DATA analysis for exam 3 bio 406
1/22
Earn XP
Description and Tags
chapter 11 ,12, 13
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
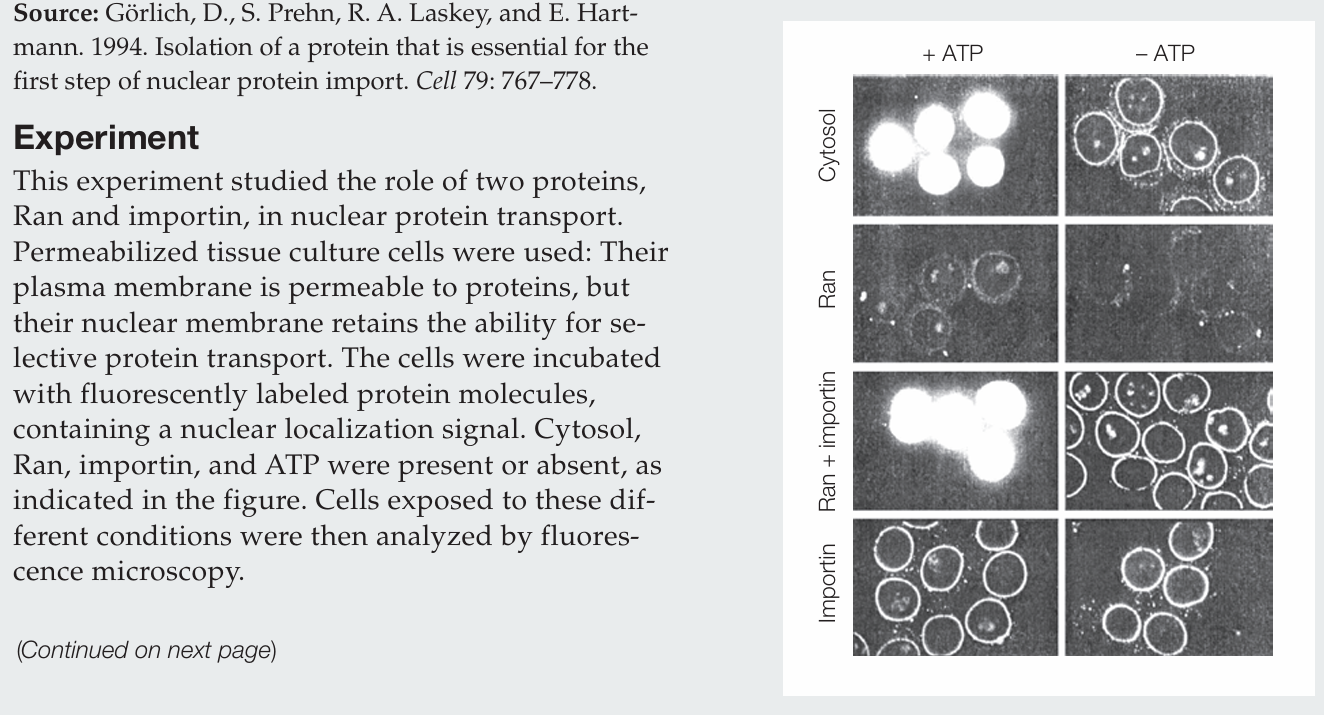
What actions does Ran perform by itself?
Ran protein alone has no activity.
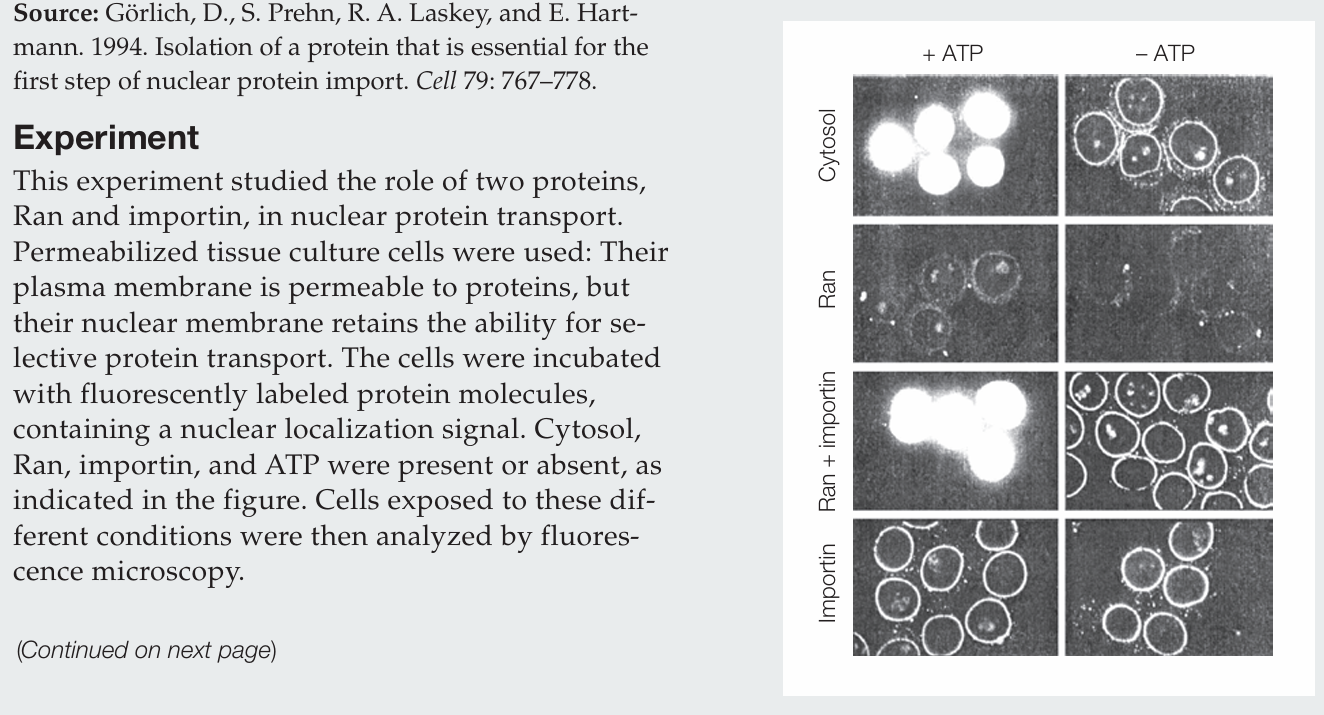
What actions does importin perform by itself?
