Bio 320 - Exam 3
1/123
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
124 Terms
mutation
Permanent change in DNA that is inherited due to lack of repair
point mutations
Alter one base
Two types of point mutations
transition (ex. pyrimidine for a pyrimidine) or transversion (ex. pyrimidine for a purine)
silent mutation
point mutation that has no effect on AA
missense mutation
point mutation that changes AA identity (if AA is essential for function, could kill cell)
nonsense mutation
point mutation that changes AA to a stop codon
indels
insertions and deletions that add or remove bases, result in frameshifts if one or two bases are changed
DNA lesions
changes in the DNA sequence
What are some large DNA mutations that can change chromosome structure?
Large deletion, duplication, inversion, or translocation events due to errors in chromosomal alignment during meiosis. Can lead to gene fusions (hybrid of two genes)
How can water cause mutations?
When a H dissociates from water, the reactive hydroxyl group (OH-) can cause spontaneous hydrolysis rxns or abasic sites which impact H-bond formation
spontaneous hydrolysis reaction
OH- from water attacks amino (NH2) group on base changing its identity. C→U base change most common. Reminder: U is foreign in DNA
abasic sites
OH- from water attacks bond between deoxyribose sugar and base, leaving an intact DNA backbone missing a base
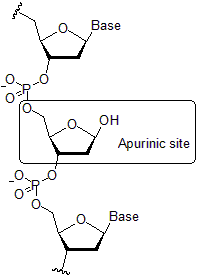
abasic site
How does oxidation cause mutations?
O and N containing compounds can cause oxidative damage by interrupting base pairing. Vitamins prevent this by being an antioxidant (ex. food preservatives can react with A and C residues)
Ames test
Uses mutant Salmonella that requires histidine in culture medium. Plate it on normal medium in the present of potential mutagen, if compound is carcinogenic bacteria experiences reversion mutations that allow them to grow without histidine
Why are mutagens used as cancer therapies?
When applying a mutagen, tumor cells that are rapidly dividing are most impacted whereas normal slow dividing cells are able to repair damage before dividing
What are the adverse side effects of chemotherapy due to?
Hair follicle, blood, and digestive tract epithelium cells naturally divide faster than normal cells, increasing their chances of being impacted by mutagens
How can ionizing radiation cause mutations?
UV light causes pyrimidine cross linkages (dimers) that creates kinks in strand where DNA pol stalls until repaired. Higher energy waves generate ROS that create single or double strand breaks
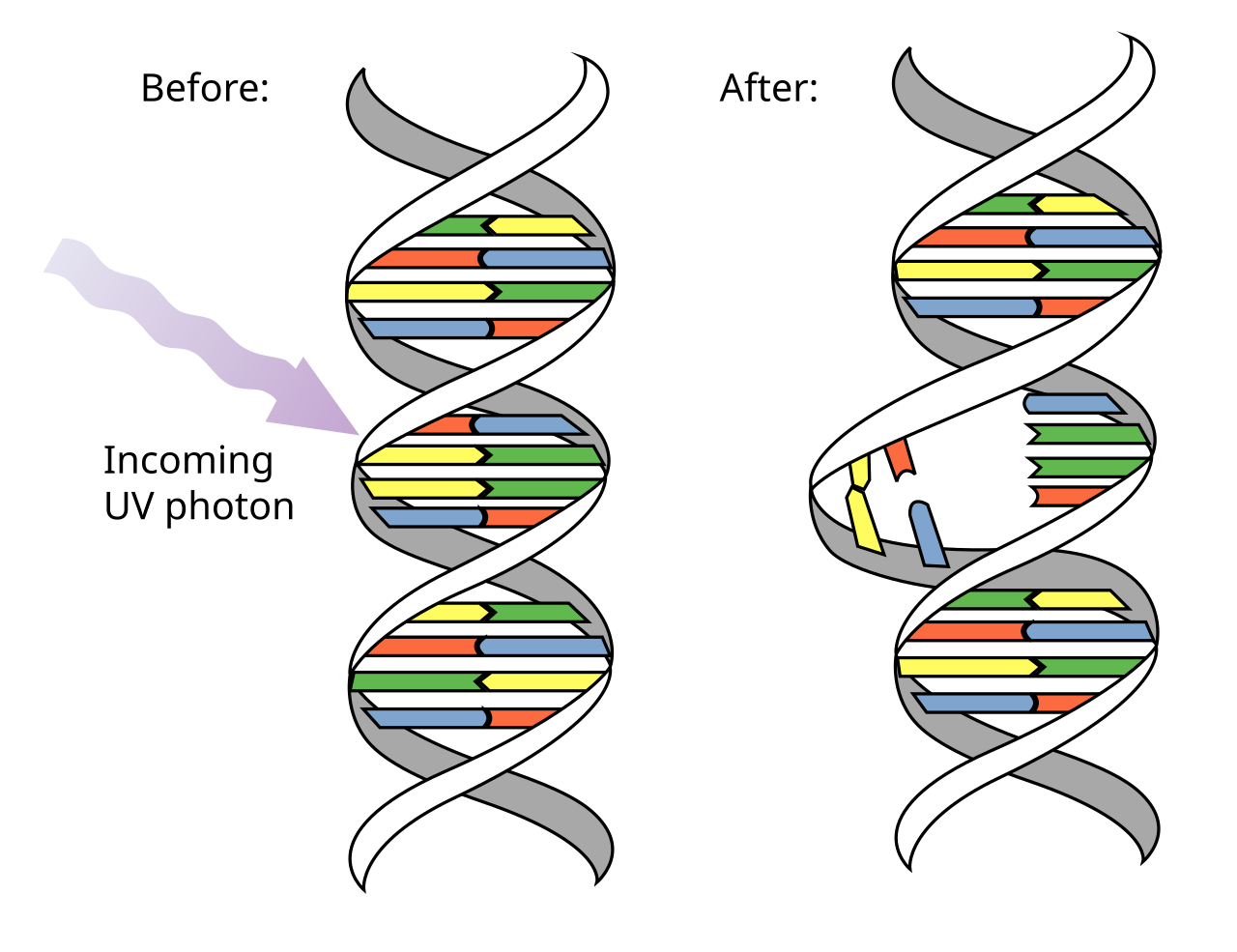
dimers
What are errors occur in DNA at random?
Tautomeric shifts (brief incorrect base pairing), indels due to pol slipping on repetitive base sequences, lower fidelity of polymerases not for replication, errors in homologous recombination due to repeated regions of homologs or non-homologs
