BI1014 - nucleic acids and edge of life
1/200
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
201 Terms
significance of DNA
- DNA (deoxyribonucleic acid) is the molecule that carries the genetic instructions
- It serves as the hereditary material in almost all organisms.
- DNA contains genes, which are specific sequences that code for proteins and functional RNA
- DNA ensures stability of genetic information and enables variation through mutation, supporting evolution
role of DNA
Protein synthesis (via transcription and translation)
Cell division and replication
Inheritance of traits from one generation to the next
structure of DNA
- DNA is made of nucleotides, which are the basic subunits
- The phosphate and sugar form the backbone of the DNA strand.
- The bases project from the backbone and are involved in base pairing.
structure of nucleotides
A phosphate group
A deoxyribose sugar (5-carbon sugar)
A nitrogenous base (one of four types):
nitrogenous bases
Adenine (A)
Thymine (T)
Cytosine (C)
Guanine (G)
formation of double stranded DNA
- DNA is double-stranded: two polynucleotide chains run in opposite directions (antiparallel).
- The strands are held together by hydrogen bonds between complementary nitrogenous bases
- Complementary base pairing ensures accuracy in replication and transcription.
- The formation of the double strand results in a stable helical structure.
base pairs
Adenine (A) pairs with Thymine (T) via 2 hydrogen bonds
Guanine (G) pairs with Cytosine (C) via 3 hydrogen bonds
key features of double helix
Antiparallel strands (one 5′ → 3′, the other 3′ → 5′)
Major and minor grooves: Allow proteins to interact with bases without unwinding DNA.
10 base pairs per turn of the helix (in B-DNA)
Diameter of helix: ~2 nanometers
key features of double helix
- DNA typically adopts a right-handed double helix known as B-DNA.
- The double helix is compact and stable, ideal for long-term storage of genetic information.
- The structure allows for easy unwinding during replication and transcription due to hydrogen bonding.
B-DNA
- Most common form under physiological conditions.
- Right-handed helix with ~10 base pairs per turn.
- Wide major groove and narrow minor groove
A-DNA
- Forms in dehydrated conditions or in DNA-RNA hybrids.
- Right-handed helix, more compact than B-DNA.
- ~11 base pairs per turn; deep major groove, shallow minor groove.
Z-DNA
- Left-handed helix with a zigzag backbone.
- Forms in regions rich in alternating purine-pyrimidine sequences (e.g., GC repeats).
- May play a role in gene regulation and chromatin remodelling
triplex and quadruplex DNA
Unusual structures formed in specific sequences or under special conditions
triplex DNA
Three-stranded structures often found in regulatory regions
other structural features
- Hairpin loops -> in palindromic sequences
- Cruciform structures -> from inverted repeats
- Triple-stranded DNA (triplex)
- G-quadruplexes
G-quadruplexes
- Found in telomeres and regulatory regions
- Involves stacked guanine tetrads
denaturation
separation of double-stranded DNA into single strands
key factors
- temperature
- GC content
- salt concentration
- pH levels
- chemical agents
temperature
- Higher temps → denaturation
- Melting temperature (Tm): temp at which 50% of DNA is denatured
GC content
- GC pairs (3 H-bonds) more stable than AT pairs (2 H-bonds)
- Higher GC content → higher Tm
salt concentration
- Stabilizes DNA by shielding negative phosphate charges
- Lower salt → easier denaturation
pH content
Extreme pH (acidic or basic) disrupts hydrogen bonding
chemical agents
Urea and formamide disrupt H-bonds → lower Tm
sugar
DNA: deoxyribose.
RNA: ribose (extra hydroxyl group at 2').
bases
DNA: adenine (A), guanine (G), cytosine (C), thymine (T).
RNA: uracil (U) replaces thymine.
strandedness
DNA: double-stranded (usually).
RNA: single-stranded (but may fold into secondary structures).
stability
DNA: more stable due to fewer reactive groups.