Importin alone allows labeled protein to accumulate at the nuclear membrane, evidenced by the fluorescence around the nuclei, and it does so whether ATP is present or absent.
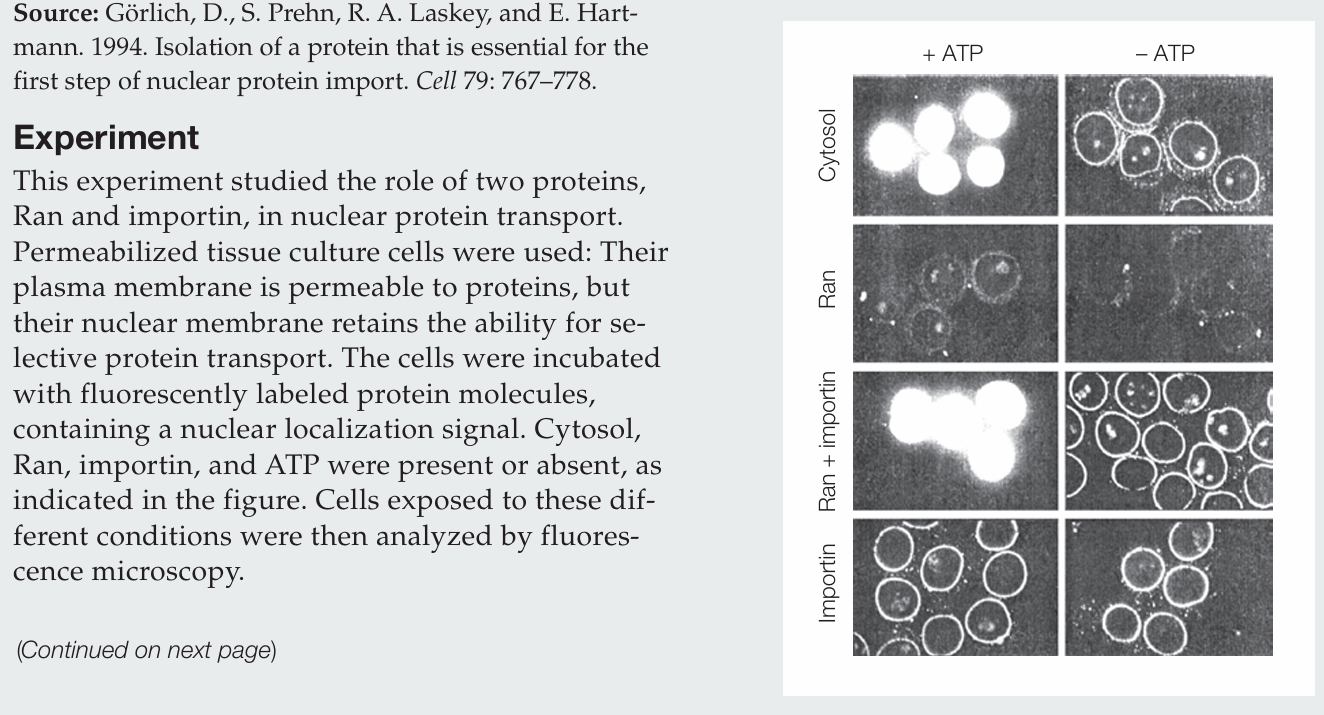
How do Ran and importin act together in the nuclear import of protein?
Ran and importin together mediate nuclear import only in the presence of ATP. Importin brings proteins to the nuclear envelope, and Ran drives their transport into the nucleus. In the absence of ATP, proteins accumulate at the nuclear envelope.
