Week 9 - Non-coding RNAs eg siRNAs
1/121
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
122 Terms
human genome actively transcribed into RNA
60 -70%
protein-coding genes
less than 2% of the human genome
non-coding RNA (ncRNA) molecules
Most of the ncRNAs have been widely implicated in regulating cellular processes
- rRNA, tRNA, microRNAs, long non-coding RNAs (lncRNAs), and other regulatory RNAs
Regulate protein synthesis by regulating mRNA
mRNA is degraded before translation begins, the encoded protein will not be produced
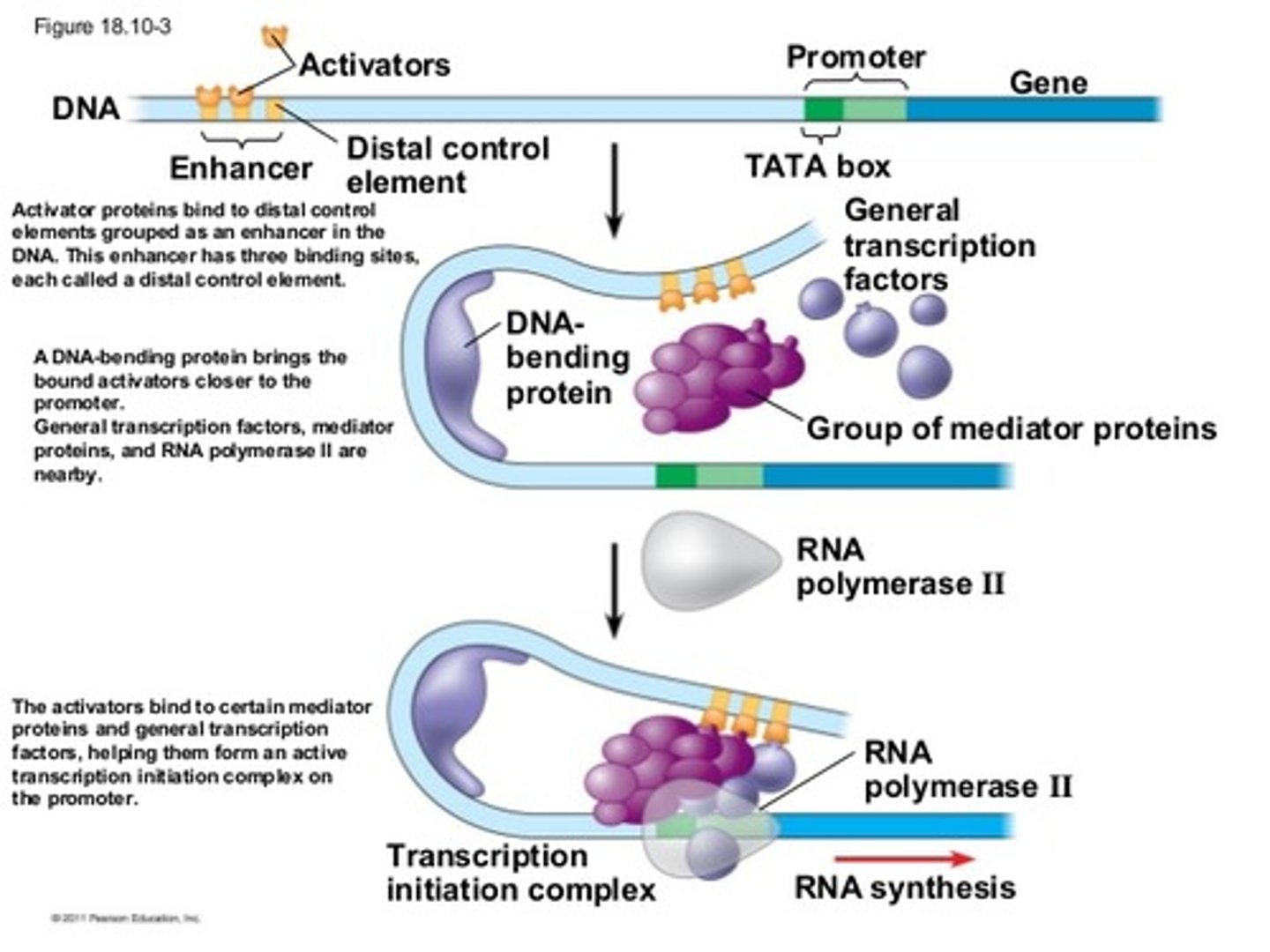
Long non-coding RNAs (lncRNAs)
lack an open reading frame and do not code for protein
Exhibit a wide range of secondary and tertiary structures compared to the coding transcriptome (mRNA)
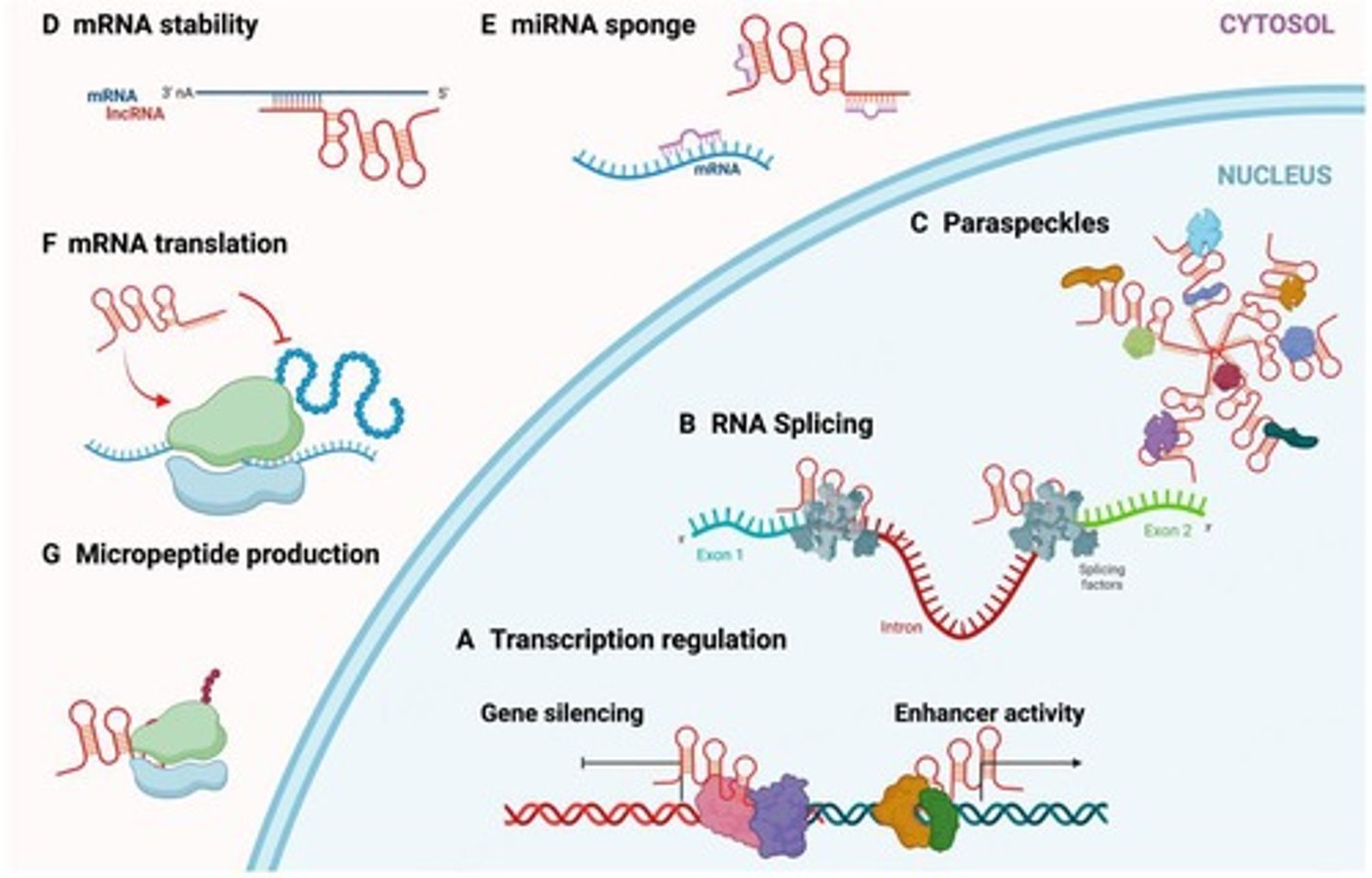
Long noncoding RNAs (lncRNAs) have some features in common with coding RNAs (mRNAs)
can be transcribed by RNA polymerase II or RNA polymerase II
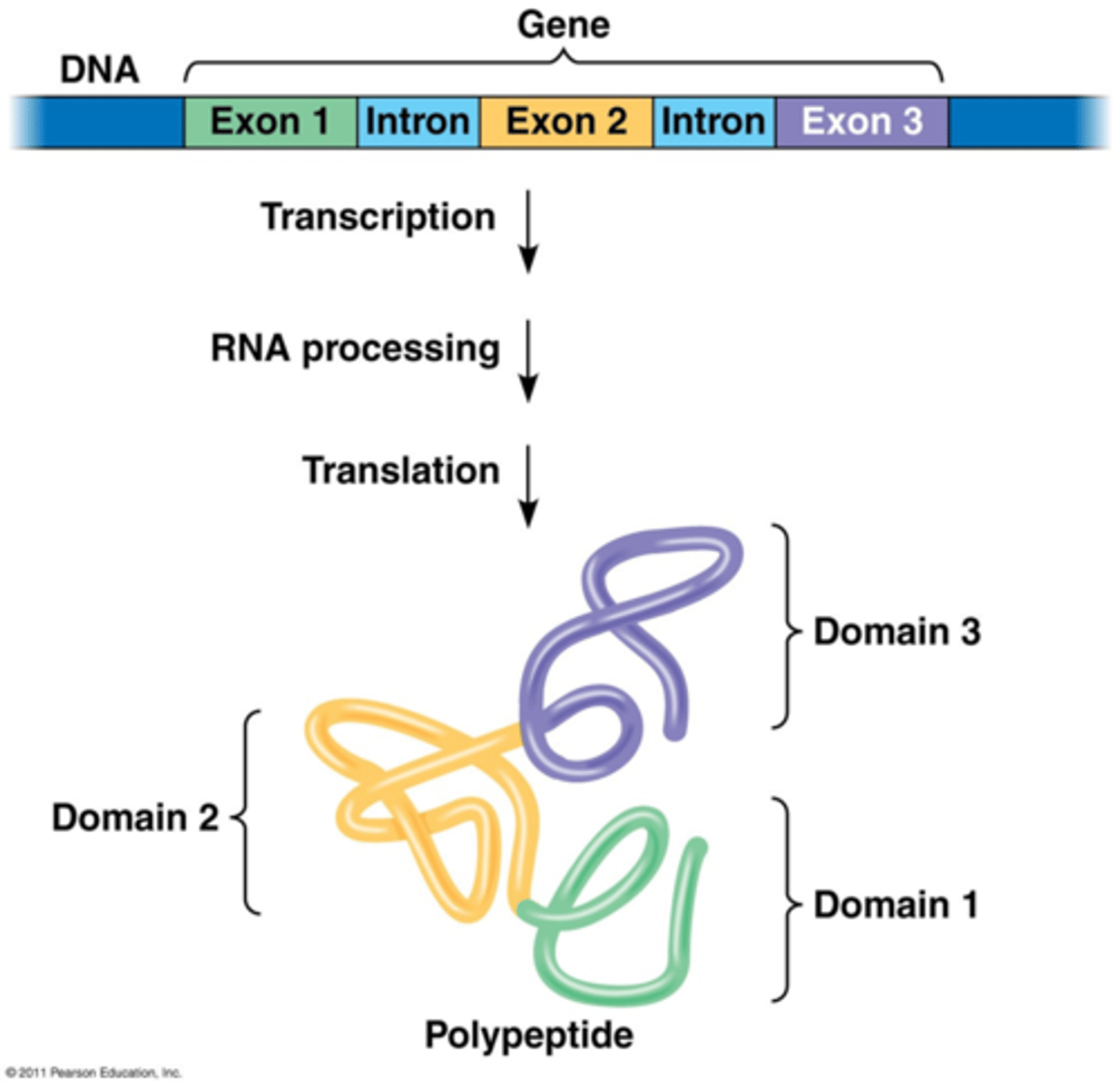
introns
exons
splice variants (alternative splice forms)
transcription is driven by promoter elements
transcription factors- 5'-cap, PolyA tail
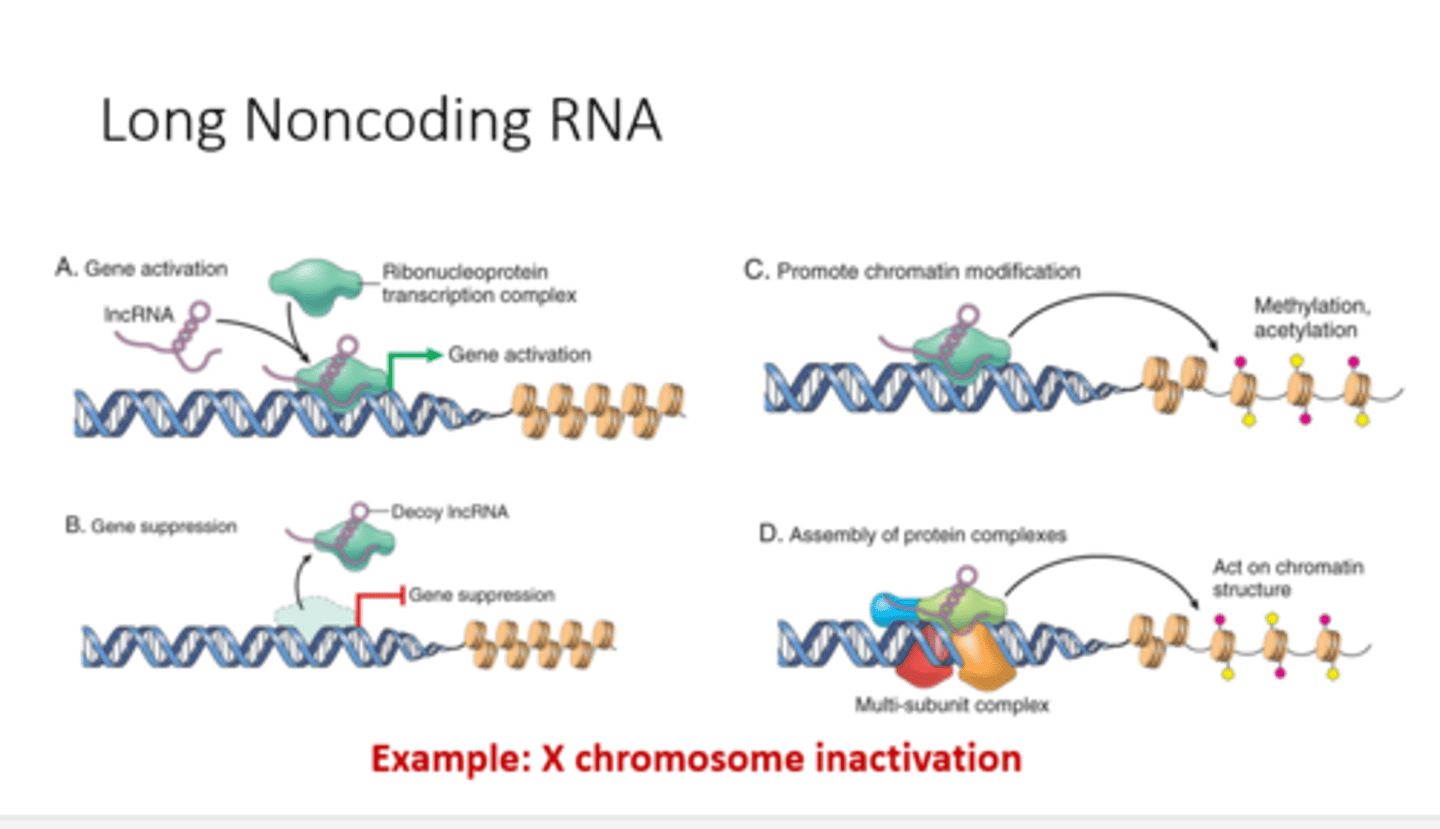
tertiary structure of the LncRNAs
lncRNAs are folded after transcription to create tertiary RNA structures and the tertiary structure determines the function of the LncRNAs
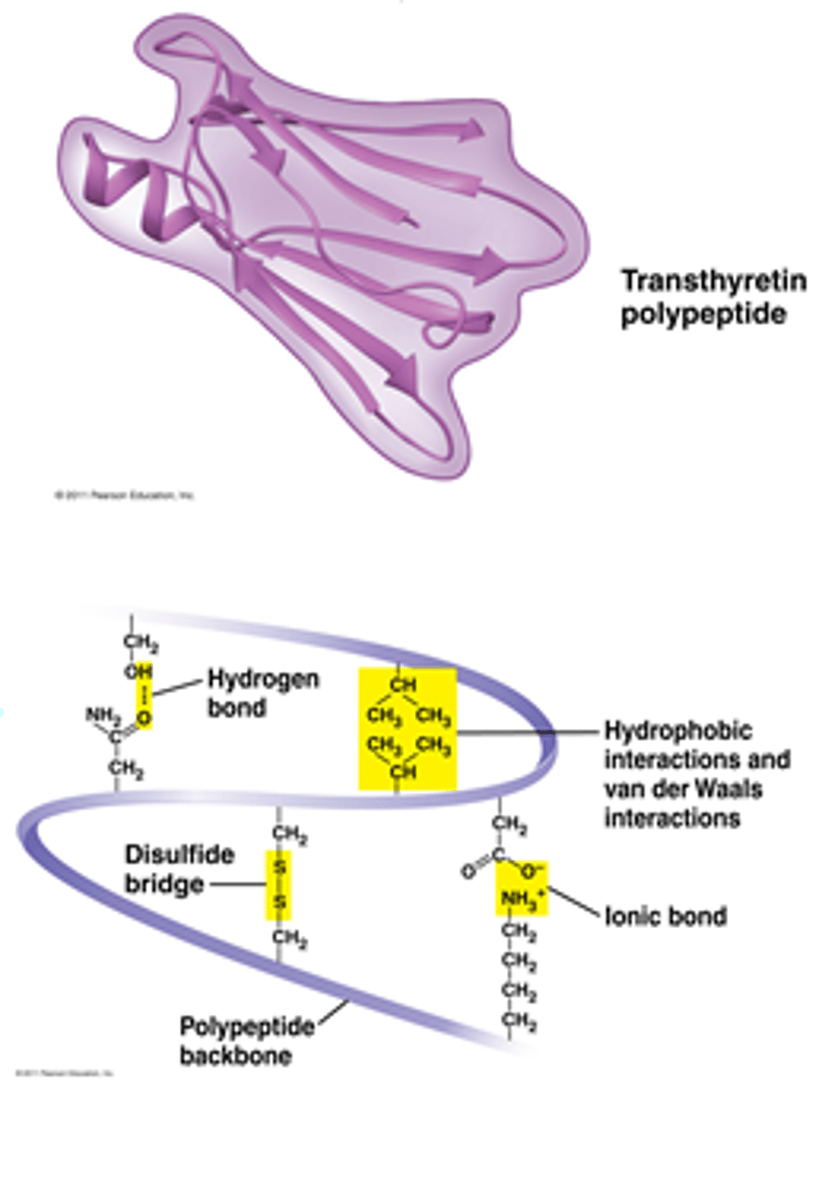
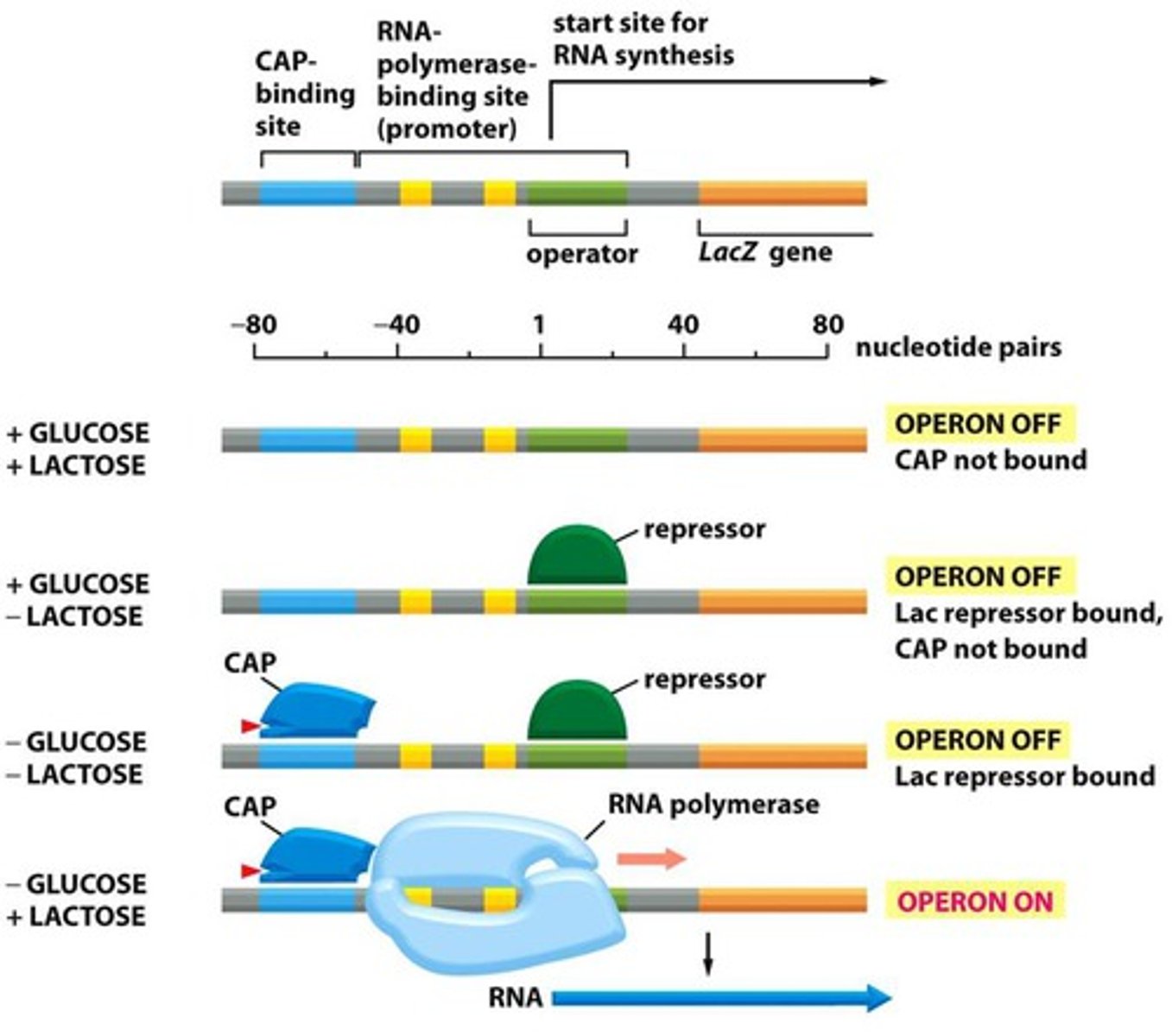
Gene silencing
switching off or turning off a protein coding gene that would normally be turned on (expressed)
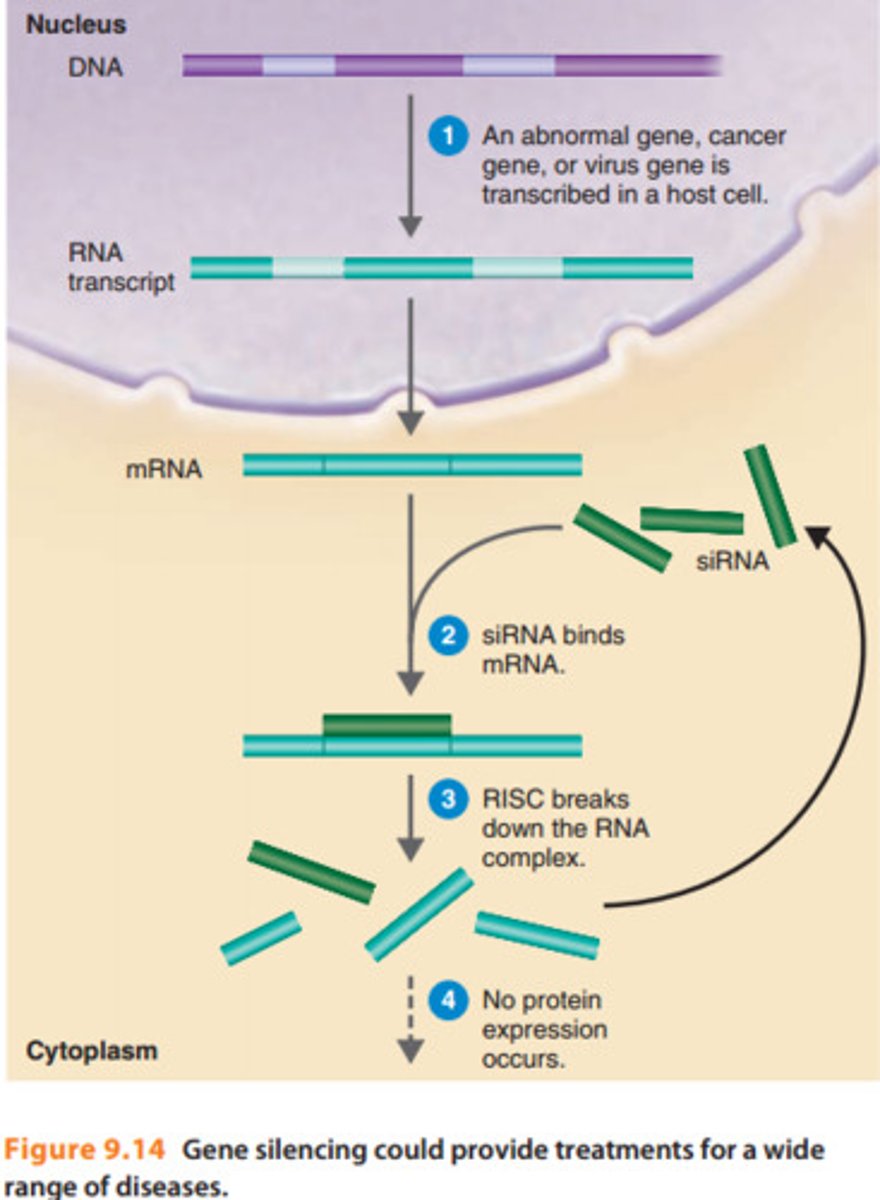
Transcriptional gene silencing (TGS)
silence genes at the level of transcription, preventing the production of mRNA
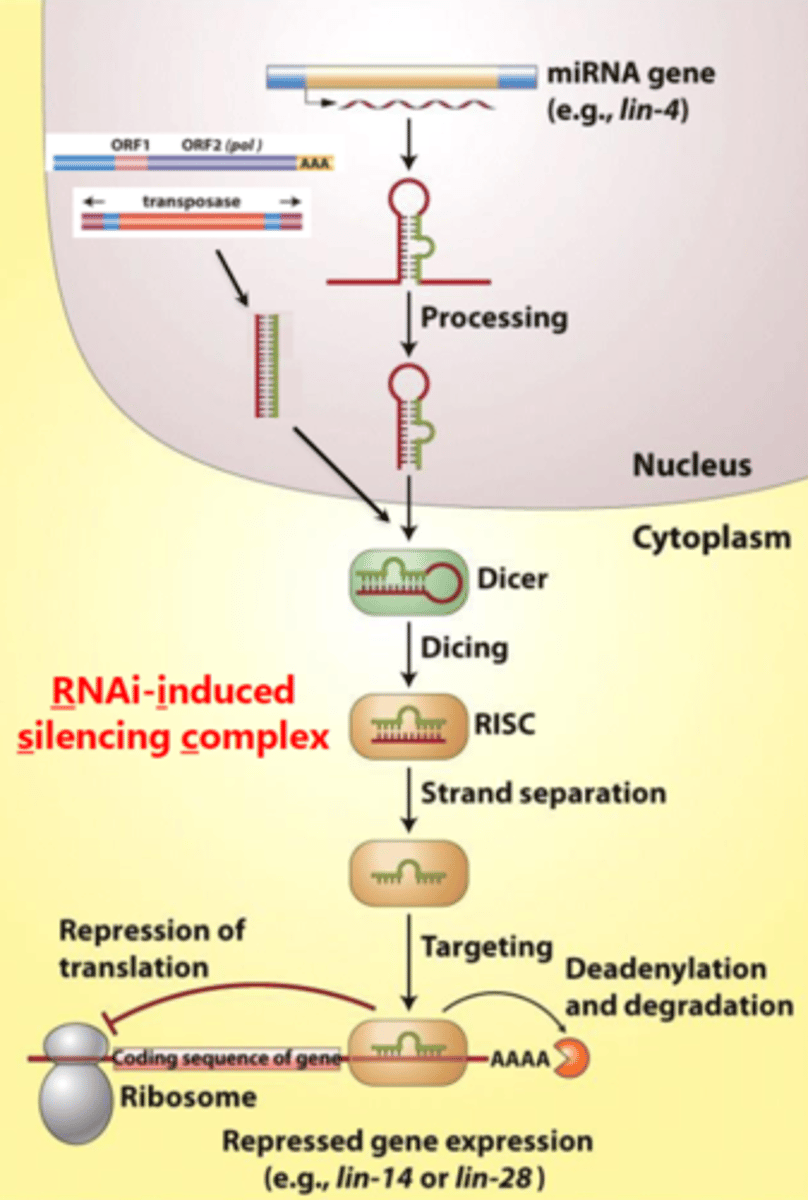
How TGS occurs
processes such as DNA methylation, histone modification, etc., which alter chromatin structure to repress transcription
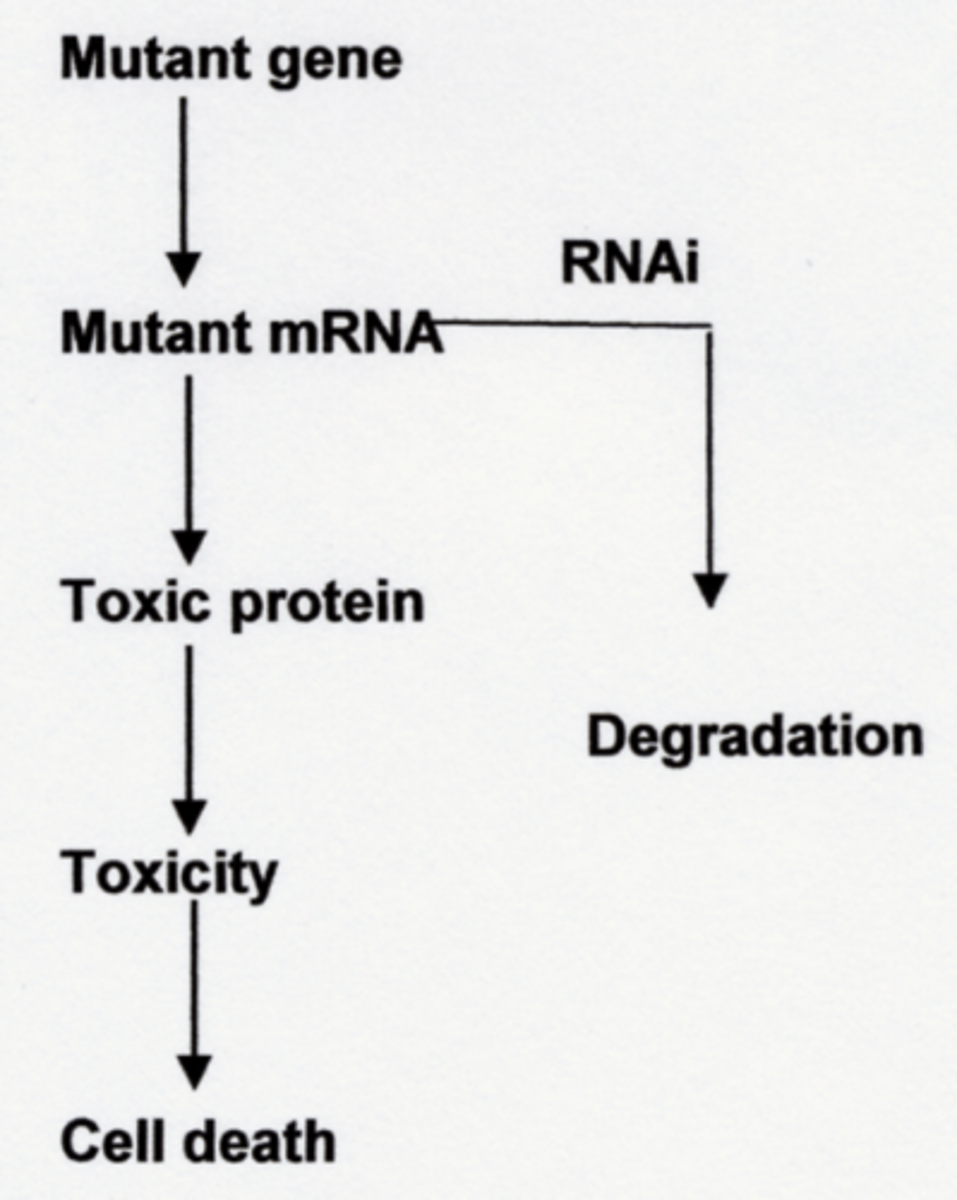
