Week 6 - RNA Polymerase II and GTF, Mechanism of transcription
1/89
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
90 Terms
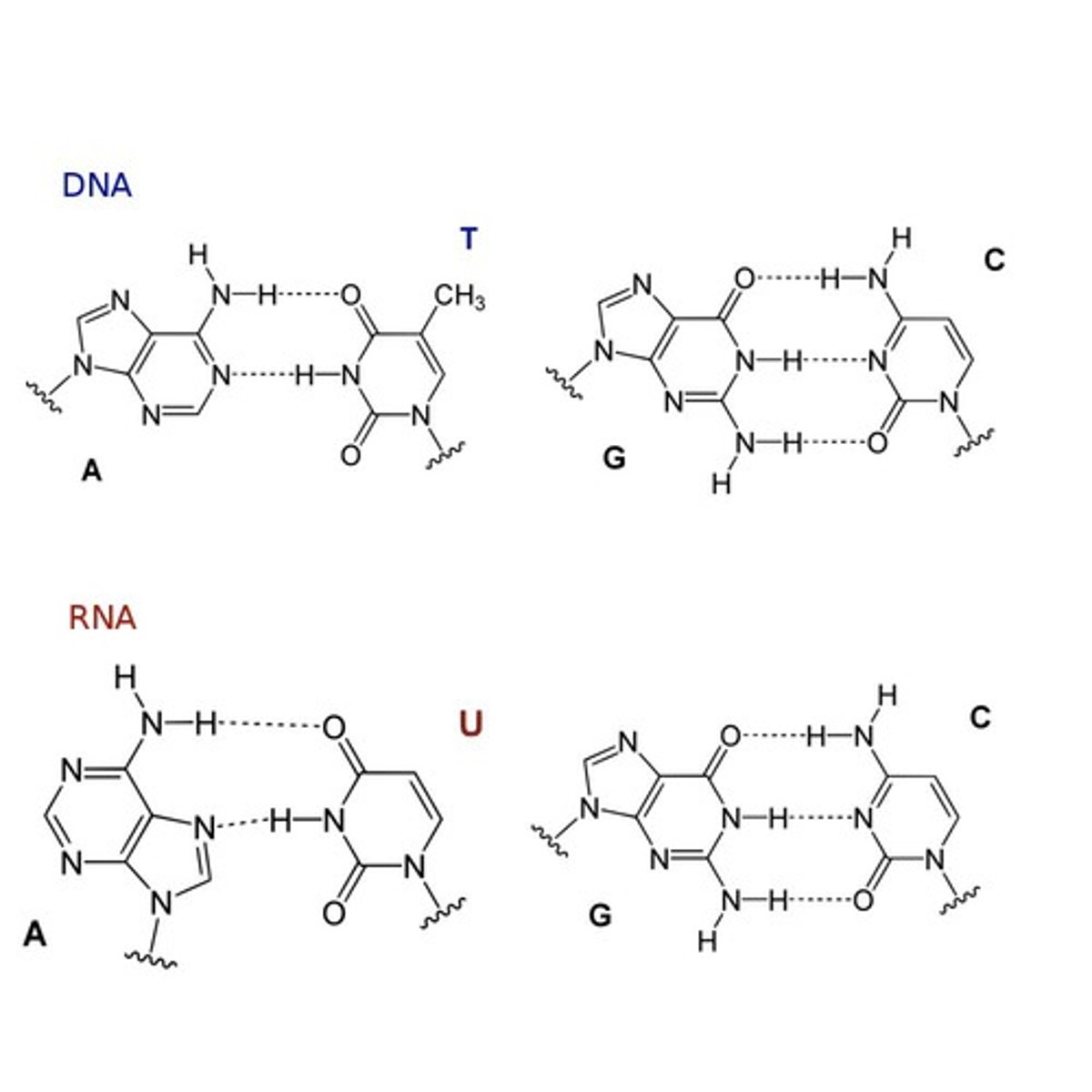
RNA base pairing
Adenine - Uracil
Guanine --- Cytosine
ribonucleotides
raw material - nucleoside triphosphate; linked by phosphodiester bonds.
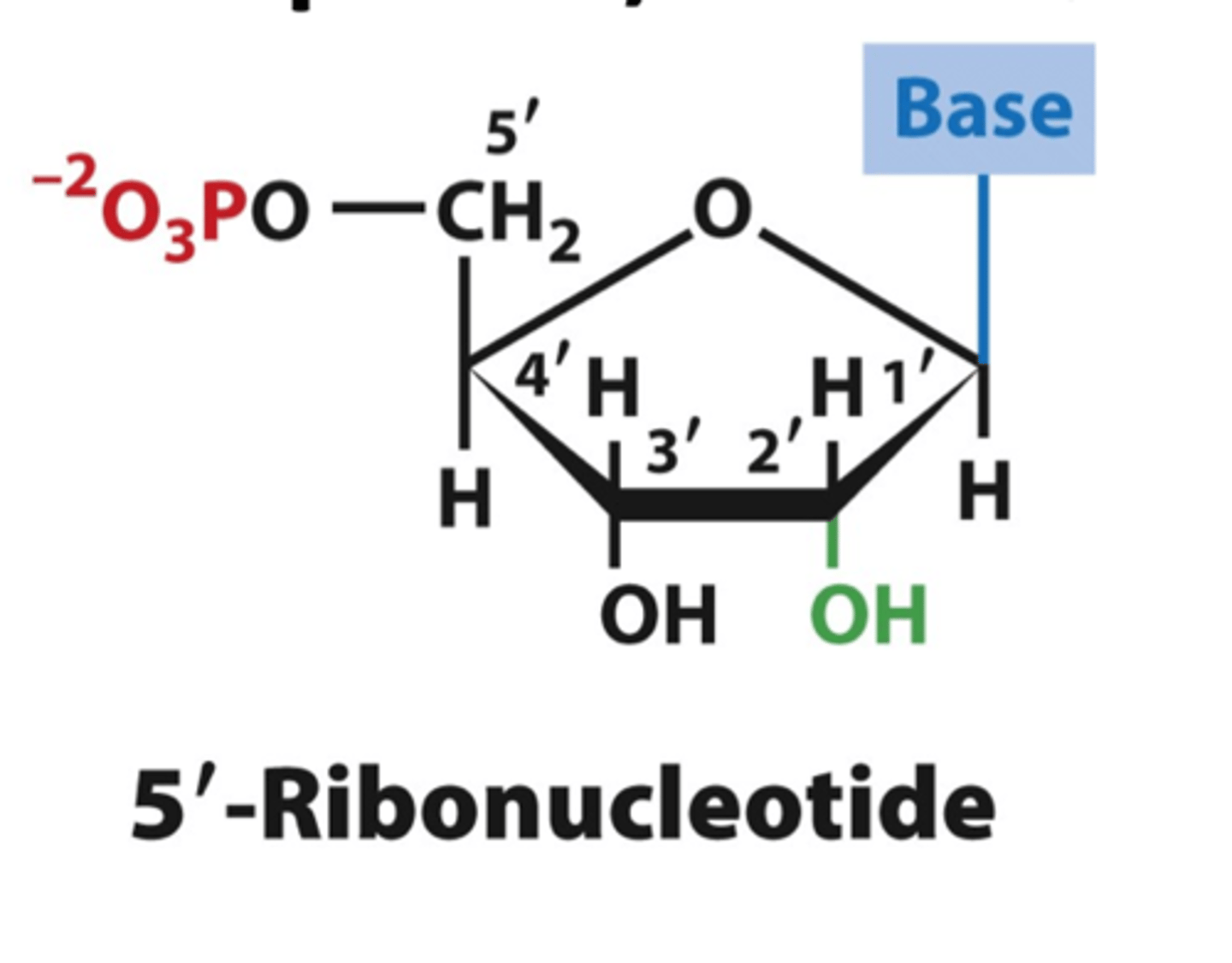
ribose's 2' hydroxyl
gives RNA a relative instability that makes it more suitable for its more short-term functions
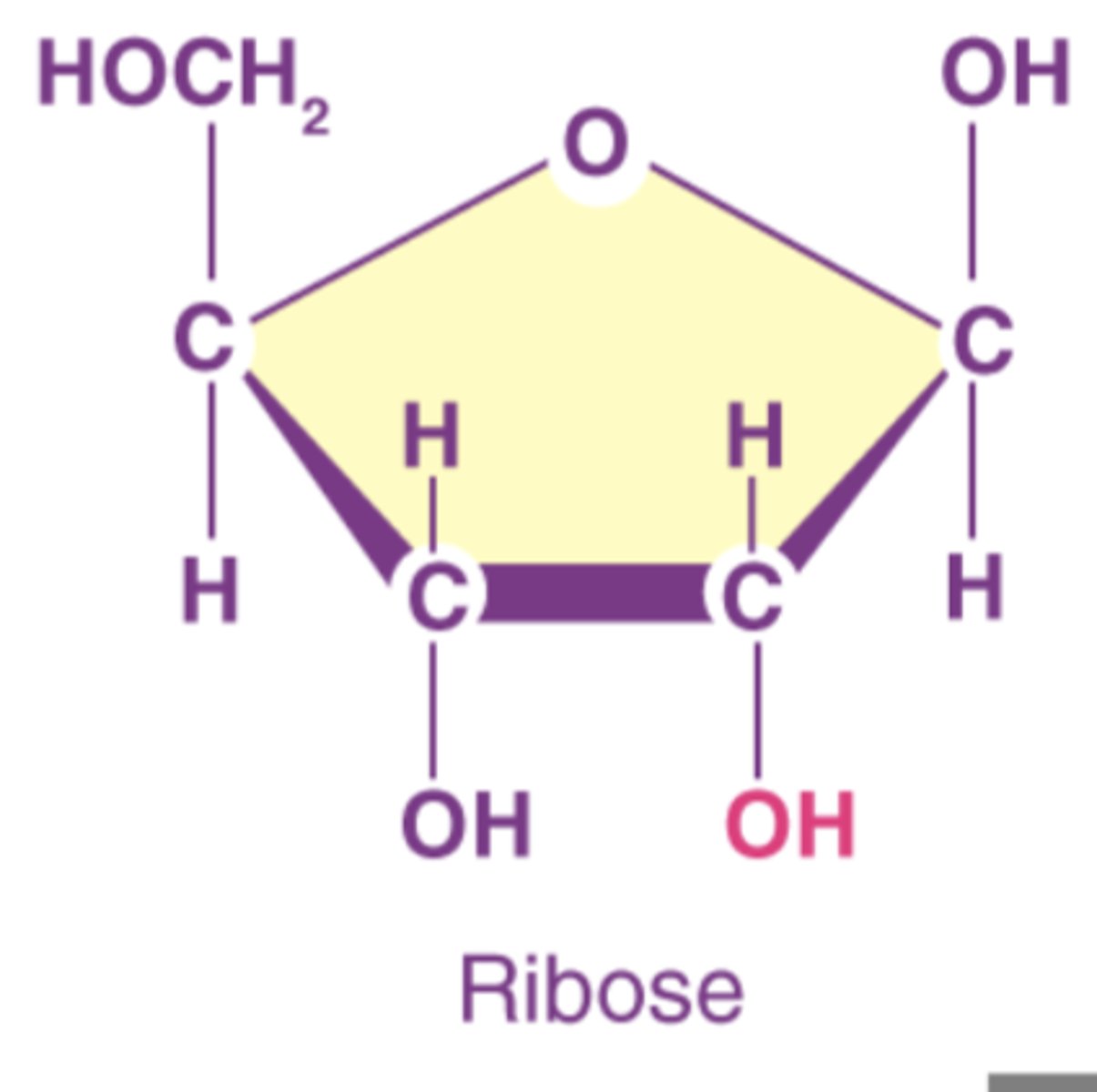
RNA differs from DNA
consists only of one polynucleotide strand
sugar is ribose
has uracil, not thymine
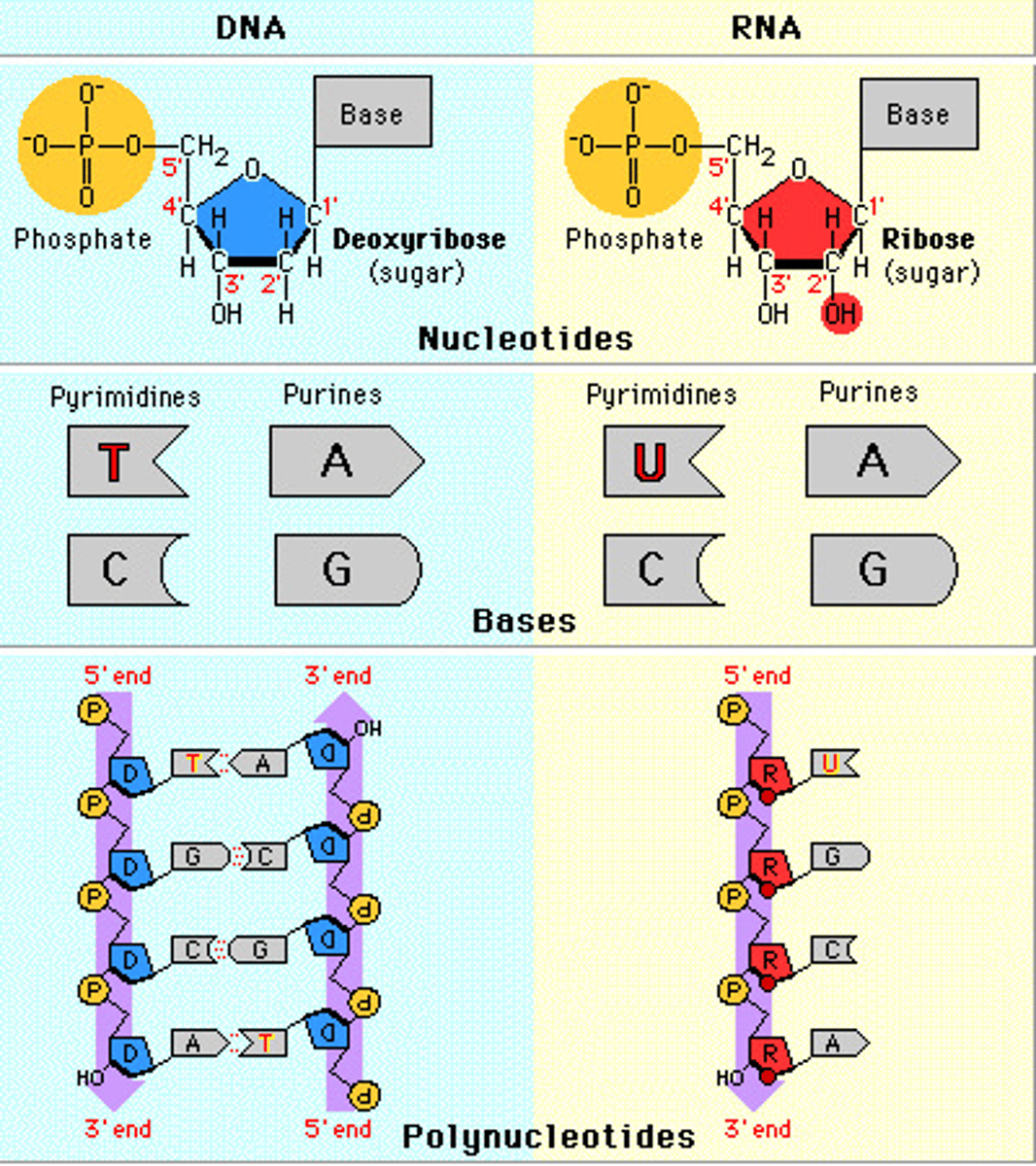
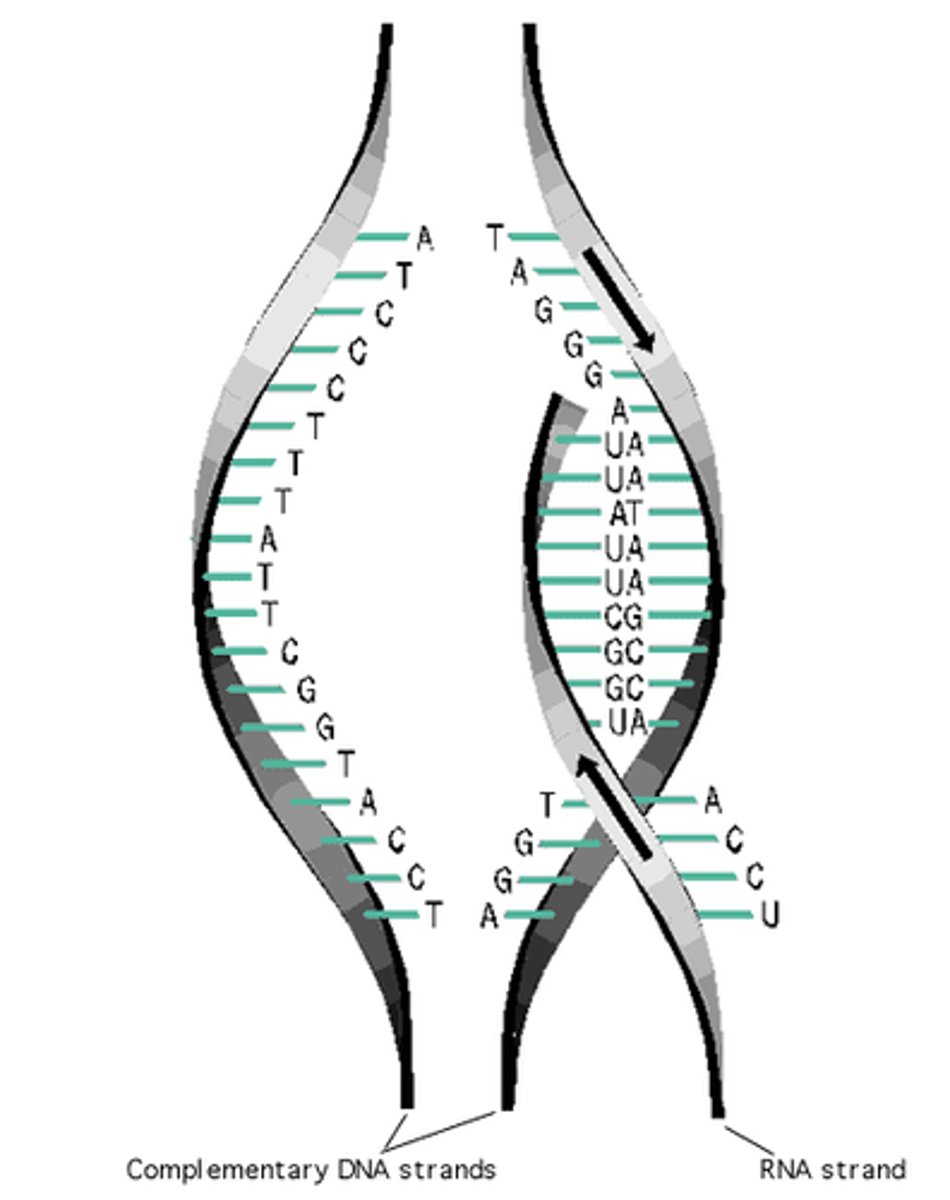
RNA can base-pair with single stranded DNA
crucial for biological processes like transcription, where mRNA is synthesized from a DNA template
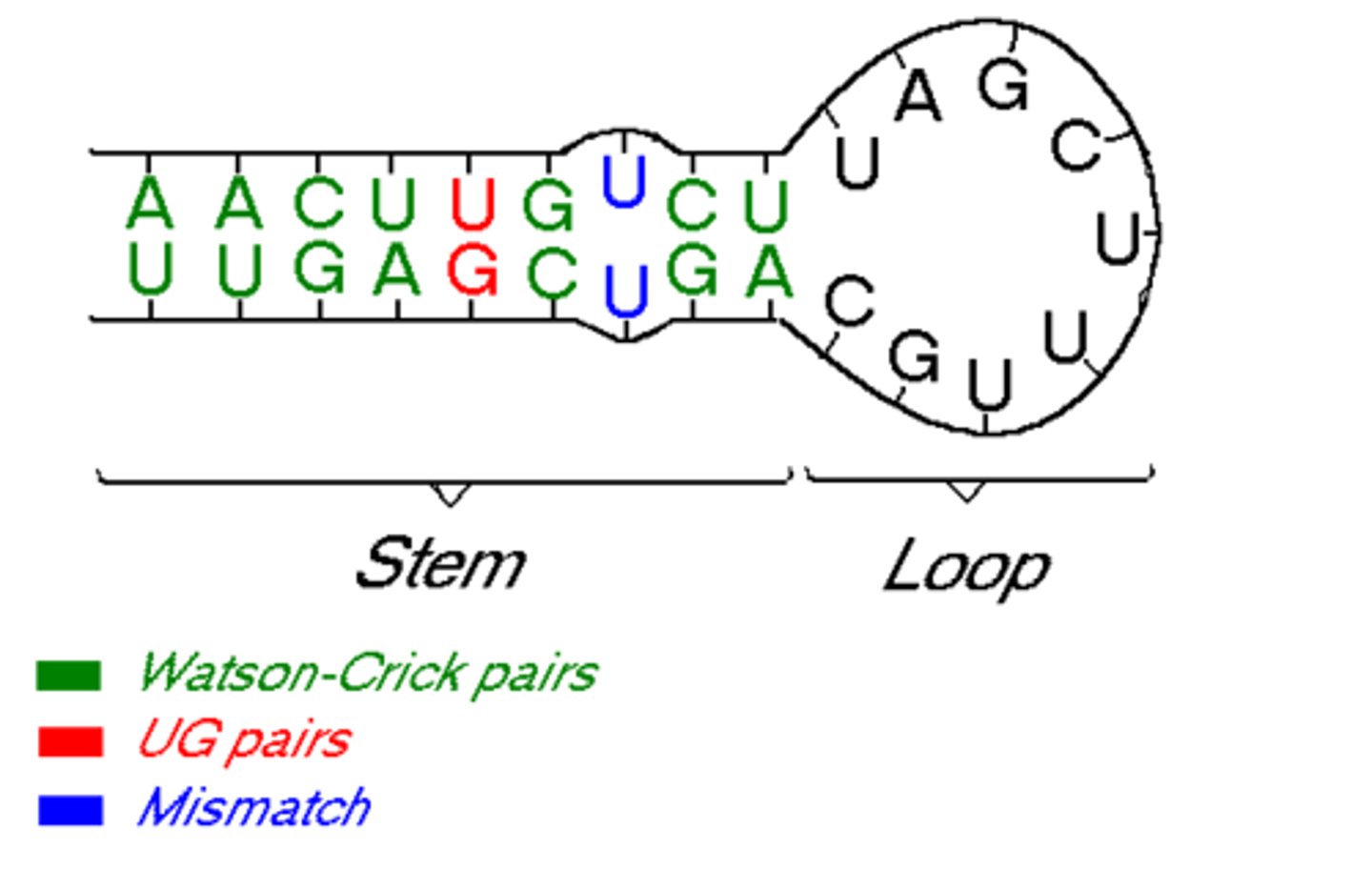
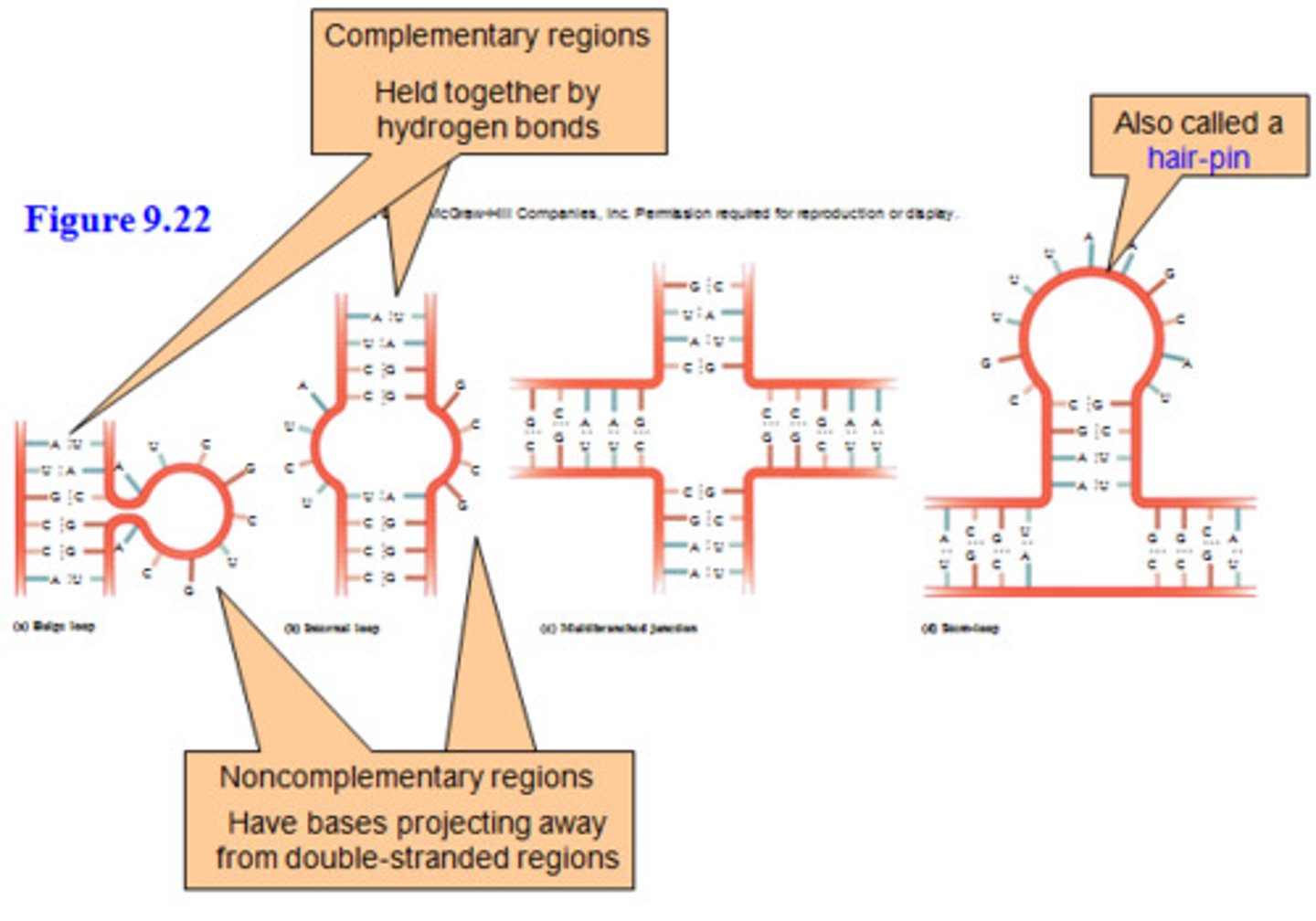
Intramolecular base pairing
RNA can fold over and base pair with itself
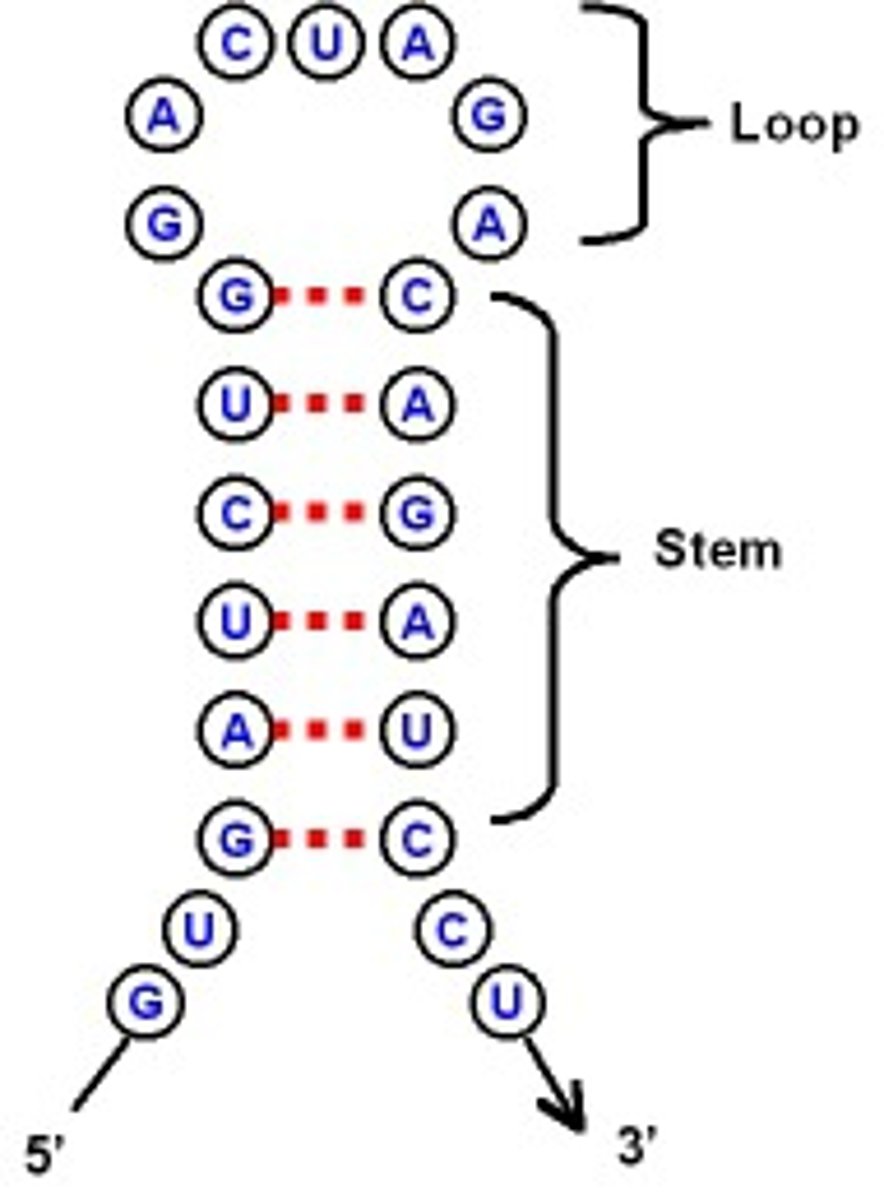
extensive intramolecular base pairing
creates a predictable three-dimensional structure essential for their function.