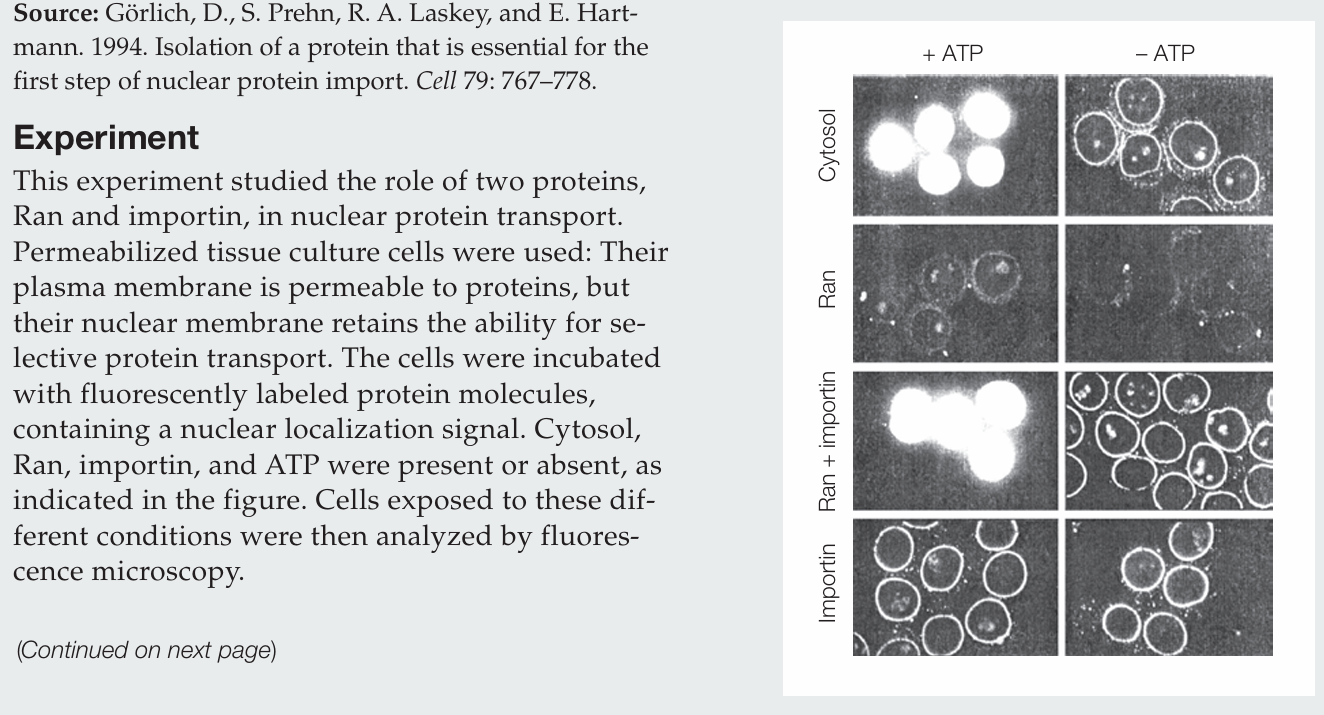
Why is ATP required?
ATP is required to generate GTP, which establishes the Ran GTP/GDP gradient that drives nuclear import.
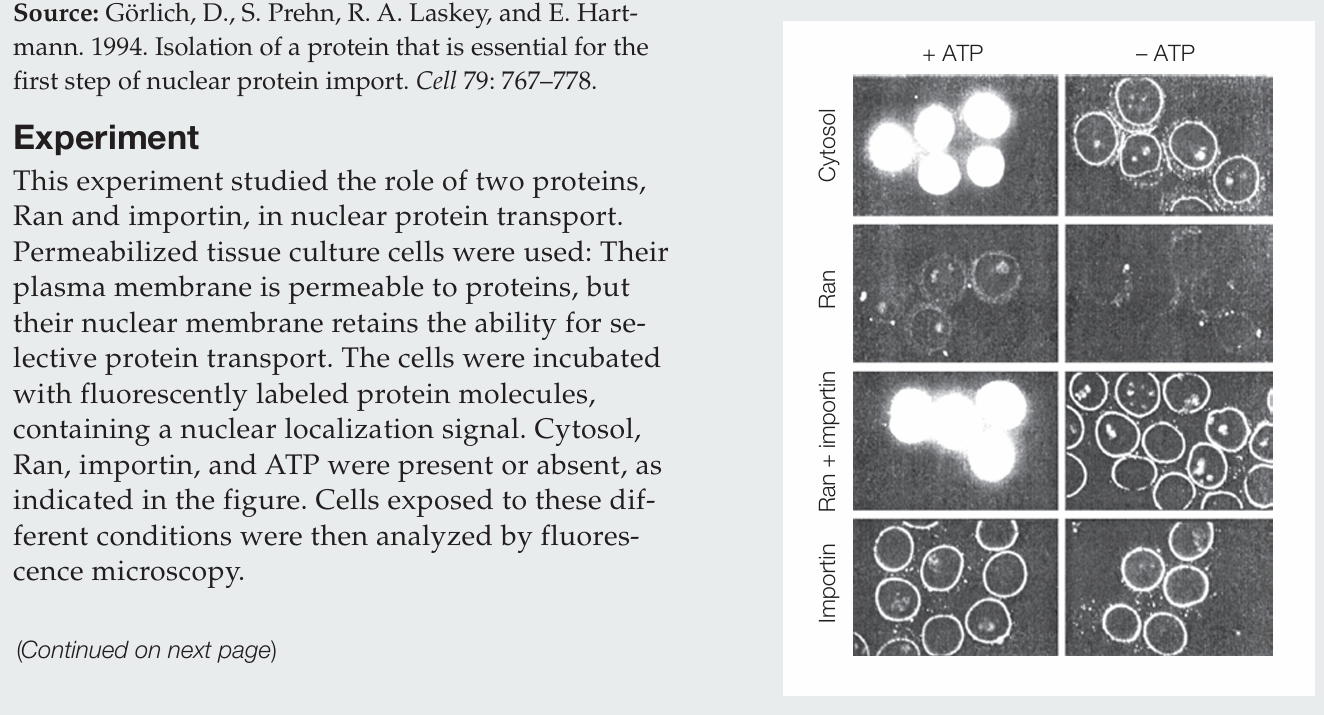
What conclusion can be drawn regarding the cytosol sample?
The cytosol contains the necessary proteins for nuclear transport, including Ran and importin, since labeled proteins are imported into the nucleus in the presence of ATP.
![<p>Why was [α-32P]GTP used in this experiment?</p><p></p>](https://assets.knowt.com/user-attachments/3dbca85e-5581-4dc4-8a7c-228193387b41.png)
Why was [α-32P]GTP used in this experiment?
[α-32P]GTP was used to label newly synthesized RNA molecules.
