Lecture 8: Plant disease resistance
1/43
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
44 Terms
why is plant breeding needed- what traits are need to be aknowledged
- Increasing population- agricultural area is the same—> increase in productivity is necessary→ eg from plant breeding certain traits eg for drought, salt resistance, plant disease resistance as well as herbicide resistance etc
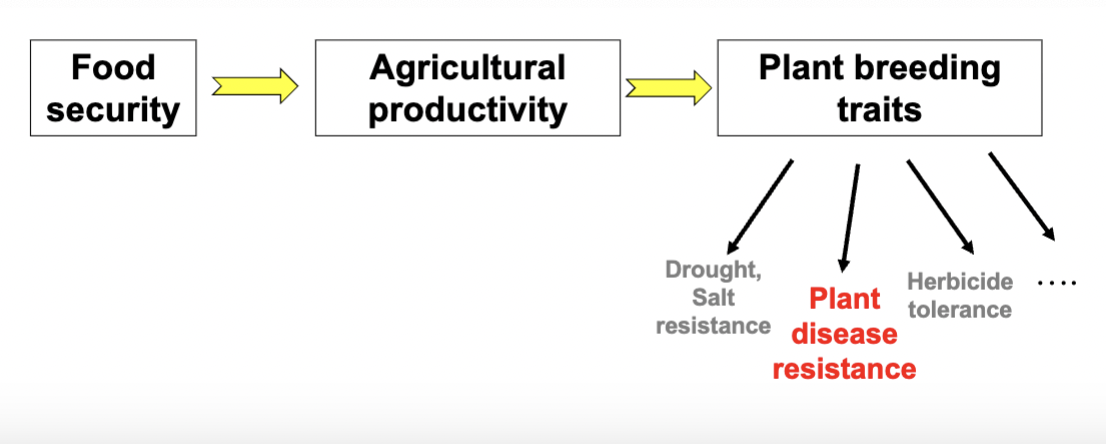
how is food security dependent on plant breeding?
Food security is closely dependent on agricultural productivity which heavily is dependent on plant breeding traits (determining agricultural yield)- plant breeding traits that can favour yield include resistance to abiotic stress, herbicide tolerance as well as plant disease resistance.
what questions need to be considered in disease resistance assesments?
Who are the enemies (pathogens)?, How do they make plants sick? How do plants sense pathogens? How do plants defend themselves? How can we help increase disease resistance? Which genes? How to express them?
explain what kinds of threats can cause disease in plants? what ways?
- Pathogens and disease greatly affect productivity. (pathogens include viruses, bacteria, fungi, insects, nematodes)
- From the field: crop disease can for instance only affect the leaves- negative impact because it inhibits photosynthesis- even though the leaves themselves are not harvested it impact the function and survival of the plant through inhibiting photosynthesis
- Other diseases affect not only the leaves, but other parts as well- even more serious: eg potato blight, tomato bacteria spot
Various types of fungi, viruses and bacteria can cause a range of different disease and cause crop yield loss to different extent and in different ways
what is Agrobacterium tumefaciens
a soil bacterium: pathogen for wide range of dicots plants: one major pathogen.
- Parasite!
- Cause crown gall
- Disease/tumours are caused by a fragment of DNA (T-DNA) from the Ti-plasmid in the bacterium
- Wide range of hosts: some common ones are different types of fruit plants, rhododendron etc
- Can bee seen by galls on the stem/roots which are the tissues primarily targeted—> however the plant will suffer in other ways too such as floral display or fruit production becoming suppressed. Severely diseased plants are more vulnerable to secondary pathogens that enter through decaying galls
what is the infectious part of the Agrobacterium tumefaciens genome- how can it be used in plant transformations?
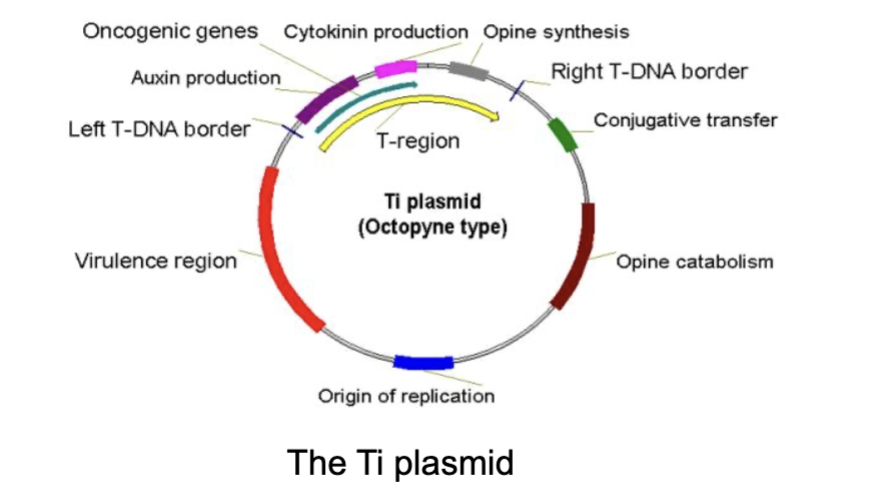
- The bacterium contains a Ti-Plasmid which in turn possess T-DNA genes which encode for excessive cell division/growth - through excessive synthesis of auxin and cytokinin- as well as additional opine synthesis- which the bacteria uses can use as nutrients = parasitism.
- The Ti plasmid can be used when transforming plant cells: clone useful gene into Ti plasmid by replacing genes in the T- DNA (but keeping the left and right borders of the T-DNA intact). —> Then put engineered Ti plasmid back into Agrobacterium and allow it to infect plant cells in culture.
