Bioinorganic Chemistry
1/87
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
88 Terms
Which elements are most important for life?
H, C, O, N, S, Fe, Ni
What is the key factor in availability for life?
Assuming life began in the oceans, the solubility of the elements (and therefore their availability) is the key factor in availability for life.
How do biological molecules interact with metal ions?
They coordinate with a lone pair (Lewis base), forming a dative bond to the metal ion (the Lewis acid).
What is a hard/soft HSAB?
A hard HSAB (Hard Soft Lewis Acid or Base) has a high charge but small size.
A soft HSAB will have a lower charge and a larger size.
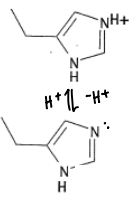
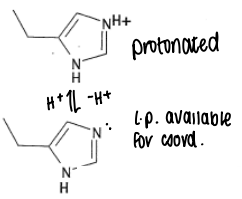
What are the properties of Histidine?
The pKa of protonated His is 6.5, therefore it will become deprotonated within the body. Deprotonated His has a lone pair available for coordination, and so has a pKa of 14.
Histidine is a borderline HSAB, so will coordinate to a range of metals.
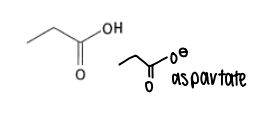
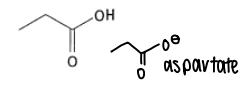
What are the properties of aspartic acid?
It has a pKa of 4.5 and can be deprotonated into aspartate in the body.
It is a hard HSAB.
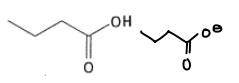
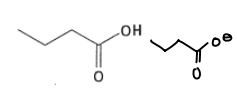
What are the properties of glutamic acid?
It has a pKa of 4.5 and can be deprotonated in the body.
It is a hard HSAB.
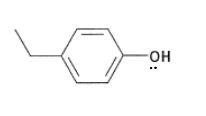
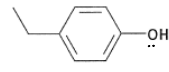
What are the properties of tyrosine?
It has a pKa of 10, therefore it is not deprotonate in the body.
It is a hard HSAB.
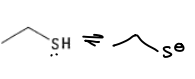
What are the properties of cysteine?
It has a pKa of 8.5, so is sometimes deprotonated.
It is a soft HSAB.
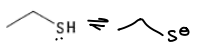
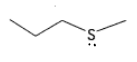
What are the properties of methionine?
It has no H+, therefore it has no pKa.
It is a soft HSAB.
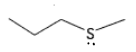
Why is it important that amino acids have a variety of coordinating properties?
To control the properties of the metal, e.g. REDOX potential.
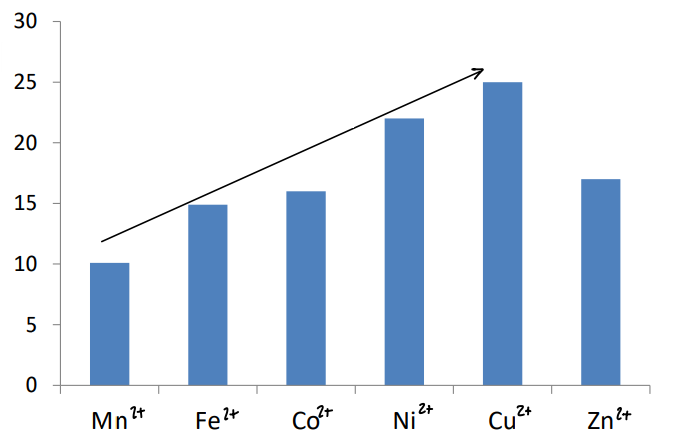
What is the Irving Williams series?
It shows for any fixed ligand, going left to right on the periodic table in the 1st row transition metals increases stability.
y axis is a logarithmic scale.
Zn2+ is an outlier, therefore biology needs to find a way to form stable complexes with this ion.
How are stable complexes formed using Zn2+?
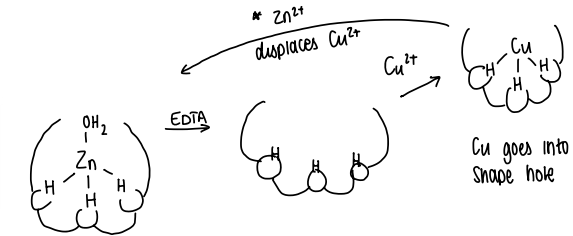
Via preorganisation:
The relative positions of the His are locked in place by the surrounding protein structures.
EDTA can be added to remove the Zn2+ metal, however the protein shape remains.
Cu2+ can be used to replace the Zn
Adding Zn2+ can then displace the Cu2+, forming the original structure.
This counters the Irving-Williams series.
What are tetra-aza macrocycles?
Can give the metal ion other properties.
They are found in many metalloproteins.
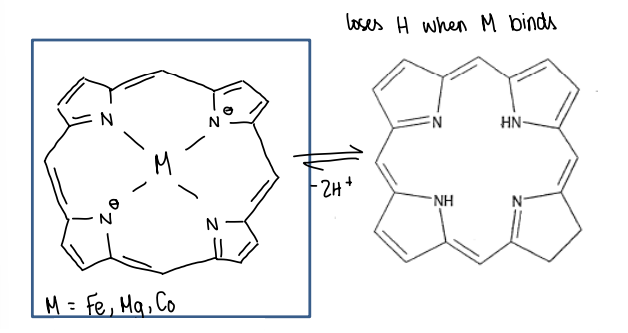
What is haem?
A tetra-aza macrocycle which has an Fe as the metal.
The Fe can often exist in many O.S. (+2, +3, +4), this is a critical function of haem.
Why is it hard to study metalloproteins?
We are studying a highly complex system, therefore when studying by IR for example, the spectrum becomes too complicated to get any information on the metal ion.