RNA: less stable, especially in alkaline conditions.
function
DNA: genetic information storage.
RNA: diverse roles (mRNA, tRNA, rRNA, regulatory RNAs).
location
DNA: mainly in nucleus (eukaryotes).
RNA: nucleus and cytoplasm.
genome
the complete set of genetic material in an organism
plasmid
small, circular DNA molecules separate from the chromosomal DNA
operon
a cluster of genes under the control of a single promoter (common in prokaryotes)
gene
a DNA sequence that codes for a protein or functional RNA
prokaryotic genome properties
- Typically circular, double-stranded DNA.
- Often haploid (one copy per cell).
- Compact - very little non-coding DNA.
- Genes often organized in operons
prokaryotic genome properties
- DNA is supercoiled and associated with non-histone proteins.
- Some have plasmids for antibiotic resistance, virulence, etc.
- No membrane-bound nucleus - DNA located in nucleoid region.
challenge
Eukaryotic cells contain large amounts of DNA (~2 meters per cell) that must fit inside a tiny nucleus (~10 μm)
levels of DNA packaging
1. nucleosome
2. chromatin fibre
3. looped domains
4. higher-order folding
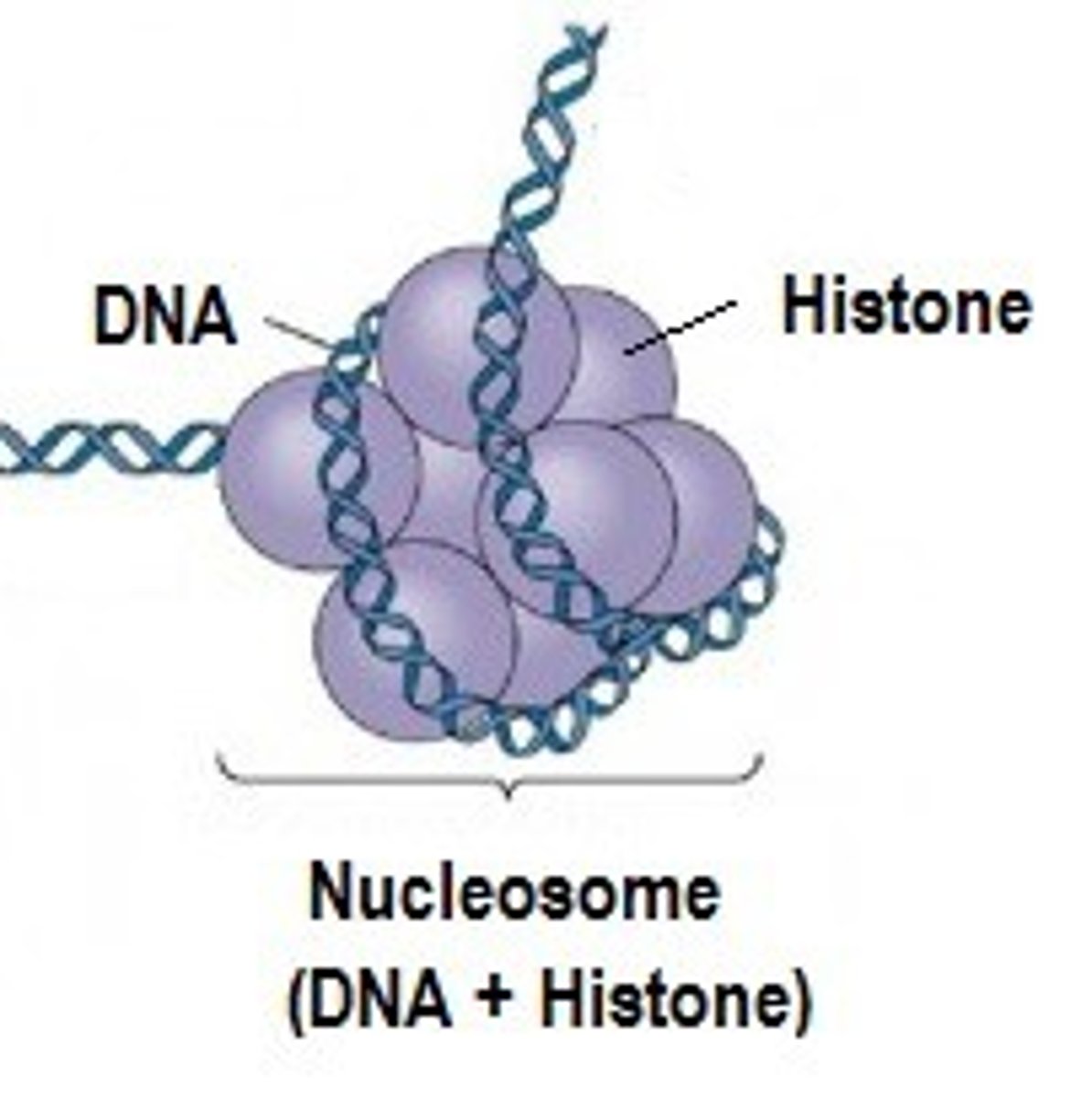
nucleosome
(1st level):
- Basic unit of DNA packaging.
- ~147 base pairs of DNA wrapped around a histone octamer (2 each of H2A, H2B, H3, and H4).
- Resembles "beads on a string" under electron microscopy.
chromatin fibre
(30 nm fibre):
- Nucleosomes are coiled into a solenoid or zigzag structure.
- Stabilized by Histone H1.
looped domains
Chromatin forms loops attached to a protein scaffold (e.g., scaffold-associated regions or SARs)
higher order folding
- Loops further fold and condense, especially during mitosis.
- Results in metaphase chromosomes, the most condensed state
types of chromatin
- euchromatin
- heterochromatin
euchromatin
- Loosely packed.
- Transcriptionally active.
heterochromatin
- Densely packed.
