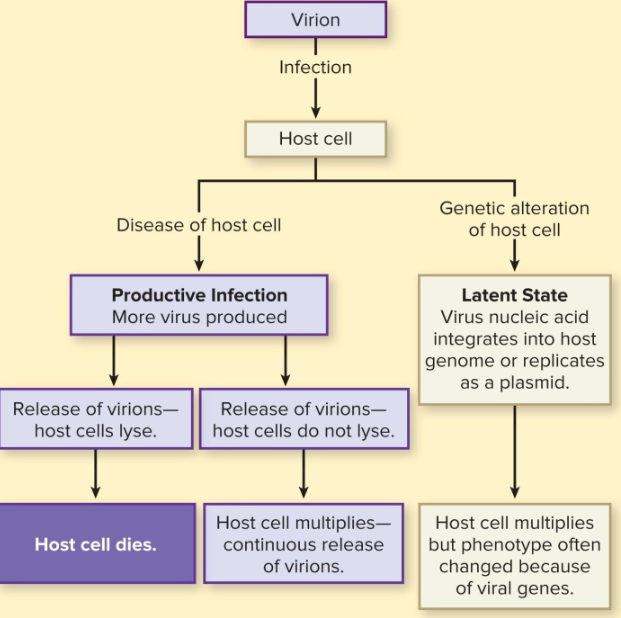
Lytic Phage
lytic/virulent phage (productive infection)
productive infections → new viral particles
30 min cycle
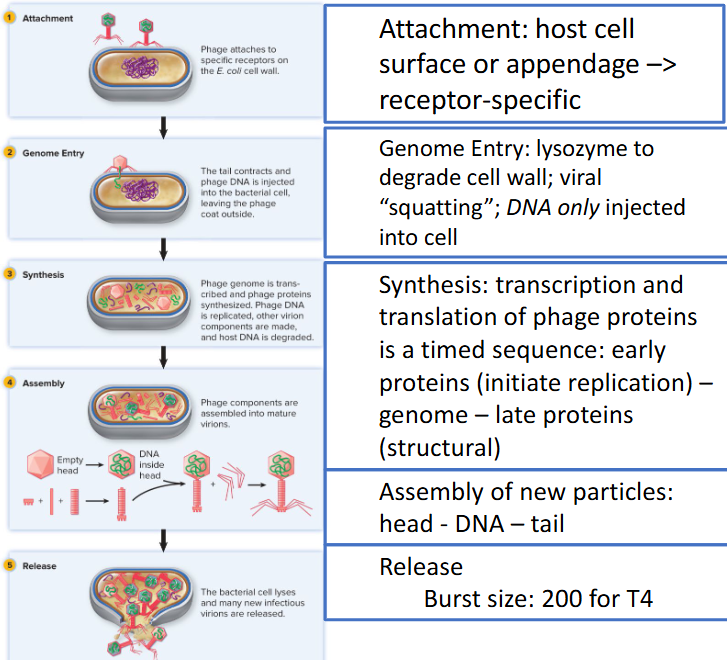
T4 model — dsDNA; E. coli
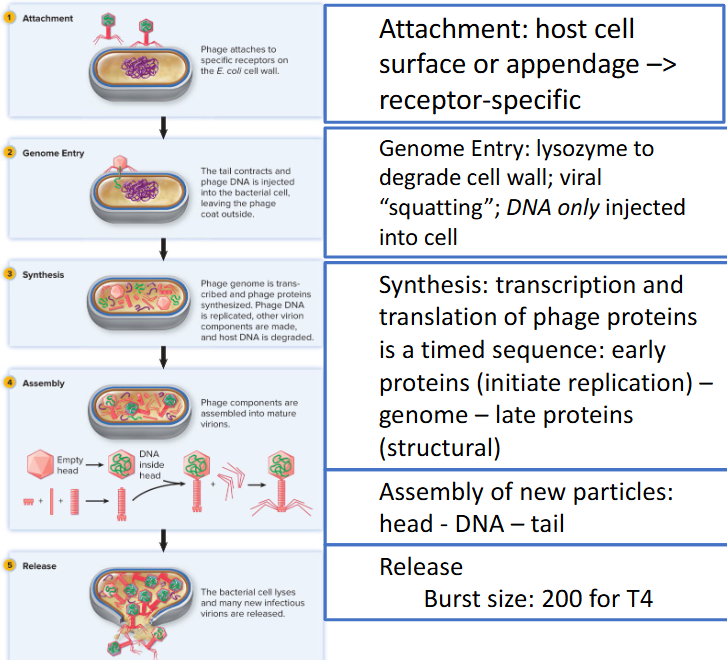
Lytic/virulent Phage
productive infection
exit host at end of infection via cell lysis
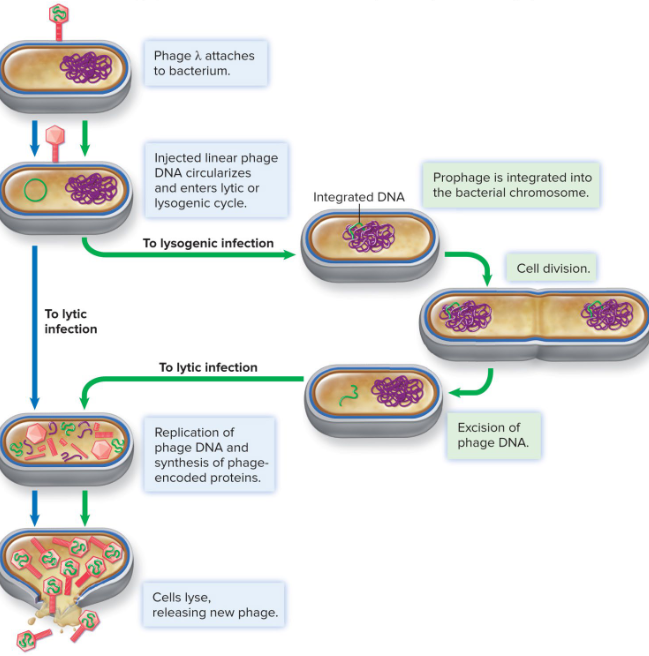
Temperature Phage Infections
2 choices = Lytic vs Lysogenic
Lysogenic Phage Infection
incorporation of phage genome into host genome (no damage in host cell)
Lysogen
prophage — integrated phage DNA
prophage passed to bacterial daughter cells
metabolic state of cell can influence which cycle occurs
Lambda as Temperate Model
linear chromosome that circularizes upon entry
integration via phage integrase
repressor protein regulars excision gene and maintains lysogenic state
role of DNA-damaging agents in phage induction
Filamentous Phage
M13
productive infections but do NOT kill host
M13 = F pilus of E.coli
single stranded DNA upon entry