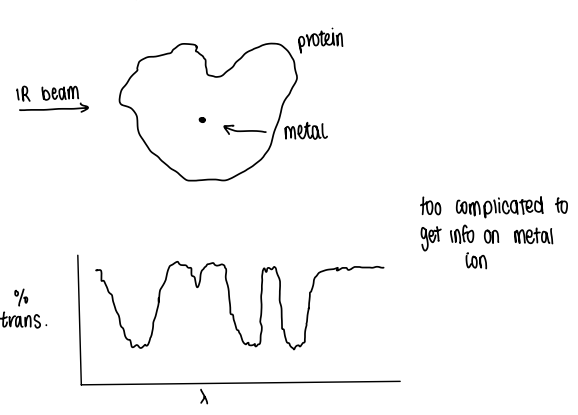
What kinds of methods are needed to study metalloproteins?
Need a method which simplifies complex info.
Need a method which ignores everything except the metal.
What is single crystal X-ray diffraction?
A powerful technique which gives a full 3D map of electron density of a molecule (protein).
However, this info is often at low resolution and crystals of the protein have to be grown, which is difficult.
How do you use XRD results?
It produces a cage contour map, this shows the electron density.
You then fit the expected protein structure to the electron density.
A higher resolution map is easier to interpret, whereas if it is a lower resolution XRD it can be much harder to interpret.
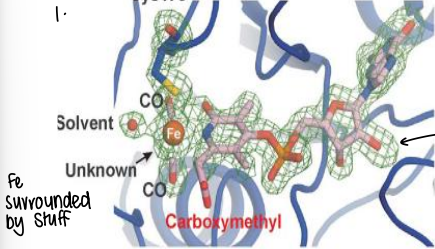
What is Extended X-ray Absorption Fine Structure (EXAFS)?
An X-ray removes a 1s electro from the metal. This tells us about electron density in the immediate vicinity of the absorbing atom.
How does EXAFS work?
X-rays are absorbed by the sample.
The absorption causes the 1s electron ionisation.
An absorption v Energy graph is plotted, giving the EXAFS fine structure (if the edge is more to the right, it has a higher O.S.).
After the ionisation, the lost electron causes a 3D ripple effect to surrounding atoms, which back scatter the ripple.
The ripples constructively and destructively interfere at the central metal.
The amount of interference depends on the type of backscatterer (more electrons means more backscatter) and distance from the metal.
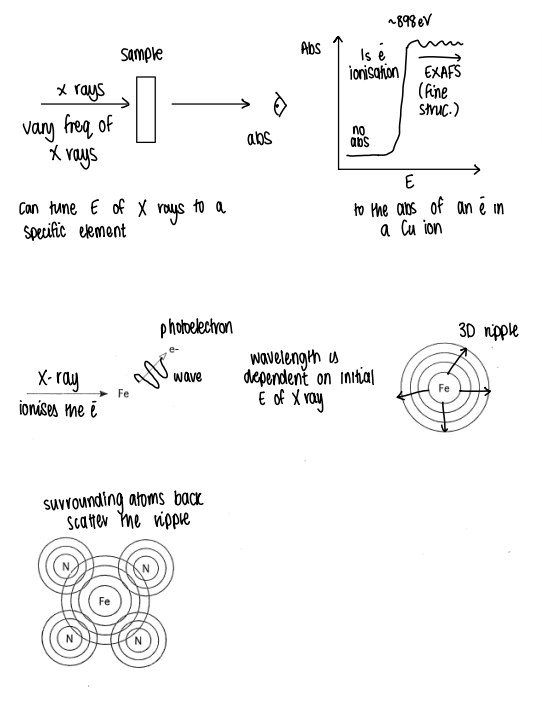
What kind of graphs does EXAFS give?
Produces graphs which tell us about electron density around the metal as a function of distance.
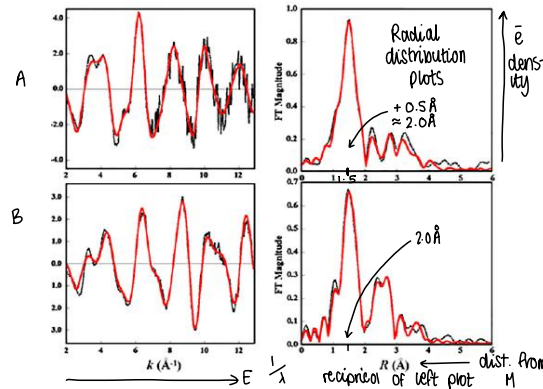
What are the typical metal atom distances for common elements?
M-O, M-N: ~2.O A
M-S: ~2.3 A
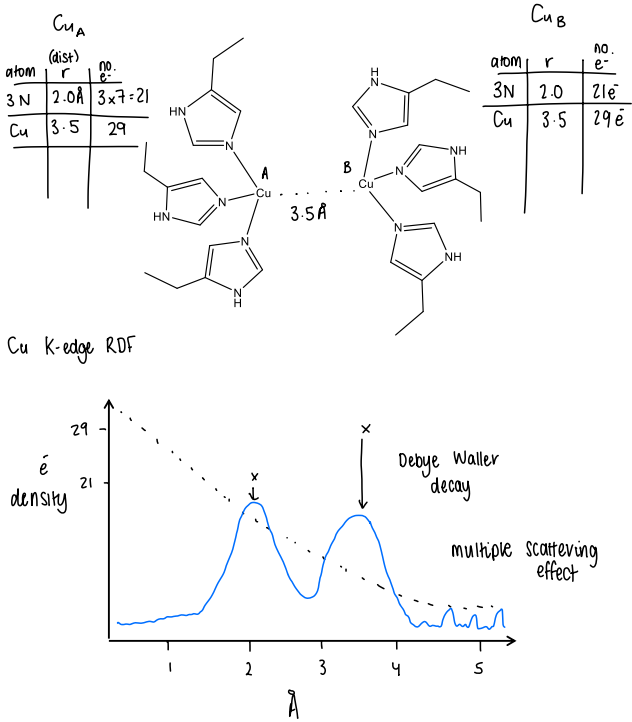
How would you predict the Cu-EXAFS spectrum of a molecule?
First identify the metal atom and the number of surrounding atoms.
