molecular biology 1
1/93
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
94 Terms
cell
basic unit of life
simple organisms can consist of a single cell (autonomous cells)
living organisms can be viewed as a collection of cells organised in specialised structures (e.g. organs)
cells and evolution
all organisms are related and most probably evolved form a common single-celled ancestor
they can be arranged in the evolutionary tree - eubacteria branch then arches then eukaryota
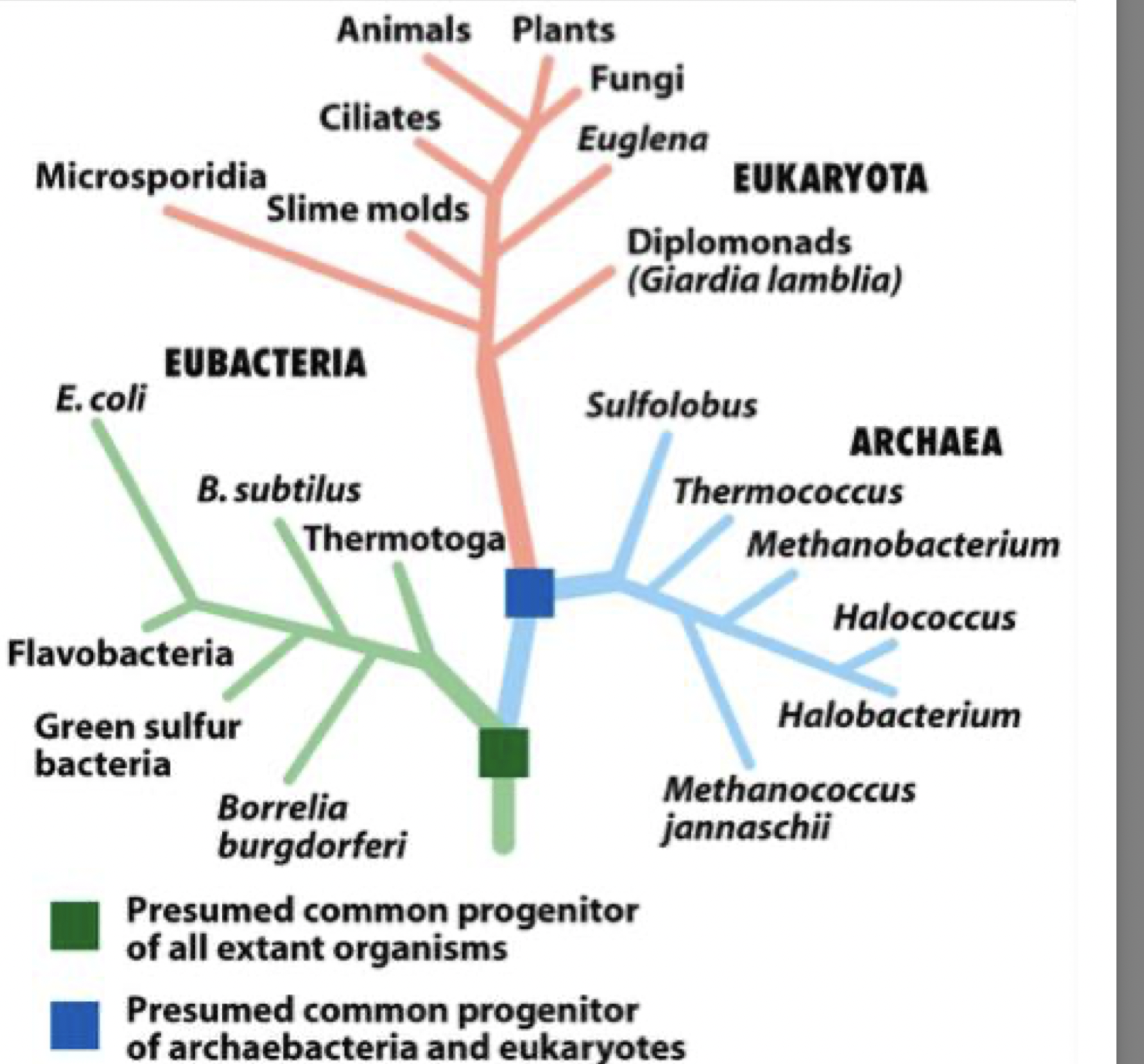
conservation through evolution
key cellular components are conserved through evolution
example: conservation of the globin protein:
hemoglobin and myoglobin in vertebrate and leghemoglobin in other species all from an ancestral oxygen-binding protein
example of key motifs being conserved throughout evolution
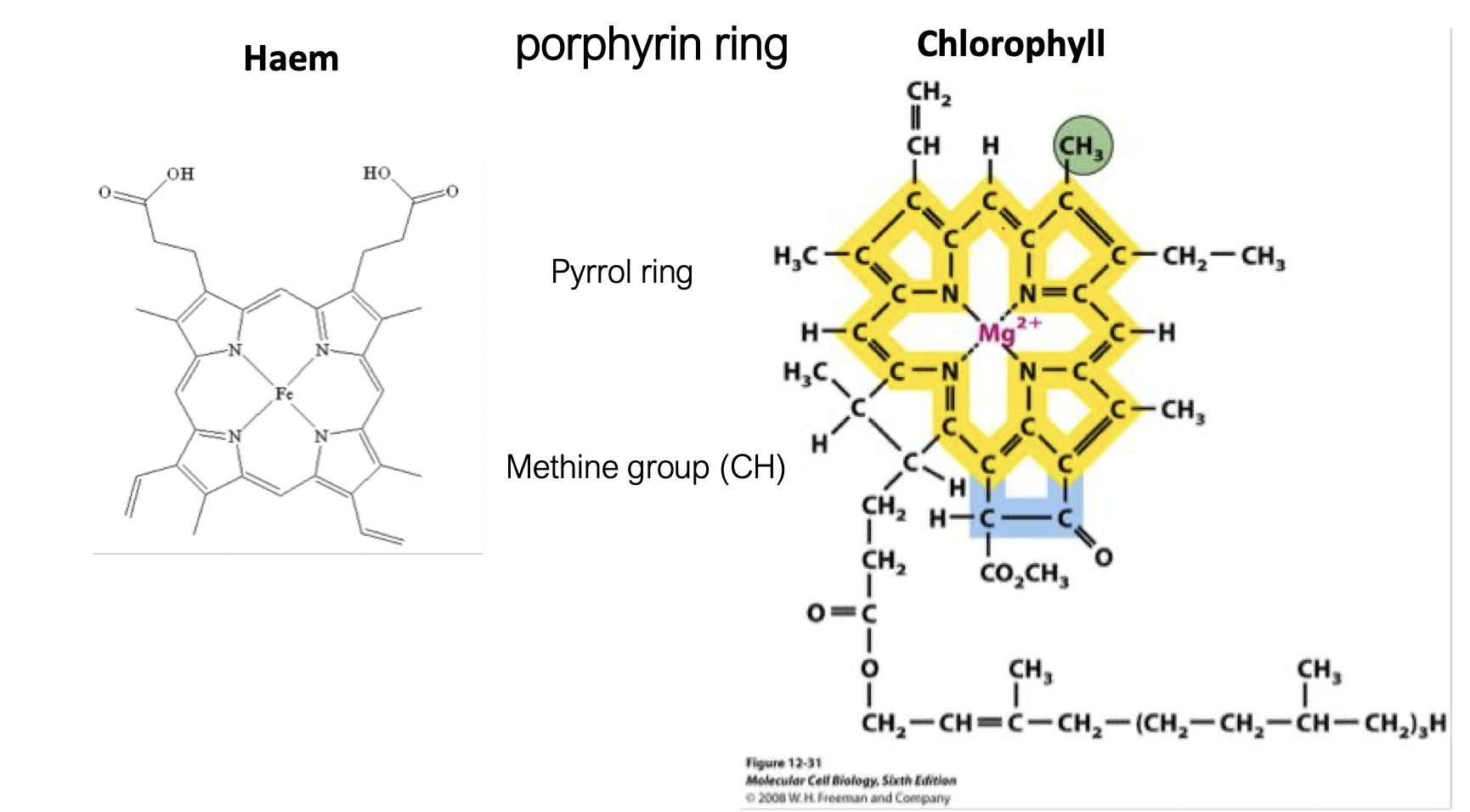
haem and chlorophyll have the a similar structure with the porphyrin ring but different central metals coordinated (haem has Fe2+ and chlorophyll has Mg2+)
what is the porphyrin ring
4 pyrrole rings (each has a nitrogen)
linked by methine bridged (-CH-)
to form a flat, conjugated ring
the nitrogen atoms can form a coordinate bond with a metal atom
central metal in haem
Fe2+
the iron can switch oxidation stated (between Fe2+ and Fe3+), which is good for its function for electron transfer
(primary function of haemoglobin is oxygen transfer but also sometimes involved in electron transfer, cytochromes are involved in electron transfer)
central metal in chlorophyll
Mg2+
prosthetic group
a non-peptide (protein) component tightly associated with a protein and required for its function