Post-transcriptional gene silencing (PTGS)
suppresses gene expression after transcription has occurred, primarily by degrading mRNA or blocking its translation
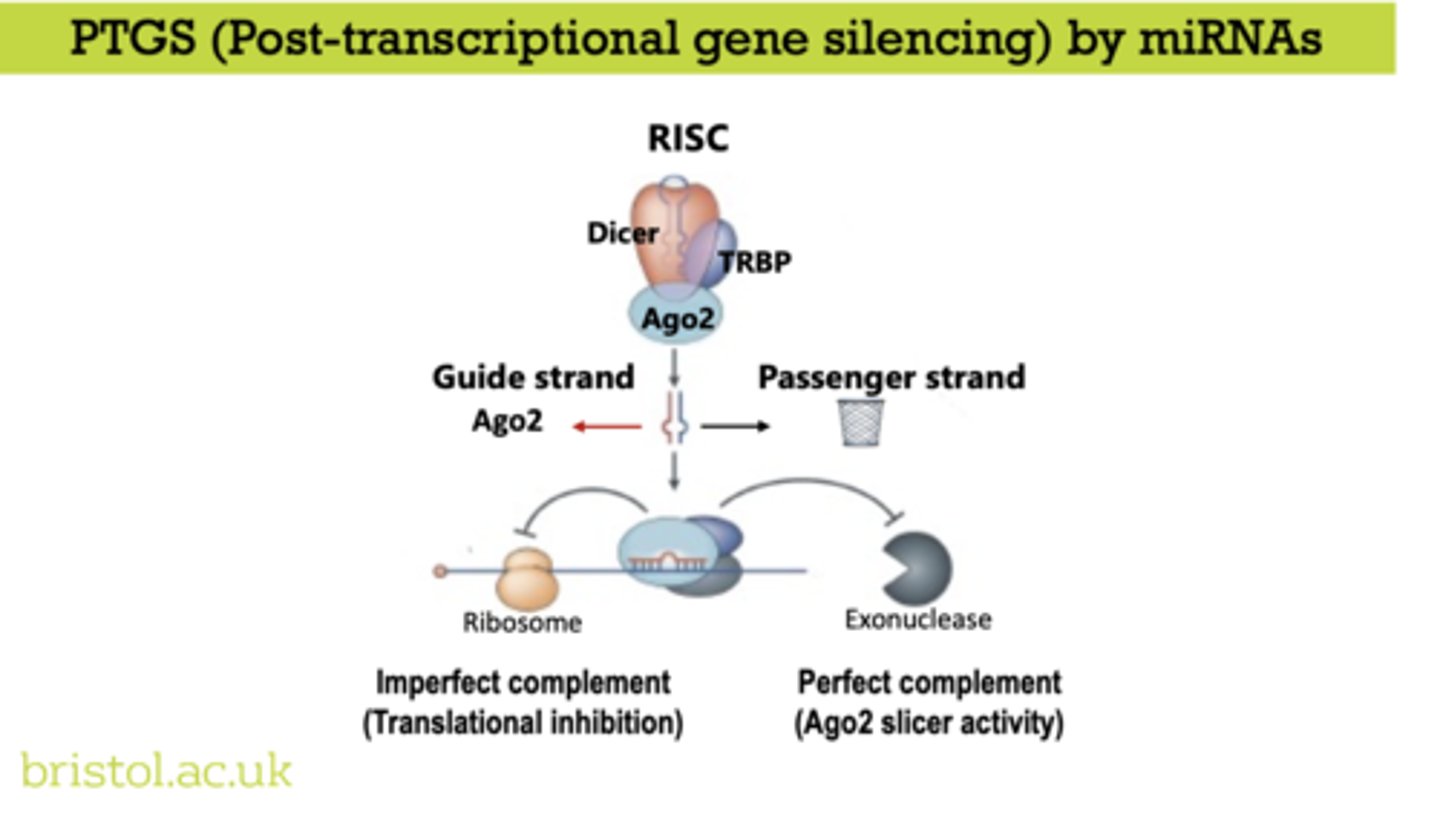
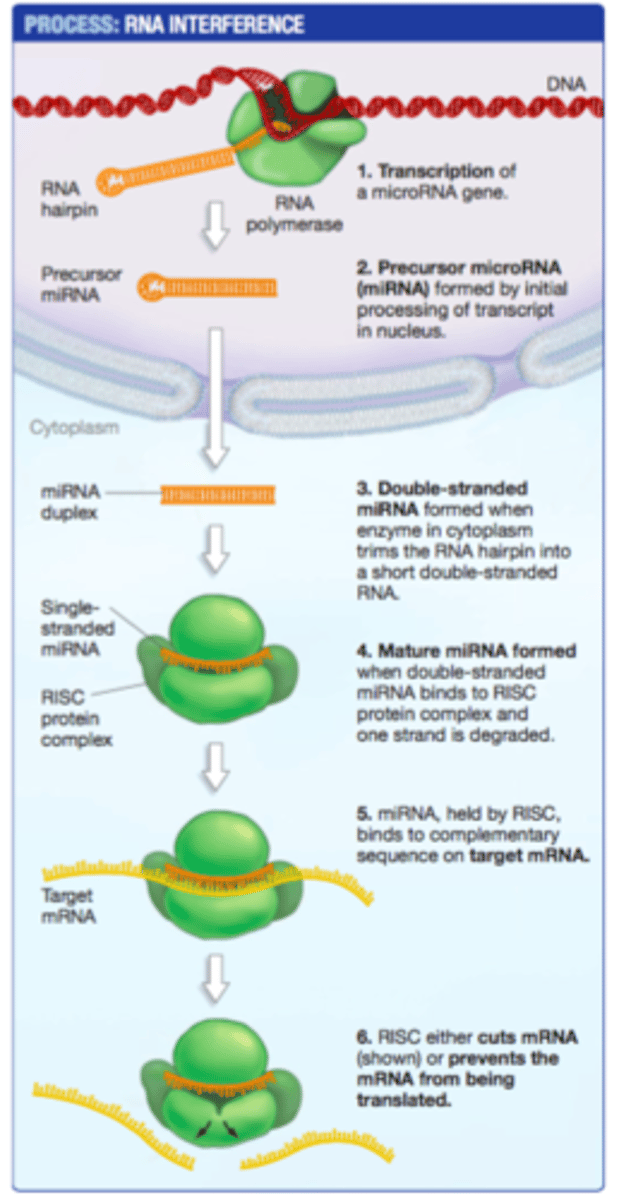
RNA interference (RNAi)
prevents the production of the encoded protein
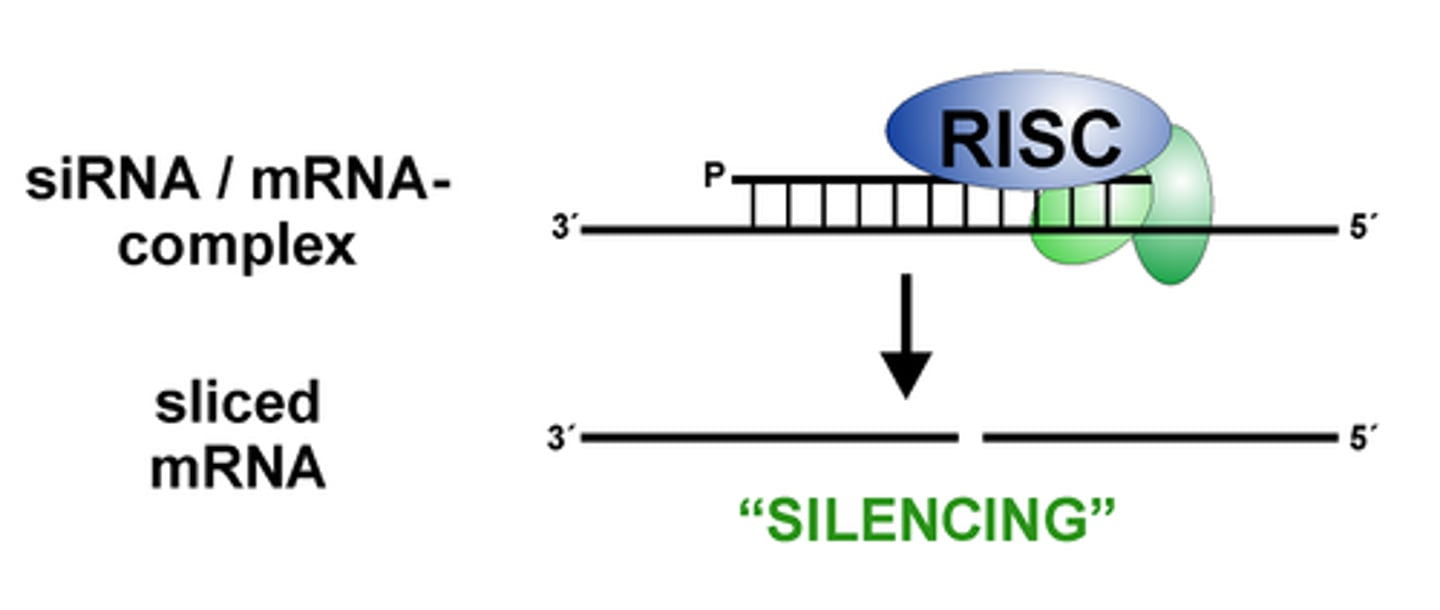
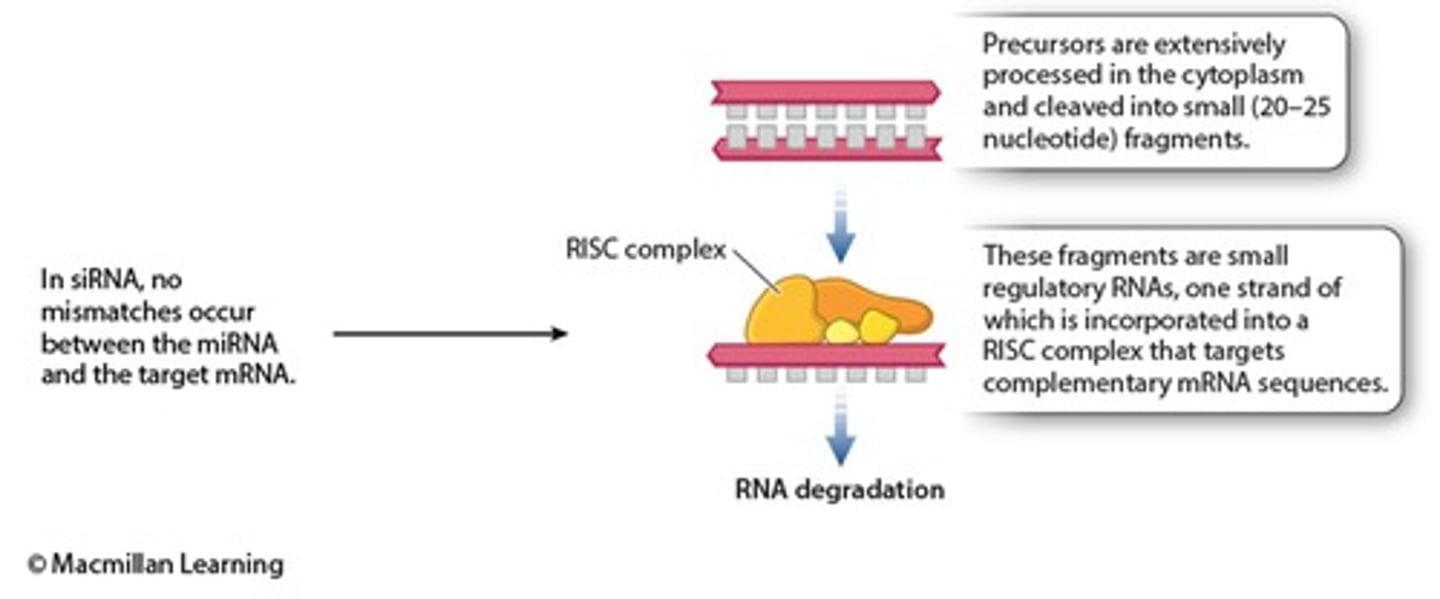
PTGS is often mediated by small RNAs
small interfering RNAs (siRNAs)
microRNAs (miRNAs)
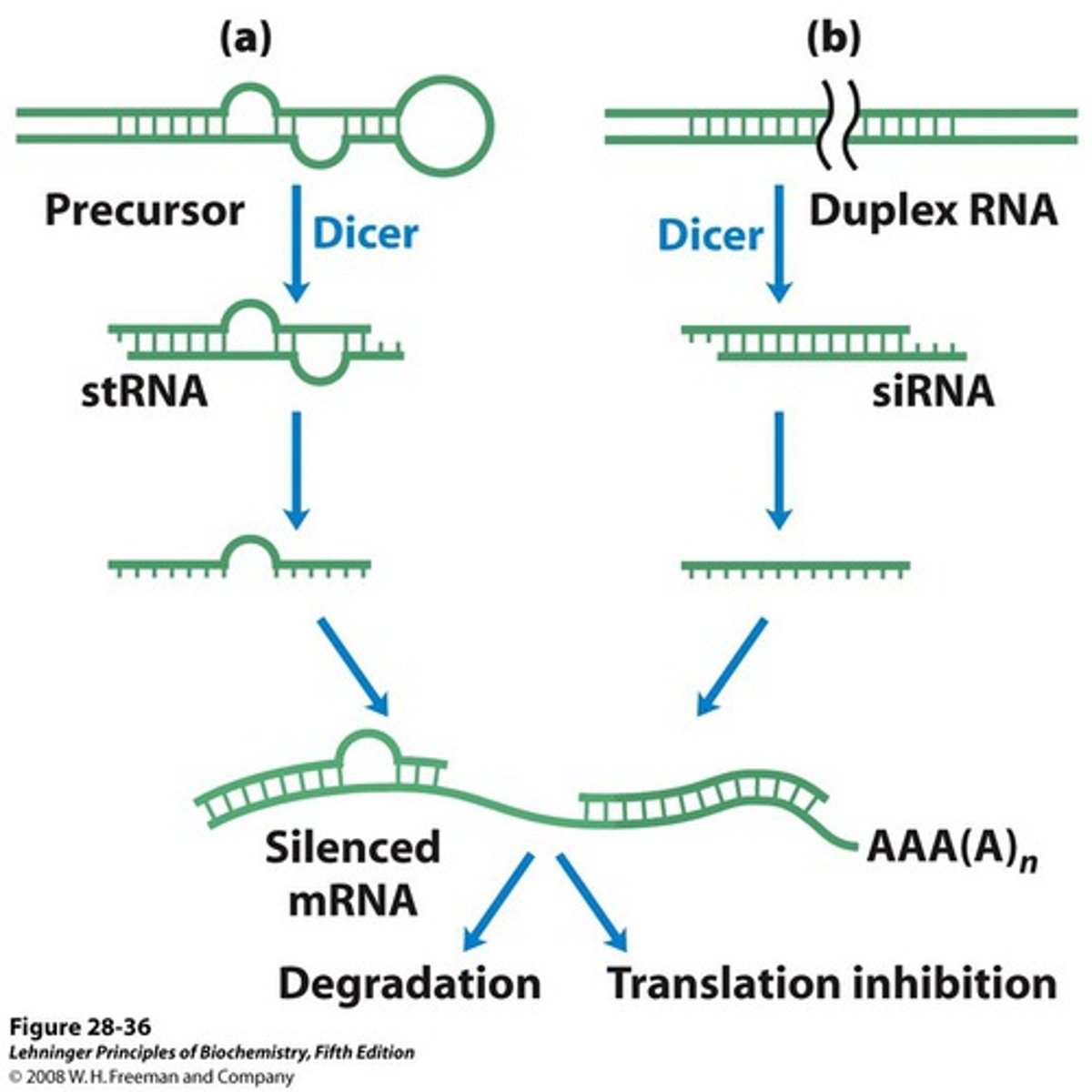
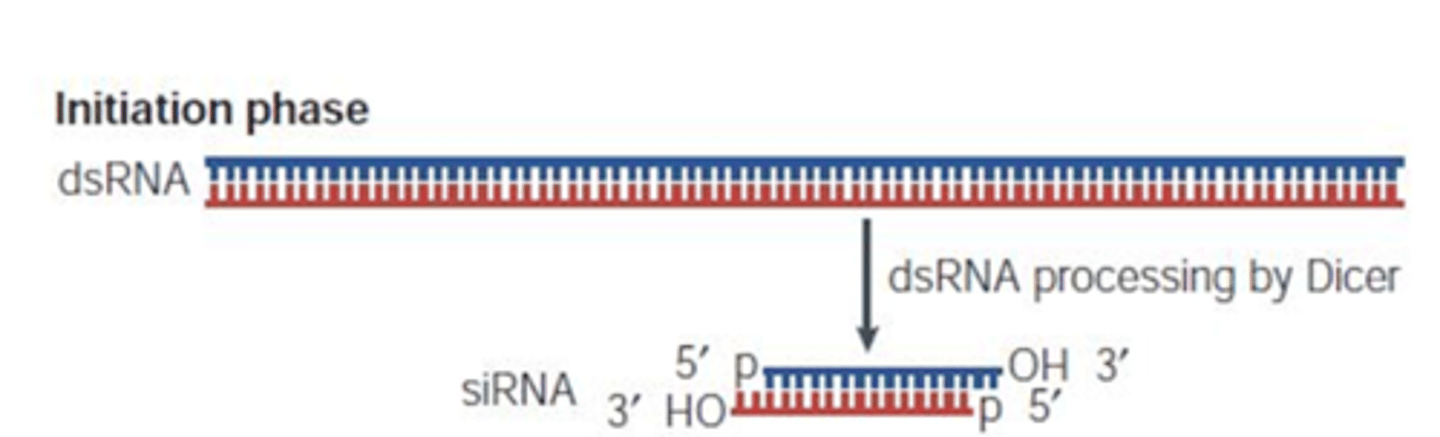
double-stranded RNA serves as the trigger
RNAi mechanism by siRNA recognizes the dsRNAs and works to degrade them
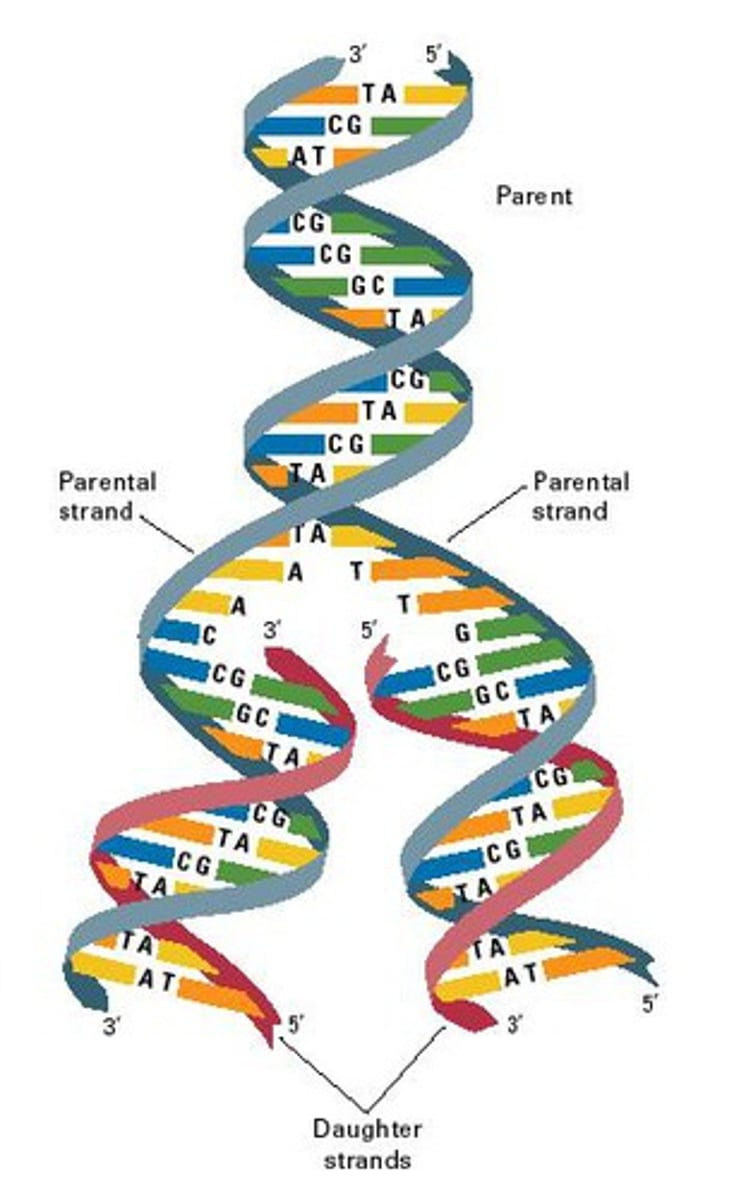
PTGS or RNAi mechanism
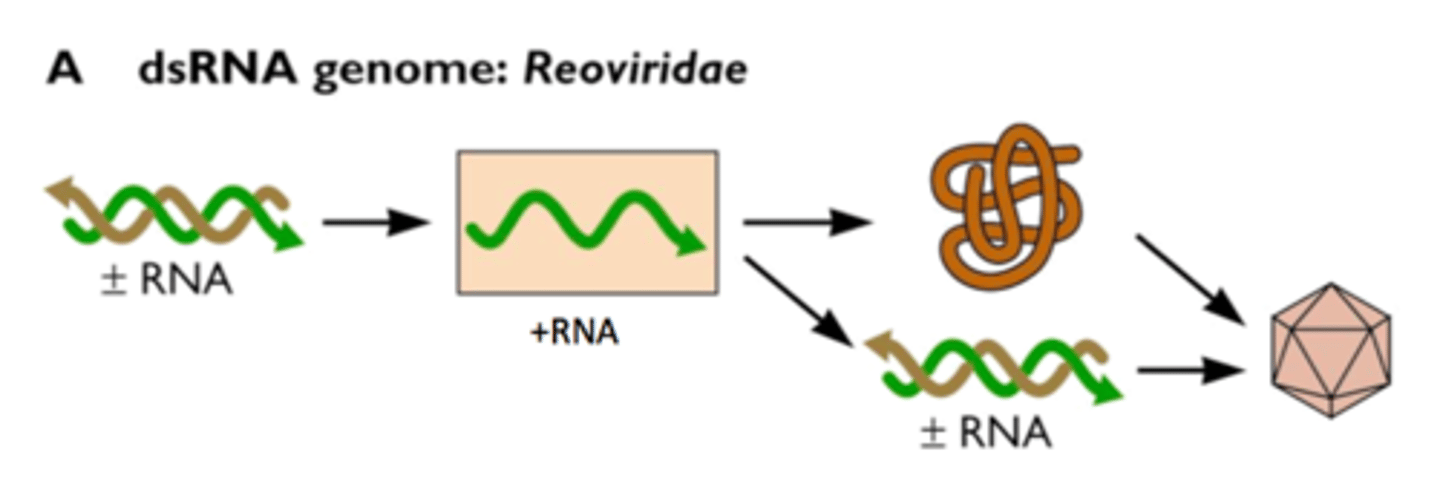
• Offers protection from dsRNA viruses
• Regulate gene expression by silencing genes
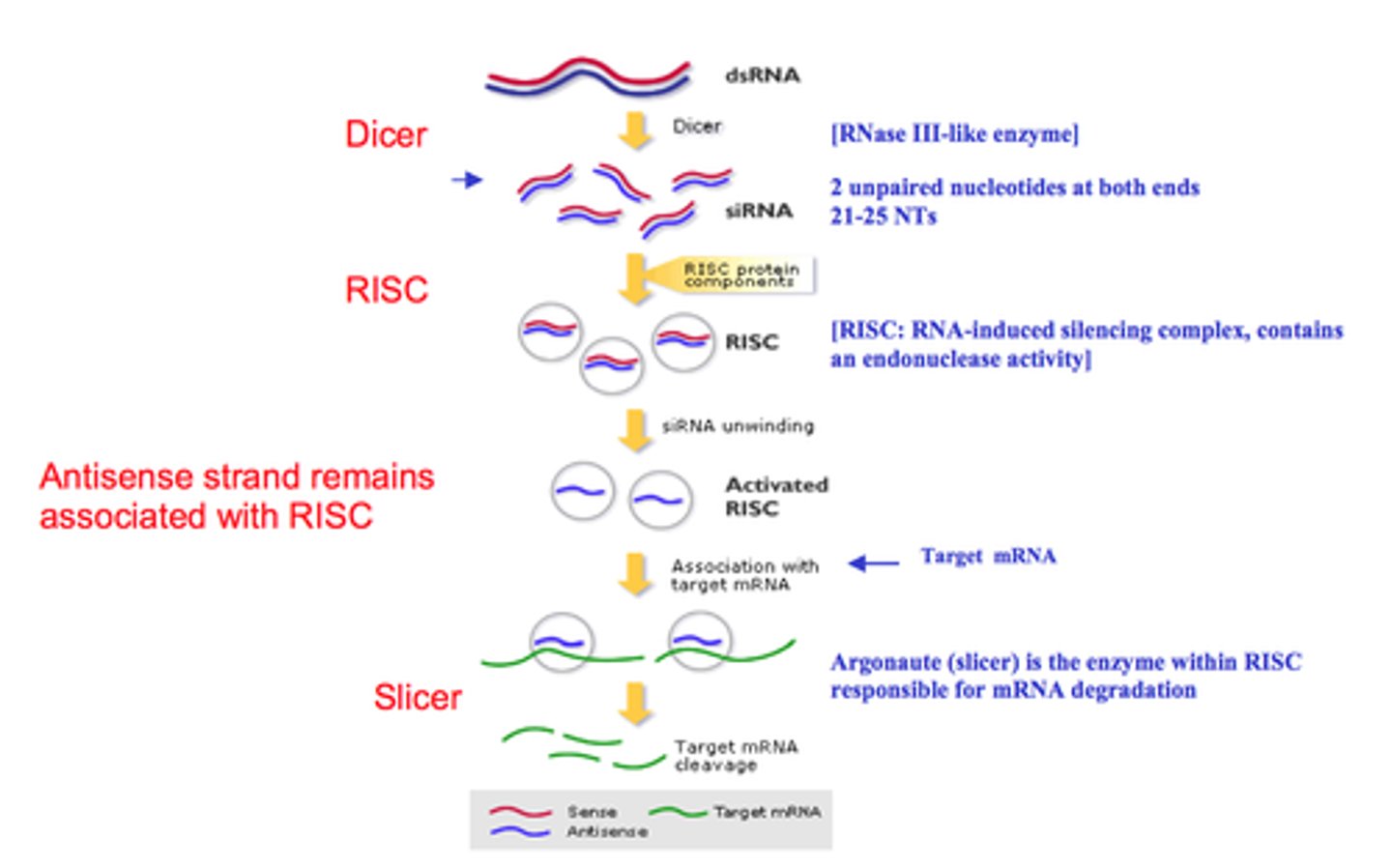
RNAs expressed from genes are single stranded so...
Double-stranded RNAs are recognized as foreign by the cell, activating an endogenous mechanism that destroys them
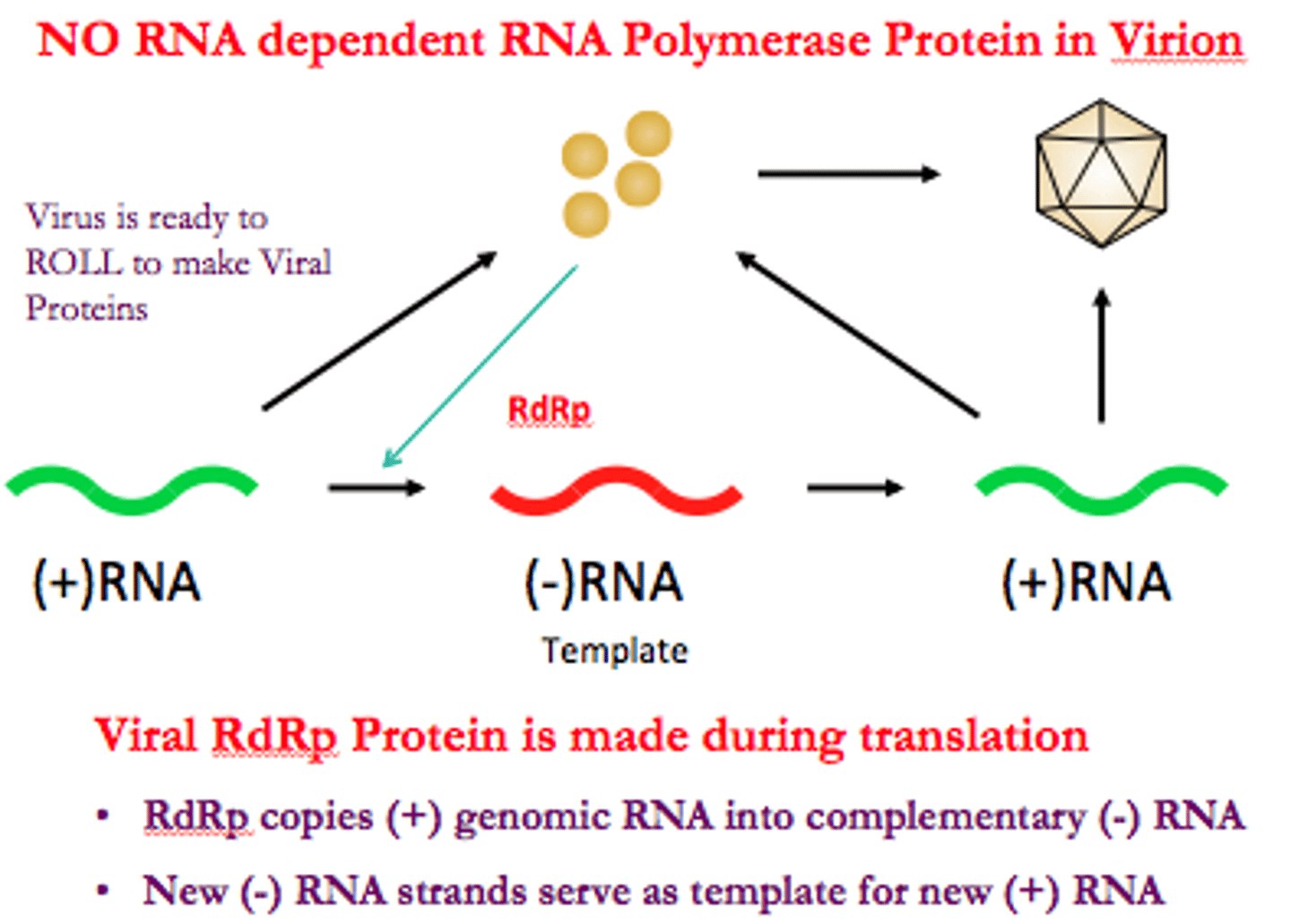
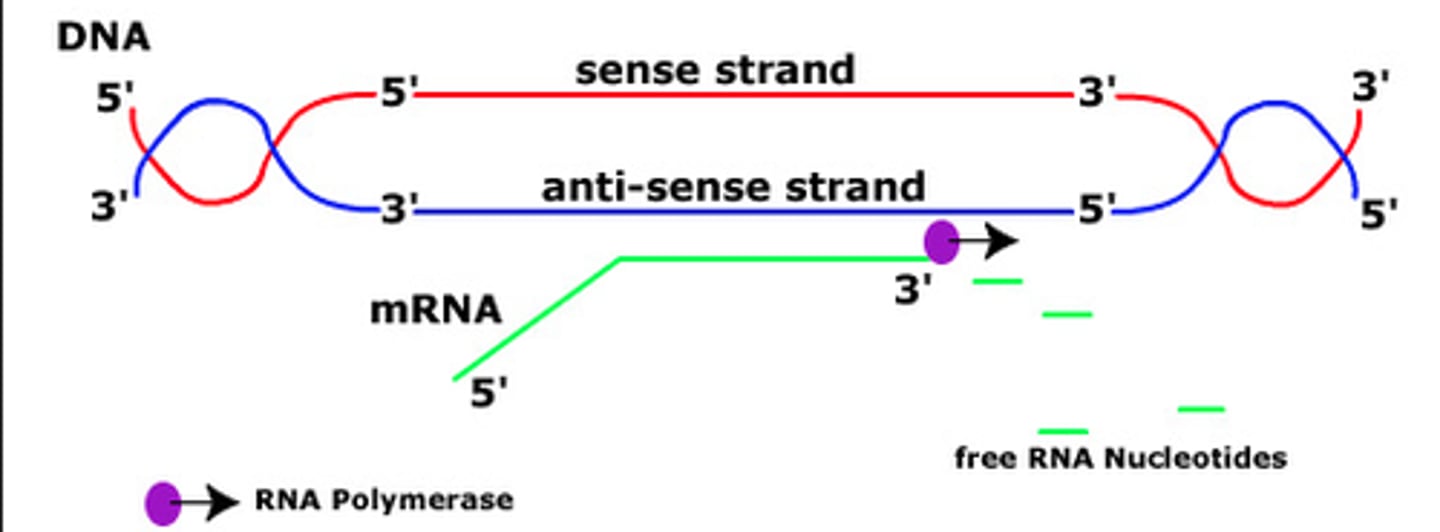
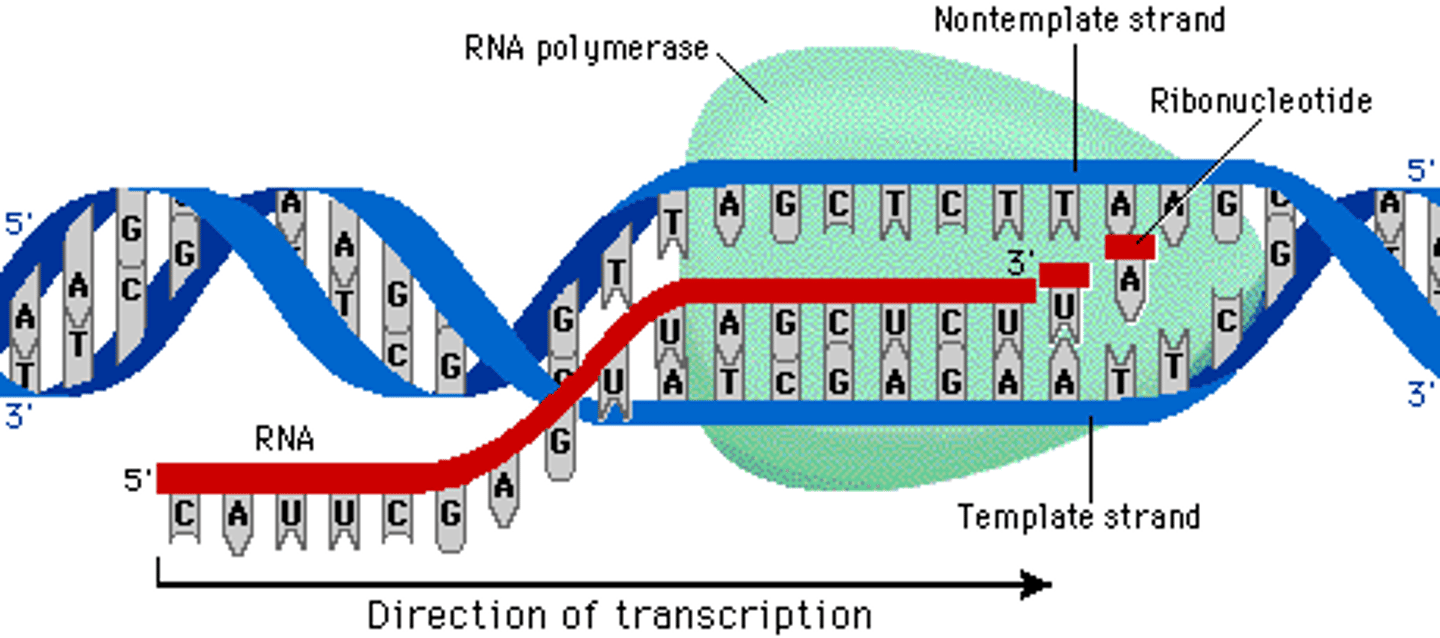
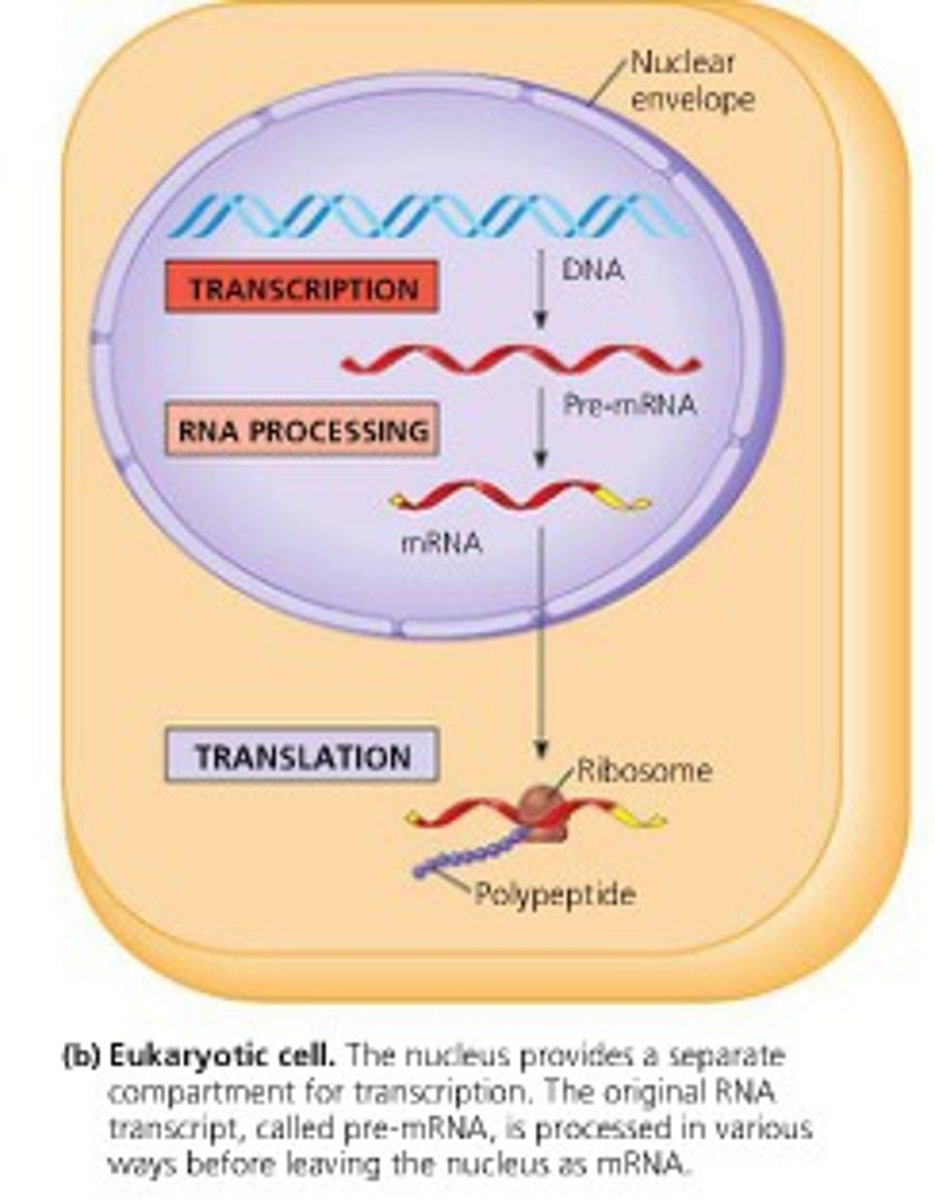
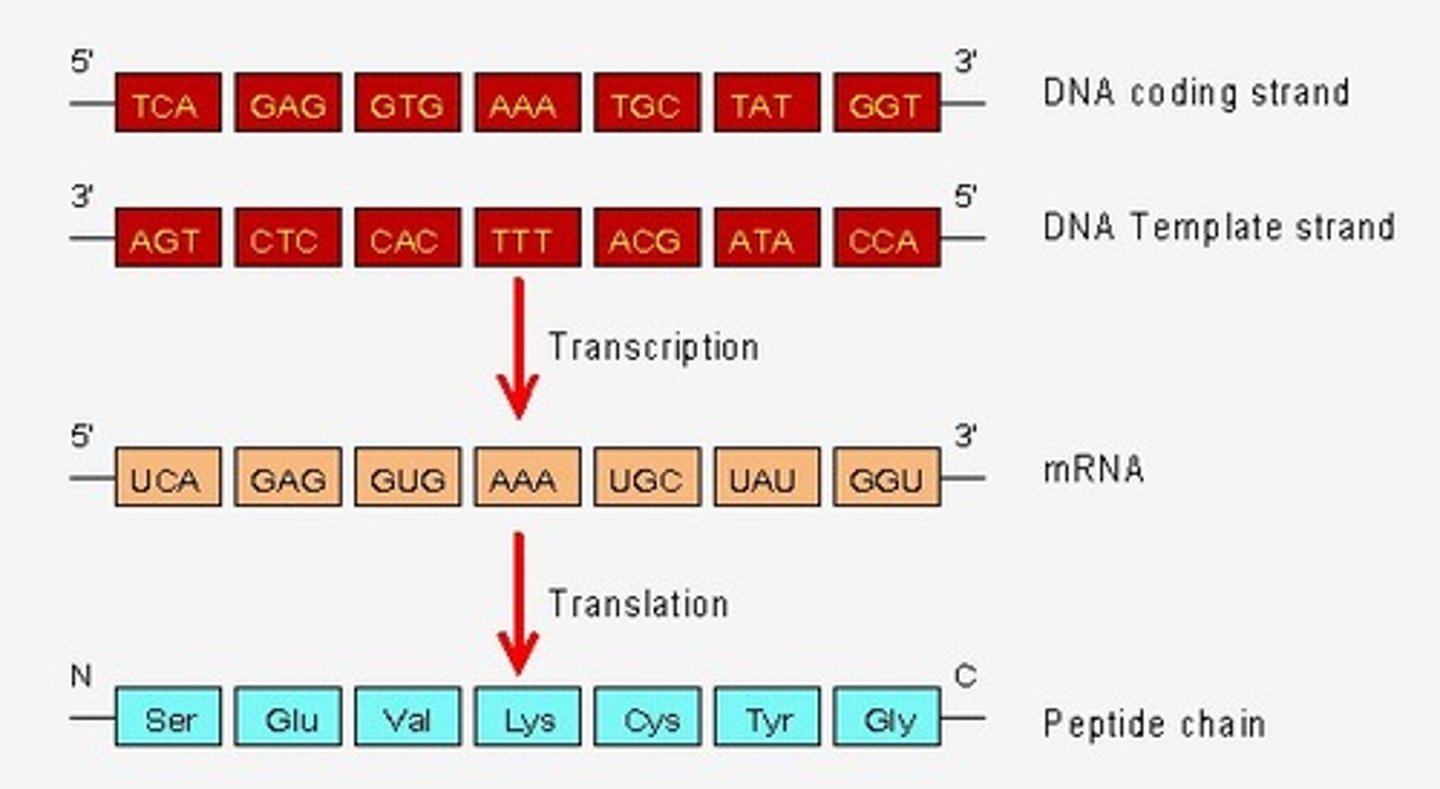
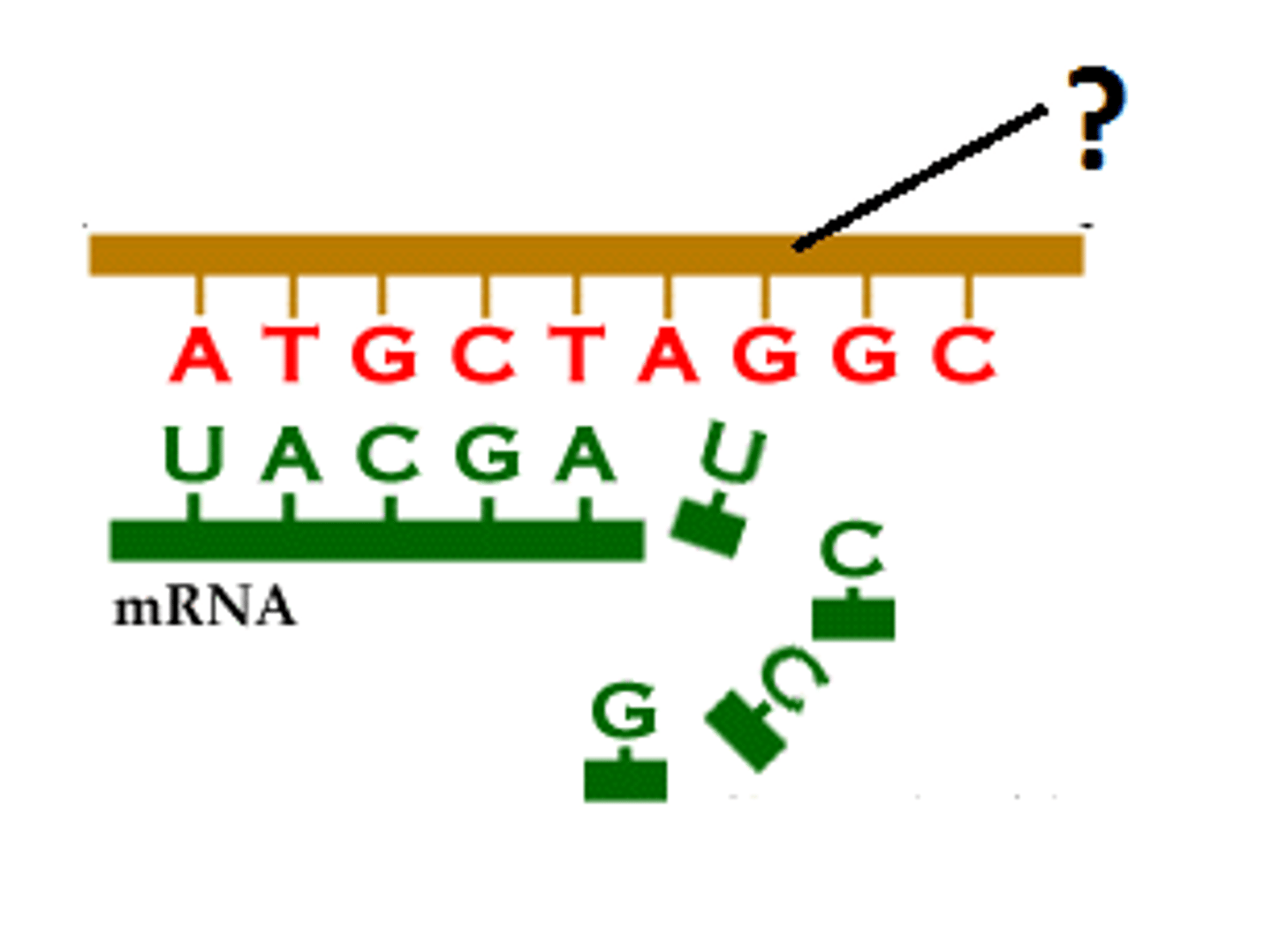
Normal mechanism of transcription
antisense DNA strand (3' to 5') is transcribed to produce a sense mRNA strand (5' to 3')
synthesize RNA exclusively in the 5' to 3' direction, requiring a 3' to5' DNA strand as a template
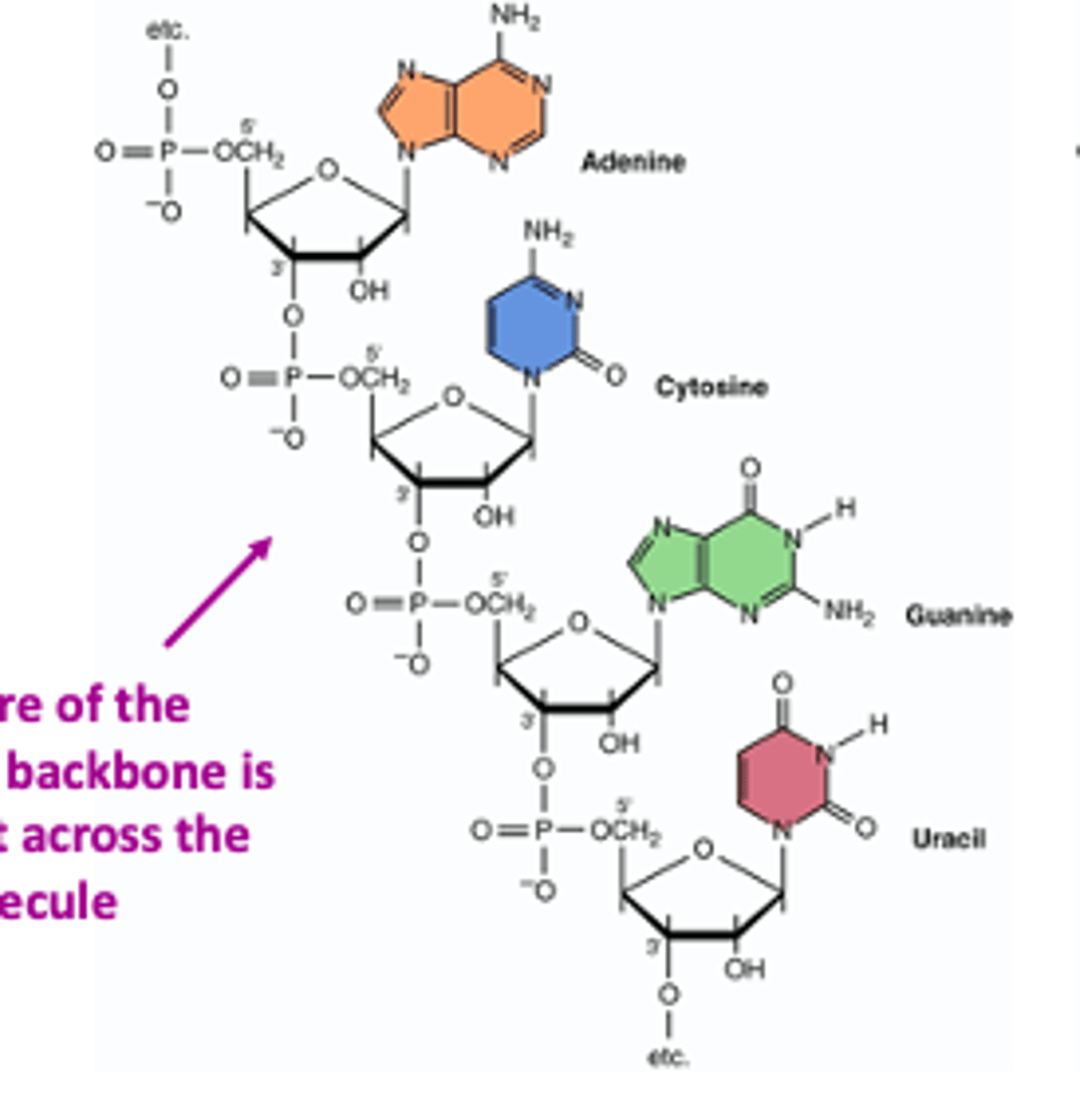
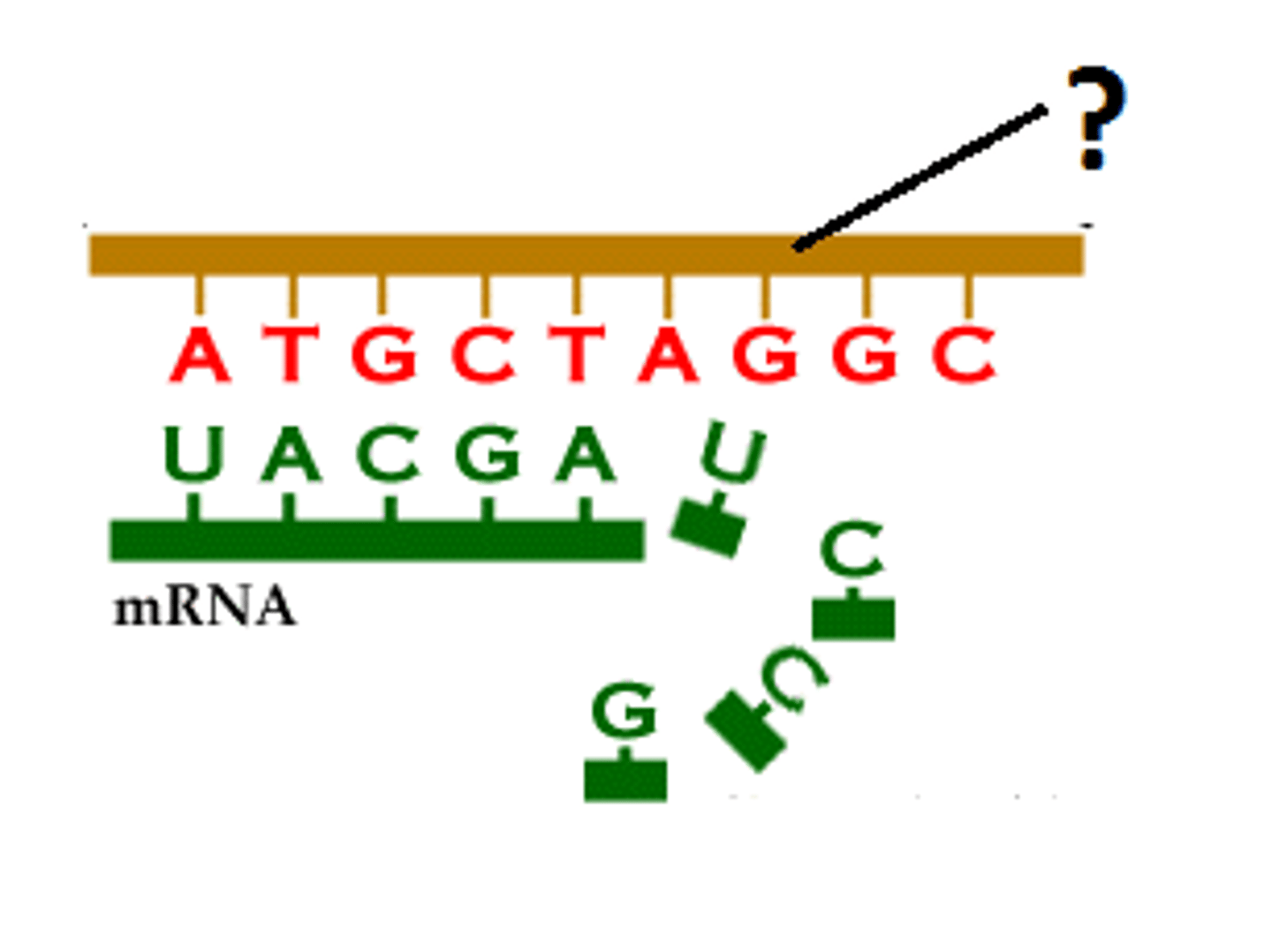
RNA nucleic acid change reminder
Always replace T with U in your RNAs
RNA Polymerases
DNA-dependent enzymes that catalyze transcription; cannot carry out synthesis from the 5' to3' DNA strand
mRNA (sense RNA, 5' to 3') bound to an anti-sense RN