Then find the distance between the metal atom and the neighboring atom.
Then find the distance between the metal and its surrounding atoms.
Calculate the number of electrons for both the metal and its surrounding atoms.
Plot a graph of electron density v distance and plot the previously determined values.
Account for the Debye Waller decay, causing the peaks to shift downwards.
The multiple scattering effect also causes some small peaks further away from the metal.
What are phase shift considerations?
Due to phase corrections, observed distances in EXAFS are slightly shorter than actual bond lengths
What is the multiple scattering effect?
Scattering from more than one neighboring atom before returning to the absorber atom.
Contributions from multiple scattering effects give small peaks and shows possible long-range order in the active site.
What increases the multiple scattering effect?
There will be increased multiple-scattering effects due to higher polarisability, potentially altering spectral fine structure, at longer distances in the radial distribution plot.
What affects the Debye-Waller effect?
There will be a higher Debye-Waller factor due to the fact that Zn–S bonds being more flexible than Zn–N bonds, leading to increased thermal disorder and peak broadening
Why can we not use isotropic NMR?
In solution state, everything will tumble and so averages out.
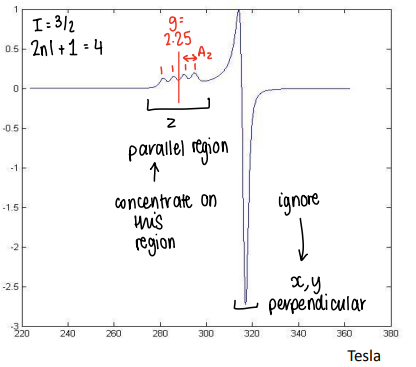
How does anisoptropic EPR work?
The EPR detects any unpaired electrons.
The g-value is split into x, y and z components.
The splitting (A) is also split into x, y and z.
Can ignore the perpendicular regions (the x and y regions), and focus only on the parallel region (z region).
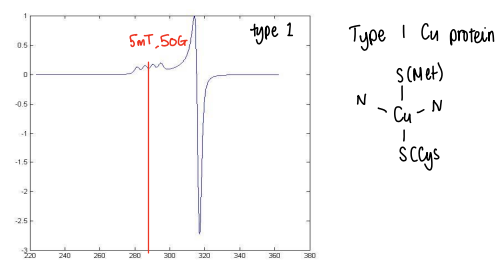
What are type 1 Cu proteins and what does their EPR look like?
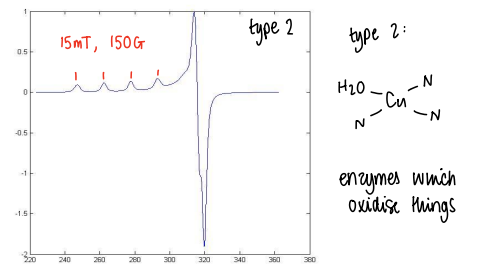
What are type 2 Cu proteins and what does their EPR look like?
These proteins are enzymes which oxidise things.
What are type 3 Cu proteins and what does their EPR look like?
These Cu proteins have active sites like deoxyhaemocyanin.
These proteins have no EPR spectrum as, after oxidation, Cu(2+) ions are formed with opposite spin states.
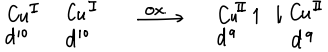
How do you differentiate a type 3 Cu protein?
Do an EPR; an EPR is expected due to u.p.e, however a type 3 Cu protein will be EPR silent.
What is Mossbauer spectroscopy and how is it carried out?
A nuclear spectroscopy technique which measures energy transitions within the nucleus of a selected metal.
Gamma rays are used to cause element specific transitions:
A fixed wavelength gamma ray is shone at the sample.
The sample is moved back and forth.
The absorption is measured as a function of wavelength.
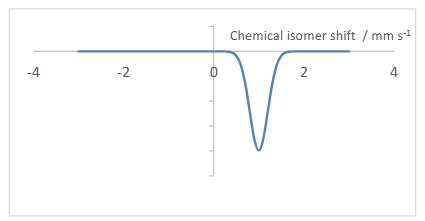
What does Mossbauer spectroscopy tell you?
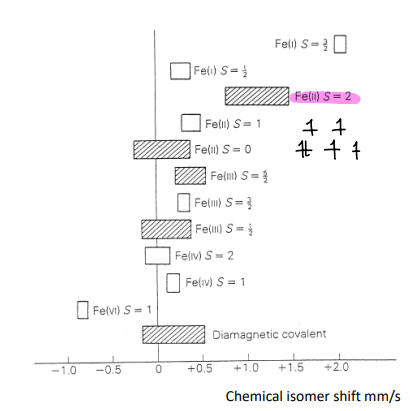
The chemical isomer shift is dependent on O.S. and spin state of Fe.
The chemical isomer shift can therefore be read off from known values, telling you about the Fe nucleus.
Fe(II) with S=2 has a shift of around +1 mm/s.
What can a protein sequence tell you?
Can tell you about the type of metalloprotein, and the potential metal that can be associated with that protein.
What is a consensus sequence?
An identifiable sequence of amino acids that coordinate to a metal.
If a particular amino acid is conserved between many consensus sequences, this means it is an important amino acid for the specific function.

Why is oxygen needed in biology?
Oxygen has a high oxidising power. It will react with low energy electrons and protons to generate water in a highly exothermic reaction.
Oxygen allows life to live at high energy as oxygen is able to oxidise low voltage electrons.
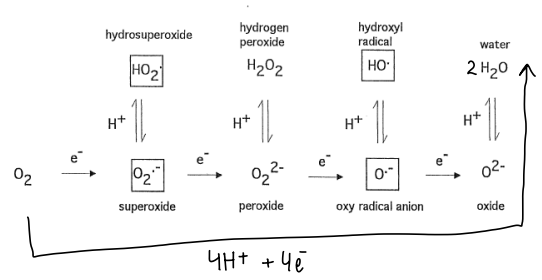
What is the problem with oxygen as an oxidiser?
All of the oxygen intermediates in the chain are highly toxic, therefore we need a means of reducing oxygen with 4 electrons instantaneously to form water.