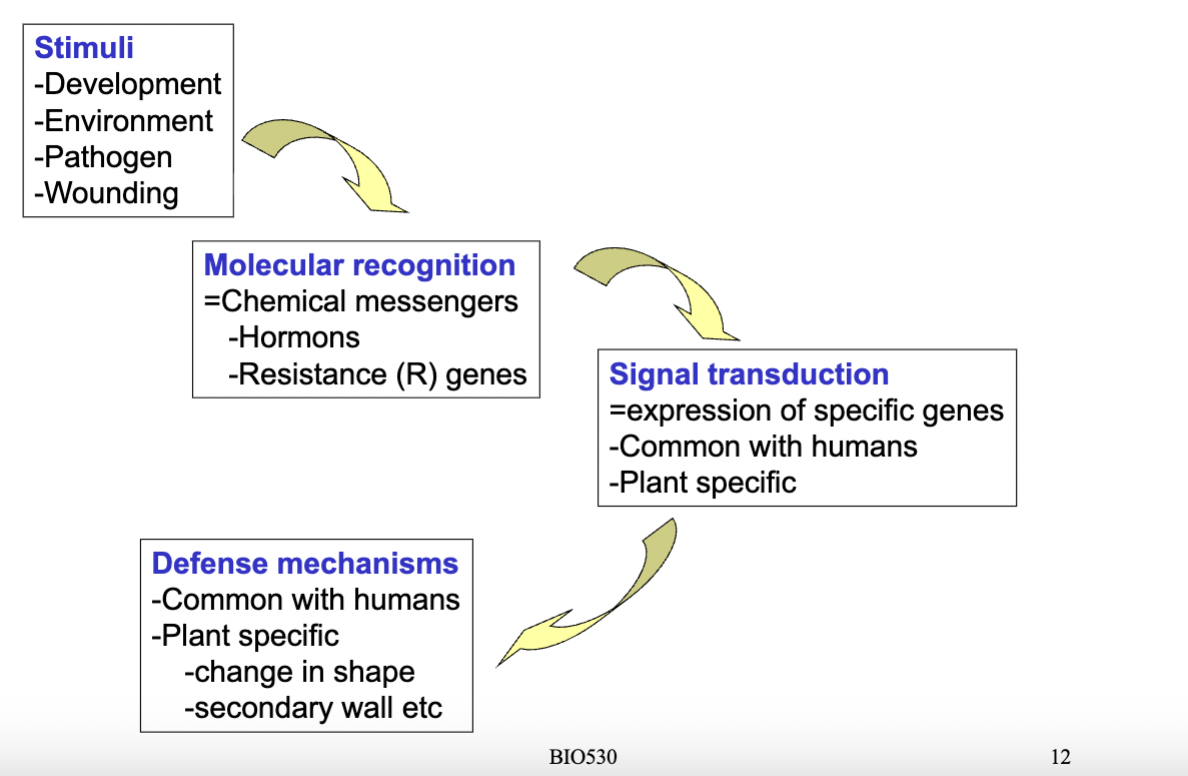
how can the environment be sensed by plants? overwiev
- Environmental factors can be recognized by the plant through receptors (in the case of pathogens there are resistance genes that can recognize pathogen presence)- lead to signal transduction and activate defence mechanisms eg change in shape of the plant eg thicker cell wall to further isolate the plant from the environment- plants recognize and respond to pathogens
more specifically explain what happens from stimuli to defence mechanism response
- The stimulus (could be pathogens, wounding, environmental factors etc) is first recognised at the molecular level through chemical messengers like hormones or resistance (R) genes, which then trigger signal transduction cascades that lead to the expression of specific genes — some of which are shared with animals and some plant-specific. This ultimately results in a defence or adaptive response, which again can be universal (like immune-type responses) or plant-specific, such as physical changes in cell shape or reinforcement of the cell wall
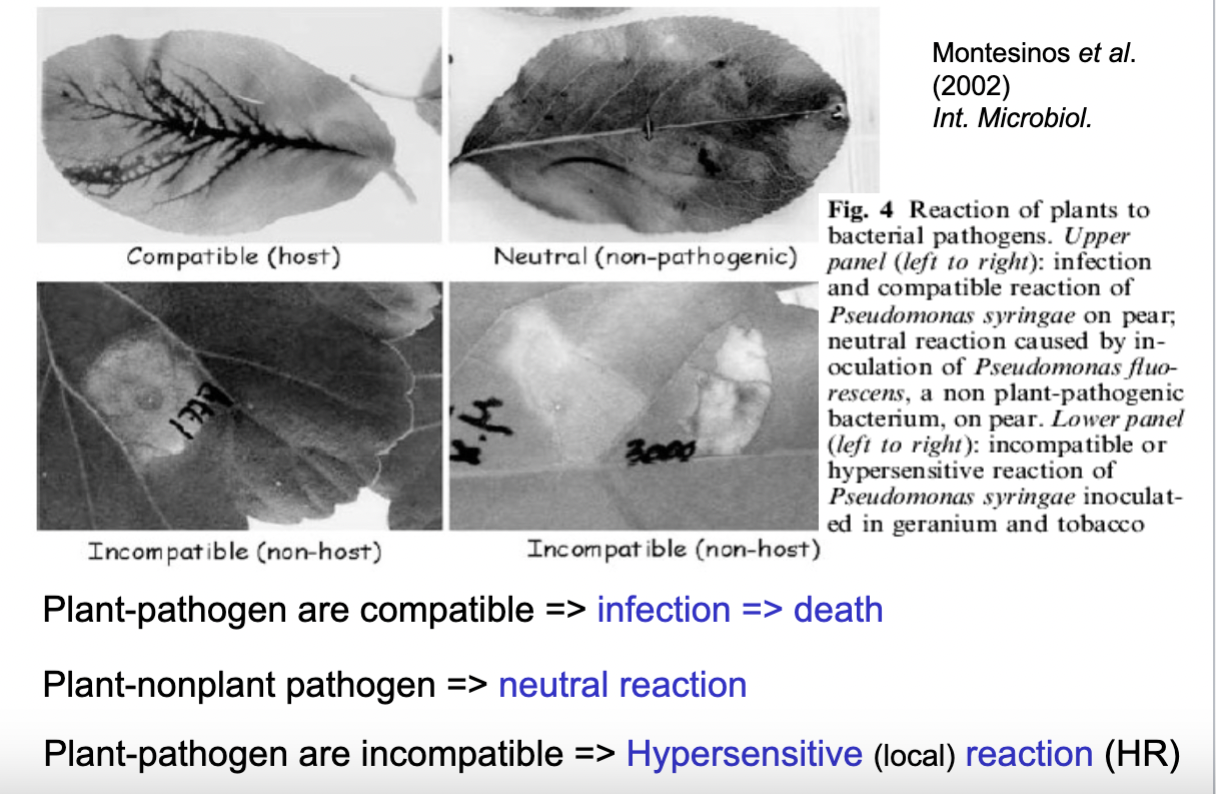
explain 1. compatible, 2. neutral and 3. incompatible plant-pathogen relations- and how the response would look like in three different examples
1. Veins are invaded by bacteria -the plant and pathogen are compatible and hence an infection will occur and lead to death. 2. Non plant pathogen in the plant will cause a neutral reaction because the pathogen is a non plant pathogen. 3. Plant and pathogen are incompatible- the plant recognize the pathogen and respond locally- a hypersensitive (local) reaction (HR) is caused and protect the plant.
- Shortly; when a pathogen attacks the plant there are three possible outcomes:
1. If the pathogen is compatible with the plant — meaning it has evolved to infect that specific host — the plant cannot defend itself effectively and the infection spreads, potentially killing the plant.
2. If the bacterium is simply not a plant pathogen at all, nothing happens and the plant shows a neutral reaction.
3. When the pathogen tries to infect a non-host plant there is an incompatible interaction — here the plant mounts a rapid and aggressive local response called the Hypersensitive Response (HR), essentially killing off the infected cells immediately to prevent the pathogen from spreading. - The plant sacrifices a small area to save itself. (The HR response is possible because the plant is a non-host to the pathogen, hence the pathogen has not evolved mechanisms to undergo the plants immune system and is recognised- triggering the HR/programmed cell death)
what is the purpose of HR? what is it
= A localized reaction in which the invaded cells and perhaps adjacent cells quickly die presumably limiting nutrient uptake by the pathogen and further growth.