dsDNA by host DNA polymerase = “replicative form”
extrusion
Extrusion
process by which virus exits cell
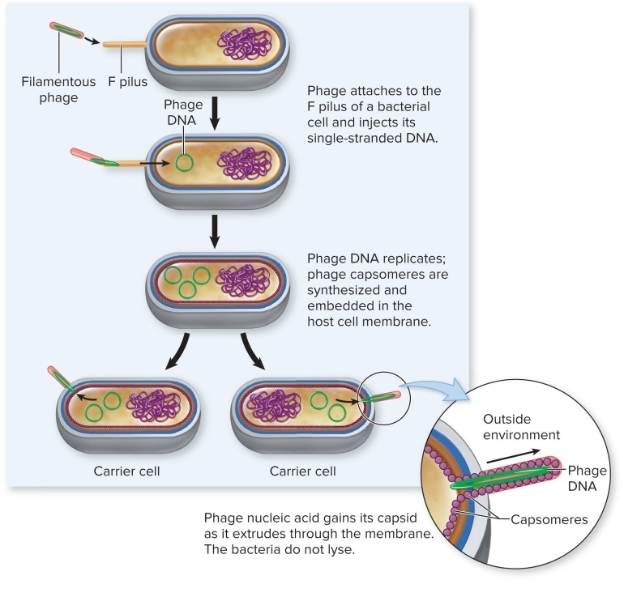

Filamentous phages…
are extruded (formed) from the host cell

All phages must have the ability to…
1. have their nucleic acid enter the host cell
have their nucleic acid replicate in the host cell
Animal Virus Replication
viral enzymes can serve as targets for antiviral drugs
infection cycle similar to that of phages:
attachment
penetration and uncoating (Genome Entry)
synthesis of proteins & genome replication
assembly
release
Attachment of Viruses
spikes bind to specific receptor on cytoplasmic membrane with receptor specificity
Tissue Tropism
viruses can only enter specific cell types
Host Range
the variety of species (hosts) a virus can infect
Measles
2 cell surface receptors
Cd46 = widely expressed in all human nucleated cells
SLAM — signaling lymphocyte activation molecule
Rabies
mammalian nervous system = nicotinic acetylcholine receptor
humans, dogs, horses, skunks, foxes, etc.