What is the structure of a mitochondria?
It has a folded membrane with high surface area.
Embedded in its membrane are complexes, such as cytochrome c oxidase.
What is cytochrome c oxidase?
It is an Fe containing complex which allows the conversion of oxygen into water instantaneously.
Fe is coordinatively saturated with approximate octahedral geometry in a porphyrin ring with axial ligands (His and Met). It can adopt two different oxidation states.
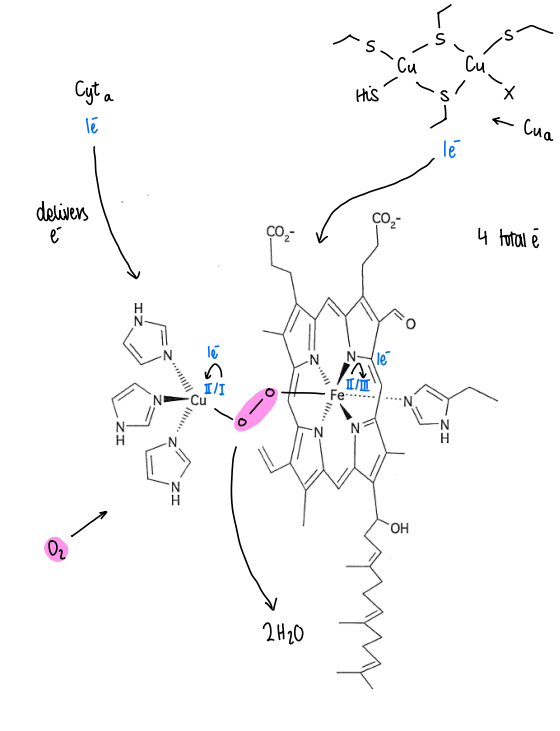
How does cytochrome c oxidase store electrons?
Cyta delivers an electron to the copper centre, converting it from Cu(I) to Cu(II).
Cua protein delivers an electron, converting the iron centre from Fe(II) to Fe(III).
Oxygen binds to both the Cu and Fe, storing 2 electrons.
This means 4 electrons are stored inside CCO, which can be rapidly released for the reduction of oxygen into water.
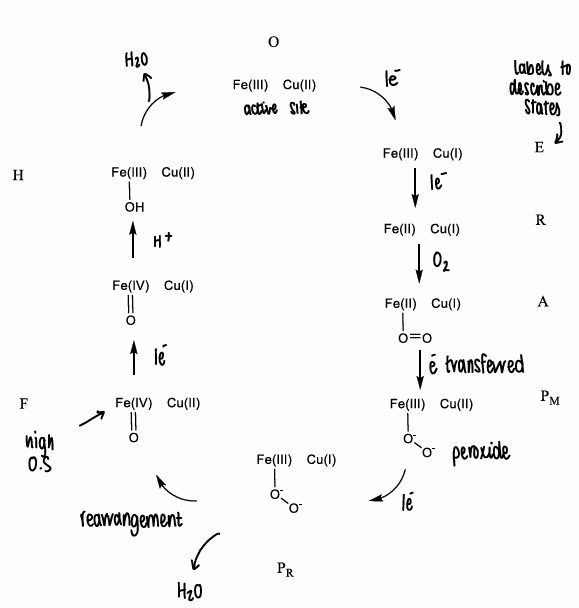
What is the mechanism for CCO?
The CCO metal ions are able to react directly with oxygen, which exists in a triplet spin state (has 2 u.p.e) and is paramagnetic.
The oxygen react quickly with paramagnetic metal ions, forming water.
What is a problem with the use of CCO?
Sometimes CCO can go wrong, especially under times of oxidative stress. This produces, O2-, O22- and -OH instead.
Copper-zinc superoxide dismutase is required to remove these toxic compounds.
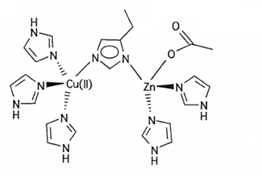
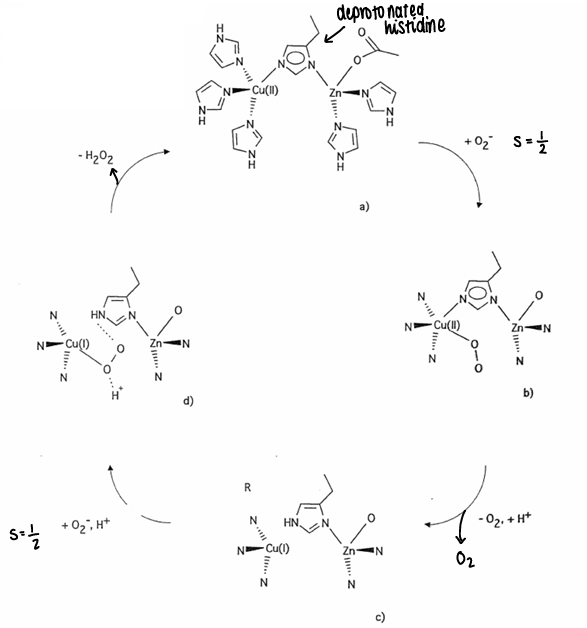
What is the mechanism for copper-zinc superoxide dismutase?
It is a metalloprotein used to remove O2-.
2O2- + 2H+ → O2 + H2O2
What is photosynthesis?
The process of turning water into oxygen.
2 H2O → O2 + 2H+ + 2e-
Plants get rid of oxygen as it is toxic to them.
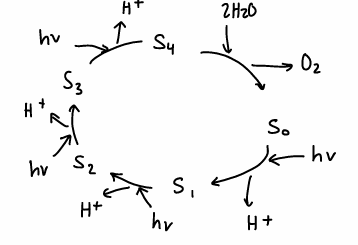
What is the Kok cycle?
A photosynthesis cycle containing 5 states which occurs in plants at a metal containing active site. Converts water into H2O.
The active site contains 4 Mn atoms forming the oxygen evolving complex (OEC).