- Transcriptionally silent.
- Includes constitutive heterochromatin (always inactive, e.g., centromeres) and facultative heterochromatin (can be activated or inactivated).
epigenetic regulation
- DNA packaging is dynamic and regulated by histone modifications (e.g., methylation, acetylation) and DNA methylation.
- These changes affect gene accessibility and expression.
purpose of DNA replication
To create identical copies of DNA before cell division.
key feature of DNA replication
Semi-conservative: Each daughter DNA has one old (parental) strand and one new strand
size of eukaryotic genome
- Much larger than prokaryotic genomes.
- Size varies widely across species (not directly proportional to organism complexity).
structure of eukaryotic genome
- Linear chromosomes (in nucleus).
- DNA packaged into chromatin with histones.
- Typically diploid (two copies of each chromosome).
coding DNA
- Genes
- Only ~1-2% of the human genome codes for proteins.
genes
- DNA sequences transcribed into RNA, many translated into proteins.
- Contains exons (coding sequences) and introns (non-coding regions within genes).
non coding DNA
introns, regulatory sequences, intergenic regions, repetitive DNA
introns
Removed from pre-mRNA during splicing
regulatory sequences
Promoters, enhancers, silencers - control gene expression
intergenic regions
DNA between genes; includes regulatory elements, repetitive DNA, and unknown functions
repetitive DNA
- Tandem repeats (e.g., microsatellites, minisatellites).
- Interspersed repeats (e.g., transposable elements like LINEs and SINEs).
- Over 50% of human genome is repetitive.
mtDNA
Mitochondrial DNA (mtDNA)
- Circular, double-stranded DNA.
- Encodes genes for components of oxidative phosphorylation, rRNAs, and tRNAs.
- Inherited maternally.
- Replicates independently of nuclear DNA.
chloroplast DNA
(in plants and algae)
- Also circular.