Purpose: programmed cell death of the very sight of attack- to limit the nutrient uptaken of the pathogen in order for it to leave/die.
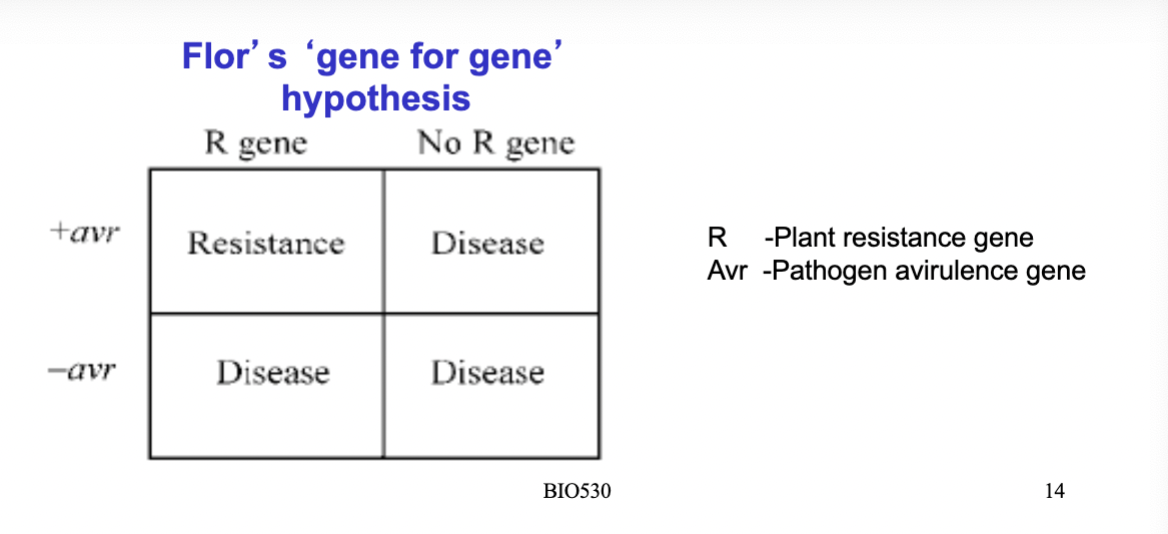
explain how the HR works on gene level- what is needed to get a HR in the plant cell?
resistance gene (R gene) in plant and avirulence gene (Avr gene) in the pathogen:
if the R gene in the plant recognizes the avr gene in the pathogen- signal transduction occur and lead to defence
if no avr gene in the pathogen is recognized by the plants R gene, no defence will occur and infection will be possible.
—> So, in order for the plant to initiate a defence, the plant need to 1. have the R gene that 2. can recognize the pathogens Avr gene

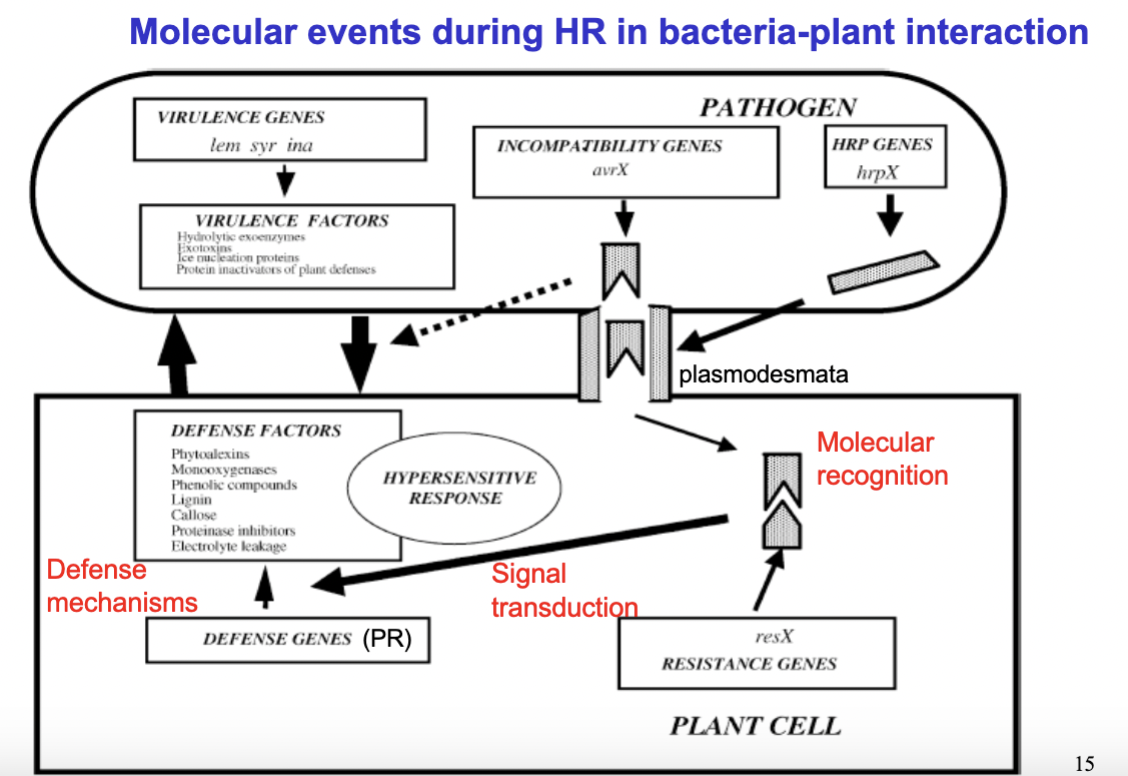
explain the molecular events during HR in plant-bacteria interaction:
- Plant cells: Have a rigid cell wall with openings called plasmodesmata- these only allow transportation of very small molecules- not proteins etc- however, pathogens can enter the plant cells through these plasmodesmata by producing certain protein encoded by the HRP genes which widen the plasmodesmata.
—> Pathogens have virulence genes encoding for virulence factors (toxins, inhibitors of plant defence etc), as well as incompability genes (avrX) which encode avirulence proteins (Avr).
—> The plant has resistance genes (resX) which encode R-proteins, which in turn can recognise the pathogens Avr proteins (in non-host plants).
The molecular recognition (R-proteins that recognize the pathogens Avr protein) lead to signal transduction which ultimately trigger activation of the Hypersensetive response (HR) —> leads (bla) to production of defense factors — phytoalexins, phenolic compounds, lignin, callose and others — which neutralize the pathogen and wall off the infected area. The infected cells rapidly die, starving the pathogen and preventing it from spreading further.
what are the similarities with the resistance (R-) proteins in plants and animal cells?