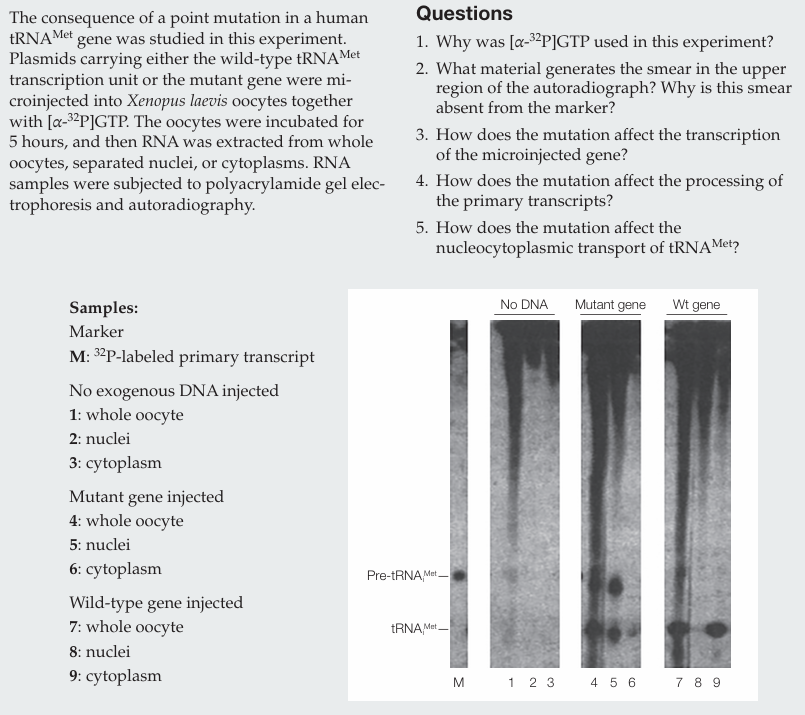
What material generates the smear in the upper region of the autoradiograph? Why is this smear absent from the marker?
The smear is generated by newly synthesized RNAs (mRNAs and rRNAs) of many different sizes. It is absent from the marker because the marker contains only labeled pre-tRNAᵐᵉᵗ.
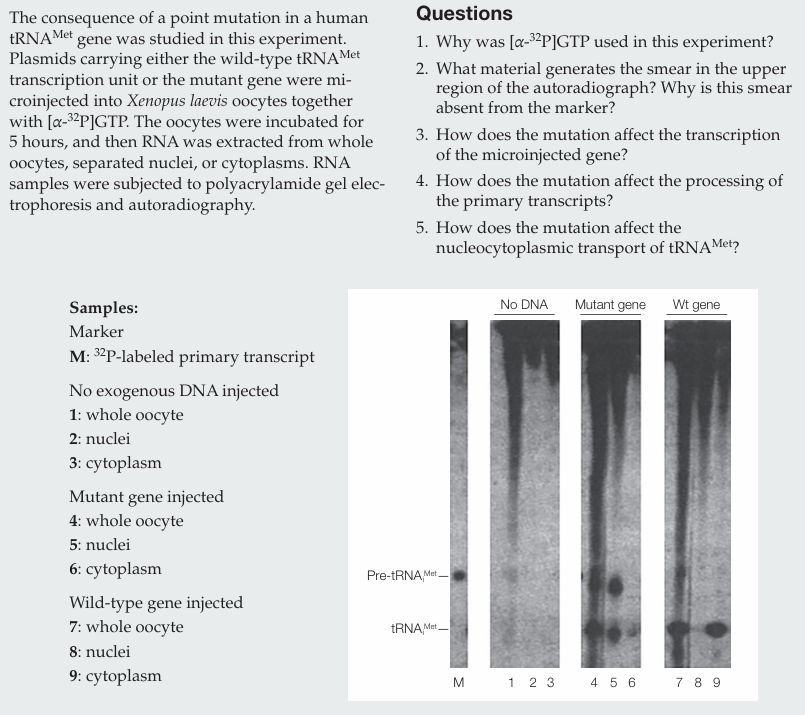
How does the mutation affect the transcription of the microinjected gene?
The mutation does not affect transcription, as similar amounts of RNA are produced.
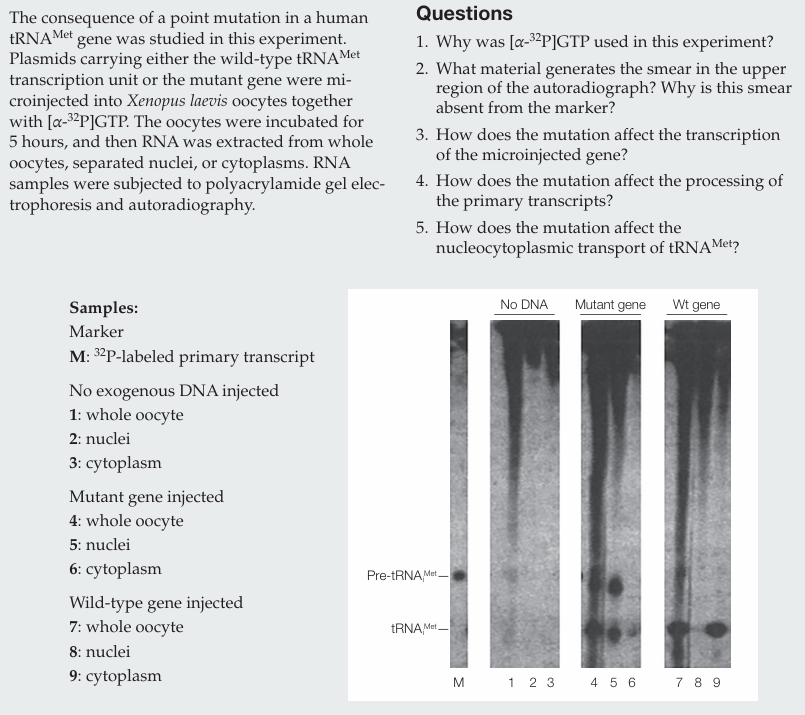
How does the mutation affect the processing of the primary transcripts?