cannot go on to make protein. Thus, the gene from which this mRNA comes is silenced since it cannot express itself by making a protein
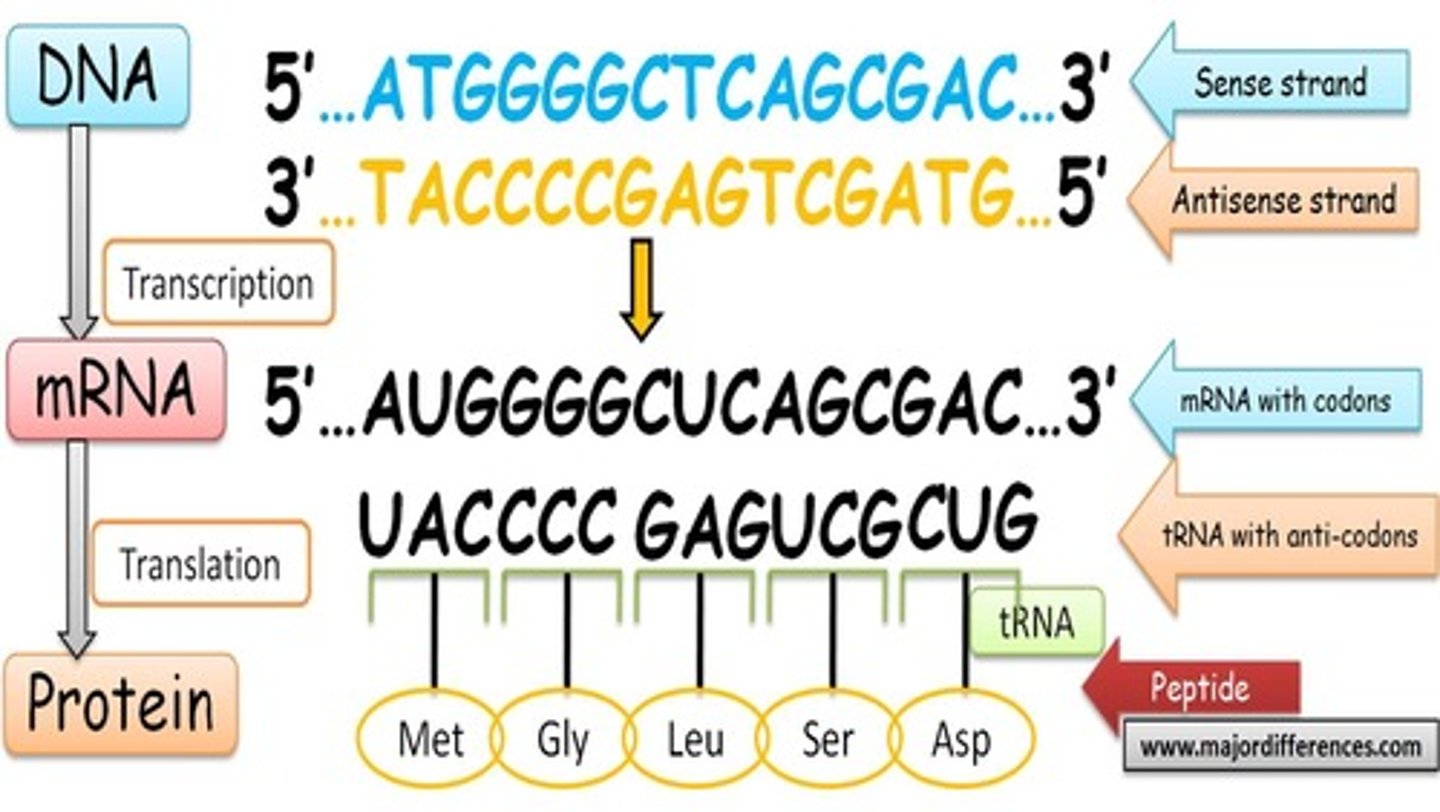
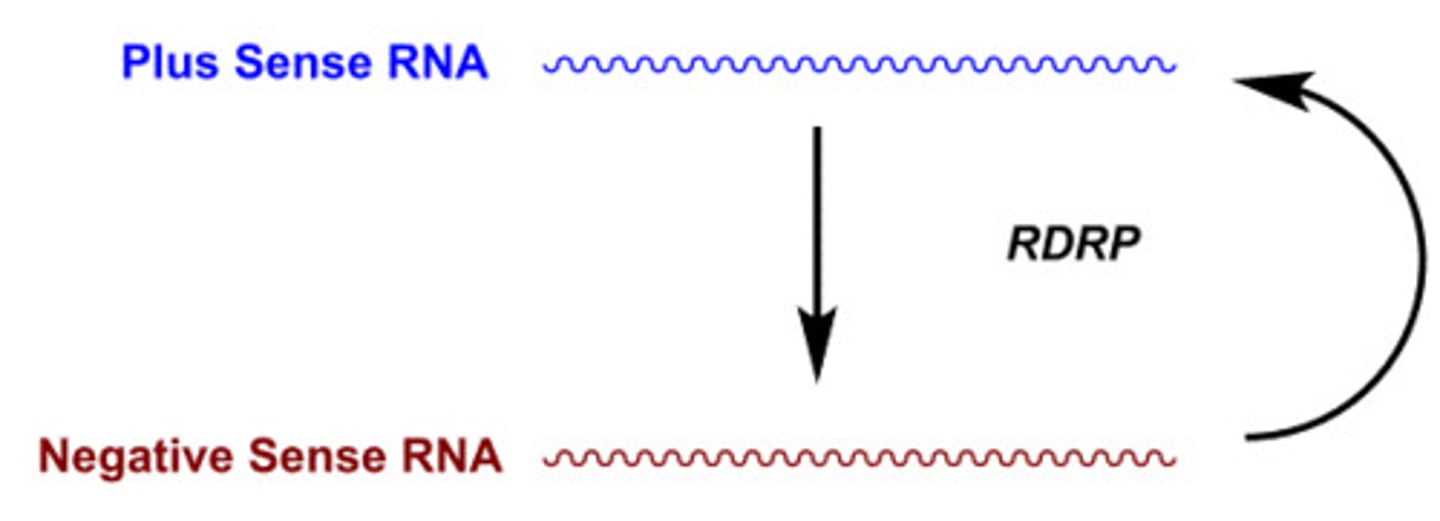
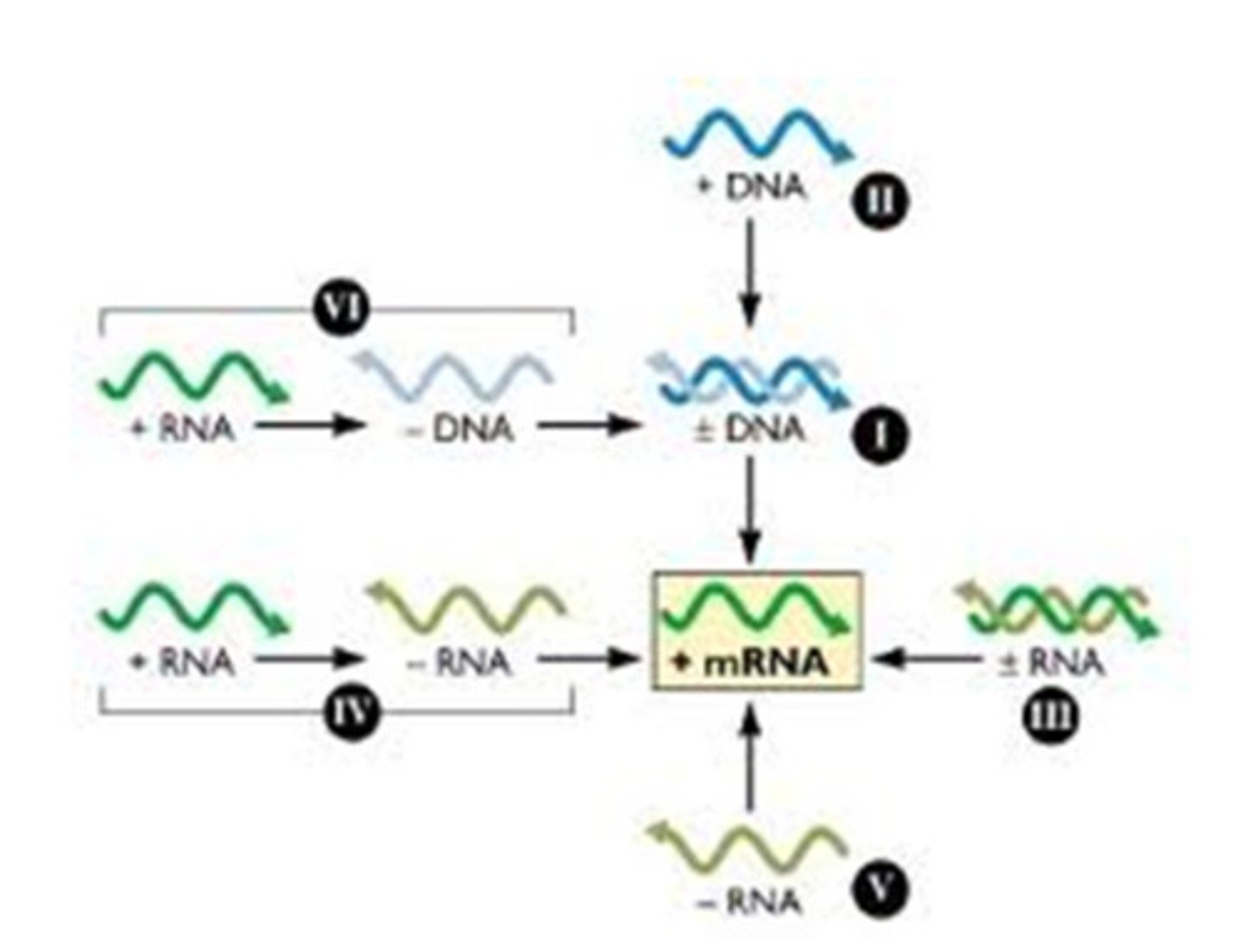
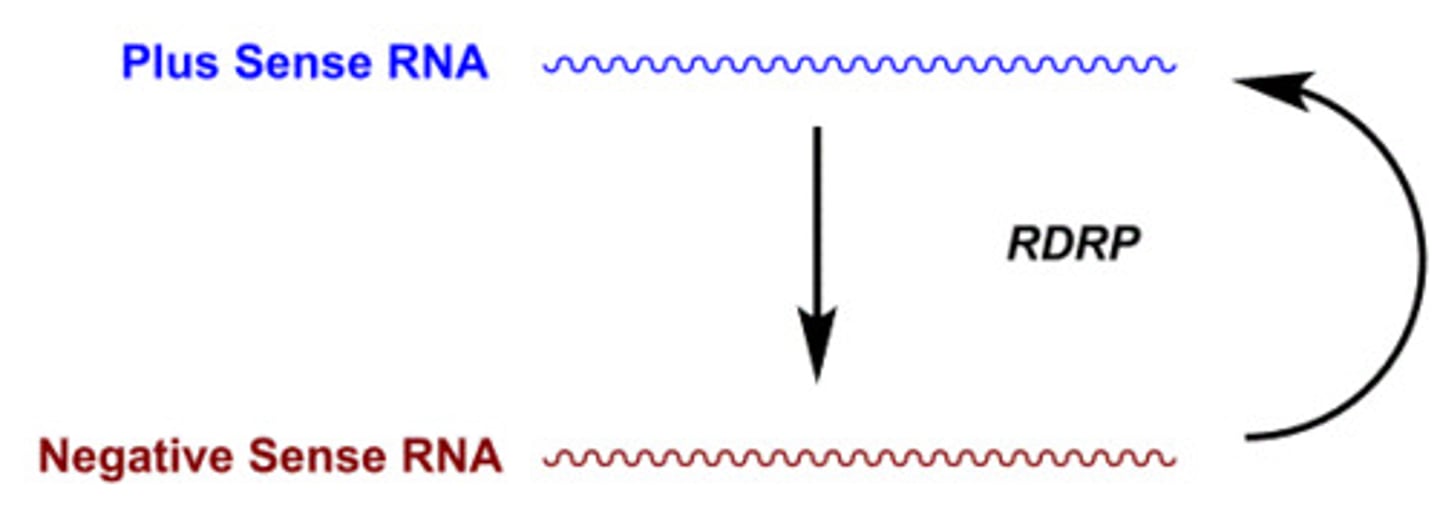
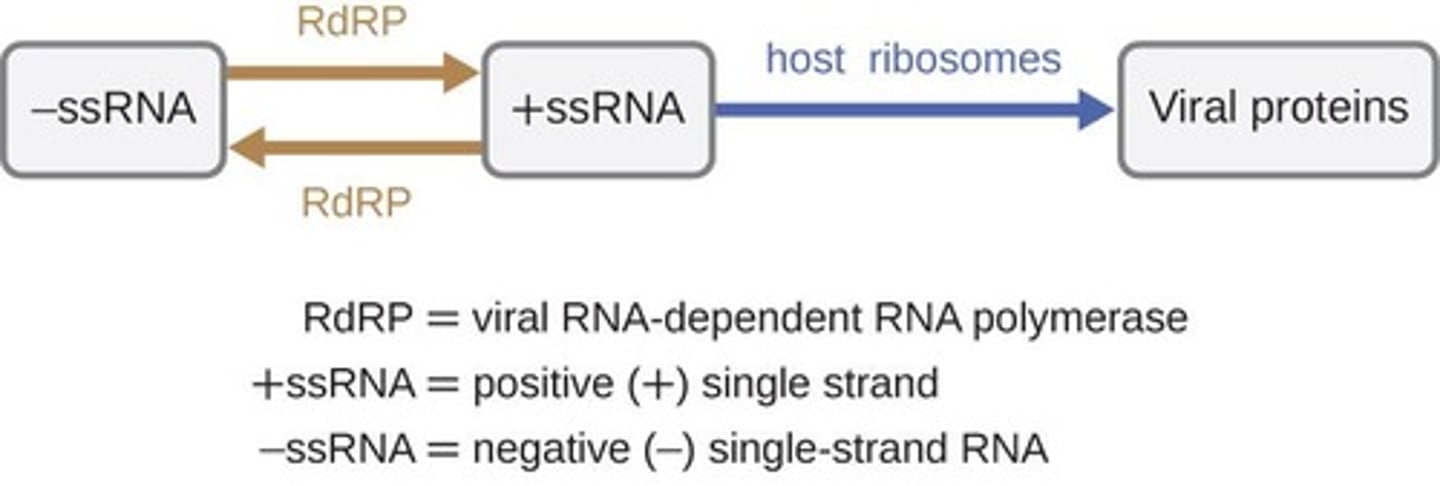
RNA-dependent RNA polymerase (RdRp)
synthesize RNA from an RNA template, playing a crucial role in the replication of RNA viruses and certain cellular processes
- present in viruses, plants, fungi, protists, and some animals
antisense RNA
• does not stay base-paired with the template long-term
• RdRP unwinds the antisense RNA during synthesis by strand displacement
• released as a separate single strand
Sense + antisense RNA can anneal
form dsRNA
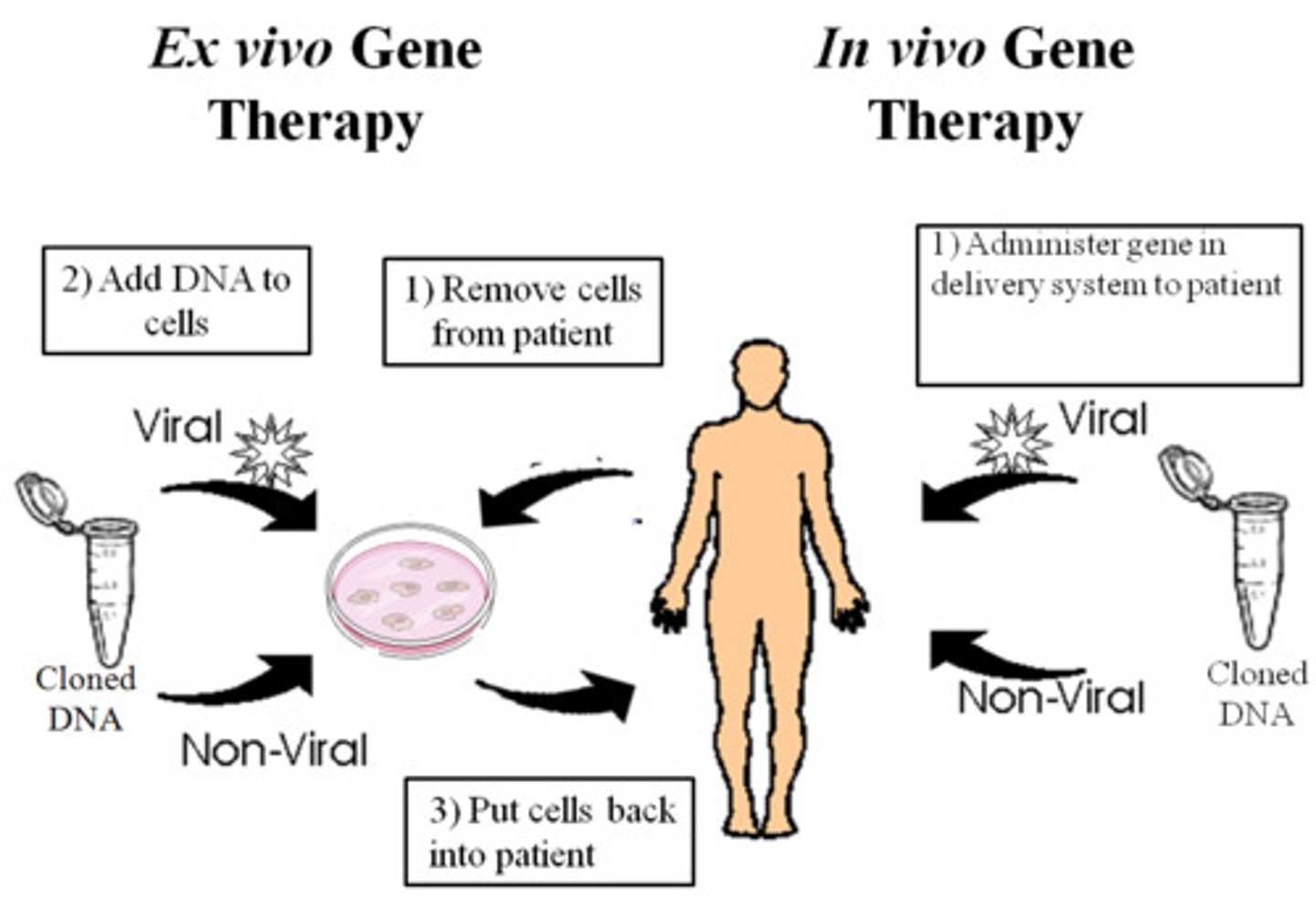
genetic engineering (cloning technology)
introduce antisense RNAs into human cells that are complementary to mRNA
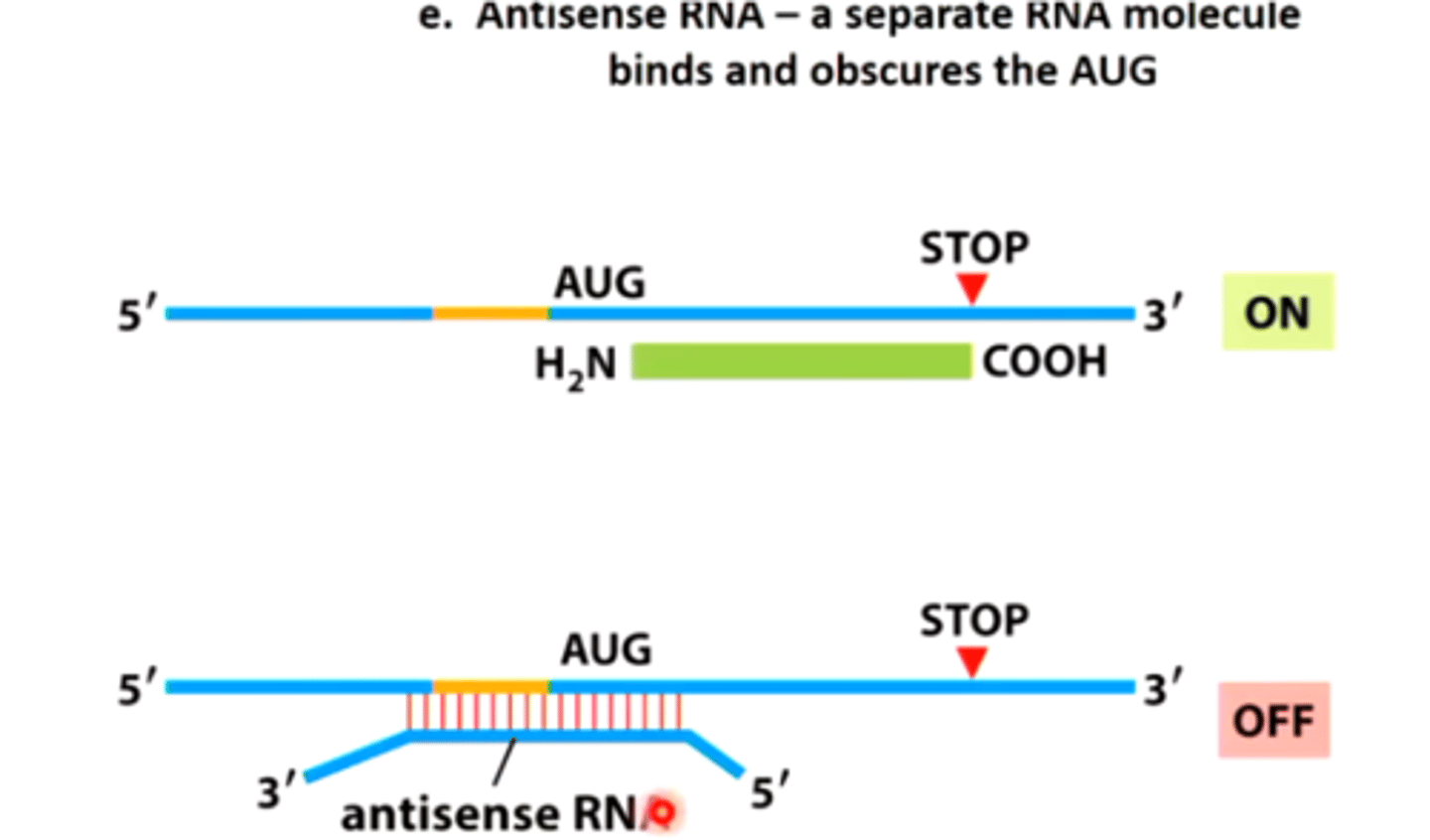
What is inserted in genetic engineering
provide animal cells with antisense RNA that is complementary to the mRNA
Result of genetic engineering
mRNA will be destroyed, and the protein that should be made from the mRNA will not be made
Gene is silenced
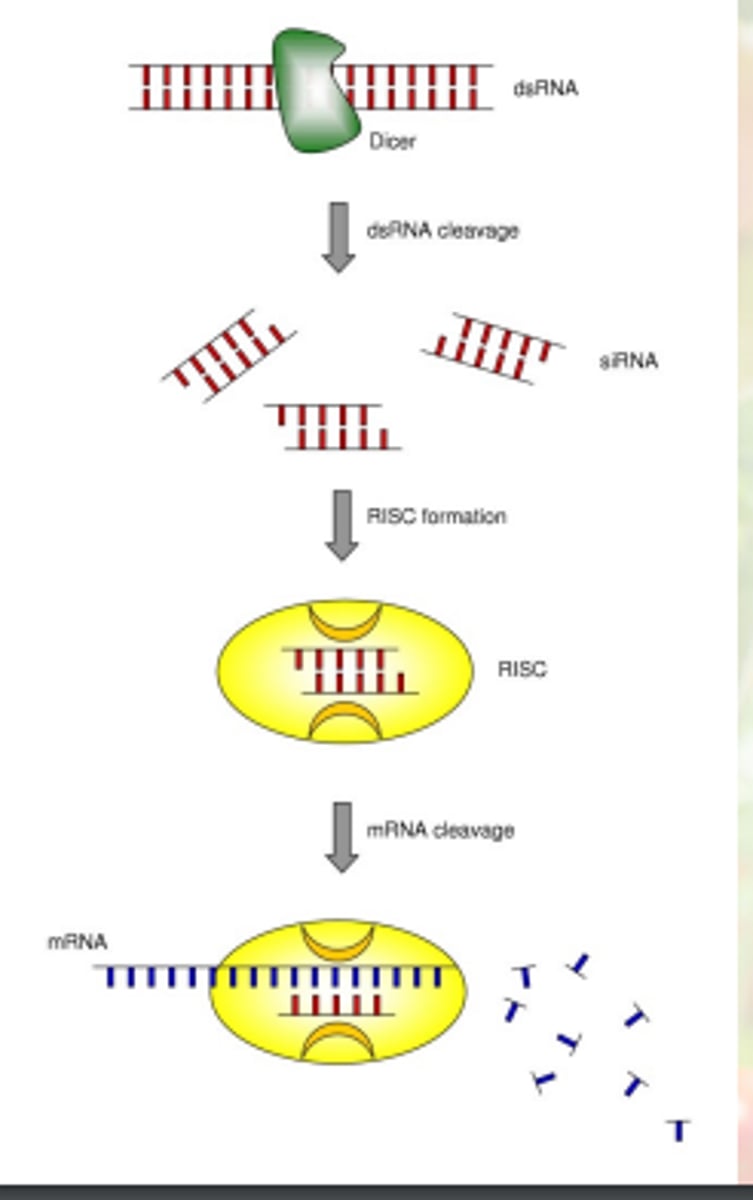
dsRNA is not degraded directly in this form
it is cleaved/cut into small pieces of dsRNAs called small interfering RNA(siRNA) through a cellular pathway
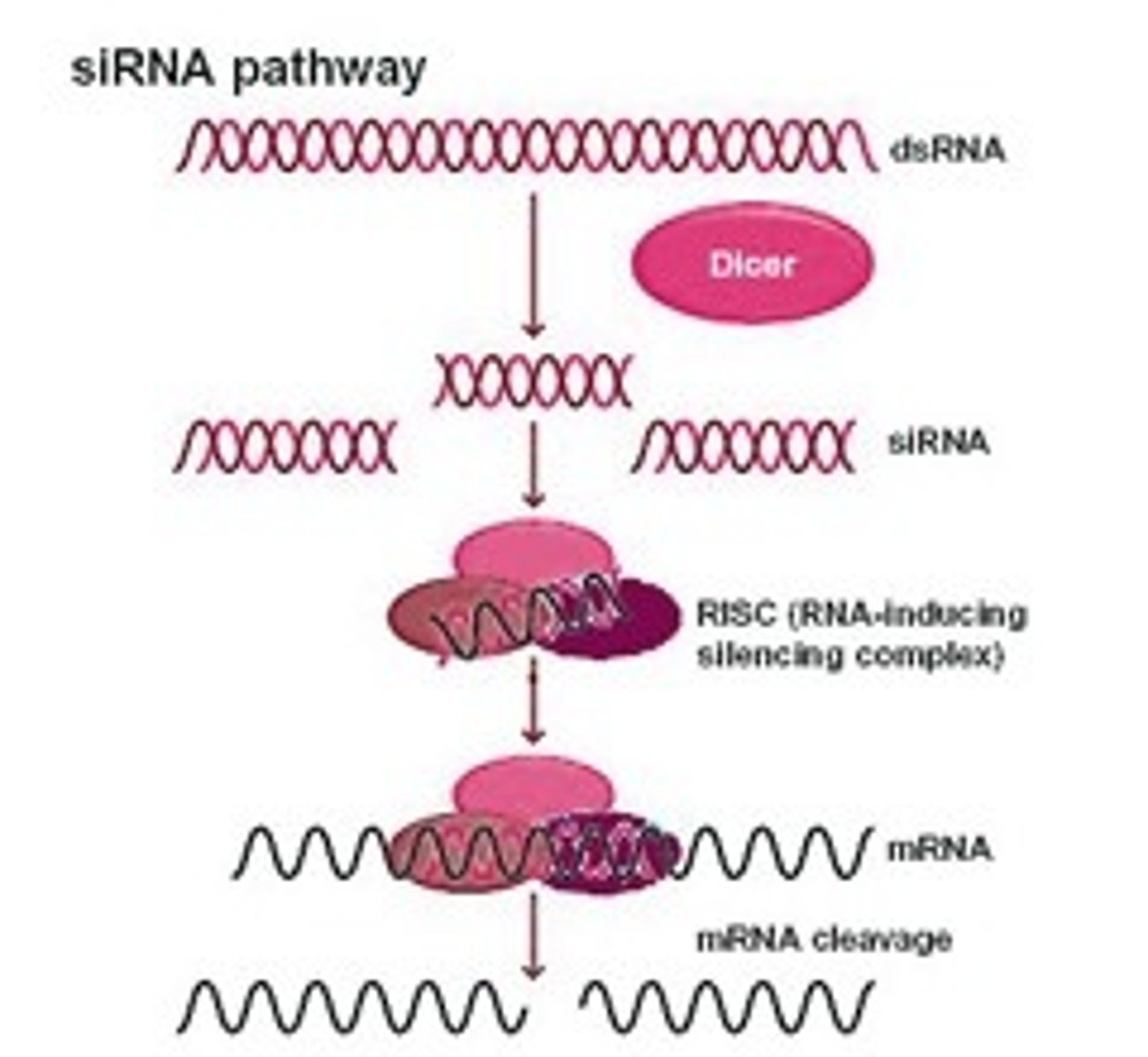
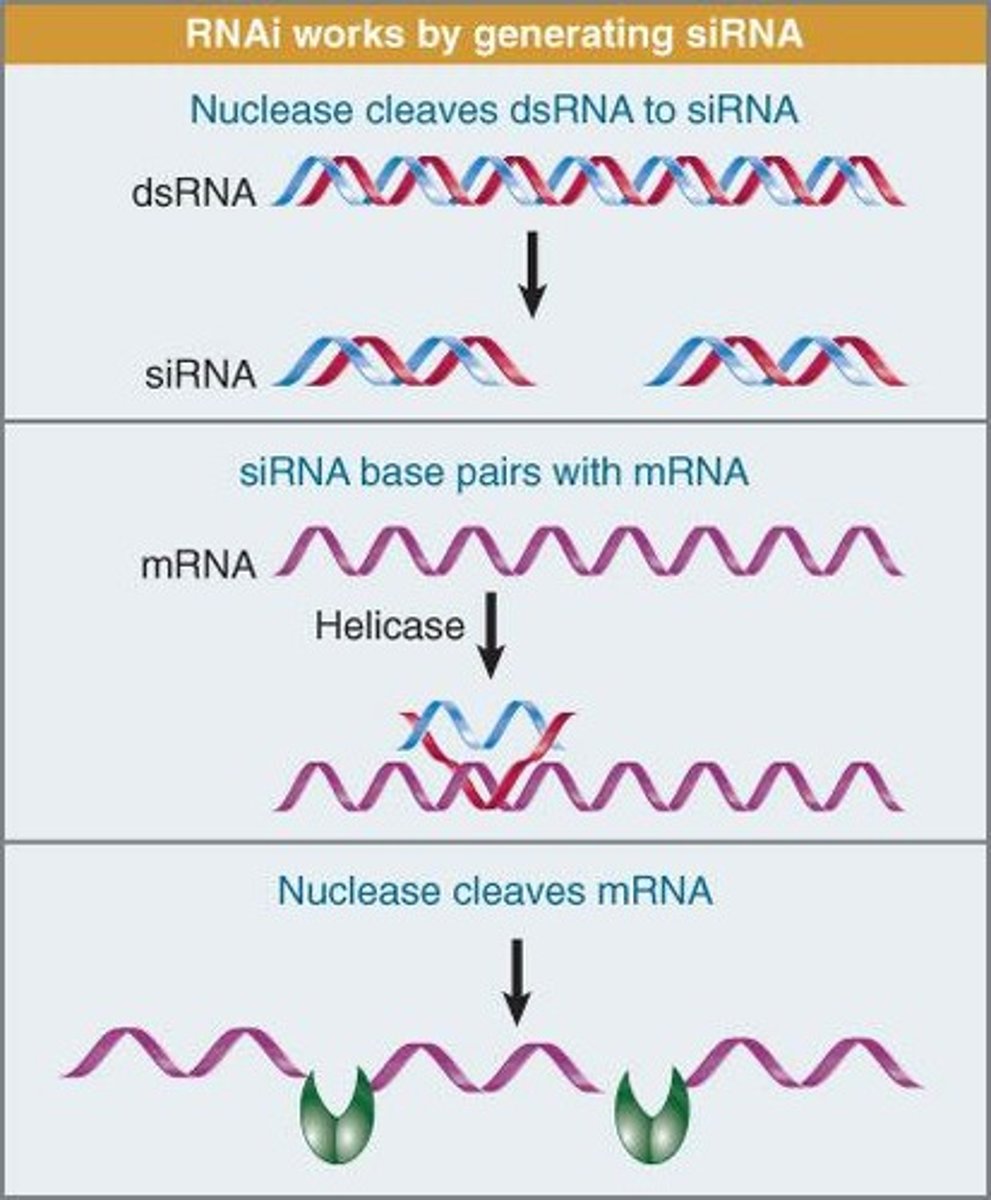
siRNA pathway
long dsRNA is cleaved into ds siRNA
dsRNA, siRNAs made into single strand siRNA; one strand is degraded
other ss siRNA bind to complementary mRNA and the mRNA is cleaved/degraded
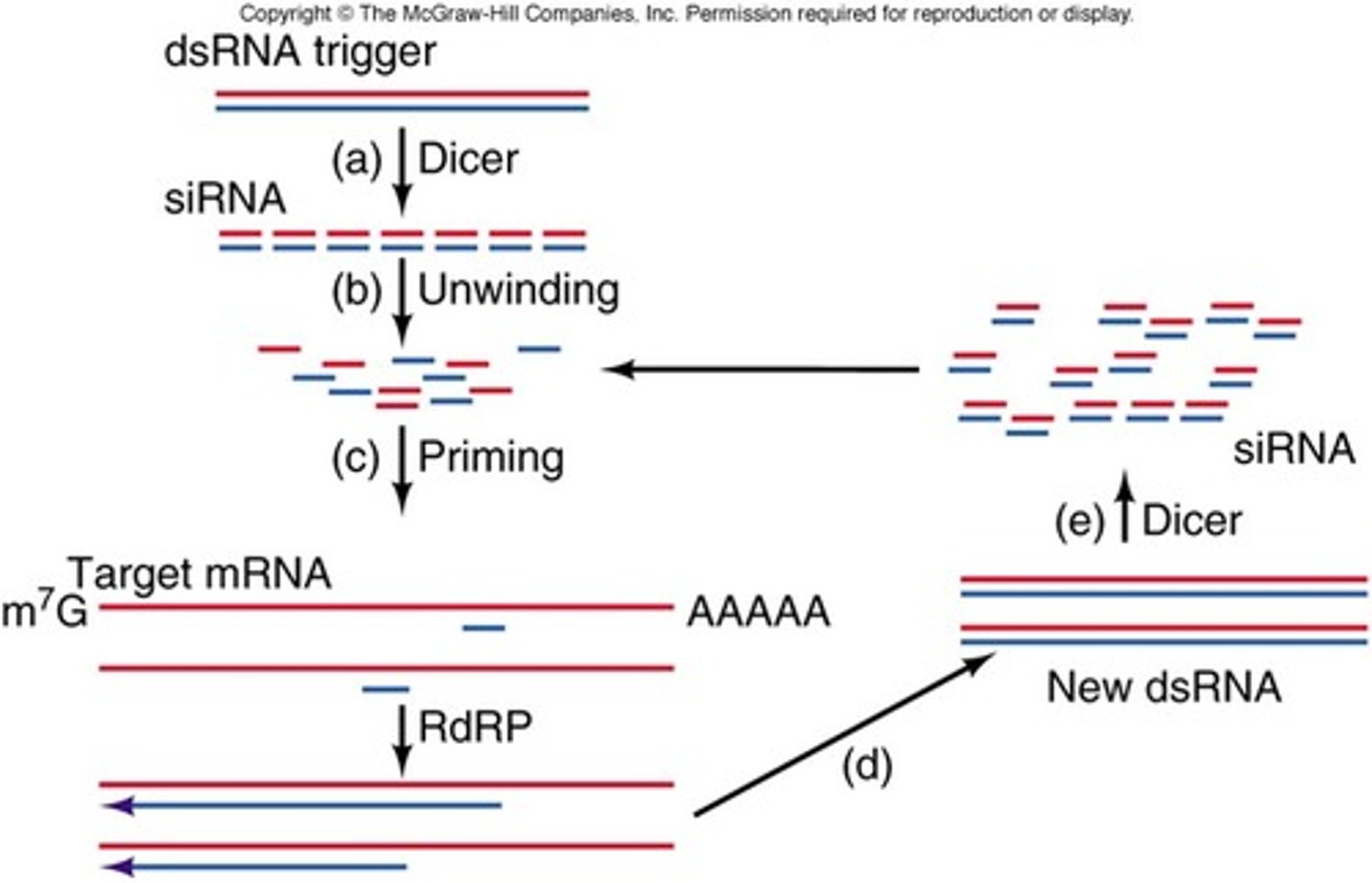
siRNA amplification
RdRP synthesizes antisense RNA using target mRNA, forming dsRNA that is processed into secondary siRNA
ENDOGENOUS
Genes already present in organism

Endogenous Sources of long dsRNA