Penetration & Uncoating (Genome Entry)
entry dependent on envelope vs non-enveloped
different from phage — entire virion enters cell
enveloped viruses can enter via (1) Membrane Fusion or (2) Endocytosis
non-enveloped viruses = endocytosis only
all viruses = uncoating after entry
Membrane Fusion
mechanism of penetration in which viral envelope fuses with cytoplasmic membrane and the nucleocapsid is released into the cytoplasm
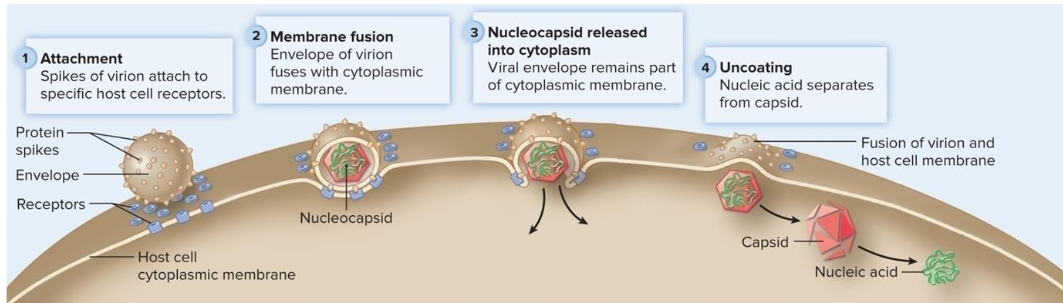
Endocytosis
mechanism of penetration in which viruses capitalize on receptor-mediated endocytosis, a normal cellular process
viruses bind to receptors that trigger endocytosis
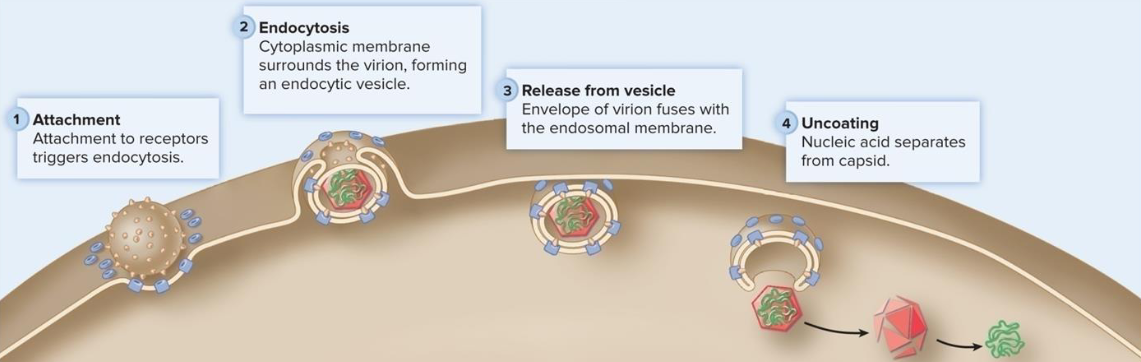
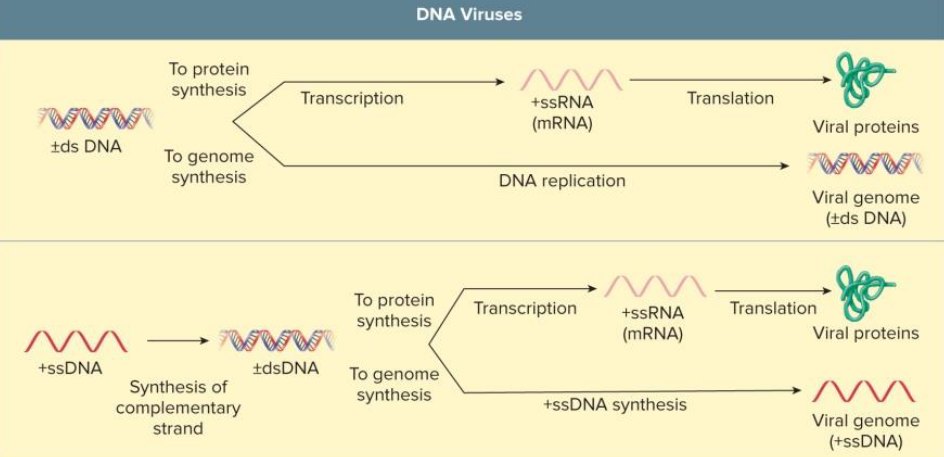
DNA Virus Replication
in nucleus
host cell separates and carries own DNA polymerase
recall (-) DNA strand is the template for mRNA
ssDNA → 2nd strand must be synthesized
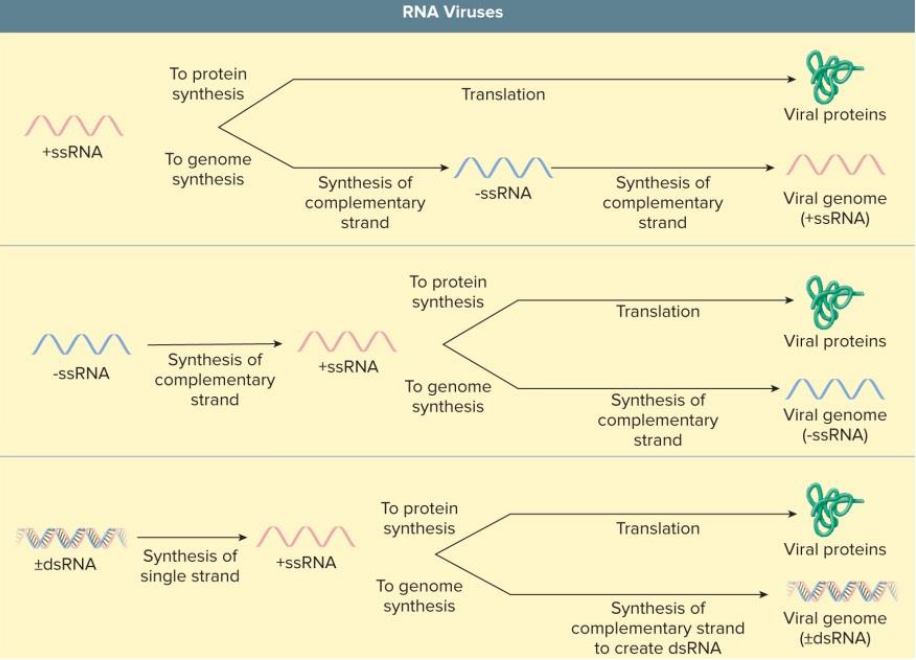
RNA Virus Replication
in the cytoplasm
requires viral RNA polymerase
replicase = RNA-dependent RNA polymerase
antigenic variation due to lack of proofreading
influence and antigenic drift
antigenic shift in segmented genome
Reverse Transcribing Virus (Retrovirus) Replication
retroviruses encode reverse transcriptase = RNA-dependent DNA polymerase
DNA from RNA template
HIV = +RNA = carries RTase in virion
permanent integration into host chromosome