mRNAs must remain free of secondary structures
otherwise, it cannot be properly decoded during translation
60 -70% of the human genome
actively transcribed into RNA
less than 2% of the human genome
codes for protein (protein-coding genes)
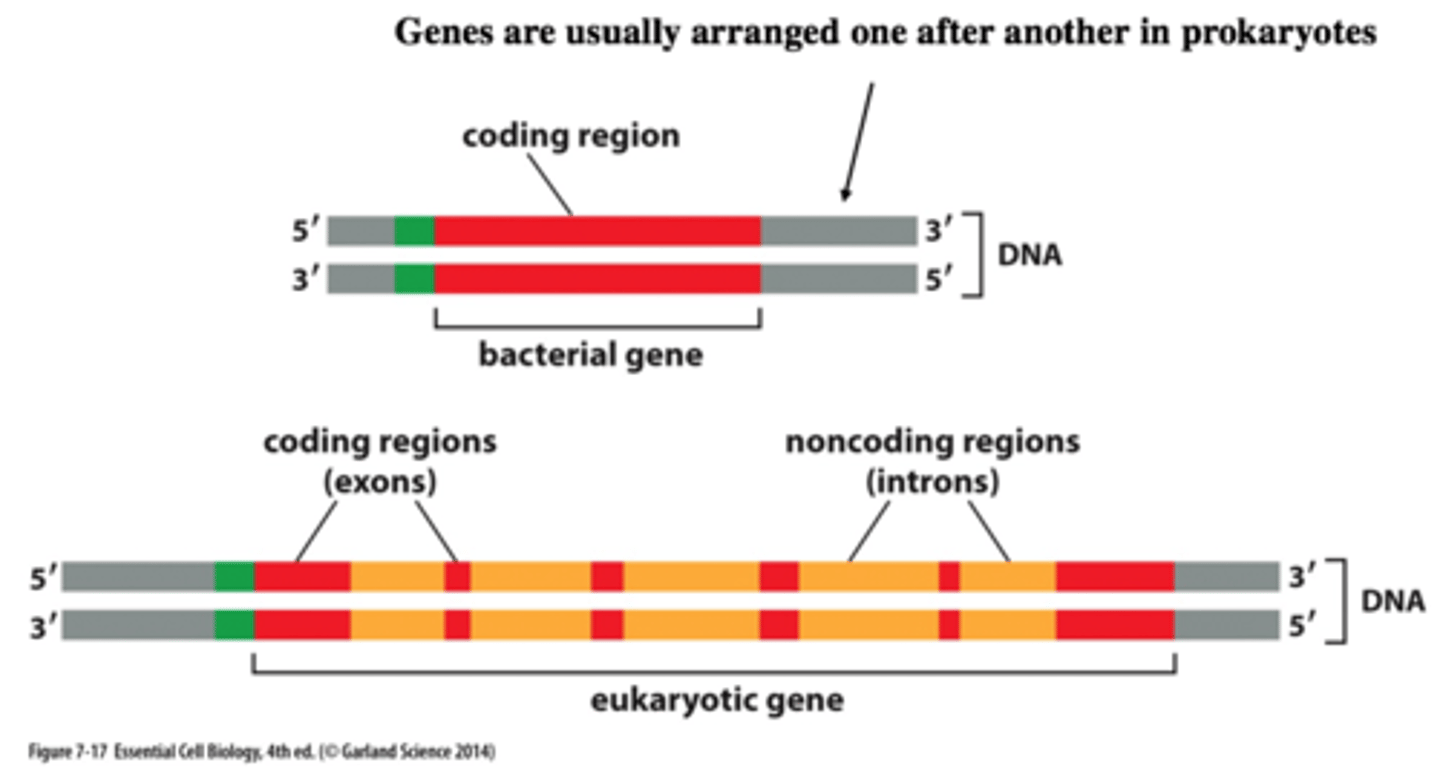
remaining genome %
non-translated RNA/transcript = non-translated or non-coding RNA(ncRNA) molecules
non-coding RNAs
including rRNA, tRNA, microRNAs, long non-coding RNAs (lncRNAs), and other regulatory RNAs
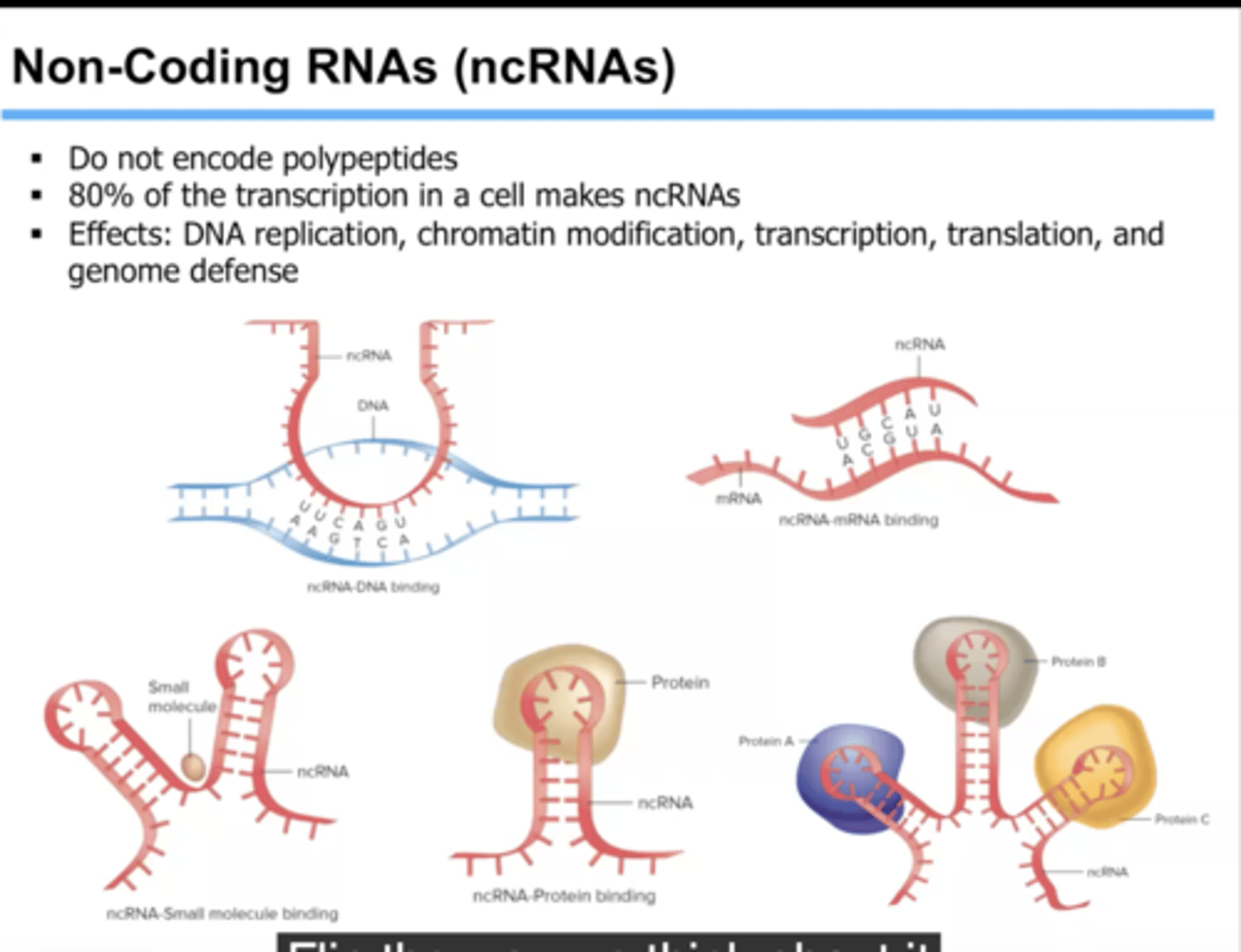
RNA Polymerase enzymes
biosynthesize RNAs in the process of transcription
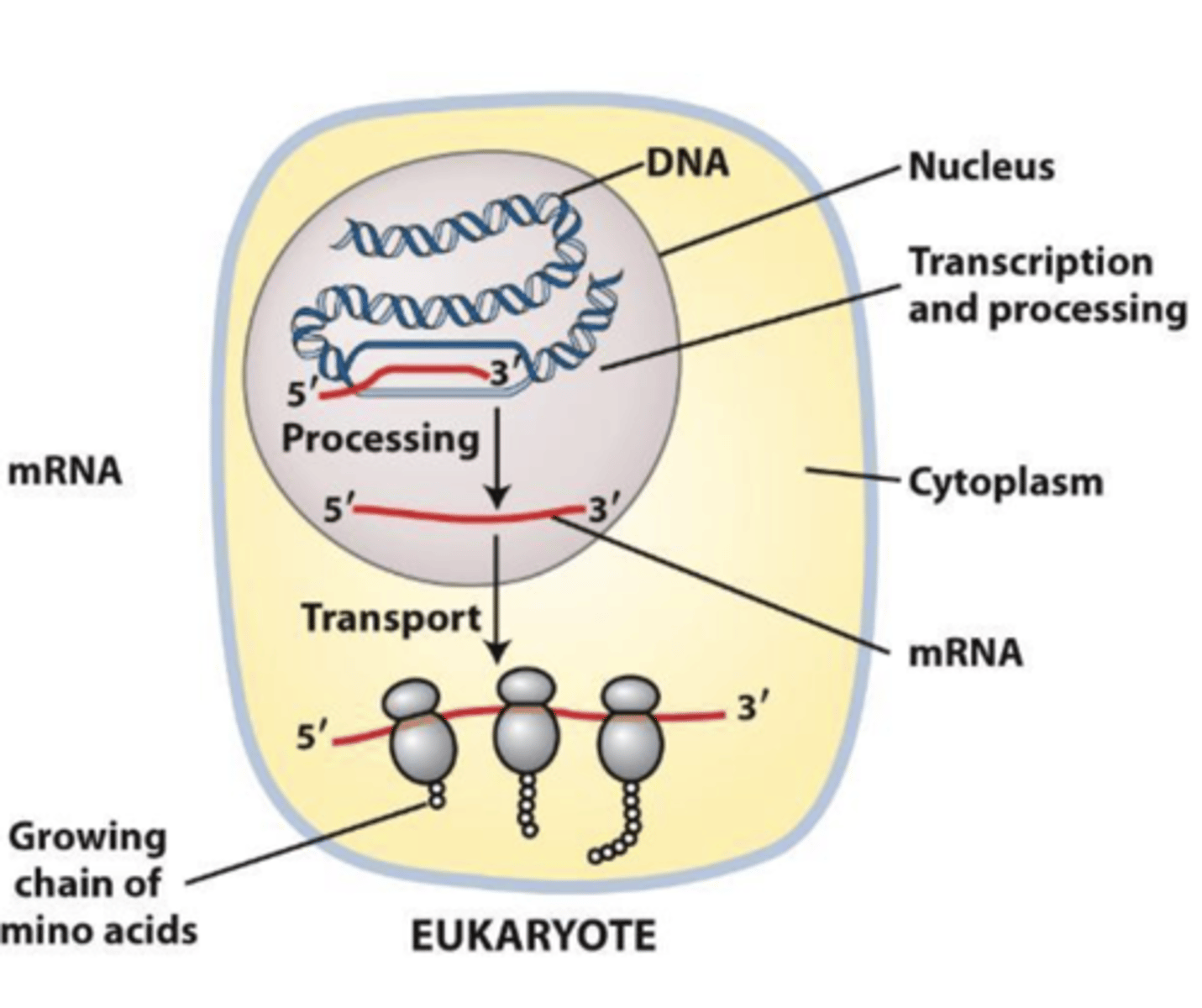
eukaryotes, RNA Polymerases function in the nucleus
transcription and translation are physically separated
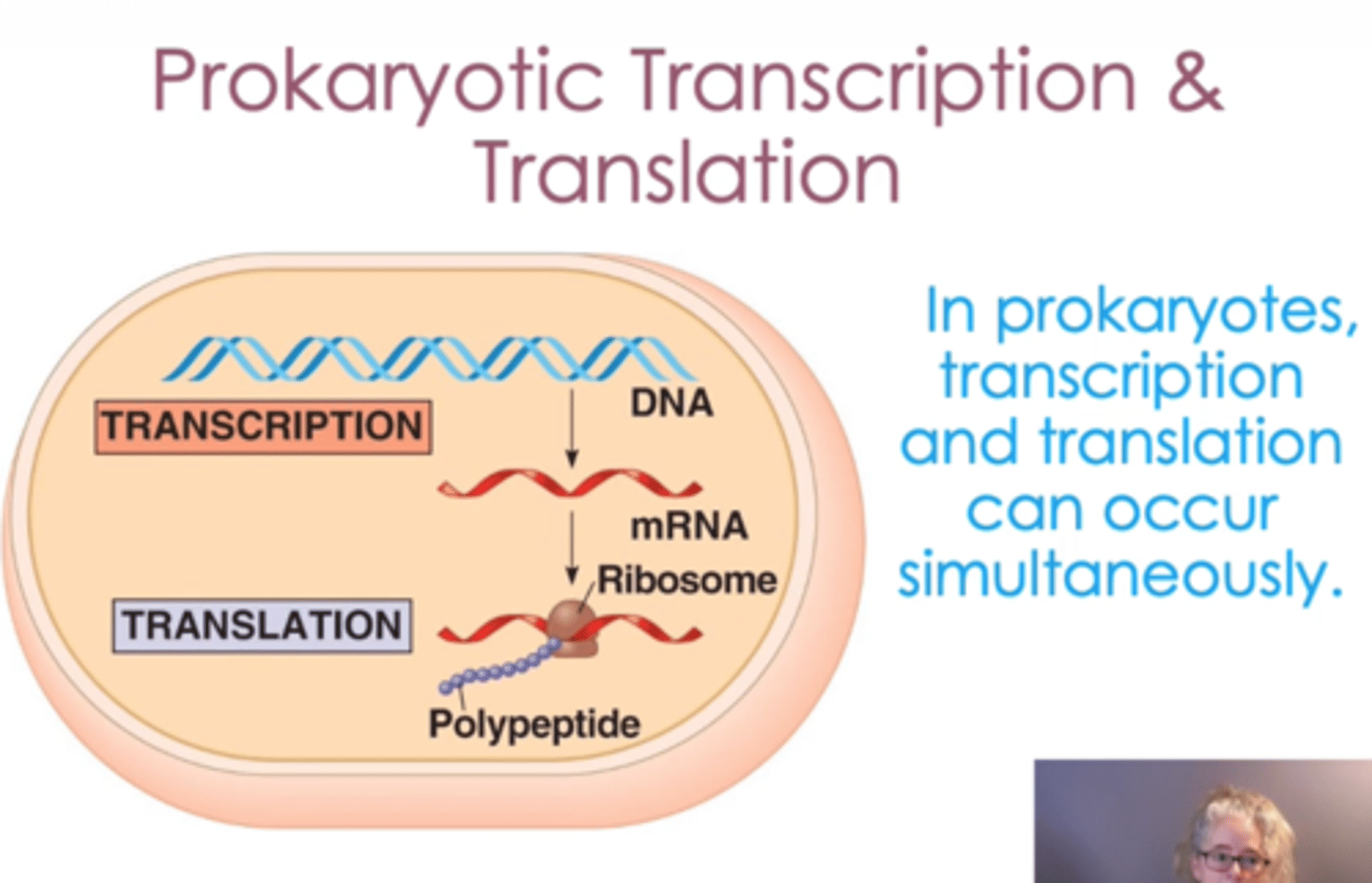
Prokaryotic, RNA Polymerases function in the cytoplasm
transcription and translation occurs simultaneously
Genomic
Gene, Transcription, RNA Processing, Translation and Post translation events
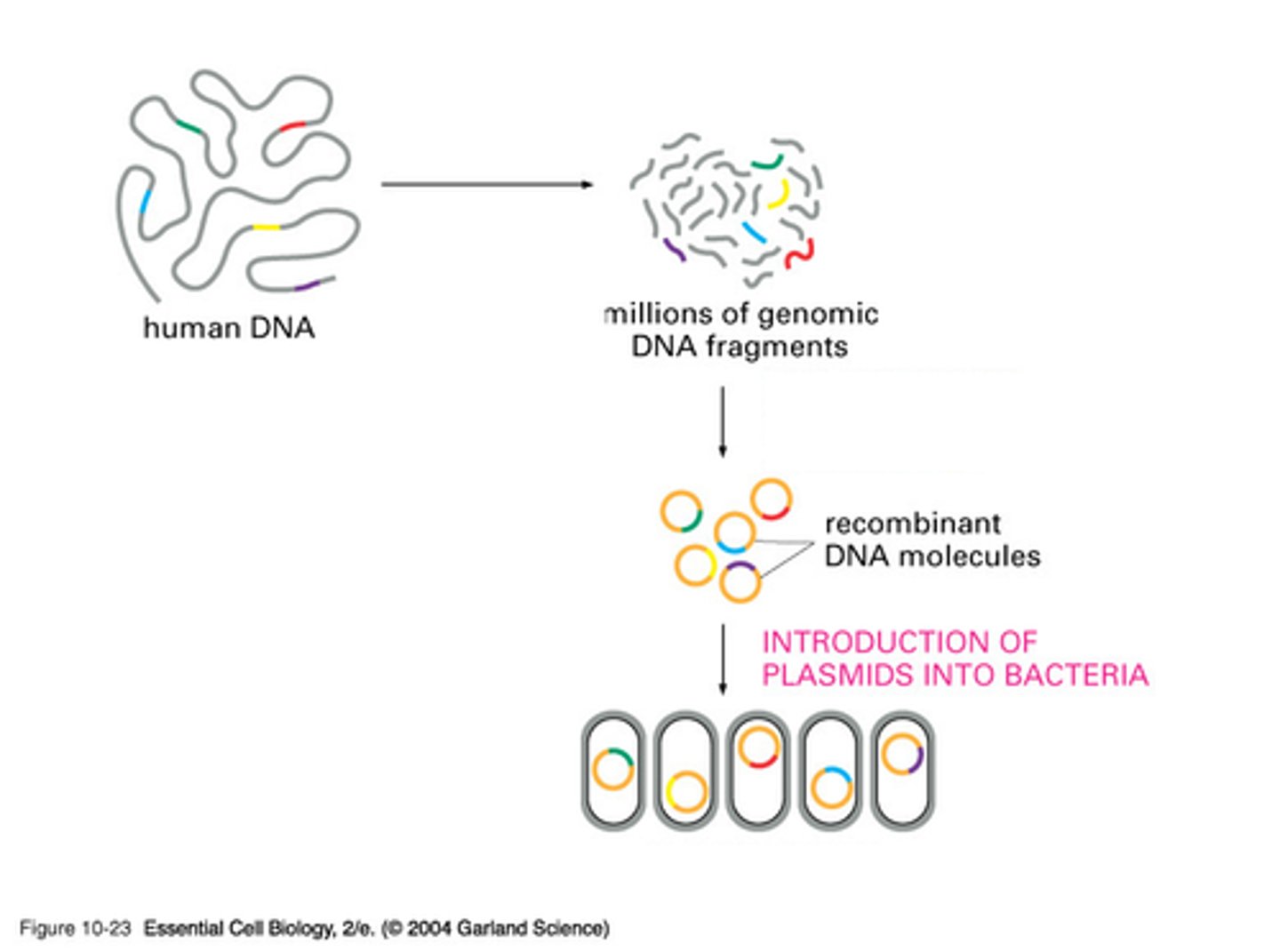
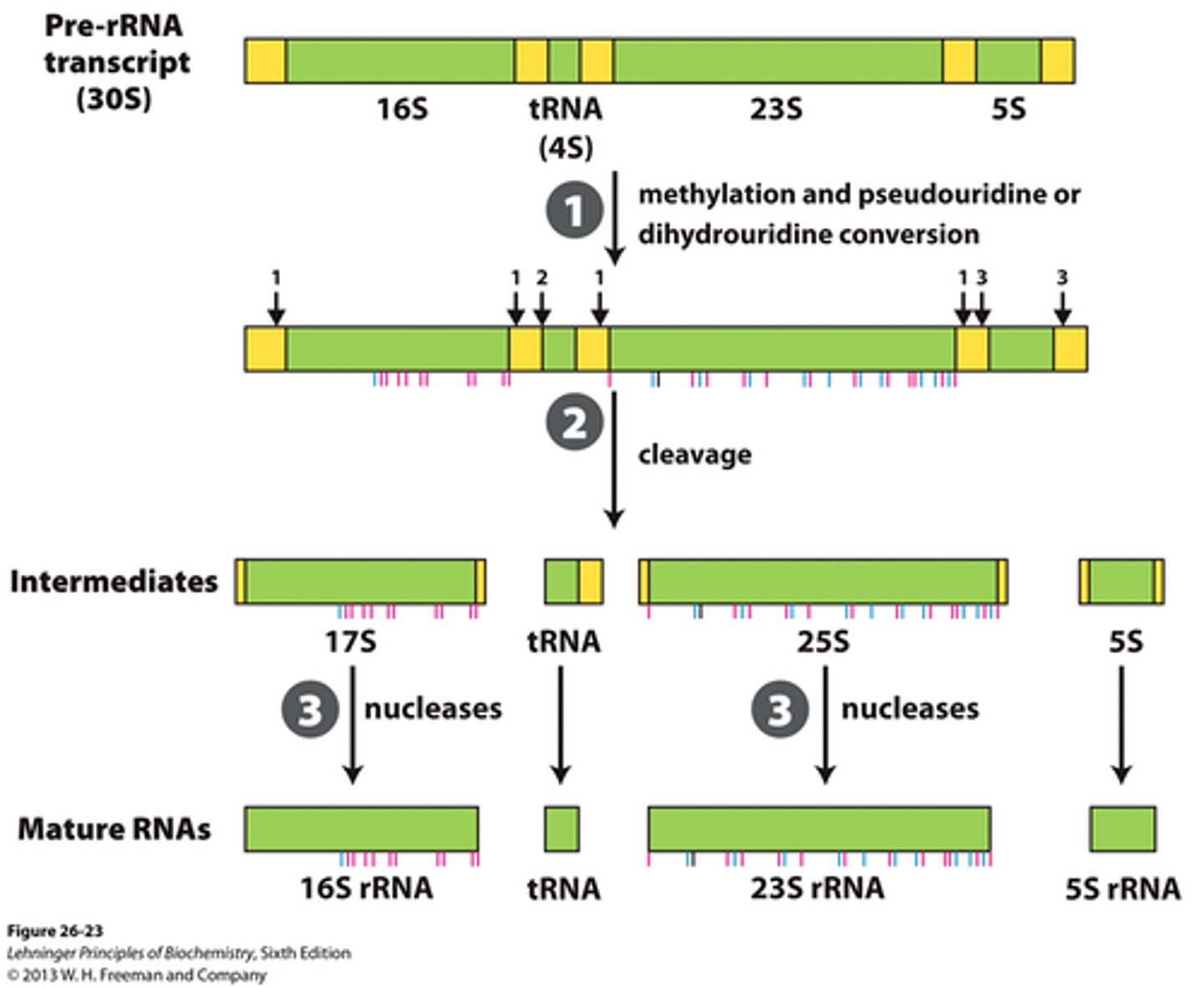
RNA polymerase I
in all eukaryotes; transcribes large rRNAs
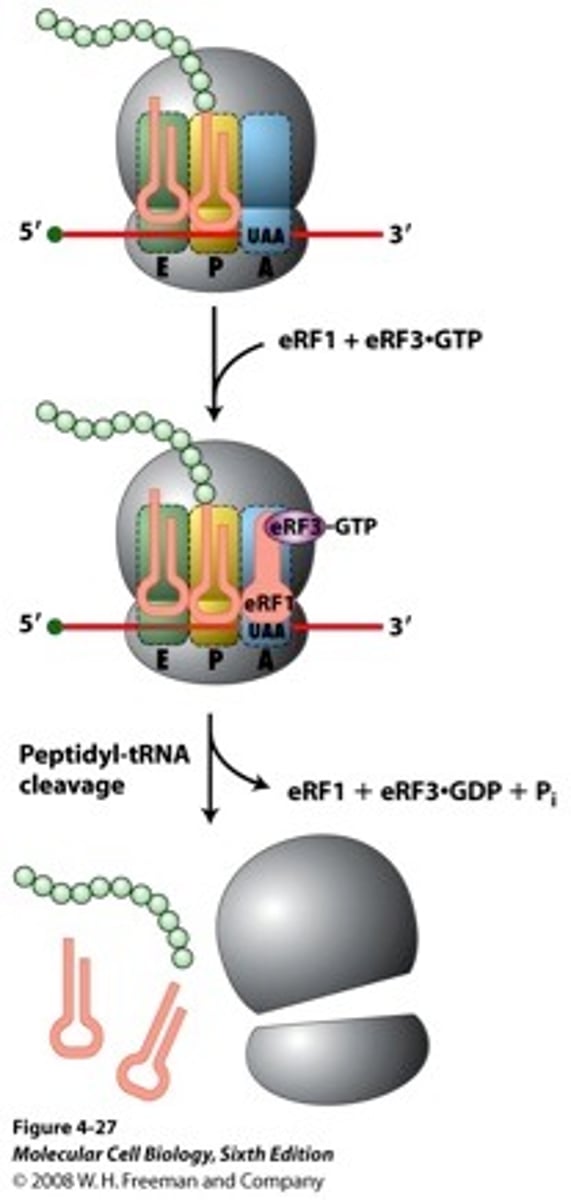
RNA polymerase II
in all eukaryotes; transcribes mRNAs, snoRNAs, snRNAs, and miRNAs
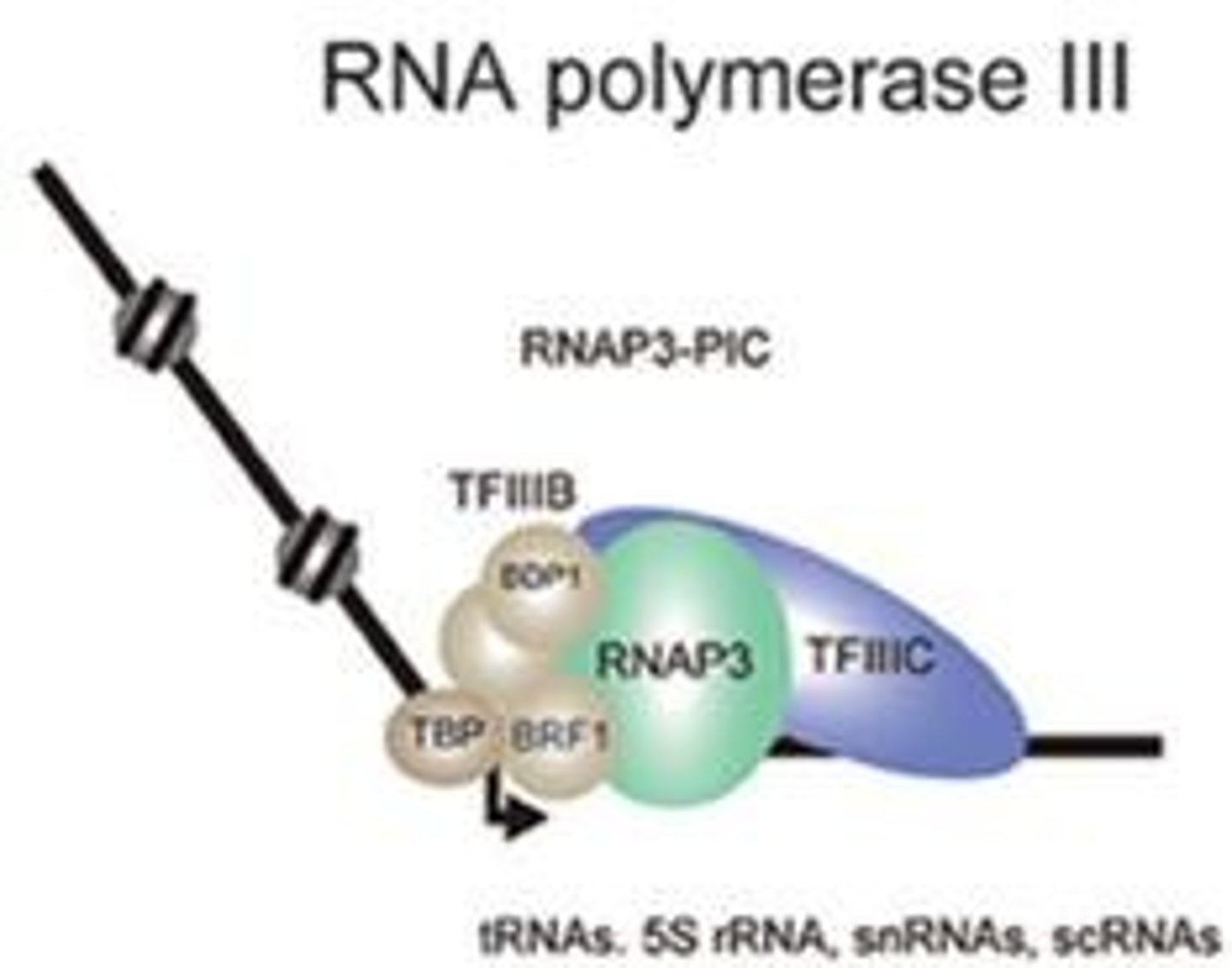
RNA polymerase III
in all eukaryotes; transcribes tRNAs, small rRNAs, snRNAs, and miRNAs
RNA polymerase IV
in plants; transcribes some siRNAs
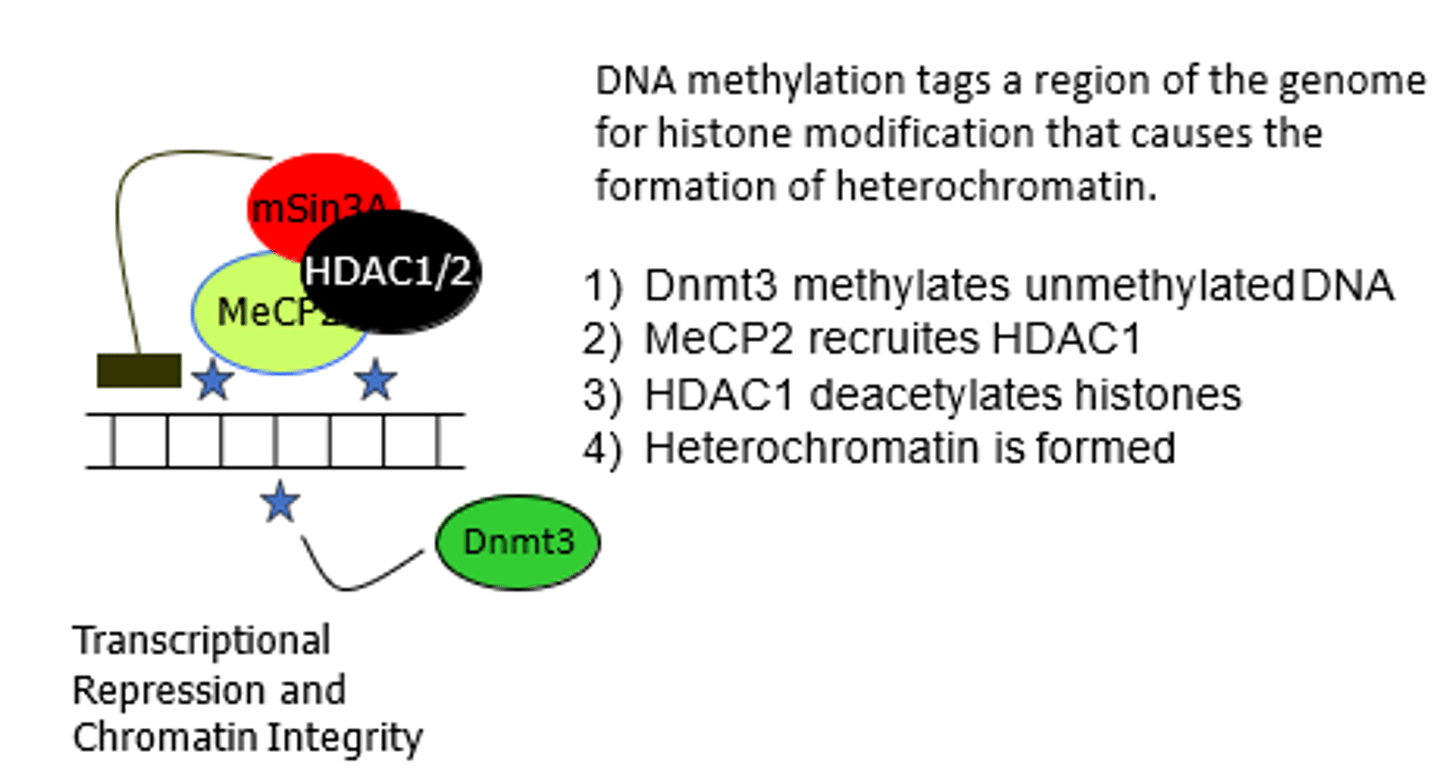
RNA polymerase V
in plants; transcribes RNAs important in heterochromatin formation
RNA Pol II are multi-subunit or multi-domain enzymes
In eukaryotes, RNA Pol II is typically composed of 12 subunits (RPB1‐12)
catalytic core
Ten subunits required for transcription initiation and regulation
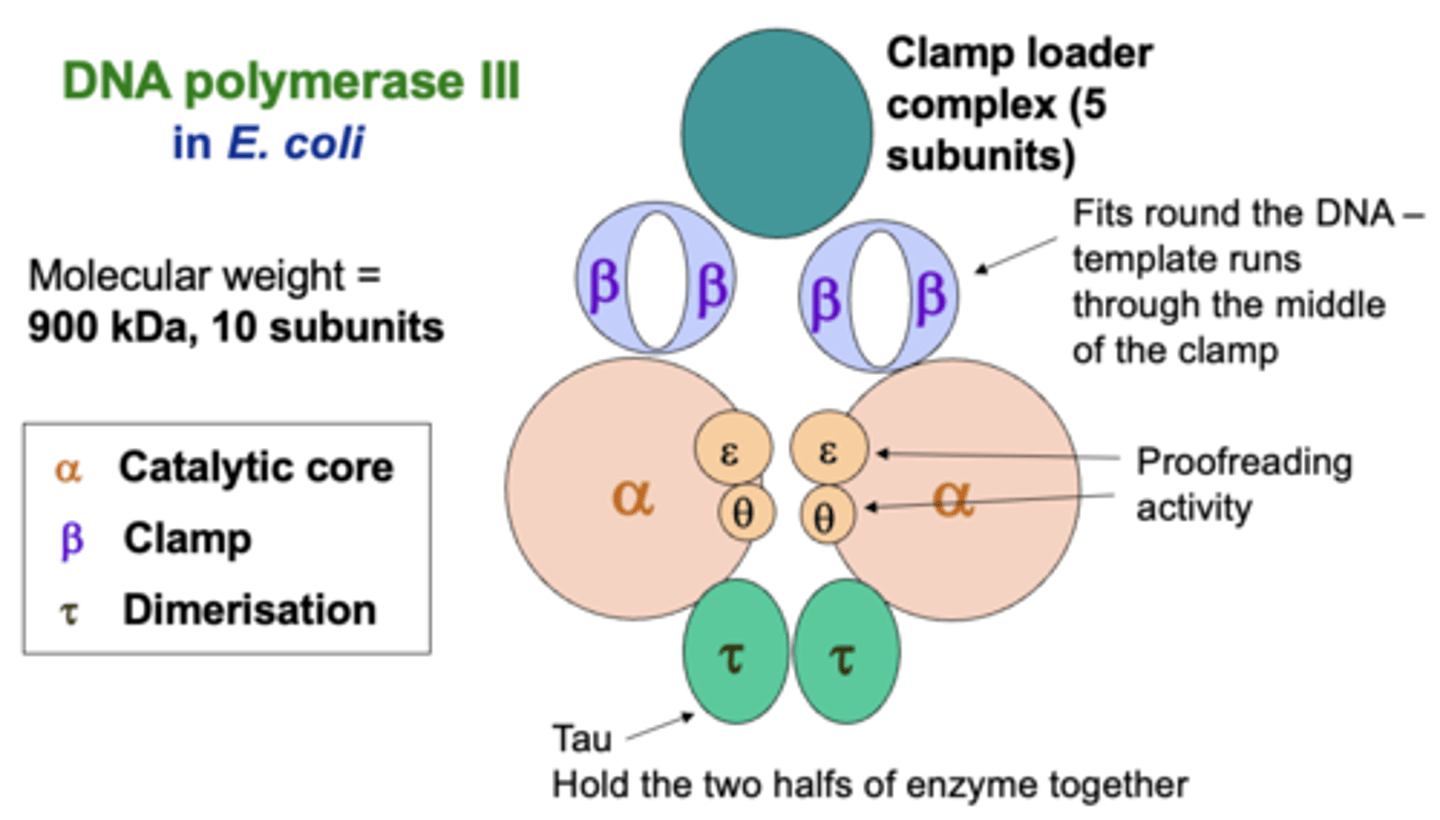
core enzyme