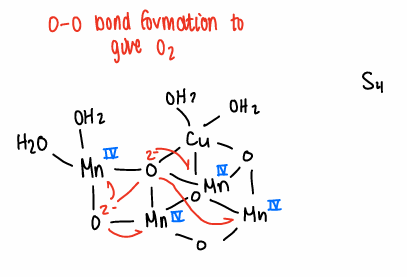
What happens at the active site of the OEC?
On addition of hv, there is a sequential increase in O.S. of the Mn ions.
O-O bond formation occurs to give O2.
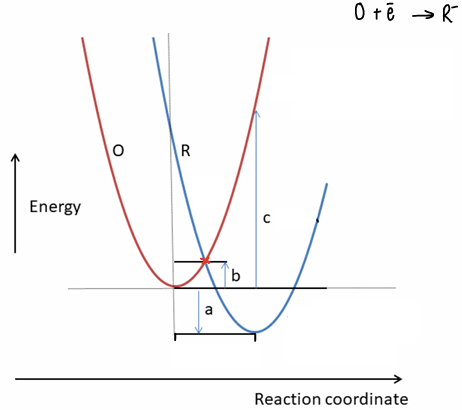
What is Marcus theory?
A basic theory to calculate rate of electron transfer.
The parabola represents the vibration of the molecule as an electron passes from a donor (O) to an acceptor (R).
The intersect represents the point at which the electron can transfer.
a = ΔG
b = activation energy
c = reorganisation energy (λ)
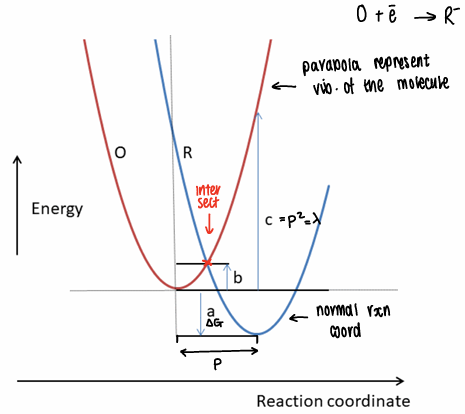
What is the Marcus equation?
p2 = λ
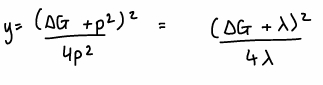
How is the Marcus equation inserted into the Arrhenius equation?
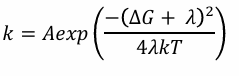
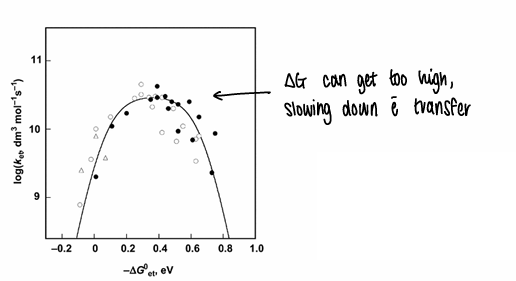
What is the Marcus inverted region?
The rate of electron transfer depends on ΔG and λ.
Fast electron transfer occurs at low values of λ.
Rapid biological electron transfer depends on the reactive site barely changing structure upon electron transfer. This means if ΔG gets too high, distortion occurs in the molecule and electron transfer slows.
Molecules need to be rigid.
What are the three main classes of biological electron transfer sites?
Cytochromes
Fe-S proteins
Type 1 Cu proteins
These all take advantage of a lack of reorganisation energy.
What are cytochromes?
Fe containing porphyrins (contain flat haem groups).
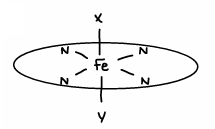
What are the properties of cytochromes?
They are 1e- transfer sites, Fe(II) ←> Fe(III).
The Fe is coordinatively saturated (has 6 ligands), meaning the Fe in its 2+ state doesnt react with O2.
The haem group is rigid and doesnt move structure as the e- goes through (e- transfer is rapid, low λ).
What is rubredoxin?
An Fe-S electron carrier protein in bacteria which has a single Fe at its active site.
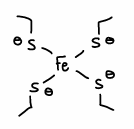
What are the properties of rubredoxin?
The cysteinates are weak ligands and so don’t split the energy of the d-orbitals on Fe by much.
Fe is in the 2+ O.S., therefore it is high spin and so it is easier to remove an electron upon electron transfer as they occupy the higher energy orbital.
The Fe-S bond is also quite covalent, therefore the electron is spread out along the bond and not a point charge (so lower reorganisation energy), so is less attracted towards charges and so more rigid. Delocalisation is also needed for rapid electron transfer.
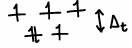
What are ferredoxins?
Photosynthesis proteins containing iron sulfur sites, facilitating electron transfer.
The electrons are fully delocalised between Fe atoms, forming Fe with a 2.5+ charge.
High delocalisation means faster electron transfer.
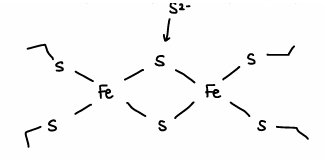
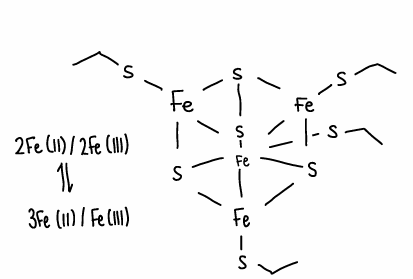
What is the structure and properties of a 4Fe-4S ferredoxin complex?
The electrons ae delocalised in a cube.
What are cupredoxin copper proteins?
A blue type 1 Cu containing protein used for electron transfer within plants.
The Cu can exist as 1+ or 2+.
Cu (I) exists as d10 in a tetrahedral geometry.
Cu (II) exists as d9 in square planar geometry.
These O.S. states are bad for electron transfer due to the large reorganisation energy between states (changes geometry).
What is the entatic state?
A coordination geometry for cupredoxin held rigid by a protein that is halfway between square planar and tetrahedral.