The mutation does not affect processing, since mature tRNAᵐᵉᵗ is present.
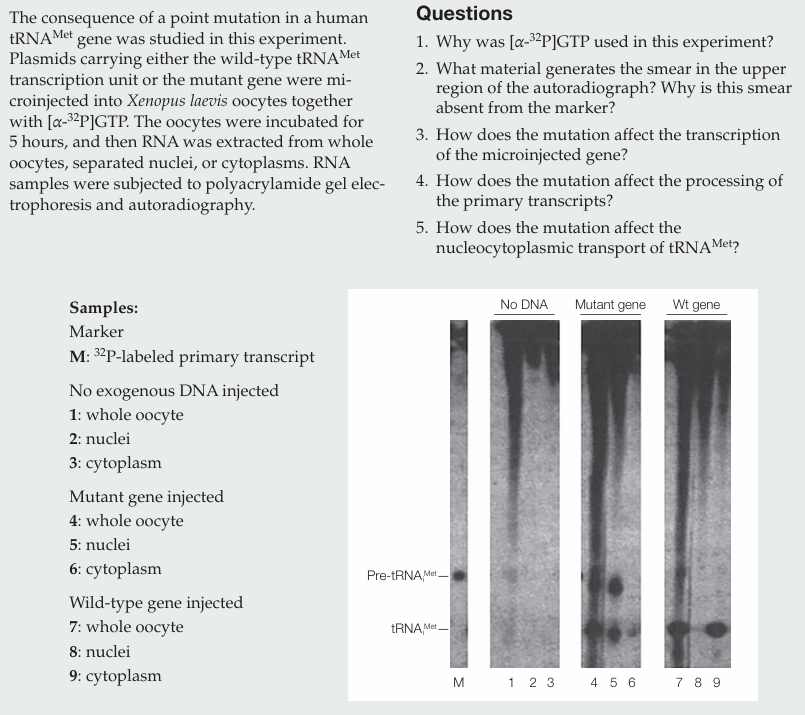
How does the mutation affect the nucleocytoplasmic transport of tRNAᵐᵉᵗ?
The mutation impairs nucleocytoplasmic transport, as less mature tRNAᵐᵉᵗ is found in the cytoplasm compared to the wild type.

Explain how the bands pPM and PM were resolved and suggest a possible relationship between them?
Proteins were separated by SDS-PAGE based on size, with pPM having a higher molecular weight than PM. This indicates that pPM is a precursor that is processed into PM by cleavage of an N-terminal signal peptide by signal peptidase at the rough ER membrane.

What conclusion can be drawn from comparing samples 1 and 2?
The presence of rough microsomes allows conversion of pPM to PM, indicating that the protein is translocated into the rough ER where its signal peptide is cleaved by signal peptidase. This process occurs via the cotranslational pathway, in which the signal recognition particle (SRP) directs the ribosome–nascent chain complex to the ER membrane for translocation through a translocon.

What conclusion can be drawn from comparing samples 1 and 3?
The absence of PM when translation is inhibited shows that translocation into the ER occurs co-translationally.

What conclusion can be drawn from comparing samples 1 and 4?
Inhibition of the SRP receptor prevents formation of PM, demonstrating that ER targeting requires the SRP–SRP receptor pathway.

What conclusion can be drawn from comparing samples 4 and 5?
Protein degradation by proteinase K indicates the protein remained outside the microsomes, confirming that translocation failed when SRP function was inhibited.
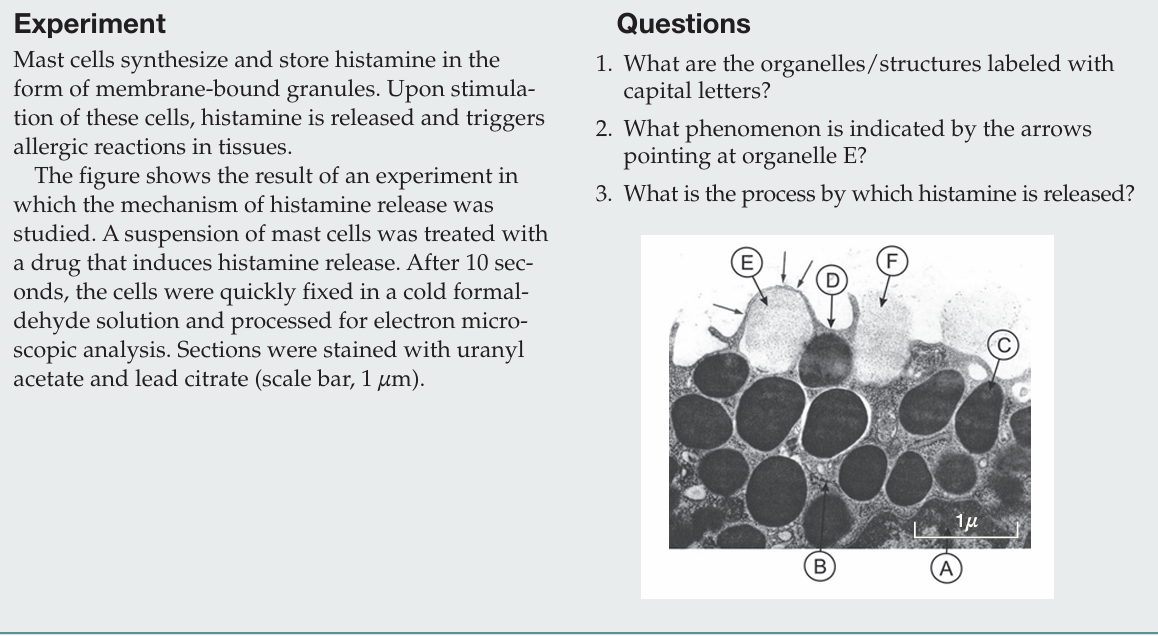
What are the organelles/structures labeled with capital letters?
