Biology Exam 2 Review
1/64
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
65 Terms
phospholipid bilayer
double layer of phospholipids that make up the cell membrane
membrane interior
fatty acids (hydrophobic, nonpolar) facing inward & interacting via Van der Waal forces
membrane exterior
phosphate group heads (hydrophilic, polar) facing away from each other
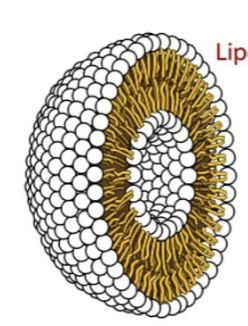
liposomes
spherical lipid bilayers with an internal aqueous compartment and a hydrophobic layer in the middle
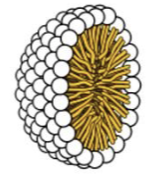
micelles
single-layer structures with a hydrophobic core and a hydrophilic exterior

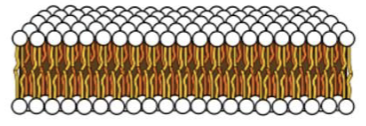
bilayer sheet
basic structural arrangement of the cell membrane before curving into a closed shape
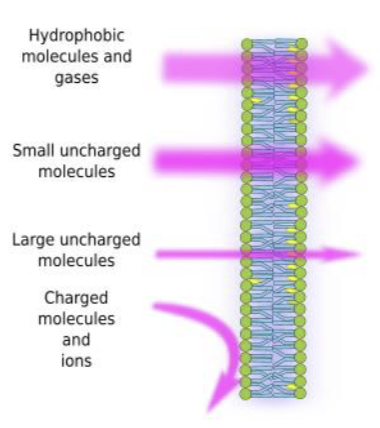
cell membrane structure
semipermeable; nonpolar region bigger than polar region (harder to get through)
small molecules = pass easily
polar molecules & ions: need protein help to cross
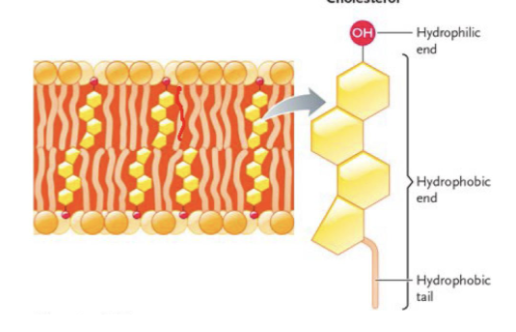
cholesterol
in the cell membrane; regulate membrane fluidity
low temperatures = increased fluidity, preventing crystalization
high temperatures = decreased fluidity, preventing membrane from letting everything through
channel proteins
proteins that form pores to let specific ions or molecules cross the membrane by facilitated diffusion
carrier proteins
proteins that bind a molecule, change shape, and release it on other side of the membrane
receptor proteins
proteins that detect signals and trigger a response inside the cell
enzymes
proteins that speed up chemical reactions at or near the membrane
anchor proteins
proteins that attach to the interior or exterior of the cell membrane for stability
protein pumps
proteins that actively move ions or molecules across the membrane using energy (ATP)
passive transport
uses energy found in a molecule’s electrochemical gradient to move itself; requires no external energy for transport
simple & facilitated diffusion
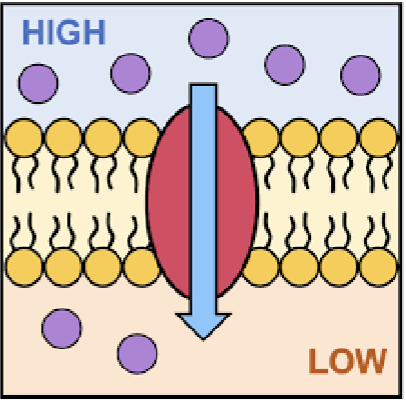
simple diffusion
moves down an electrochemical gradient; goes through the lipid bilayer alone; no protein help
facilitated diffusion