can be metals or organic
acts to support the structure of the protein
can also act as an electron carrier
haem = prosthetic group in haemoglobin
chlorophyll = prosthetic group in photosystems
why has the porphyrin ring been conserved evolutionarily
the conjugated ring has electron delocalisation which makes it stable and chemically versatile so evolution keeps it and saps the metal for different functions
key defining feature of a eukaryotic cell
presence of a membrane-bound nucleus and organelles
what replaces the nucleus in prokaryotes
the nuceloid (DNA not enclosed by a membrane)
how is DNA organised in prokaryotes vs eukaryotes
prokaryotes - circular, free in cytoplasm
eukaryotes - linear, enclosed in nucleus
what membranees are present in prokaryotic cells
inner (plasma0 membrane
cell wall
often an outer membrane (gram-negative bacteria)
periplasmic space between membranes - contains enzymes and transport proteins
why are prokaryotes more limited in complexity
lack of compartmentalisation - all reactions occur in the same space, less regulation of biochemical processes
size of prokaryotic cells
~1-5 micrometers
size of eukaryotic cells
~10-100 micrometers
plasma membrane
selective barrier controlling ion, nutrient and signal exchange
phospholipid bilayer with proteins embedded - transport via channels, carriers, pumps
mitochondria
ATP production via oxidative phosphorylation
lysosomes
degrade macromolecules and organelles
acidic lumen (pH of 4.50) and hydrolytic enzymes
nuclear envelope
double membrane separating nucleus from cytoplasm