A: heterochromatin in the nucleus;
B: rough endoplasmic reticulum;
C: secretory granule;
D: plasma membrane;
E: secretory granule undergoing exocytosis;
F: degranulated vesicle.
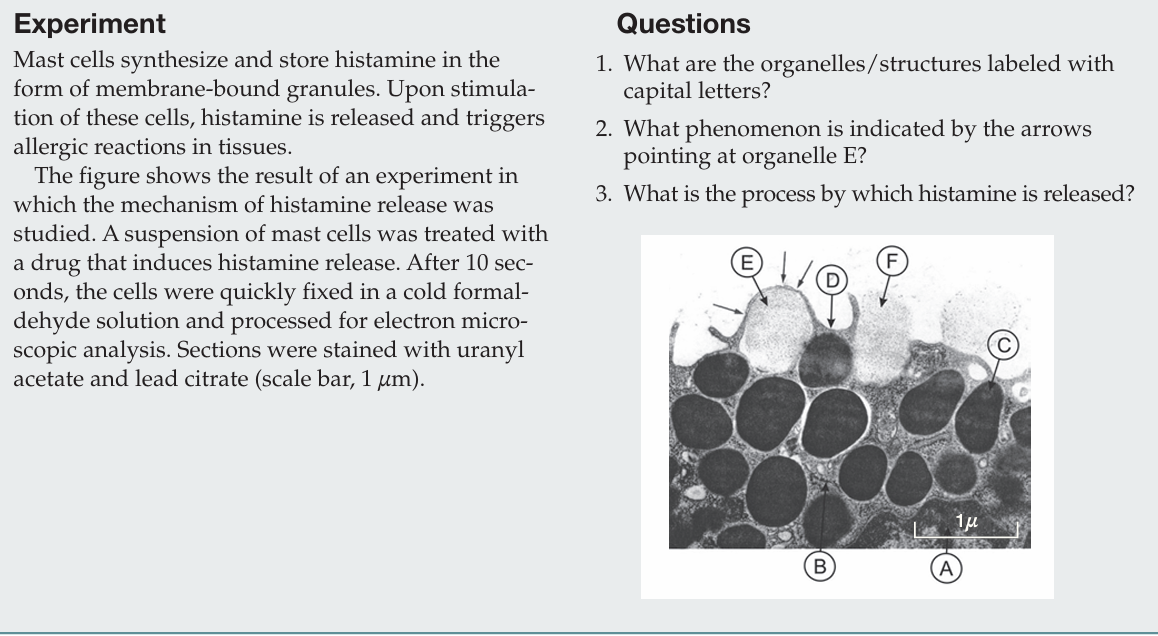
What phenomenon is indicated by the arrows pointing at organelle E?
The arrows indicate membrane fusion between a secretory vesicle and the plasma membrane.
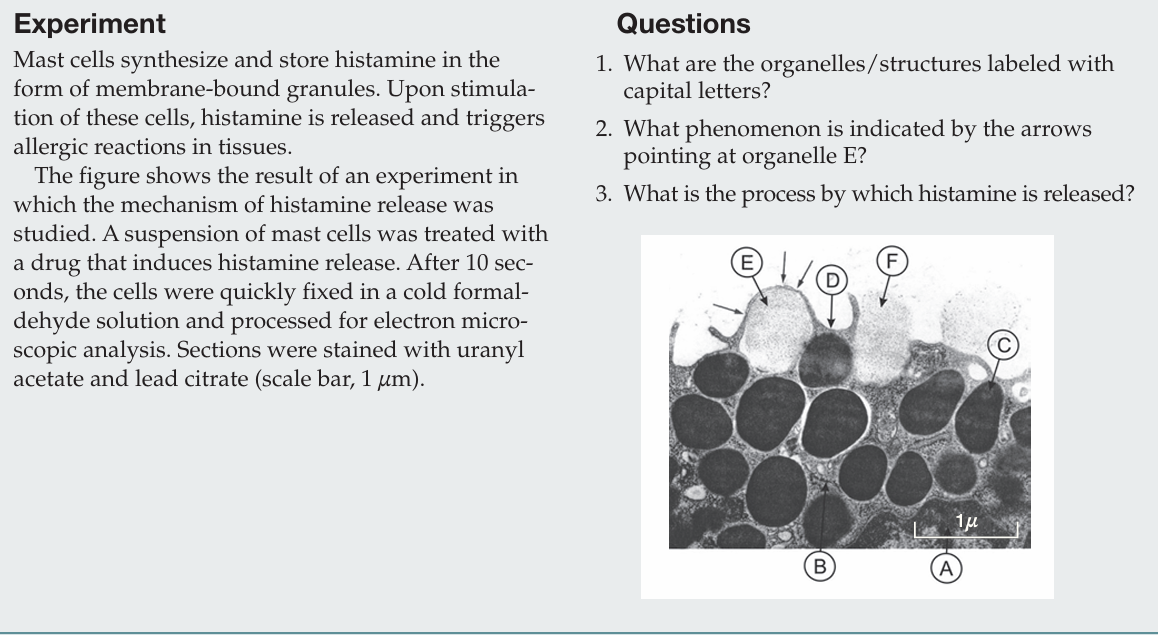
What is the process by which histamine is released?
Histamine is released by exocytosis, in which secretory granules fuse with the plasma membrane and release their contents from the cell.