contains the active site where phosphodiester bonds are formed, mainly by the RPB1 and RPB2 subunits
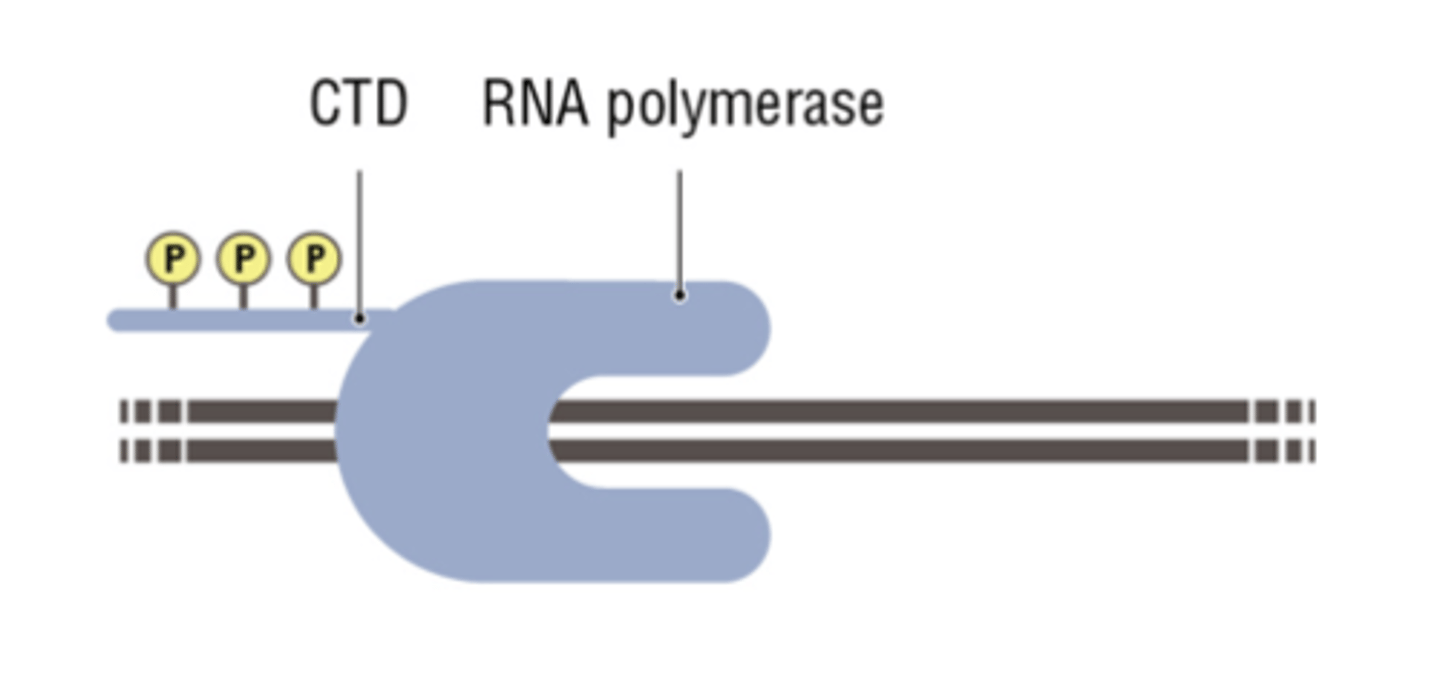
RPB1
the largest subunit, is essential for polymerase activity through its carboxy terminal domain (CTD)
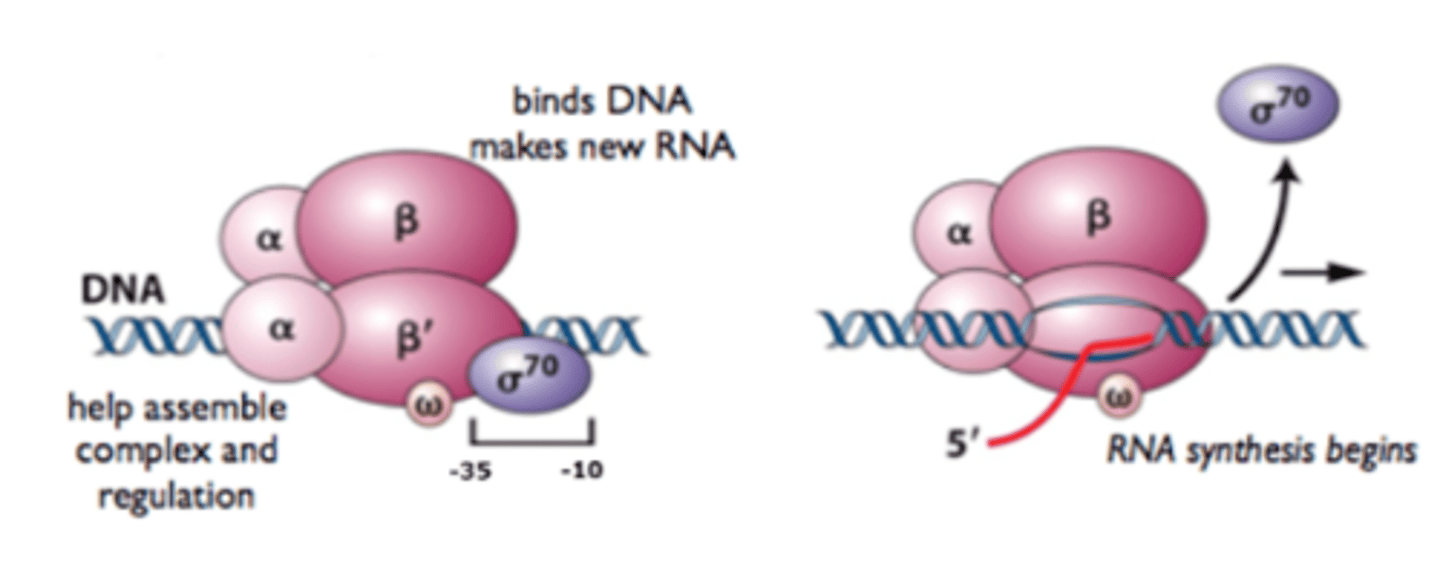
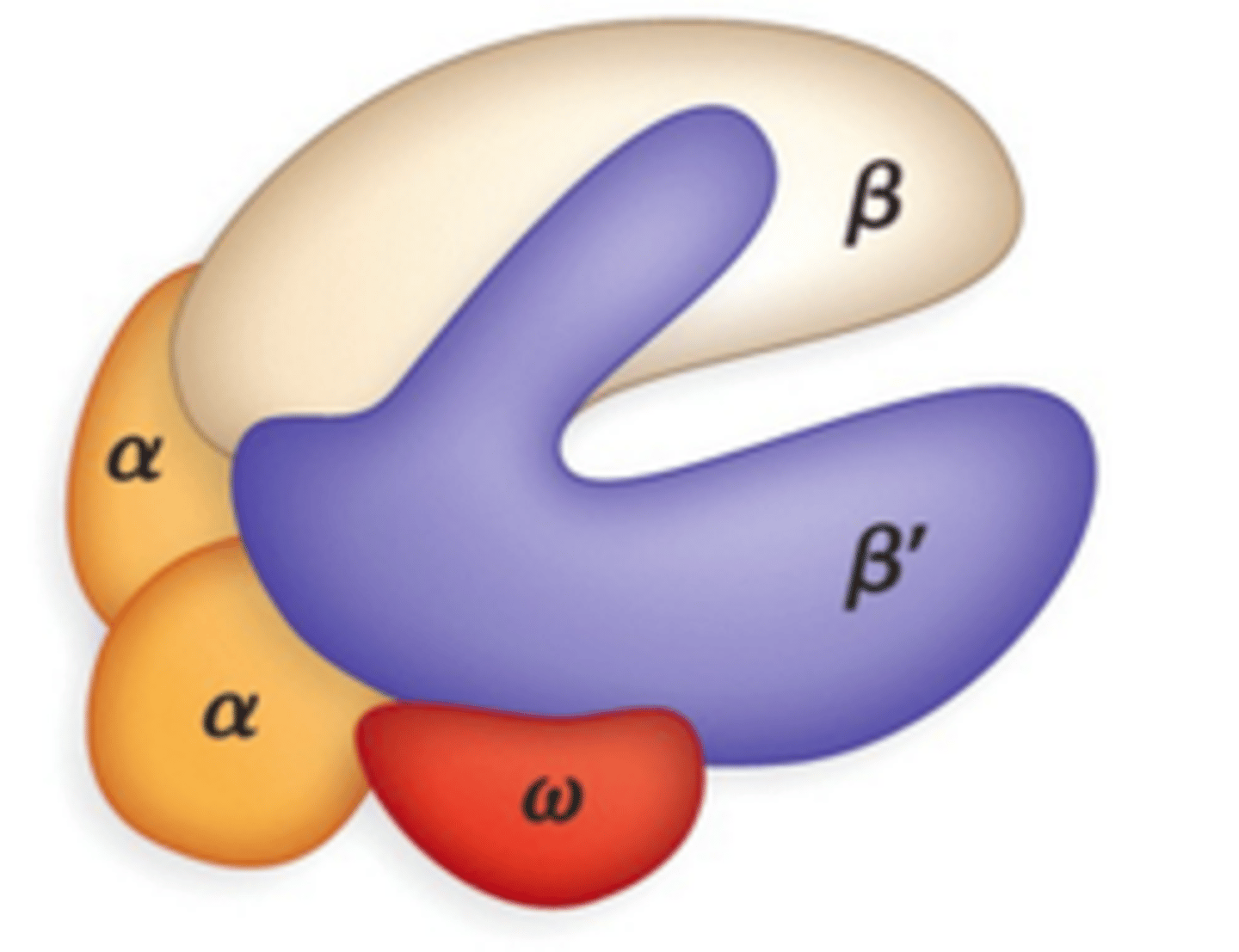
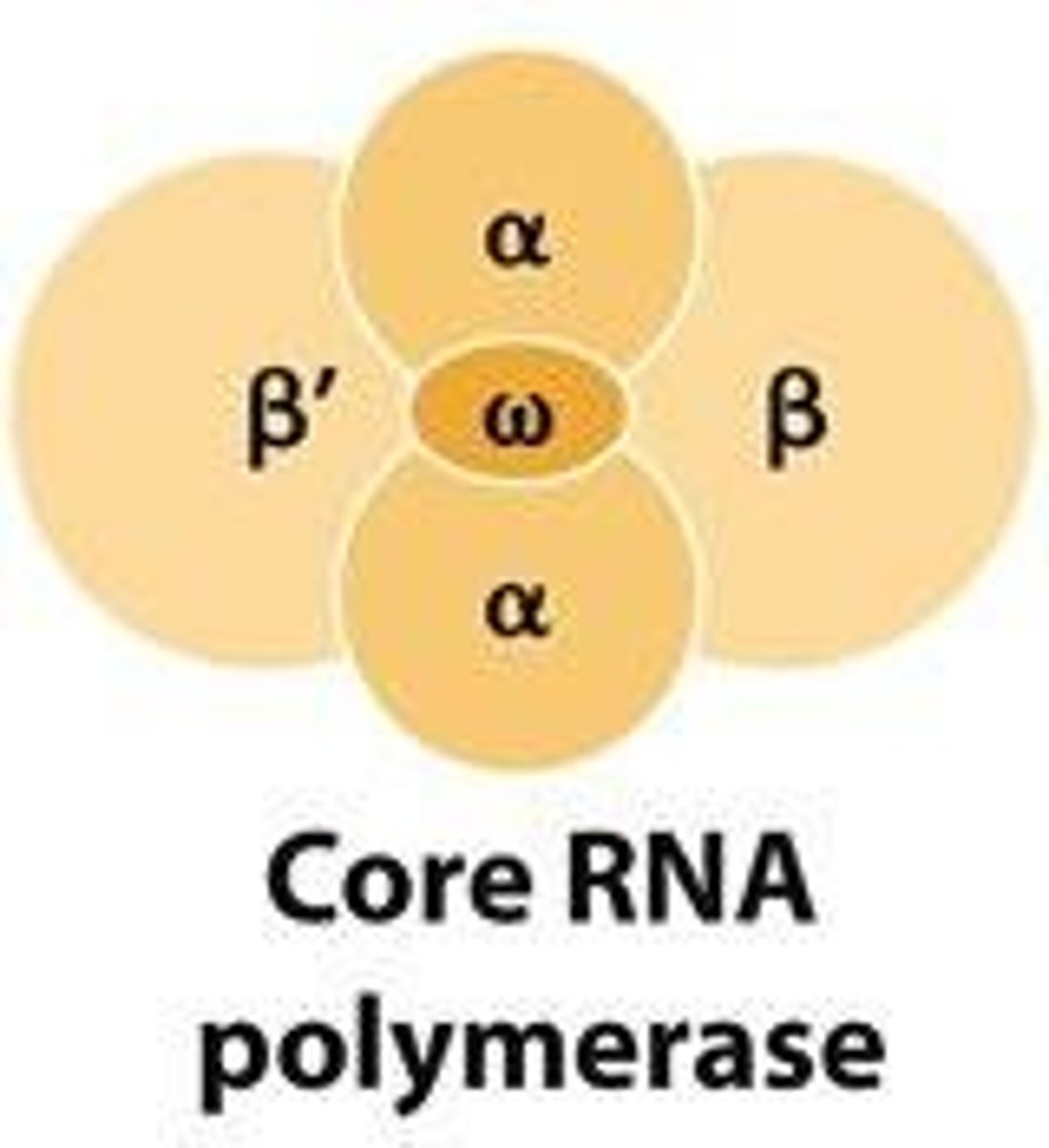
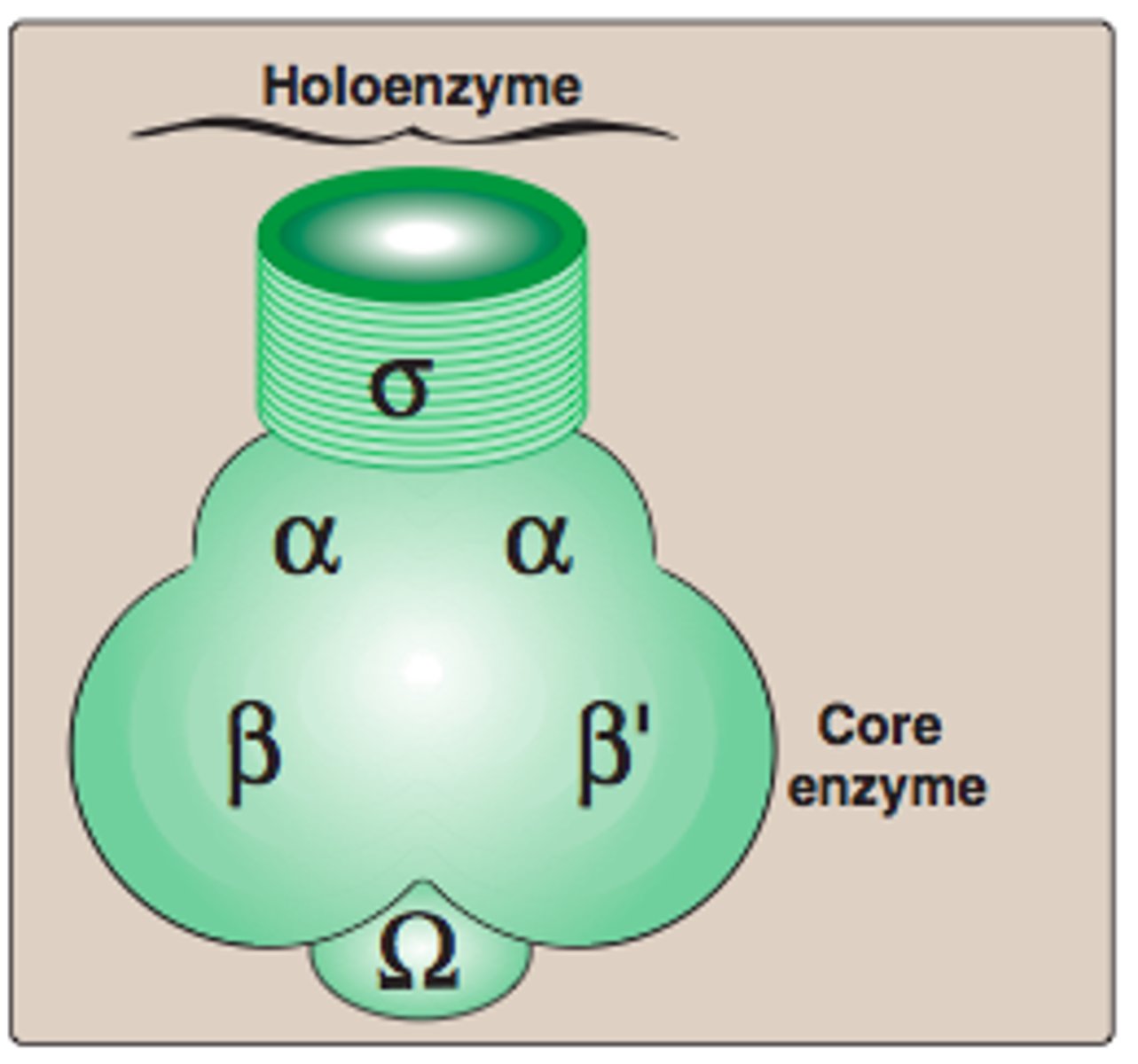
Bacterial Core RNA polymerase's
enzyme that catalyzes phosphodiester bond formation during RNA synthesis
Bacterial Core RNA polymerase's 5 subunits
2 α (alpha) subunits
1 β (beta) subunit
1 β′ (beta prime) subunit
1 ω (omega) subunit
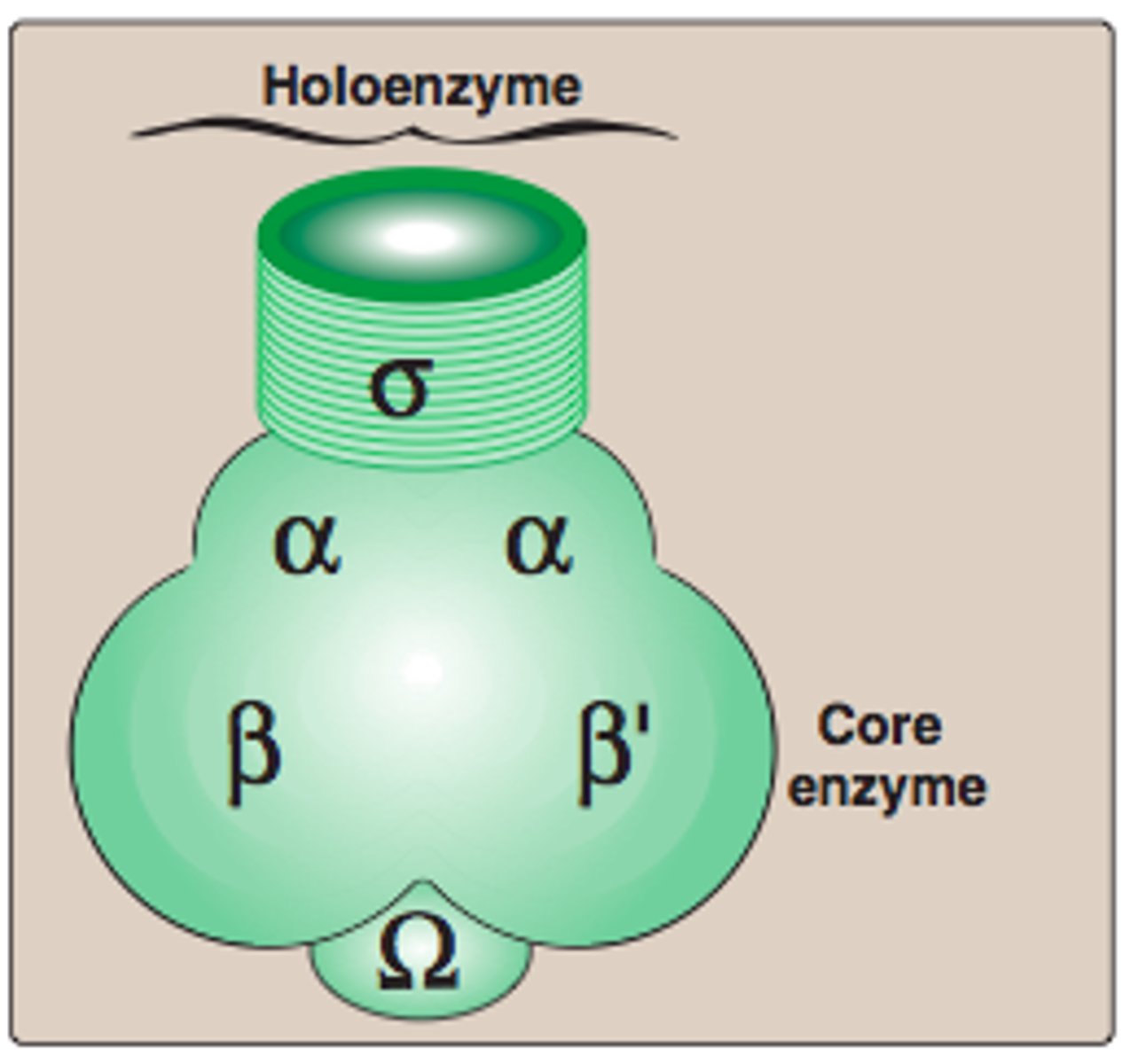
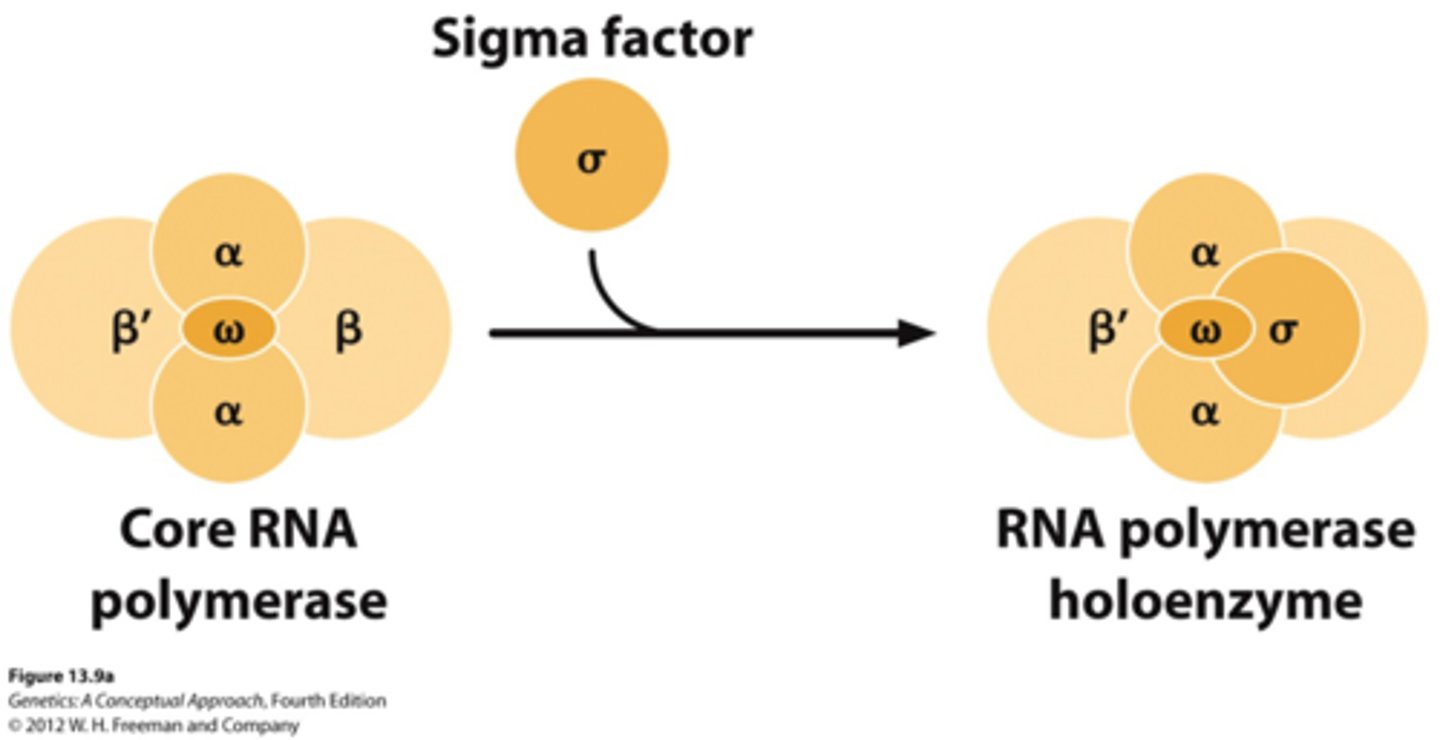
Holoenzyme (6 subunits)
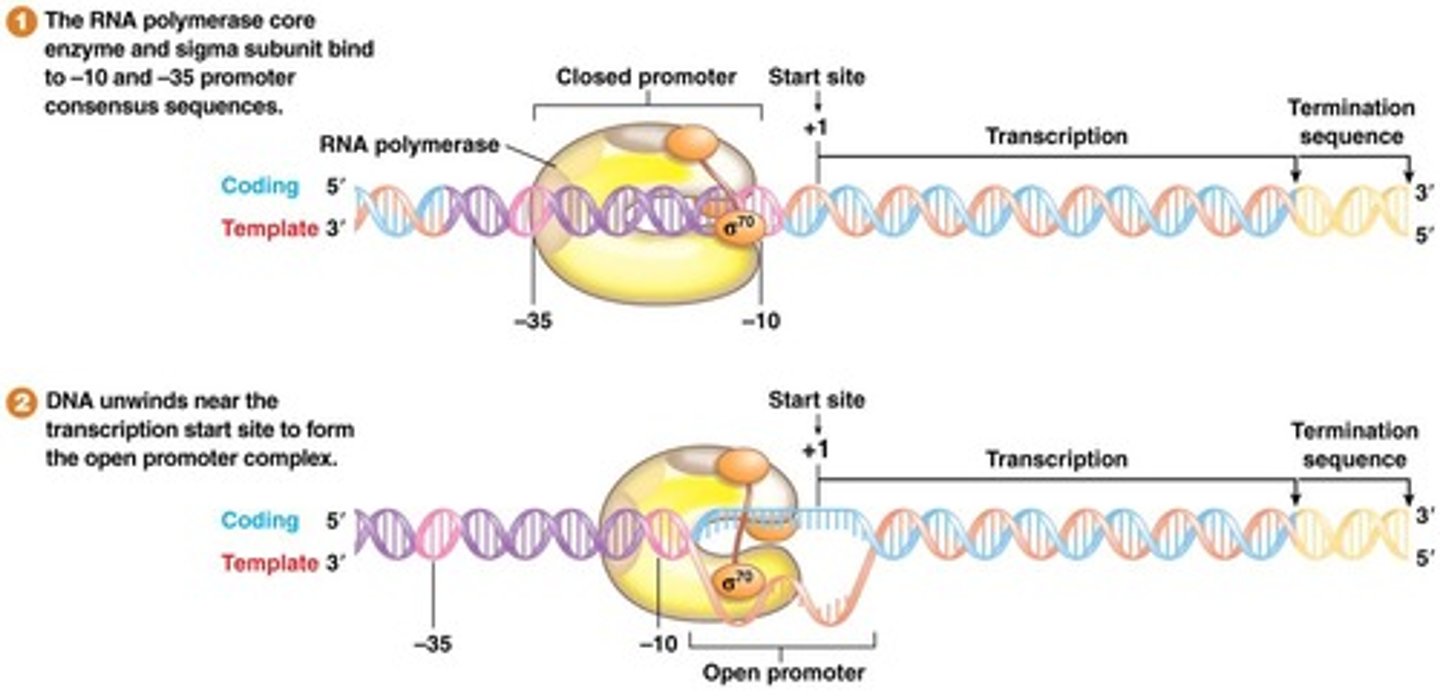
For transcription initiation, the core enzyme associates with a σ (sigma)factor
- α₂ β β′ ω + σ
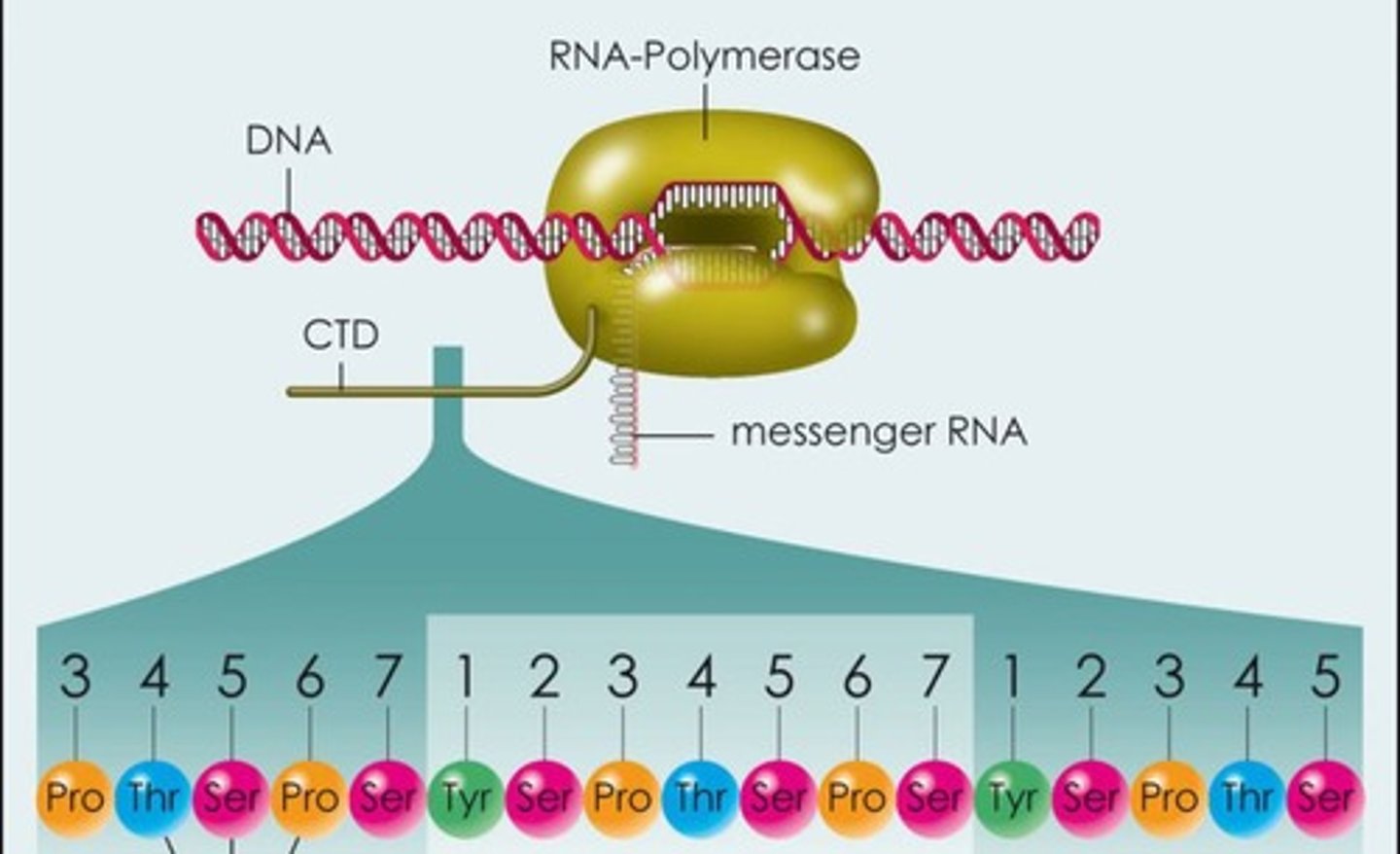
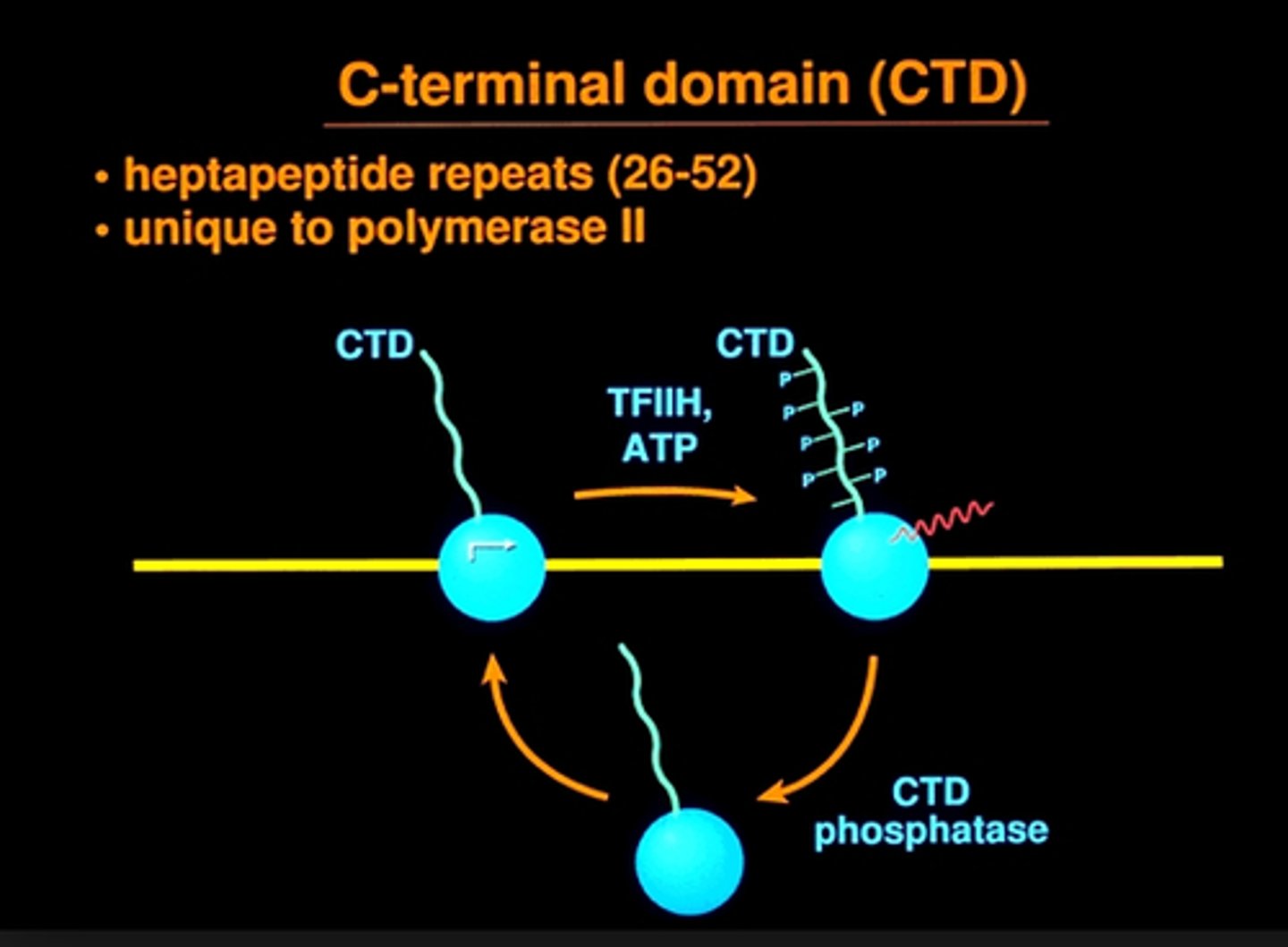
Carboxyl-terminal domain (CTD)
composed of tandem heptad amino acid repeats that constitutes a unique feature of RNAP II and distinguishes it from all other polymerases
CTD important roles
enhancing or modulating the efficiency of all of the RNA processing reactions required for completion of synthesis of the mature RNA
what determines CTD activity
The phosphorylation state
kinases
add phosphate groups on specific proteins, in a process called phosphorylation
- tells RNA pol II to start
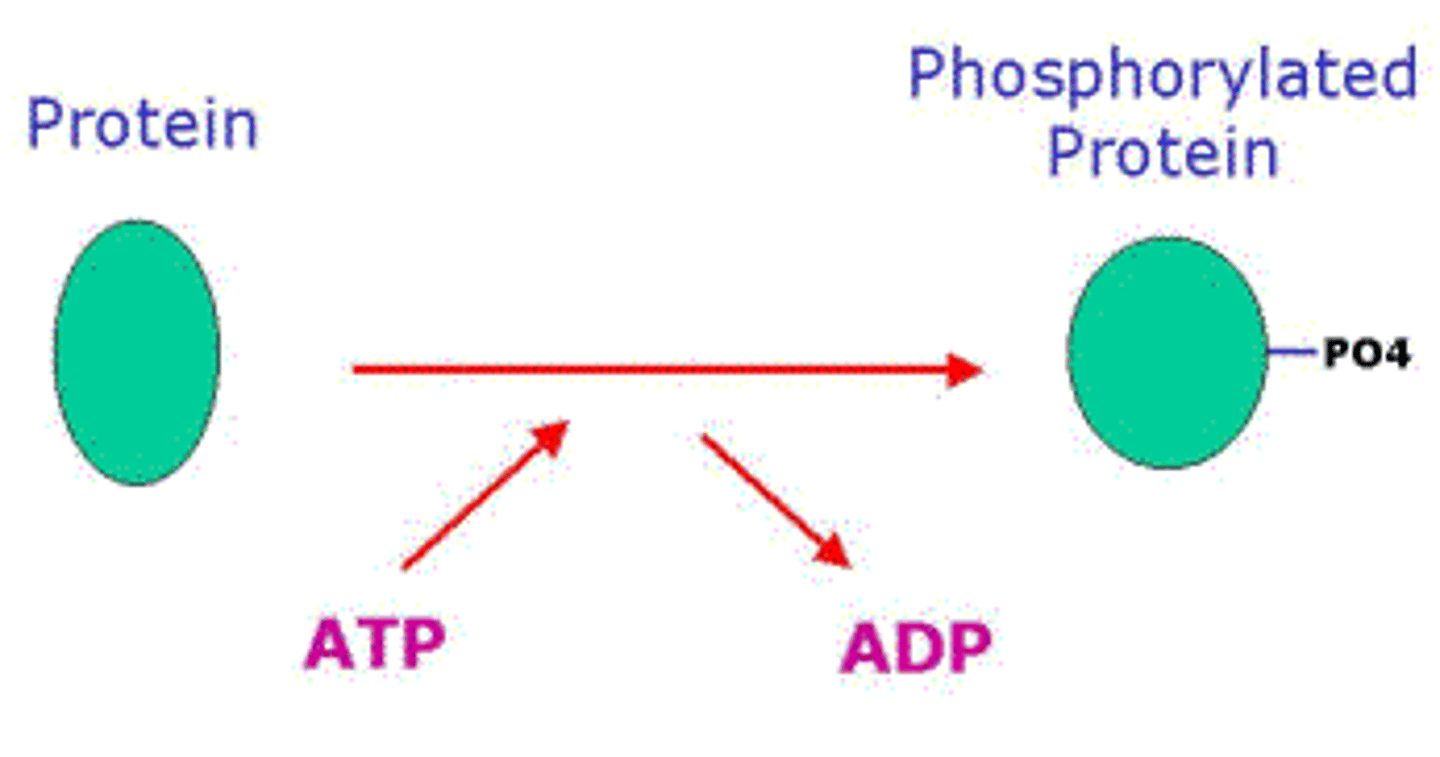
phosphatase
remove phosphate groups (dephosphorylation) from a specific protein
- tells RNA pol II to stop
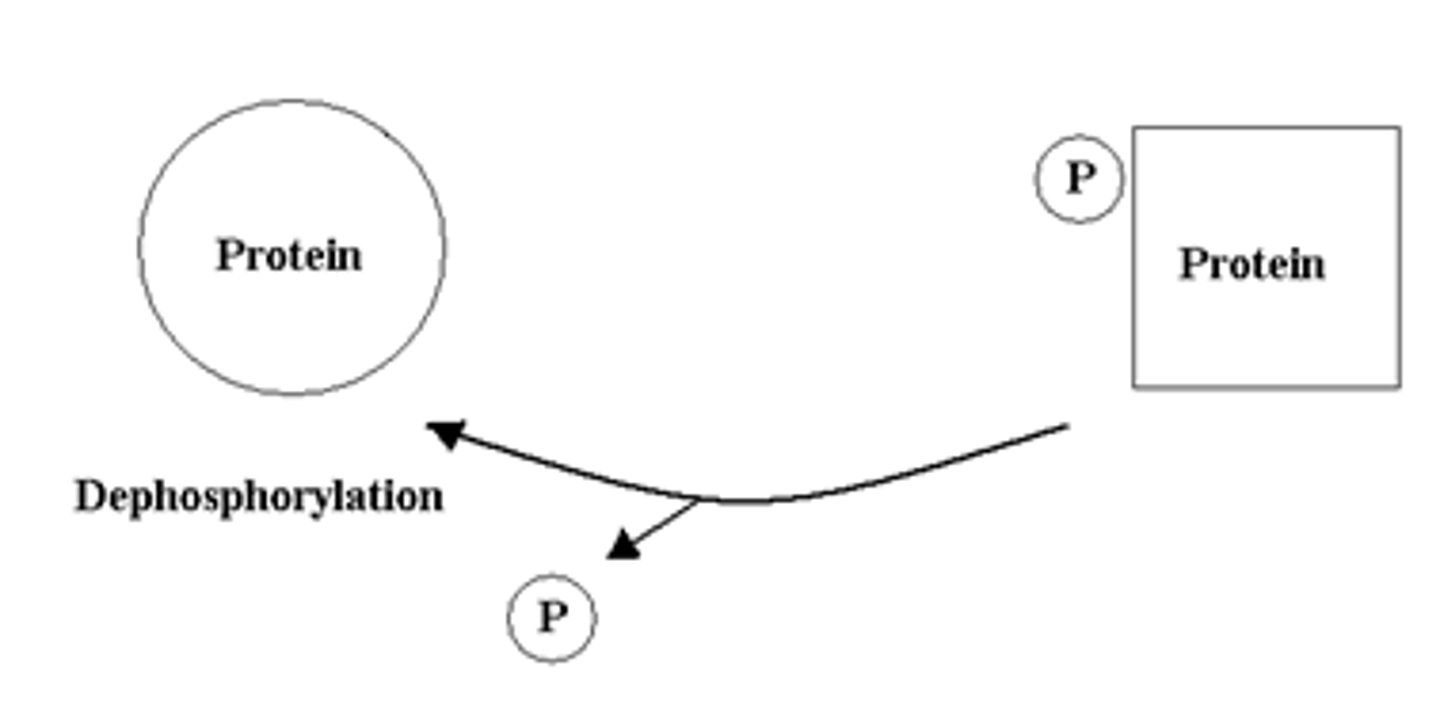
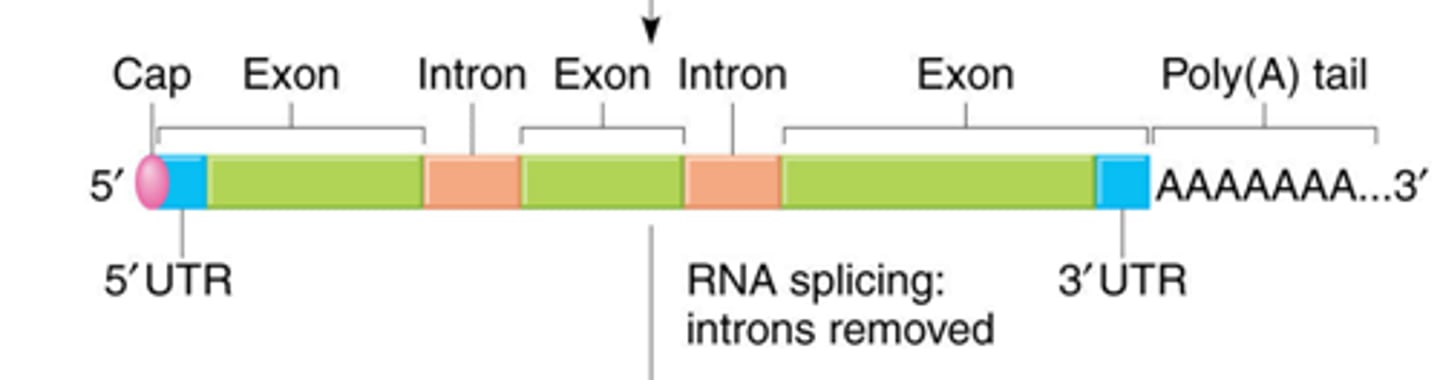
mRNA processing
introns removal, 5'-Capping, Poly A tail addition
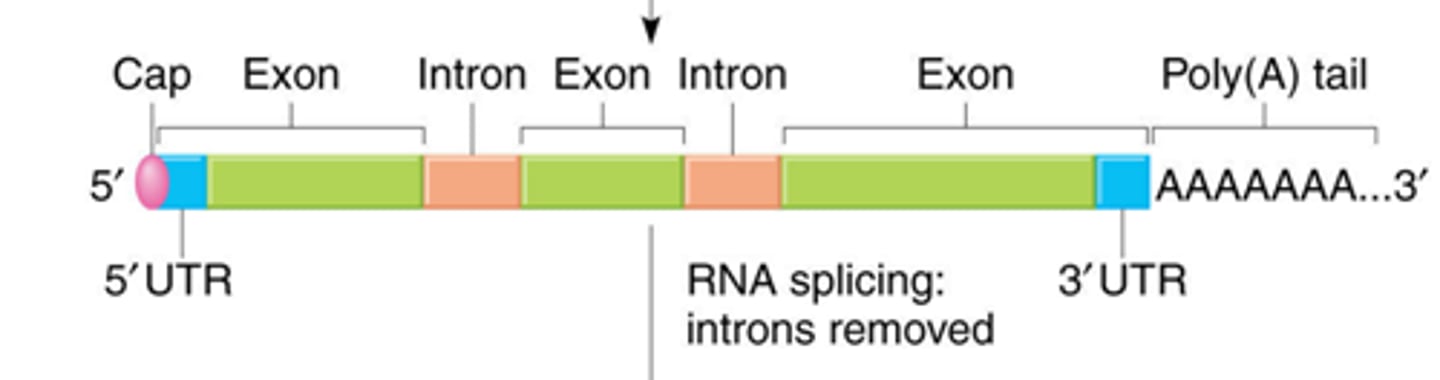
Enzymes required for mRNA processing
attach to the CTD tail of RNA Pol II Rpb1.