RdRp can use mRNA (5 to 3') to make antisense RNA (3' to 5')
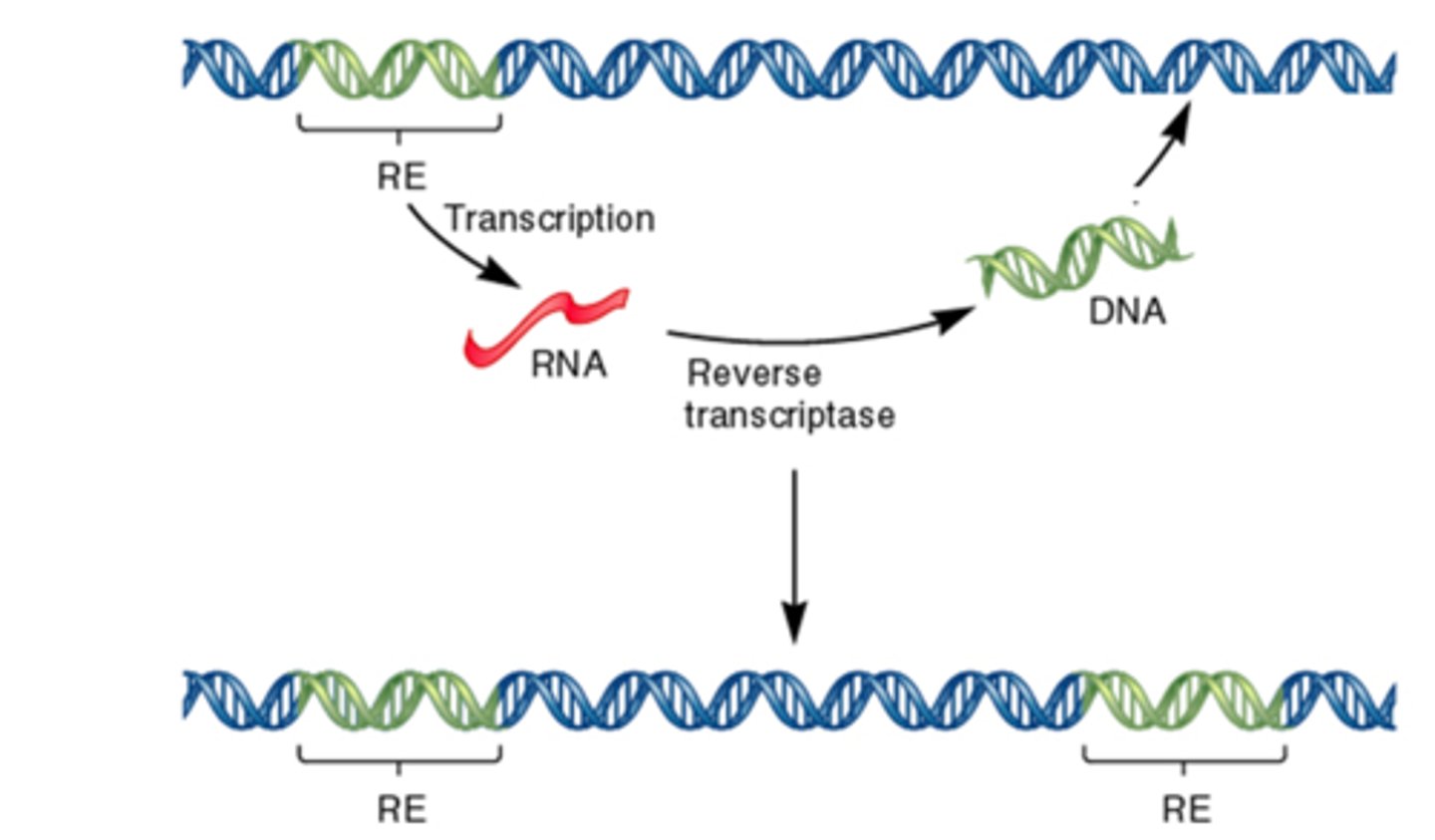
Transposable elements
EXOGENOUS
Adding more copies of already endogenous genes into organism
Exogenous Sources of long dsRNA
Viral infection
Artificial introduction
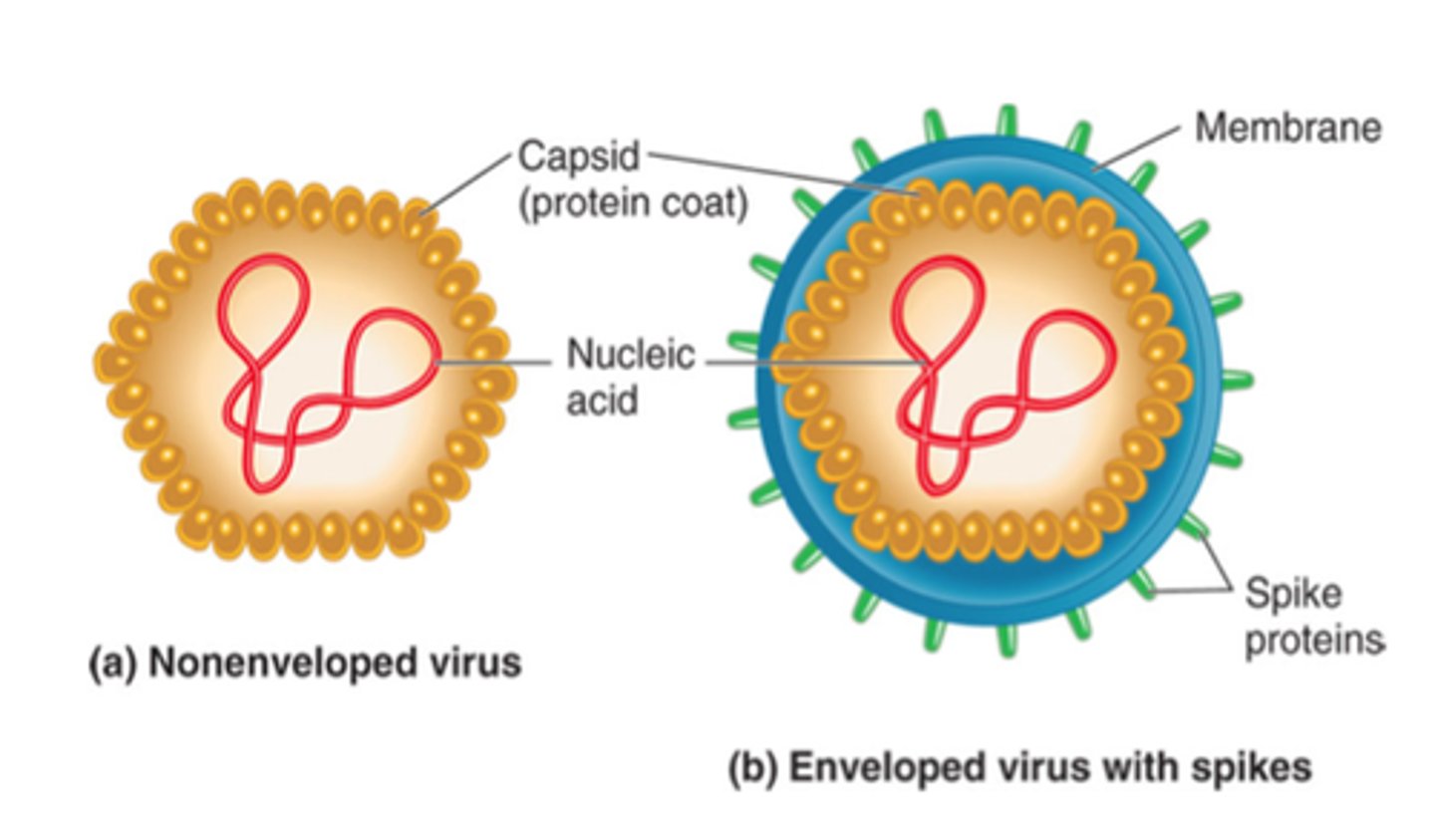
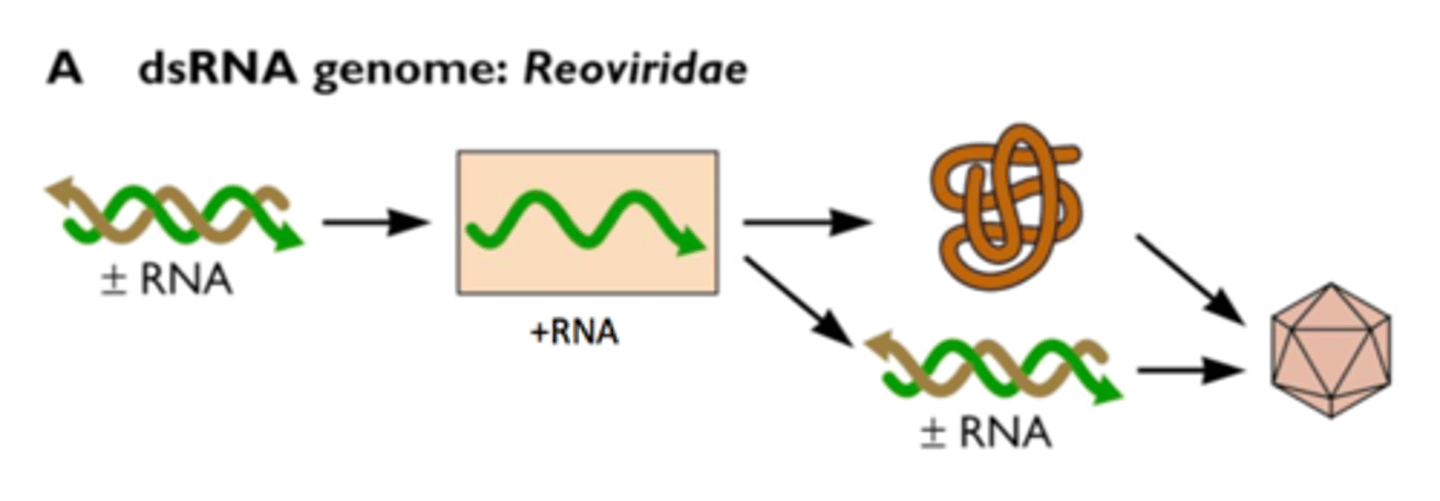
Viral infection
Many RNA viruses generate dsRNA intermediates during replication, which can activate the host RNAi response
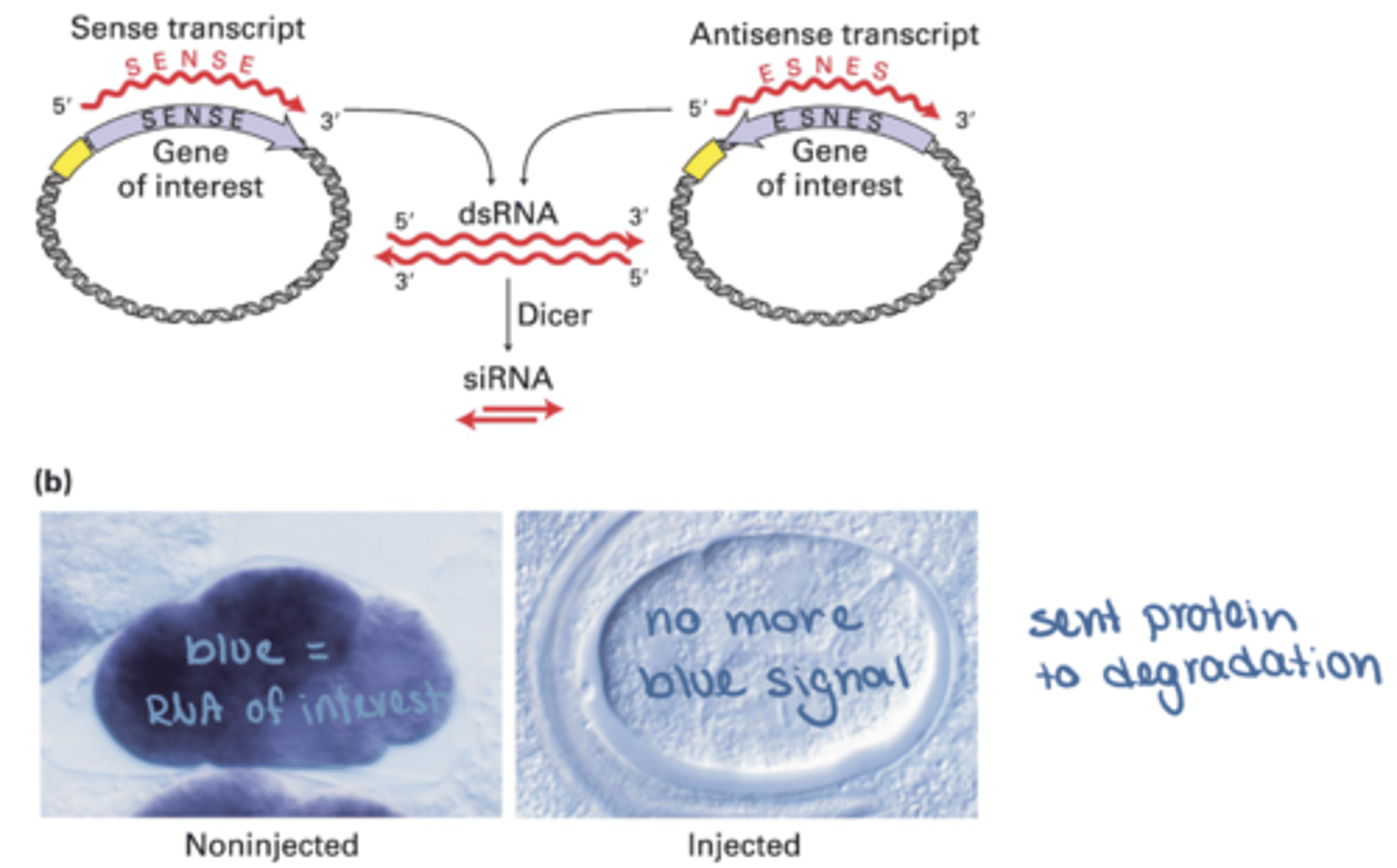
Artificial introduction
Researchers can introduce dsRNA into cells
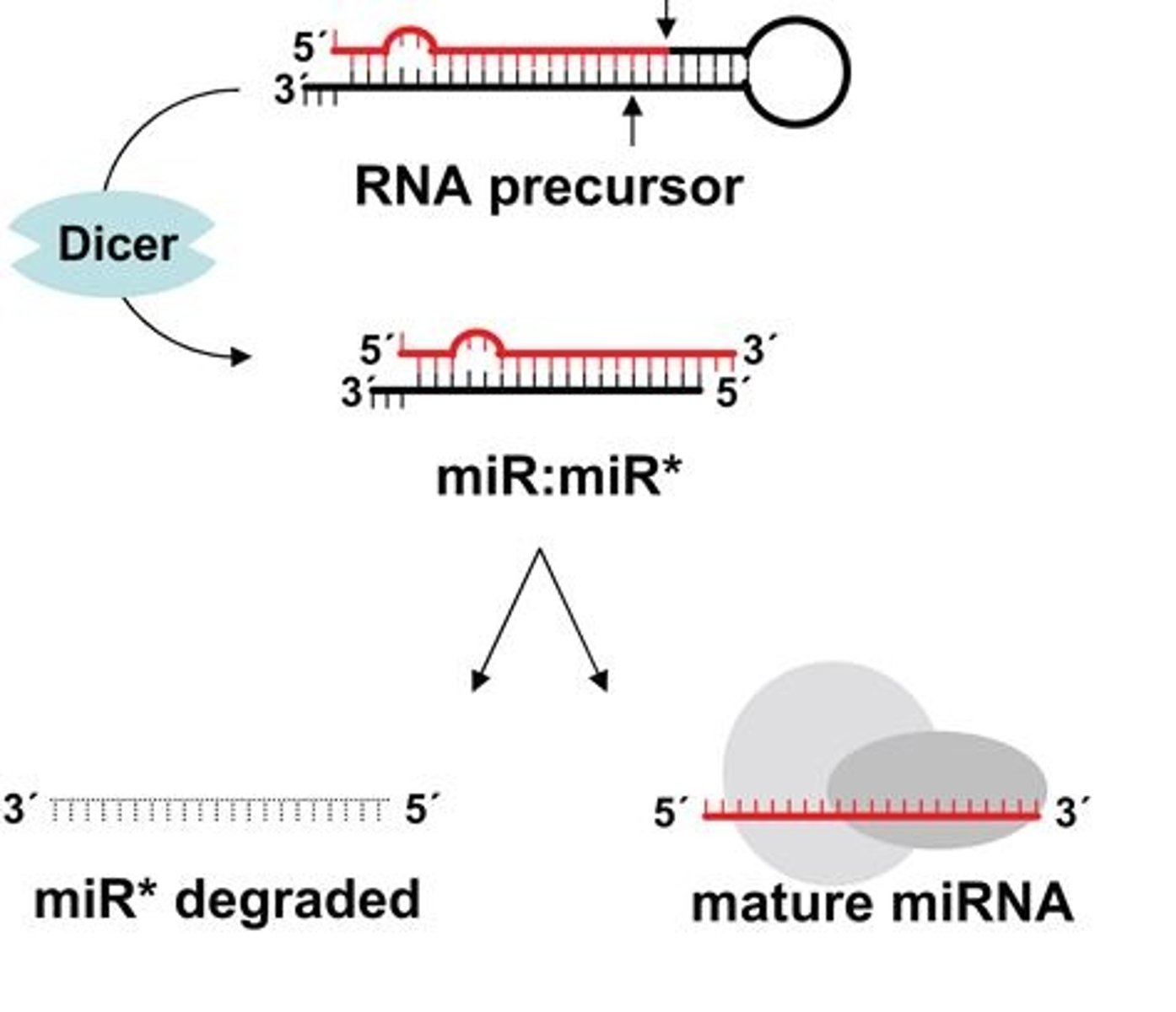
Synthetic siRNAs
Chemically synthesized siRNAs that mimic naturally processed siRNAs
Short hairpin RNAs (shRNAs)
Engineered RNA molecules that form dsRNA hairpins and are processed into siRNAs by the RNAi machinery
Vector-based expression systems
Plasmids or viral vectors can be engineered to produce dsRNA in cells
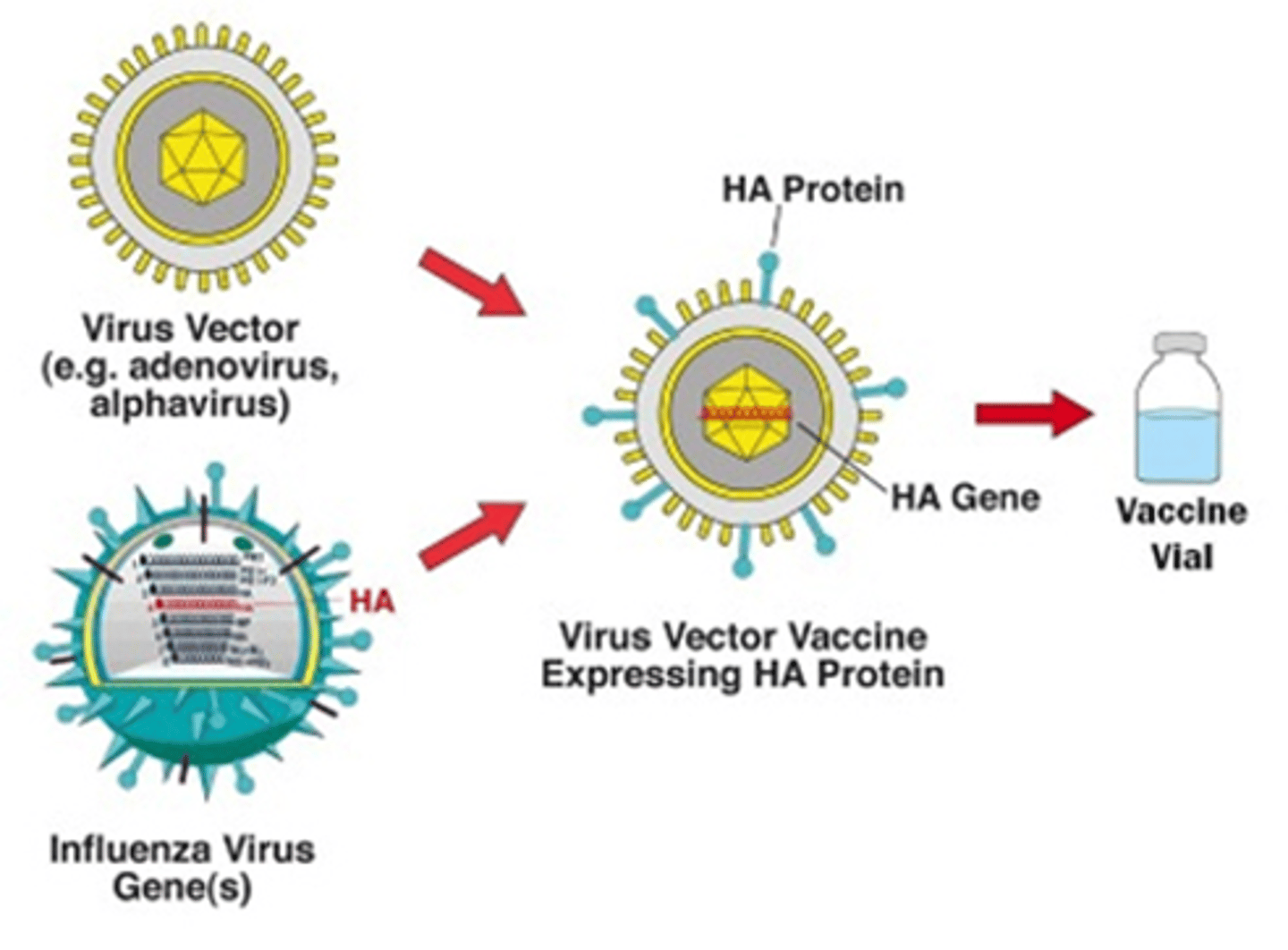
natural RNAi mechanism by siRNA is sequence-specific
a piece of RNA (~22) binds to a complementary sequence on the mRNA perfectly, which leads to degradation of the mRNA
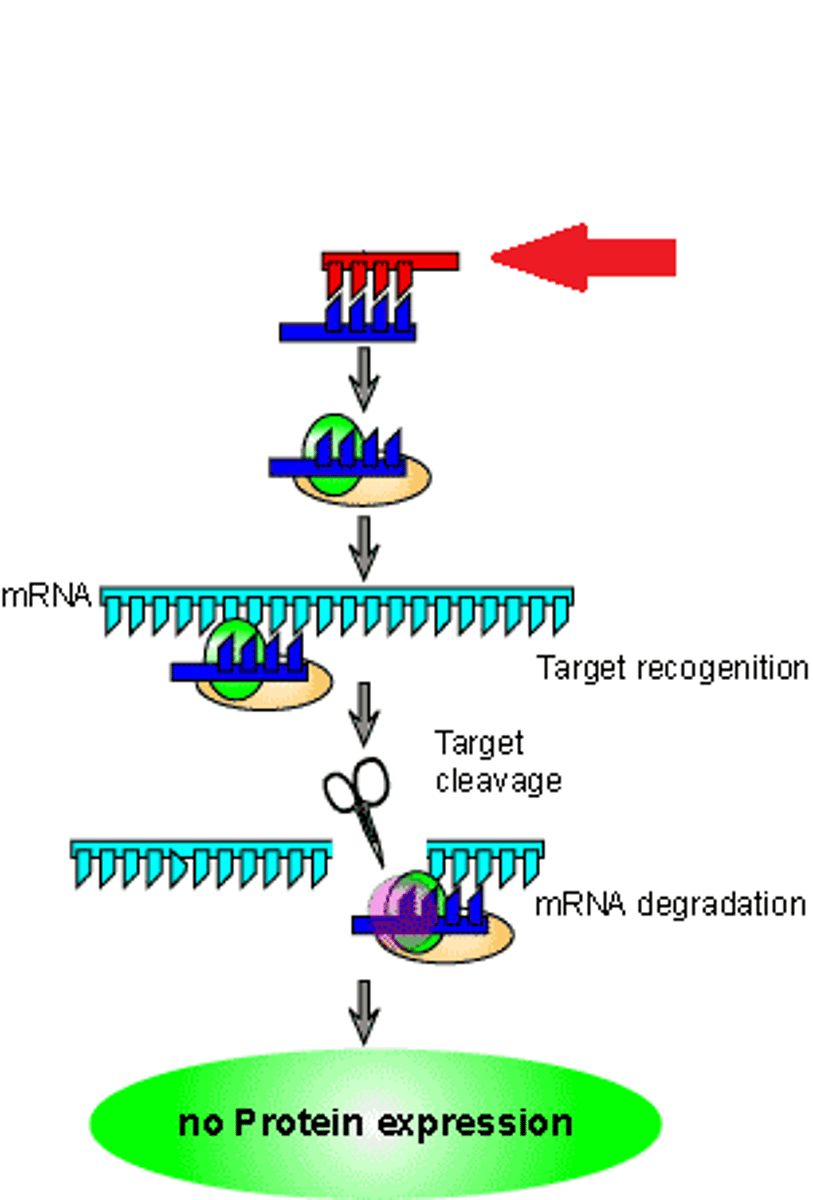
RNAi by siRNA is a conserved biological response
RNAi mediated by siRNA have been found in many organisms, including plants, fungi, worms, Drosophila, mice, and humans
siRNA destroys dsRNA, and by doing so
mediates protection to both endogenous parasitic and exogenous pathogenic nucleic acids
regulates the expression of protein-coding genes
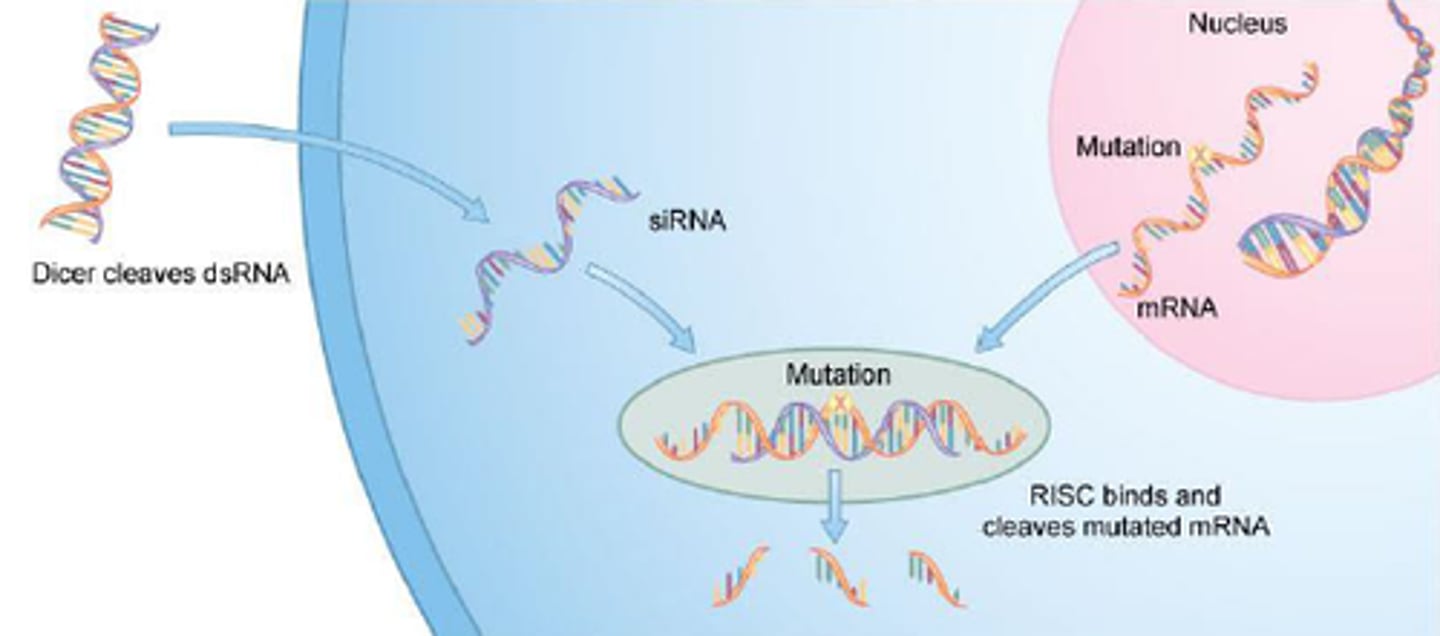
Discovery of PTGS mediated by small interfering RNAs (siRNA)
discovered as a naturally occurring pathway accidentally in experiments with TRANSGENES
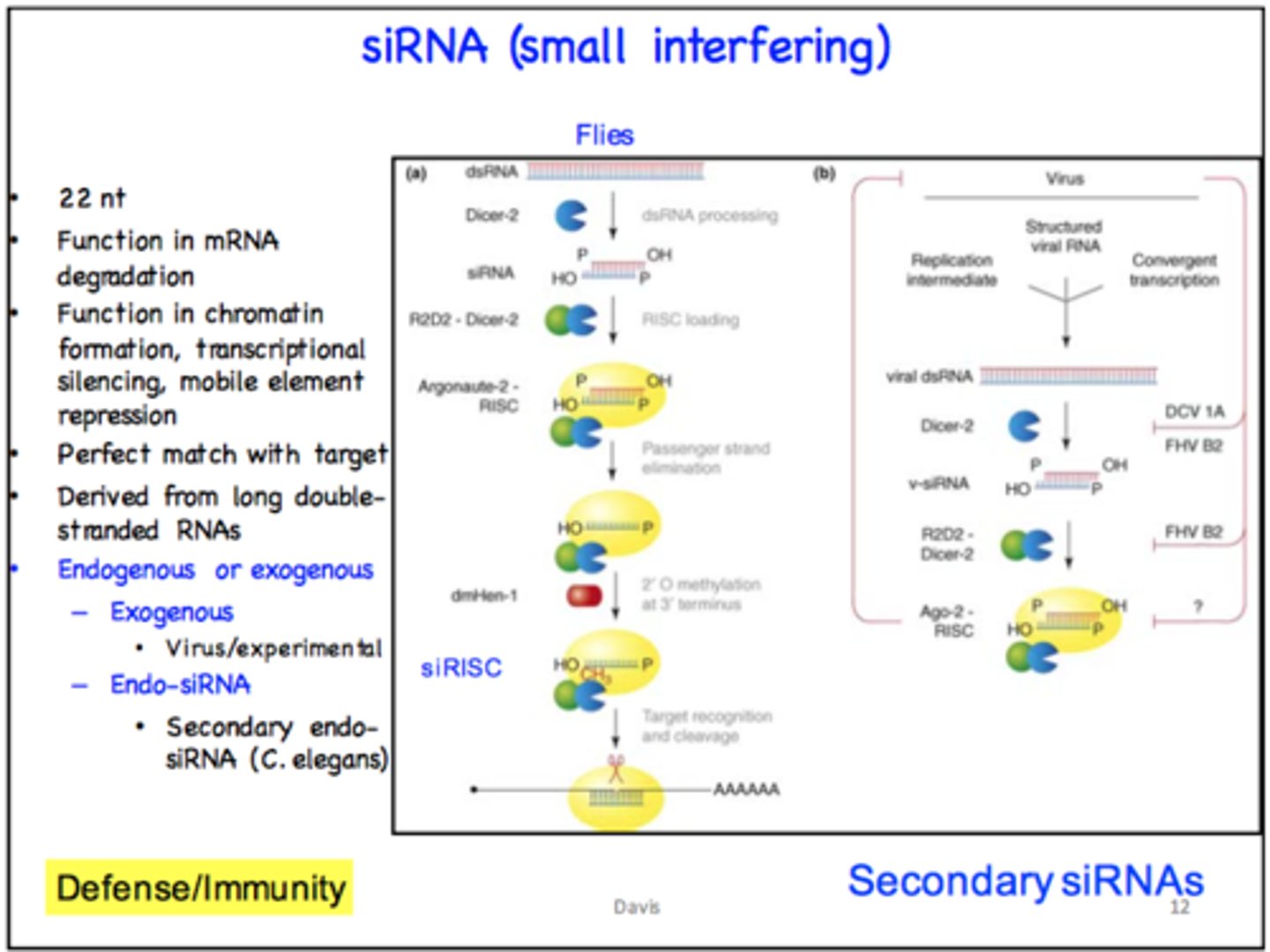
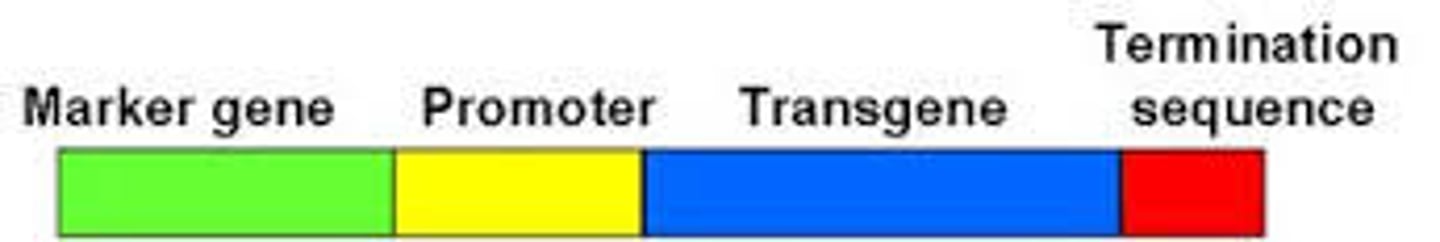
What is a transgene?
gene introduced into an organism using genetic engineering techniques, either from another organism or as additional copies of an existing gene to enhance the production of its encoded protein