- Resistance proteins: plant R (Resistance) proteins are structurally and functionally similar to innate immunity receptors found in insects (Drosophila) and mammals —> not unique to plants- have similarity to innate immunity recepters to insects and mammals. plants and animals use the same molecular building blocks (LRR, TIR, kinase domains etc.) for pathogen recognition, but they are combined and arranged differently in each organism to form their specific immune receptors.
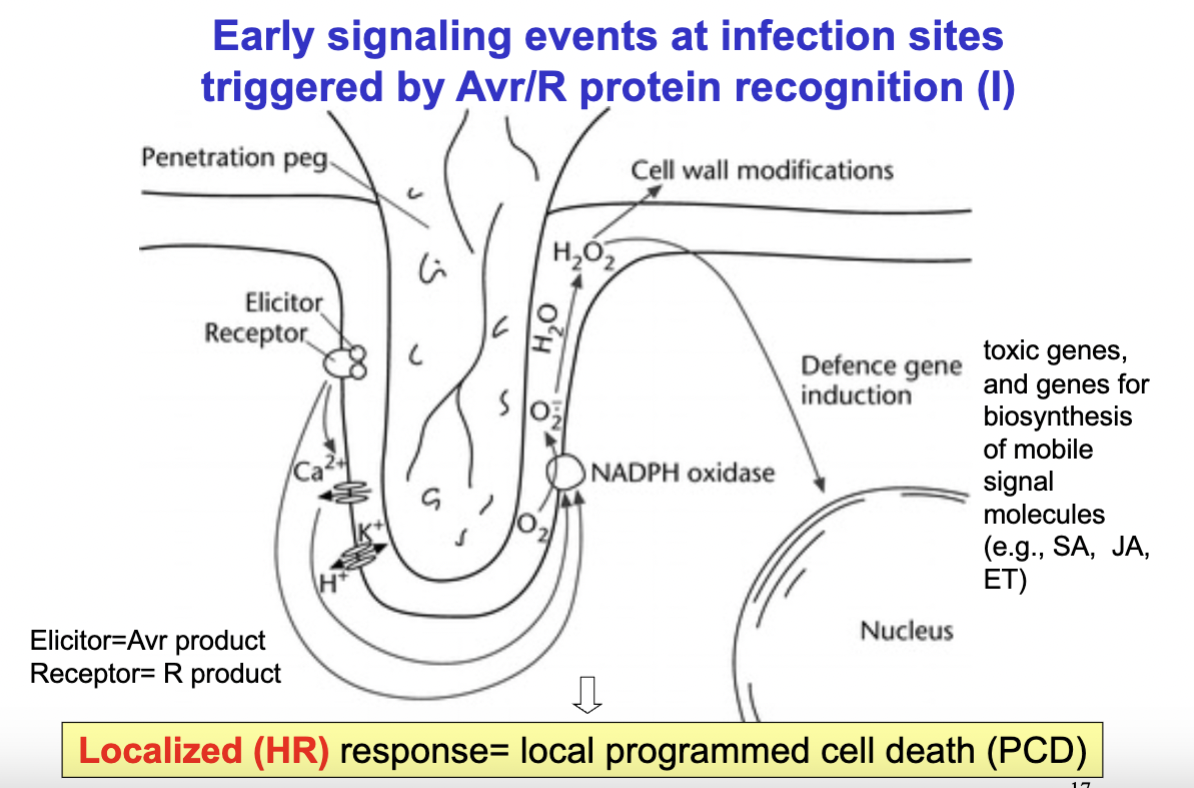
explain what happens in the Early state of infection (bacteria-plant interaction) on the molecular level in detail- what are the products/result
- bacteria tries to penetrate cell wall- bacteria flagell/or surface proteins are recognized by the cell wall receptors- lead to opening of Calcium channels- increase in calcium concentration activate channels and lead to acidification in cytosol which activate NADPH oxidase which lead to creation of reactive oxygen species and can lead to cell wall modification or lead to defence gene induction in the nucleus and lead to the synthesis of certain molecules- mobile signal molecules (bla):
—> in short: pathogen's Avr protein is recognized by the plant's R protein at the plasma membrane—> this triggers an ion flux — calcium (Ca²⁺) flows in and K⁺ and H⁺ flow out, which causes acidification of the cytosol. The calcium signal then activates NADPH oxidase in the membrane, which converts O₂ into reactive oxygen species (ROS), specifically H₂O₂. This is called the oxidative burst. —> The H₂O₂ then has two effects— it triggers cell wall modifications to physically reinforce and block the infection site, and it signals to the nucleus to induce defense genes. These defense genes produce toxic compounds and also biosynthesis of mobile signal molecules — salicylic acid (SA), jasmonic acid (JA), and ethylene (ET) — which can travel to other parts of the plant to warn them and prime their defenses too.- The end result of all of this is programmed cell death (PCD))
what are mobile signal molecules, give examples
- local response lead to local programmed cell death- to continue to next events- mobile molecules are able to move to other regions of the plant and warn those cells.
- mobile molecules= hormones—> eg Salicylic acid (SA), Jasmonic acid (JA), Ethylene (ET)
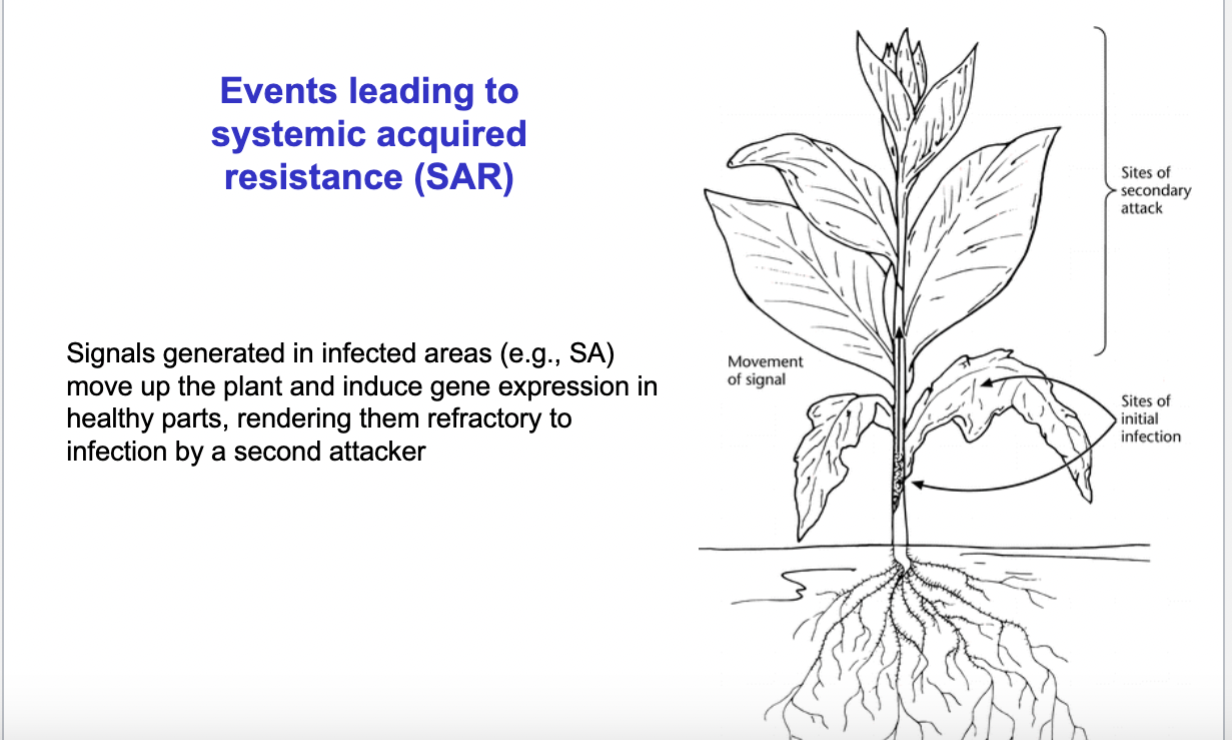
what is SAR?