How do plants make electron transfer in cupredoxin faster?
They use the entaic state, and the copper atom is coordinated typically by two Histidine residues (a borderline HSAB), one methionine and one cystenate (soft).
This delocalises the electrons.
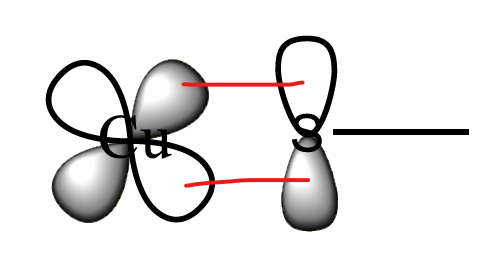
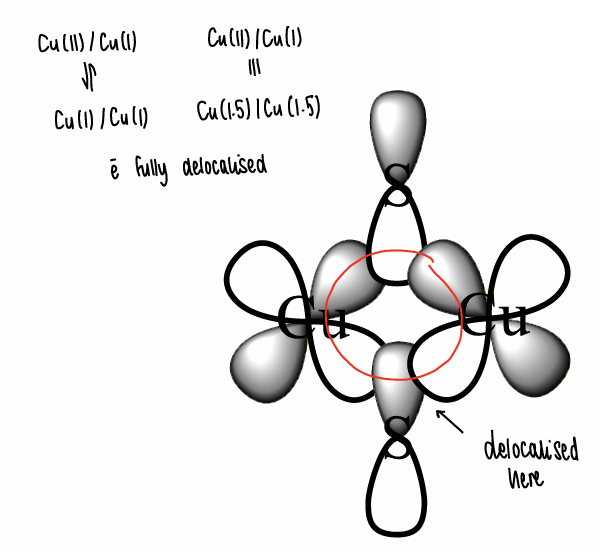
What is the CuA site?
A Cu containing electron transfer site in humans (CuA is found in CCO).
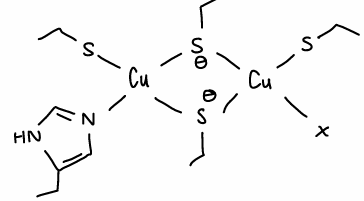
What are the properties of the CuA site?
The electrons are fully delocalised around the Cu2S2 core part of the active site, therefore electron transfer is rapid.
What are the properties of zinc in biology?
It has a single O.S., Zn(II), d10, therefore it is redox inactive.
It is a LA, therefore is a way to make hydroxides at a normal pH.
What are the differences between Zn and Co chemistry and how is this used?
Zinc is spectroscopically silent, it has no paramagnetism (no EPR) and no d-d transitions (no UV-Vis).
Co(II) can often substitute for Zn (II), as it is d7 it can be seen in EPR and UV-Vis.
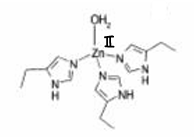
What is human carbonic anhydrase II?
The most important zinc protein and one of the most efficient enzymes. Zinc has an approximate tetrahedral coordination geometry and a fixed oxidation state.
HCA is able to take a reaction with a rate constant of 0.1 M-1s-1 and increase it by 109.
HCA catalyses the conversion of CO2 into HCO3-.
It is diffusion limited.
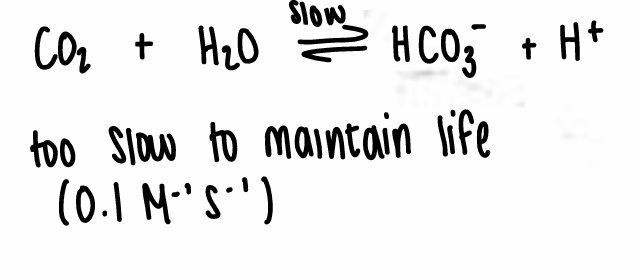
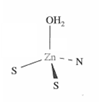
What is the mechanism for HCA?
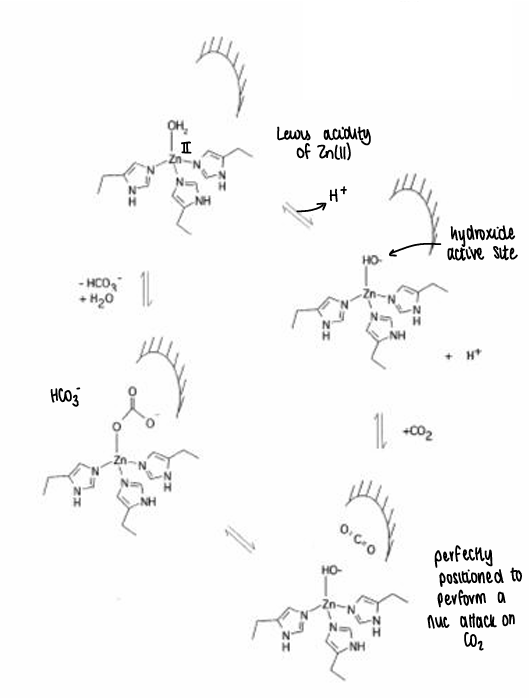
What is liver alcohol dehydrogenase?
A Zn(II) dependent enzyme that oxidises ethanol into methanal.
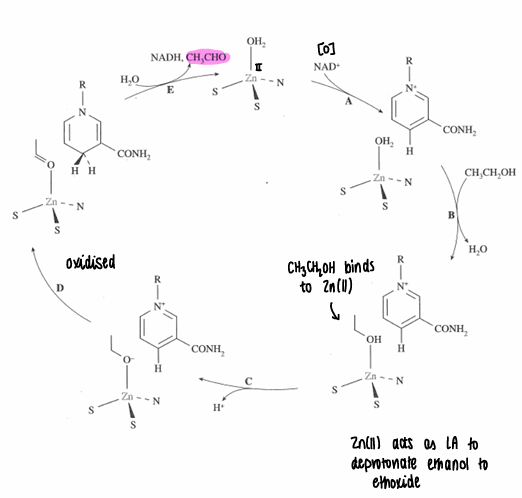
What is the mechanism for liver alcohol dehydrogenase?