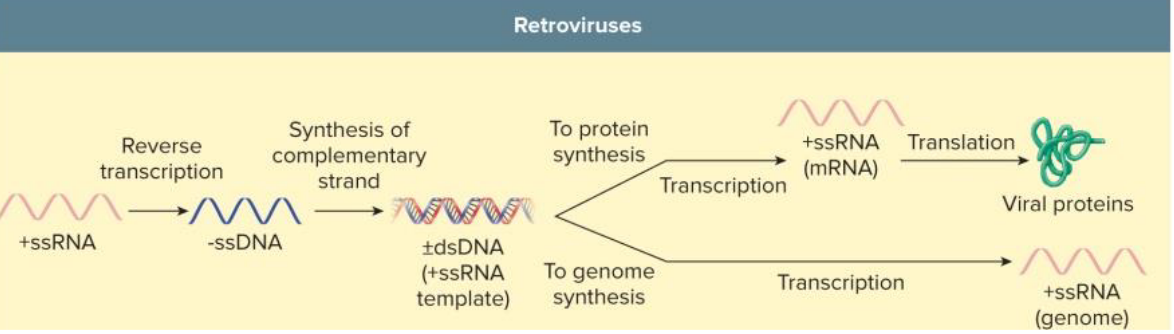
Assembly & Maturation of Viruses
packaging nucleic acid with capsid proteins
spontaneous self-assembly
different sites of assembly and maturation based on virus type
non-enveloped = cytoplasm
enveloped = part of maturation can occur as virion exits cell
DNA viruses = nucleocapsid in host cell nucleus
Release of Viruses
mechanism dependent on whether virus enveloped or not
enveloped viruses → budding
non-enveloped → apoptosis
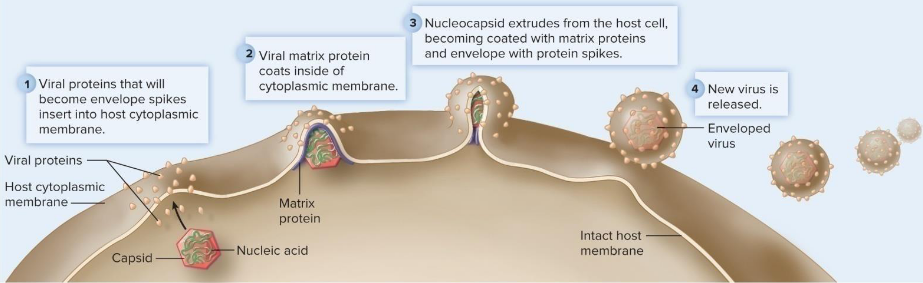
Viral Spread
influenza = coughing and sneezing
norovirus & rotavirus = fecal-oral (contaminated food & water)
HIV = sexual contact
vectors = organisms that carry the virus
Host Range
specificity determines the host range
receptor-binding is major determinant of host range (specific binding btwn viral capsid proteins & specific host cell receptors)
receptor recognition by viruses = 1st and essential step of viral infections of host cells
Narrow Host Range
HIV — humans
surface protein gp120 interacts with CD4 molecule on human leucocytes
Broad Host Range
cucumber mosaic virus — 1000 plant species
influence A virus — mammals and birds
Sialic Acids (SAs) of cell surface glycoproteins and glycolipids are the receptors for the influenza virus, recognized by the viral glycoprotein hemagglutinin
Coronavirus
large family of viruses with broad host range