- Contains genes for photosynthetic proteins, tRNAs, and rRNAs.
- Similar to prokaryotic genomes, supporting the endosymbiotic theory.
unique features of eukaryotic genomes
alternative splicing, gene families, pseudogenes, epigenetic modifications
alternative splicing
allows one gene to produce multiple proteins.
gene families
similar genes with related functions (e.g., haemoglobin genes)
pseudogenes
former genes that have lost their function due to mutations.
epigenetic modifications
like DNA methylation and histone modification regulate genome accessibility and expression
function of DNA polymerase
DNA polymerases catalyse the synthesis of new DNA strands during replication
features of DNA polymerase
direction of synthesis, template requirement, primer requirement, substrates
direction of synthesis
always 5' -> 3'
template requirement
Needs a single-stranded DNA template
primer requirement
Needs a pre-existing 3'-OH group to add nucleotides (RNA primer synthesized by primase)
NOT IN HUMANS
substrates
Deoxynucleoside triphosphates (dNTPs)
catalytic activity of DNA polymerase
- Adds nucleotides by forming phosphodiester bonds between the 3'-OH of the growing strand and the 5'-phosphate of the incoming dNTP.
- Releases pyrophosphate (PPi) as a byproduct
proofreading
- Most DNA polymerases have 3' → 5' exonuclease activity for error correction.
- Enhances fidelity of DNA replication
different types of DNA polymerase
(example in E. coli):
DNA Pol I: Removes RNA primers and fills in gaps.
DNA Pol III: Main replicative enzyme; highly processive and fast.
dAMP
Deoxyadenosine monophosphate
dCMP
Deoxycytidine monophosphate
dGMP
Deoxyguanosine monophosphate
dTMP
Deoxythymidine monophosphate
supercoiling
- If DNA is underwound or overwound, it will become supercoiled - the molecule twists around itself
- Underwinding generates negative supercoils
- Overwinding generates positive supercoils
nucleosomes
- When interphase nuclei are lysed, and their contents viewed through the electron microscope we see this 'beads on a string' structure is sometimes known as the 10nm filament
- The string is DNA
- The beads are nucleosomes
nucleosome structure
A nucleosome us 145bp DNA wrapped around "core particle" containing
- 2 molecules of histone H2A
- 2 molecules of histone H2B
- 2 molecules of histone H3
- 2 molecules of histone H4
Nucleosomes can be thought of as subunits of chromosomes and chromatin
Meselson Stahl experiment
- Grew E. coli in growth medium containing a heavy isotope of nitrogen (14N) -> 15N, the DNA in these cells is therefore denser than normal
- Switched the E. coli to a medium containing 14N, all DNA synthesised after the switch will therefore be less dense than pre-existing DNA
- Measures the density of DNA after DNA replication
dna polymerases
The enzymes that synthesise DNA are called DNA polymerase
DNA replication in e coli
replication origin, initiation, elongation, termination
replication origin
- Starts at a single origin called oriC (~245 bp long).
- Recognized by the initiator protein DnaA.
