MBB lec 21 eukaryotic DNA
1/21
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
22 Terms
when is dna in most condensed/visible form
metaphase chromosome (X-shaped)
functions of dna being packaged compact
protect
contribute to regulation of replication/transcription
name the levels of dna packaging from most condensed to most simple
-metaphase chromosome (1400nm) (fully condensed during division)
-condensed chromatin (700nm) (folded/compact)
-looped domains (300nm) (looped chromatin for organization)
-chromatin fiber (30nm) (coiled nucleosomes, form thick fiber)
-nucleosomes (11nm) (dna wrapped around histones)(beads on string)
-dna (2nm)
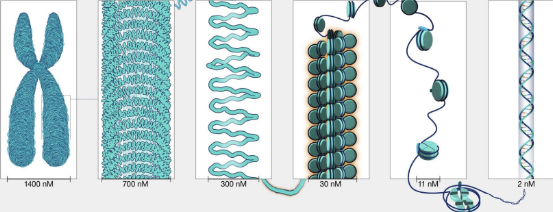
label each dna packaging level
right to left: dna, nucleosomes (circles is histone in middle), chromatin fiber, looped domains, condensed chromatin, metaphase chromatin)
describe the first level of chromatin packaging
Nucleosome
-dna wrapped around histone proteins (bead on string structure)
-aka 11nm fiber
-even tho dna packed, still accessible (important for transcription)
what are histones? are they charged? why? how assembled
-major PROTEIN component of CHROMATIN
-positive charge (rich in Lys and Arg) binds to negative DNA
-assembly requires histone chaperones (highly conserved)
5 types of histones and which one is in nucleosomes
H2A, H2B, H4, H3 (core histones)
H1 (clamps dna wrapped around nucleosome)
how does histone core protect dna in nucleosome
protect from degradation by nuclease
what is the 2nd level of chromatin organization
-30 nm fiber (aka solenoids or chromatin fiber) made by coiled 11nm fibers
-chromosomes in this structure most of time (interphase)
-H1 histone on interior of 30nm fiber
what interactions help pack the 11nm fiber into more compact 30nm fiber
positive N-terminal tail BIND negative sugar-phosphate backbone of neighbouring nucleosomes
what are the more open, less compact regions with less H1. REGIONS that are more accessible to transcription machinery
actively transcribed regions (eu-chromatin)
in solenoids (30nm fiber), what repositions/removes histones to increase dna accessibility
atp-dependent chromatin remodeling complexes
dynamic chromatin structure allows dna to switch between ___ and ___ states. which is active and which isnt
condensed (inactive) and open (active) states
what 2 processes increase accessibility for chromatins dynamic structure to switch between closed/open states (expose dna and allow for transcription replication repair)
Chromatin remodeling complexes (use ATP to reposition/remove nucleosomes)
Histone-modifying enzymes (ex. histone acetyltransferases HAT and deacetylases) alter histone charge and dna interaction
further compacted 30nm fiber forms the next level, what is it and what is it attached to. this organization allows dna to be compacted up to about ____ fold
looped domains
-attached to a nuclear scaffold (nuclear scaffold proteins like H1, topoisomerase II stabilize high-order structure)
-compacted up to 10k-fold
describe size and appearance of chromatin in interphase vs M phase mitosis
interphase: less condensed (30nm fiber) and exist as induvidual chromosomes
M phase: highly condensed into visible chromosomes
what is the site where dna relication begins during s phase
replication origin
where do splindle fibers attach to pull chromatids apart during cell division
centromere
whats 2 functions of telomeres
protect from degradation
solve end replication problem
explain the end replication problem and its solution
-dna poly cant start synthesis without rna primer
-at end of lagging strand, final rna primer removed
-PROBLEM: no upstream 3’ OH available so dna cant be filled, leaving primer gap
-RESULT: chromosomes get shorter each replication
-Solution: telomerase extends 3’ end of chromosome
steps: 1. bind to 3’ end of strand overhand and its rna base pairs with telomere dna
extends 3’ end (acts as reverse transcriptase, uring rna template to add dna repeats)
once extended, primase adds rna primer for dna pol to finish dna synthesis
how are ssdna ends of euk chromosomes protected from degradation
-telomeres form protective T-loop at chromo ends
-3’ overhand folds back and invades dna (D-loop formation) that hides chromo end, reventing it from being recognized as dna damage
-telomere-binding proteins stabilize structure and protect against nucleases