As RNA Pol II is transcribing the mRNA these enzymes can "jump" off the CTD tail and carrying out their mRNA processing function
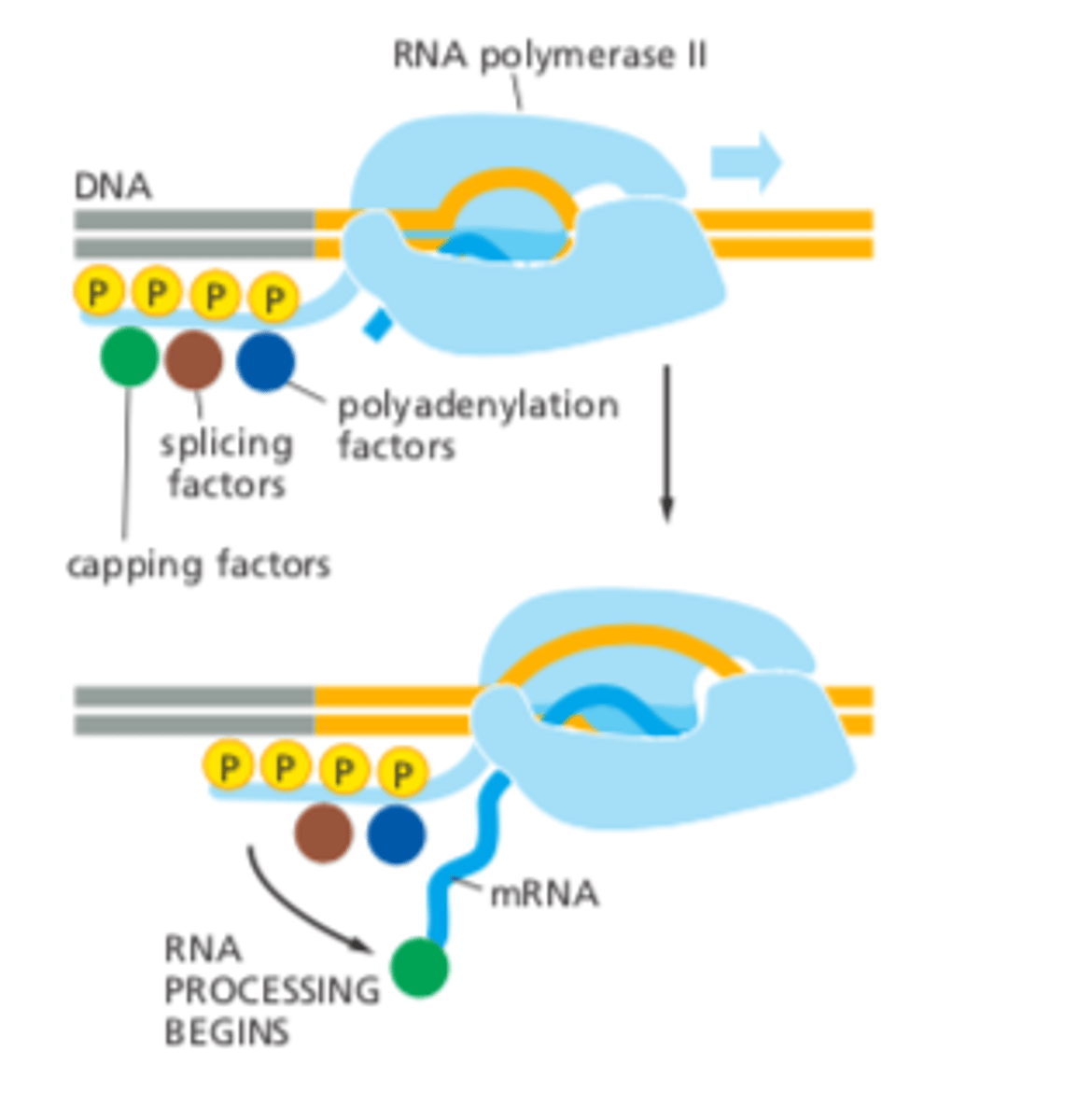
Dynamic Phosphorylation of the CTD tail during the transcription cycle
phosphorylation patterns changing to recruit specific factors during initiation, elongation, and termination
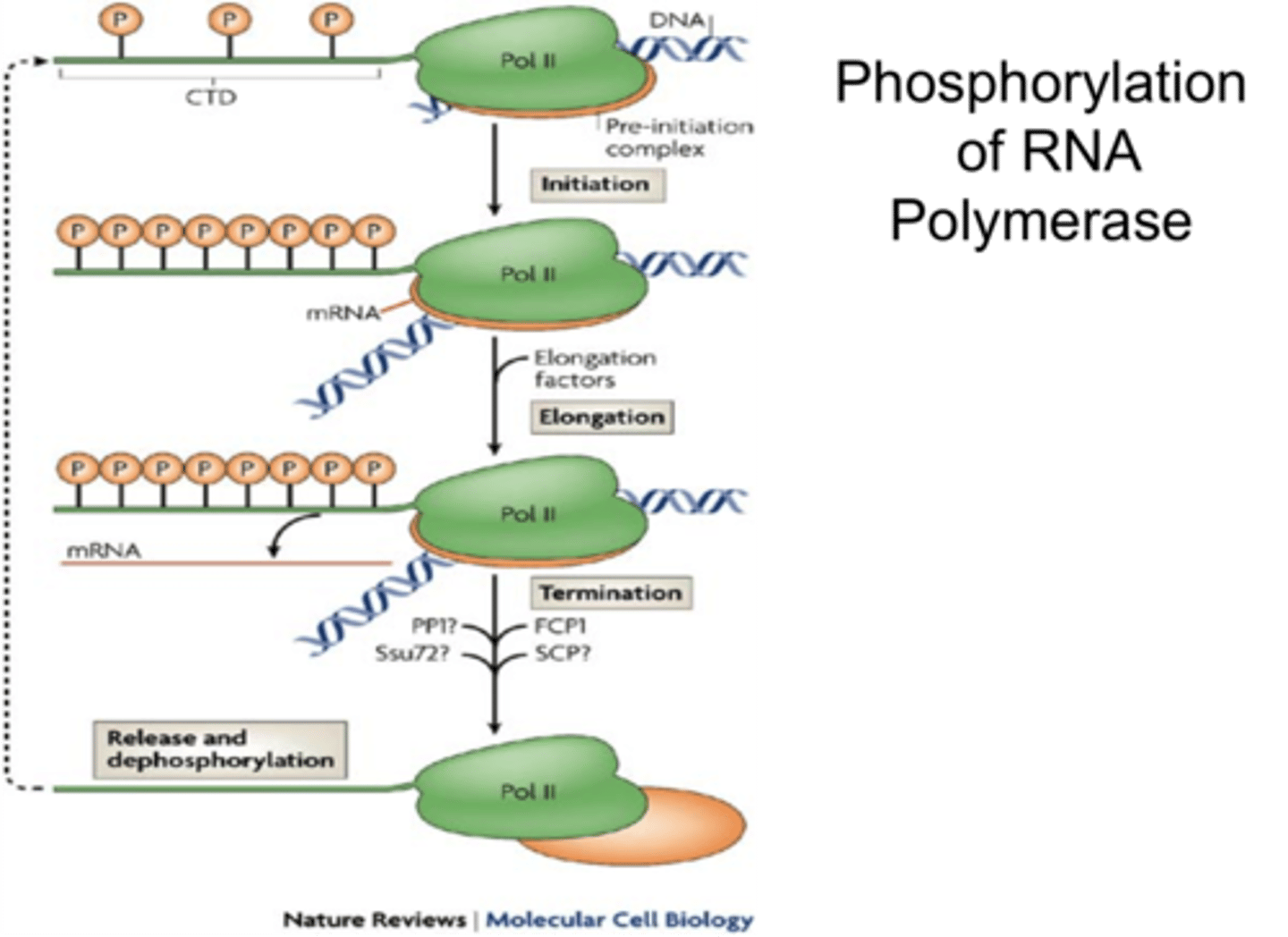
Essential molecular component for transcription:
Transcription factors (TFs)
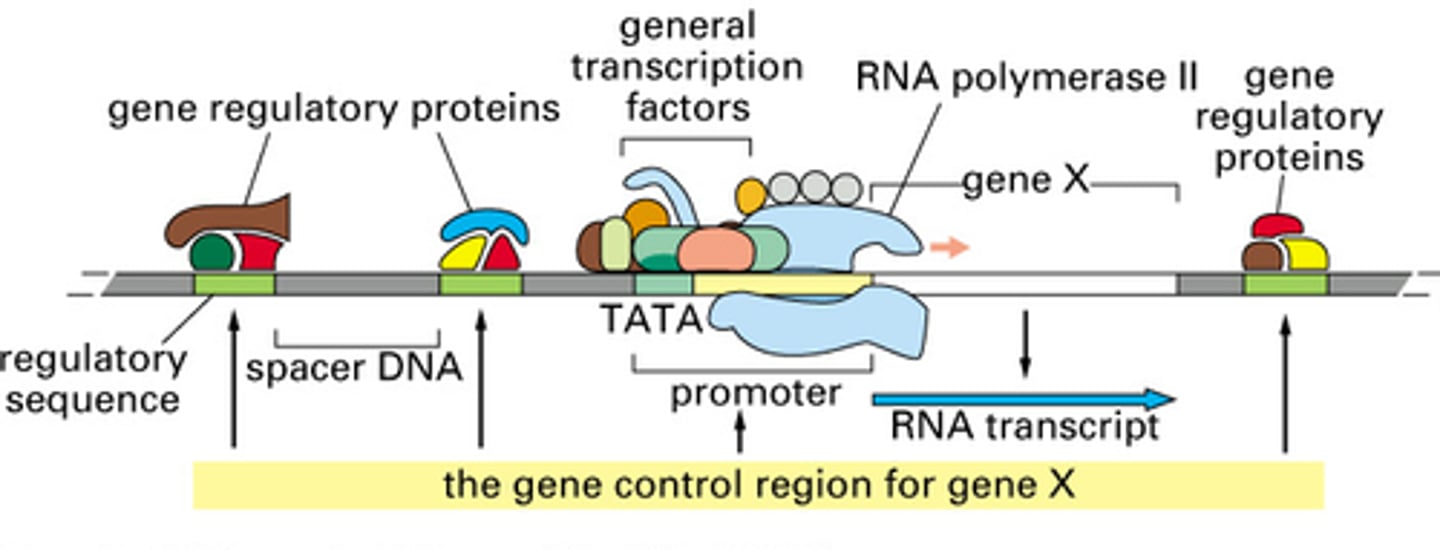
Transcription factors
proteins that regulate gene transcription
Cis transactivation
Regulatory elements / factors that regulate genes on the same chromosome(enhancers, promoters)
trans transactivation
Regulatory elements / factors that regulate genes on a different chromosome from their origin
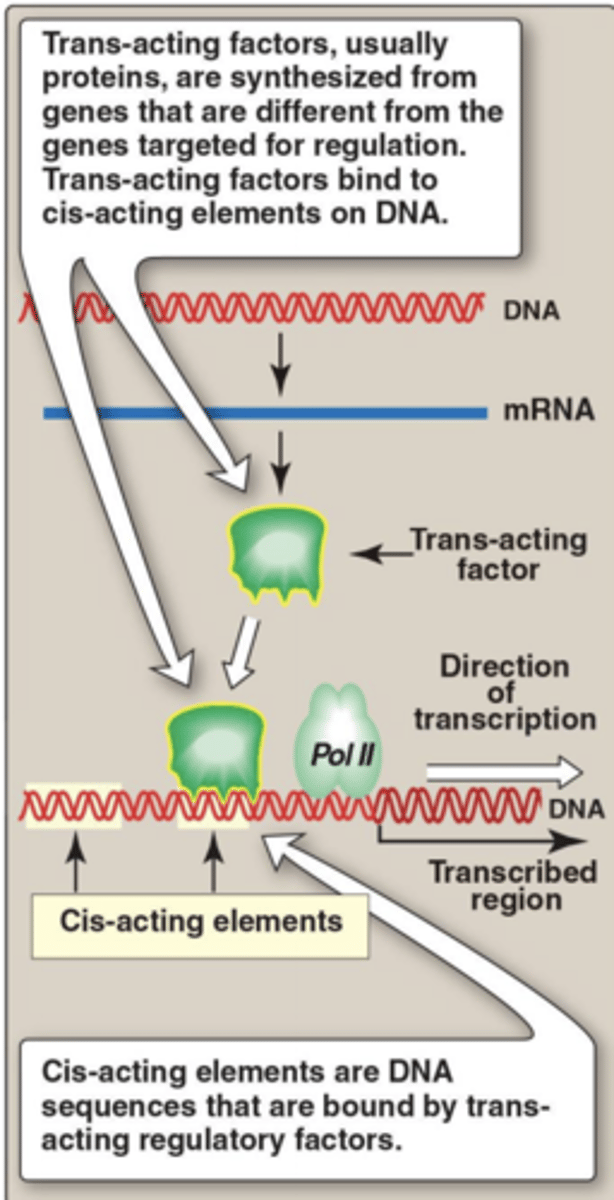
basal transcription factors/General transcription Factors (GTFs)
required to initiate transcription of a gene
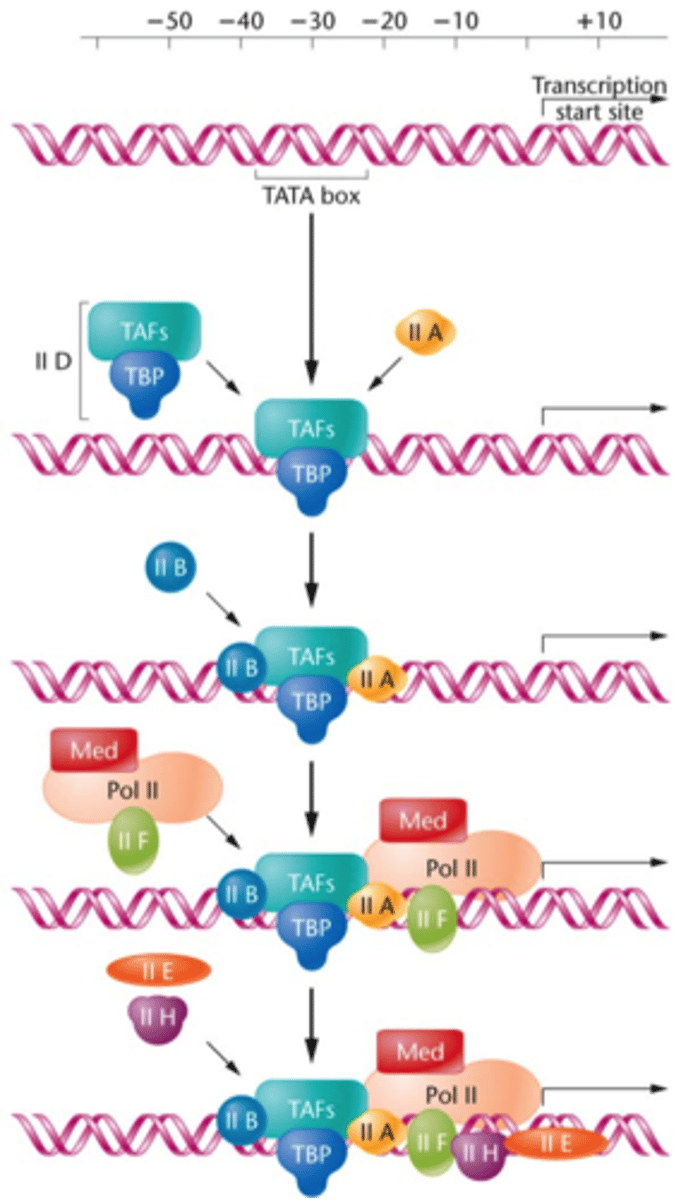
GTFs
multi-subunit protein complexes involved in core promoter recognition
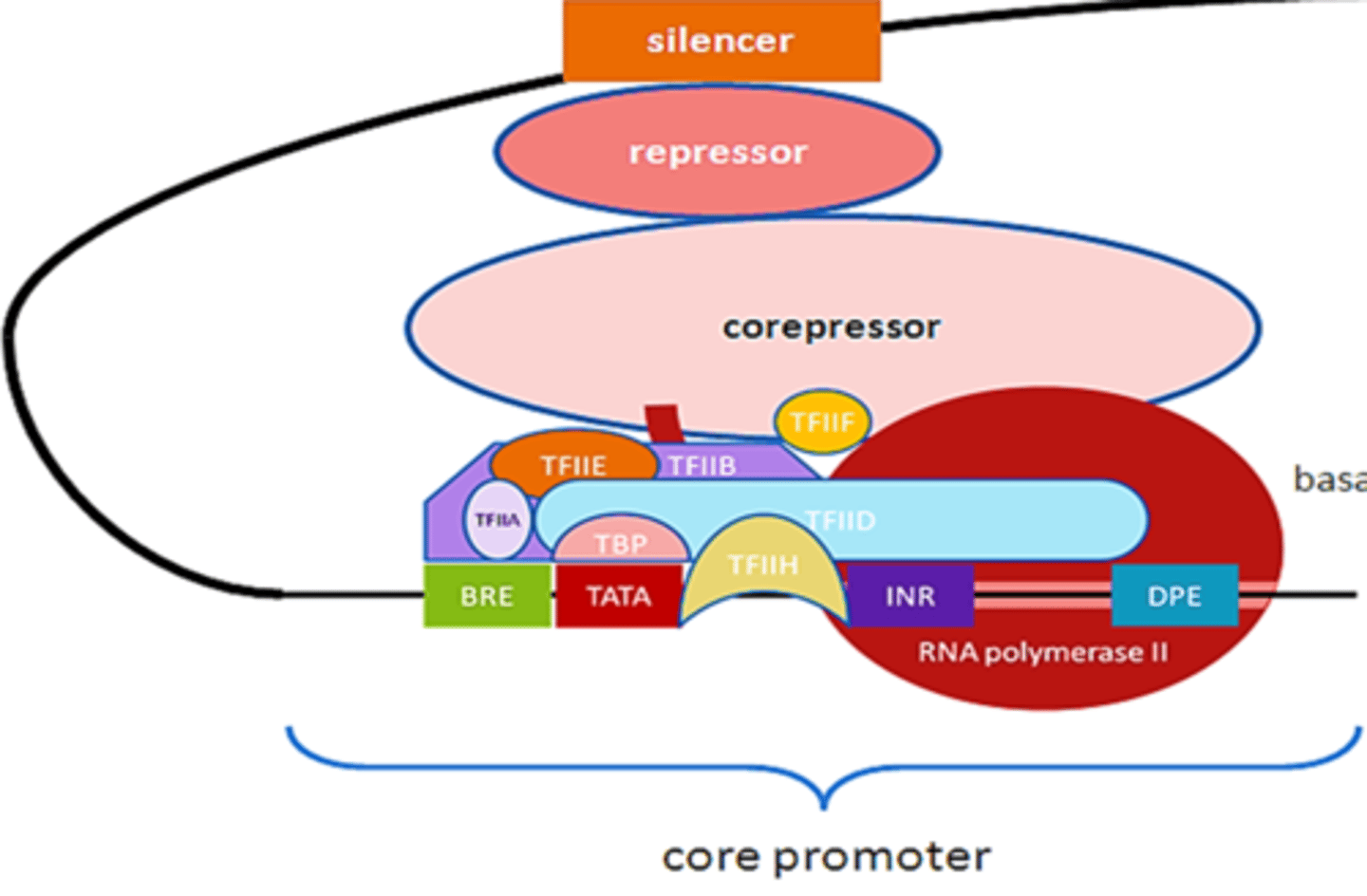
RNA polymerase II in transcription
synthesizes all eukaryotic mRNA precursors
by itself it is incapable of binding to DNA and initiating transcription at specific sites - needs GTFs
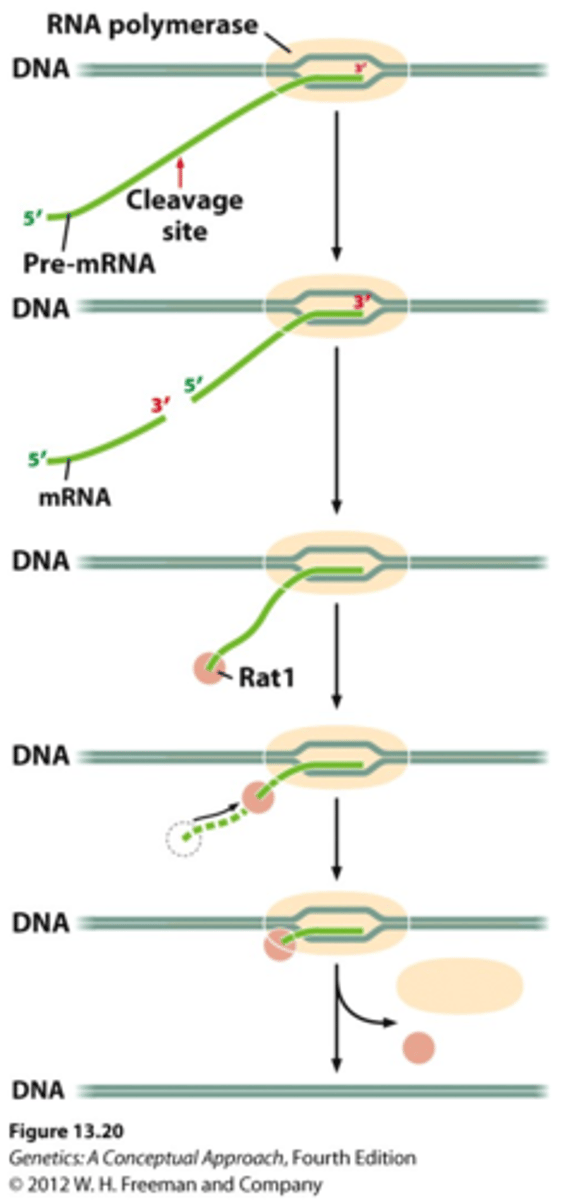
TFII
Transcription Factor for RNA Pol II
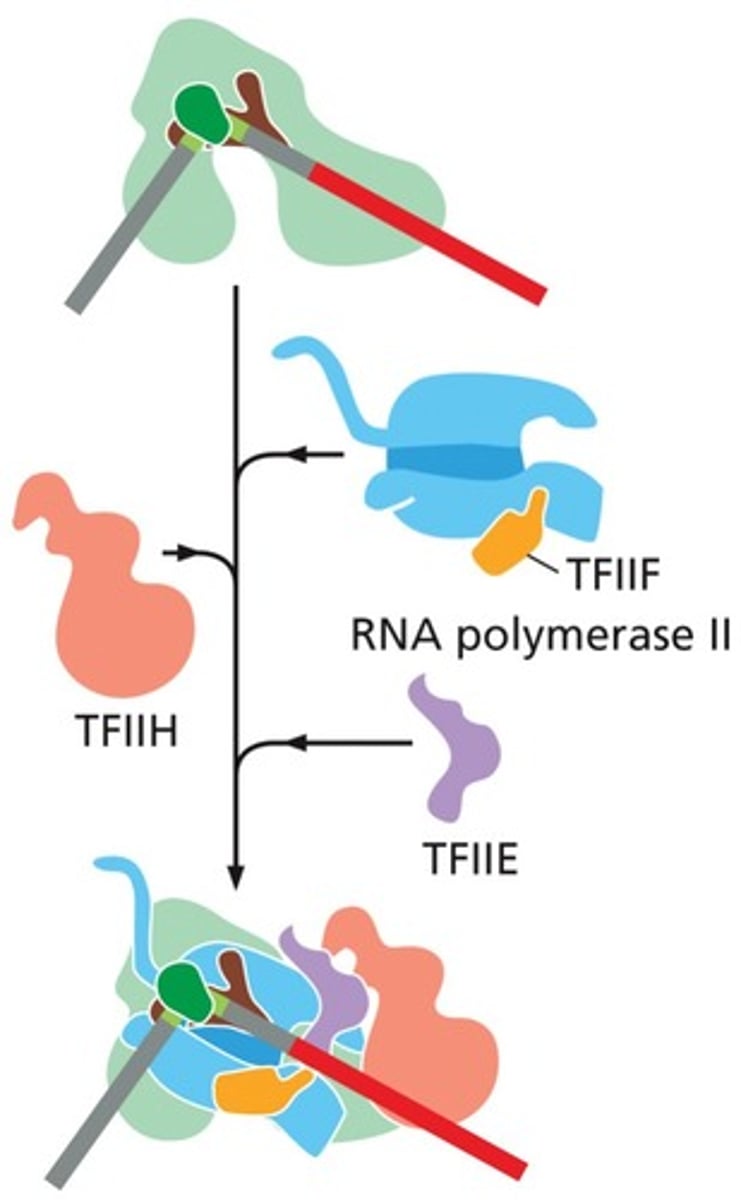
Six general transcription factors
TFIIA, TFIIB, TFIID, TFIIE,TFIIF, and TFIIH
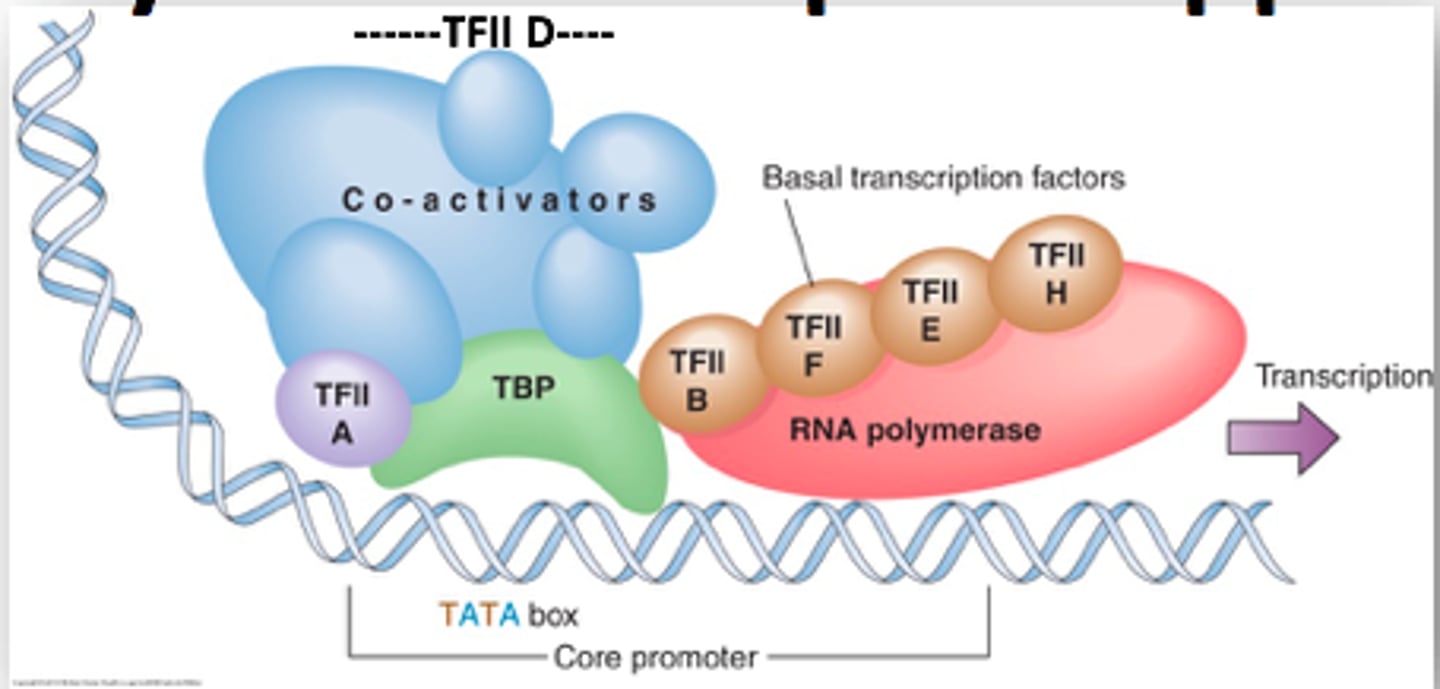
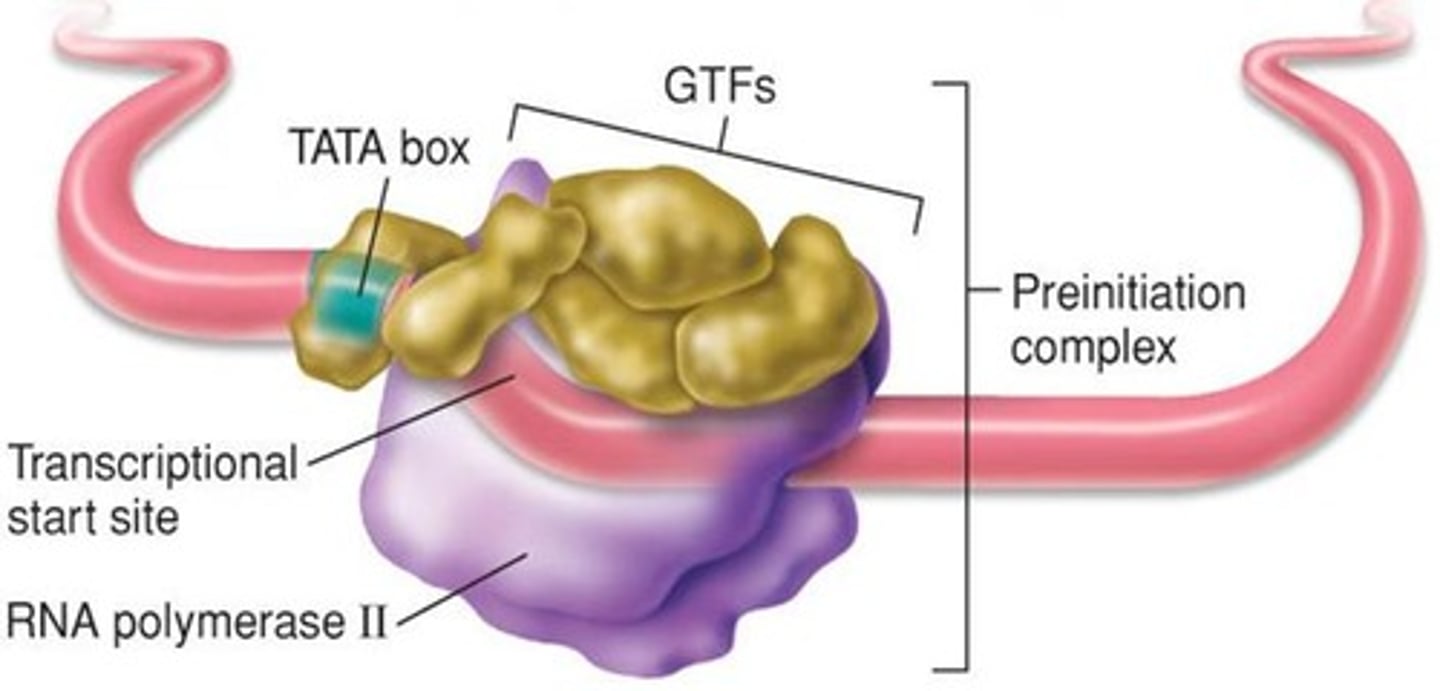
RNA Pol II + GTF
= basal transcriptional machinery or preinitiation complex
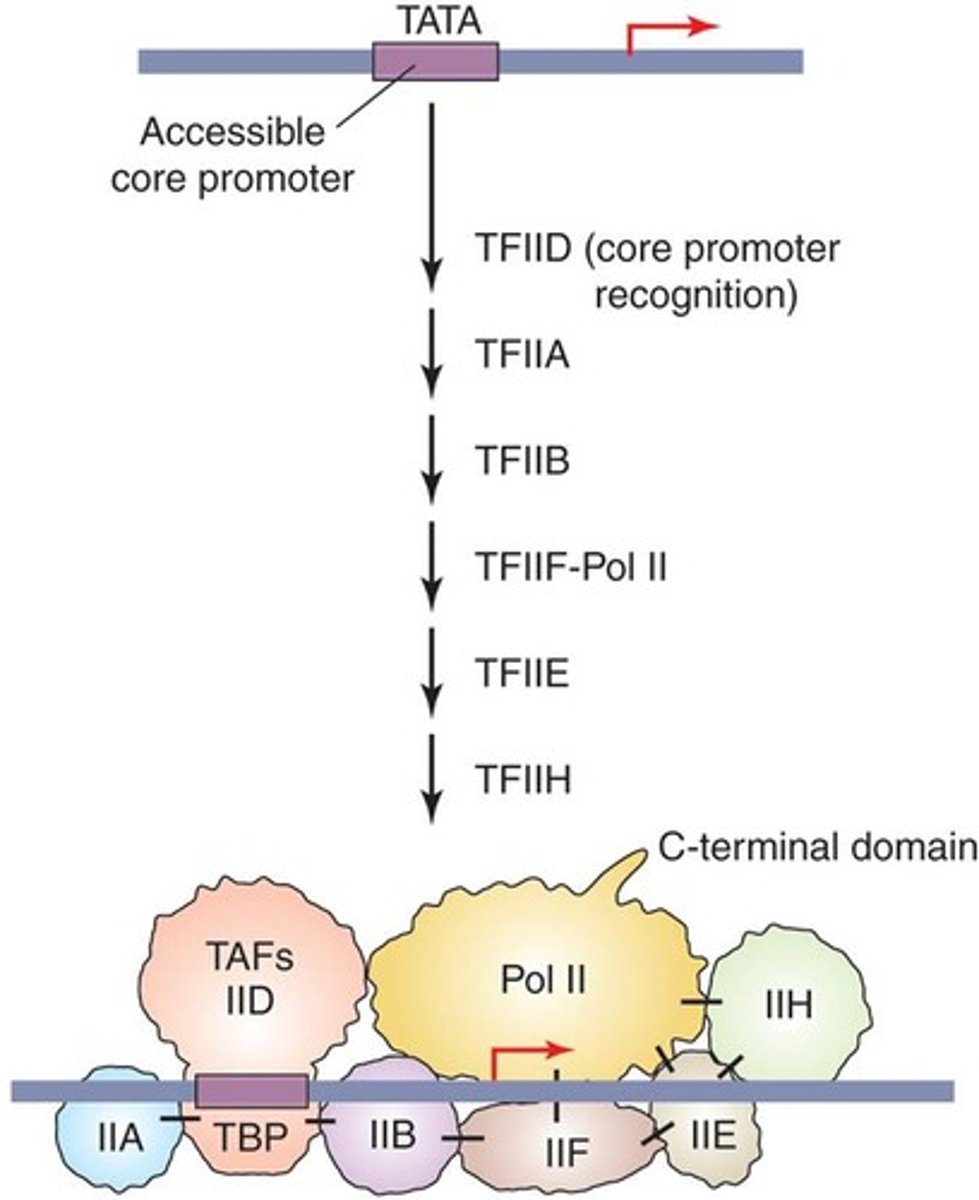
basal transcriptional machinery
assembles on the promoter.