What is the importance of DNA repair?
There’s high selective pressure for enzymes that maintain DNA integrity even if its incredibly energy intensive. Methods of repair in all cells with little divergence indicate its early origin
direct repair
repair error without removing DNA
indirect repair
must remove DNA to repair error
direct DNA mechanisms that repair in one step
photolyase repair of UV light damage and methyltransferases that remove improper methylated bases
photolyase DNA repair
In all cells except for placental mammals. Photolyase absorbs photon (energy from light), passes e- to excite FADH which then reduces pyrimidine dimer to reverse crosslinks, and e- goes back to regenerate FADH-
How are improperly methylates bases repaired?
Methyltransferases remove methyl group from improperly methylated bases and then are degraded
Mismatch repair system in prokaryotes
MutS/L protein complex recognizes DNA lesion in daughter strand and binds. Forms a loop to scan template DNA sequence for GATC site. MutH cuts the unmethylated GATC sequence, helicase binds to separate strands and exonuclease degrades incorrect DNA. DNA pol replaces bases in single strand gap, ligase seals nick
base excision repairs
DNA glycosylate recognizes damage and hydrolyzes bond b/t sugar and base. AP endonuclease makes a nick next to the abasic site. In prokaryotes pol I binds and ligase seals nick, in eukaryotes a short or long repair is made.
nucleotide excision repair for prokaryotes
Uvr proteins mediate removing DNA around large lesions. UvrAB recognizes lesion and separates strands, UvrC cuts phosphodiester bonds 5 bases downstream and 8 bases upstream of error. UvrD (helicase) binds to remove fragment, pol I fills gap, ligase seals nick
excinuclease activity
Displayed by UvrC, hydrolyzing more than one phosphodiester bond at once to remove a large fragment
nucleotide excision repair for eukaryotes
XP proteins mediate removing DNA around large lesions. XPC recognizes damage and XPBD (helicases) separate strands. XPF and XPG bind to create 5’ and 3’ nicks around a 24-32 fragment. DNA pol fills gap, ligase seals nick
transcription coupled repair
RNA polymerase can repair DNA if stalled in a region, is displaced by helicase, and repair proteins are recruited
Repairing DNA breaks
Breaks can disrupt replication fork and repair is required for division. Uses homologous recombination or non-homologous end joining
translesion synthesis
Allows DNA rep at the cost of integrity. If a lesion is present in ssDNA there’s not a template base for guidance. Some polymerases have ability to read across error to allow DNA replication which results in a mutation most of the time.