moves down an electrochemical gradient; requires a transport protein
active transport
requires an external addition of energy to facilitate transport ffor a molecule up/against its gradient
primary & secondary
primary active transport
involves use of ATP to provide energy for proteins to pump molecules or ions against their concentration gradient
secondary active transport
uses an existing concentration gradient to move another molecule
transported solute
moves with concentration gradient
coupled ion
moves against concentration gradient
endergonic
requires an input of energy; nonspontaneous
exergonic
releases energy; spontaneous
catabolic
breaks down molecules to release energy
anabolic
builds molecules and requires energy
nonspontaneous
requires energy input to occur
spontaneous
occurs without energy input
hydrolysis
breaks bonds by adding water
dehydration synthesis
forms bonds by removing water
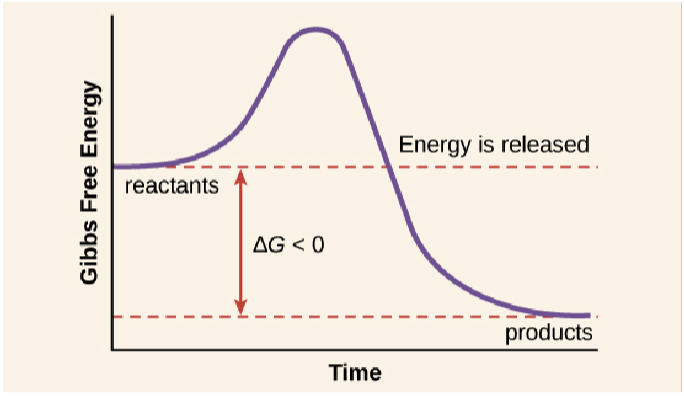
exergonic, spontaneous, catabolic, hydrolysis
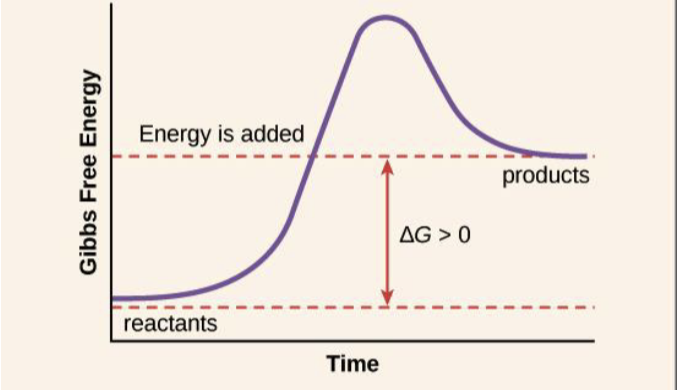
endergonic, nonspontaneous, anabolic, dehydration synthesis
ATP
cellular energy currency; can directly energize proteins in PAT; can energize cellular processes through energy coupling
ATP hydrolysis
spontaneous reaction that breaks off phosphate group from ATP; negative ΔG = reaction releases free energy
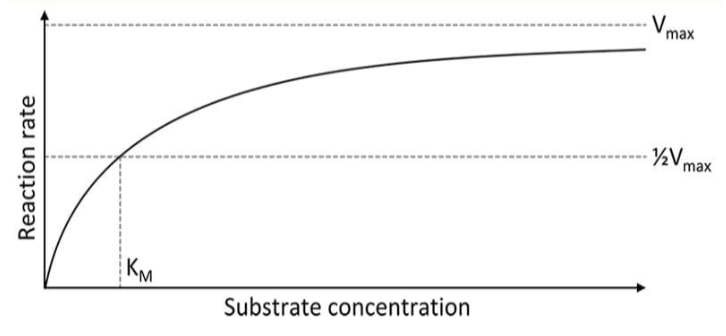
michales-menten model
graph used to show the rate of an enzyme reaction based on substrate concentration
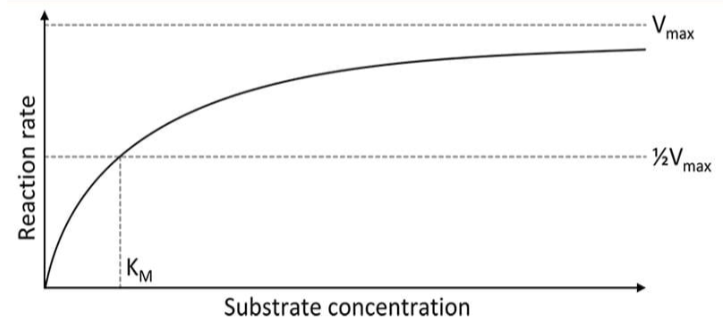
Km
the substrate concentration at which we have reached half of the maximum velocity
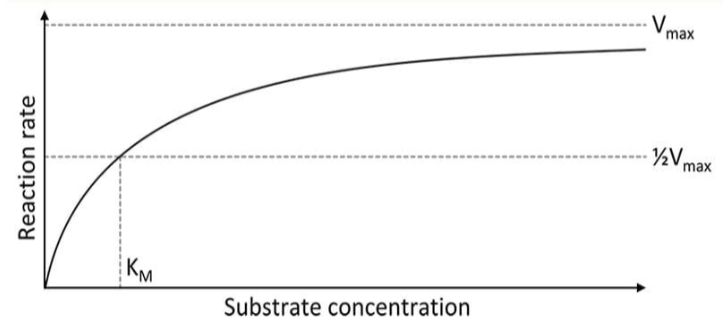
Vmax
the maximum rate of the reaction that can be achieved with a given amount of enzymes
competitive inhibition
binds active site; ↑ Km, Vmax same; substrate increased to reach Vmax
noncompetitive inhibition
binds allosteric site; Vmax ↓, Km same; enzyme activity reduced
speeds up reaction
head, increasing substrate concentration, increasing enzyme concentration
slows/stops reaction
low temperatures (molecules not moving fast enough), denaturation, unsuitable pH, mutations, competitive & allosteric inhibitors
metabolism
