Post Quiz 4 Content
1/41
Earn XP
Description and Tags
lec 45 - end
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
42 Terms
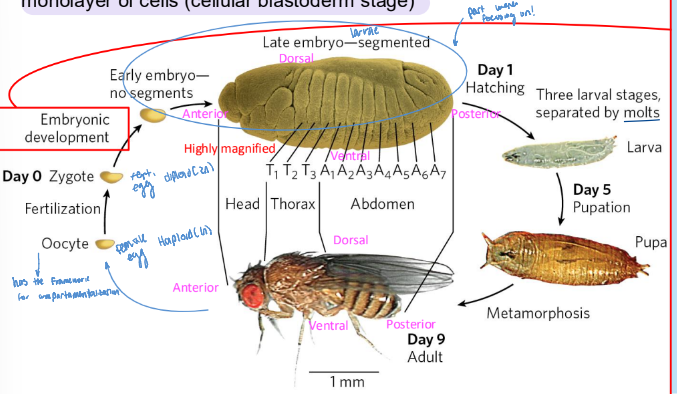
Drosophila developmental cycle
oocyte → zygote → early embryo- no segments → late embryo (segmented) → larval station → pupa → adult
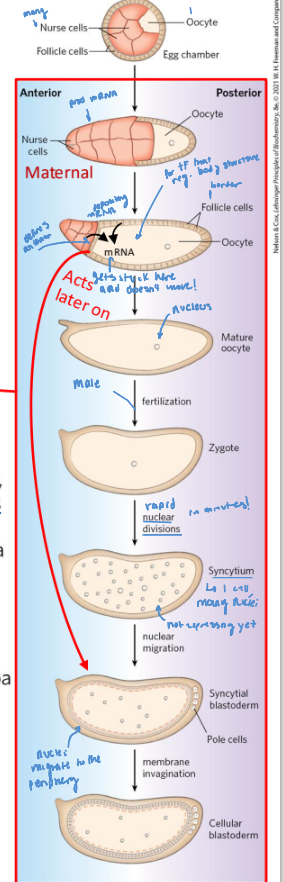
embryonic development stages
1 oocyte with many nurse cells
oocyte exbands, retrackting the nurse cells as follicle cells develop
mature oocyte can be fertilized (becomes a zygote)
rapid nuclear division in 1 cell
membrane begins to invaginate
forms a cellular blastoderm
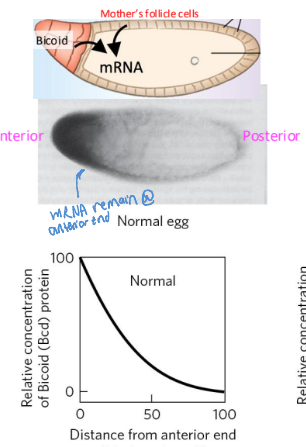
Bicoid - maternal gene product
biocoid mRNA is synthesized by nurse cells and deposited in the unfertilized egg near its anterior pole
after fertilization, biocoid mRNA is translated to make Bicoid, acting as an anterior TF
makes a concentration gradient from anterior to posterior
5 main TF cascades that drive cell fates
bicoid (anterior → posterior)
kruppel & hunchback (gap genes)
Eve & ftz (pair-rule gene)
Caudal (posterior → anterior)
Nanos (posterior → anterior)
both bicoid and nanos comes from the mother laying mRNA
the opossing TF gradients + cooperative binding sharpen transitions of gene expression!