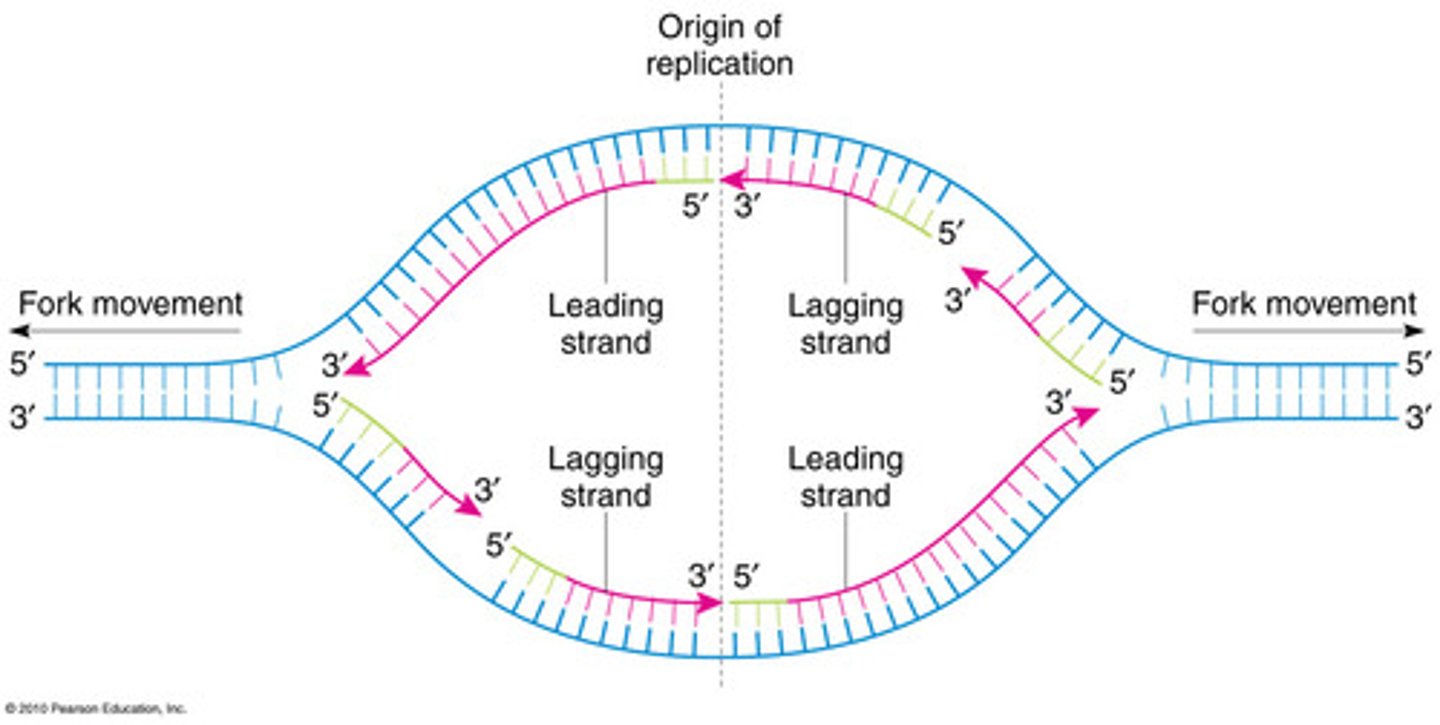
initiation
- DnaA binds oriC → DNA unwinds.
- Helicase (DnaB) is loaded with help of DnaC → unwinds the helix.
- Single-stranded binding proteins (SSBs) prevent reannealing.
- Primase (DnaG) synthesizes short RNA primers.
elongation - enzyme 1
DNA Pol III holoenzyme:
- Leading strand: synthesized continuously in 5’ → 3’ direction.
- Lagging strand: synthesized discontinuously in Okazaki fragments.
elongation - enzyme 2
DNA Pol I:
- Removes RNA primers (via 5’ → 3’ exonuclease activity).
- Fills in gaps with DNA.
elongation - enzyme 3
DNA ligase:
- Joins Okazaki fragments by sealing nicks in the sugar-phosphate backbone.
termination
- Replication ends at ter sequences.
- Tus proteins bind ter sites and block replication fork progression.
- Result: Two identical circular DNA molecules.