Where must a transgene get into for transcription to occur?
The nucleus of the cell.
transgenesis/recombinant DNA technology
has the potential to change the phenotype of an organism
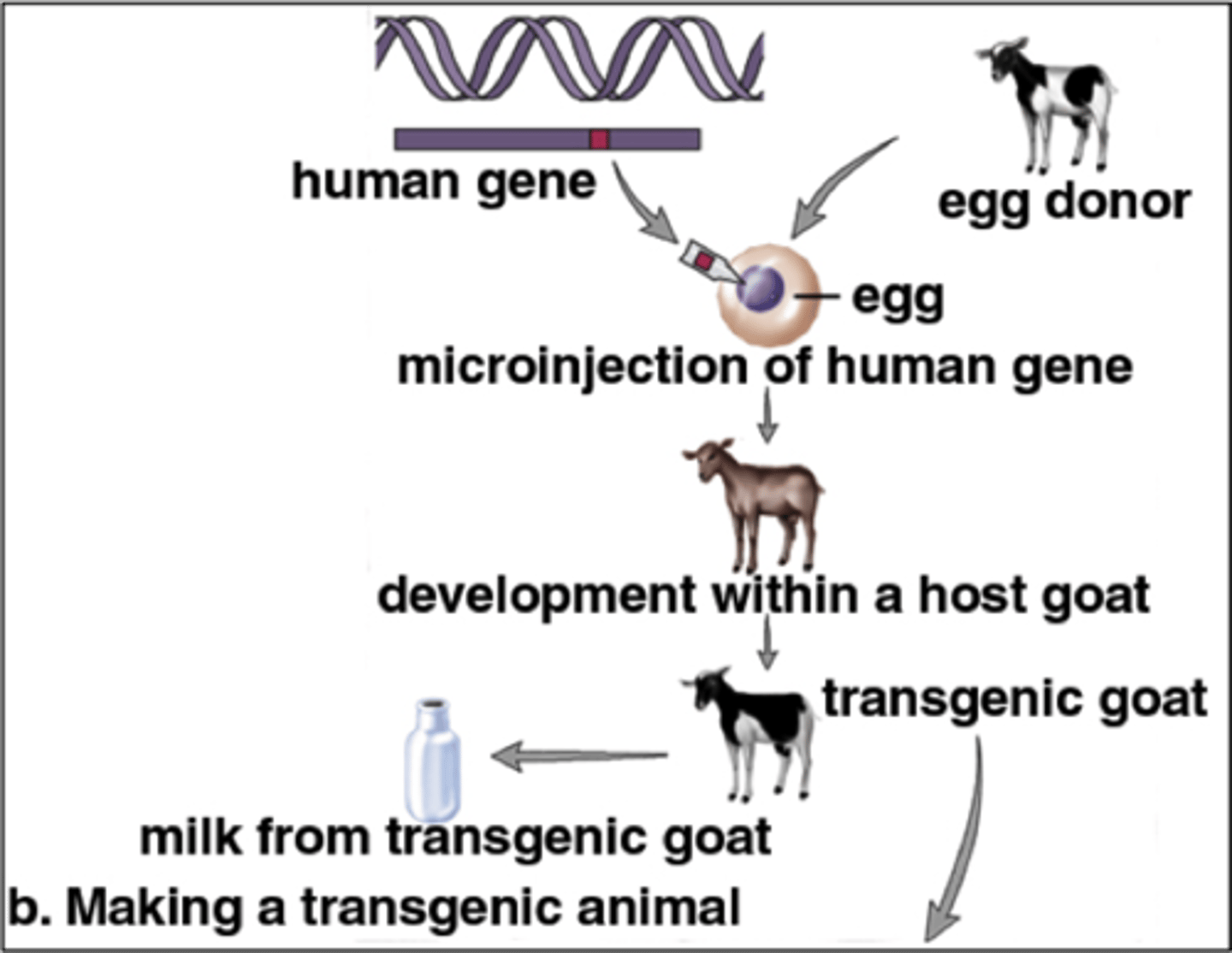
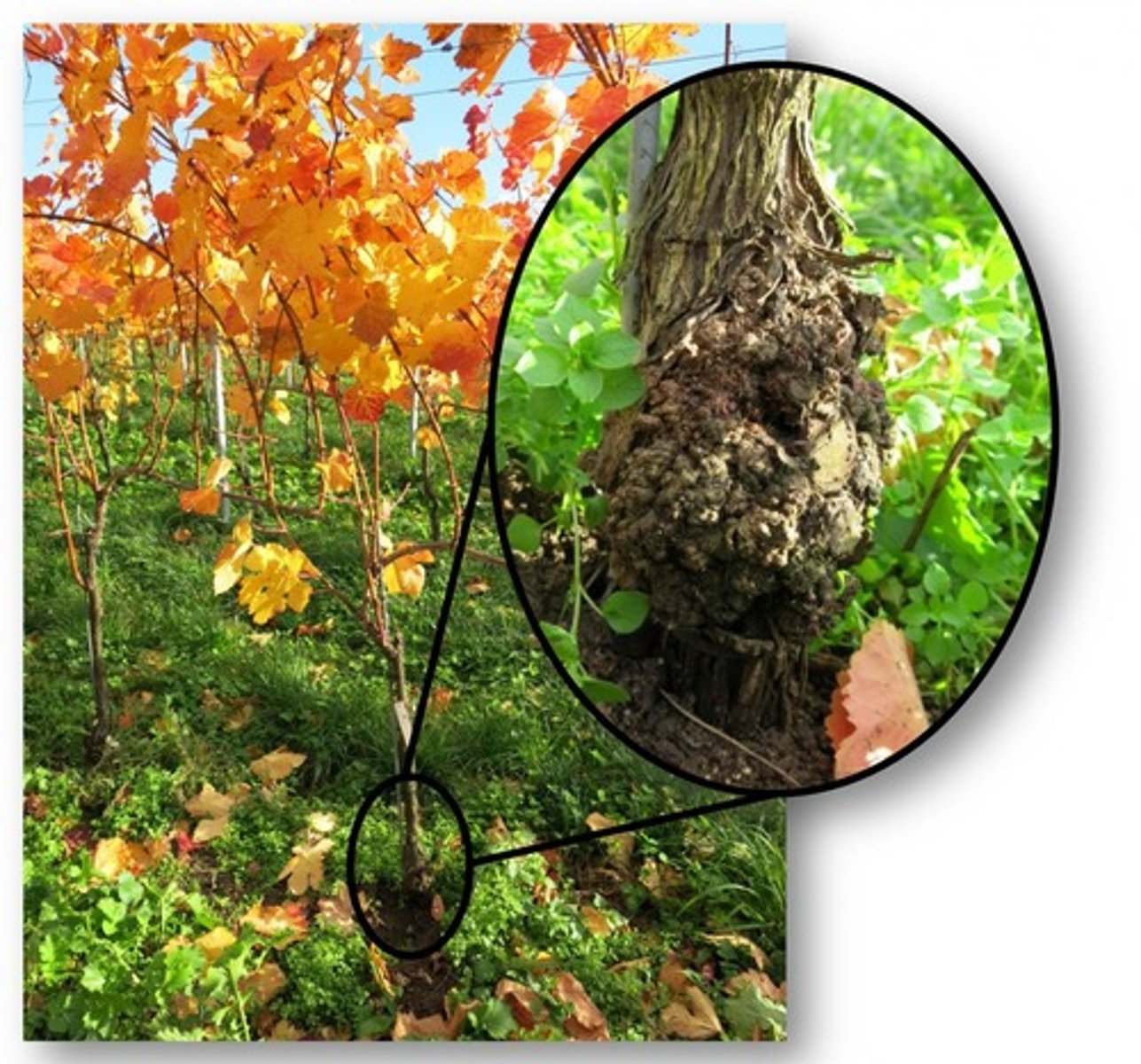
agrobacterium tumefaciens
used to introduce genes into plant genomes

Transgene induced post transcriptional gene silencing
Transgene X makes antisense RNA X
Endogenous gene X makes mRNA X
RNA X from the transgene X binds to the endogenous mRNA X
mRNA does not make a protein
Experiment that determined TGS
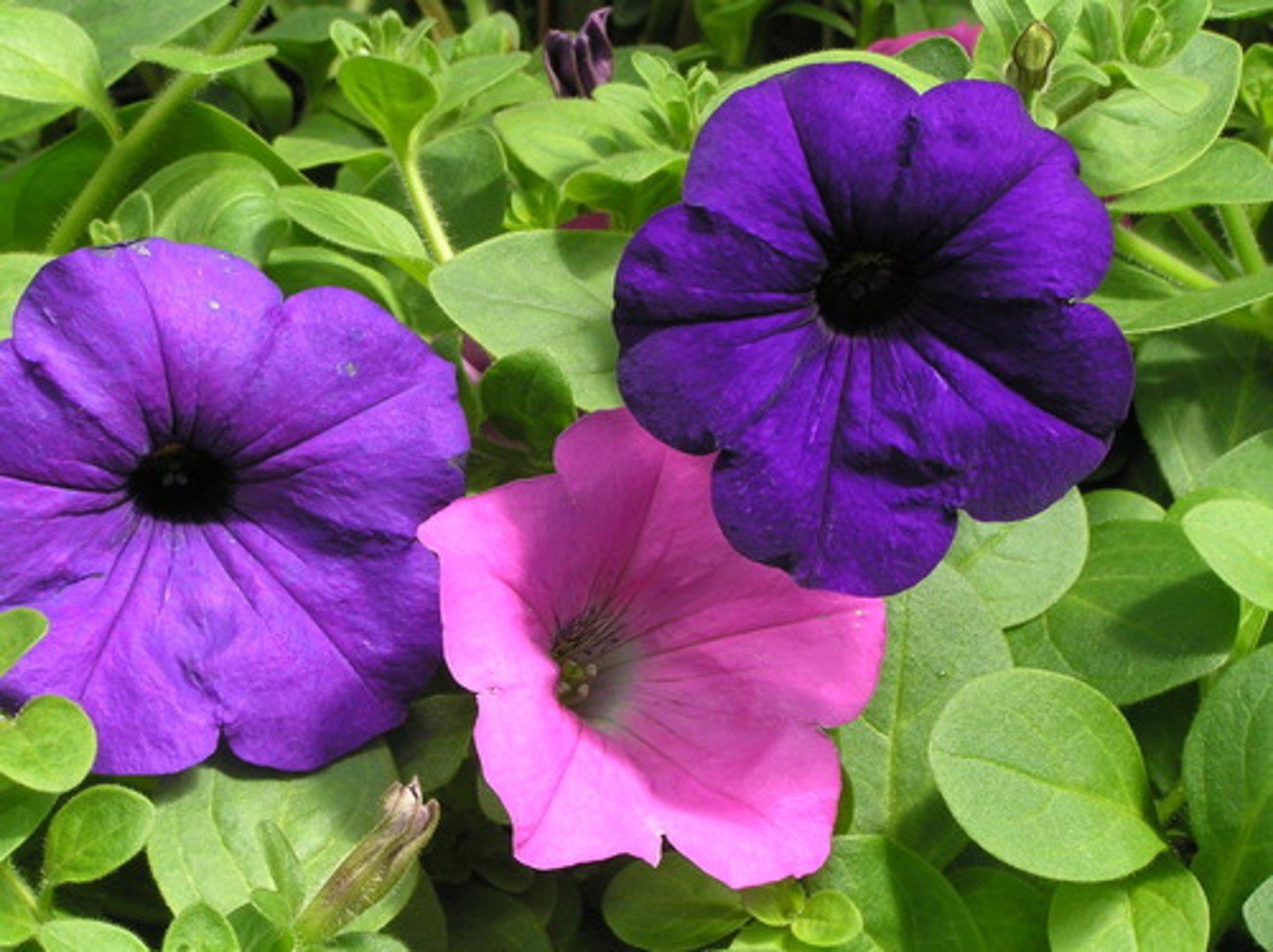
They wanted to make Intensely purple petunia flowers and White petunia flowers

Chalcone synthase
anthrocyanin pigment gene in petunia
enzyme at start of biosynthetic pathway for anthocyanins
More copies of CHS gene
more CHS mRNA --> more CHS enzyme to convert --> more substrate (coumaric acid) into chalcone --> leading to more anthocyanins --> deeper coloration

antisense CHS mRNA will bind to sense CHS mRNA
less CHS enzyme --> less anthocyanins -->white / whitish coloration
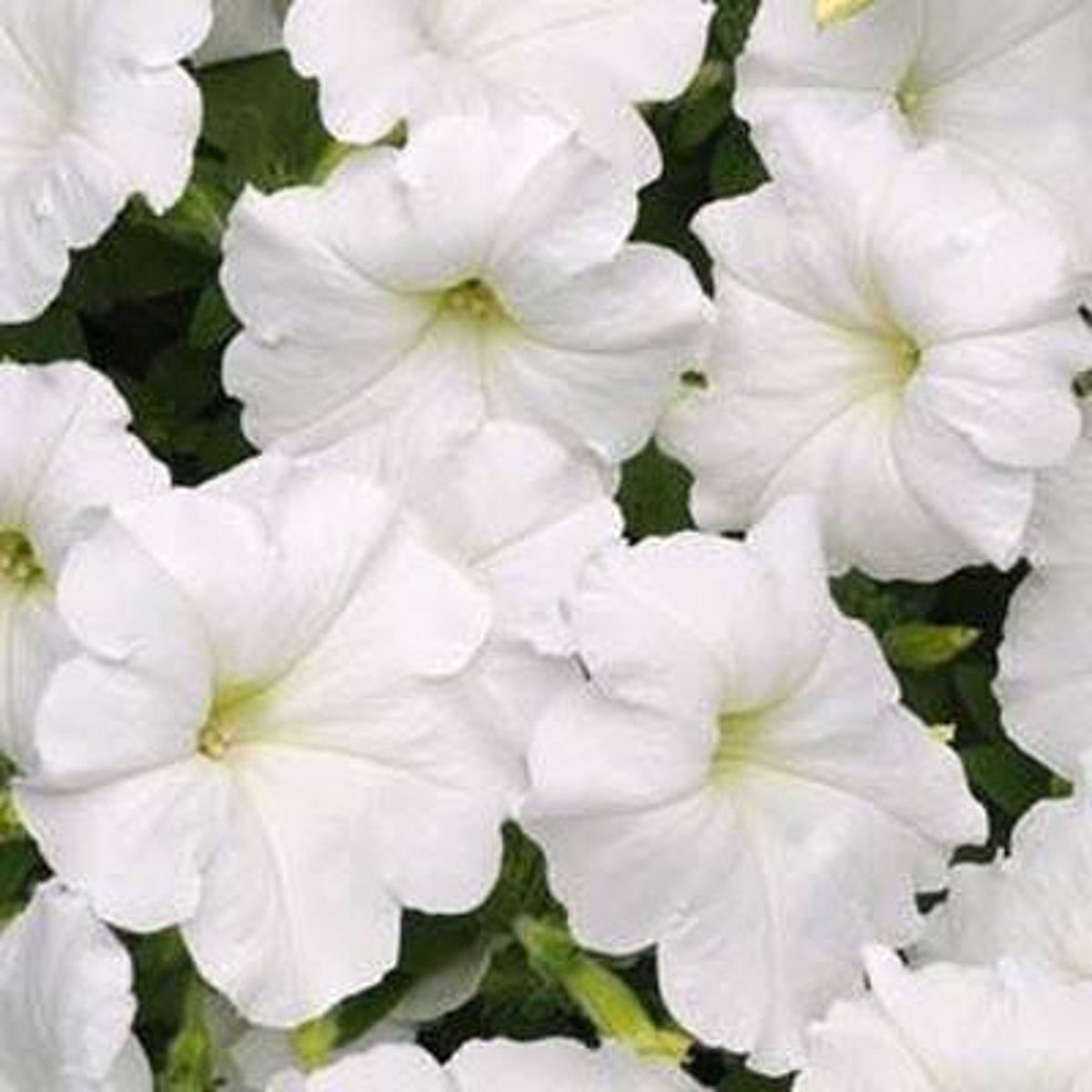

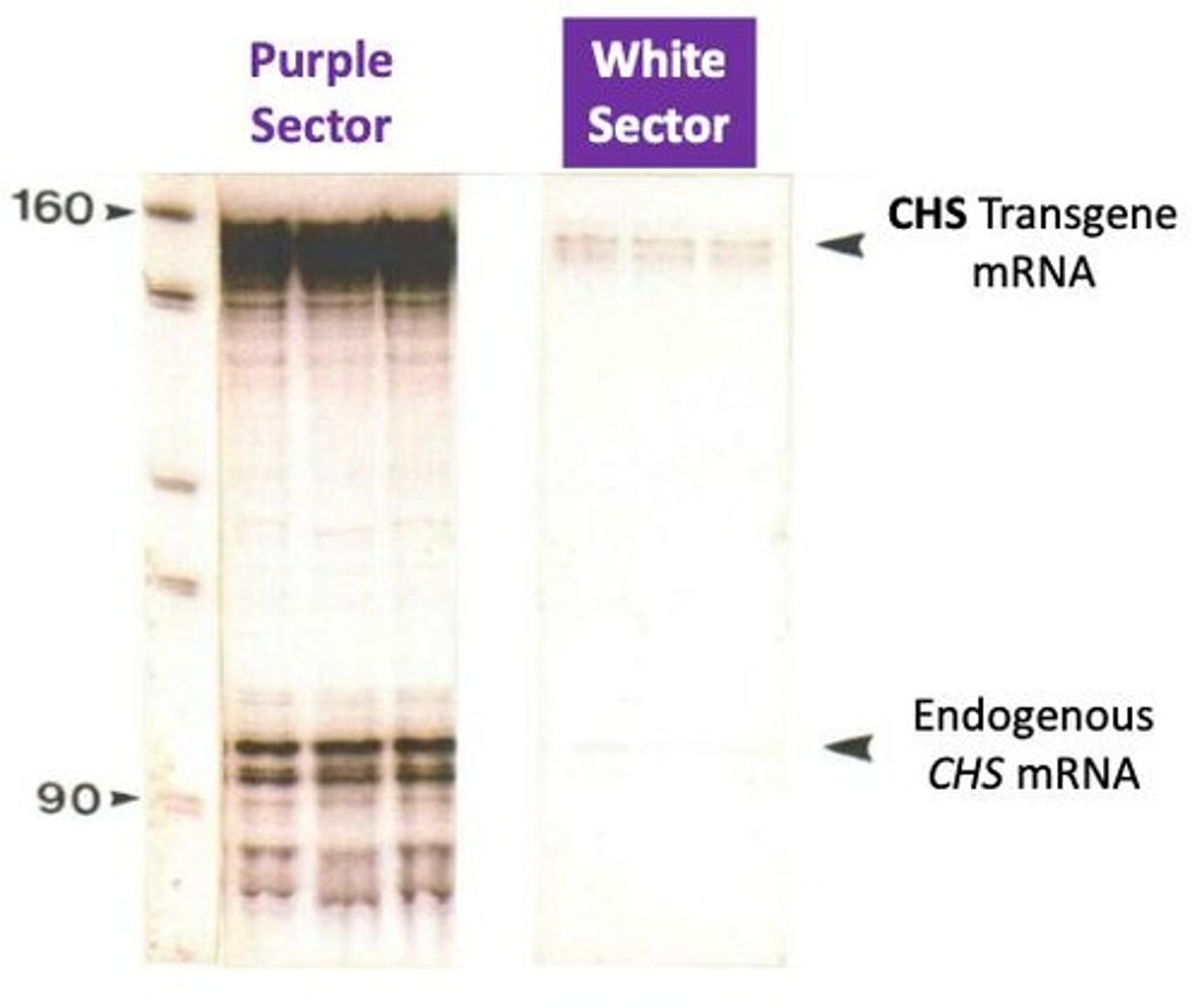
Result of experiment
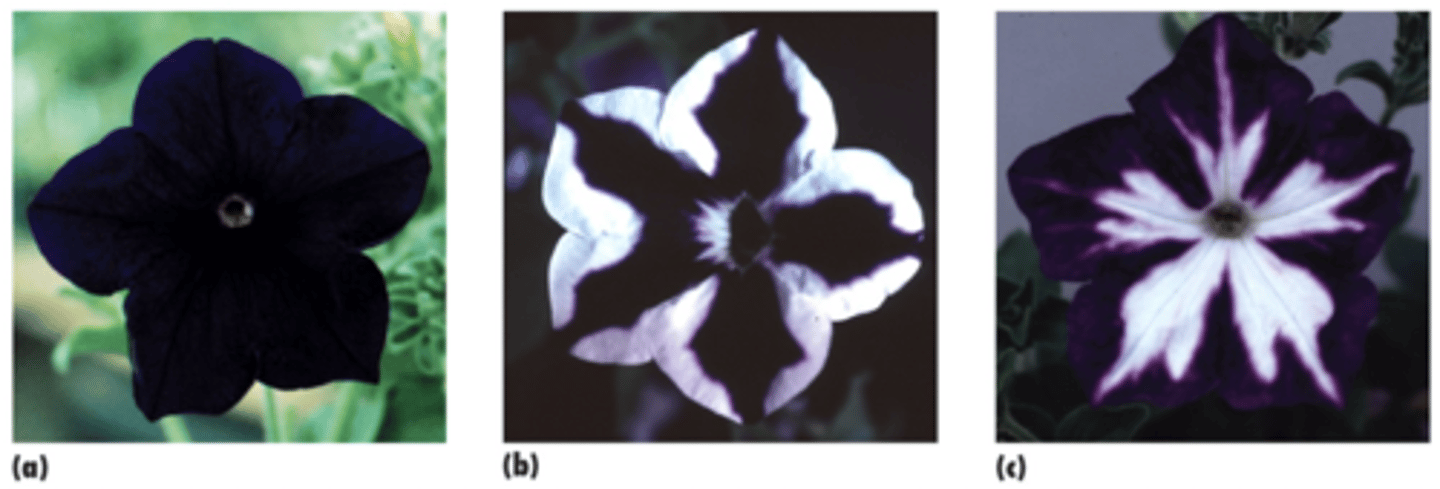
they saw a mix of deeper colored flowers (expected) loss of pigment (unexpected)
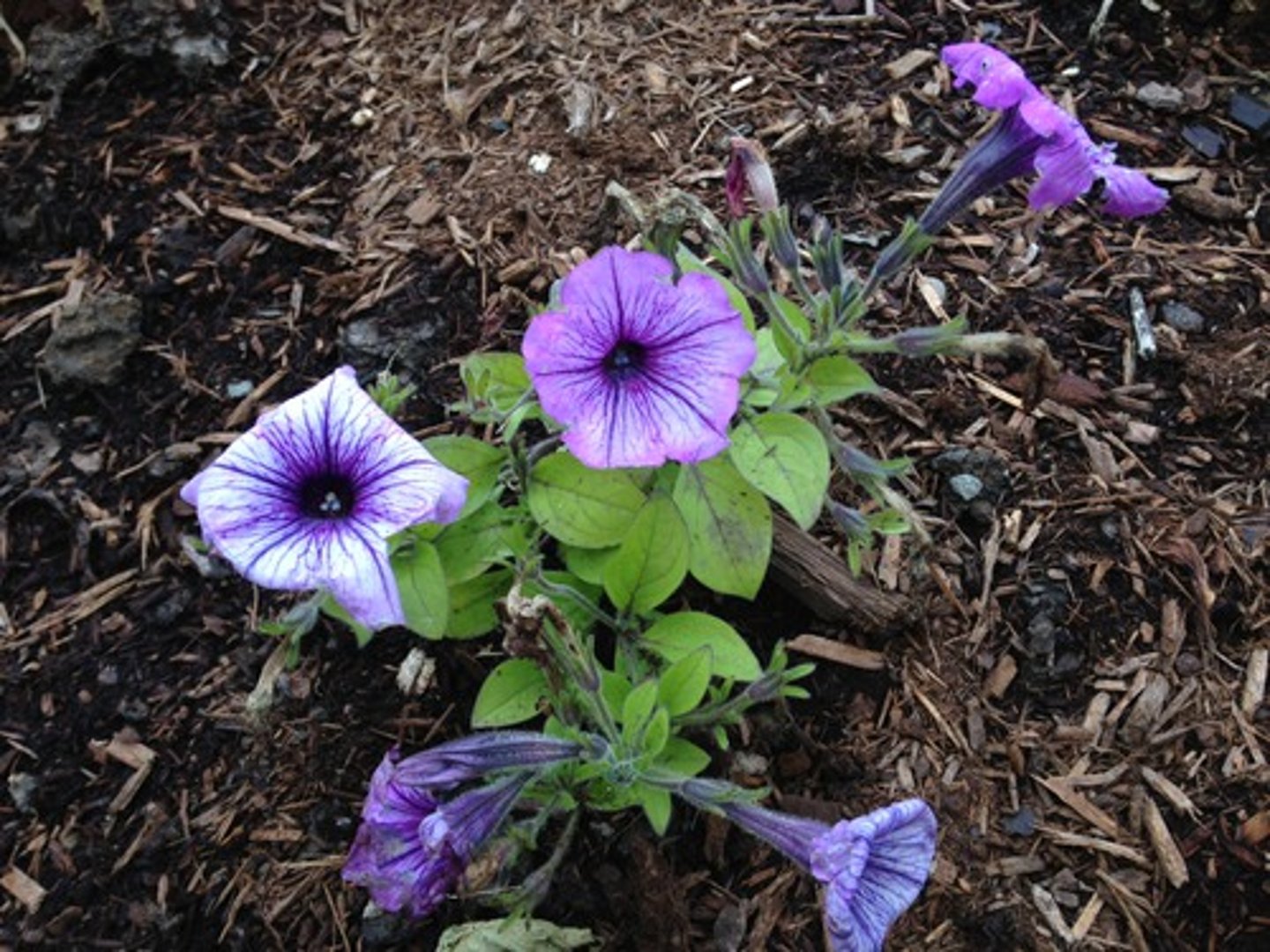

the sense construct
Instead of exhibiting more pigment, most petunias displayed reduced color, with some areas completely white
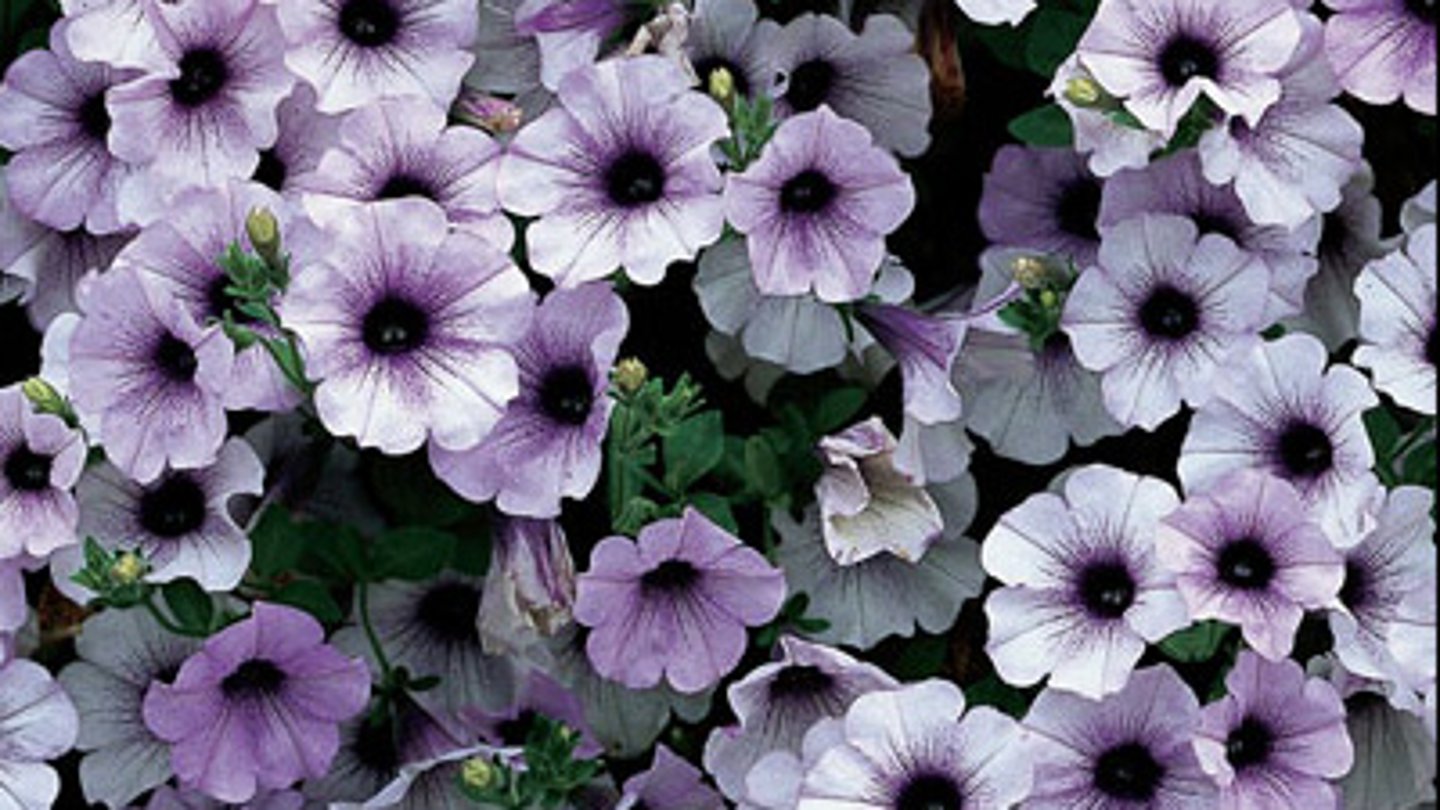

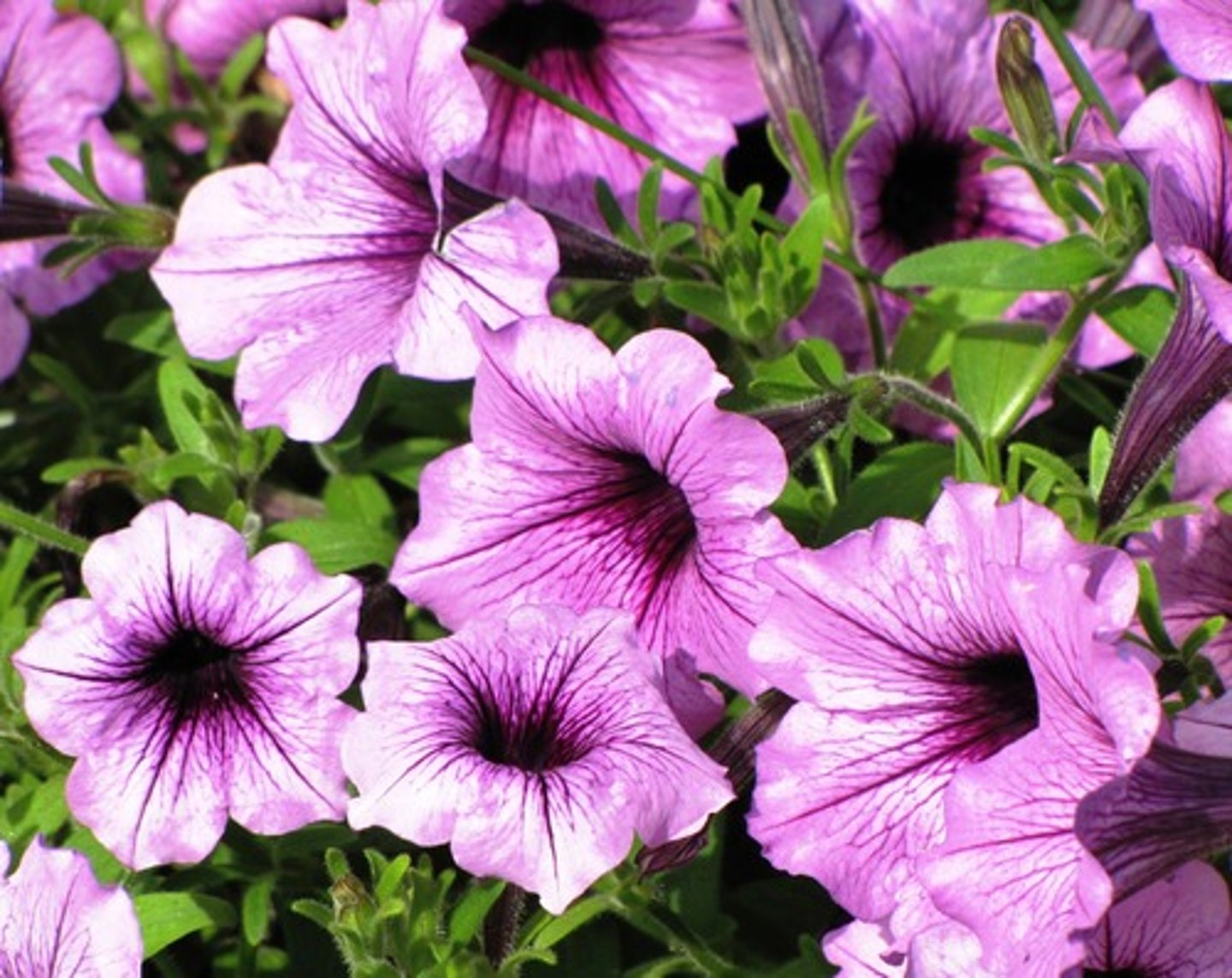
the antisense construct
instead of exhibiting less pigment, most petunias displayed areas of pigment

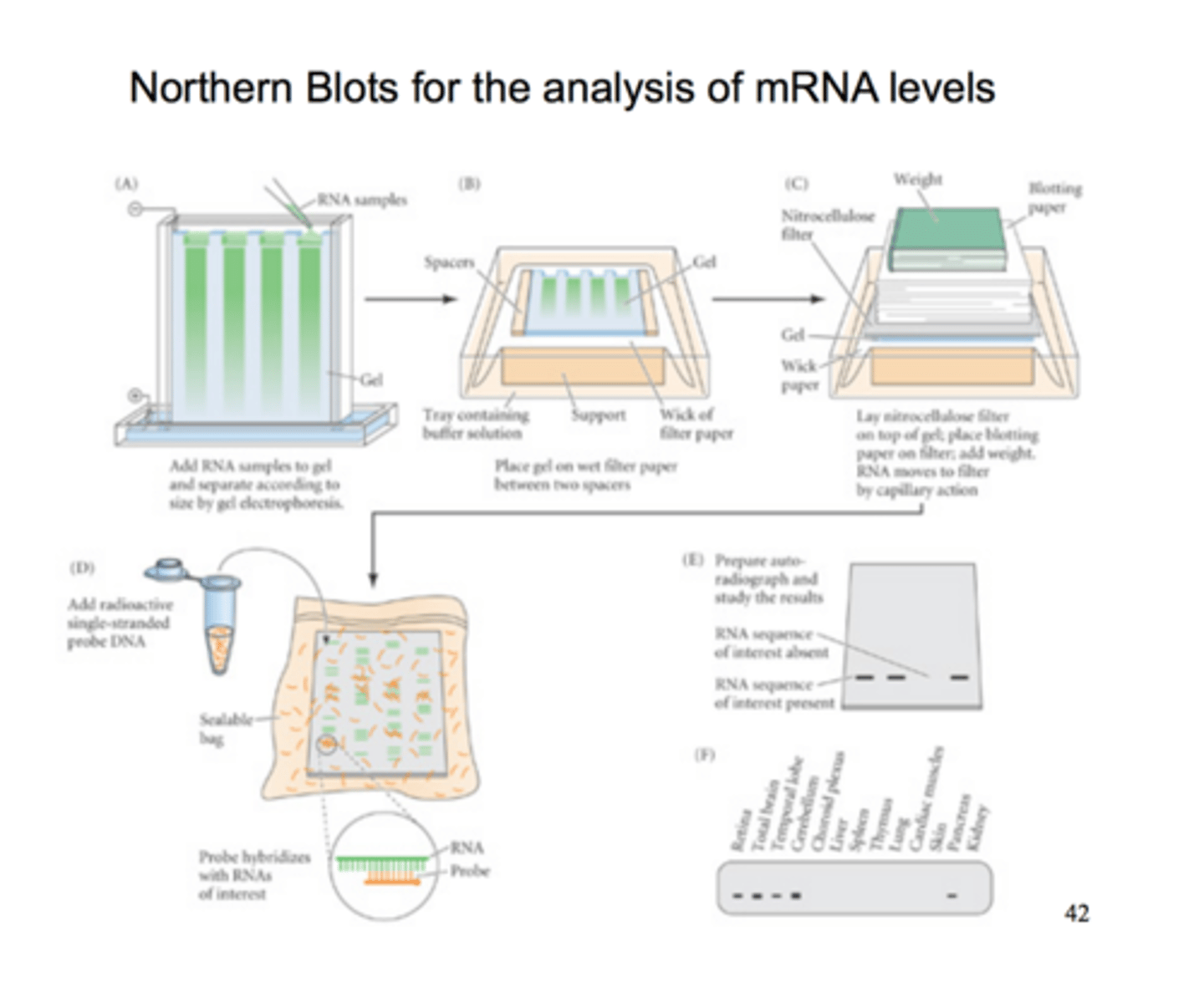
Northern blots
a technique used to assess gene expression by determining whether a gene is producing mRNA and measuring the quantity of mRNA being synthesized