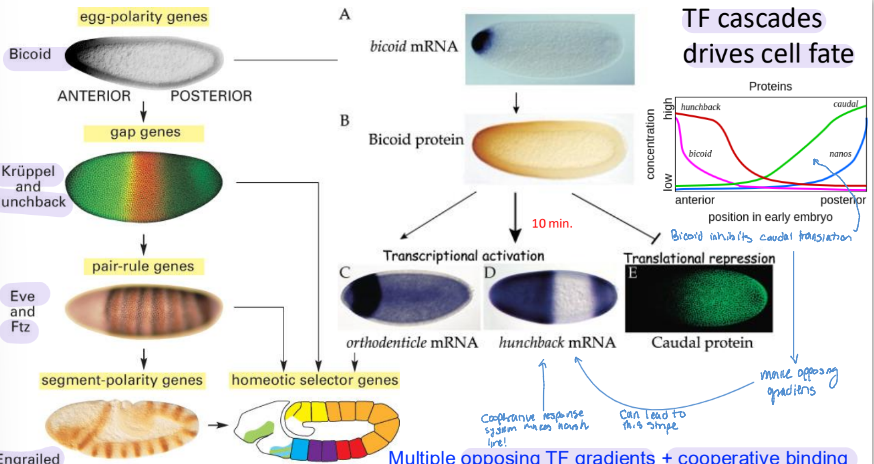
Segmentation Genes
direct the formation of the proper number of body segments
transcribed after fertilization
gap genes
divide the developing embryo into several b road regions
ex: kruppel and hunchback
pair-rule genes + segment polarity genes
define 14 stripes that become the 14 segments of an embryo
ex: Eve and Ftz
Homeotic genes
Specify which organs/appendages will develop in particular body segments
typically encode TFs that contain a homeodomain
encode Hox genes
Homeodomain
bind specific DNA enhancer sequences
Hox gene clusters
responsible for the development of structures in a defined part of the body
Drosophila have 1 Hox gene
humans have 4!
There is a lot of conservation between Hox and mammalian hox complexes!
stem cells
cells that can differentiate into various tissues
2 main functions!
replenishing themselves
providing cells that can differentiate
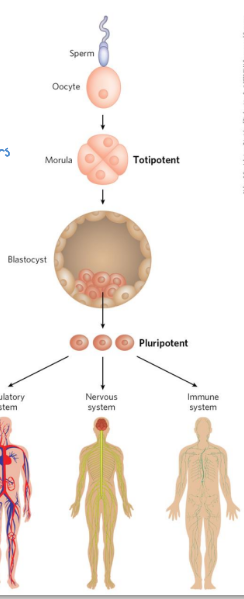
totipotent cells
cells that can differentiate into ANY TISSUE or a complete organism
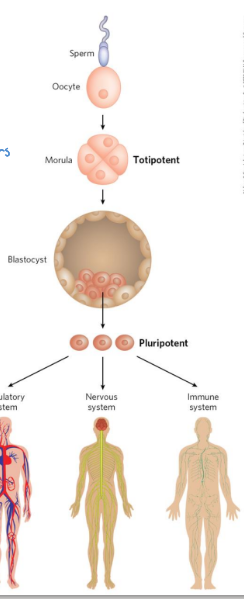
pluripotent cells
can give rise to cells of all 3 germ layers as well as many tissue types
cannot differentiate into a complete organism
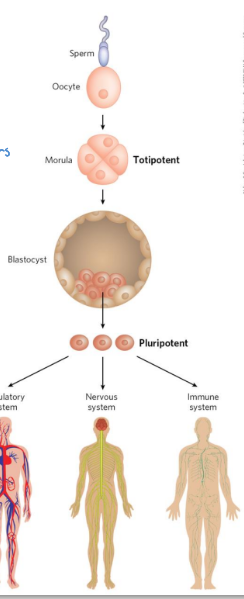
embryonic stem cells
pluripotent cells of the blastocyst
unipotent cells
