Cell Bio Ch. 18: The Cell Division Cycle
1/43
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
44 Terms
4 phases of the cell cycle
G1: preparation, growth, protein synthesis, organelle duplication
S: DNA replication
G2: ensure replication is proper before mitosis
M: mitosis (nuclear division) and cytokinesis
3 major checkpoints
G1-S: are we ready to duplicate?
G2-M: is all DNA replicated and repaired before we can divide?
Anaphase: are all the sister chromatids aligned correctly and attached to mitotic spindle?
what other eukaryotic cells are most used to study the cell cycle
yeast cells
CDKs and cyclins
CDKs: cyclically activated protein kinases that control cell cycle
not always active/phosphorylating
cyclins: proteins that bind and activate CDK
regulating cyclin concentration regulates CDKs
general cell checkpoint concept: different cyclin-CDK complexes
different complexes of cyclins and kinases trigger different steps in the cell cycle
G1-S and G2-M checkpoints are regulated by what which complexes?
G1-S: active G1/S or S cyclin-CDK complexes
G2-M: active M cyclin-CDK complex
proteolysis (what it is and what effects)
degradation of proteosomes to breakdown cyclin, preventing cell cycle (transcription) from continuing
silencing CDKs through proteolysis decreases cyclin concentration
how do proteins end up in the proteasome for degradation?
active CDK is bound to cyclin
tagging with ubiquitin and DAG silences the complex
tagging signals protein to be sent to proteasome for proteolysis
proteasome degrades cyclin and inactivates CDK to stop cell cycle
APC (anaphase promoting complex)
adds ubiquitin
promotes the separation of sister chromatids
wants to ensure cell doesn’t repeat mitosis
regulates the M-CDK complex (stop cell from entering M)
indirectly promotes anaphase and mitosis, but also prevents cell cycle from re-entering mitosis in the same cycle
how do cyclin-CDK complexes depend on phosphorylation? (Wee1 and Cdc25)
kinases are inhibitory
phosphorylation inactivates CDKs
Wee1 phosphorylates CDK
phosphatases are activators
dephosphorylation activates CDKs
Cdc25 reverses the action of Wee1
phosphorylation determines whether or not a cyclin-CDK complex is active
If I have a CDK bound to a cyclin, do I know for sure the CDK is active?
NO
Need know if it is phosphorylated or not
three regulations of CDK
controlling cyclin concentration
(de)phosphorylation of CDK
inhibitory proteins
inhibitor proteins on CDK
important for stopping and fixing errors
bind onto CDK complex like a clamp to inactivate the complex
p27, p21, p58 are inhibitory proteins
3 ways the cell cycle control system can pause the cycle
G1-S: CDK inhibitors (p21,27,58) block entry to S phase
G2-M: kinases (Wee1) and phosphatases (Cdc25)
Anaphase: ubiquitylation driven by APC
G0
rest stop: slow down and assess
cell can hold here for a while to fix or go through apoptosis
terminal differntiation
cell doesn’t want to divide anymore
resting favored
CDKs indefinitely silenced
cyclin concentration wiped
G1
cell grows, replicates organelles, makes proteins (cyclins, CDKs), prepares for replication
mitogens
external chemical ligand that promotes cell cycle (proliferation) and triggers G1
promote the production of cyclins that stimulate cell division
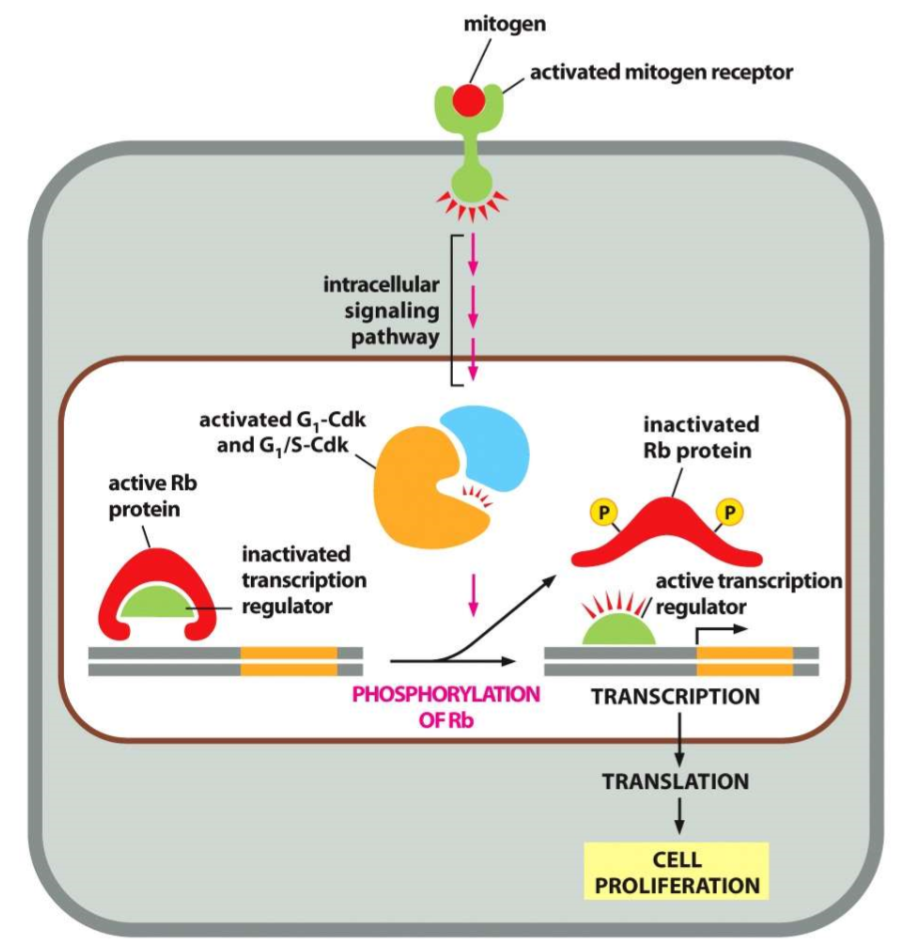
mitogens and Rb protein
mitogen activates a receptor that activates an intracellular signaling pathway
G1-CDK and G1/S-CDK is activated to phosphorylate the active Rb
phosphorylation inactivates the Rb and it releases from the transcription regulator, thus activating it
transcription can take place, leading to cell proliferation
retinoblastoma
mutant Rb (or none)
constant transcription due to constantly activated transcription regulator causing cancer in the eye
what happens when DNA is damaged
DNA damage signals kinases that phosphorylate p53 and activate it