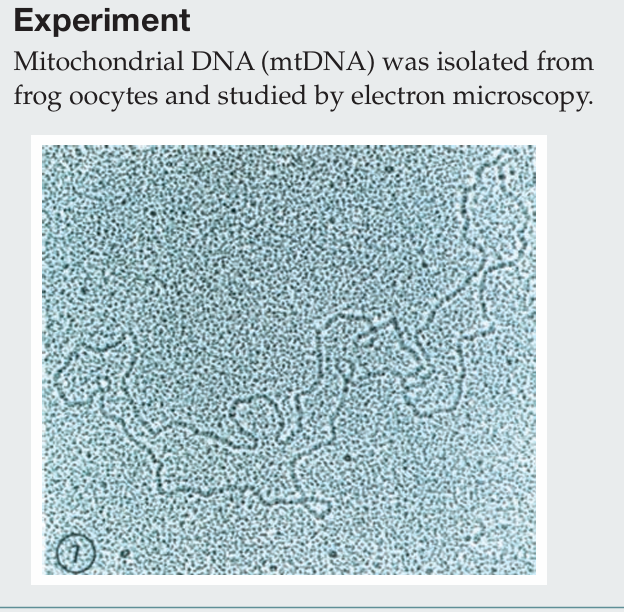
Briefly describe the procedure for isolating mitochondria.
Cells are homogenized and subjected to differential centrifugation. A low-speed spin pellets nuclei, and the supernatant is then centrifuged at a higher speed to pellet mitochondria. The mitochondrial fraction can be further purified using density gradient centrifugation. Mitochondria are then lysed, and mtDNA is isolated through organic extraction, ethanol precipitation, RNase treatment, and final purification by CsCl gradient centrifugation.
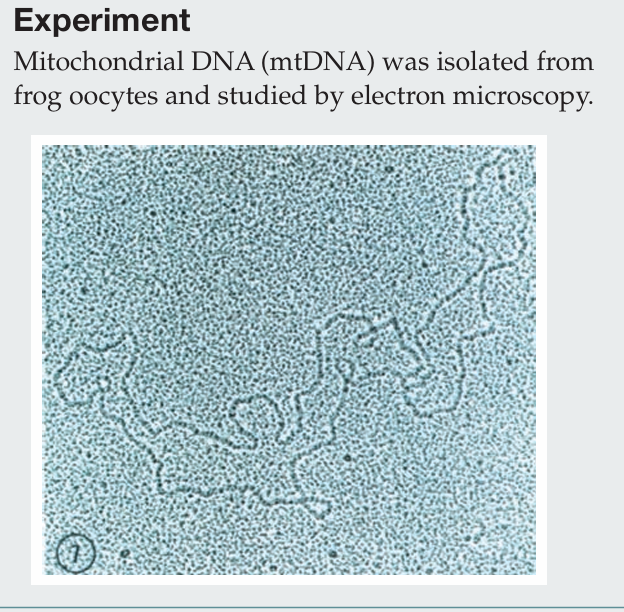
Which electron microscopy technique was used to visualize this mtDNA molecule?
Transmission electron microscopy (TEM) was used, in which heavy metal staining creates electron-dense regions, allowing the DNA to appear as a dark filament against a granular background.
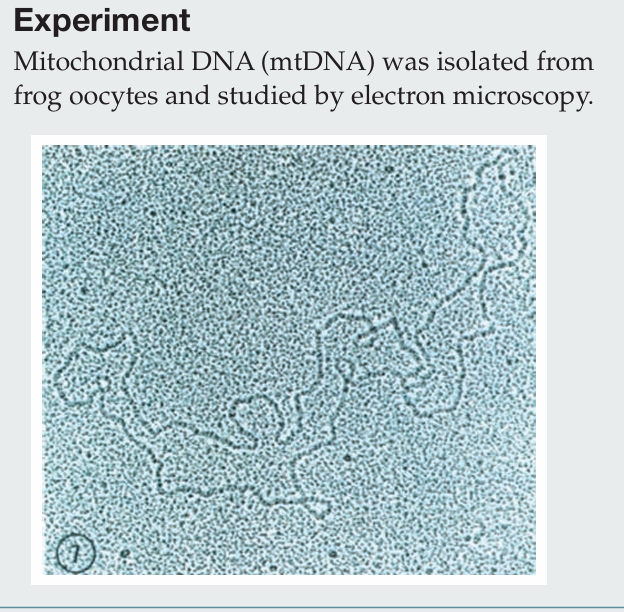
What property of the mtDNA is similar to bacterial genomes?
Both mitochondrial DNA and bacterial genomes are circular.