can develop into only 1 type of cell/tissue
adult stem cells
more limited potential than embryonic stem cells
considered multipotent
have a niche: microenvironment that promotes stem cell maintenance while allowing differentiation of some daughter cells
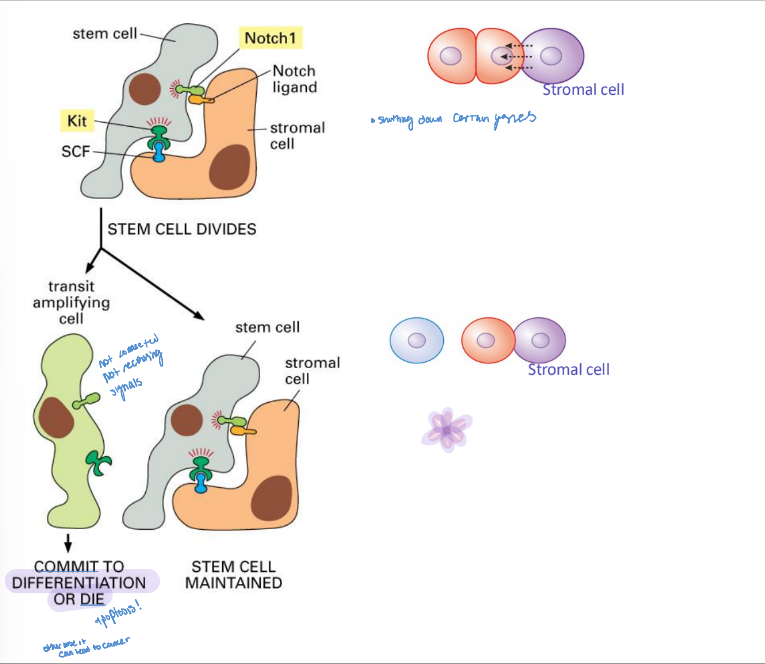
niche
microenvironment that promotes stem cell maintenance while allowing differentiation of some daughter cells
receives signals from neighboring cells to maintain the stem cell lineage
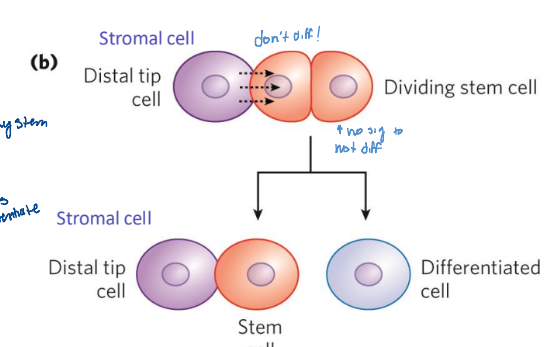
signaling of neighboring cells
can cause a cell to commit to differentiation or die through apoptosis
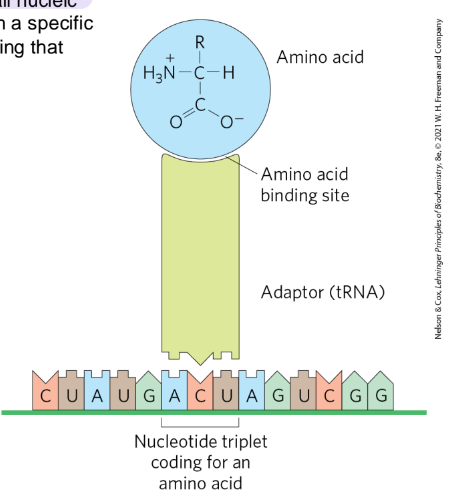
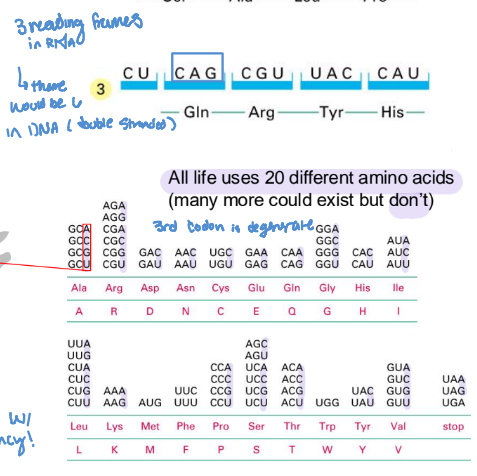
codon
triplet of nucleotides that codes for a specific amino acid with specificity
How are codons degenerate?
There are 4 code letters and 3 spots within a codon (4³=64) however there are only 20 AA
multiple different codes encode for 1 AA however, the 1 code NEVER codes for multiple AAs.
specificity of reading frames
open reading frames have a start codon that have the longest sequence without a stop codon.
evolved to remove sequences that could prematurely stop UNLESS there is a frameshift
then it would likely encounter a premature stop codon, forming a truncated protein
How were AA codon codes found?