contains nuclear pores for regulated transport
nucleolus
ribosome production
rRNA synthesis and ribosomal subunit assembly
nucelus
DNA storage and gene regulation
transcription occurs here - mRNA is exported out through pores in the nuclear membrane
smooth ER
lipid synthesis, detoxification, Ca2+ storage
enzymatic reactions in membrane
rER
protein synthesis for secretion/membranes
ribosomes translate proteins into ER lumen
Golgi apparatus
modify, sort and package proteins
glycosylation (enzyme-cata;ysed post-translational modification process of attaching carbohydrates to proteins or lipids) and vesicle trafficking
secretory vesicles
transport proteins to the membrane for secretion
fuse with the plasma membrane - exocytosis
peroxisomes
detoxification and fatty acid oxidation
produce and break down H2O2 via oxidases/catalase
cytoskeletal fibres
structural network for shape, transport, movement
microtubules, actin, intermediate filaments
microvilli
increase surface area for absorption
actin-based protrusions of membrane
cell wall
provides structural support and protection
made of cellulose in plants, peptidoglycan in bacteria and chitin in fungi
vacuole
storage and maintenance of turgor pressure
water influx keeps cell rigid
chloroplast
photosynthesis
light reactions (thylakoids) and Calvin cycle
which organelles are involved in the protein trafficking pathway
nucleus
rER
Golgi
vesicles
membrane.lysosome/secretion
which organelles are involved in detox
sER and peroxisomes
which organise generate energy
mitochondria - ATP
chloroplast - glucose (plants)
what is the fundamental purpose of cellular compartments?