cause respiratory, gastrointestinal, and central nervous system diseases
contain and envelope, helical capsid, and single-stranded, positive-sense RNA genome
3 major genera = a, B, y (alpha beta gamma)
a = human coronavirus NL63, porcine respiratory coronavirus
B = severe acute respiratory syndrome coronavirus (SARs-CoV), Middle East respiratory syndrome (MERS-CoV), bovine (BCoV)
y = avian infectious bronchitis virus (IBV)
MERS-CoV, SARS-CoV, and SARS-CoV-2 are the exceptions of animal to human infection
Coronavirus Specificity
coronaviruses recognize variety of host receptors
receptor binding domains = S1 subunit of spike proteins both N-term (S1-NTD) and C-term (S1-CTD)
recognize 4 protein receptors & 3 sugar receptors
coronavirus spike protein binds to receptor on host cell surface then fuses viral and host membranes
spike has 3 parts = ectodomain, transmembrane anchor, tail
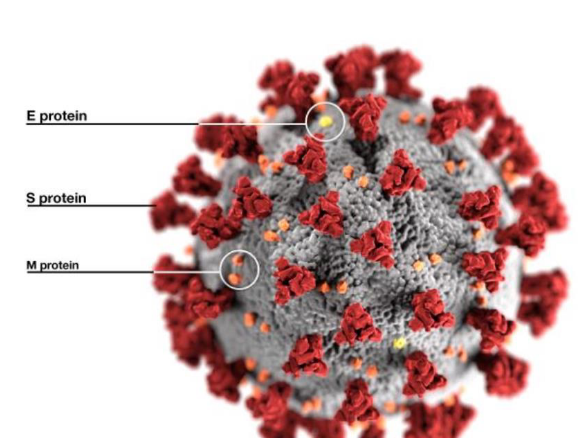
SARS-CoV-2 Virus Characteristics
S protein protrude from the viral surface resembling a crown or corona
an enveloped virus with an unsegmented single-stranded positive-sense RNA genome
4 main structural proteins
spike (S) glycoproteins
envelope (E) glycoproteins
membrane (M) glycoproteins
nucleocapsid (N) proteins
genome codes for 16 nonconstructural proteins involved in viral replication, maturation, and release
Ectodomain
S1 subunit binds to receptor