- After the local HR response: The mobile signal molecules from the infection site — mainly salicylic acid (SA) — travel through the plant to healthy tissues. SA activates a protein called NPR1, which moves into the nucleus and switches on PR (Pathogenesis Related) genes. These PR genes produce defensive proteins like chitinases and glucanases that can directly attack and degrade pathogen cell walls.--> end result is called Systemic Acquired Resistance (SAR).
—> basically: the local response HR lead to signaling (hormones) spreading throughout the entire plant to healthy cells which then are primed with a range of antimicrobial defence mechanisms (SAR)- make them less vulnerable and “prepared” against the pathogen.
give one example of the role of salicylic acid
Salicylic acid (SA) can interact with NPR1- one kind of protein that ultimately switches on the expression of PR genes.
- the mobile warning molecules eg SA spread to non affected cells in the plant and lead to induction of defence responses in those- that can inhibit the pathogen to infect the healthy cells.
—> induces a Cascade of kinases and ultimately lead to transcription of PR genes.
—> some enzymes encoded by the PR genes attack the pathogen cell wall. Results in SAR response
what happens if you create high levels of NPR1
plants become more tolerant to other pathogens- much lower dose of fungicides is needed to be effective (NPR1 is a target for crop engineering — plants with higher NPR1 expression are more disease resistant and also respond better to fungicides, meaning lower doses of chemicals are needed)
expression of NPR1 can be induced by SA
NPR1 is a major component of SAR in plants
can you in short explain how HR salicylic acid NPR1 and SAR work in plant defence against bacteria?
When bacteria are detected, the plant can trigger a Hypersensitive Response (HR) at the infection site — a rapid, localised programmed cell death that essentially walls off and kills the pathogen. This local attack also triggers salicylic acid (SA) accumulation, which binds to the receptor NPR1, causing it to move into the nucleus and activate PR genes that produce antimicrobial proteins. Simultaneously, a mobile signal travels through the phloem to the rest of the plant, priming the same SA–NPR1–PR gene pathway system-wide — a state called Systemic Acquired Resistance (SAR), where the entire plant becomes faster and stronger at defending itself against subsequent infections.
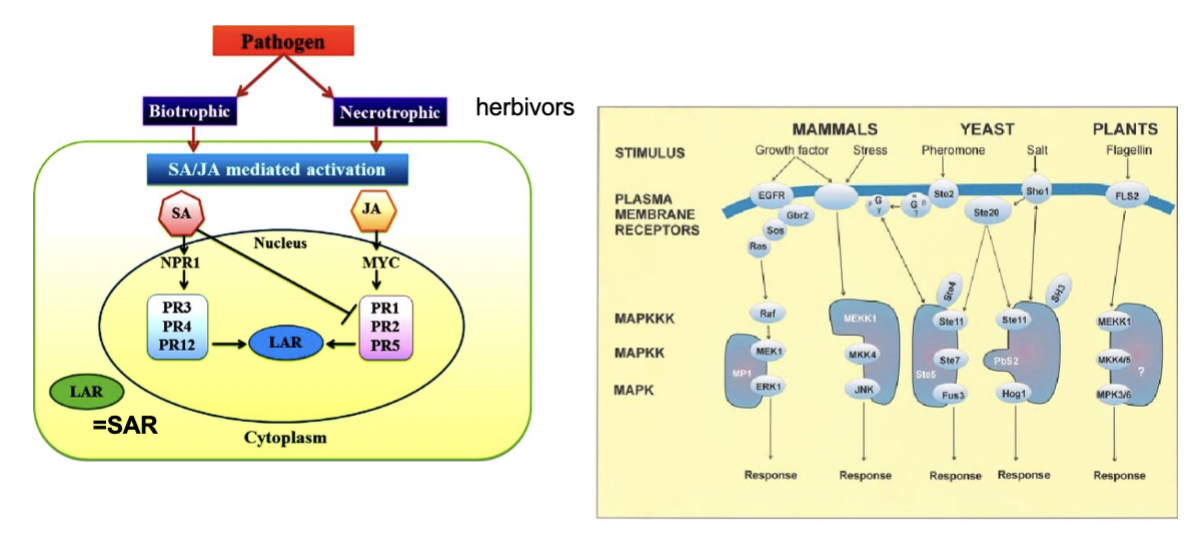
what is MAPK
In plant defence, MAPKs work as a rapid signalling cascade (called the MAPK cascade): an external signal (like detecting bacterial flagellin) activates a chain reaction — MAPKKK → MAPKK → MAPK — which quickly amplifies the signal and passes it on to transcription factors that activate defence genes
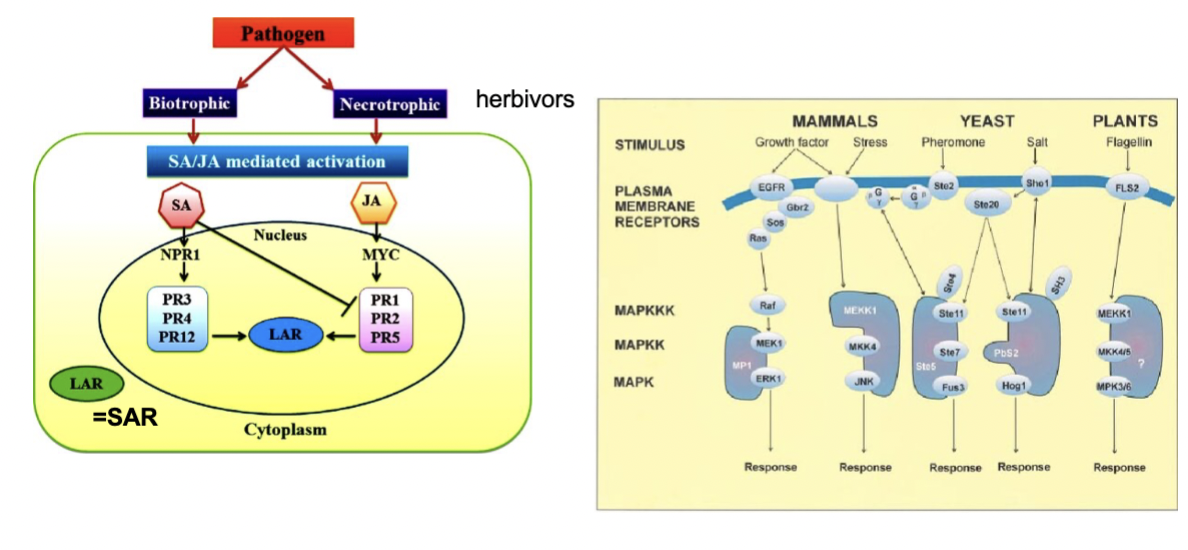
what does biotrophic and necrotrophic mean, explain
- Pathogens are either biotrophic (they keep the plant alive and feed off living tissue) or necrotrophic (they kill the tissue and feed off the dead matter). The plant uses different hormones to fight each: SA is primarily used against biotrophic pathogens — it activates NPR1 which switches on PR1, PR2, PR5 genes, leading to SAR. JA is primarily used against necrotrophic pathogens and herbivores — it activates a transcription factor called MYC, which switches on a different set of PR genes (PR3, PR4, PR12)
could you very briefly explain the molecular events of 1) HR in plants, and 2) SAR in plants
1) HR (Hypersensitive Response) Pathogen effector detected by NLR receptor → MAPK cascade activated → burst of ROS and NO produced → rapid programmed cell death at infection site → pathogen contained.