poly(u) and 20 radioactive amino acids were fed to E.Coli and it found that UUU codes for Phe
same approach with CCC for proline, AAA for lysine
The 3 termination codons
UAA, UAG, UGA
also called nonsense codons
do not code for any AA
Open reading Frame
starts with AUG (Met), initiation codon, followed by at least 50 codons, and ends with a stop codon
anticodon
A 3-base sequence on the tRNA that base pairs with mRNA
pairing occurs via hydrogen bonding
it is an antiparallel alignment
Only 32 tRNAs (not 61) are needed, due to the wobble effect
wobble effect
the third codon is a degenerate psotition
much weaker H-bonding
if there is a mutation at the 3rd position, mutations are likely to be silent (encode for the same AA)
Resiliency in code mutation
Mutations of a codon usually produce a conservative substitution
ex: valine → alanine
both are nonpolar!
missense vs. nonsense
missence - mutations that change the encoded amino acid
nonsense - mutations that make a stop codon
Indels
insertion or deletion. Only indels with a length the multiple of 3 will maintain the reading frame.
when out of frame, a stop codon is likely to be encountered
encoded translational frameshifting
aka “hiccupping” of ribosomes
allows 2+ related but distinct proteins to be produced from 1 transcript
occurs during translational frameshifting
2+ related but distinct proteins can be produced from 1 transcript
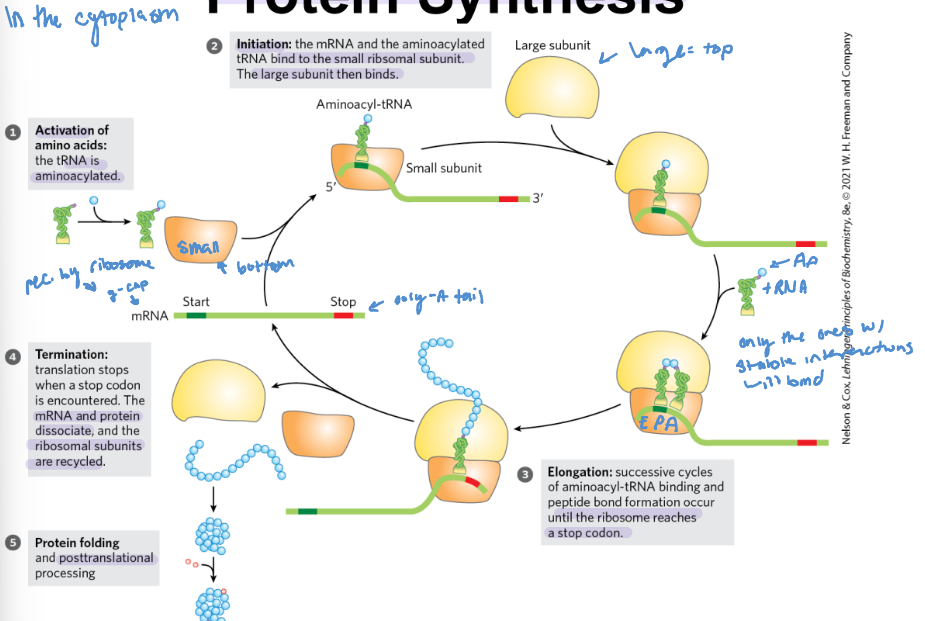
Five stages of protein synthesis
1) activation of amino acids (when tRNA is aminacylated)
2)Initiation: the mRNA and the aminoacylated tRNA bind to the small ribosmal subunit, then the large ribosomal subunit binds
3) Elongation: successive cycles of aminoacy-tRNA binding and peptide bond formation occur until the ribosome reaches a stop codon
4) Termination: translation stops when a stop codon is encountered. The mRNA and protein dissociate, and the ribosomal subunits are recycled
5) Protein folding: posttranslational processing
essential components for the activation of amino acids in E. Coli
all 20 AA, 20 aminoacyl-tRNA synthases and 32 or more tRNAs
ATP and Mg2+
Step1 for AA activation