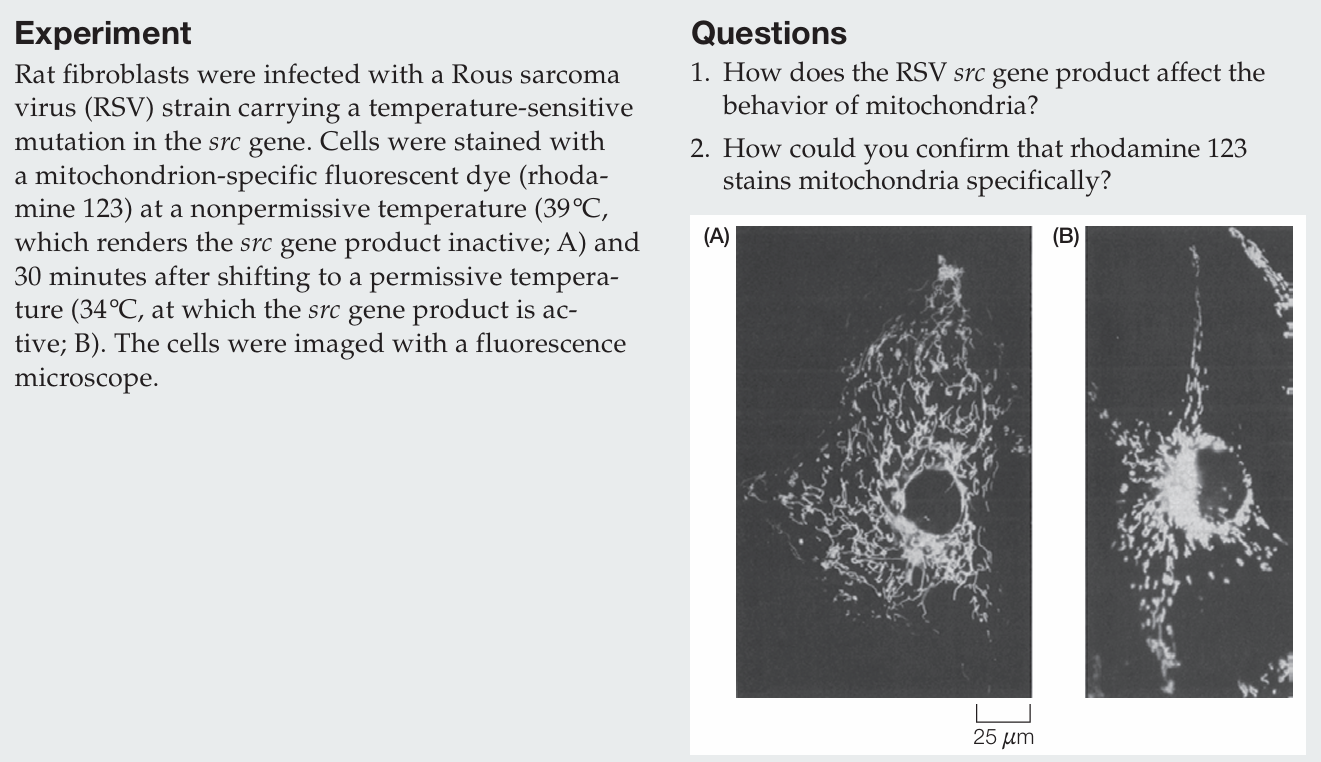
How does the RSV src gene product affect the behavior of mitochondria?
When the src gene product is inactive, mitochondria are distributed throughout the cytoplasm; when it is active, mitochondria cluster around the nucleus.
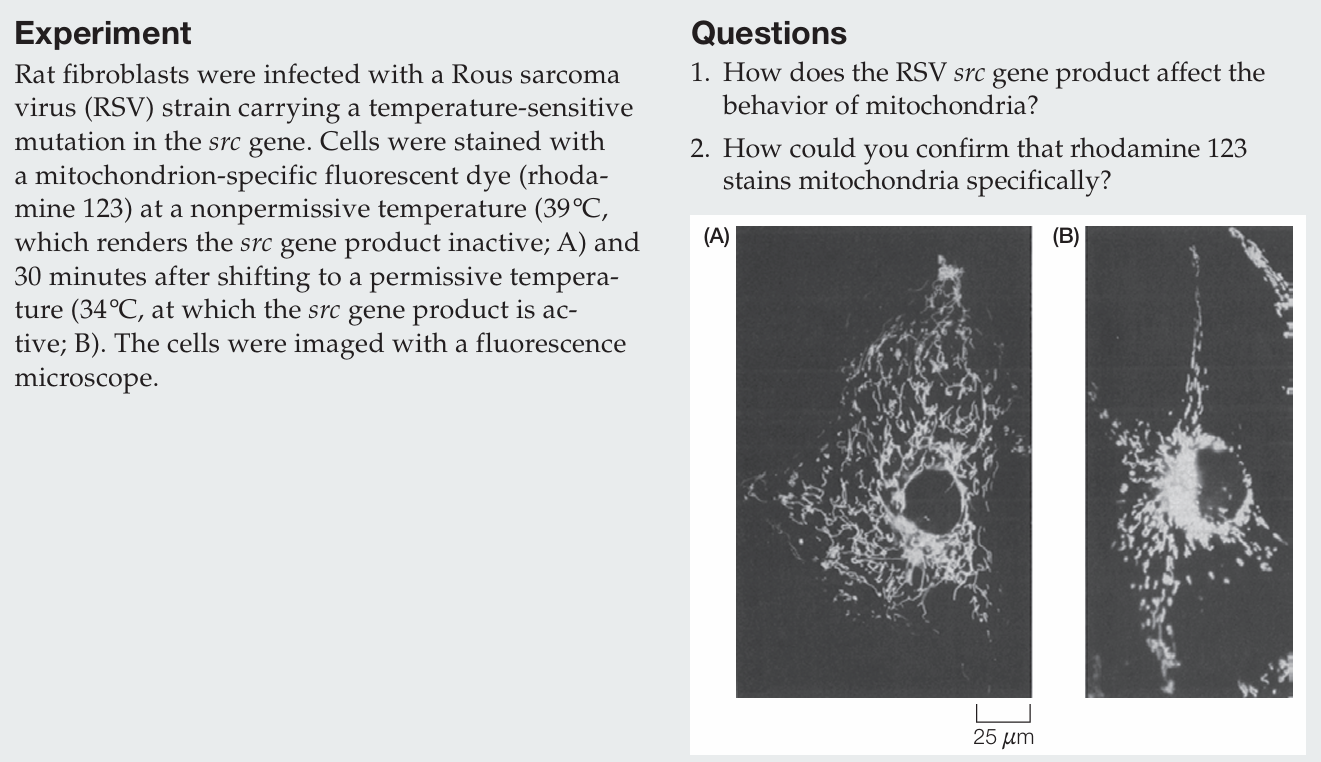
How could you confirm that rhodamine 123 stains mitochondria specifically?
Specificity can be confirmed by co-localization with a known mitochondrial marker, such as a fluorescently labeled antibody against a mitochondrial protein or a GFP-tagged mitochondrial protein; overlapping signals would indicate specific staining.