2) SAR (Systemic Acquired Resistance) HR/local infection → SA accumulates locally → NPR1 activated → PR genes expressed → MeSA and other mobile signals travel via phloem to distal tissues → SA rebuilt from MeSA in healthy cells → NPR1 and PR genes primed system-wide → broad-spectrum resistance established.
what is the aim with engineering pathogen resistance in plants? what questions need to be adressed
Pathogen resistance in plants can be engineered using biotechnologies for eg to achieve plants with increased and durable resistance however, the bioengineering may result in poor quality and lead to less crop yield- the plants will use resources for defence instead of producing harvestable organs- need to accquire a middle ground- what genes should be engineered? From which organisms? But especially how should the genes be expressed in order to limit the costs of resistance to actual infection sites (ie not “wasting” energy on gene expression in non infected tissues) in order to achieve, suitable promoters need to be chosen.
- One clever solution can be to have very specific promotors that only is activated when there is an attack- inhibit the defence mechanisms to be constant and use up a lot of energy.
what is meant by candidate genes? which ones are there?
the genes that are transformed into plants to gain pathogen resistance. “candidate genes” can be manipulated and transformed into the plant, they can be taken from: eg other plants that are relatives to those who will be engineered.
Candidate genes in the transformation: useful genes can come from three sources:
1. From the plant itself — master switch genes that encode for transcription factors, kinases and NPR1 that control the whole defense response, PR genes that directly attack pathogens, or R genes that improve recognition
2. From the pathogen — Avr genes from the pathogen can be inserted into the plant so it constantly "pre-recognizes" the threat and is always primed to respond improving recognition of the pathogen by the plant
3. From other organisms entirely — antifungal peptides, antibodies, or toxic genes that the plant wouldn't normally have at all
what are the four main approaches to engineering resistance:
1. Constitutive expression — the defense genes are always switched on
Local expression — defense genes only activate when a pathogen is detected (using pathogen-inducible promoters)
RNAi — silencing essential pathogen genes so the pathogen can't function
Gene knockouts — knocking out negative regulators of defense so the plant's own response is stronger
—> Each has tradeoffs between effectiveness, specificity and potential fitness costs to the plant.
what does constitutive expression mean, what are the good and bad?
The genes that come from plants are commonly constitutively expressed: always expressed and expressed everywhere: due to very strong promoters:
- Resistance genes: pyramiding (you cane have r-genes for different plants/pathogens to get a durable resistance)
- Durable defence agains different pathogens: disadvantages however: there is a price that genes are expressed constantly- more energy is needed for the defence and less for yield in crops eg- fitness cost
Problems:
•To demonstrate specificity and activity against pathogen
•Risk to activate defence pathway in the absence of pathogen due to strong promoters
name other forms of expression (other than constitutive)
- Some genes are only expressed locally- in the target part (for the pathogen) of the plant: good for energy preserving
- RNAi: interference RNA= silence pathogen essential genes- you can construct it so that you can target all/several pathogens
- Knockouts: known negative regulators of defence- these are knocked out
what are the two constitutive expression ways?
- monogenic resistance: one R gene recognizes one specific Avr protein from one specific pathogen. —> the plant is only protected against that one particular pathogen.