tRNAs are charged when attaching their amino acid (aminoacylated)
aminoacyl-tRNA synthetases esterify the 20 AA to their corresponding tRNAs
each is specific for 1 amino acid and 1 or more corresponding tRNAs
occurs in the cytosol
ATP activates the carboxyl group of each AA
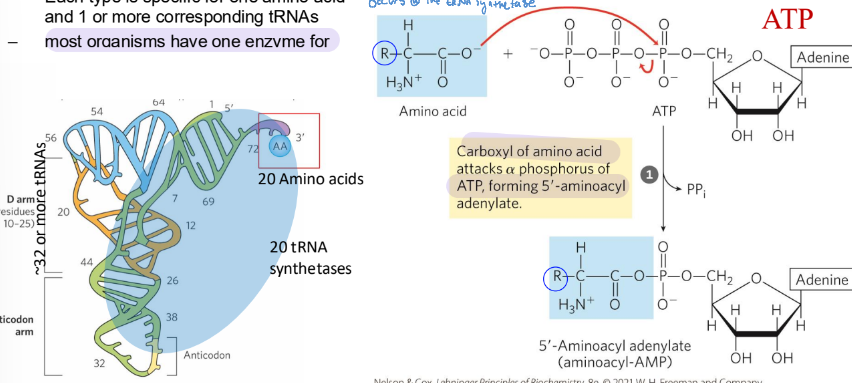
Step 2 for aminoacyl-tRNA synthetases
attach the correct amin oacid to their tRNA as defined by their anticodon
transfer the aminoacyl group from teh enzyme0bound aminoacyl-AMP to the corresponding specific tRNA
The aminoacyl group is esterified to the 3’ position of the terminal A m=nucletodide of the tRNA
the ester linkage activates the AA and joins it to the tRNA
More on Aminoacyl-tRNA
aminoacyl-tRNA synthetases must match
Essential components for protein synthesis initiation in E. Coli
mRNA
fMet in prokaryotes
Initiationcodon (AUG)
50S ribosomal subunit
Initiation factors (IF1,2,3)
GTP
Mg2+
Essential components for protein synthesis elongation in E. Coli
functional 70S ribosomes (initiation complex)
aminoacyl0tRNAs specified by codons
elongation factors (EF-Tu, EF-Ts, EF-G)
GTP
Mg2+
Essential components of termination and ribosome recycling in E. Coli
termination codon in mRNA
Release factors (RF1,2,3, RRF)
EF-G
IF3
Folding and posttranslational processing
chaperones and folding enzymes (PPI, PDI)
specific enzymes, cofactors, and other components for the removal of initiating residues and signal sequences
modifications of terminal residues
attachments of acetyl, phosphoryl, methyl, carboxyl, carbohydrate, or prosthetic groups
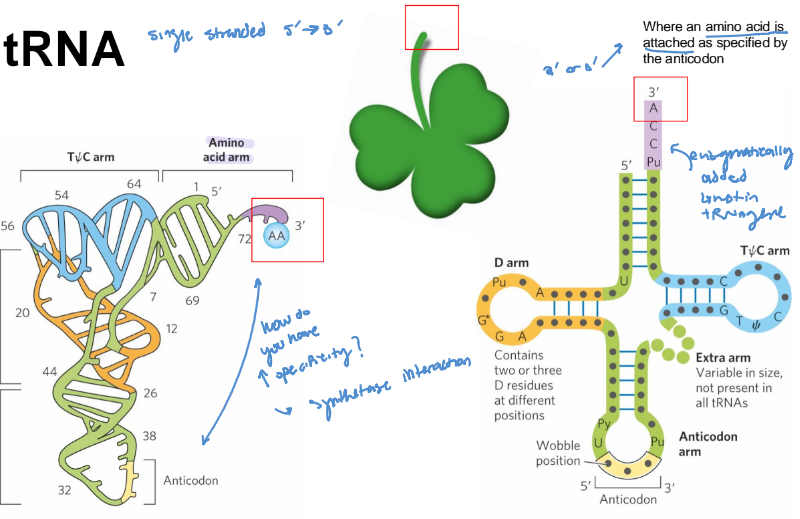
tRNA structure
AA arm - carries a specific amino acid esterified by its carboxyl group to the 2’-OH or 3’-OH group of the A residue at the 3’ end of the tRNA
emphasis on esterified AA!
anticodon arm
contains the anticodon
Crick’s adaptor hypothesis
A small nucleic acid could act as an adaptor, binding to both a specific AA and the mRNA econding that AA
amino acids are activated for protein synthesis
aminoacyl-tRNAs: tRNA attached to an amino aicd
aminoacyl-tRNA synthetases
catalyzed the formation of aminoacyl-tRNAs