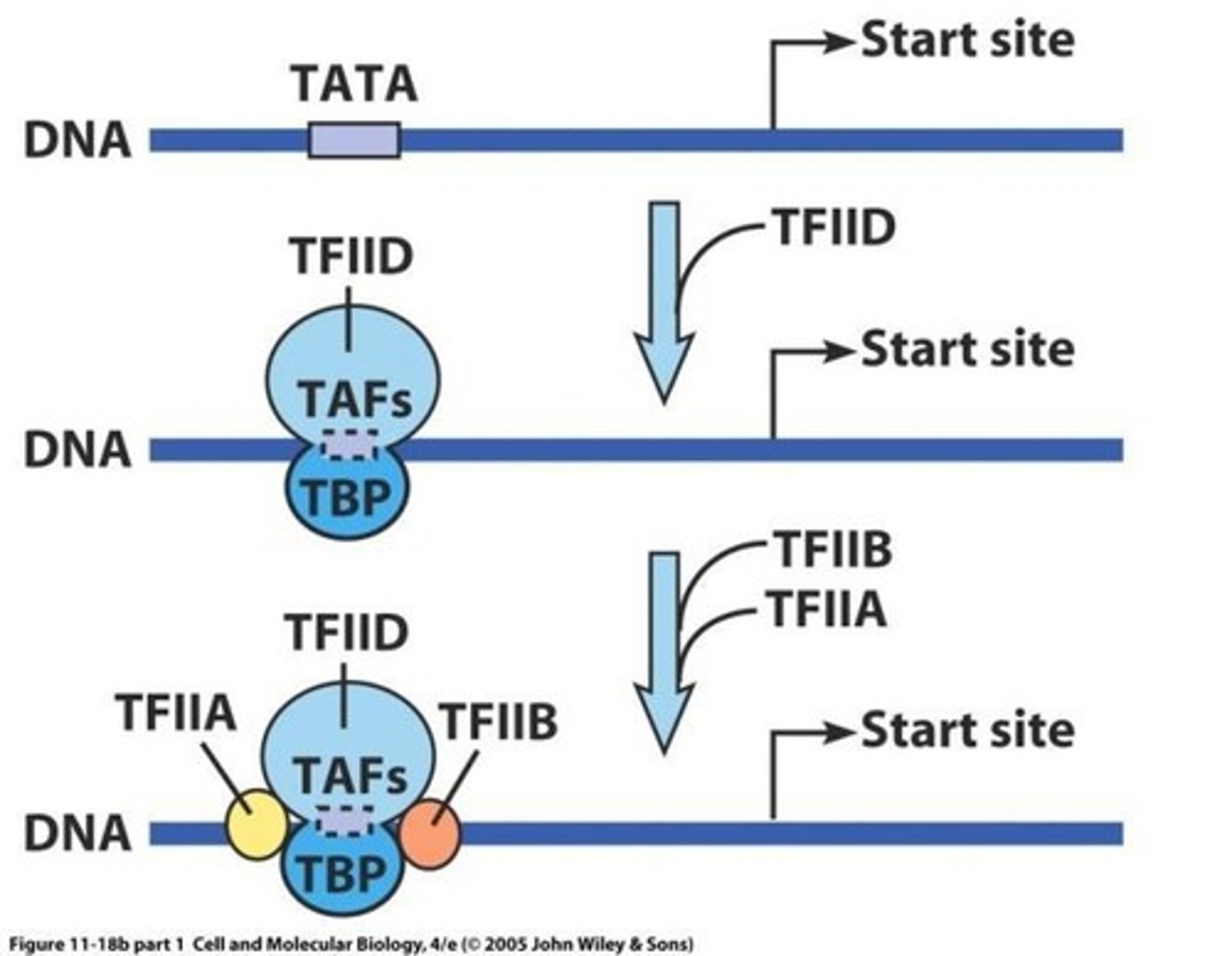
preinitiation complex
the assembly required before transcription can start
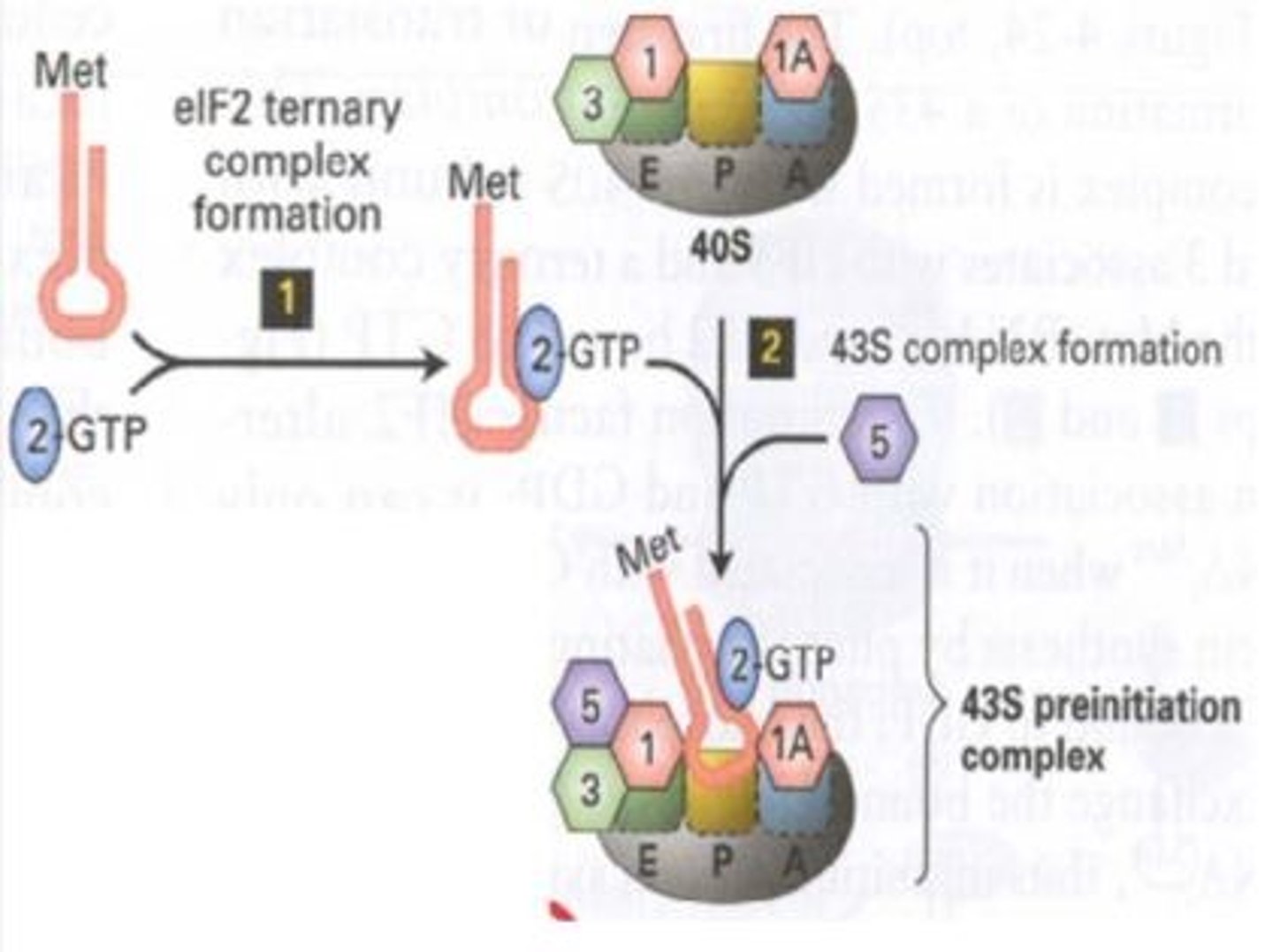
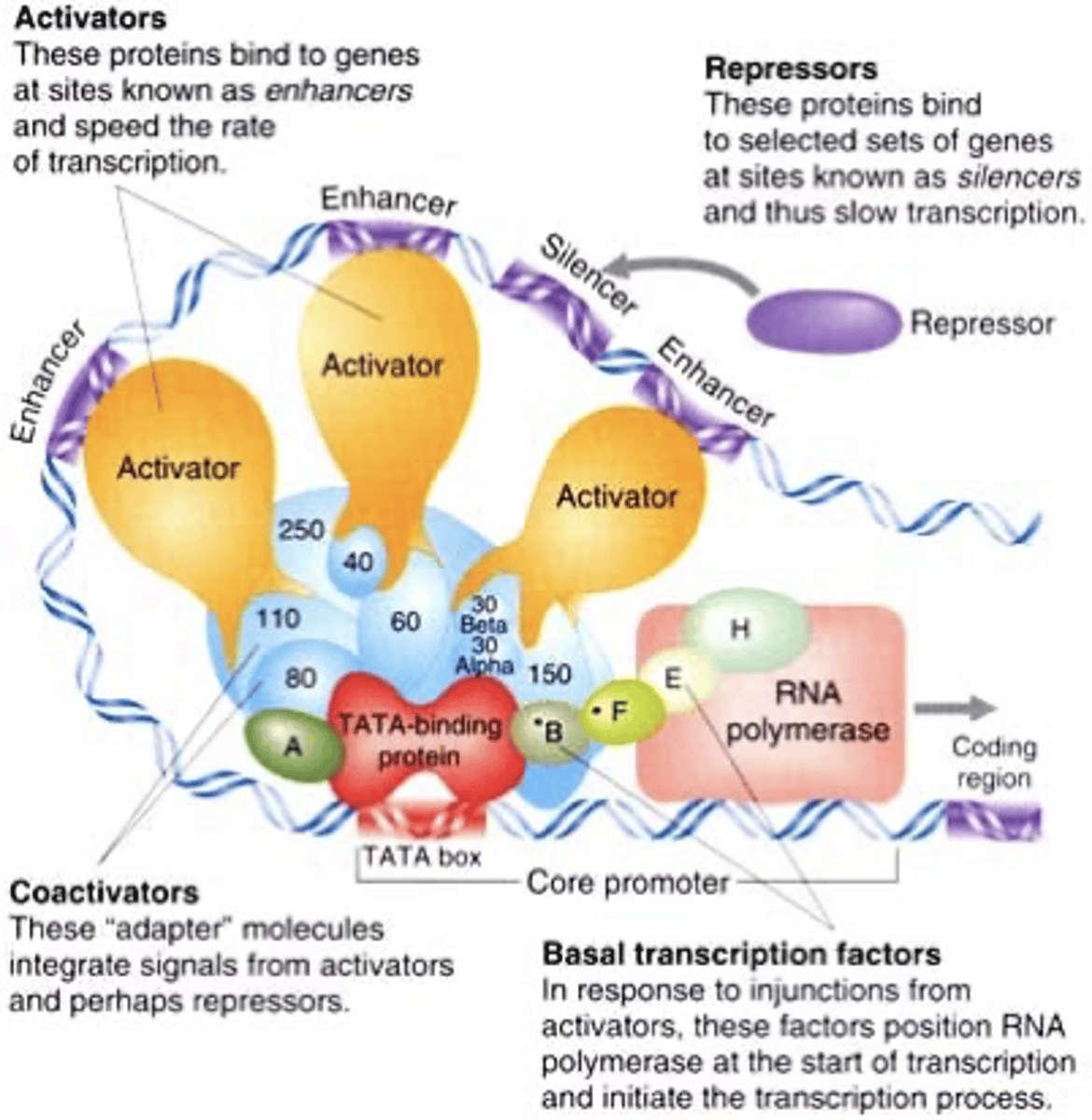
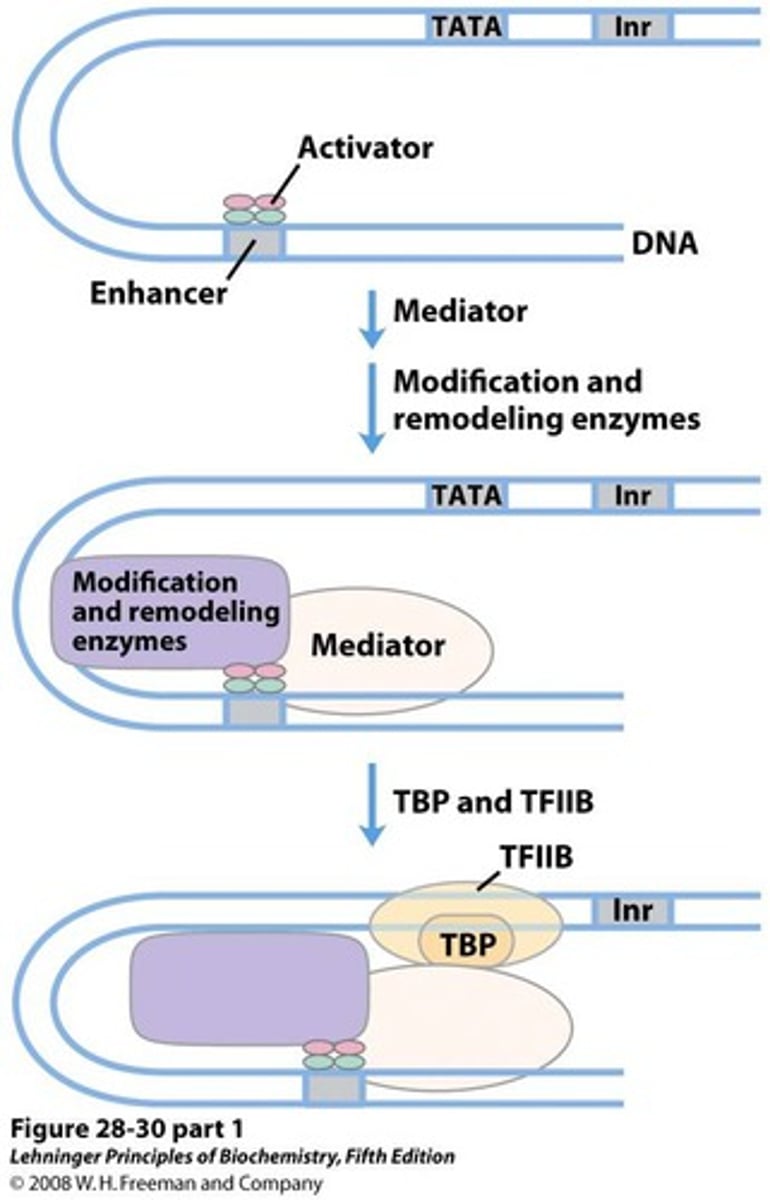
TFs (activators)
bind distal promoter sequence (enhancers) and increase transcription efficiency from basal levels
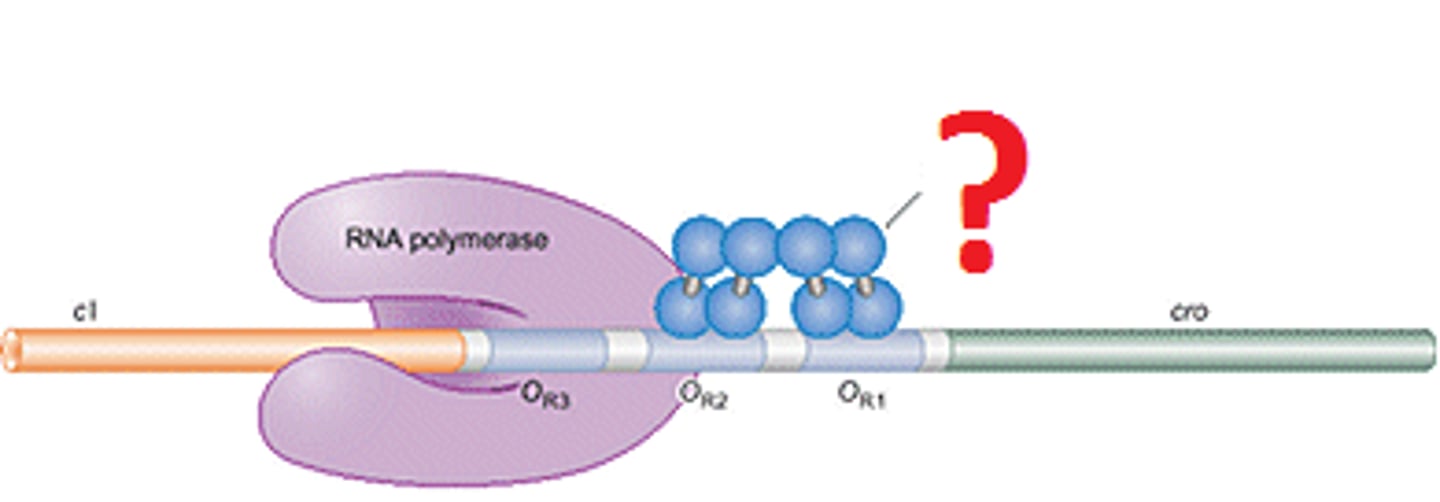
8-10 nucleotides of the newly synthesized RNA strand remain base-paired with the DNA template strand at any given time
How many nucleotides are synthesized when RNA polymerase II carries out transcription?
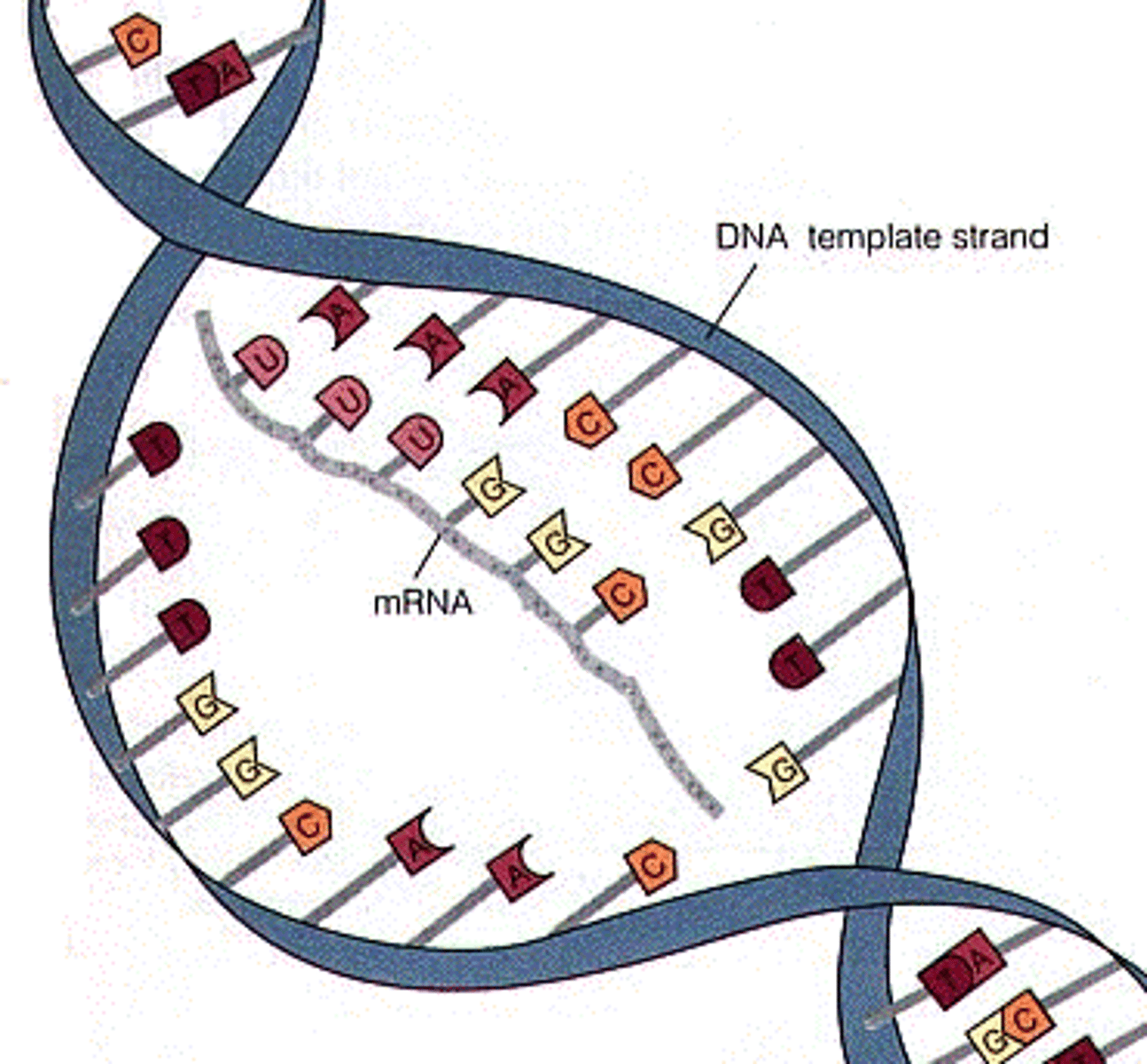
Why are these short strands Important?
This short RNA-DNA hybrid within the transcription bubble helps stabilize the RNA and prevents it from falling off prematurely
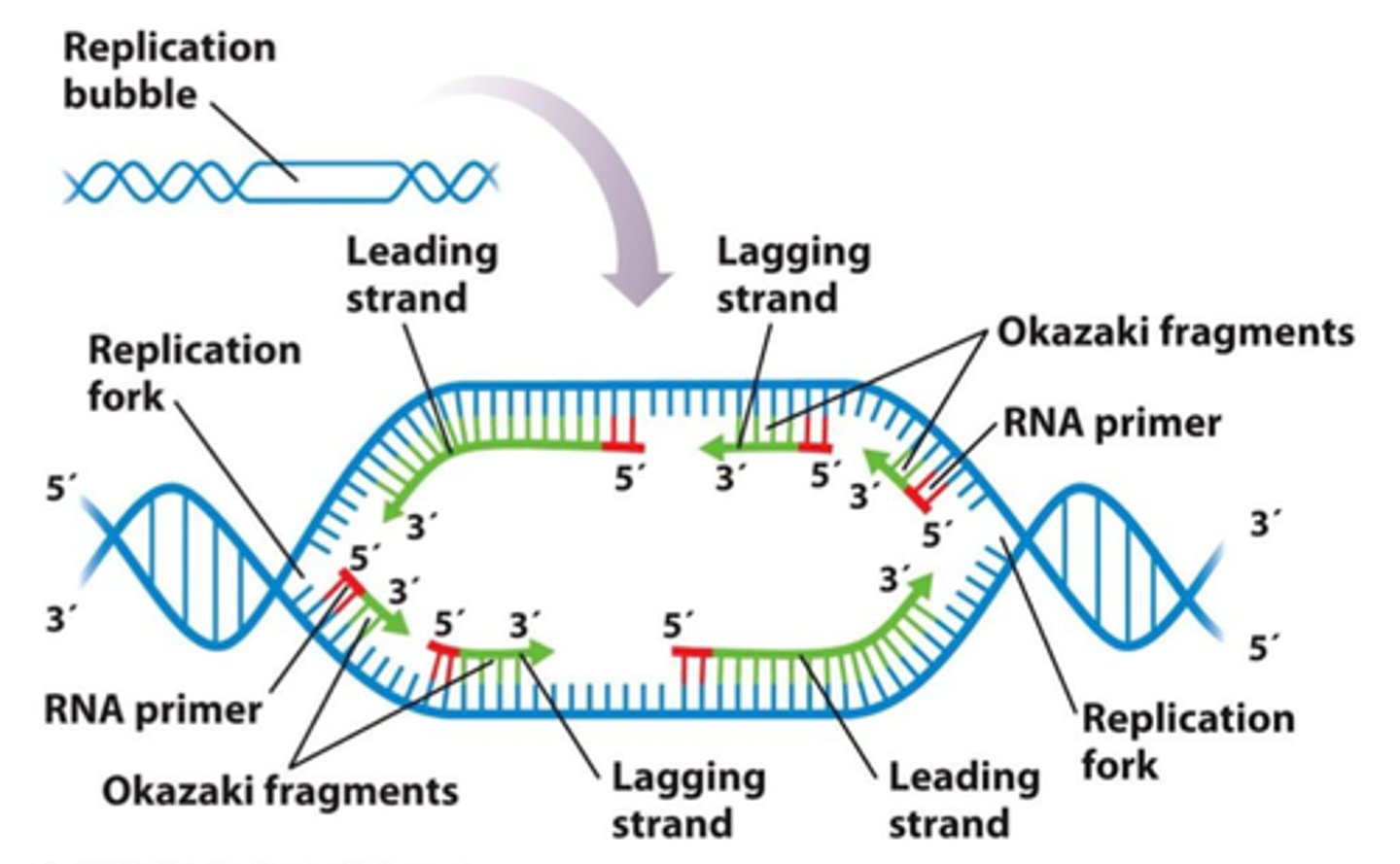
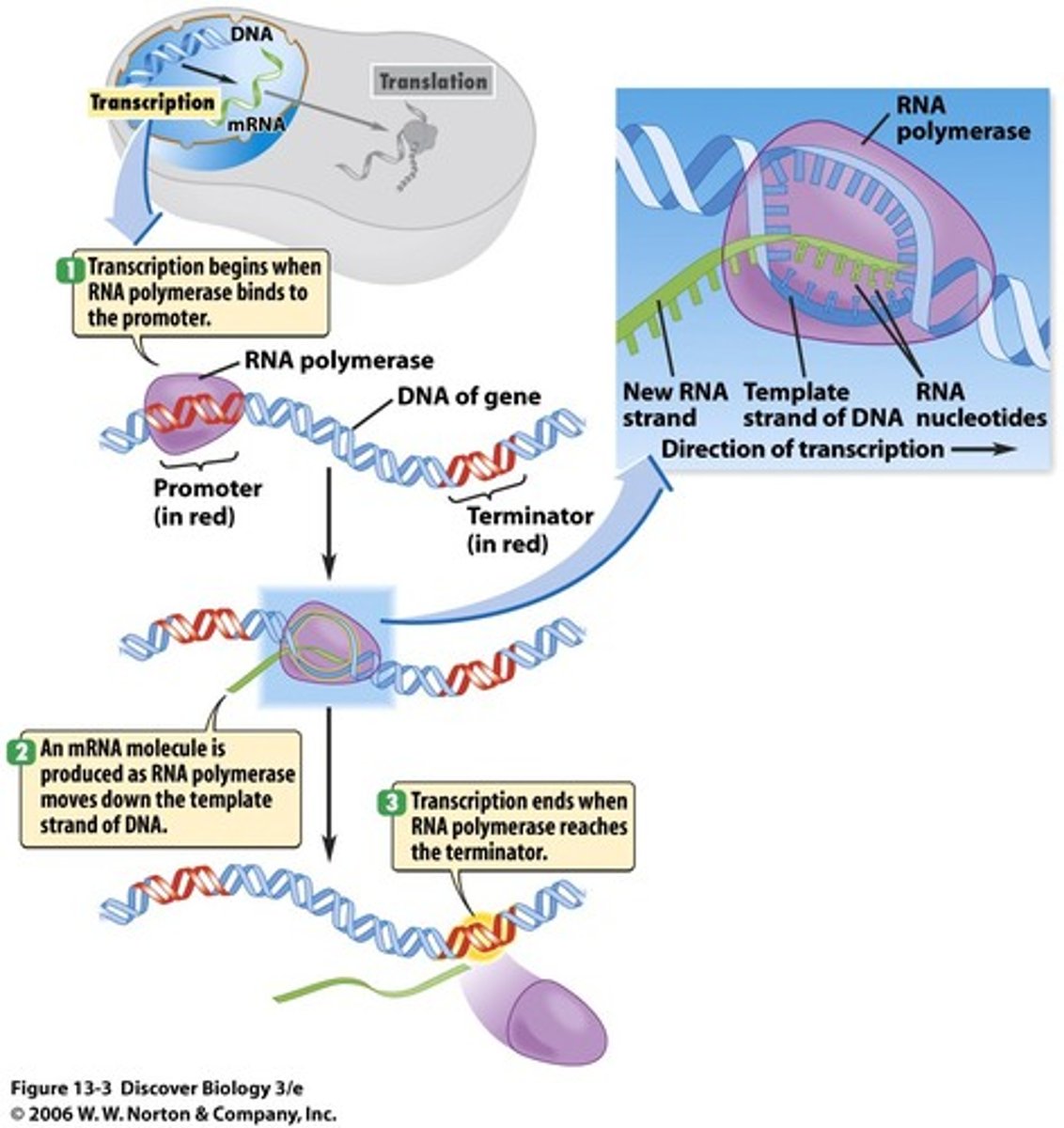
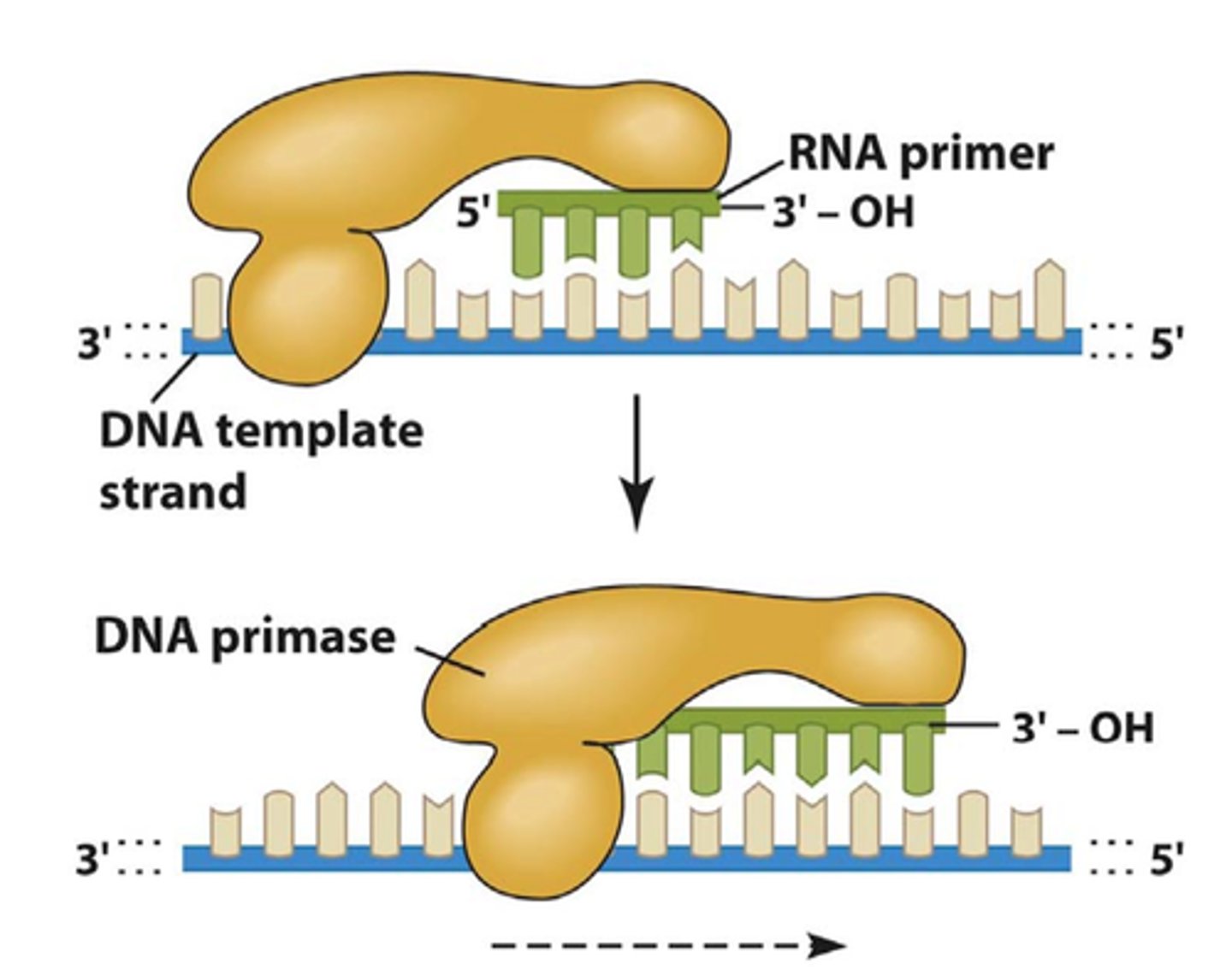
RNA Polymerases like DNA polymerases can only synthesize
5' to 3' direction
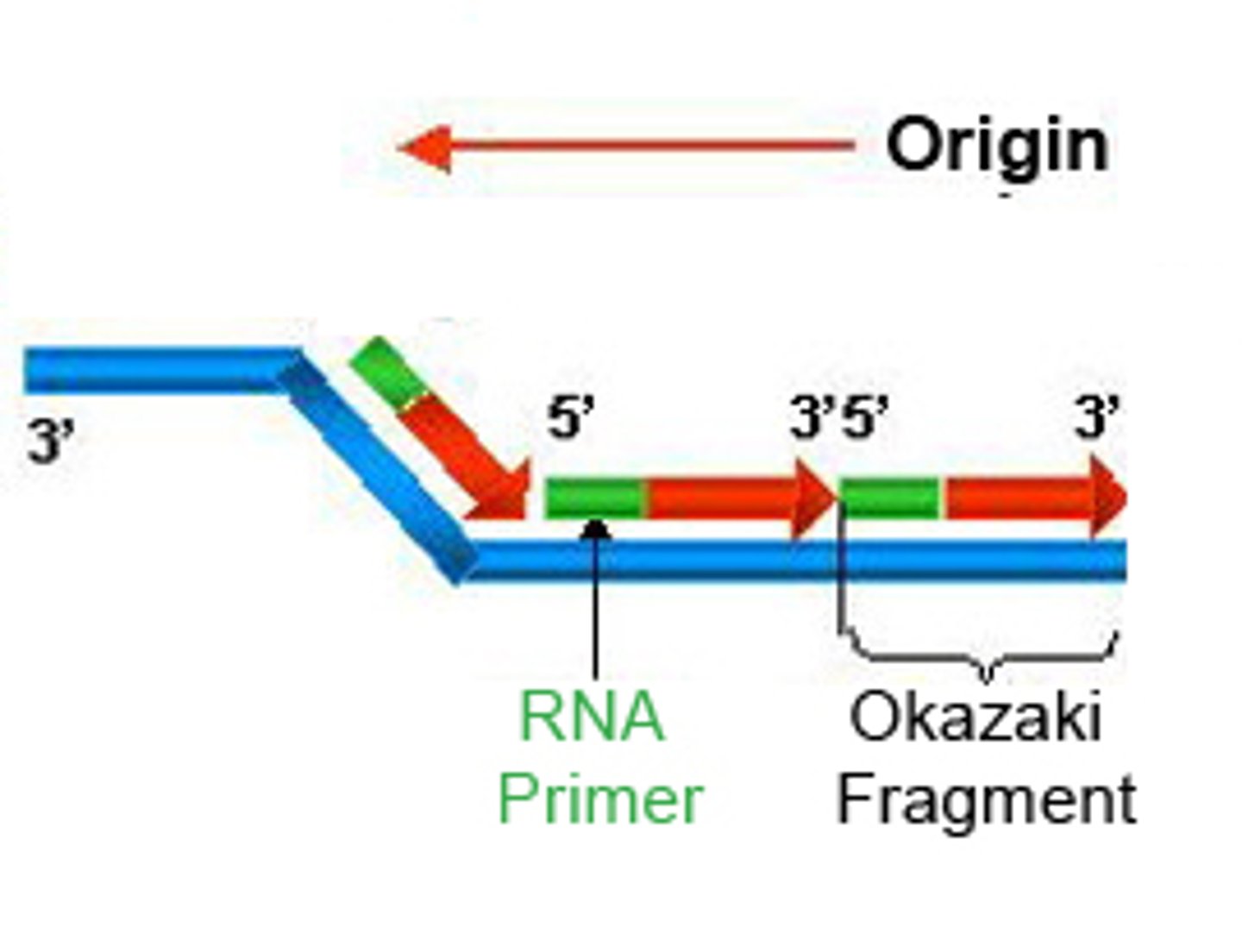
Transcription of mRNA by RNA Polymerase II occurs in 4 steps
Pre-initiation
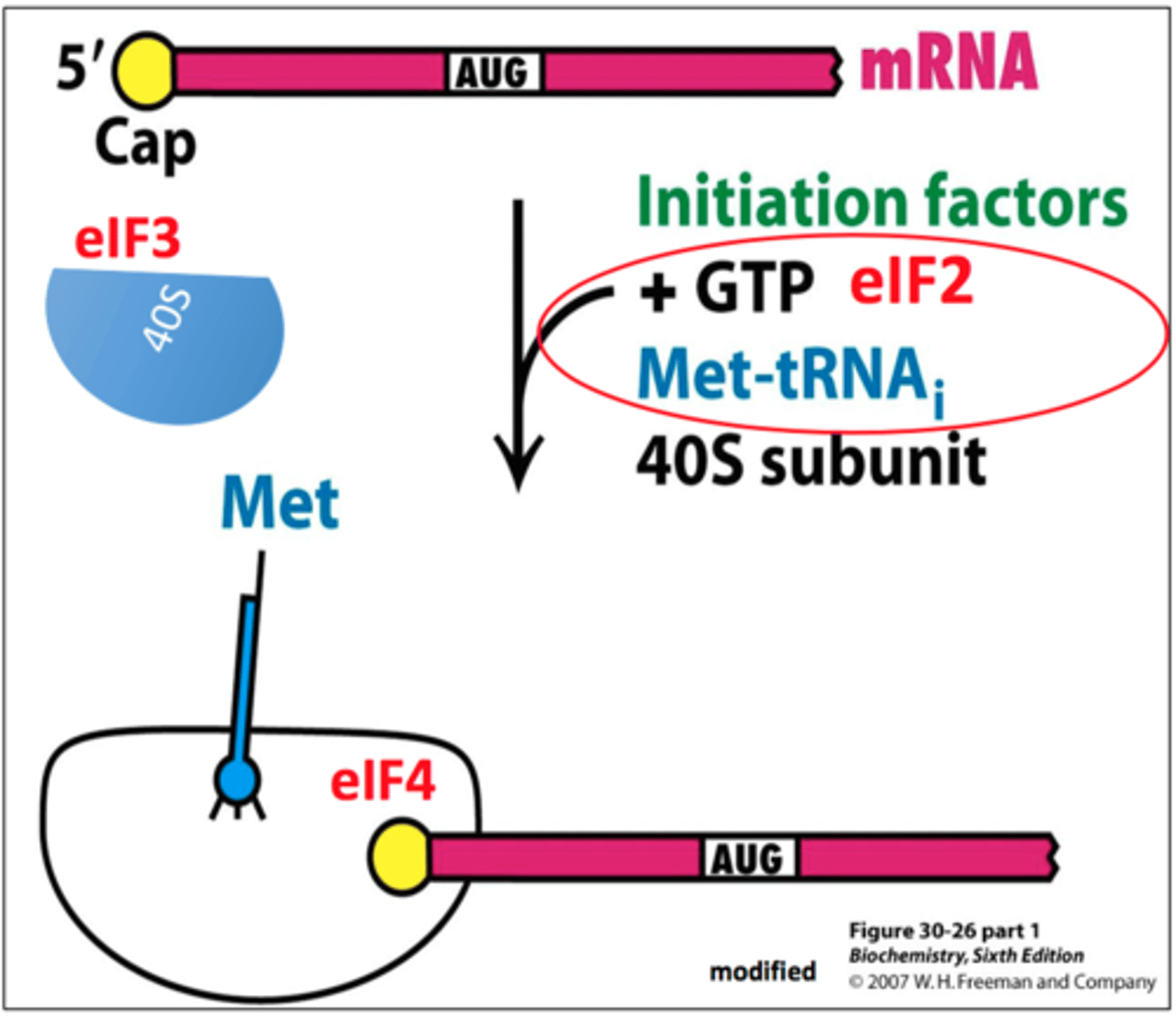
Initiation
Elongation
Termination
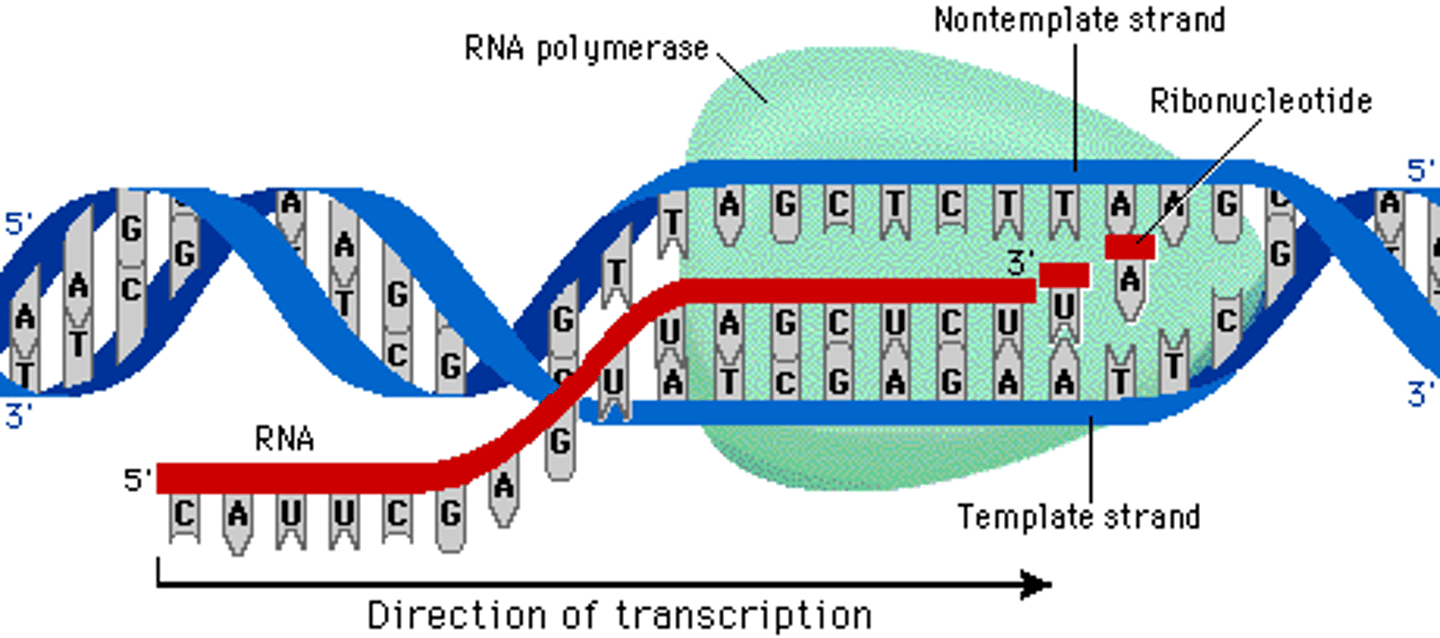
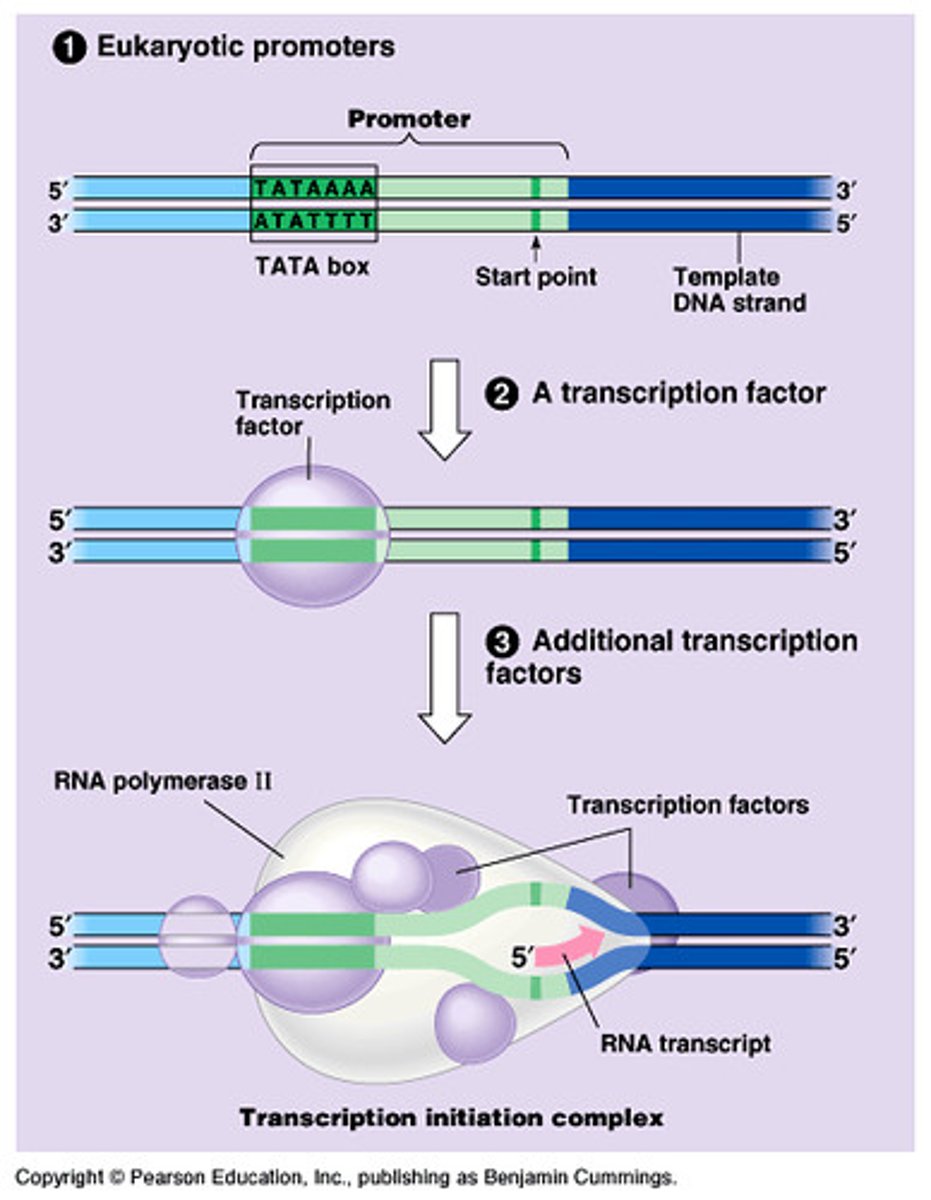
pre-initiation complex (PIC)
(general transcription factors and RNA Pol II) at the core promoter of the gene
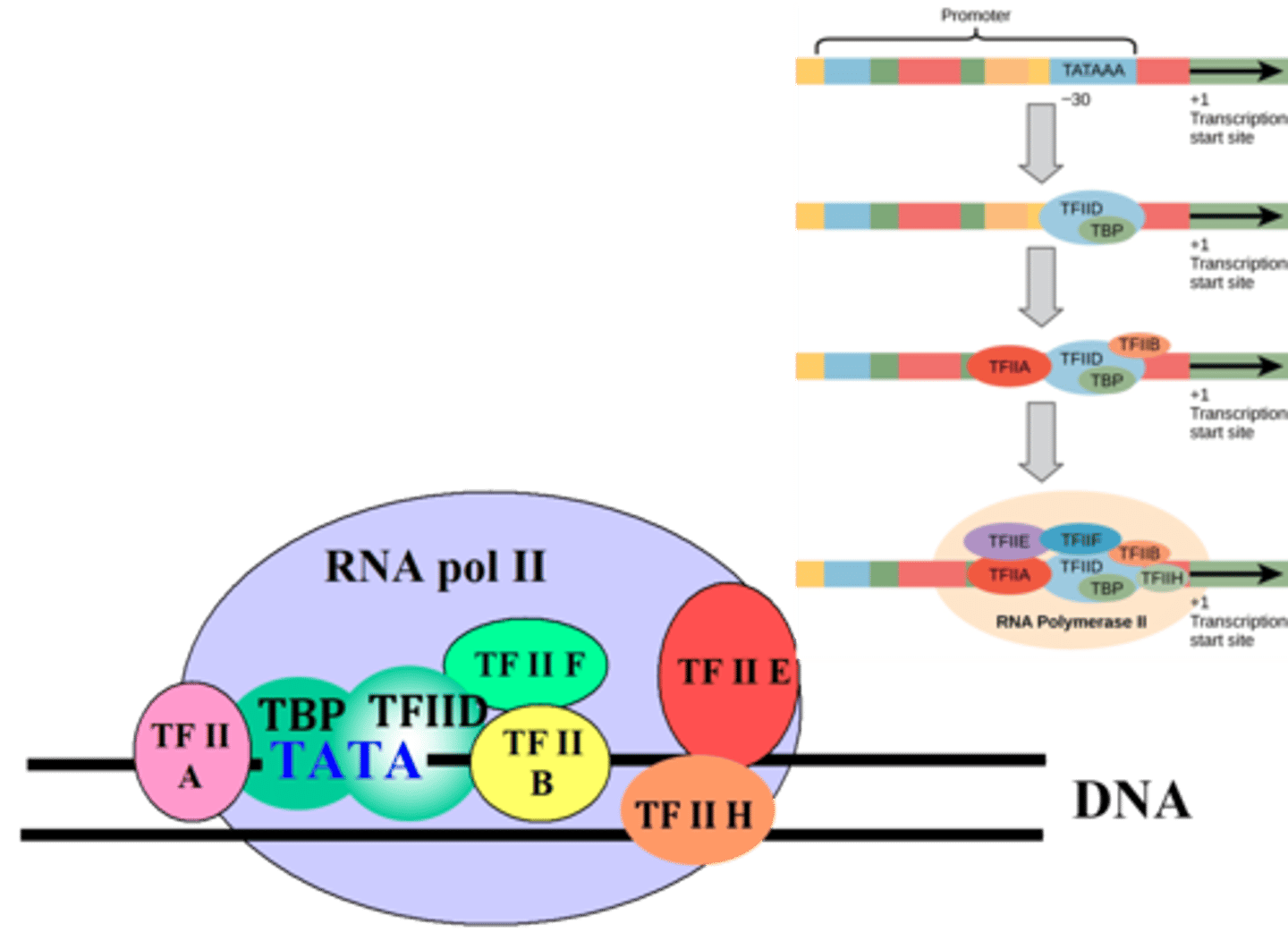
Formation of PIC
sequential assembly of the six general transcription factors (GTFs)
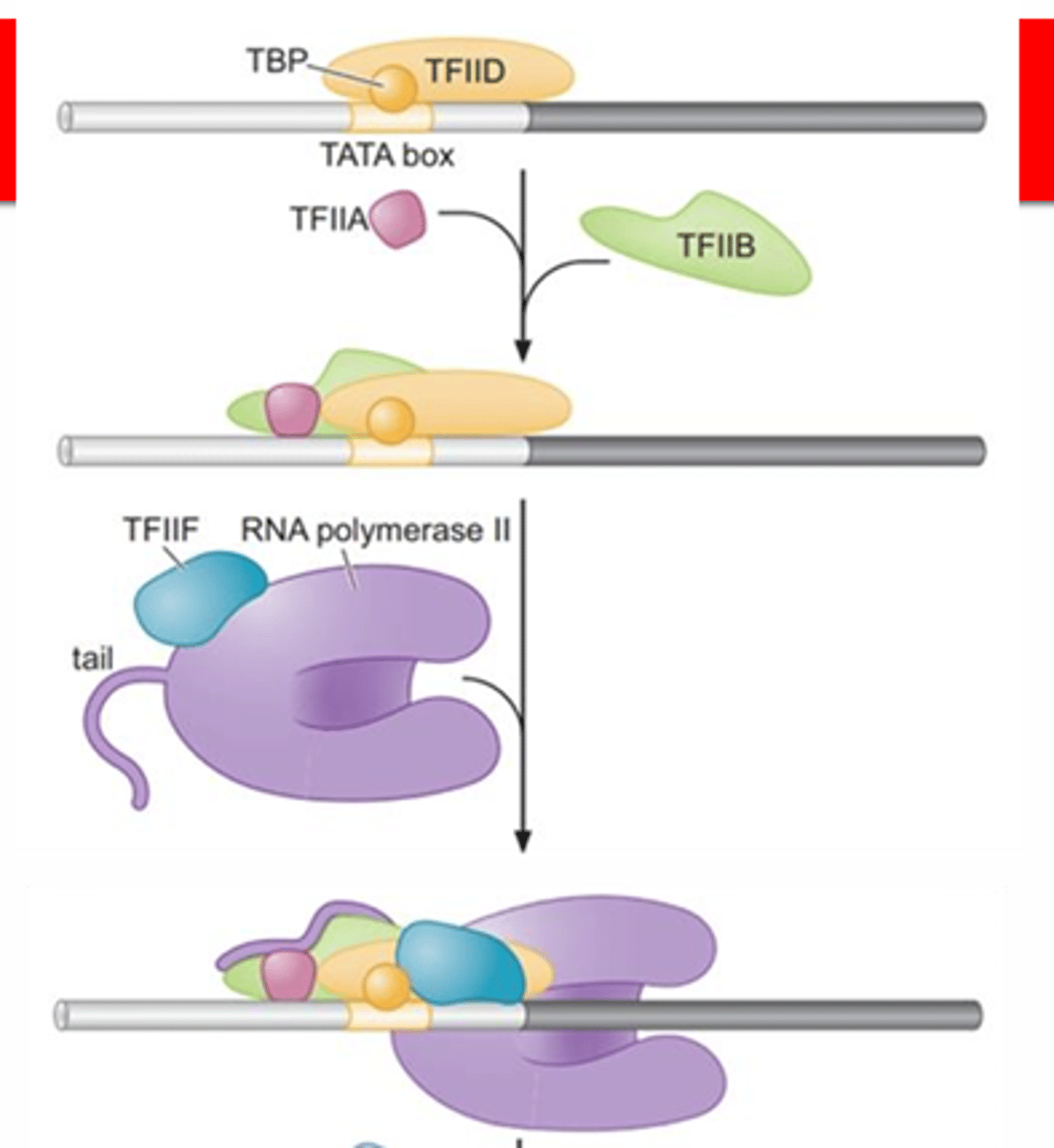
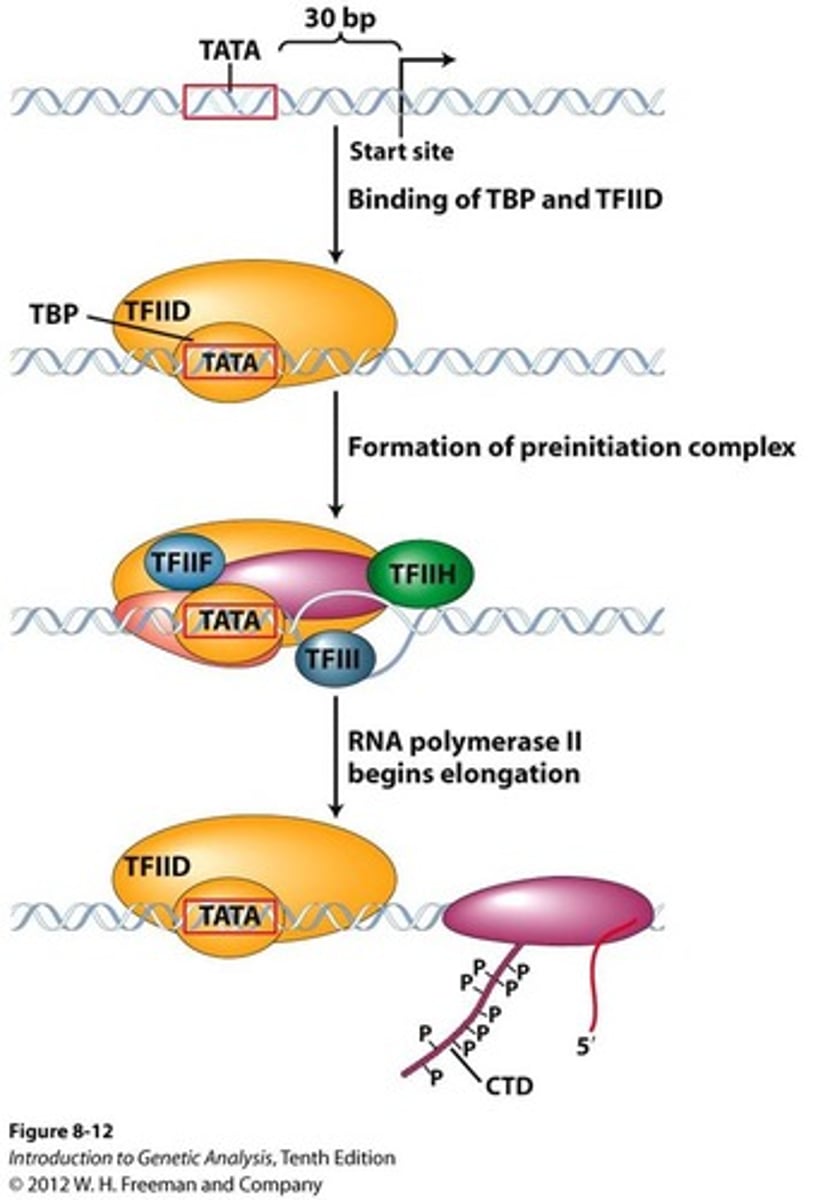
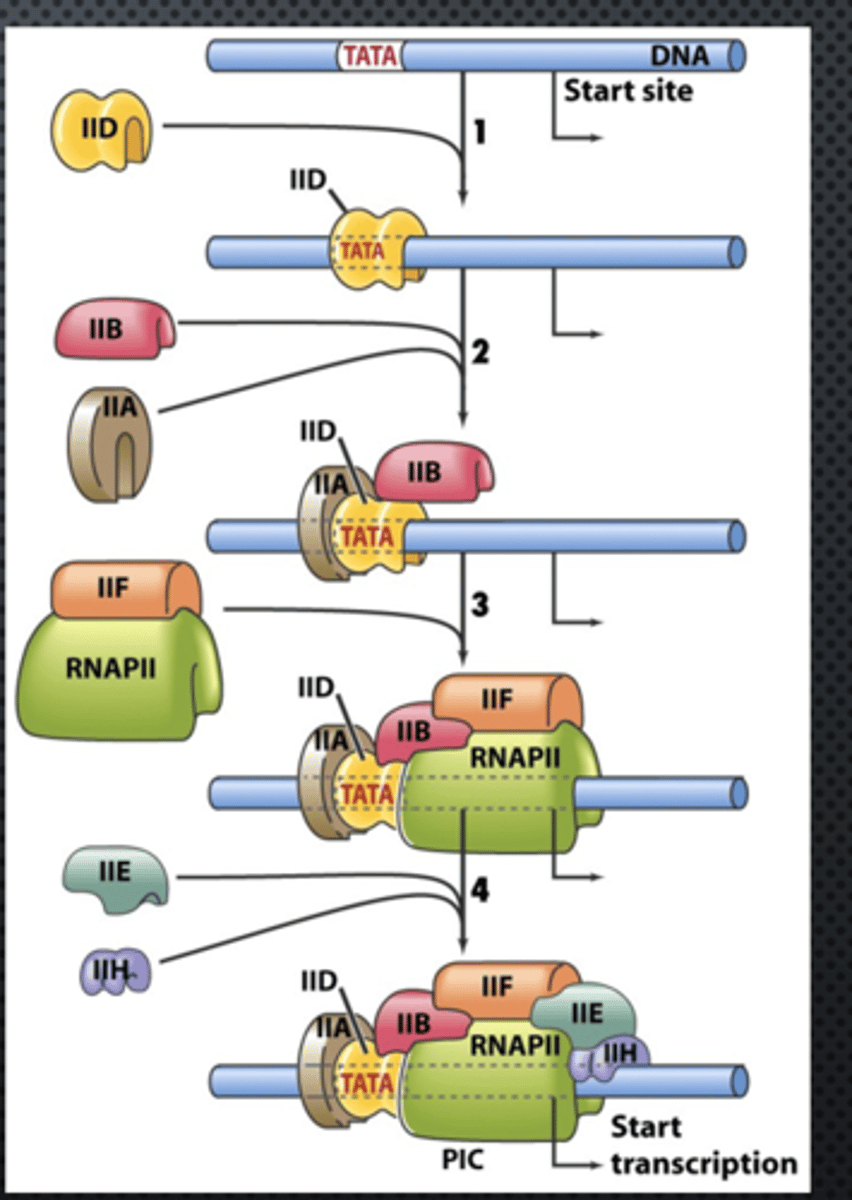
STEP 1 (Pre-initiation)
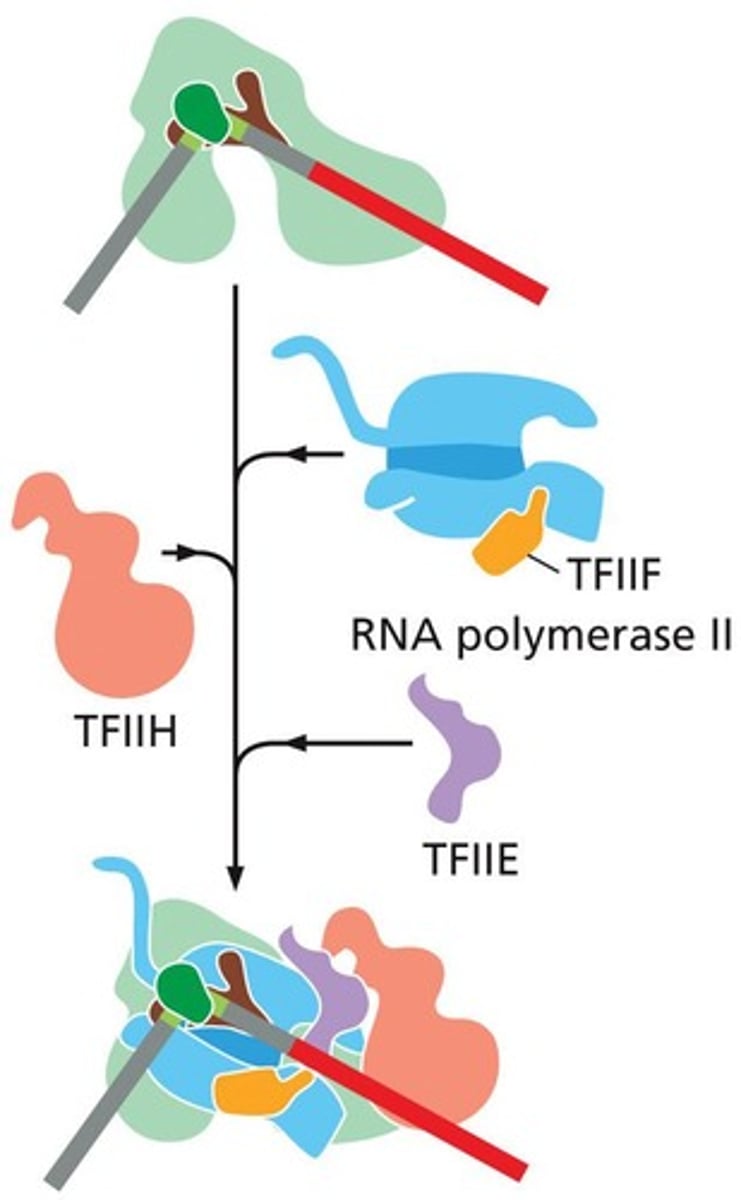