- R genes for several pathogens: broad spectrum of resistance: pyramiding of r-genes. —> Multiple stacked R genes into the same plant give multiple signalling pathways and thus broad-spectrum resistance against a much wider range of pathogens
what are the Two problems with bioengineering disease resistance in plants (CONSTITUTIVE PROMOTERS)
- Each R gene has to be proven to actually work against its specific pathogen — demonstrating that specificity is technically challenging
- Because constitutive expression means the defense genes are always switched on, there is a risk that strong promoters trigger the defense pathway even when no pathogen is present — which wastes the plant's energy and can reduce growth and yield, essentially a fitness cost
explain the strategy to induce resistance through RNAi and chemicals:
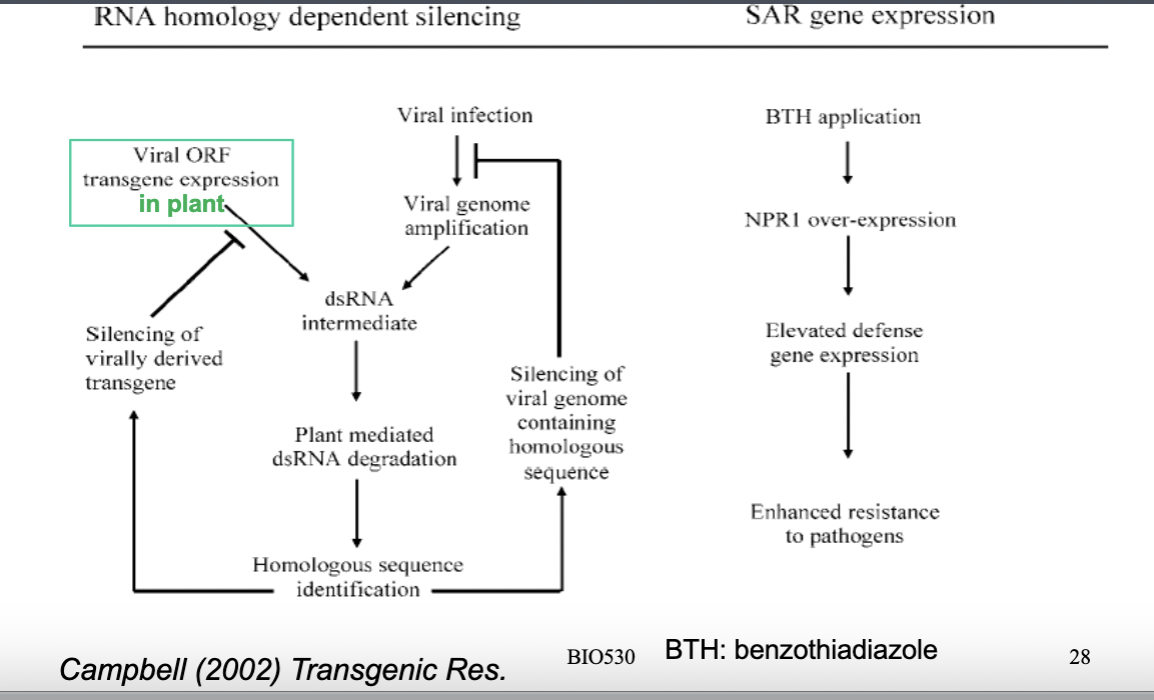
- RNA homology dependent silencing: a viral gene sequence (ORF) is inserted into the plant as a transgene. When a virus then infects the plant, the virus replicates and produces double-stranded RNA (dsRNA) as an intermediate — and crucially, the plant's own transgene also produces RNA that is complementary/homologous to the viral genome. The plant's machinery recognizes this dsRNA as foreign and dangerous, and degrades it, effectively silencing the virus.— the viral genome gets destroyed thanks to the dsRNA recognition which trigger the degradation
- BTH: chemical strategy: BTH is a synthetic chemical that mimics SA — spraying it on crops activates NPR1 and triggers defense gene expression without any genetic modification. BTH spraying on the plant—> stimulate expression of defence genes—> enhanced resistance to pathogens- non GMO strategy to enhance defence against pathogens. However surface level and can hence be rinsed off by rain for eg
what is the goal in plant engineering pathogen resistance
You want defense genes to switch on only at the infection site to avoid the fitness cost of constitutive expression. This requires finding the right pathogen-inducible promoters — either discovered through transcriptomics studies or newly designed synthetically
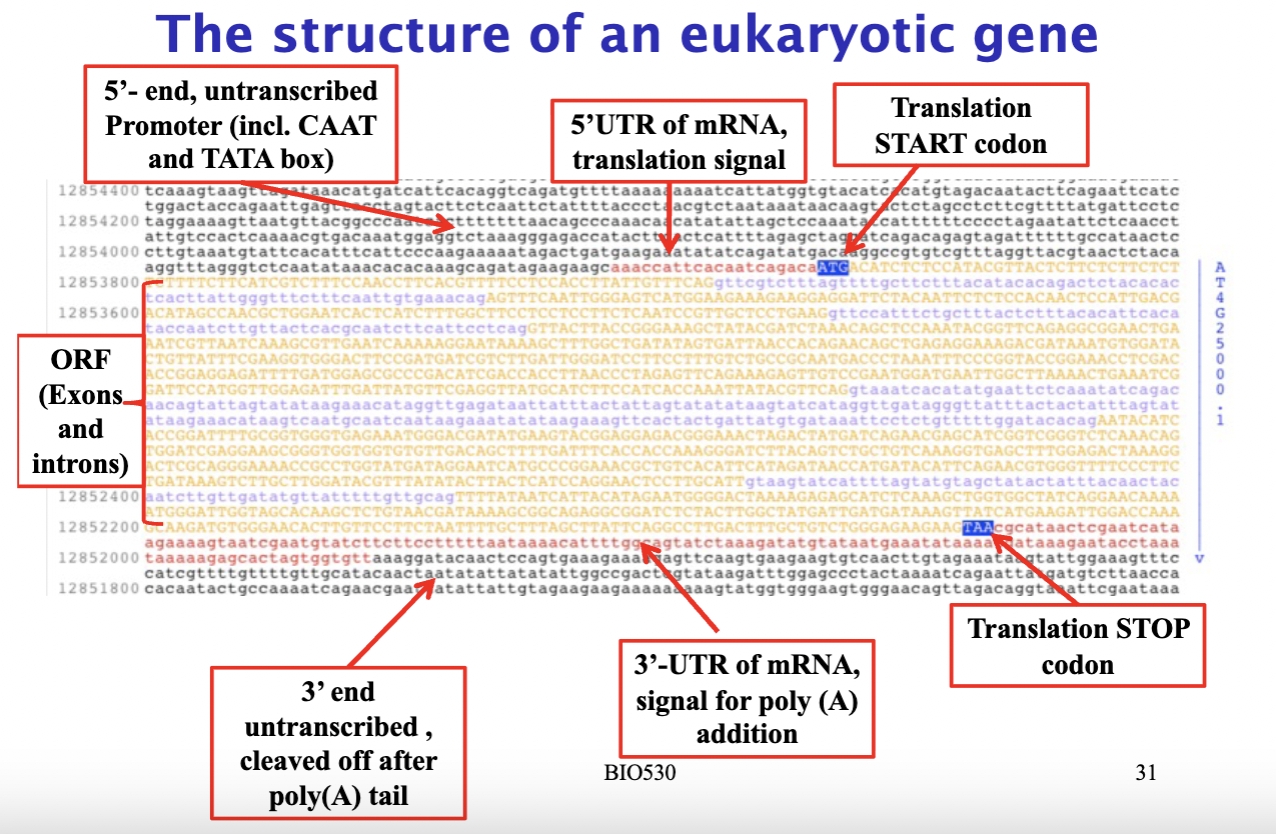
what do all eukaryotic genes have- and hence need to be included in the designed resistance genome
- promoter at the 5' end (containing TATA and CAAT boxes) that controls when and where the gene is expressed- determines when, where and how much the gene is expressed
- ORF (open reading frame): containing the actual genes that encode for proteins- exons and introns,
- UTR regions (flankes the ORF)- regulate translation,
- ending with a stop codon and poly(A) signal.