2 domains in which either or both function as receptor-binding domain
S1-N/CTD (N-term vs C-term)
S2 subunit for membrane fusion
receptor-binding domain can serve as vaccines against coronavirus infections
Virus Entry into Hosts
portals of entry = nasal, oral, conjunctiva of eyes passing through nasolacrimal duct, esophagus
once transmitted, viral S-proteins bind to host cells via angiotensin-converting enzyme 2 (ACE2) (viral tropism)
S-proteins embedded in viral envelope bind to ACE2 on host cells triggering viral endocytosis, or membrane fusion and viral genome entry
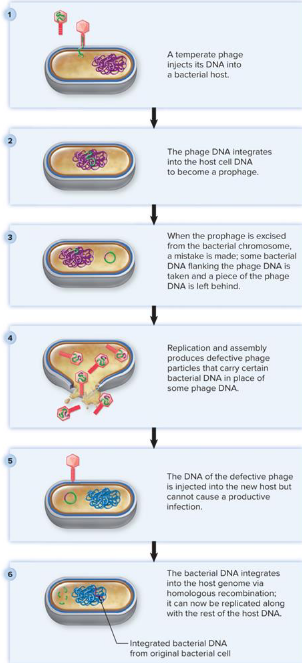
Horizontal Gene Transfer via Bacteriophages
generalized transduction
specialized transduction
differences based on type of error
Generalized Transduction (Horizontal Gene Transfer via Bacteriophages)
packaging error during assembly = bacterial rather than phage
homologous recombination
any bacterial gene can be transferred (“generalized transduction”)
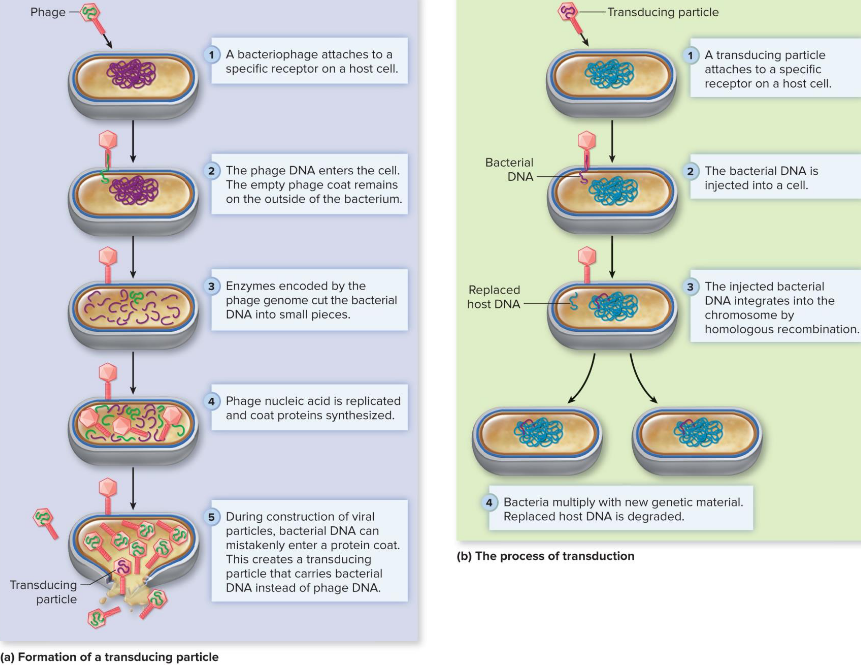
Specialized Transduction (Horizontal Gene Transfer via Bacteriophages)
excision error by prophage
bacterial + phage DNA genes
“specialized” = only bacterial genes adjacent to phage genes transferred
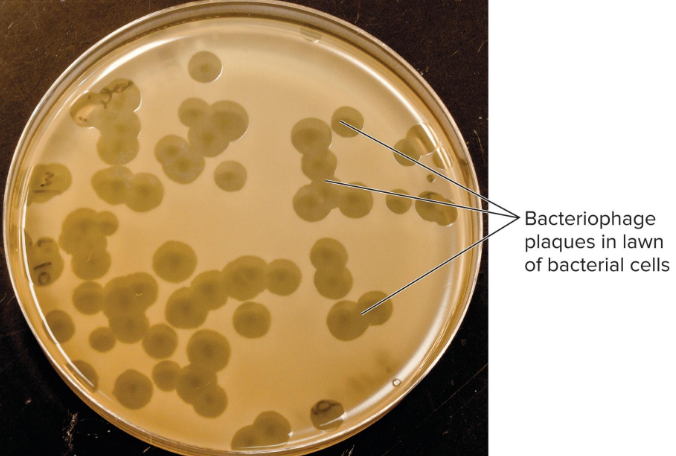
Plaque Assay
method of detecting and quantifying phages
bacterial host + phage sample
based on zone of clearing
used to determine titer within sample