Importance of DSBs in homologous recombination
DSBs stimulate homologous recombination since it can repair single or double strand breaks as well as add diversity
How did homologous recombination evolve?
First as a DNA repair mechanism prior to euk. and prok. cell divergence. Evolution of diploidy allowed it to be used to generate diversity
Why might a double strand break occur?
If replication forks encounter a single strand break from a DNA lesion or exposure to radiation or something that breaks phosphodiester bonds
What is happening if there is a DNA lesion and repair has not begun?
Translesion synthesis may occur (recomb not needed), fork stalls until repair completes, replisome may be blocked and restart downstream of lesion
When is homologous recombination appropriate?
Either a second copy of the chromosome or sister chromatid is present from replication. When cell is in S or G2 phase
When is non-homologous end joining appropriate?
Cells are in G0 or G1 and most likely somatic. Occurs outside meiosis
How does homologous recombination repair DS breaks?
Nucleases degrade 5’ ends faster than 3’ ends to create overhangs, strands separated by helicase. Enzyme retrieves intact homologous chromosome. RecA allows strand invasion and D-loop formation, 3’ overhang from DS break replaces empty spot from D-loop to create Holliday junction. More RecA mediates branch migration and 3’ ends act as primers for DNA pol
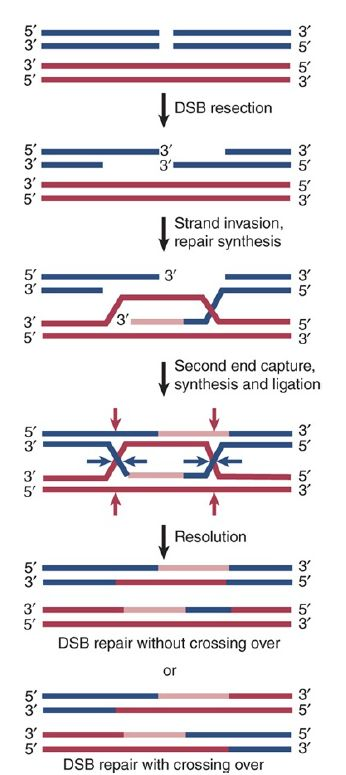
homologous recombination
How can chromosomes be separated after HR?
synthesis-dependent strand annealing or double strand break repair
Synthesis-dependent strand annealing
Helicases break H-bonds b/t invading strands so that they re-anneal to their original helix
Double strand break repair
Holliday junctions are separated by specialized resolvase enzymes (Ruv) in two ways depending on where H-junction is cut. If both outside strands are cut there is no crossover event (two strands still intact) but if one outside and one inside strand is cut there is crossover
How does HR generate eukaryotic diversity?
During meiosis I, specifically during Prophase I b/t maternal and paternal homologs. DSBs are at unpredictable locations and purposely carried out by Spo11 in yeast
Spo11 in eukaryotes
Initiates DSB for HR. Tyrosine residue on Spo11 enzyme breaks phosphodiester bonds and endonucleases create 3’ overhangs. RecA-equivalent proteins initiate DSB repair through HR
Non-homologous end joining
More common in eukaryotes, doesn’t conserve original DNA sequence. Ku proteins bind to DSB and attract repair proteins which hold broken ends together, overhangs are created, strands anneal, pol extends DNA, ligase seals nick
Can NSEJ anneal the ends of two different broken chromosomes?
Unlikely due to the tightly organized chromatin structure, but would be deleterious if it did
Why is learning about NHEJ and HR useful in biotechnology?
Genomic editing technology utilizes these to add/remove genetic info of interest
How are site-specific recombination and transposition similar?
They recombine non-homologous fragments and are mediated by enzymes related to topoisomerase that utilize phosphoryl transfers (no ATP required)
Site-specific recombination
Precise and predictable insertions, deletions, or inversions of DNA at specific regions. Utilizes recombinase, inverted repeats (binding sites), and core sequences (nonpalindromic)
Recombinase
Enzyme that binds at binding sites and cut at end of core sequence to mediate switching of two different cores
Steps to site-specific recombination
Active site of recombinase contains tyrosine or serine residue with -OH that breaks phosphodiester bonds, reattaches partner strands, rearranges, and repeats with next two strands
Uses of site-specific recombination
Insertion of viral genomes into host DNA, regulation of gene expression in bacteria to evade immune detection, in biotech to invert delete or insert sequences
transposition
The mov’t of a genetic sequence from a donor site to a target site within the same genome. Random of favors a specific site. Tightly regulated to prevent cell death
Why are transposons “molecular parasites”?