to separate and control biochemical reactions within distinct environments
what defines a cellular compartment
a membrane bilayer enclosing a distinct internal environment
structure of a membrane bilayer
hydrophilic heads facing the aqueous environment
hydrophobic tails facing inwards
this forms a selective barrier
this si formed spontaneously due to the hydrophobic tails
why is compartmentalisation essential in eukaryotic cells
allows separation go incompatible reactions, increased efficiency, independent regulation
do all organelles ahem identical membranes
no, each organelle has a distinct lipid composition and specific proteins
why do membranes differ between organelles
membrane composition is tailored to function
for example mitochondria membrane contains ETC proteins, lysosome membrane is acid resistant and contains proton pumps
what determines membrane permeability
molecule size
polarity
charge
which molecule diffuse freely across membranes
small non-polar molecules (O2, N2, CO2)
which molecule are slightly permeable across membranes
small uncharged polar molecules (e.g. H2O, urea)
which molecules cannot cross without help
ions (K+, Ca2+, Cl-, etc.)
large polar molecules (e.g. glucose)
charged molecules (e.g. ATP, amino acids)
why are ions impermeable to membranes
hydrophobic core repels charged species
how do impermeable molecules cross membranes
via membrane proteins: channels, carriers, pumps
integral membrane proteins
embedded within the bilayer (often span membrane)
peripheral membrane proteins
loosely attached to membrane surface
lipid-anchored proteins
covalently attached to lipids within membrane
why is selective permeability critical
maintains ion gradients, metabolic control, cell signalling
how do membranes enable biochemical specialisation
by localising enzymes, creating gradients, controlling substrate access
significance of different head groups and tails within phospholipids
alters packing, shape and length
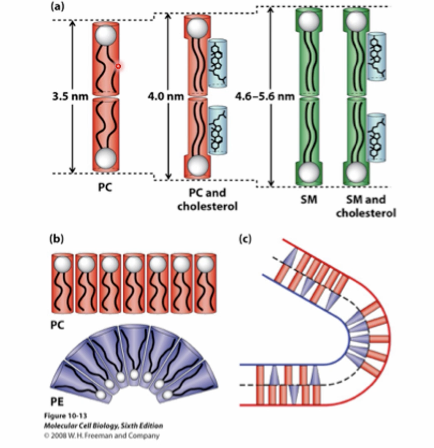
three main classes of membrane lipids
phosphoglycerides (phospholipids)
sphingolipids
sterols
basic structure of a phosphoglyceride
glycerol backbone, 2 fatty acid tail, phosphate group, variable head group
what distinguishes sphingolipids from phosphoglycerides structurally
sphingolipids have a sphingosine backbone instead of glycerol and usually one fatty acid chain attached via an amide bond
sterol and general structure
a lipid with a rigid four-ring structure that inserts between phospholipids in membranes
sterol found in animal membranes
cholesterol
sterol found in plant membranes
stigmasterol
sterol found in fungal membranes
ergosterol
head group attached fro phosphate of phospholipid
determines the identity of the phospholipid
common phospholipid head groups
PC: phosphatidylcholine
PE: phosphatidylethanolamine
PS: phosphatidylserine
PI: phosphatidylinositol
how do different head groups affect membrane properties
they alter charge, size and hydrogen bonding
this affects membrane curvature, interactions and protein binding
how does cholesterol affect membrane fluidity
buffers fluidity:
decreases fluidity at high temperatures
prevents tight packing at low temperatures - increases fluidity
cholesterol … membrane fluidity at low temperatures
increases
cholesterol … membrane fluidity at high temperatures
decreases
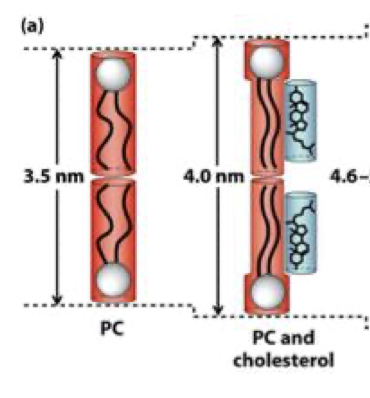