sequential assembly of the GTFs and RNA Pol II (Basal TF machinery) to the core promoter of a eukaryotic gene to form the PIC
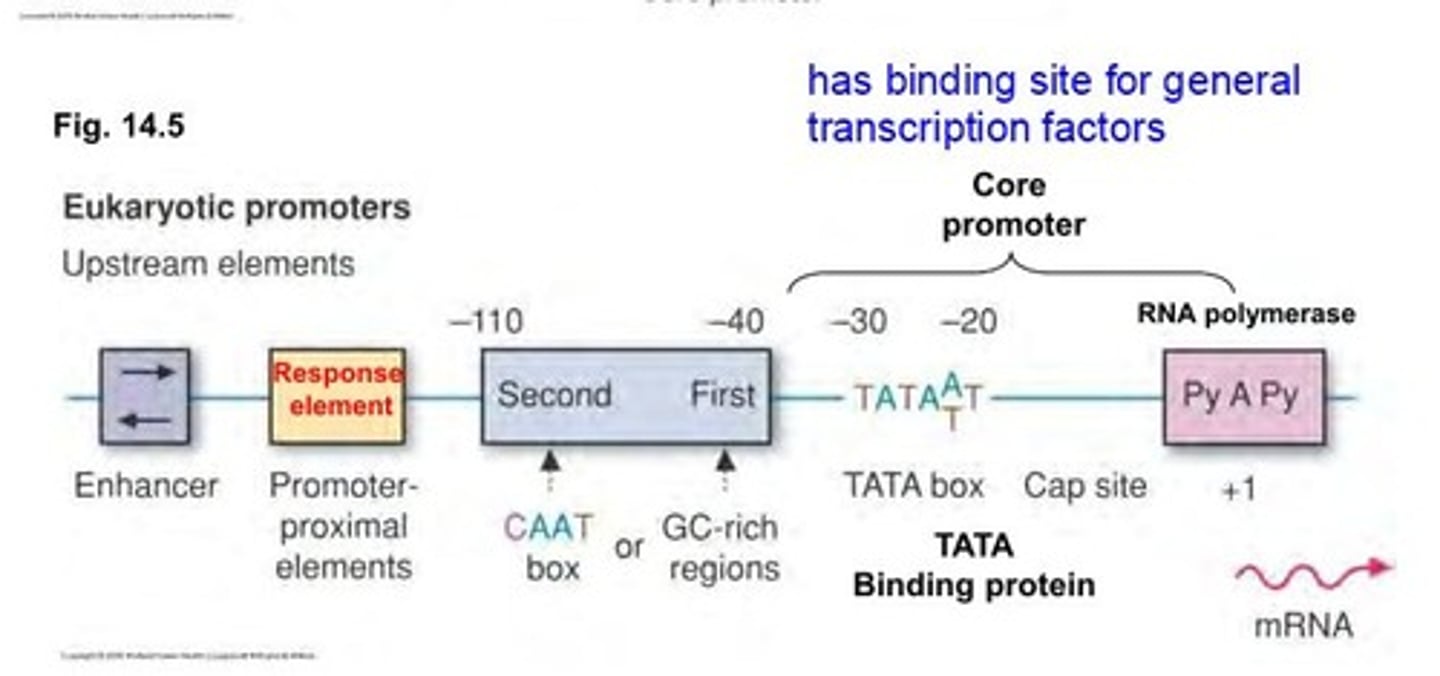
core promoter region
TATA box (commonly located -25 to -35 location)
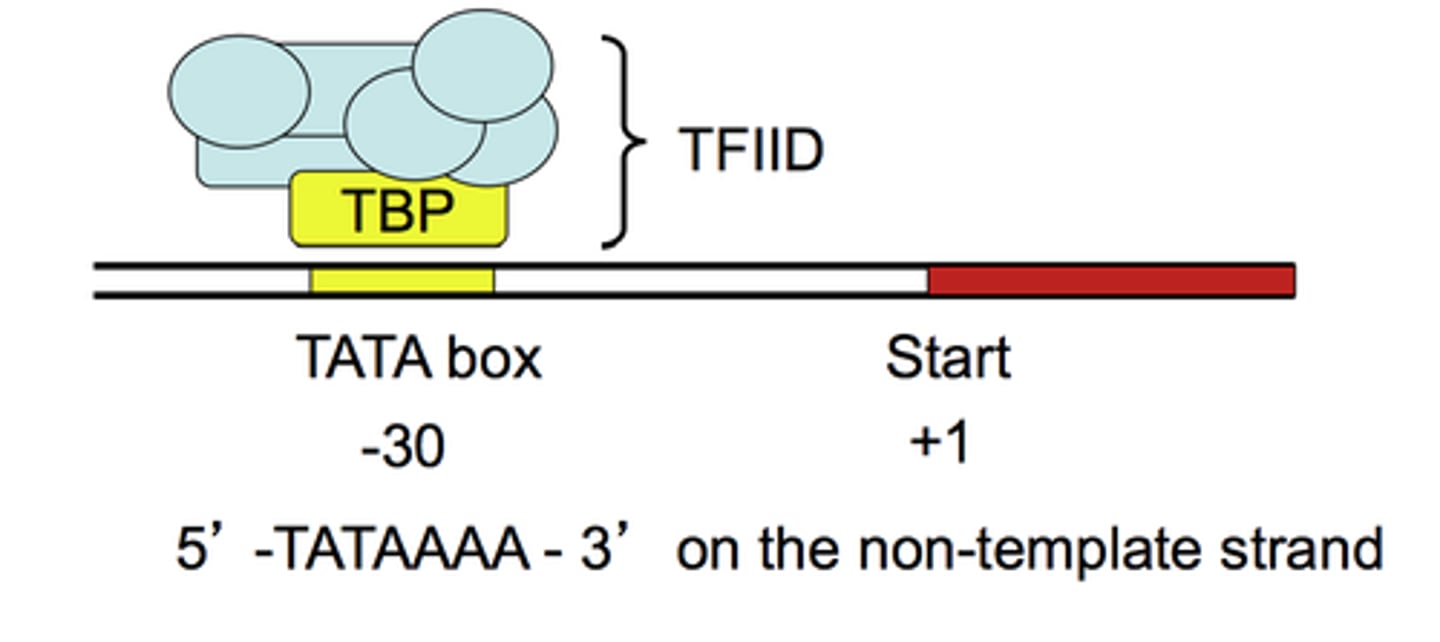
TFIID
recognize the core promoter and initiate assembly of the transcription pre-initiation complex (PIC)
TFIIE
After RNA polymerase II and TFIIF bind to the promoter, TFIIE binds to the complex and recruits TFIIH
Regulates TFIIH activity
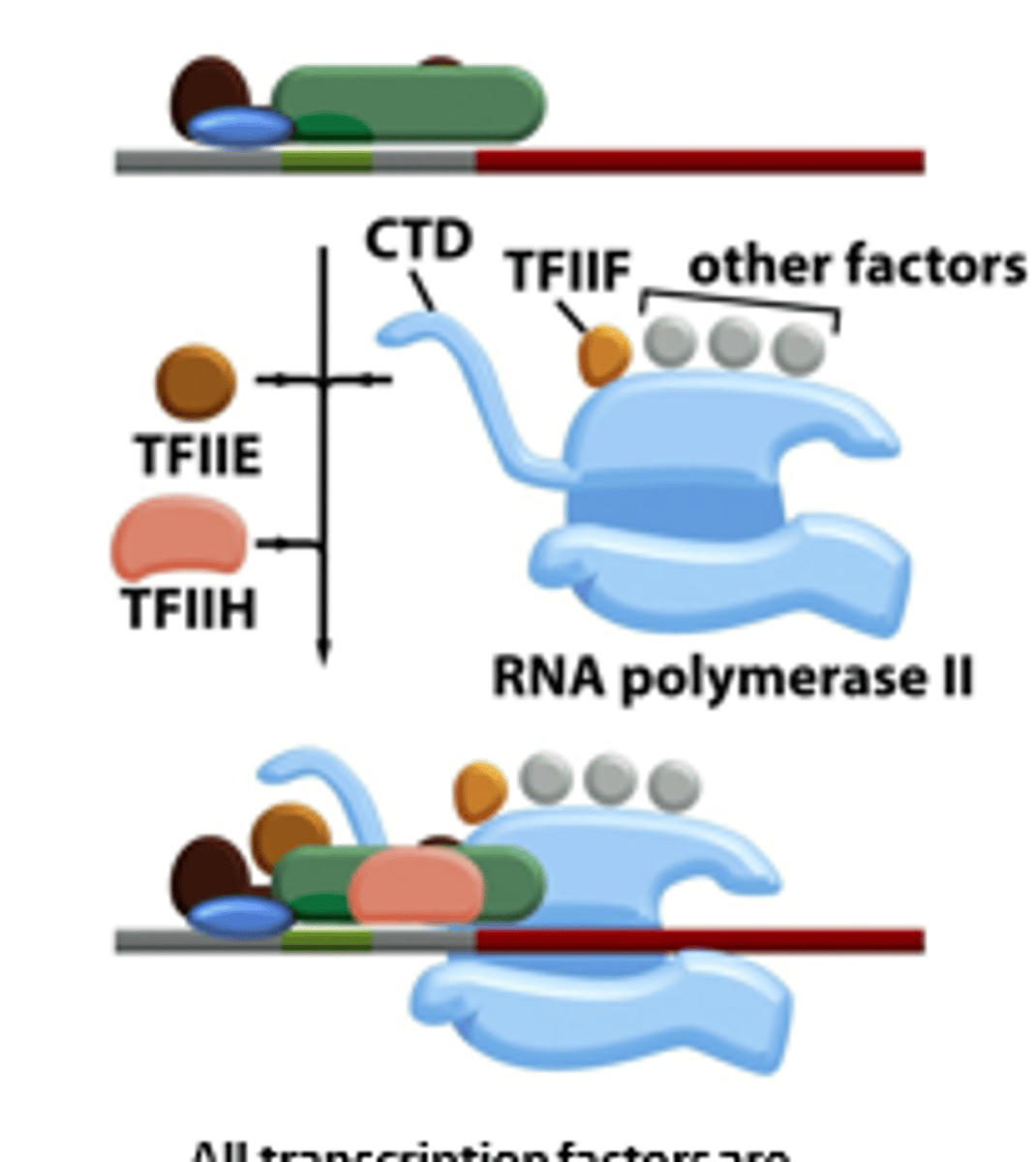
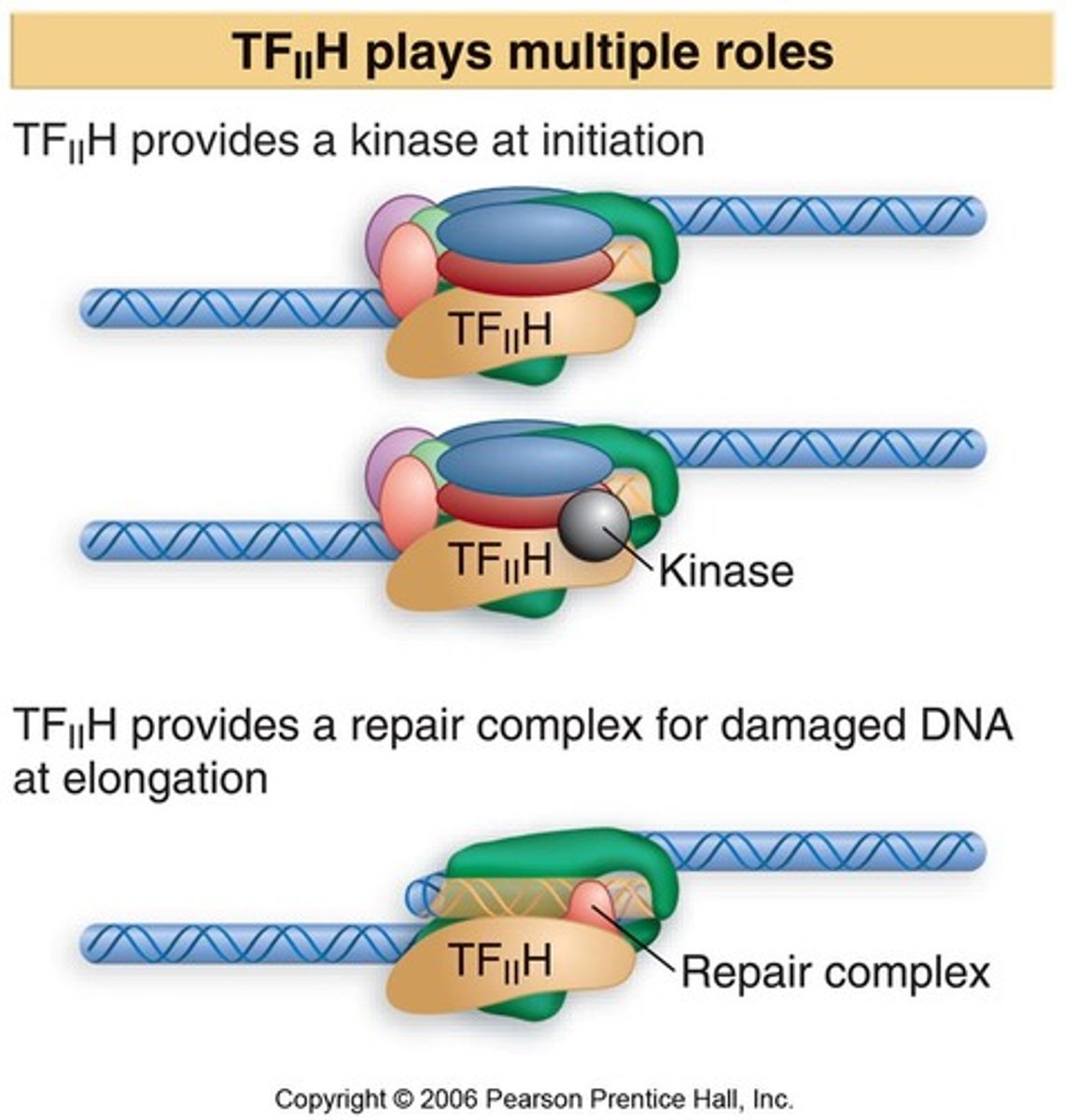
TFIIH
contains 9 subunits,3 of which possess enzymatic activities
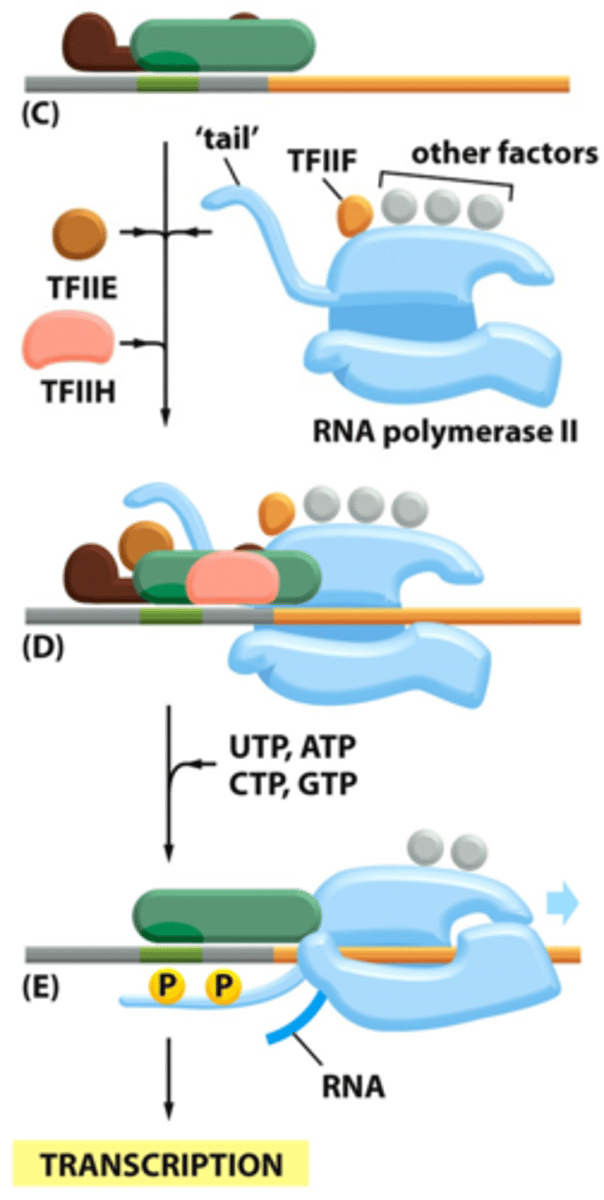
TFIIH's activities includes
1. ATPase helicase (with its helicase domain)
2. Kinase activity which it uses to phosphorylate the CTD of RNA Pol II (C-terminal kinase domain)
TFIIB
Help position the polymerase correctly for transcription initiation
TFIIA
Prevents inhibitory proteins from disrupting TBP binding
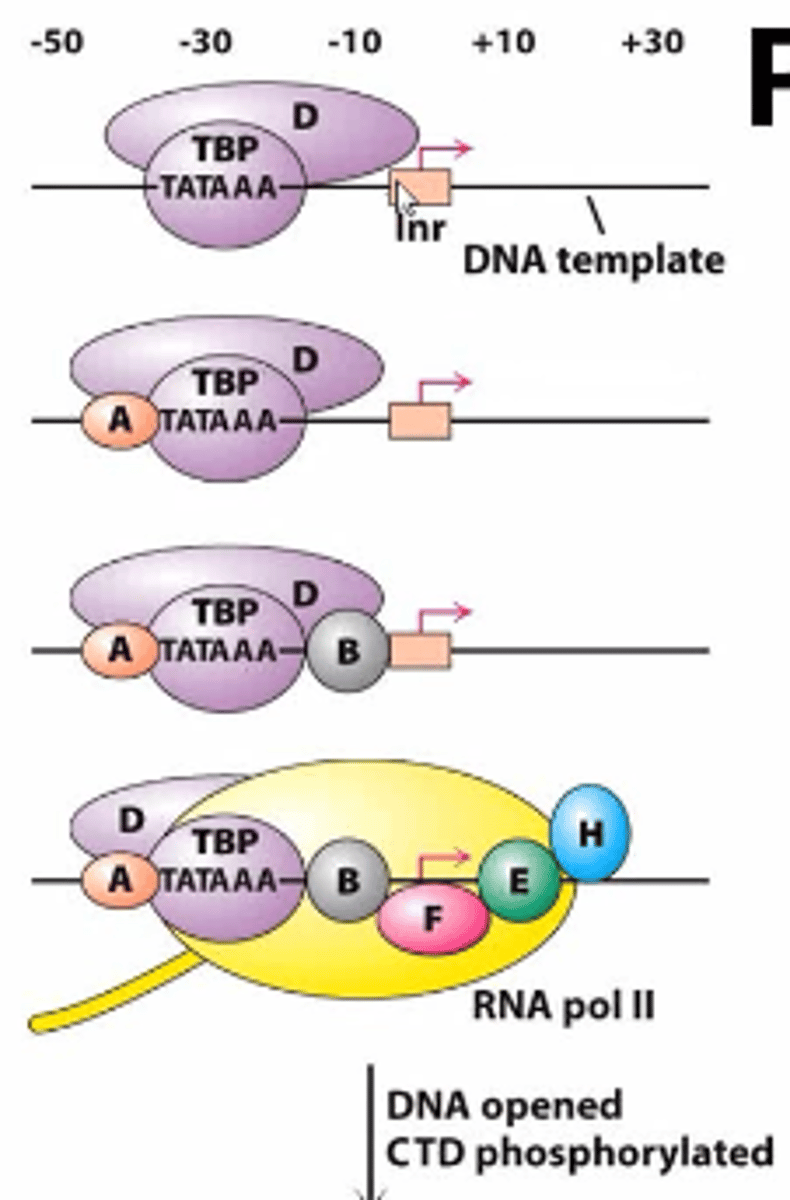
sequential assembly
1. TFIID includes the TBP subunit, which binds to TATA box
2. TFIIB binds and provide a binding site for RNA polymerase II
3. TFIIA binds directly to TBP of TFIID and stabilizes its binding to the promoter
4. TFIIF assist RNA Pol II to bind to the promoter
5. TFIIE binds to the complex and recruits TFIIH
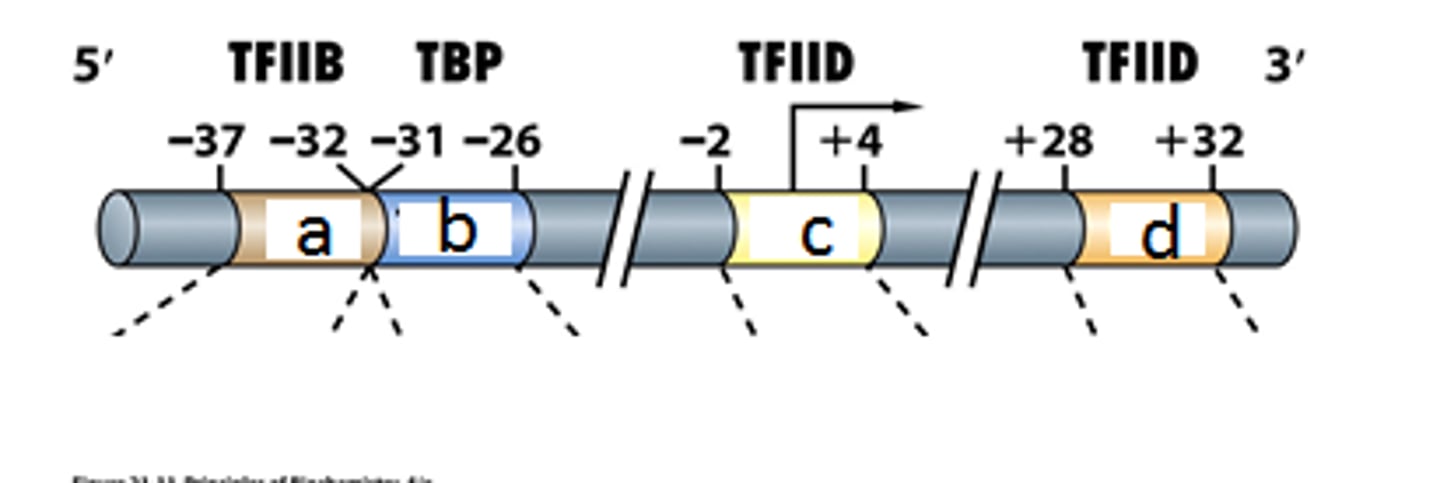
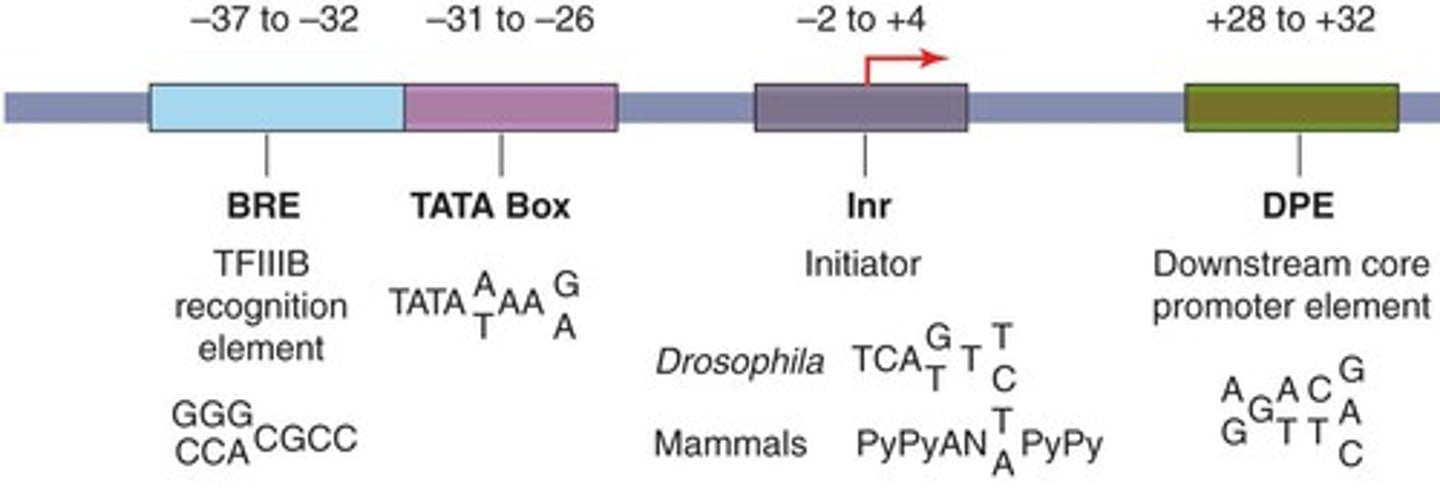
B recognition element (BRE)
found immediately upstream of the TATA box
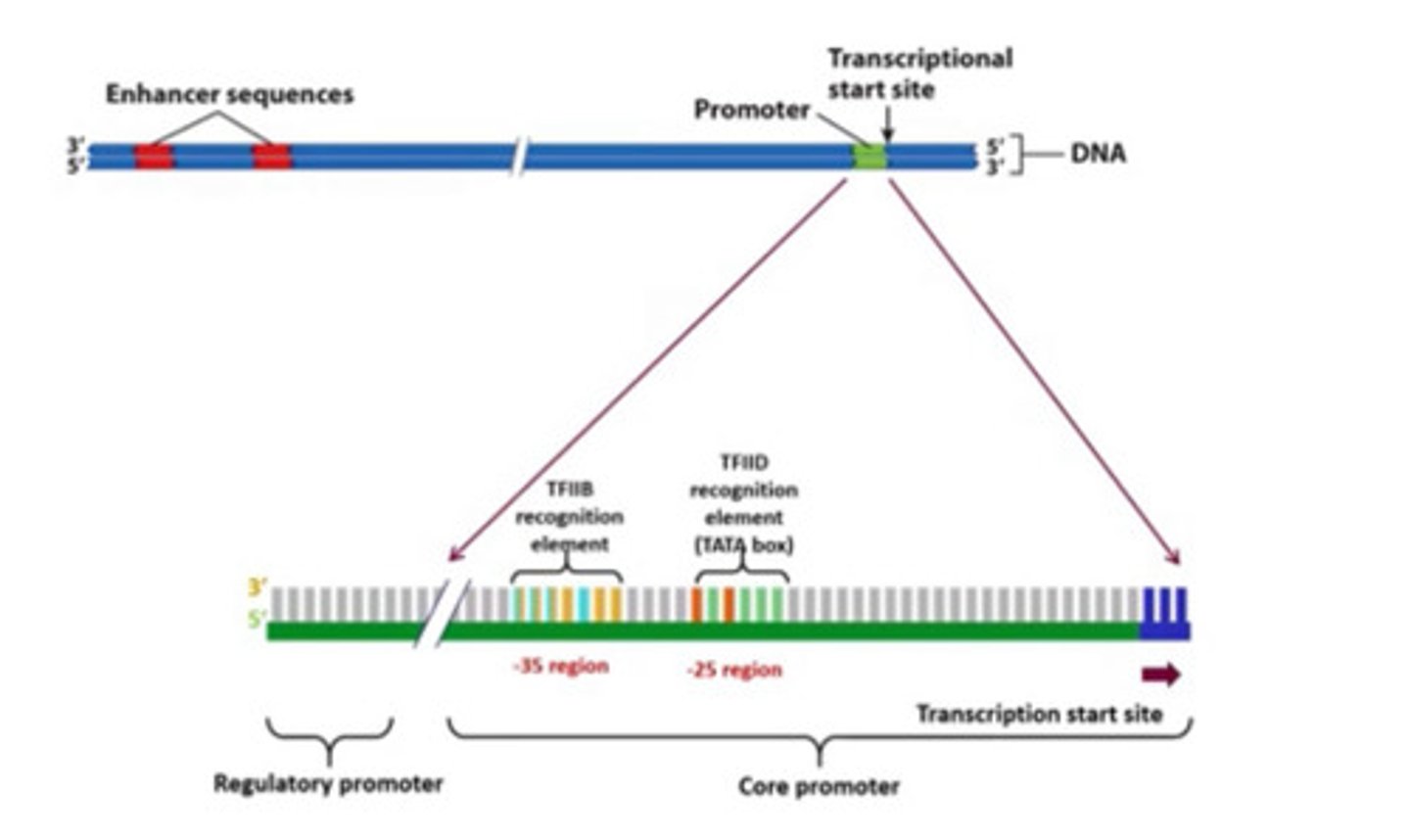
initiator element (Inr)
Some genes do not contain a TATA box and use an Inr for transcription initiation; overlaps TSS
enhances the strength of a promoter that contains a TATA box
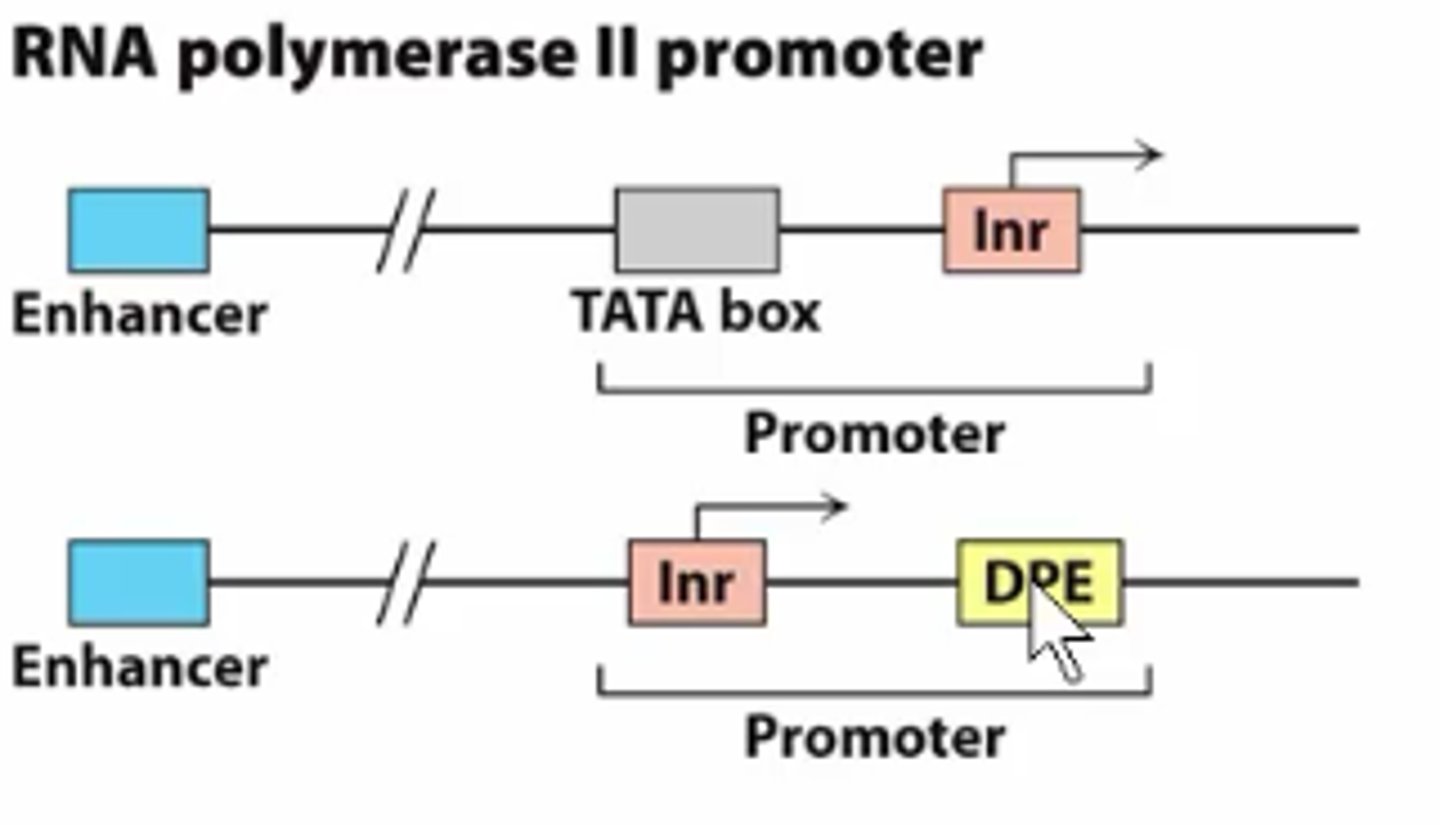