EGF and TGF genes
exhibit low expression in normal cells but show high expression in cancer cells
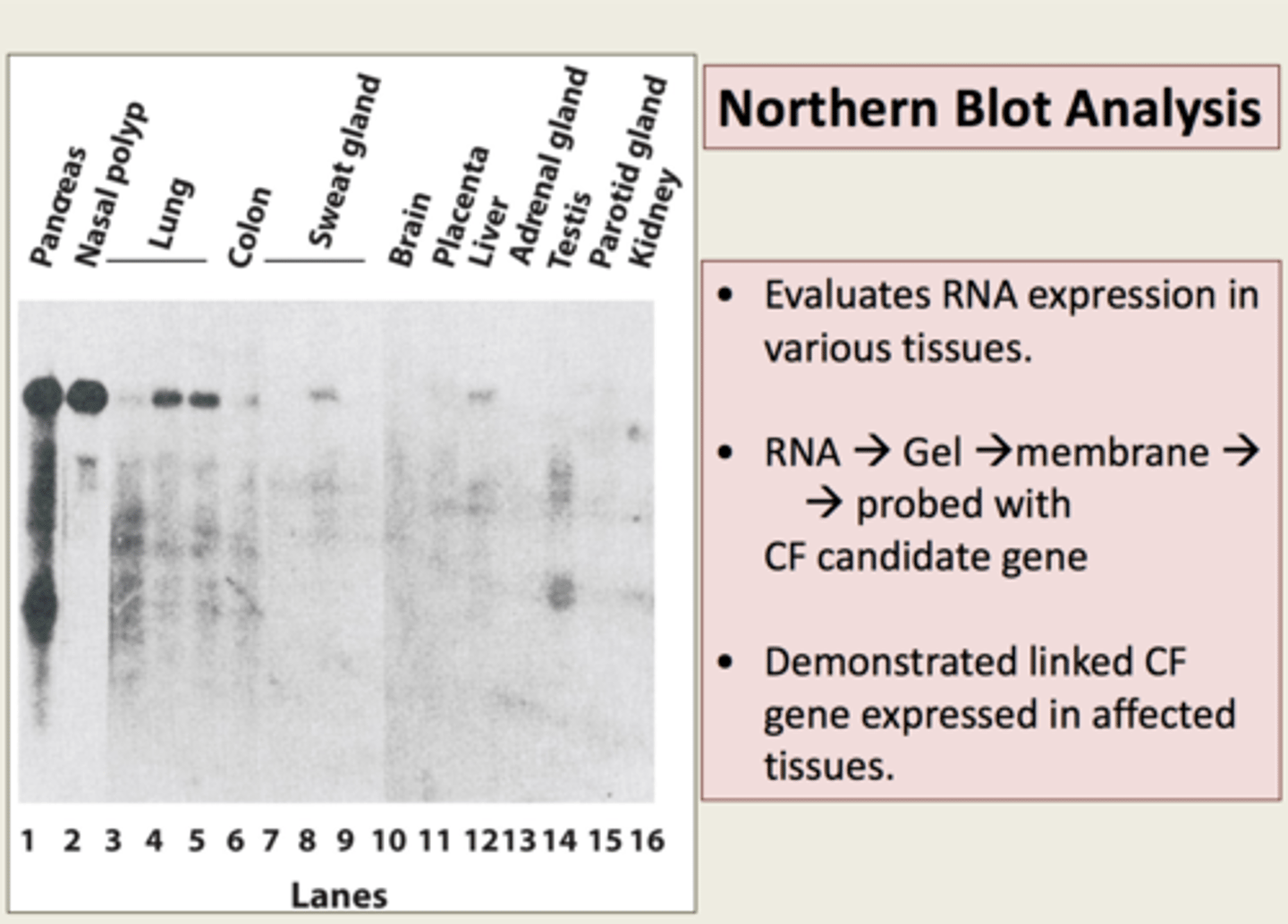
small interfering RNA (siRNA)
When there is an excessive amount of mRNA, the cells perceive it as an aberration. As a response, organisms with RdRp gene made the RdRp protein, which uses some of the 5' to 3' mRNA to generate antisense (3' to 5') mRNA
RdRp enzymes
made in response to an overabundance of a specific mRNA (in this e.g.it is CHS mRNA) because the cell view this as abnormal, so it wants to regulate this excessive levels of mRNA.
long double stranded RNA
The sense and antisense from RdRp bind
processed into small RNAs (siRNA) of about 21 to 24 bp
RNA dependent RNA Polymerase (RDRP)
uses the CHS sense mRNA to make antisense mRNA...
BECAUSE THE OVERABUNDANCE OF CHS mRNA IS VIEWED AS ABNORMAL and it wants to regulate the amount of CHS mRNA in the cell
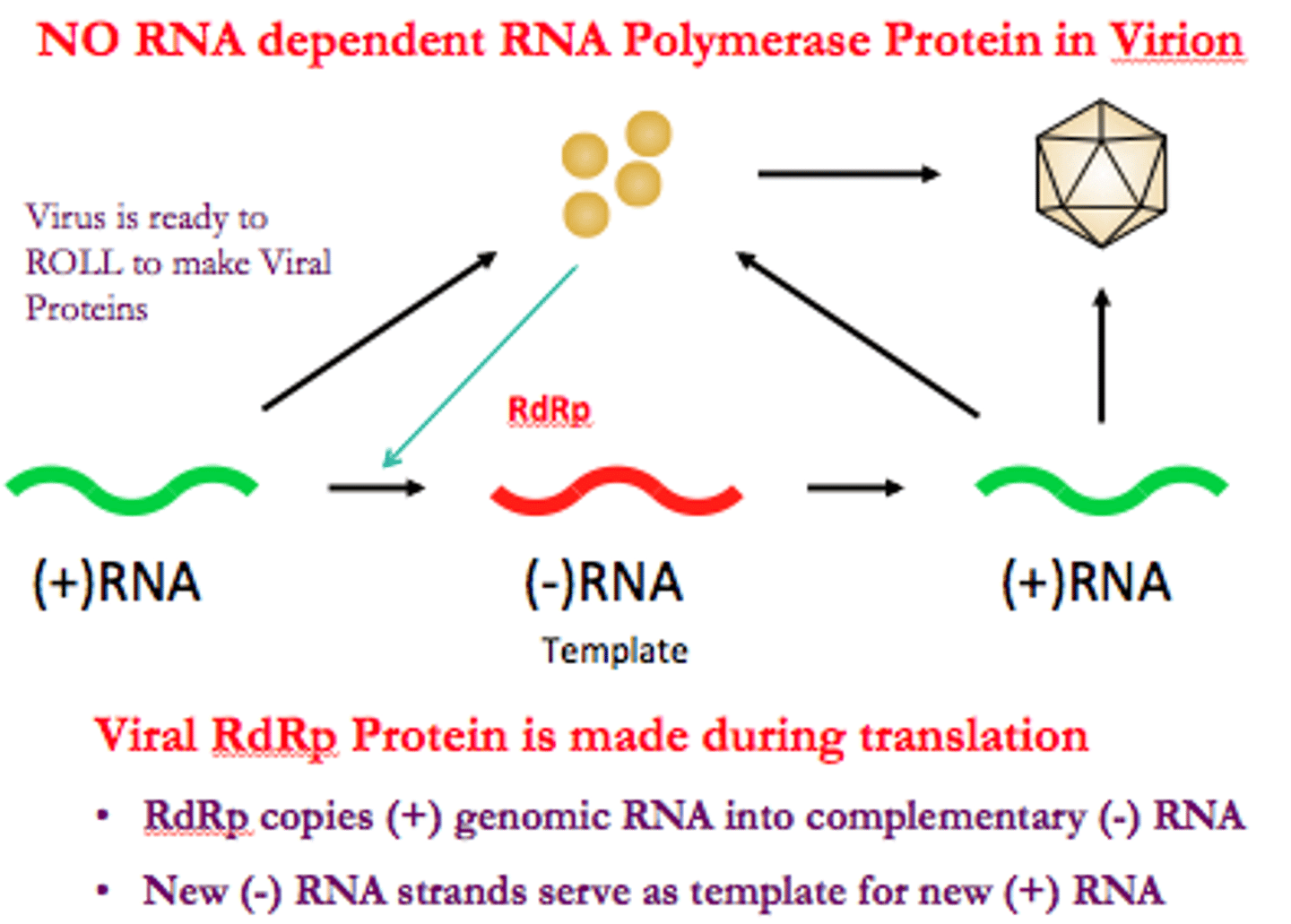
RdRP
using the 5' to 3' transcript as a template to make 3' to 5' antisense transcripts that are complementary to each other
- ubiquitous in plants
- In animals, it has been identified in only a few species
Why is silencing uneven?
1. Variable uptake of RNAi molecules
2. Differences in RNAi machinery
3. Amplification differences (in some organisms)
4. mRNA expression differences
Variable uptake of RNAi molecules
Not all cells receive the same amount of:
• dsRNA
• siRNA
Less RNA → weaker silencing
Differences in RNAi machinery
•Levels of silencing enzymes e.g.,
• Dicer
• Argonaute (RISC)
Cells with more machinery → stronger silencin
Amplification differences (in some organisms)
•Some cells produce more secondary siRNA
Stronger effect
mRNA expression differences
Cells expressing more of the target gene may show:
partial silencing instead of complete knockdown
What uneven silencing looks like experimentally:
• Patchy phenotype