p53 binds and activates p21 CDK inhibitor protein
p21 inactivates G1-S CDK / S CDK
Damaged DNA halts cell cycle progression to fix
what happens when S initiates?
S-CDK initiates DNA replication and blocks re-replication
DNA is replicated using DNA Polymerase III between two unwinding strands
How is the replication is S phase controlled?
need brakes to ensure DNA replicates only once so sister chromatids are equally separated
G1:ORC (origin recognition complex) region with Cdc6 bound
as cyclin complexes activate, the Cdc6 dissociates while 2 molecules of helicase bind to the lift of ORC to form pre-replicative complex
S: S-CDK phosphorylates ORC and allows helicase to separate strands for DNA Pol III
role of Cdc6
Cdc6 is bound at ORC during G1
dissociates so helicase can bind to initiate S replication
incomplete replication, role of Cdc25
arrests cell in G2
Cdc25 (dephosphorylates M-CDK and activates it) gets locked by phosphorylation of CDK by Wee1
condensins and cohesins
condensins: compact chromosomes and make them visible
cohesins: keep sister chromatids together until anaphase
cytoskeleton 2 roles in mitosis
mitotic spindle (form centrosome MT)
contractile ring (actin + myosin)
mitotic spindle
form centrosomes (2 on opposite sides)
usually we only have 1 so in S they replicate
initially they are very close, but they move to other sides to make spindle
aster microtubules come out like a star
cell cycle steps
interphase
prophase
prometaphase
metaphase
anaphase
telophase
cytokinesis
interphase
G1, S, G2
cell grows in size
DNA and centrosome replicates
prophase
duplicated chromosomes condense
mitotic spindle assembles as centrosomes move apart
prometaphase
nuclear envelope breaks down
chromosomes attach to spindle via kinetochores
metaphase
chromosomes align on equator of spindle
kinetochore MTs attached to kinetochores on each side of sister chromatid
anaphase
sister chromatids separate and pull toward each pole of mitotic spindle
two ways they are pulled apart
kinetochores shorten and pull
centrosomes go further apart
sisters must be attached to avoid aneuploidy
trisomy 21: Down Syndrome
telophase and cytokinesis
have 2 complete sets of chromosomes
nuclear envelope reassembles around each
contractile ring squeezes cell to divide into 2
how do centrosomes duplicate
G1: there’s only one centrosome
S: centrosomes replicate
M: asters form (star-like MTs)
centrosomes start moving apart to poles
mitotic spindle forms
spindles attach to chromosomes
three types of mitotic spindle microtubules
kinetochore MTs: attach to kinetochores on chromosomes to pull sister chromatids apart
aster MTs: star shape; anchore spindle to cell cortex to position division
interpolar MTs: overlap in middle to push poles apart
what action is needed to separate sister chromatids?
APC action required
APC tags with ubiquitin to be sent to proteosome
cohesins break down through proteolysis to allow separation
how chromosomes separate in Anaphase A then Anaphase B
A: kinetochore MTs shorten
B: interpolar MTs push poles apart
spindle assembly checkpoint
if attachment is incomplete, anaphase will halt
unattached chromosomes send a stop signal
APC inhibited
sister chromatids stay together
nuclear envelope reassembly in telophase
nuclear lamins/proteins are dephosphorylated (originally phosphorylated by M-CDKs in prometaphase)
reassemble around new chromosomes
what determines where the contractile ring will form?
mitotic spindle
what is the contractile ring made of
actin and myosin
they are perpendicular to the plane of cleavage
organelle separation during cytokinesis
nonspecific; organelles can vary by number in cells
ER
is a part of nuclear envelope
attaches and rearranges with the microtubule cytoskeleton
remains whole then MTs release the ER in daughters during cytokinesis
Golgi
cisternae fragments separate and rearrange to form independent golgi