downstream promoter element
is within the transcribed portion of a gene
STEP 2 (Pre-initiation)
After PIC formation, a "transcription bubble" is created (The 3' to 5' DNA template strand is single stranded and ready to be transcribed into RNA)
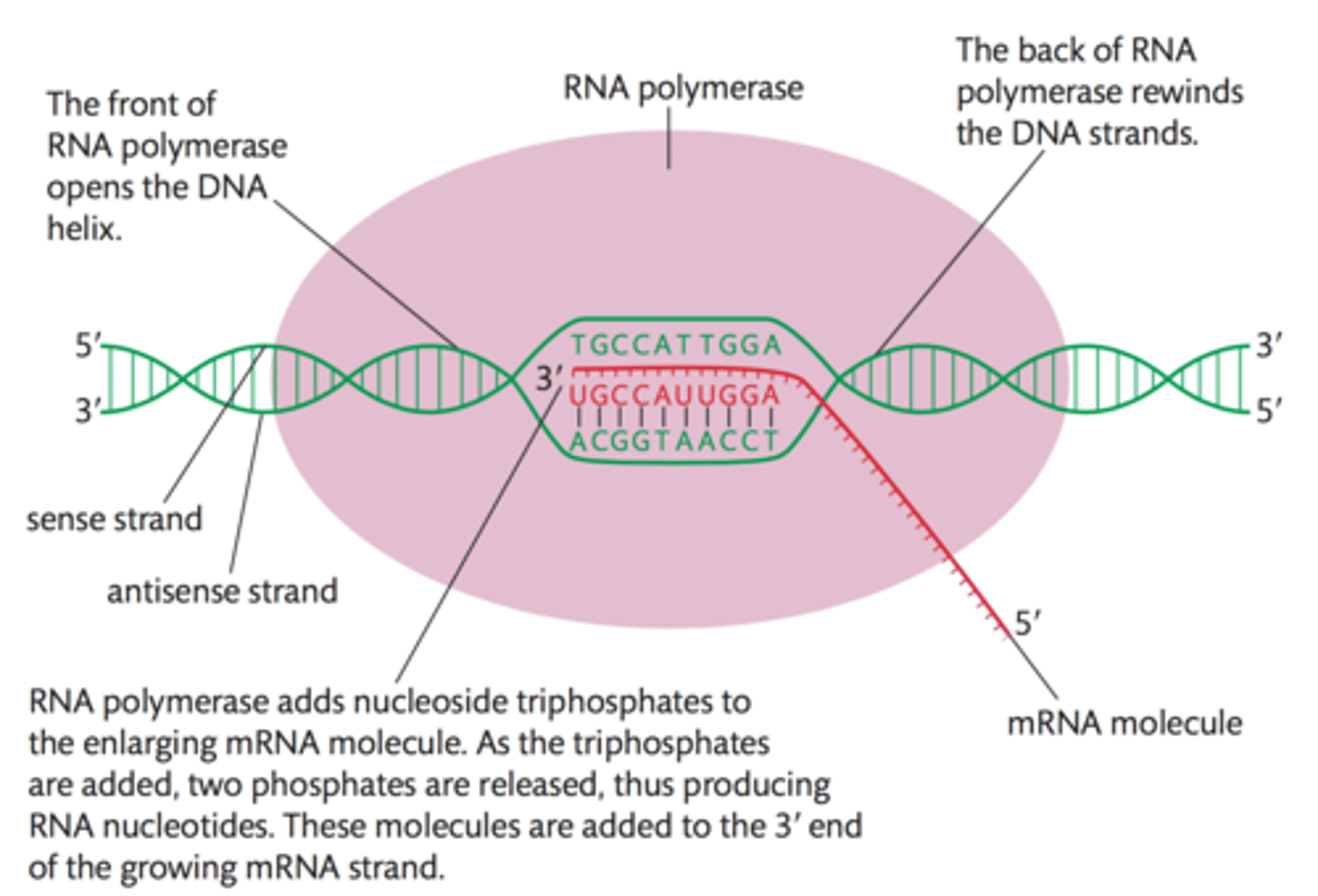
Two subunits of TFIIH
helicases; unzips ∼6 bp of DNA upstream from the TSS
RNA Pol II then catalyses further DNA unzipping, leading to a DNA bubble measuring 13 bp
STEP 3 (Initiation)
Abortive initiation - formation of an early elongation complex
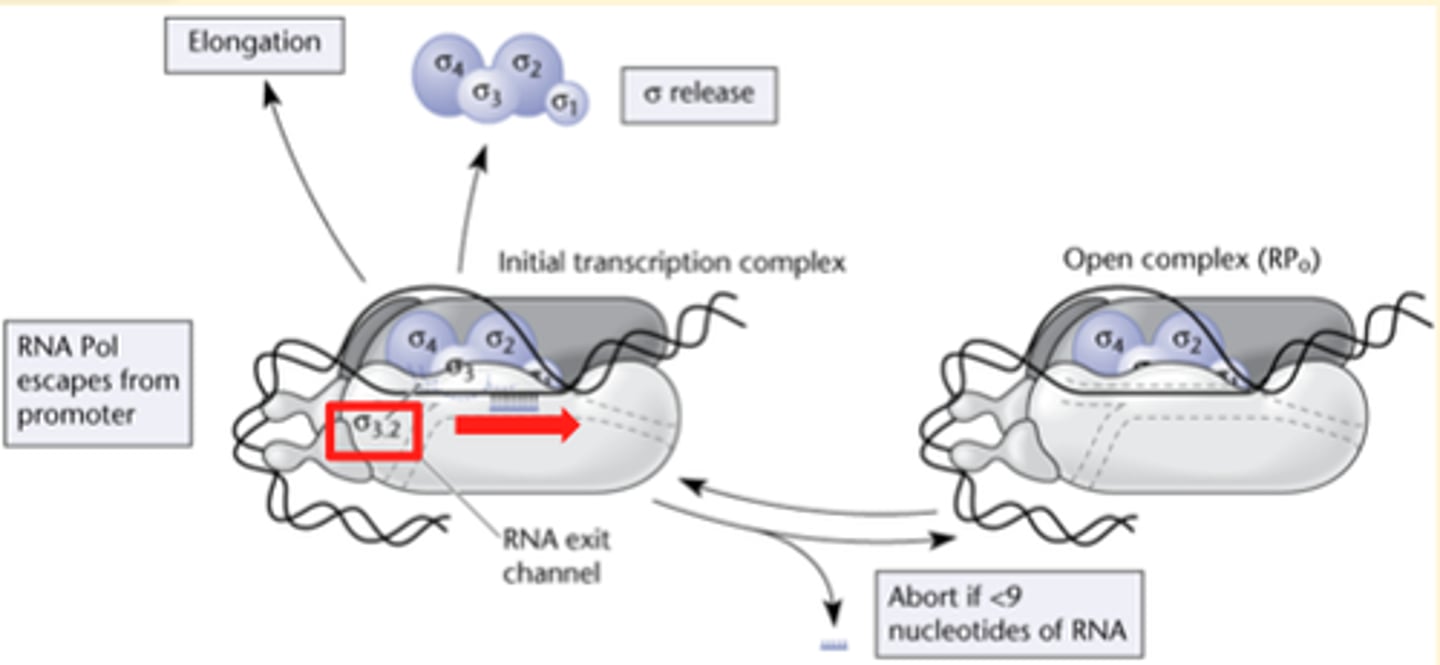
abortive initiation
releases large amounts of 2-3nucleotide long RNA transcripts
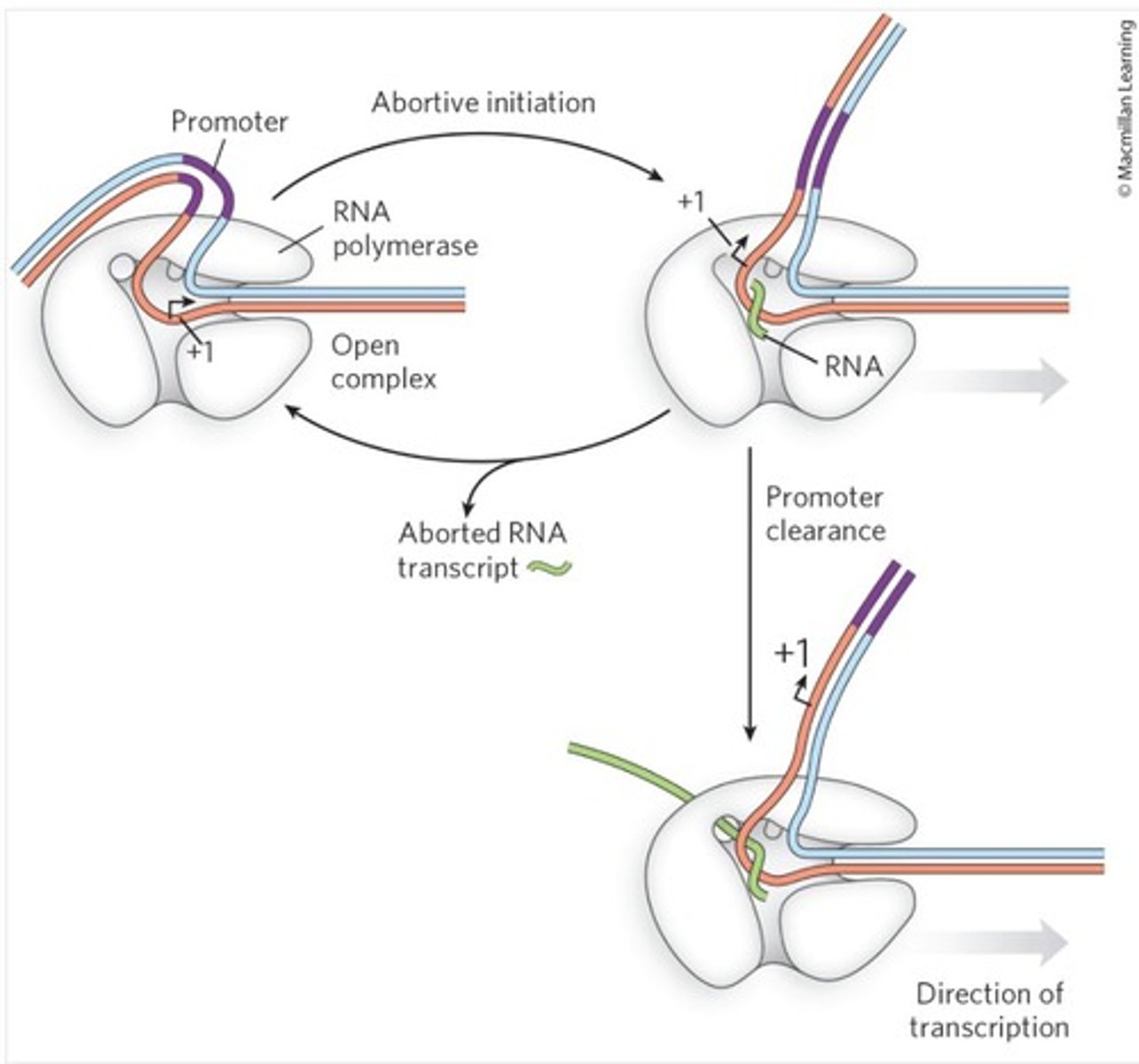
latched RNA pol II
RNA polymerase remains strongly attached to the promoter and the pre-initiation complex, so it repeatedly releases short RNA fragments
STEP 4 (Initiation)
Escape commitment (meaning a commitment to escape from the promoter)
reducing abortive initiation
When the RNA Pol II is able to synthesize 4 nucleotides, the 4-nucleotide-RNA transcript makes contact with TFIIB which stabilizes the short newly synthesized RNA
escape commitment
formation of this stable transcription complex; RNA Pol II is now committed to move away from the promoter
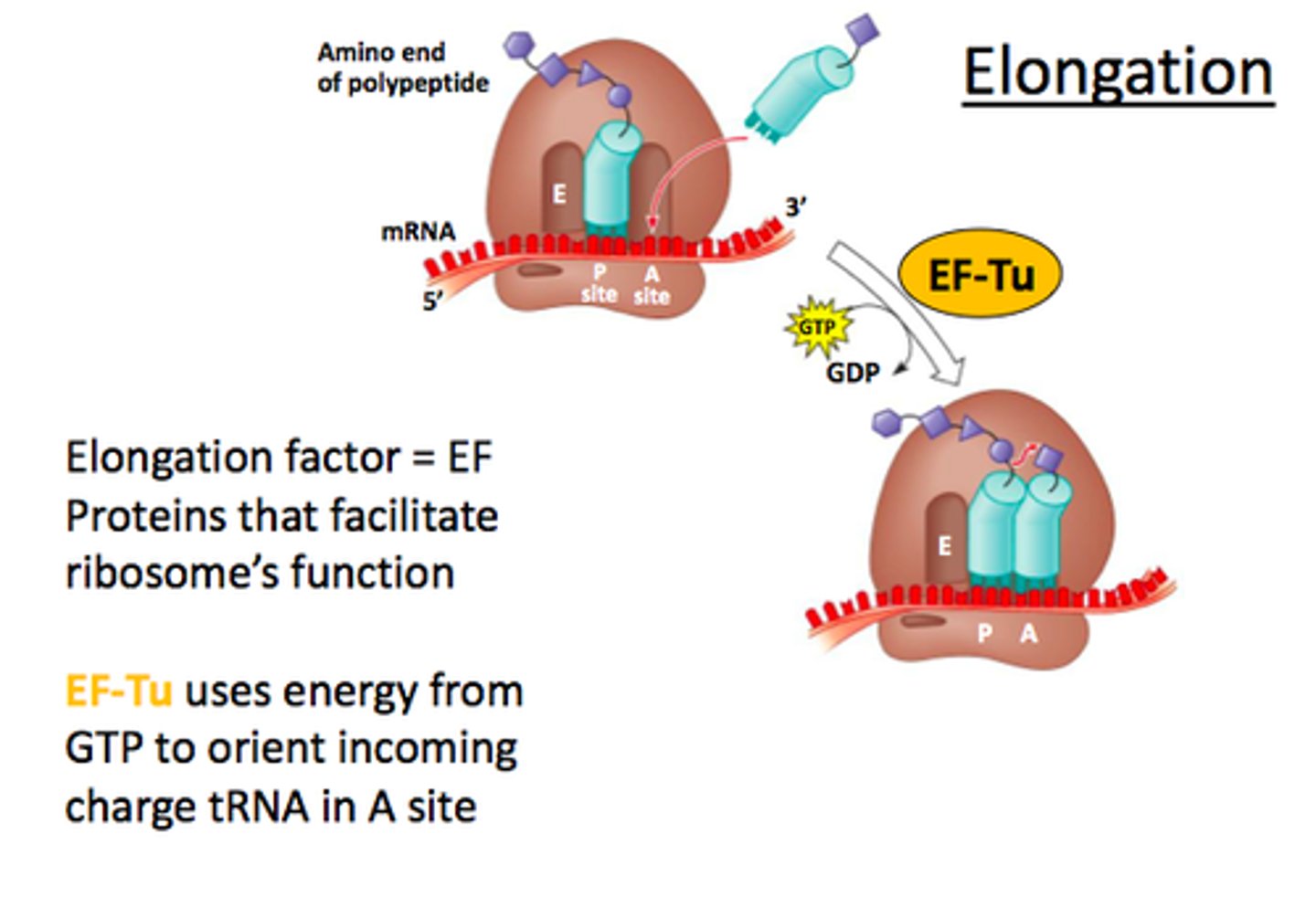
STEP 5 (Elongation)
Early elongation complex
early elongation complex
Once the RNA Pol II has committed to move away from the promoter, the RNA transcript grows to about 7-nucleotidesand the transcription complex
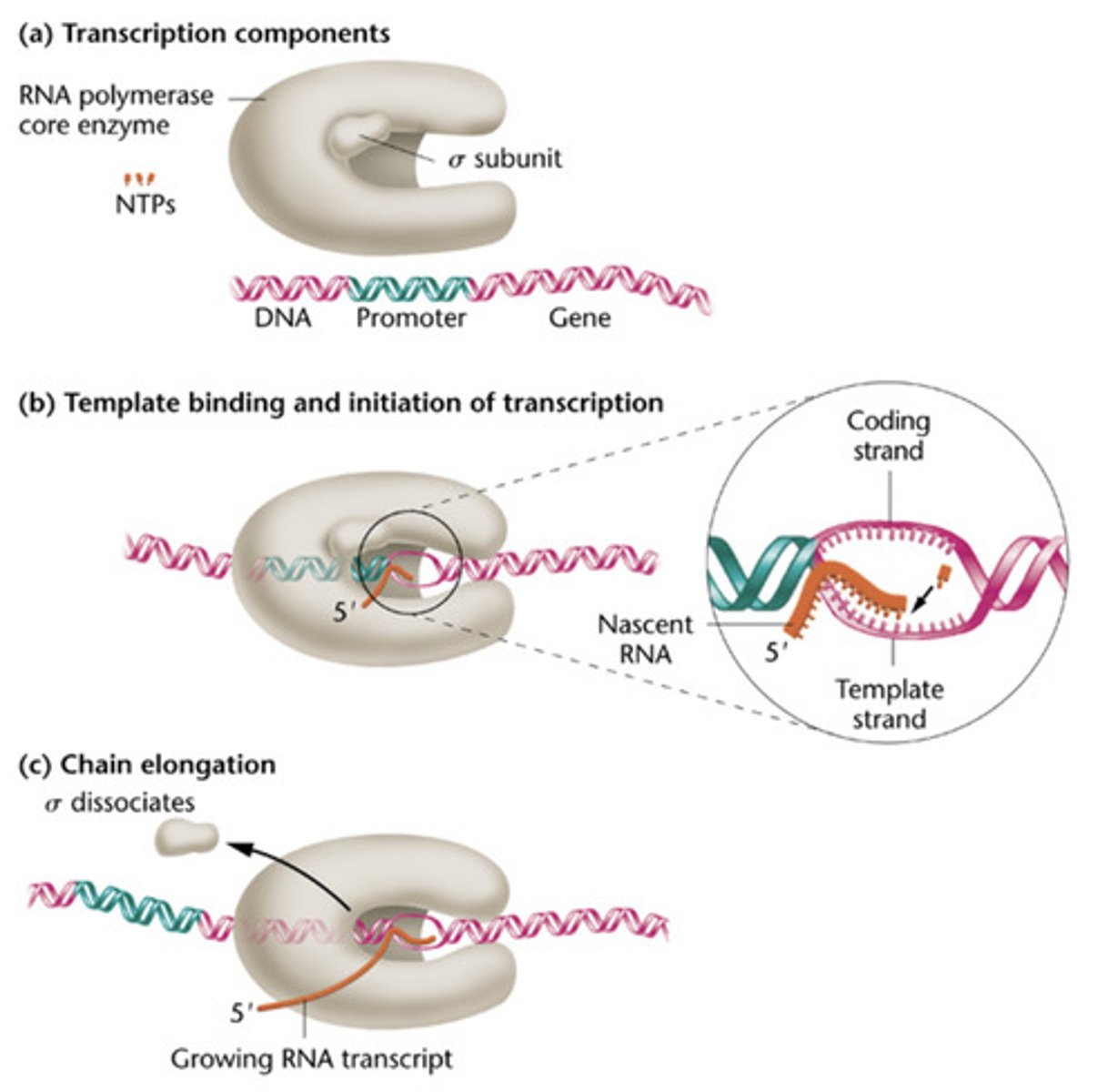
STEP 6 (Elongation)
Escape from promoter
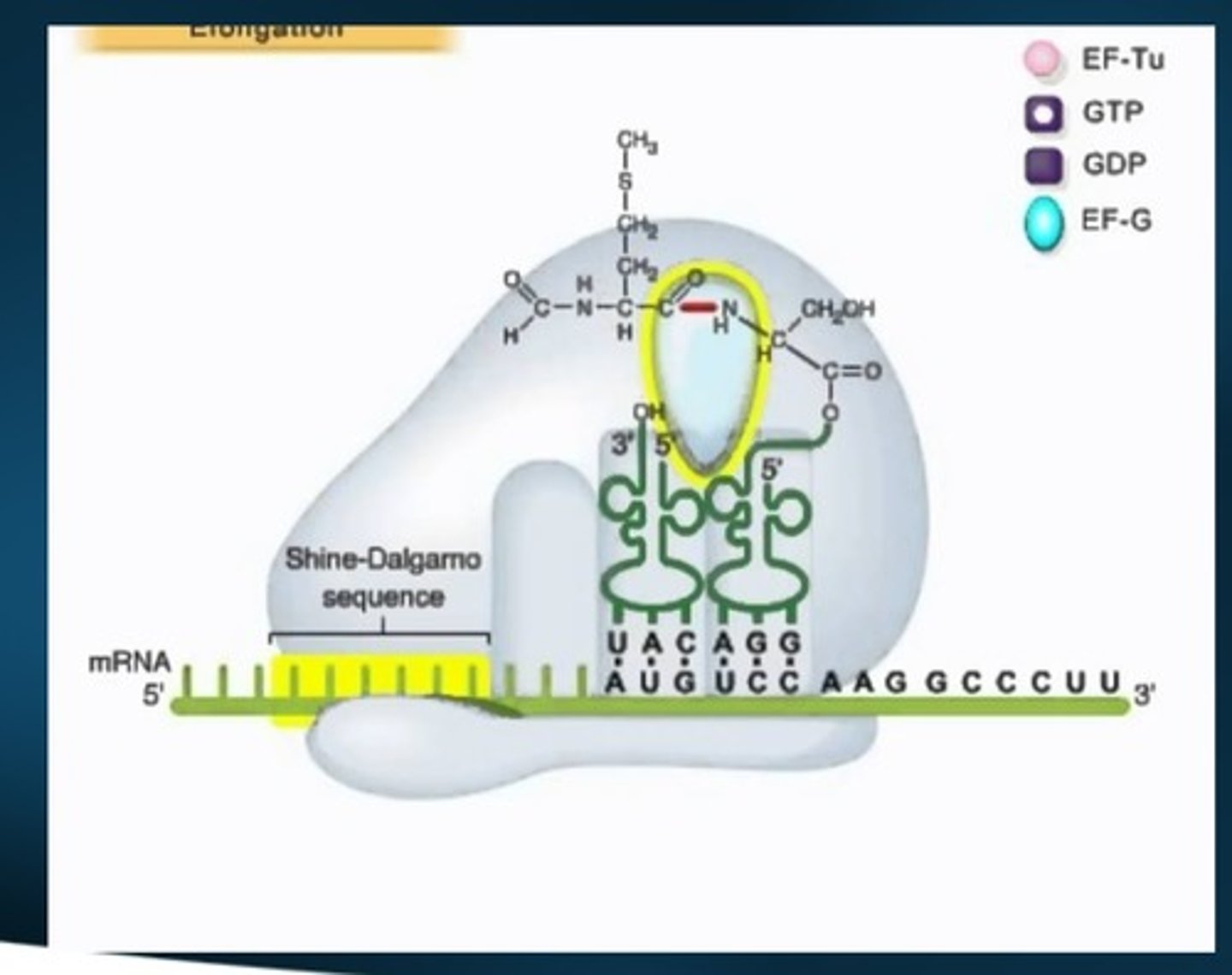
"escape from the promoter"
RNA Polymerase II to productively elongate the RNA
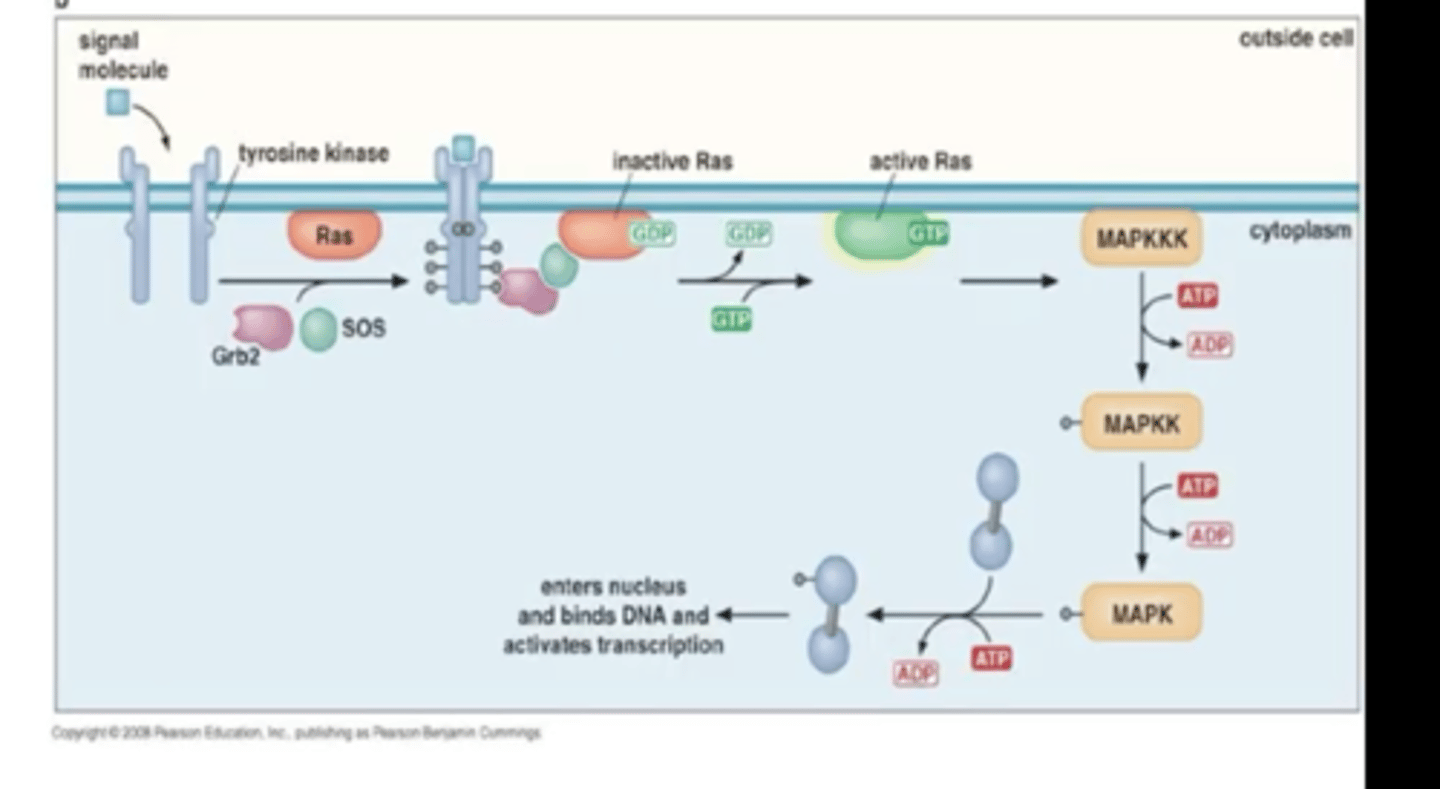