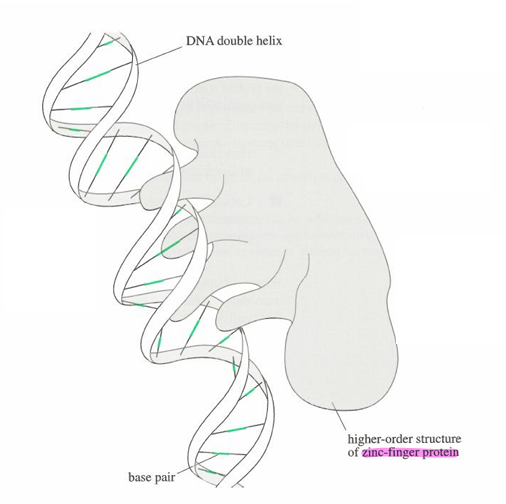
What are zinc finger proteins?
The protein grabs DNA through the grooves and is able to manipulate DNA.
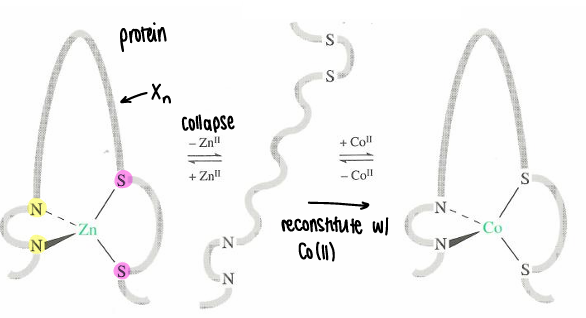
How are zinc finger proteins studied?
Upon removal of zinc (which is coordinated by 2 hsitidines and 2 cysteinates), the structure collapses. This can be reconstituted with Co(II), reforming the structure.
Co(II) is spectroscopically active, therefore UV-Vis can be used. The UV-Vis shows the Co must be in tetrahedral geometry due to the magnitude of the extinction coefficient of the d-d transition (octahedral would have a lower value).
What is the problem with using Fe?
At the great oxygenation event, O2 entered the atmosphere and drive the reaction.
Fe(III) is a LA, so the complex deprotonates repeatedly, forming Fe2O3 (rust) and Fe(OH).
Fe(OH)3 is very insoluble.
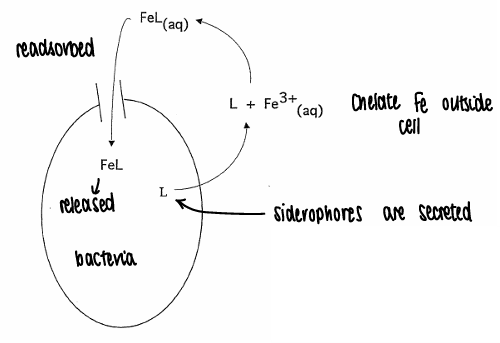
How do bacteria acquire Fe?
Using siderophores.
The Fe chelates to siderophores outside of the cell (must be good at chelating, thermodynamics determines the most stable Fe complex).
The Fe complex is readsorbed, the Fe is then released and siderophores are secreted back out of the cell.
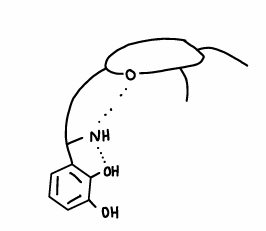
What are the properties of enterobactin (a siderophore)?
H-bonding from the ring forces catechol groups to face inwards, therefore preorganising the molecule for binding of Fe(III).
It is one of the most stable Fe(III) complexes known.
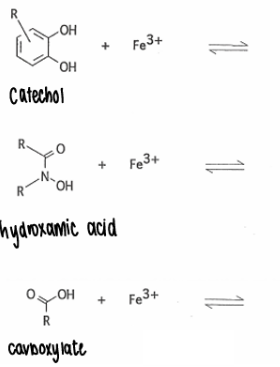
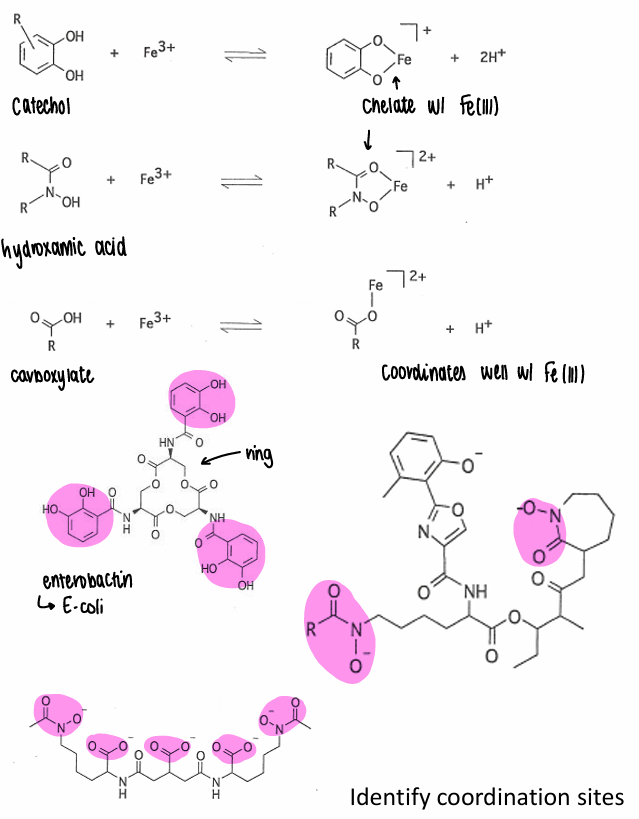
How do siderophores coordinate with Fe?
They all contain hard ligands which act as the coordination site.
How do mammals transport and store Fe?
There are dedicated proteins to move Fe from cell to cell (Transferrin).
Transferrin has a clover leaf shape protein structure, however when Fe(III) binds, the Fe becomes hidden in the middle of the protein.