• Partial reduction in gene expression
• Different levels of protein in different cells

Sources of long dsRNA
In plants, RdRP can make antisense RNA from sense RNA. The antisense mRNA will bind to sense RNA.
In humans, if antisense RNA is provided, it can bind to the sense mRNA
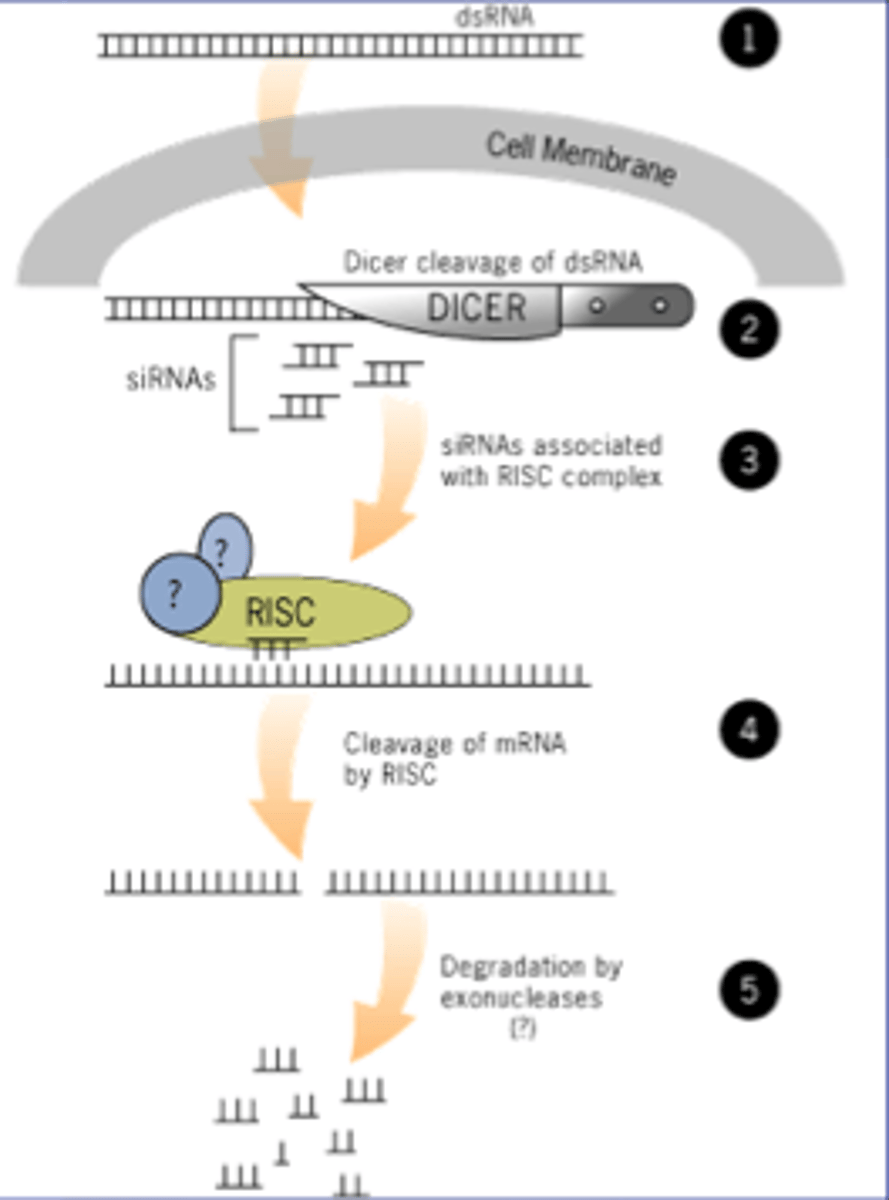
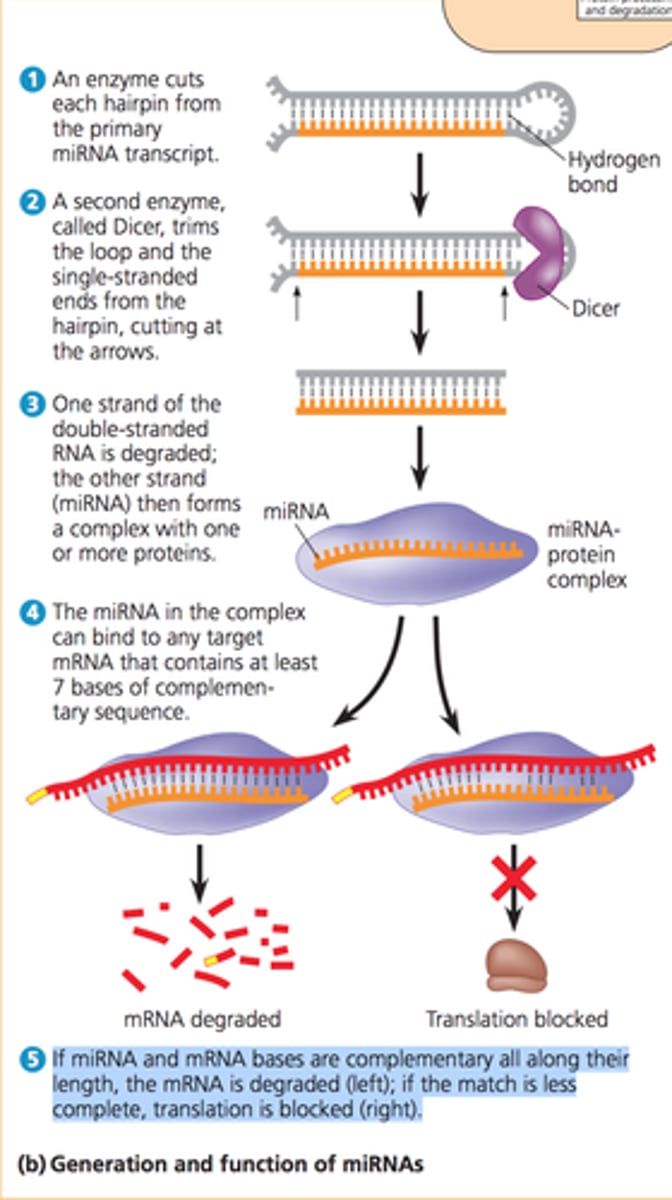
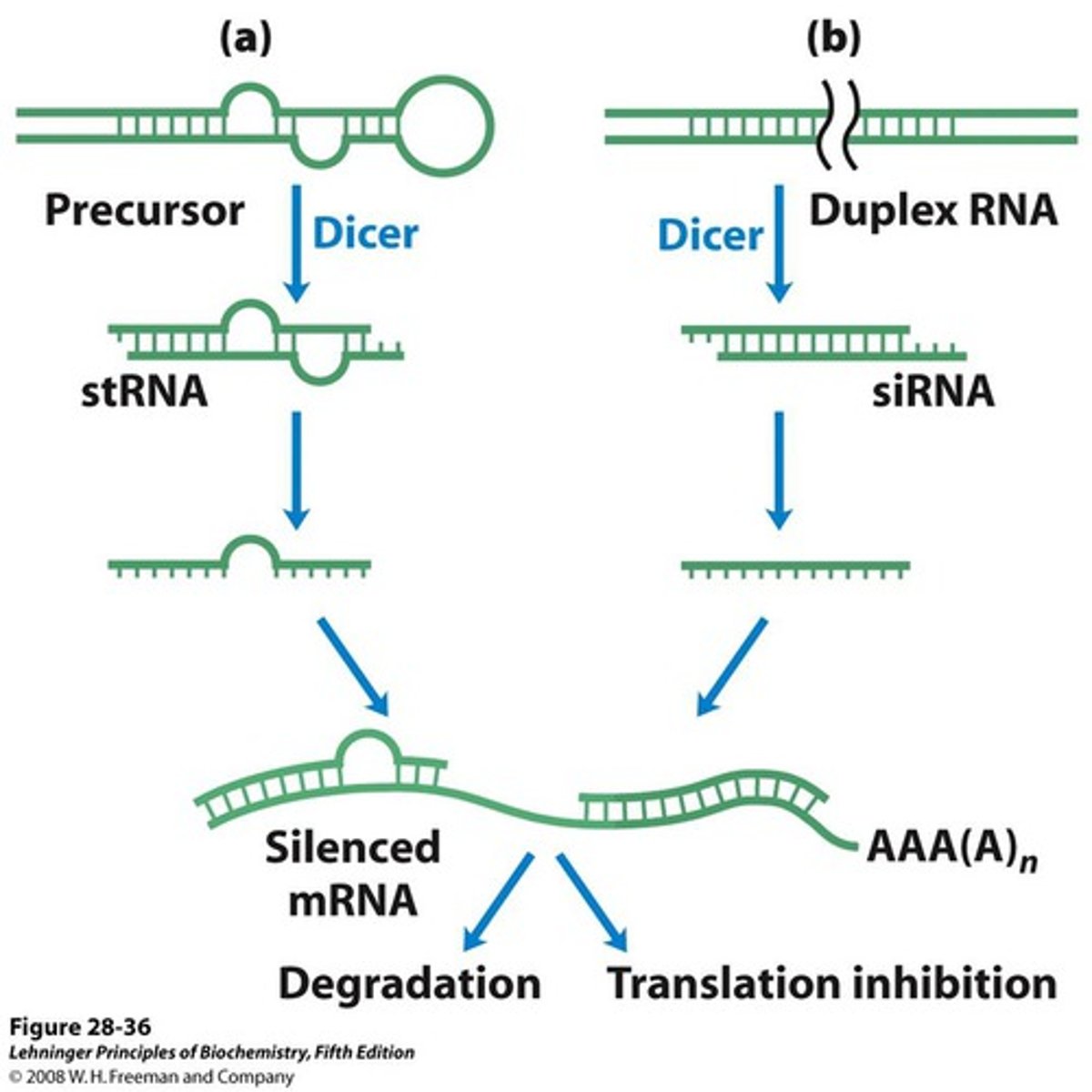
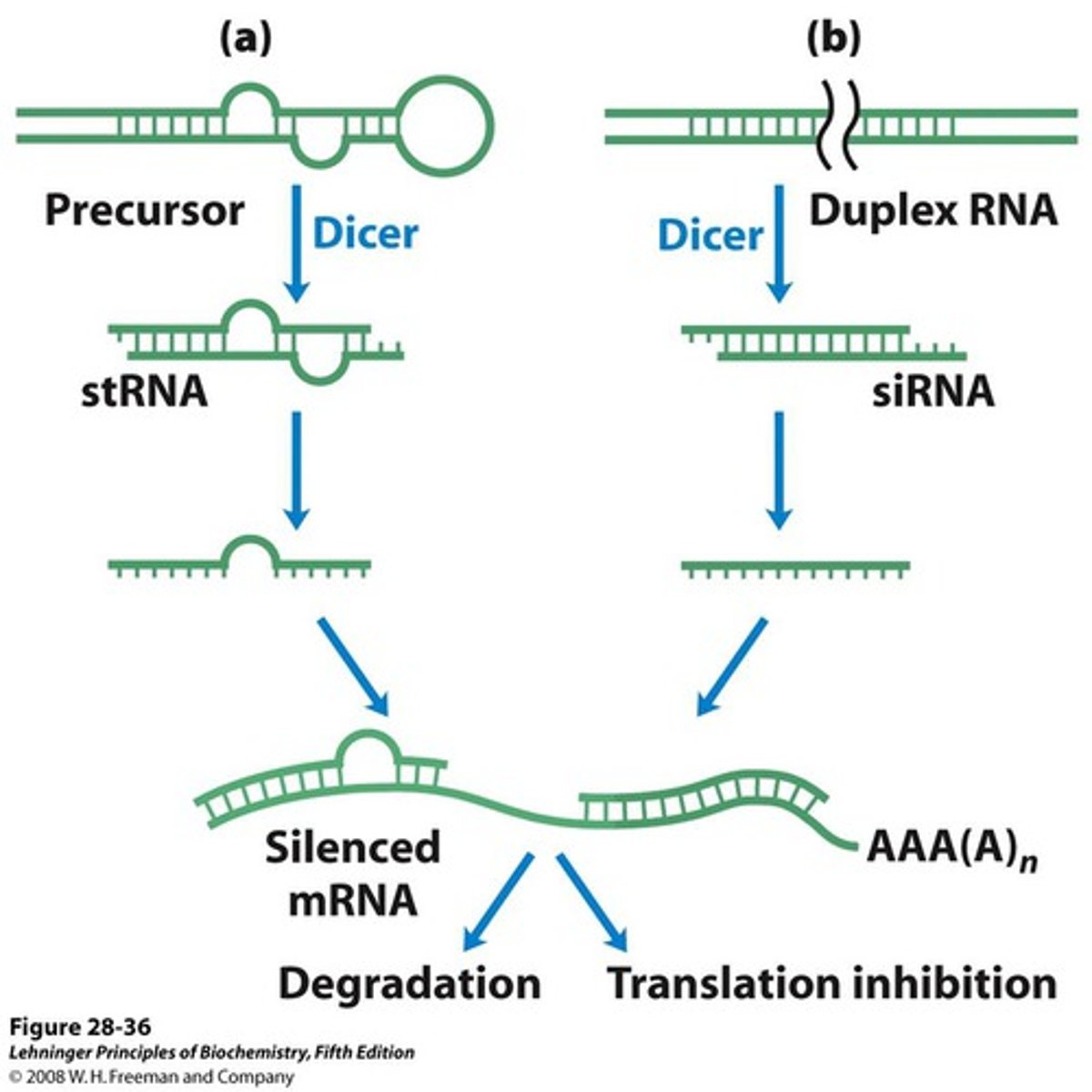
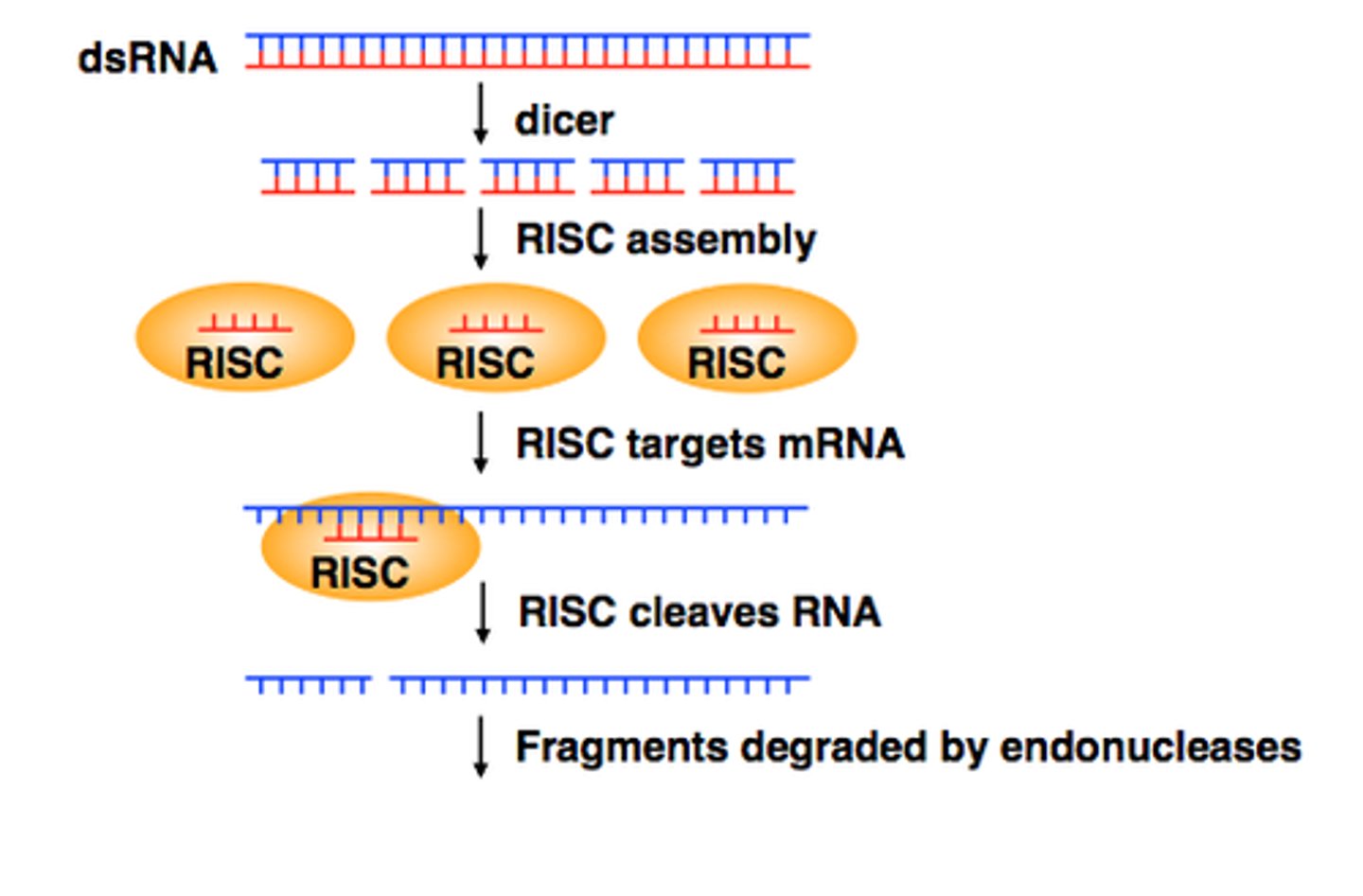
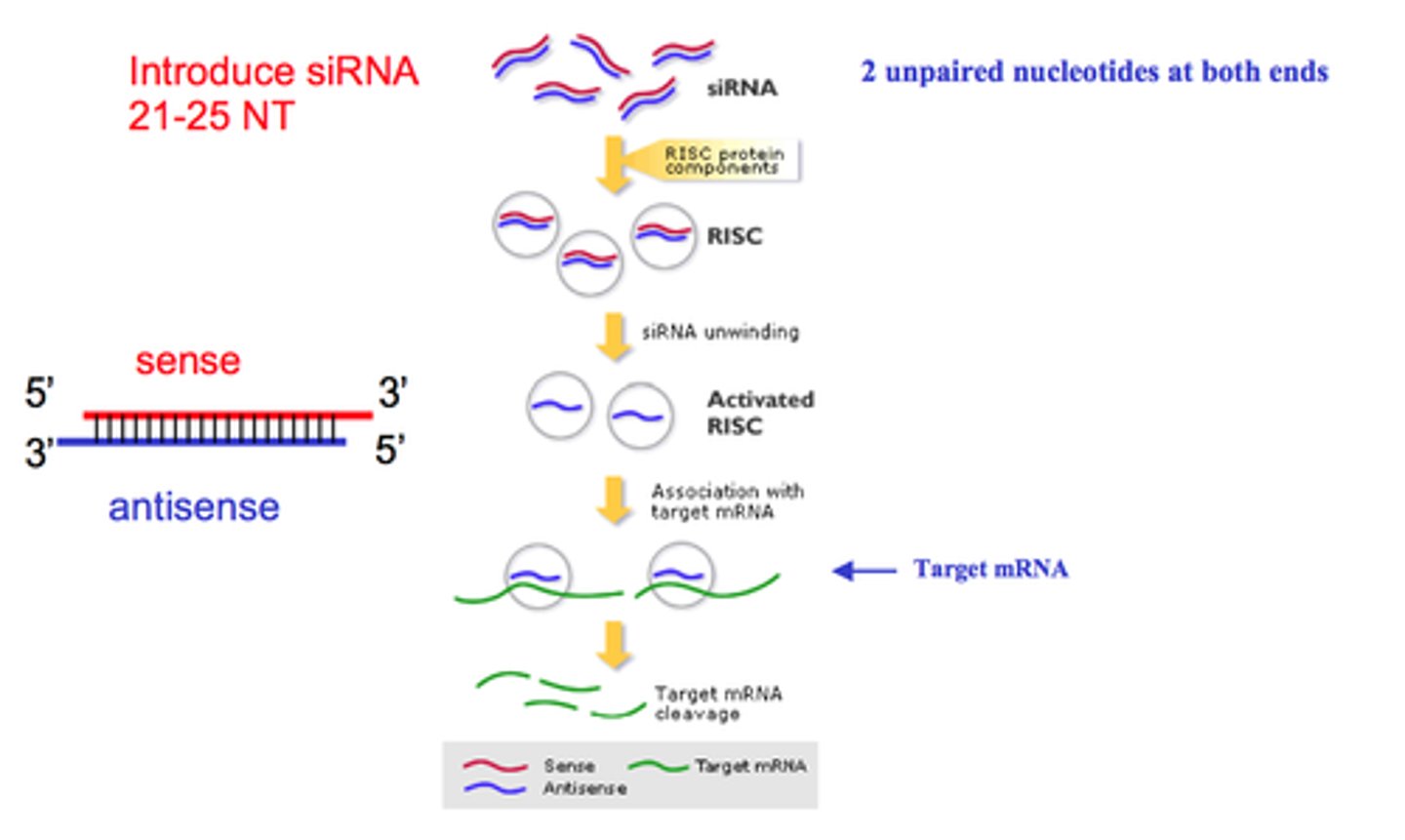
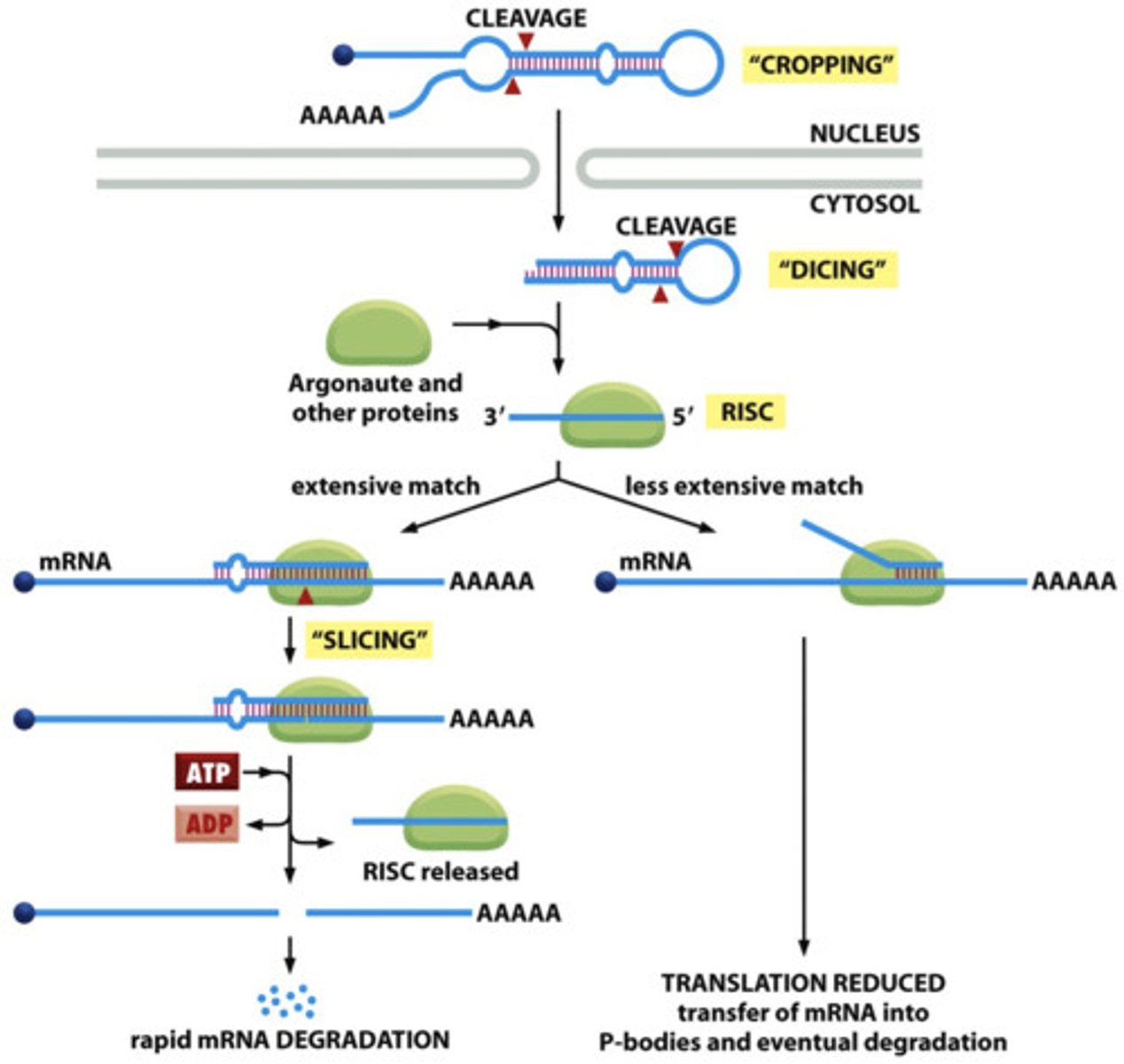
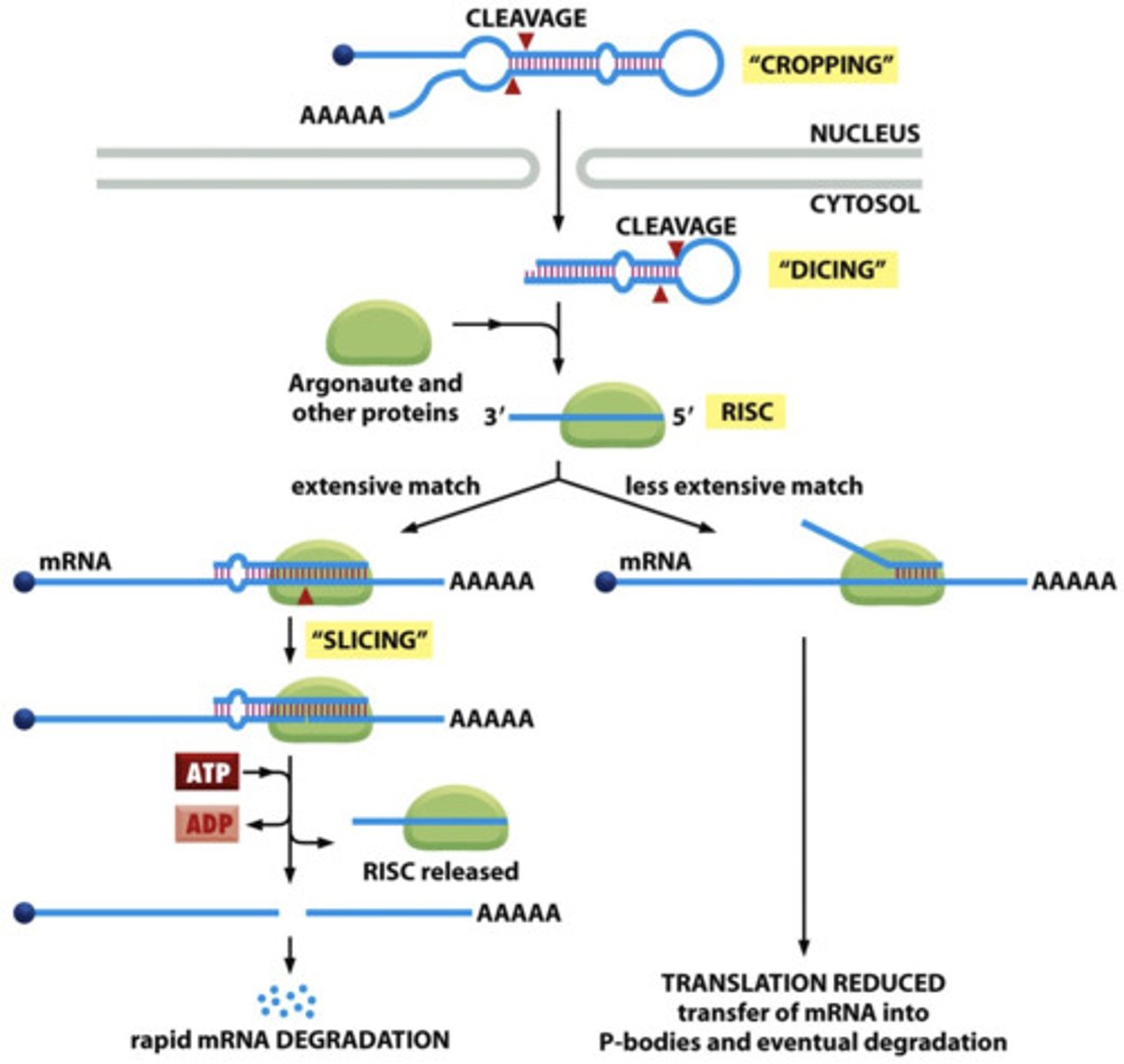
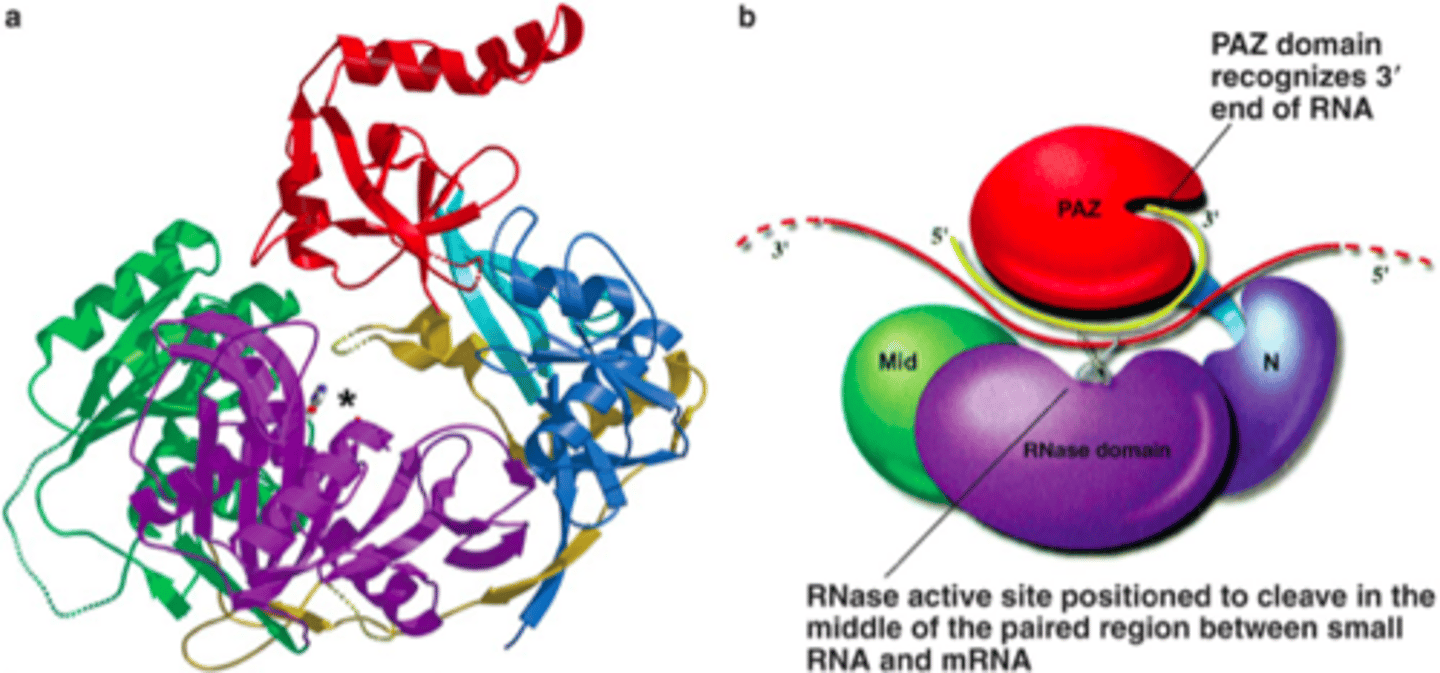
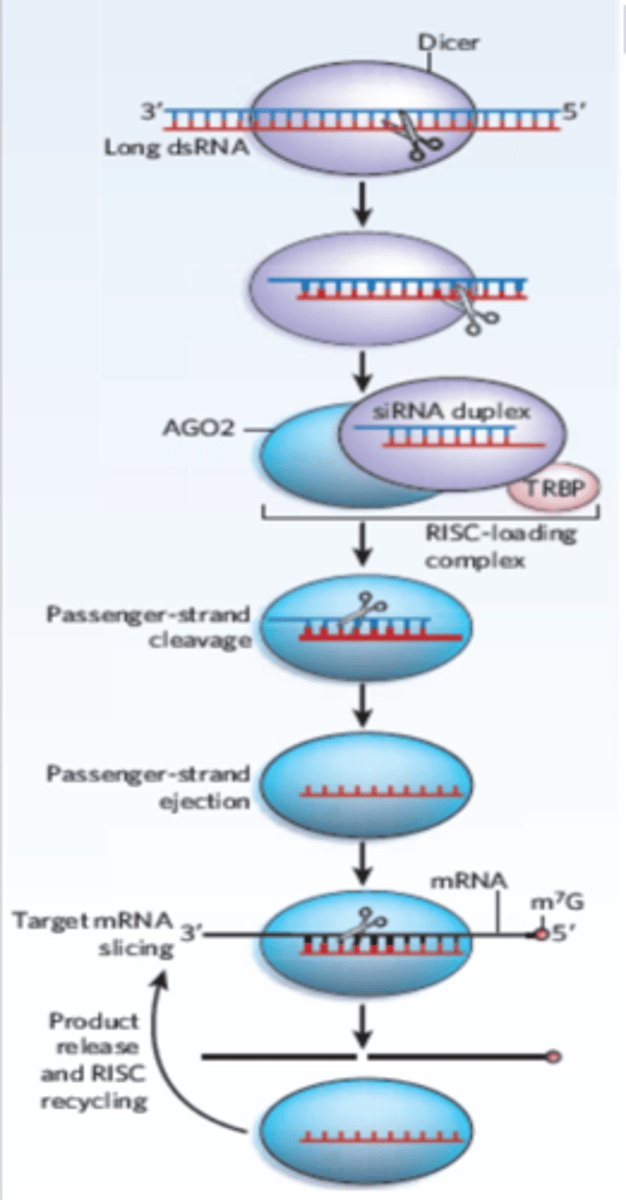
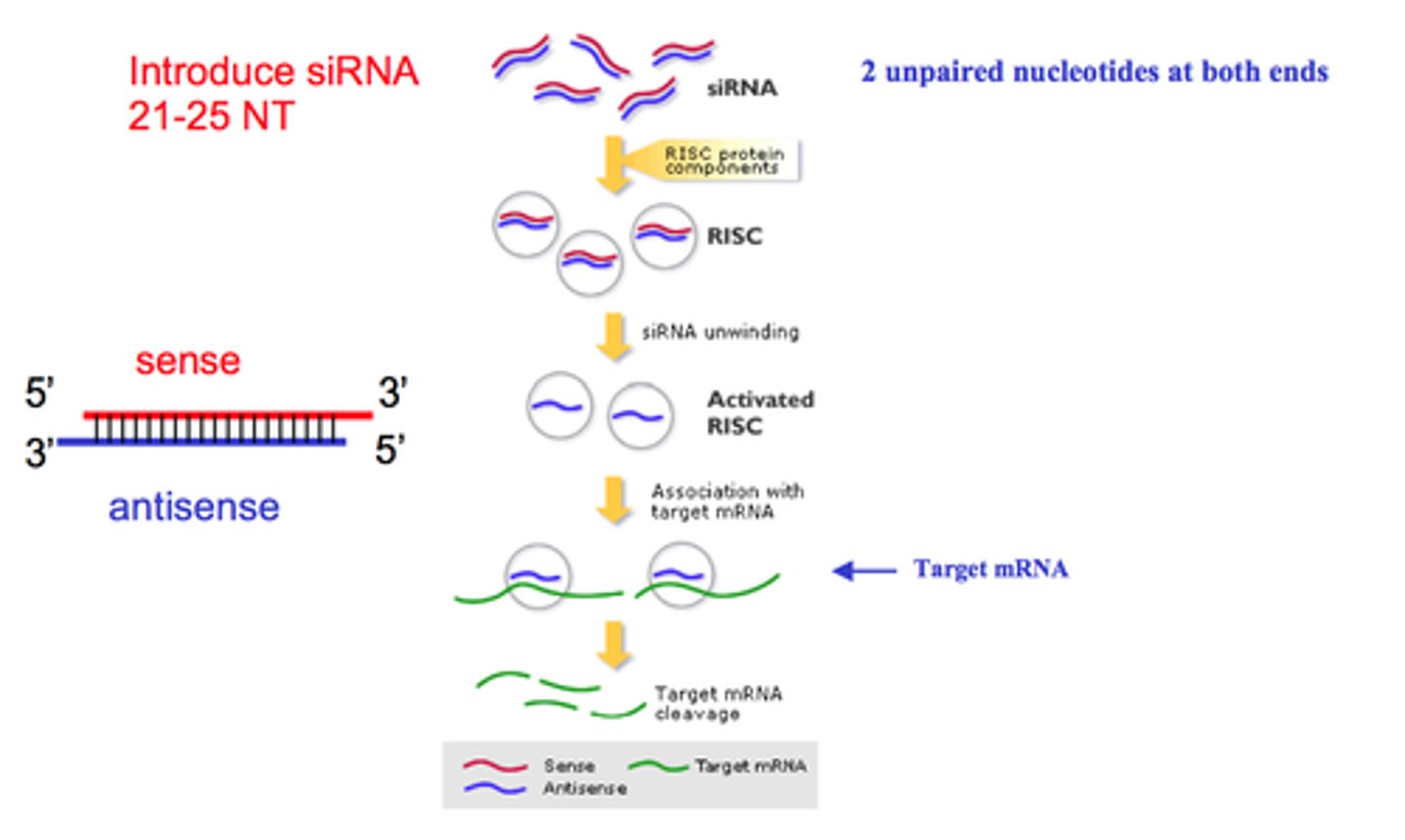
Step 1 of siRNA
The long dsRNA is cleaved by the enzyme Dicer (this is an endoribonuclease enzyme, so ribonuclease activity) into 21-24 bp small interfering RNAs (siRNAs)
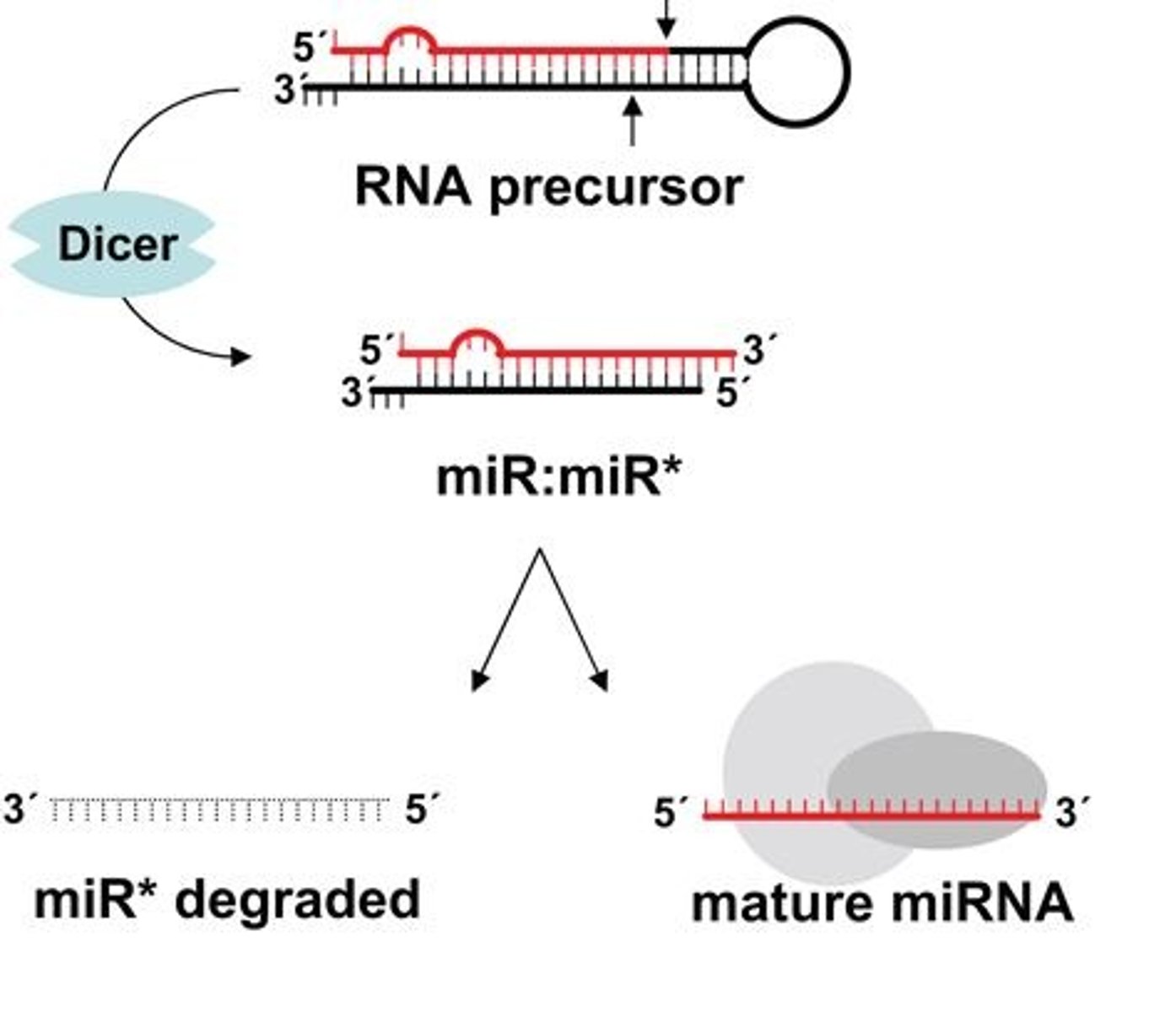
dicer
enzyme that cleaves and processes double stranded RNA to produce siRNAs or miRNAs that are 21-25 nucleotids in length
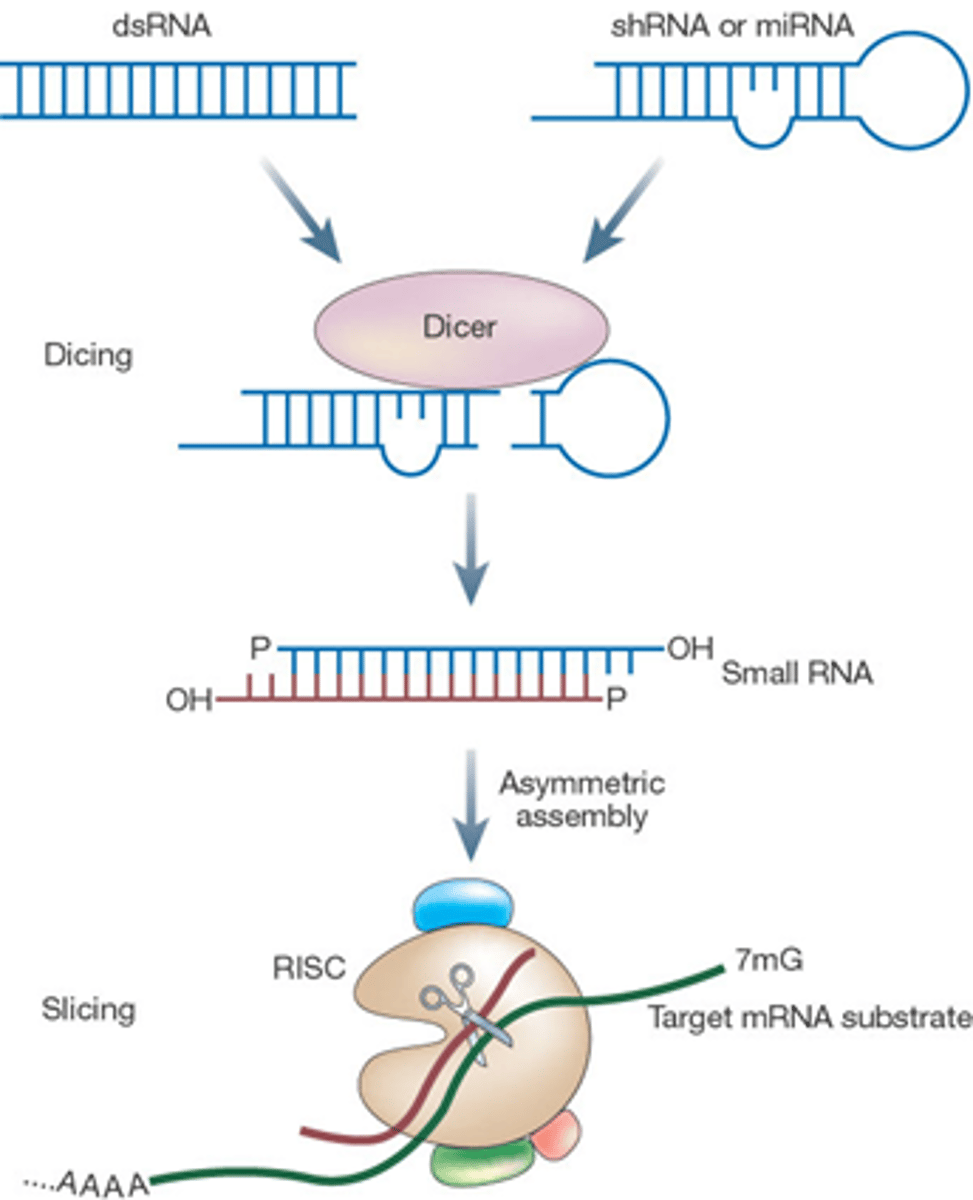
Step 2 of siRNA
The double-stranded siRNAs are loaded into RISC
In RISC is ARGONAUTE (Ago), which is an endoribonuclease.
In the RISC complex, the ds siRNA strands unwind to become single stranded and the 5' to 3' strand called the passenger strand is degraded. The 3' to 5' strand, called the GUIDE strand, binds to the AGO protein
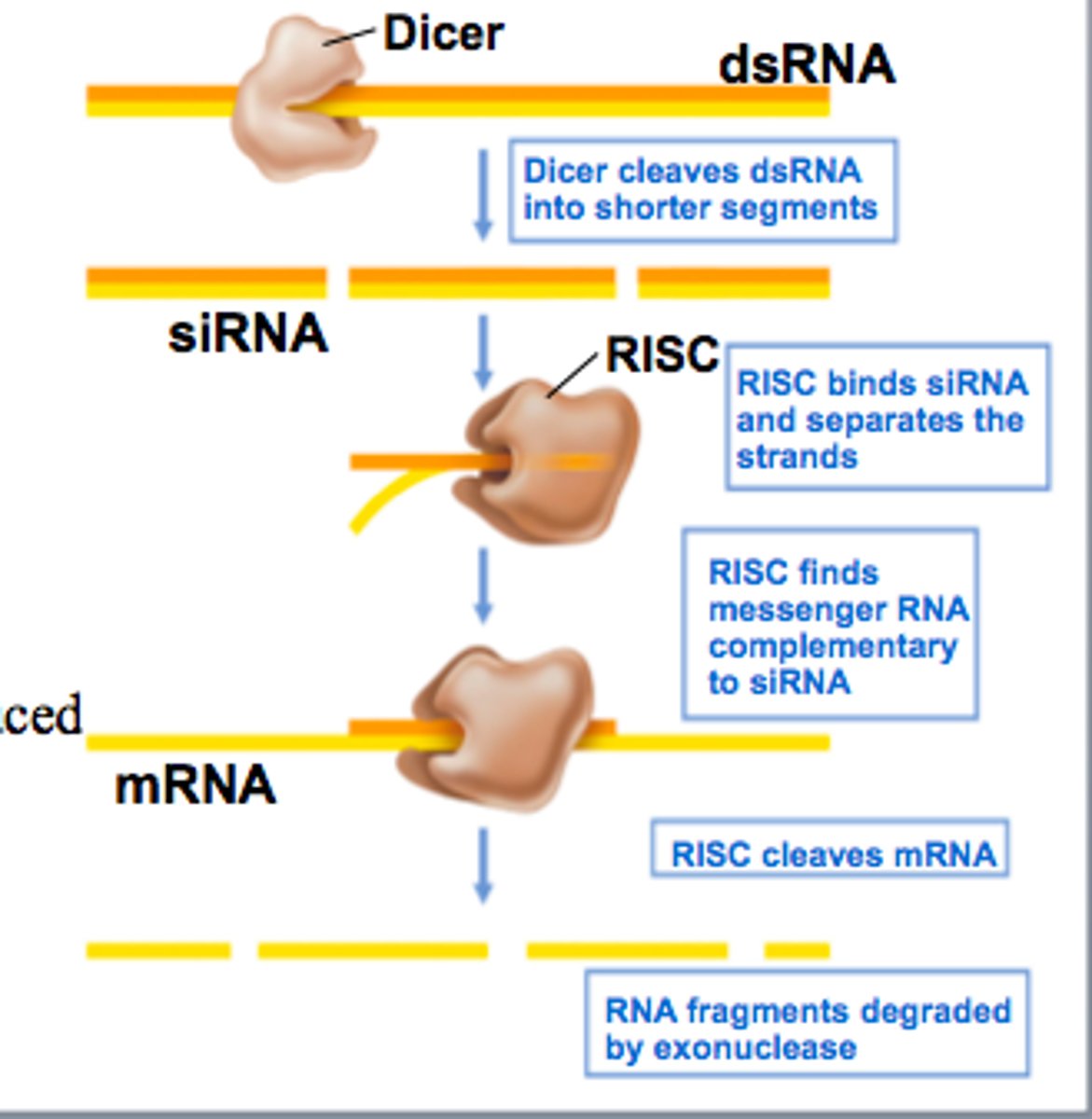
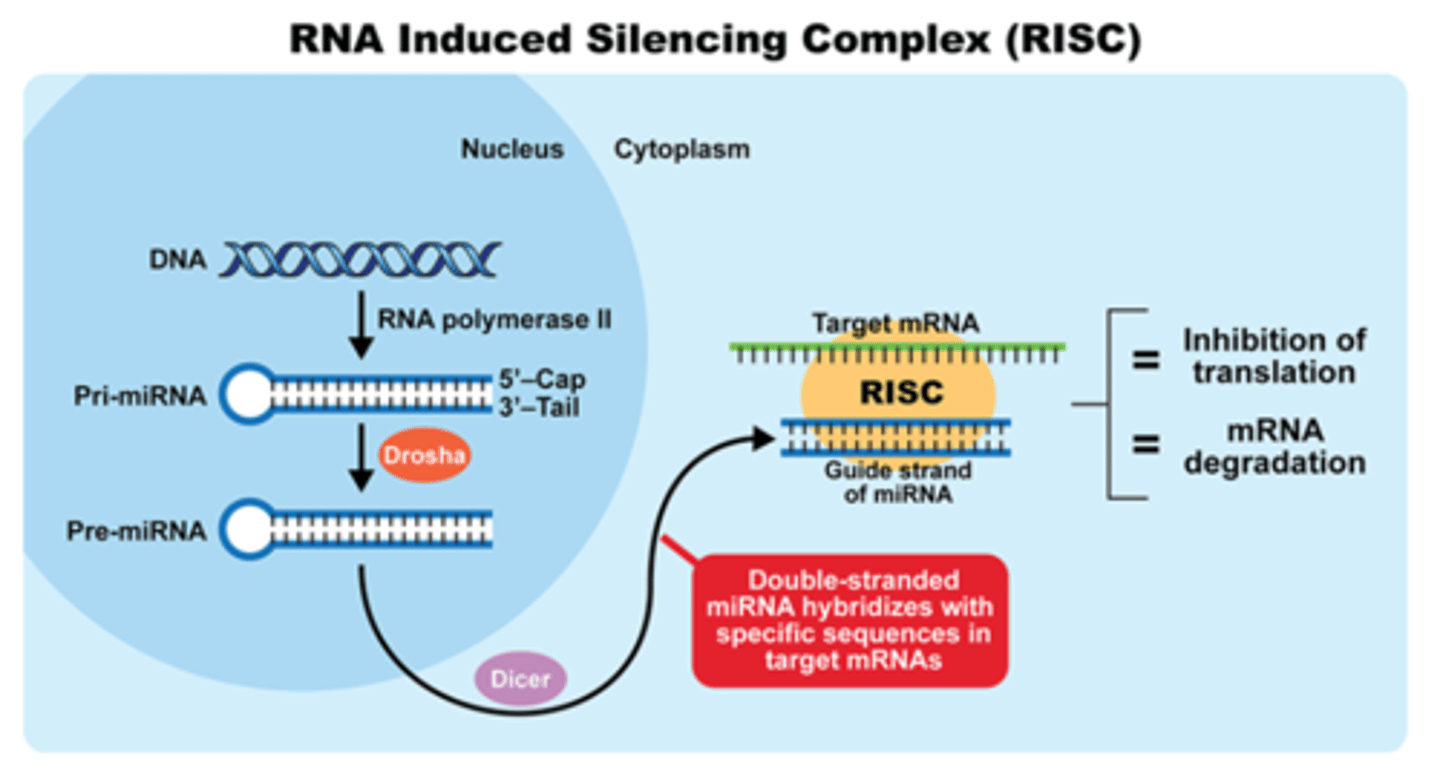
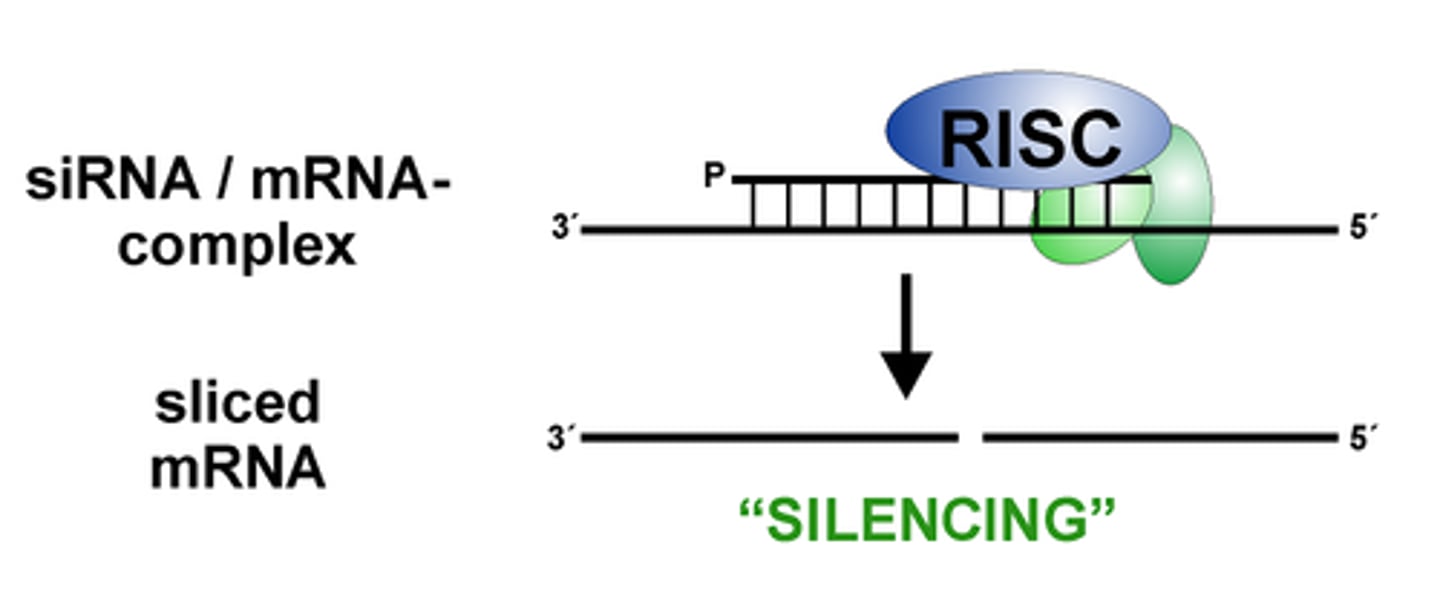
RNA-Induced Silencing Complex
uses siRNA as a guide to identify complementary mRNA, then acts as an endonuclease (via Argonaute protein) to cleave and degrade it
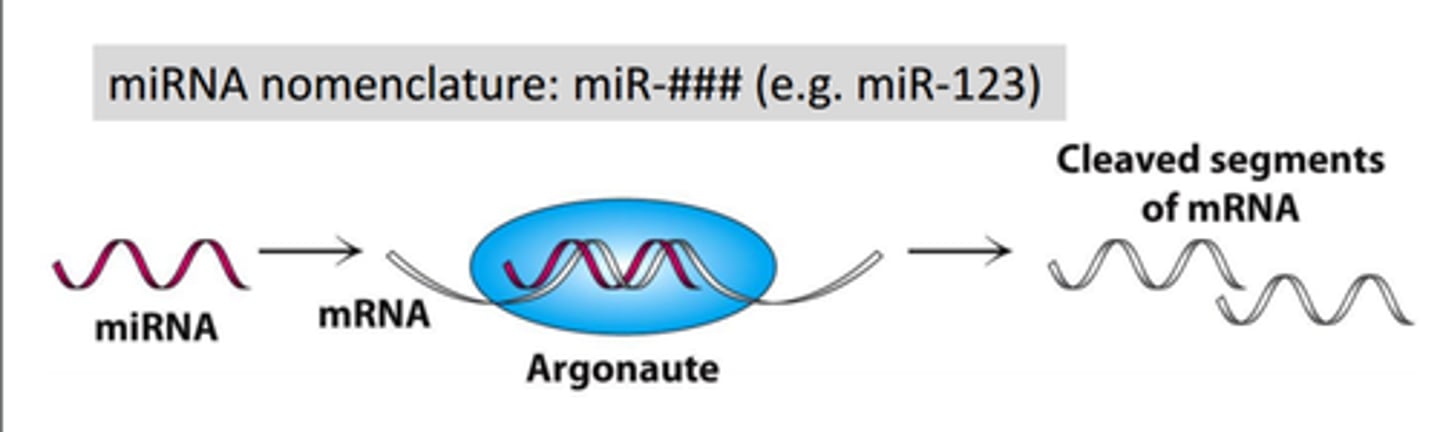
Argonaute
Catalytic subunit of the RISC complex of the RNAi machinery. Responsible for the cutting (or "slicing") activity of RISC that destroys target mRNAs.