RNA Pol II is productively elongating
(adding nucleotides) the RNA transcript, it is highly processive and it does not dissociate from the DNA template until it finishes
Pol II must break contact with the promoter-sequence to do this
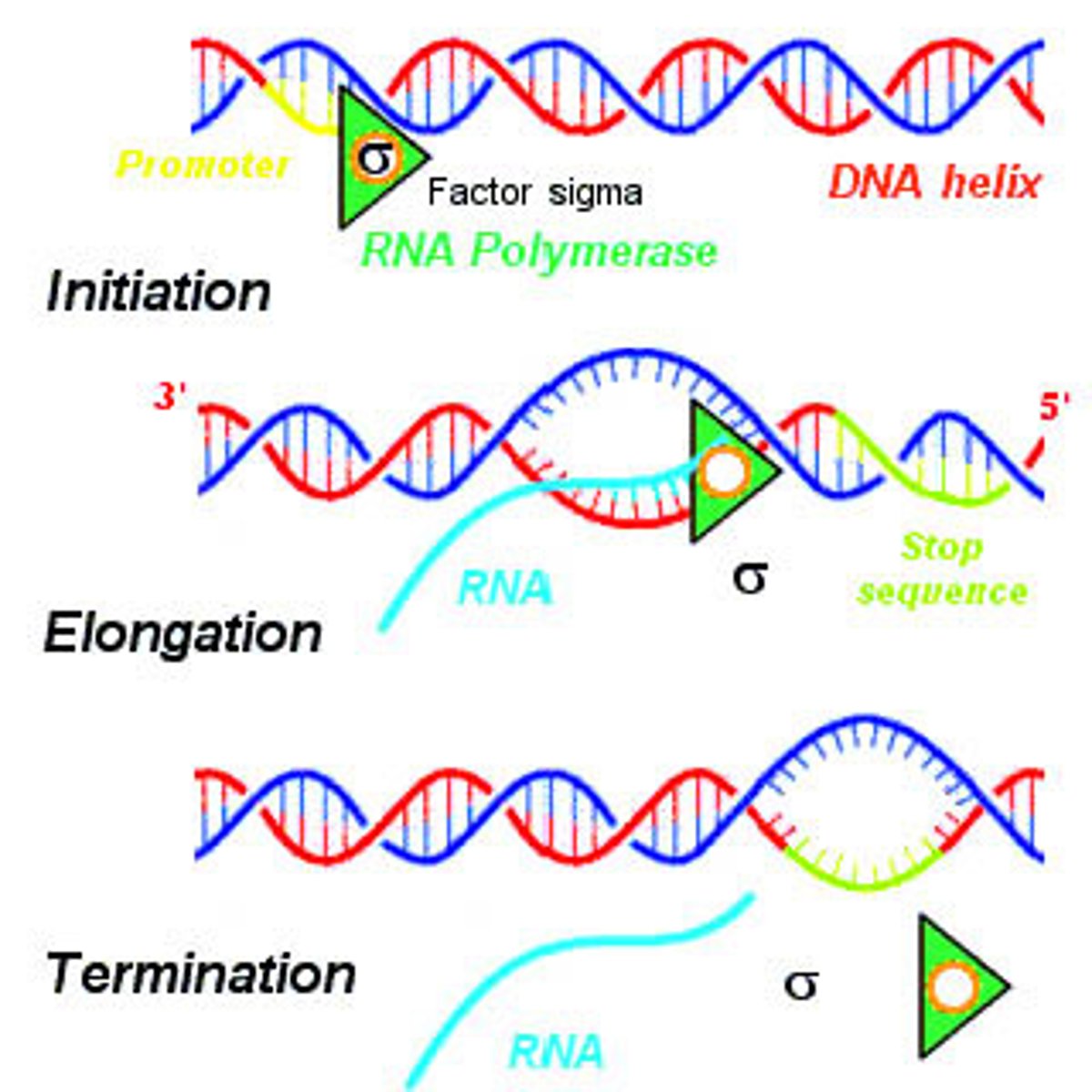
Carboxyl-terminal domain (CTD phosphorylation)
the largest subunit of polymerase II consists of seven amino acid repeats (Tyr-Ser-Pro-Thr-Ser-Pro-Ser)
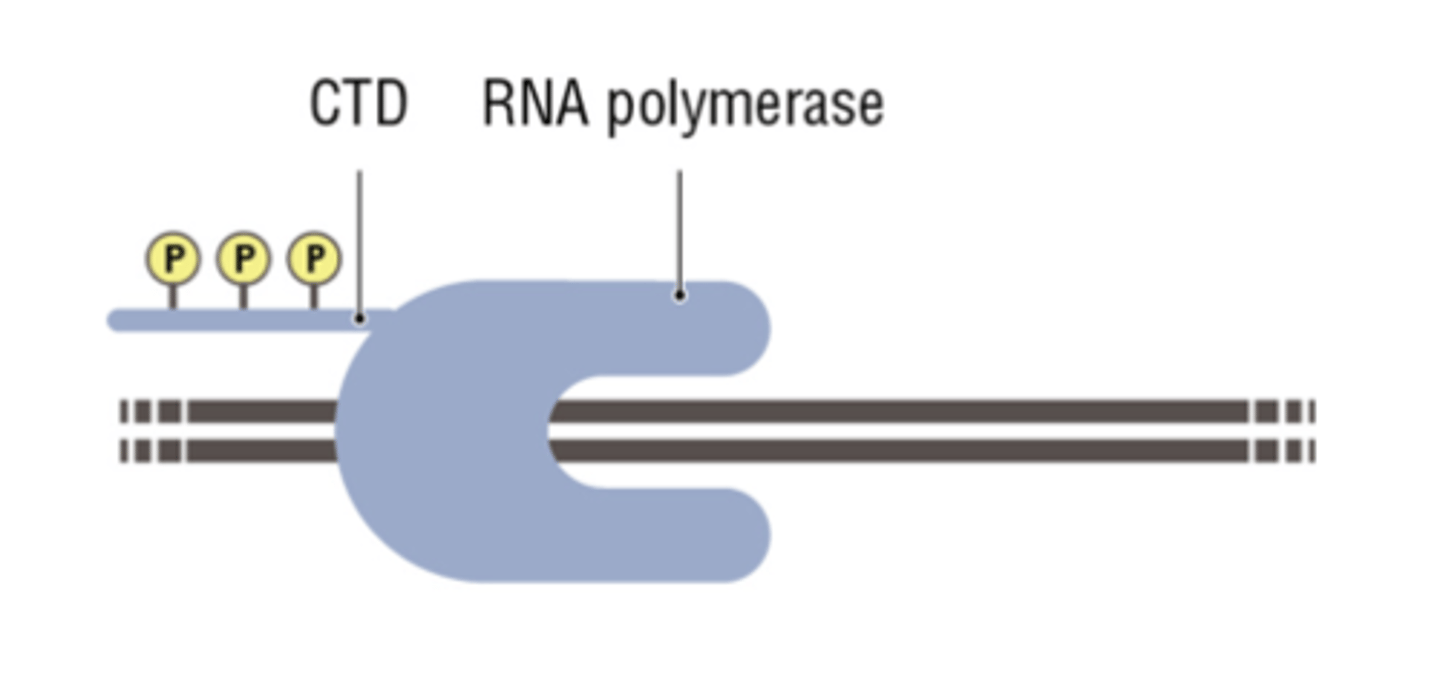
RNA Polymerase II CTD phosphorylation is required for promoter escape
The kinase domain of TFIIH phosphorylates the serine #5 on the heptad of the CTD
CTD tether
holding a variety of proteins/enzymes involved in transcription elongation (elongation factors (ef) and mRNA processing enzymes close by until they are needed
STEP 7 (Elongation)
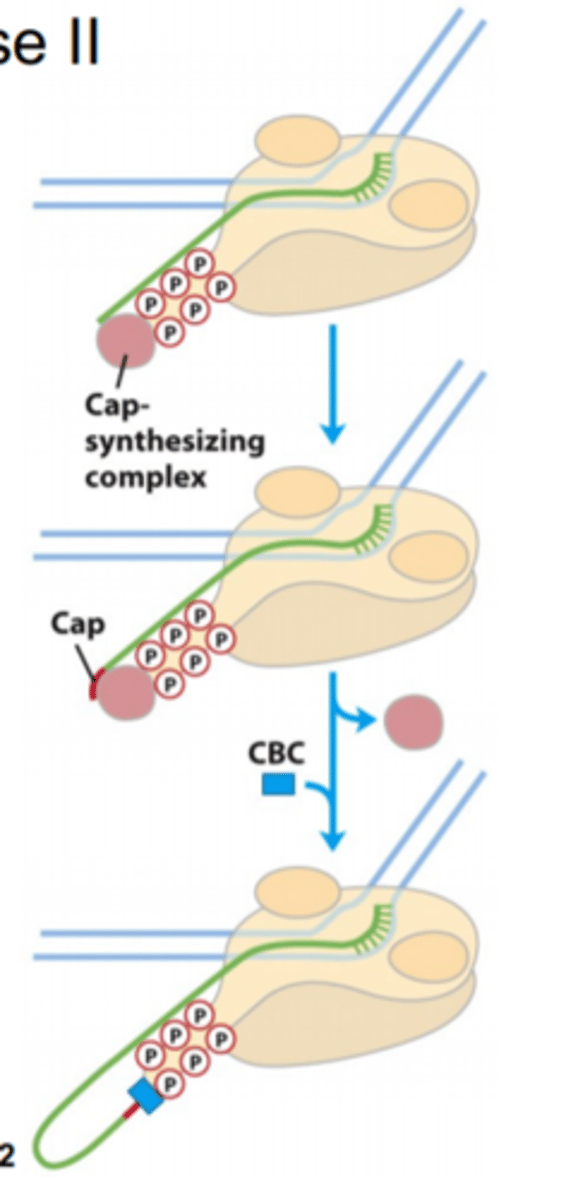
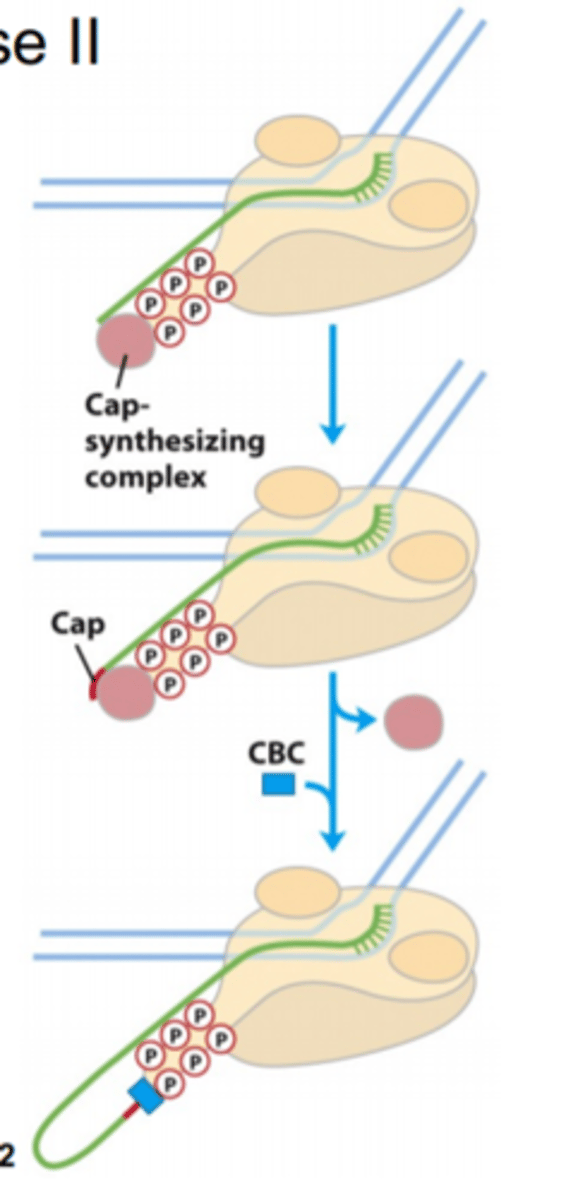
Transitioning to the pause region
- add 5' cap
- regulated (stop/go signal
P-TEFb
transcription elongation factor
phosphorylates serine #2 on the heptad CTD tail to mobilized the RNA Pol II from the pause
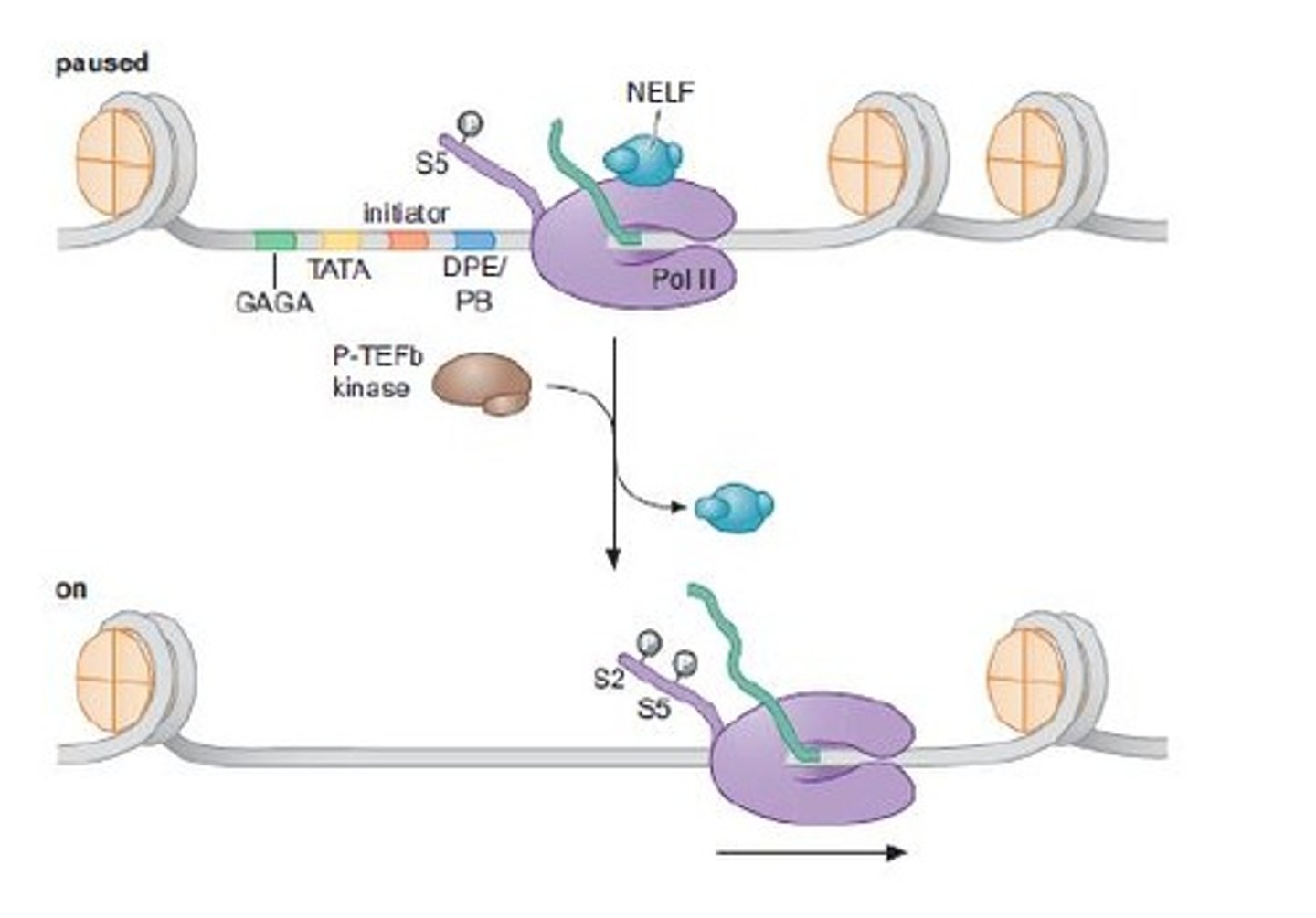
STEP 8 (Elongation)
Productive elongation
The RNA Pol II will continue productive elongation until the entire gene is transcribed
Release of paused Pol II into productive transcription
triggered by transcription factors that recruit P-TEFb, a kinase that phosphorylates the Pol II C-terminal domain and promotes elongation
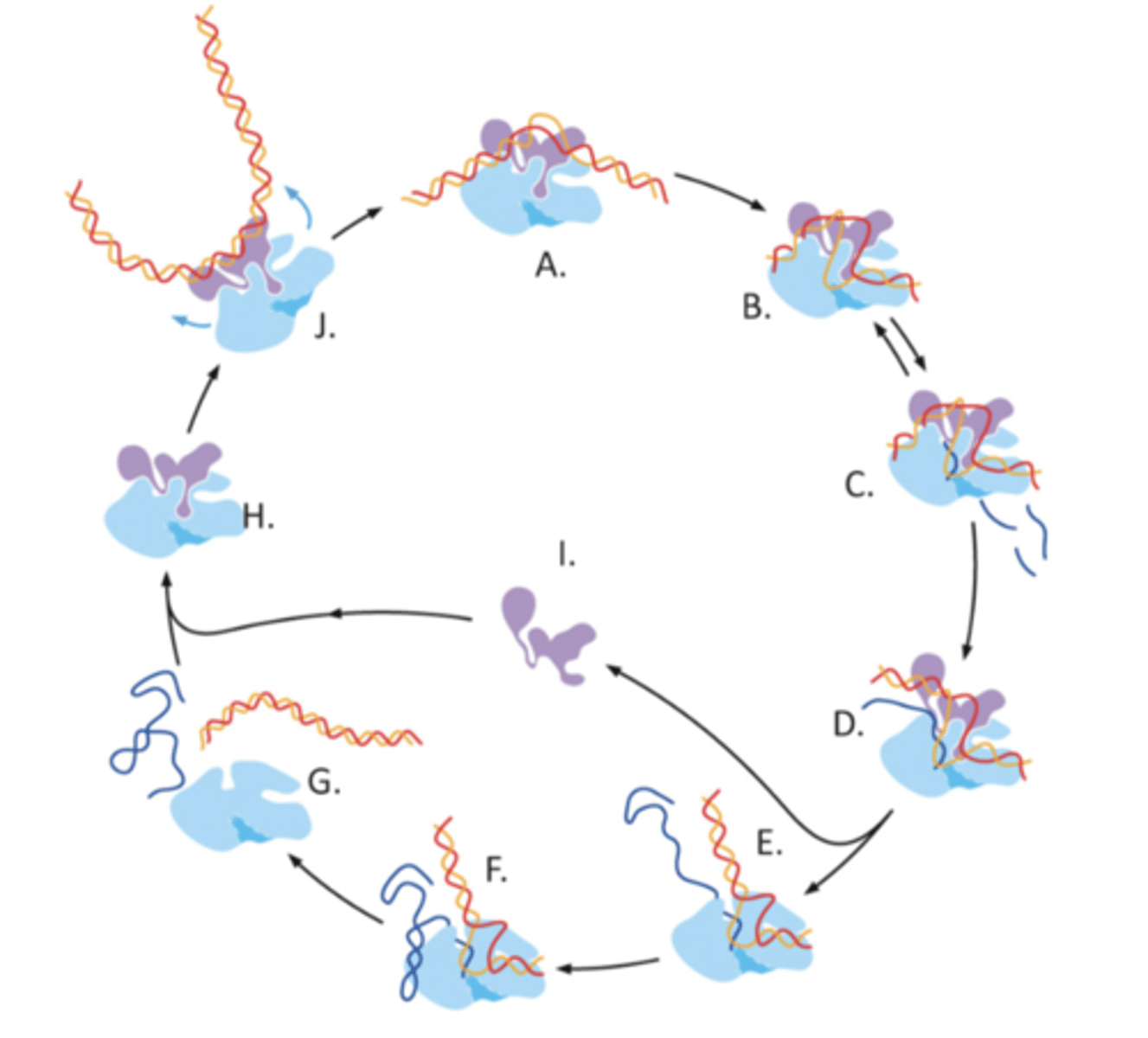
SII, ELL, and PTEFb
three of a number of elongation factors that may be associated with the polymerase as it moves along the DNA
PTEFb
kinase that phosphorylates theSer2 residues of the CTD after elongation begins
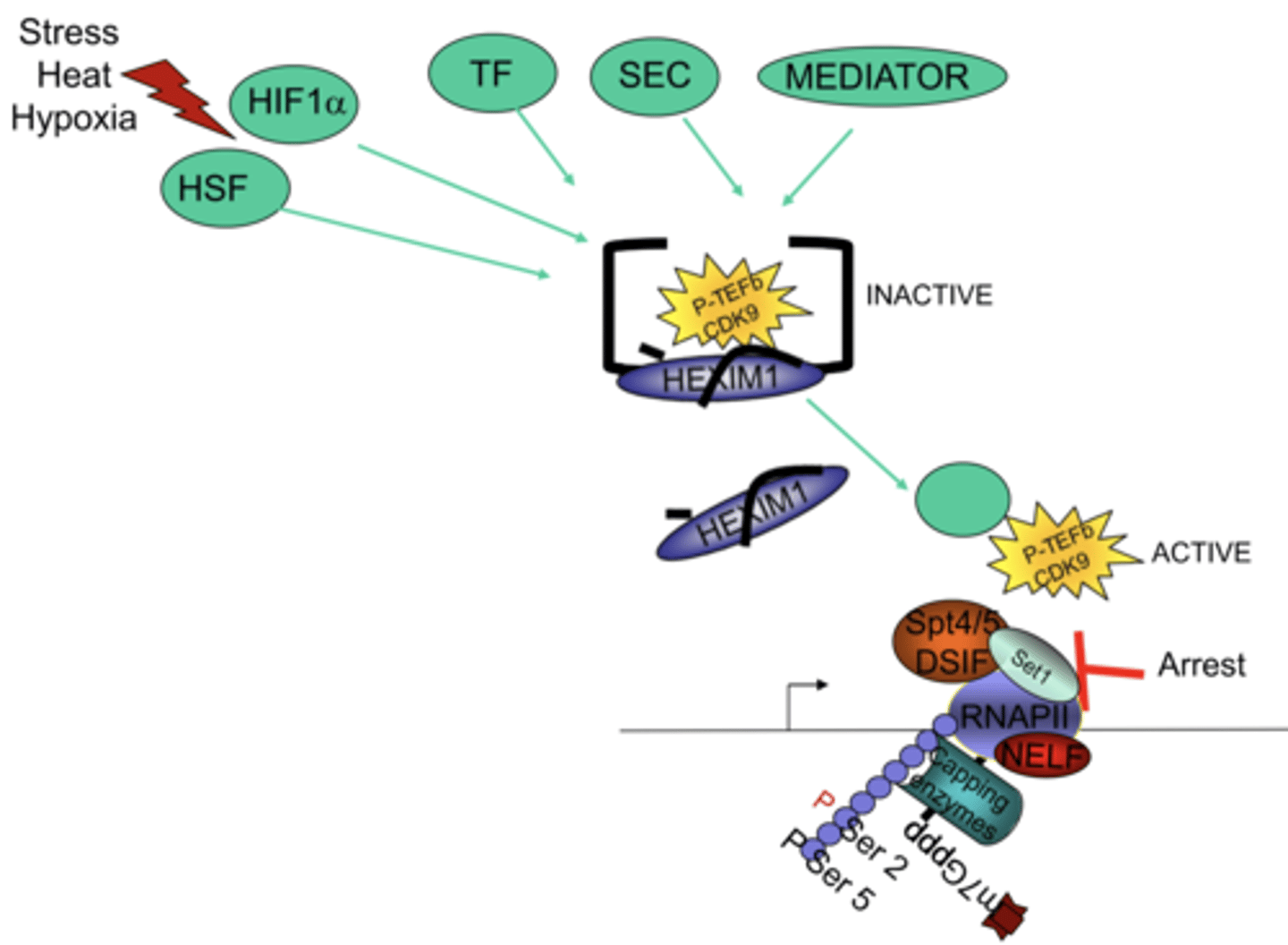
STEP 9 (Termination)