titer = laboratory measurement that determines the concentration of a specific substance, usually antibodies, in a blood sample
Innate Immunity
immunity at birth, germ-line encoded
Adaptive Immunity
immunity vis antigens and antibodies
First Line (Immune) Defenses
physical barriers, skin
mucous membrane
antimicrobial substances
normal microbiota
Epidermis
many layers of epithelial cells, keratin
Sensory Systems (Immune Defense)
sentinel cells = pattern recognition receptors, other PRRs, cytoplasm
complement system = circulates in blood and tissue in inactive form
Innate Effector Actions (Immune Defense)
interferon response
phagocytosis
complement activation
inflammatory response
fever
Phagocytes
cells that engulf and digest microbes and other foreign debris via phagocytosis
Bodily Antimicrobial Substances
lysozyme
lactoferrin/transferrin
antimicrobial peptides
defensins = + charge
AMPs and defensins → destroy microbes, other roles in immune system, vitamin D / AMP regulation
Hematopoietic Stem Cell
from which all blood cells arise
bone marrow
Immune System Cells
Hematopoietic stem cells
red blood cells (erythrocytes) → O2
platelets → clotting
white blood cells (leukocytes) → defense
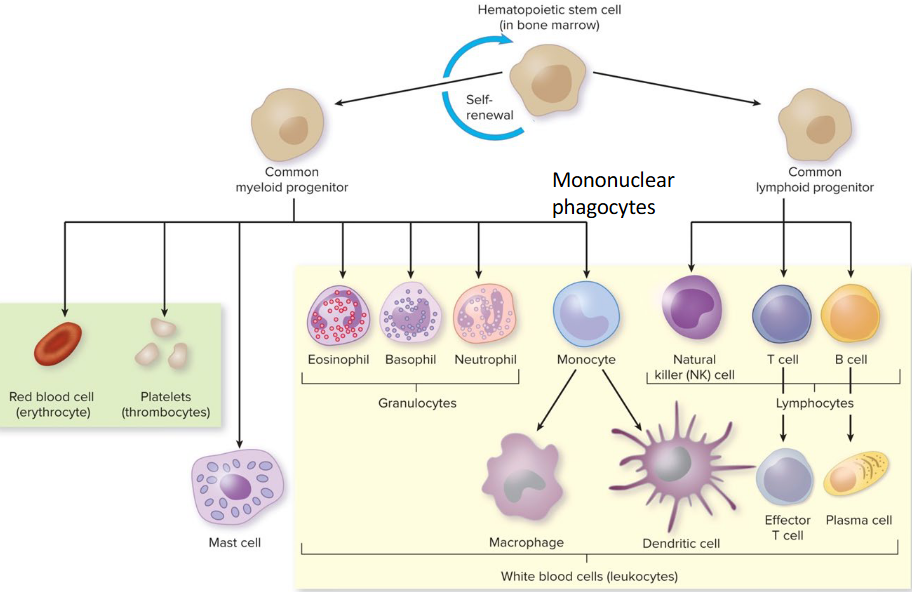
Granulocytes
type of leukocyte
cytoplasmic granules containing compounds important for cell protection
neutrophils, eosinophils, basophils