OriC
bacteria - single origin
eukaryotes - multiple origins per chromosome
replication rate
bacteria - fast
eukaryotes - slow
DNA polymerases
bacteria - fewer e.g pol I and pol III
eukaryotes - many types
initiator proteins
bacteria - DnaA, DnaB, DnaG
eukaryotes - ORC (origin recognition complex), MCM complex
primase
bacteria - DnaG (separate enzyme)
eukaryotes - Part of DNA Pol α complex
lagging strand processing
bacteria - Pol I removes primers
eukaryotes - RNase H and FEN1 remove primers
chromosome structure
bacteria - circular
eukaryotes - linear
end replication problem
bacteria - absent
eukaryotes - present (solved by telomerase)
termination
bacteria - ter sites with tur proteins
eukaryotes - no specific termination site, forks eventually meet and end replication
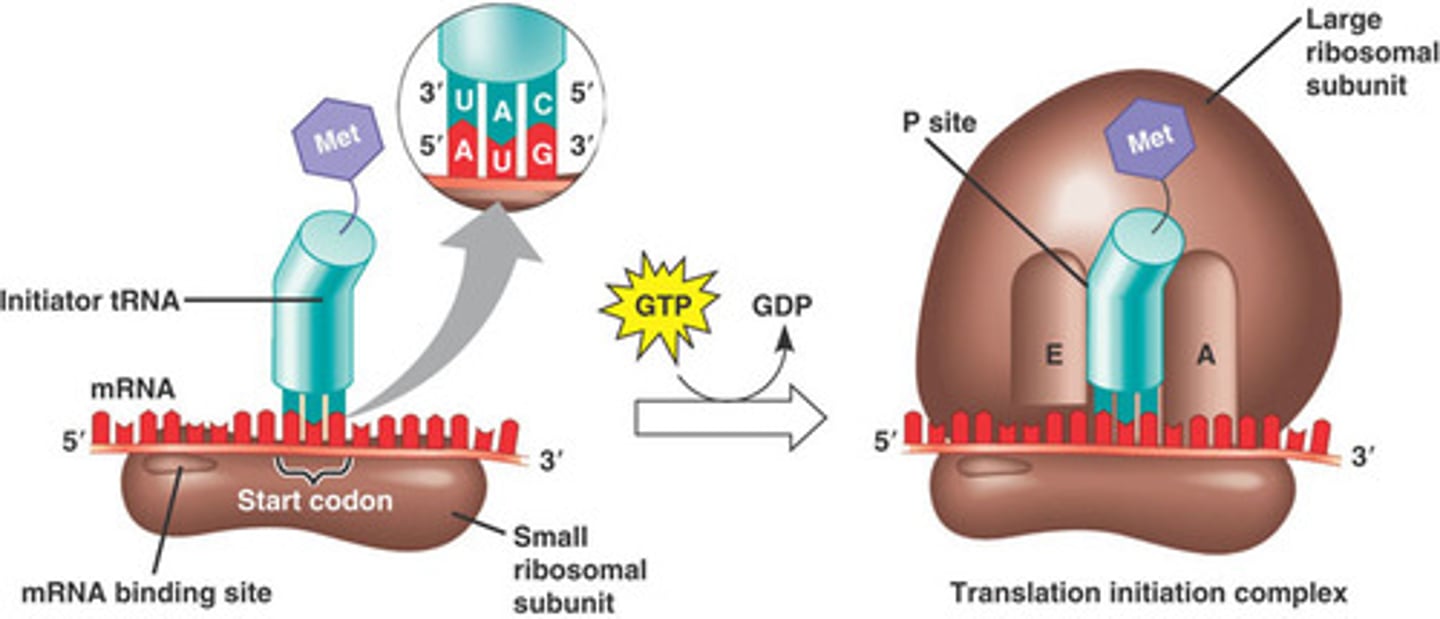
transcription in e coli
- Transcription = synthesis of RNA from a DNA template.
- Carried out by RNA polymerase.
- Occurs in cytoplasm (no nucleus in E. coli).
RNA polymerase in e coli
- Core enzyme: α₂ββ'ω – performs elongation.
- Holoenzyme: Core enzyme + σ factor – required for initiation and promoter recognition.
- Common σ factor: σ⁷⁰ (recognizes standard promoters).
stages of transcription in e coli
initiation , elongation, termination