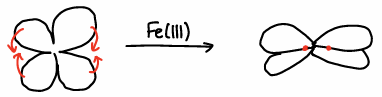
How does transferrin prevent acquisition of Fe(III) by bacteria?
The stability constant of Fe in Tf is ~1029, whereas in bacteria enterobactin has a stability of ~1049.
Transferrin kinetically stabilises the Fe, it is closed off to bacteria as it takes too long to get into the protein.
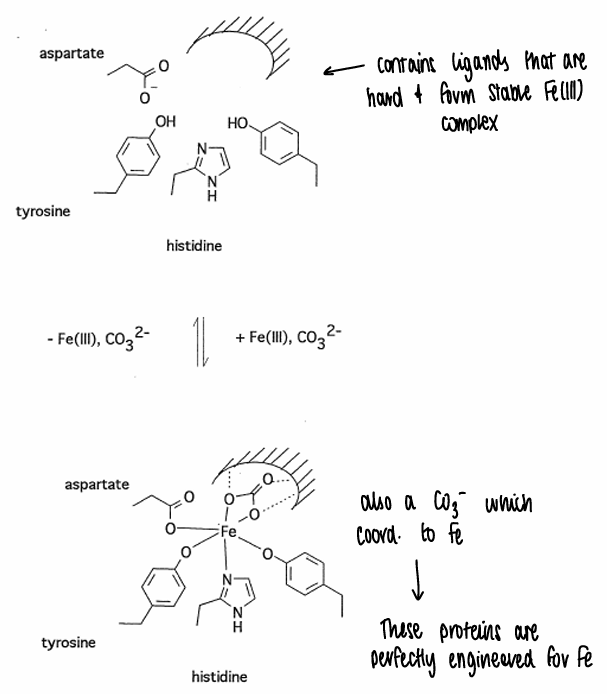
What are the active site characteristics of transferrin?
Transferrin contains ligands that are hard and form stable Fe(III) complexes.
A CO3- also coordinates to the Fe.
Fe(III) forms the most stable complex with transferrin, with other metals having either the wrong charge or wrong size.
What is ferritin?
An iron storage protein.
What is the structure of ferritin?
The ferritin strands self assemble into hollow spheres, forming a pore in the middle.
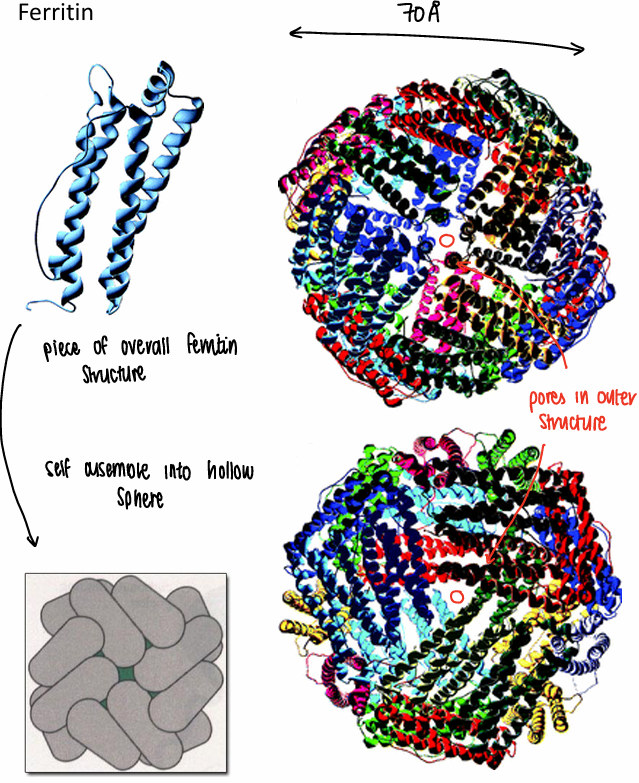
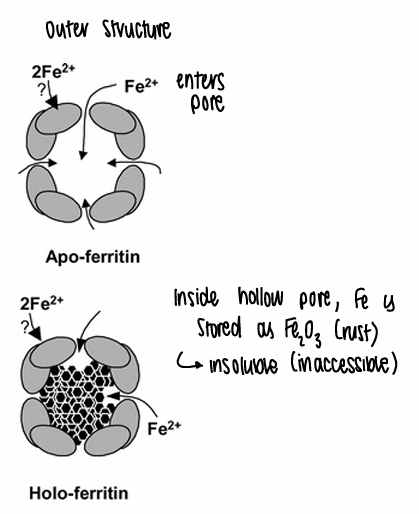
How does ferritin bind iron?
The iron enters the pore, and inside the pore the Fe is stored as Fe2O3 (rust), which is insoluble and therefore inaccessible.
What is XANES spectroscopy?
XANES is a spectroscopic method that analyzes X-ray absorption near an element’s edge to determine its oxidation state and local chemical environment.
Plots absorption v energy around absorption edge.
Higher O.S = higher energy edge
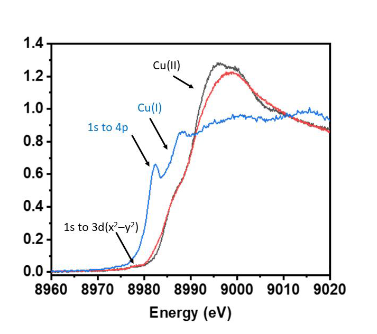
What is the advantage of using EXAFS instead of XRD?
Sample can be in any phase (liquid, solution, solid).
Sample does not need to be crystalline.
Highly specific for active site.
What does a Mossbauer spectrum look like?
Plots gamma ray transmission against chemical isomer shift.
How do the chemical properties of Fe ions affect its bioavailability?
Fe³⁺ exchange is slow, making kinetics of uptake slow.
Fe has two accessible oxidation states (Fe²⁺/Fe³⁺) the former of which has higher ligand exchange rates leading to higher uptake rates.
Fe³⁺ is a strong Lewis acid and forms insoluble hydroxides at physiological pH, necessitating transport via proteins like transferrin.