They insert themselves are are replicated passively as cell divides
What are the 3 transposition pathways?
cut-and-paste, replicative, or with an RNA intermediate
cut and paste method of transposition
Transposase creates DSB at 3’-OH of transposon, transposon is reinserted in the same or a different chromosome, DSB on empty and new site repaired
replicative method of transposition
Transposon partially cut & bonded to target sequence before excision, 3’-OH of transposon acts as primer for DNA rep to create 2 copies of transposon at each site
retrotransposition using an RNA intermediate
Transposon DNA is transcribed by RNA polymerase, RNA is reverse transcribed back to DNA and inserted into the target site
Why is retrotransposition possible?
The transposon DNA codes for a reverse transcriptase enzyme, and the transposon DNA never leaves its original site to be transferred
Transposase
Enzyme coded by the transposon that mediates its transfer using -OH group. Can cut phosphodiester bonds and create new bonds by attacking 3’-OH of target site DNA
Method of transposition most common in bacteria
cut and paste, categorized by types of genes present
Insertion sequences in bacteria
only contain the transposase
Composite transposons in bacteria
carry additional genes besides transposase (ex. abiotic resistance)
Complex transposons in bacteria
Carry many genes besides transposase that may assist transposition and other cellular processes (some are viruses)
Method of transposition most common in eukaryotes
Cut and paste (usually not active due to mutations/high disruption) and retrotransposition (90%). 46% of human genome consists of transposon sequences
retroviruses (ex. HIV) in eukaryotes
Viral RNA is converted into DNA using reverse transcriptase to be incorporated into host genome. Probably evolved from retrotransposons since rare events allow them to pass from one species to another and similarities in enzyme base sequences
Evolution of transposons
Retrotransposons gave rise to retrovirus, jumping transposons may be origins of introns in eukaryotes, transposases are eventually silenced by mutation or something else
Why can jumping/accumulating of transpositions lead to evolution?
Excision can become less precise causing gene duplication, new gene copies, and new functions. Poor excision=deletions, random insertion=disruption of other genes
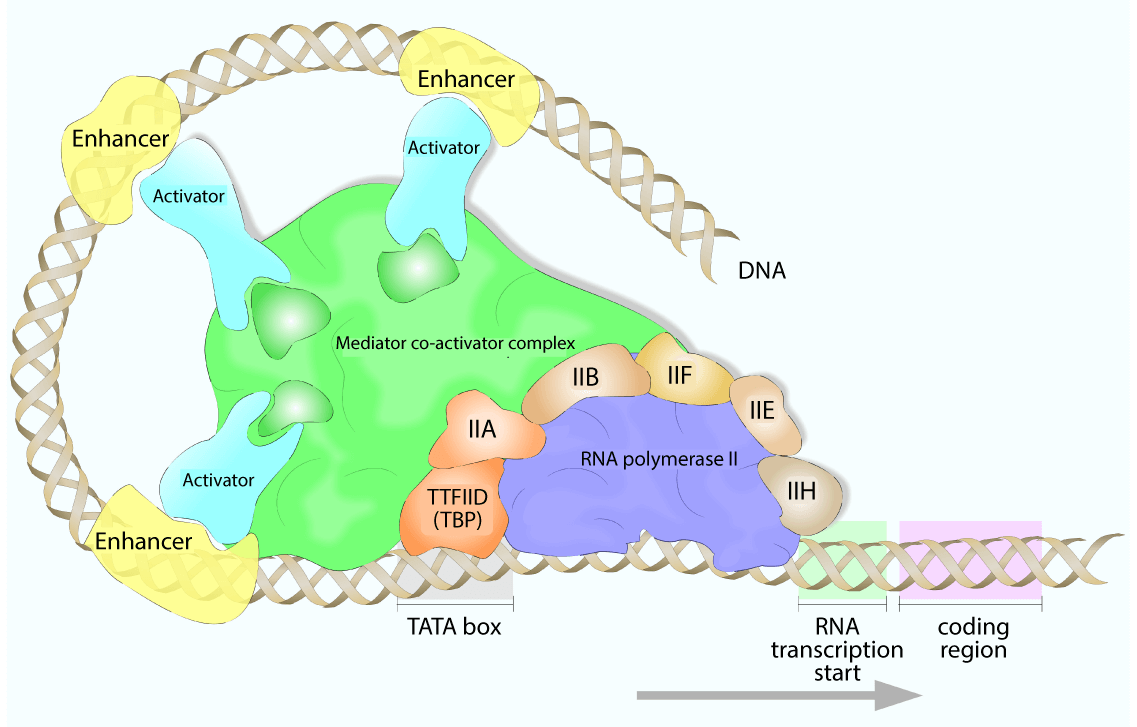
transcription mediator complex
transcription
RNA synthesis using a DNA template through initiation, elongation, and termination
RNA polymerase
Catalyzes the formation of phosphodiester bonds b/t rNTPs using Mg2+ cofactors
bacterial RNA polymerase
Made of 5 polypeptide subunits, a holoenzyme that associates with a σ cofactor
eukaryotic RNA polymerase
pol I (large rRNA precursors), pol II (most protein-coding RNA), pol III (small functional RNA like tRNA)
bacterial vs eukaryotic RNA polymerase
Structurally and functionally similar suggesting ancient origin, both create abortive transcripts
Euk pol. is larger with more proteins, must be phosphorylated, initiation requires >30 proteins
Similarities b/t RNA and DNA polymerase
Synthesize 5’→3’ antiparallel to template, require a template strand, has initiation elongation and termination steps
Differences in RNA polymerase from DNA polymerase