a series of net catabolic reactions that transfer energy carrying electrons to NAD+ and FAD, driving ATP production
oxidation in metabolism
glucose losing electrons as its broken into smaller molecules
NADH and FADH2 donating electrons in the electron transport chain and becoming NAD + and FAD
reduction in metabolism
NAD+ and FAD accepting electrons lost and becoming NADH and FADH2
substrate level phosphorylation
enzymes transfer a phosphate group directly to ADP to make ATP
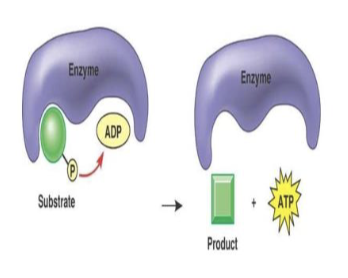
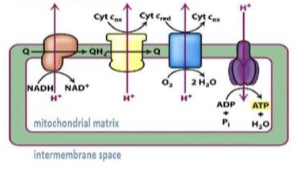
oxidative phosphorylation
uses ATP synthase and an electron transport chain to produce large amounts of ATP
6 carbons
glucose
3 carbons
pyruvate
acetyl-CoA
2 carbons
1 carbon
CO2
electron carriers
NAD+ and FAD (oxidized); NADH and FADH2 (reduced)
glycolysis
the oxidation of glucose into pyruvate, occurs in cytoplasm
pyruvate oxidation
the oxidation of pyruvate into acetyl CoA, occurs in mitochondrial matrix
oxidative phosphorylation
transfer of electrons from NADH and FADH2 through protein complexes in the inner mitochondrial membrane; releases energy to make large amounts of ATP
electron transport chain
set of proteins in the inner mitochondrial membrane where NADH and FADH2 are oxidized and electrons move through complexes
complex I
NADH is oxidized back to NAD+; transfers electrons and pumps protons
complex II
FADH2 is oxidized back to FAD
complex III
transfers electrons and pumps proteins
complex IV
transfers electrons to oxygen (final electron acceptor), forms water, and pumps protons
mitochondrial matrix
innermost space of the mitochondria where pyruvate oxidation + citric acid cycle takes place
inner mitochondrial membrane
the folded membrane where the electron transport chain + ATP synthase are located, producing ATP
proton motive force (PMF)
a gradient of protons in the inner membrane space that want to go back into the matrix
final electron acceptor
oxygen, which receives electrons at complex IV and forms water
purpose of the ETC
to create the proton motive force
ATP synthase
enzyme that uses the proton gradient to produce ATP
function: spins like a turbine to power the conversion of ADP + phosphate into ATP