why do sphingolipids make membranes thicker
longer, more saturated hydrocarbon chains - tighter packing
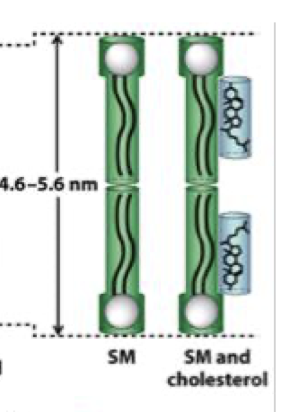
membrane thickness in presence of cholesterol
more ordered and slightly thicker
why does PE promote curvature
small head group - cone-shaped - induces membrane bending
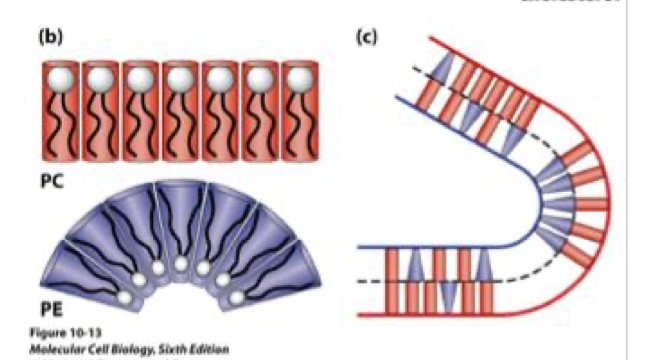
which lipid head promotes flat vs curved membranes
PC - cylindrical - flat bilayer
PE - cone-shaped - curvature
why is membrane curvature important
essential for vesicle formation, endo/exocytosis, organelle shape
how does lipid composition influence membrane function
controls fluidity, thickness, curvature and protein activity
bright-field (light) microscopy
light passes through the sample - contrast appears from the H&E staining
Haematoxylin
basic dye
binds acidic structure (DNA, RNA, acidic amino acids such as aspartate and glutamate) - blue/purple
Eosin
acidic dye
binds basic proteins (lysine, arginine) - pink
what information does H&E staining provide in tissues
reveals cellular and tissue structure, allowing distinction between norma vs pathological (e.g. cancerous) tissues
florescence microscopy
uses flourescently labeled moleculesthat bind specific cellular targets
why are cells fixed and permeabilised in flouresecne microscopy
fixation - preserves structure
permeabilisation - allows dyes/anitbodies to enter cell
what does the fluorescent probe phalloidin label
actin (microfilaments)
what does the fluorescent probe antibody label
tubulin (microtubules)
phalloidin dye colour
labelled with Rhodamine
red
antibody dye colour
green
confocal microscopy over conventional microscopy
uses optical sectioning to image only a single focal plane, eliminating out-of-focus light
why does confocal microscopy produce sharper images
excludes out-of-focus light, improving resolution and 3D reconstruction
compare bright-field, fluorescence and confocal microscopy
bright-field - general structure (low specificity)
fluorescence - specific molecules
confocal - high-resolution and depth (3D imaging)
why can’t electron microscopy be used on living cells
electron beams damage biological molecules (break bonds, generate free radicals) and require a vacuum
why does electron microscopy have higher resolution than light microscopy
electrons have a much shorter wavelength than visible light - higher resolving power
why are samples coated with heavy metals in electron microscopy
to increase electron scattering (contrast) and reduce damage, but it can reduce resolution/detail
how does cryo-EM reduce damage and improve imaging
rapid freezing preserves structure and reduces radiation damage (~6 fold) allowing near-atomic resolution (~0.2-0.3nm)
how does TEM (transmission) work and what does it show
electrons pass through thin specimen
shows internal ultrasound (2D image)
how does SEM (scanning) work and what does it show
electron beam scans surface, detects scattered electrons
shows 3D surface topography
key difference between TEM and SEM
TEM - internal structure - 2D
SEM - surface structure - 3D
why is cryo-EM a major advancement over conventional EM
preserves native structure without heavy metal staining and achieves near-atomic resolution
compare light vs electron microscopy
light - lower resolution, can image living cells
electron - very high resolution but requires dead, fixed samples