Doesn’t require a primer, only one DNA strand serves as a template, copies specific sections of DNA rather than the whole strand, processes UNTP instead of TNTP
General transcription steps
RNA pol binds to promoter
RNA pol creates open complex
First nucleotide is added to +1 site on template
Elongation (base pairing and phospho. bonds)
DNA reanneals, pol reaches terminator site
How does bacterial RNA pol bind?
Promoters contain -35 and -10 AT-rich regions that σ interacts with. UP element (third AT-rich region) is recognized by subunit to stabilize RNA pol binding
RNA polymerase openings
DNA entry, RNA exit, rNTP entry
pin
Structure in RNA pol that separates DNA strands
Bacterial transcription initiation
DNA enters closed RNA pol on the promoter, pin creates open complex. RNTPs bind template DNA strand, RNA pol forms bond
abortative initiation
There is low probability of interaction between template DNA and new rNTPs at 1+ start site, so pol may release transcript and restart until connection is stable
Bacterial transcription elongation
Starts once RNA fragment is >10bp long, RNA pol may proofread by stalling and moving backwards, 50-90 bases added per second
Bacterial p-independent transcription termination
Terminator hairpin loop forms in DNA and disrupts pol. structure to release RNA
Bacterial p-dependent transcription termination
Transcription of rut site (CA rich region) attracts a p helicase that moves 5’-3’ on the RNA to disrupt polymerase into releasing the RNA
Eukaryotic transcription core promoter
The TATA box 25-30bp upstream of +1 start site, requires binding of TATA-binding protein (TBP)
Eukaryotic transcription initiation
RNA pol and TFs bind core promoter, TFII proteins bind to their recognition sites to form closed complex, TFIIH (helicase) creates open complex in pol, RNA pol must be phosphorylated to disengage from promoter, TFs released
Why are multiple transcription factors required for eukaryotic transcription?
Chromatin structure often needs to change and genomes are larger
Why does having several pol. make eukaryotic transcription more complex?
Each pol. recognizes unique promoters and uses different transcription factors
multiprotein mediator comples
In eukaryotes, required for interaction b/t TFs and regulatory proteins
Eukaryotic transcription elongation
After 60-70 rNTPs and phosphorylation of pol, elongation factors bind and assist until termination
2 models for eukaryotic transcription termination
Hairpin formation (similar to p-independent in prok.) or torpedo model
Torpedo model of eukaryotic transcription termination
Suggests proteins cut transcript to disrupt RNA pol
Elongation factors in eukaryotic transcription
Bind to enhance RNA pol activity by suppressing pauses, enhance mismatch editing, and recruit proteins for post-transcriptional processing
RNA processing in bacteria
RNA from transcription not modified before use, RNA degradation normal and performed by RNases, transcription & translation coupled in cytoplasm
RNA processing in eukaryotes
RNA modification must occur to begin translation, not coupled, processing includes 5’ capping, 3’ poly-adenylation, splicing, degradation, editing
5’ capping (eukaryotes)
Guanylyl transferase (GTase) associates with pol II, caps each mRNA, links 7-methyl-guanosine (7-meG) to RNA by a 5’ triphosphate linkage. Protection, signals export out of nucleus, signals ribosomal binding. Occurs after first 20-30 bases form