- when engineering resistance genes into plants, you need all these components — especially the right promoter — to control how and when your gene of interest gets expressed.
explain what promoter cna be used in the transformation
We need suitable promotors to regulate when, where and how much resistance genes should be expressed- a few different kinds:
- Constitutive promoters: express genes at very high level everywhere and anytime. Eg CaMV 35S and ubiquitin promoter
- Tissue specific promoters: could target a gene only in one tissue eg in the epidermis alone
- Inducable promoters: will initiate expression only when a certain molecule will come and interact with it for eg hormones such as SA (triggered by stress and other environmental factors)
- Chemically induced promoters: these will not reach all sites of the plant- can be washed away, eg tetracycline, copper, ethanol
- Pathogen inducible promoters: potentially the most useful= DNA regulatory sequences that activate gene expression specifically in response to pathogen infection, allowing plants to activate defense mechanisms only when needed
- Native promoters: unmodified
- Synthetic promoters: elemts from other sources are gathered- artificially engineered, non-natural DNA sequences designed to control gene expression with high precision, strength, and specificity. They are created by combining cis-regulatory elements (CREs) and core promoters to achieve tailored transgene expression
- Minimal promoters: only contain TATA box and the transcription start site
One strategy to enhance pathogen defence is by using pathogen inducable promoters eg Hsr203J- explain what this is and how it works
- elicitor gene (= pathogen-derived genes that produce molecules (elicitors) capable of triggering immune responses in plants) with a TATA box in front of the gene along with a promoter which become active when the pathogen is recognised- defence is activated
- Avr/elicitor gene is inserted into the plant as a transgene, with a pathogen-inducible promoter (hsr203J in this case) in front. Under normal conditions nothing happens. When a pathogen attacks, the pathogen-induced signal activates the promoter, which switches on the expression of the elicitor transgene, which is then recognized by the plant's own R proteins — triggering a full defense response. —> essentially the plant is engineered to trigger its own HR more reliably and rapidly upon infection
explain
explain how synthetic pathogen inducible promoters work, what are advantages and disadvantages?
Synthetic promotors= transgene with promotors from different pathogens infront- can be switchen on and off. Found to be most promising- more flexible, and strength etc can be altered. Have been tested for different pathogens but also for some fungi and oocyte- if plant is wounded or affected by abiotic stress- resulted in defence—> can get broad resistance even for abiotic stress
- However: need a lot of copies of the element
- Problem: increased “background” expression in unaffected tissues because of the amount of copies needed
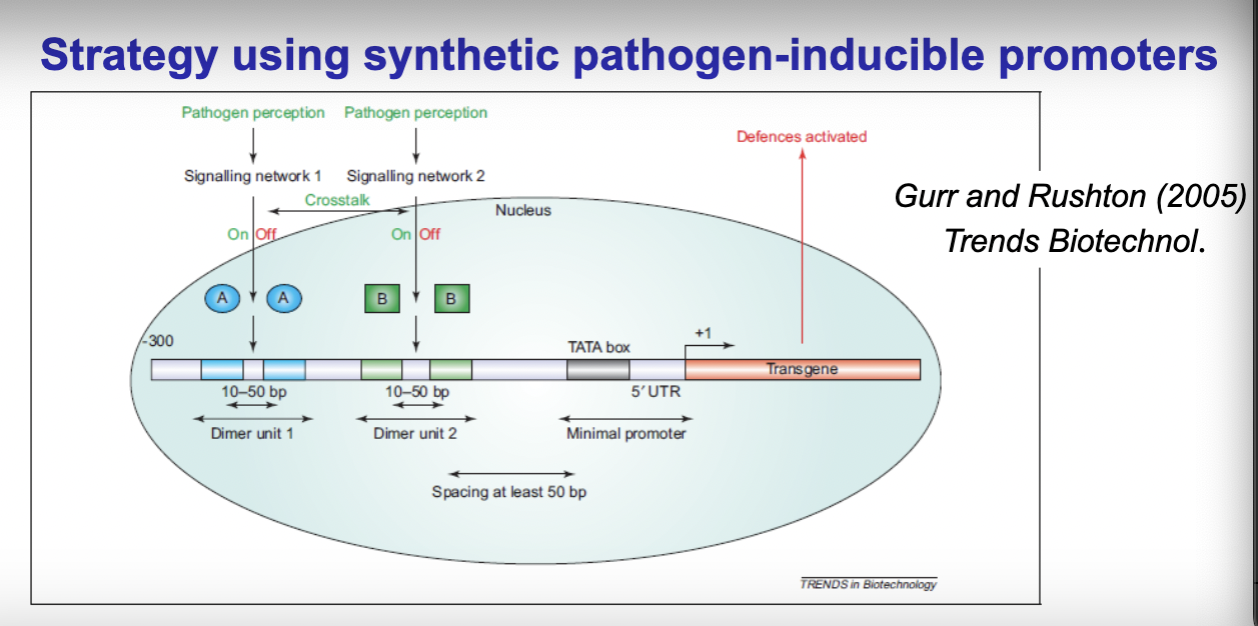
- Synthetic pathogen-inducible promoters: Rather than using a single natural promoter, synthetic promoters are built from multiple small regulatory elements (cis-acting elements/ "dimer units") combined in front of a minimal TATA box promoter. —> "lego blocks" — you can mix and match different elements to customize the promoter's behaviour. Having multiple different dimer units in the promoter means multiple signalling pathways (eg stacked pathogen inducible) can switch the promoter on, making it more broadly responsive. This gives more flexibility than natural promoters in terms of both strength and what triggers them.
- Trade off: Strength vs specificity
what are some strategies to incurr resistance to potato blight?
- Overexpressed (many copies) r-genes from a relative potato species was taken (is resistant to the pathogen) and transformed—> lead to resistance in the leaves but not in the tubers.- non sufficient
- Constitutive expression of kinase that activates NADPH oxidase and results in oxidative burst and HR-like cell death—> increased resistance to potato blight was required- however caused an increased vulnerability to other pathogens
- Added R- genes from relative: transformed potatos where planted in regular conditions—> still affected however not affected to a greater extent- partly succesful- Potatoes that carry the specific R genes show few small disease spots, but performed far better than normal potatoes. – “stacking” several resistance genes would likely give even longer-lasting protection.
explain the potential of GMO
GMO: potential to increase and stabilize yields in a world with a changing climate and a growing human population. —> potential benefits: reduced use of pesticides, reduced concentrations of, e.g., mycotoxins in the end products.
what are the most common GMO, what genes do they contain- explain
The most common GM-crops tpday are Bt and Ht. They both contain heterologous transgenes, meaning the genes come from completely unrelated organisms
Homologous = gene from a related organism, something the plant's gene pool could have had naturally
Heterologous = gene from a completely unrelated organism, something entirely foreign to the plant
what are virus resistant GM crops
h plants are purposely inoculated with a mild strain of a virus in order to establish protection against later infection by a more severe strain (similar principle as that of vaccination).
what are risks witg GM crops
spread of traits to wild relatives, spread of antibiotic resistance to other microorganisms, spread to organic crops, reduction in genetic diversity (only GM crops)