Step 3 of siRNA
RISC complex with the guide strand bound to AGO is guided to an mRNA (i.e., the target) to which the guide strand then binds complementary to the mRNA (target recognition)
guide strand
21-24 nucleotide strand will bind to a specific region of the mRNA with perfect complementarity.
passenger strand
Strand that is degraded and decomposed after cut by dicer in RNA
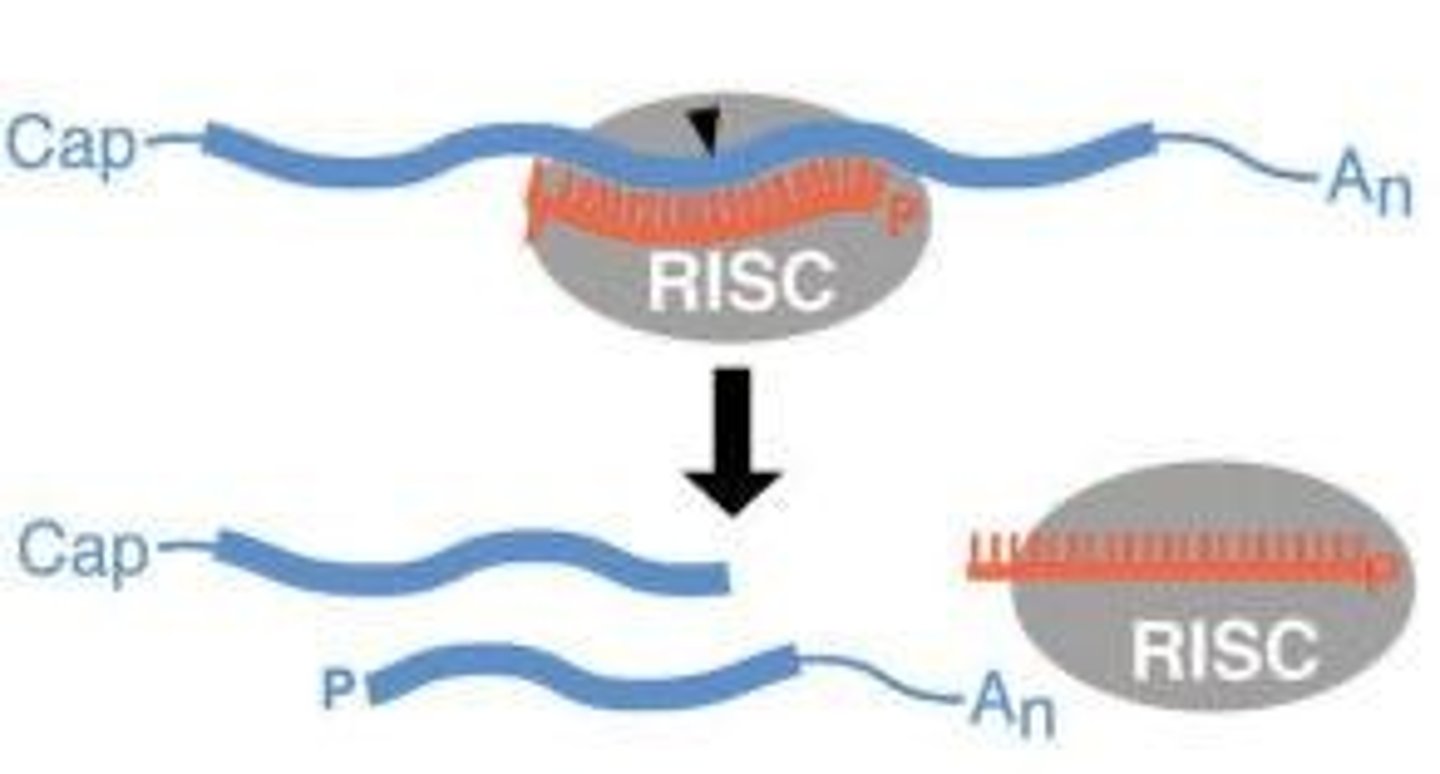
Step 4 of siRNA
The AGO, which is an endoribonuclease, cleaves the mRNA.....The cleaved mRNA is then degraded, it cannot be translated to a protein
Strand Selection within RISC
he strand with the less stable (lower GC content) 5′ end is typically selected as the guide strand, while the opposite strand is discarded
Step 1 siRNA amplification by RdRP
RISC binds the target mRNA (Step 3)
- The siRNA inside RISC pairs with the target mRNA.
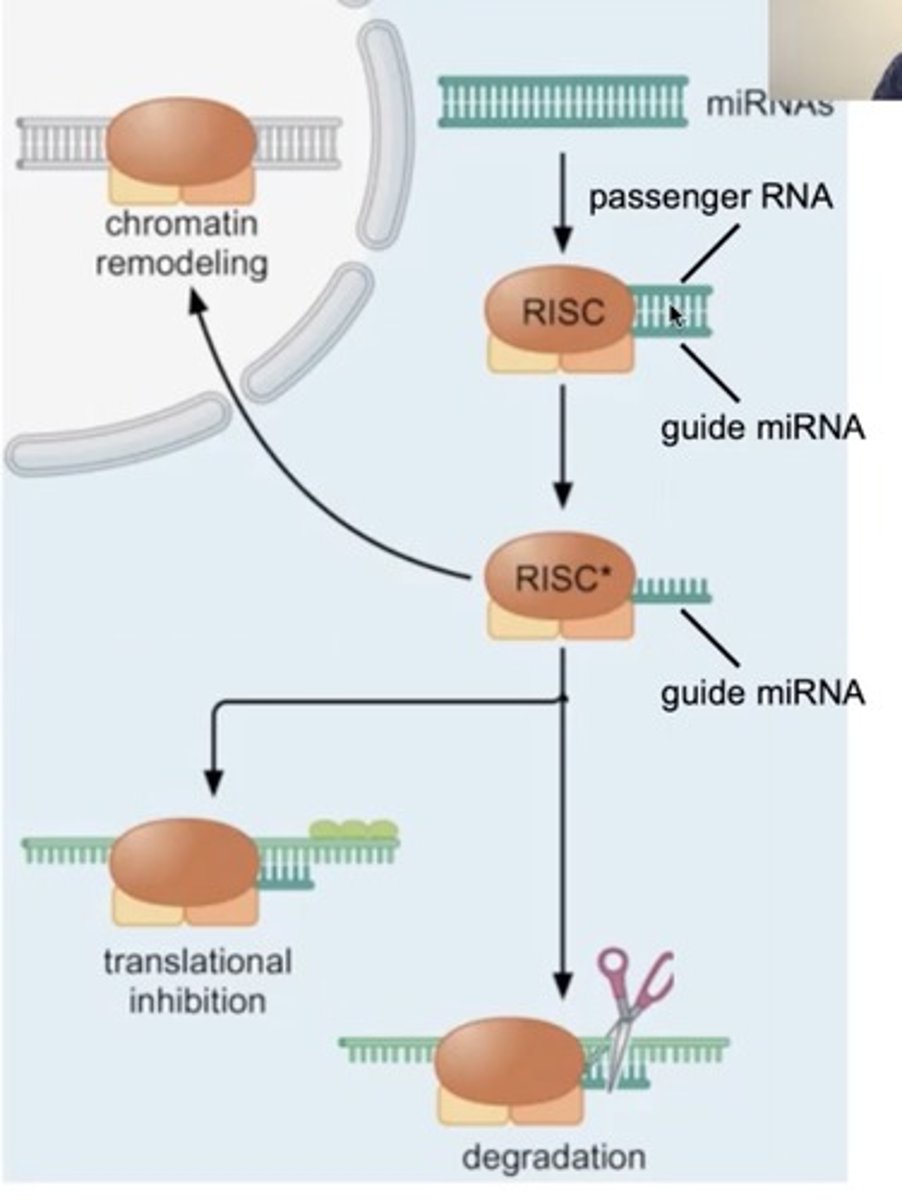
Step 2 siRNA amplification by RdRP
RdRP uses this mRNA as a template
RNA-dependent RNA polymerase (RdRP) copies the target mRNA.
This creates new double-stranded RNA (dsRNA)
Step 3 siRNA amplification by RdRP
Dicer cuts this new dsRNA again
The new dsRNA is processed into more siRNA
- SECONDARY siRNAS
Step 4 siRNA amplification by RdRP
More siRNAs = stronger silencing
These new siRNAs load into RISC
they go on to target and degrade more mRNA.
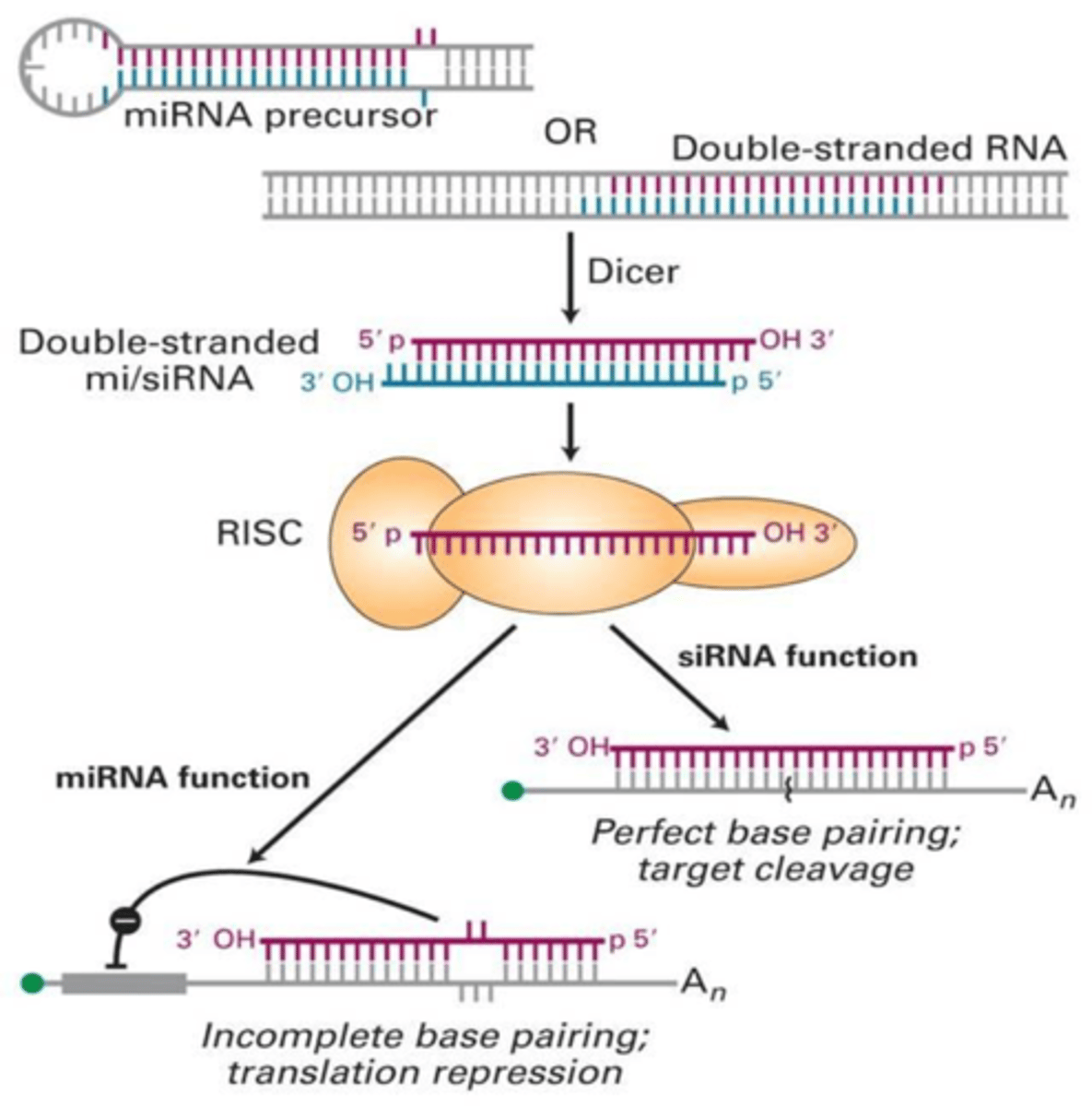
Perfect siRNA pairing
Argonaute cuts the mRNA at the site paired to positions10-11 of the guide siRNA
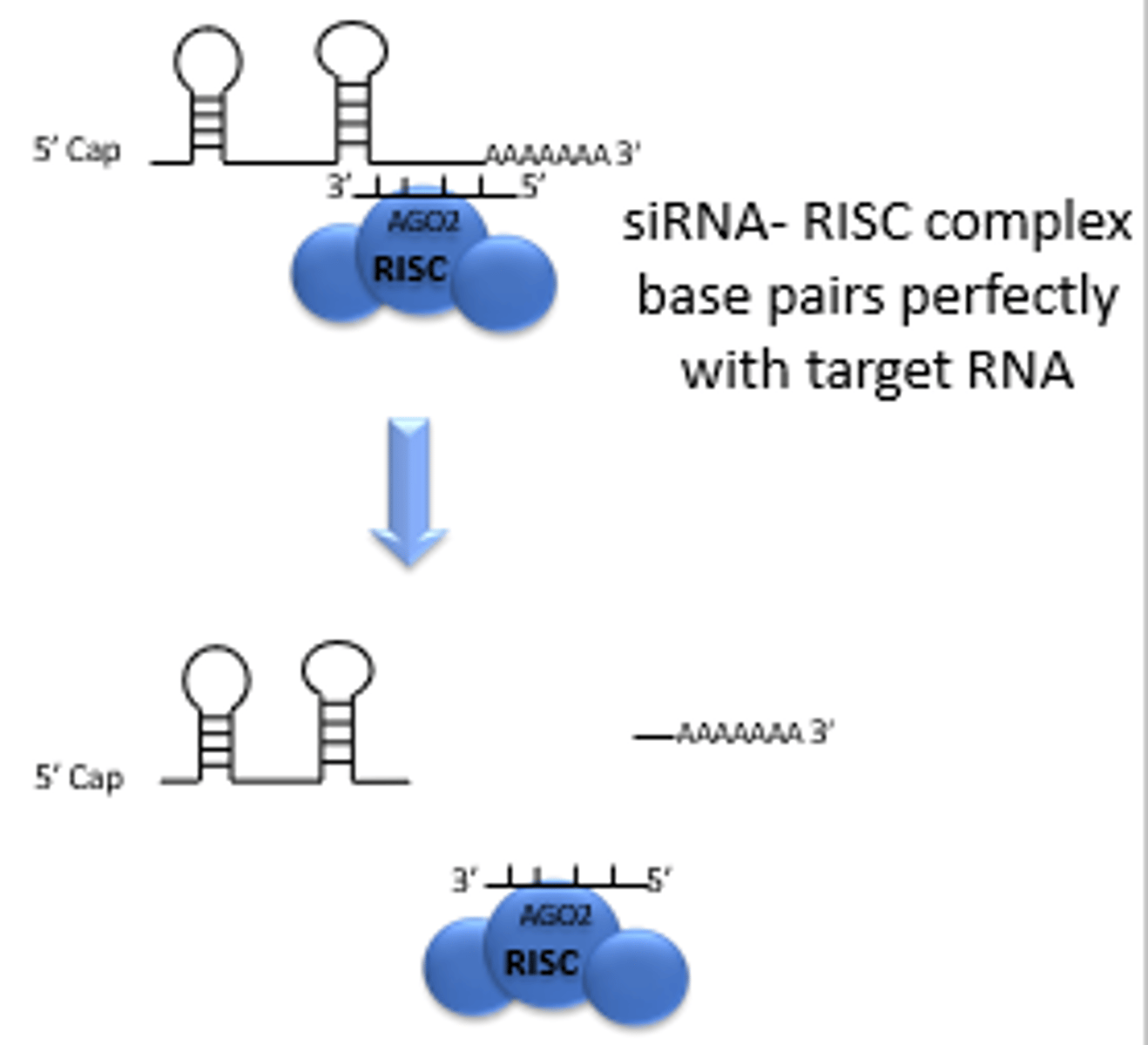
Cleavage occurs
5′ fragment (uncapped → rapidly degraded)
3′ fragment (no protection → degraded)
Antisense CHS mRNA from the transgene
Antisense CHS mRNA can be converted into double-stranded RNA (dsRNA) by RNA-dependent RNA polymerase (RdRp).
Antisense CHS mRNA can also directly base-pair with endogenous sense CHS mRNA to form dsRNA
Sense CHS mRNA from the transgene
An unusually high level of sense CHS mRNA is recognized by the cell as abnormal
RNA-dependent RNA polymerase (RdRp)uses some of this CHS mRNA as a template to make a complementary antisense strand
As a result, both sense and antisense CHSRNA are present, which pair together to form double-stranded RNA (dsRNA)
recycling of siRNA:
siRNA guide strand remains in the RISC complex and can target multiple mRNA molecules, making the process highly efficient.
siRNA: short-interfering RNA, 21-24 nt
Mostly exogenous origin (i.e., come from external sources). Other than when RdRp makes antisense RNA which then go on to form dsRNAs
- dsRNA precursors
- Target specific
- TRANS-ACTING
siRNA based therapy
Patisiran, givosiran, lumasiran, and inclisiran
Givosiran
drug used to treat acute hepatic porphyria (AHP), a rare disorder caused by overproduction of a protein (enzyme), Aminolevulinic Acid Synthase 1 (ALAS1).
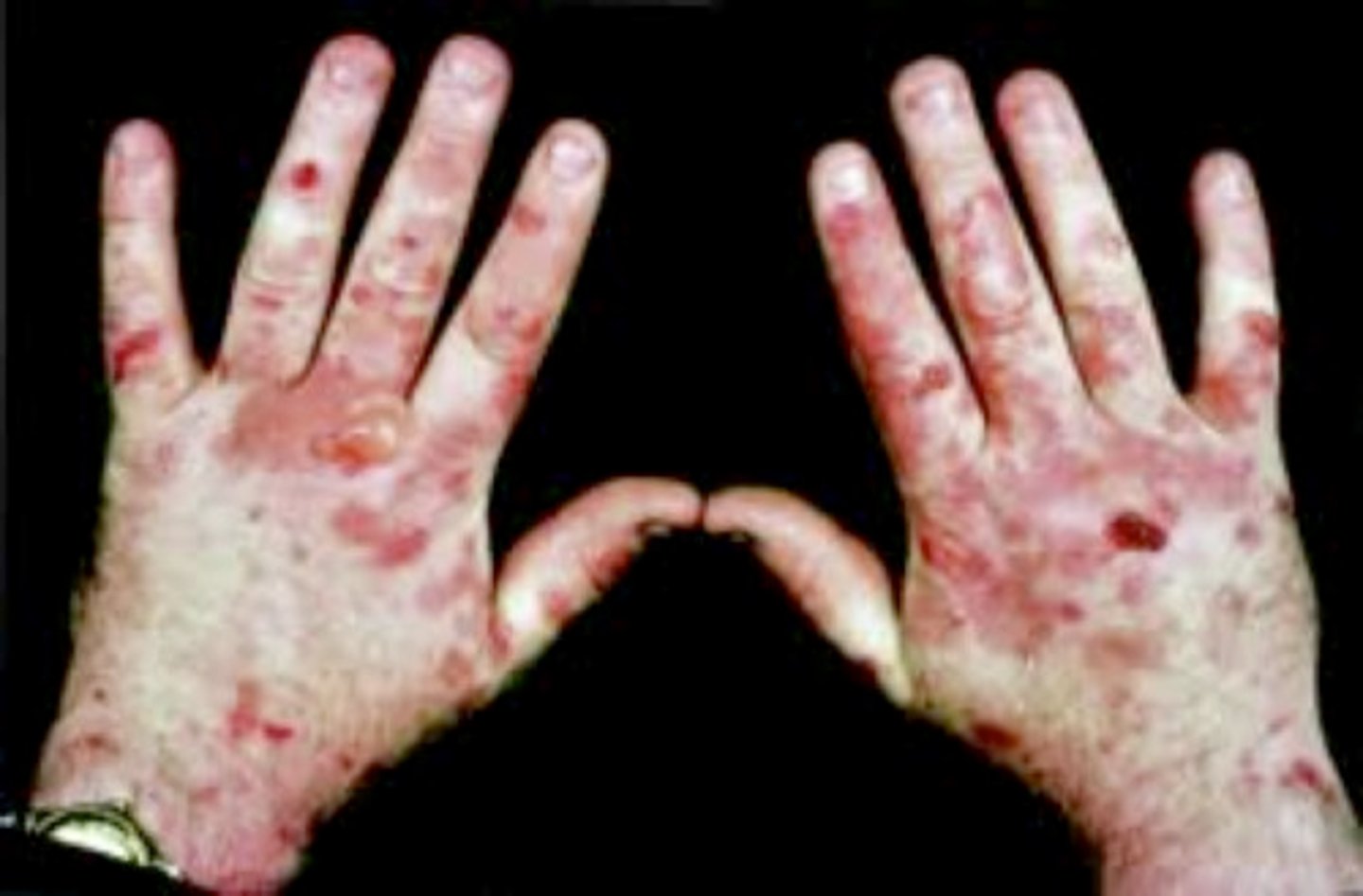
acute hepatic porphyria
Excess of the ALAS1 enzyme is produced --> accumulation of heme intermediates
severe neurological and gastrointestinal symptoms
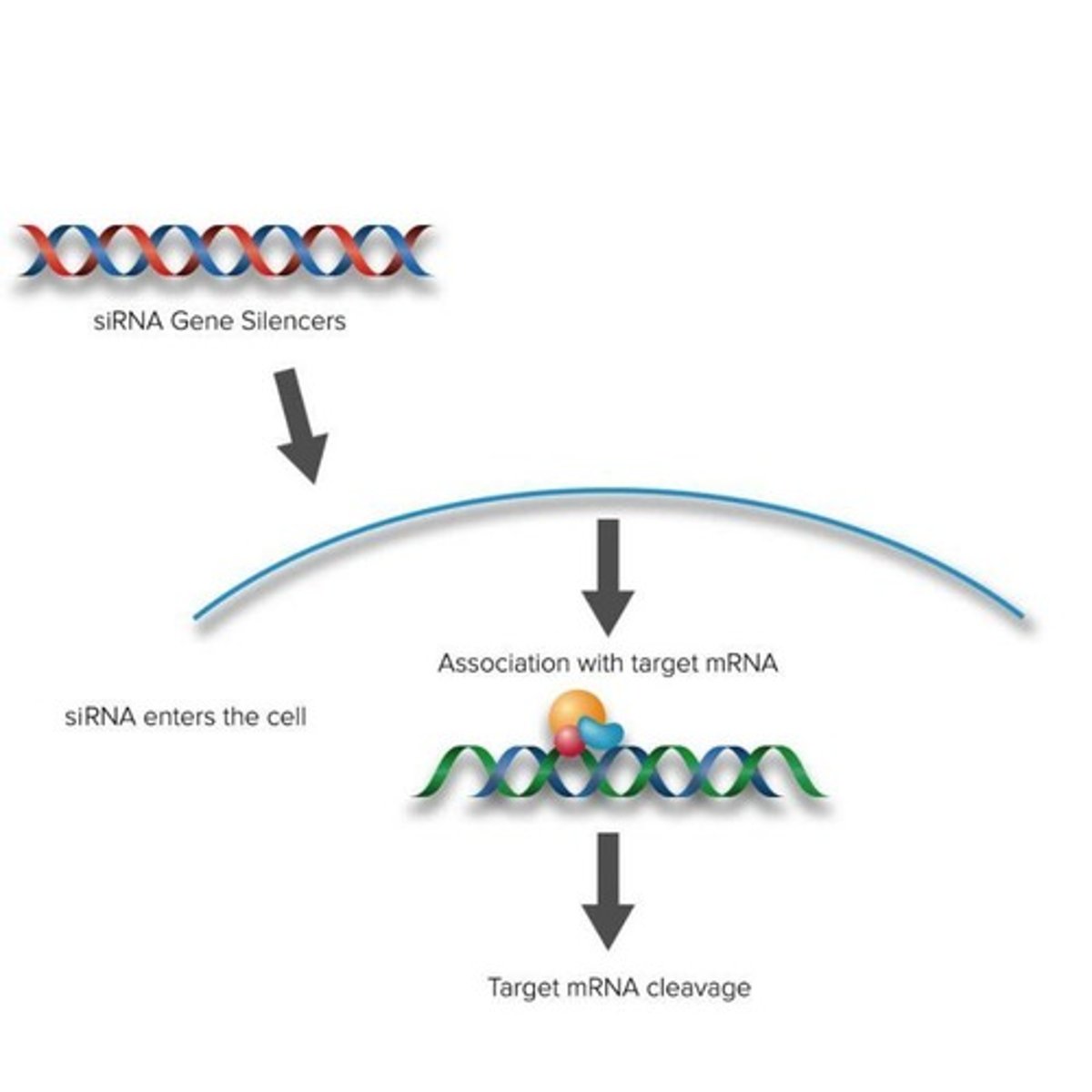
Givosiran siRNA
siRNA designed to target / bind with perfect complementarity to ALAS1 mRNA
The siRNA binds to the ALAS1 mRNA and the ALAS1 mRNA is destroyed
As a result, less ALAS1 enzyme is produced
Lumasiran
a drug used to treat primary hyperoxaluria type 1(PH1), a rare genetic disorder caused by mutations in the AGXT gene
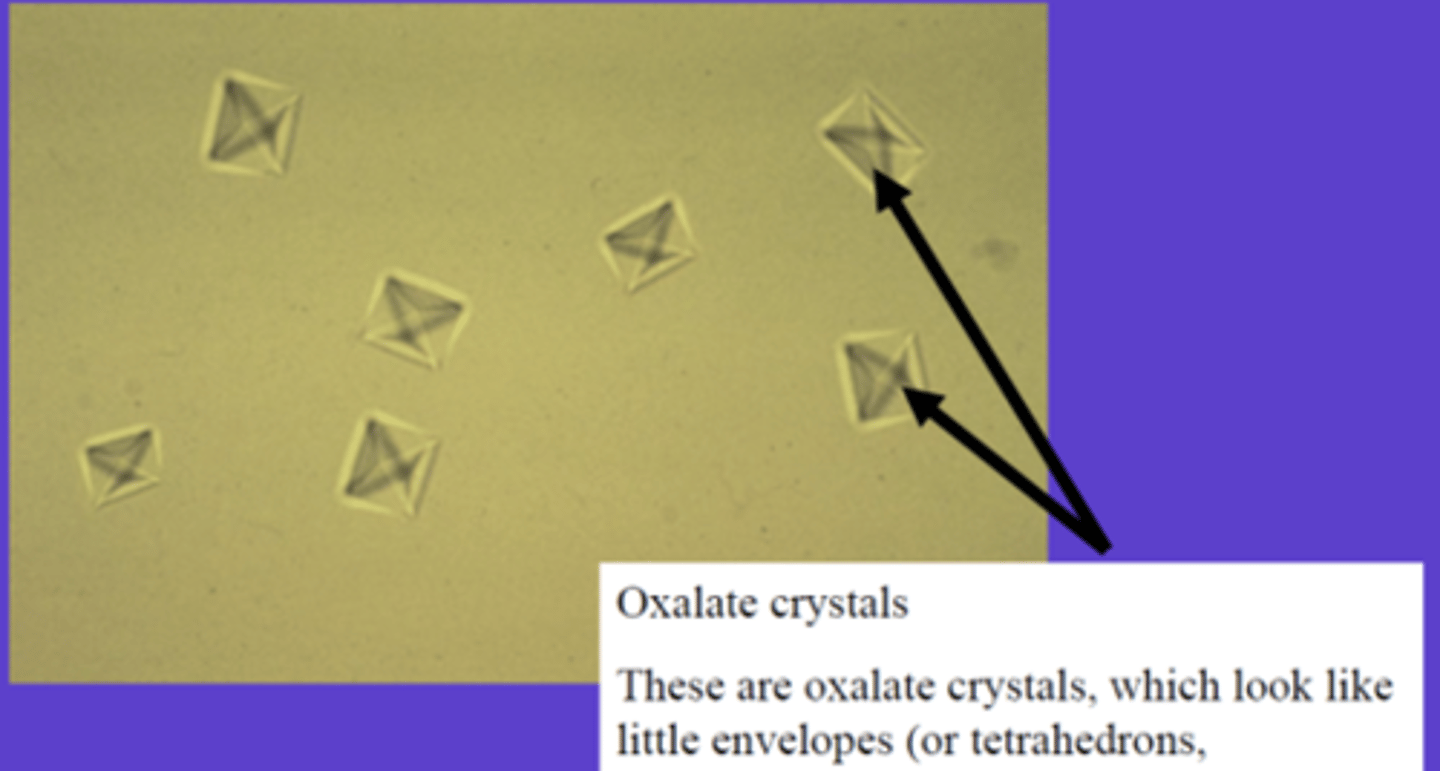
primary hyperoxaluria type 1 (PH1)
The AGT enzyme activity is absent
This leads to glyoxylate accumulation
Glyoxylate is converted into oxalate
Excess oxalate forms calcium oxalate crystals, which damage the kidneys
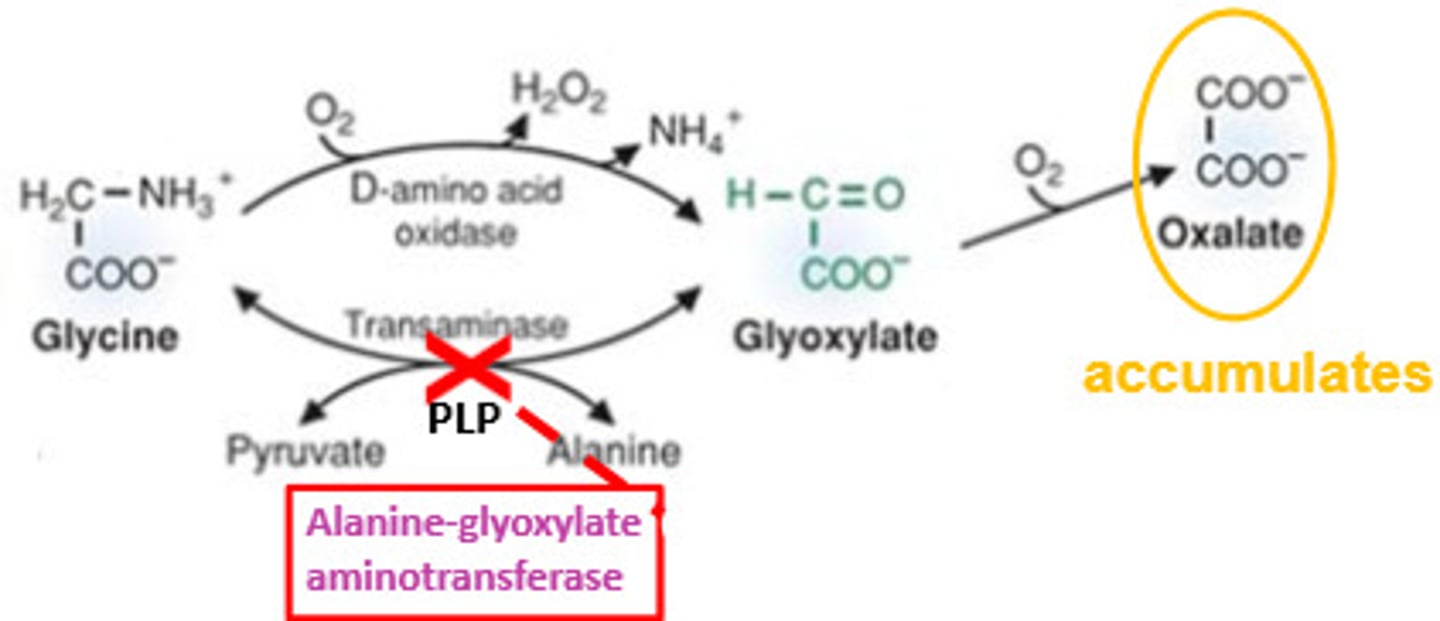
Lumasiran siRNA
siRNA designed to target glycolate oxidase mRNA
The siRNA binds to the glycolate oxidase mRNA and it is destroyed
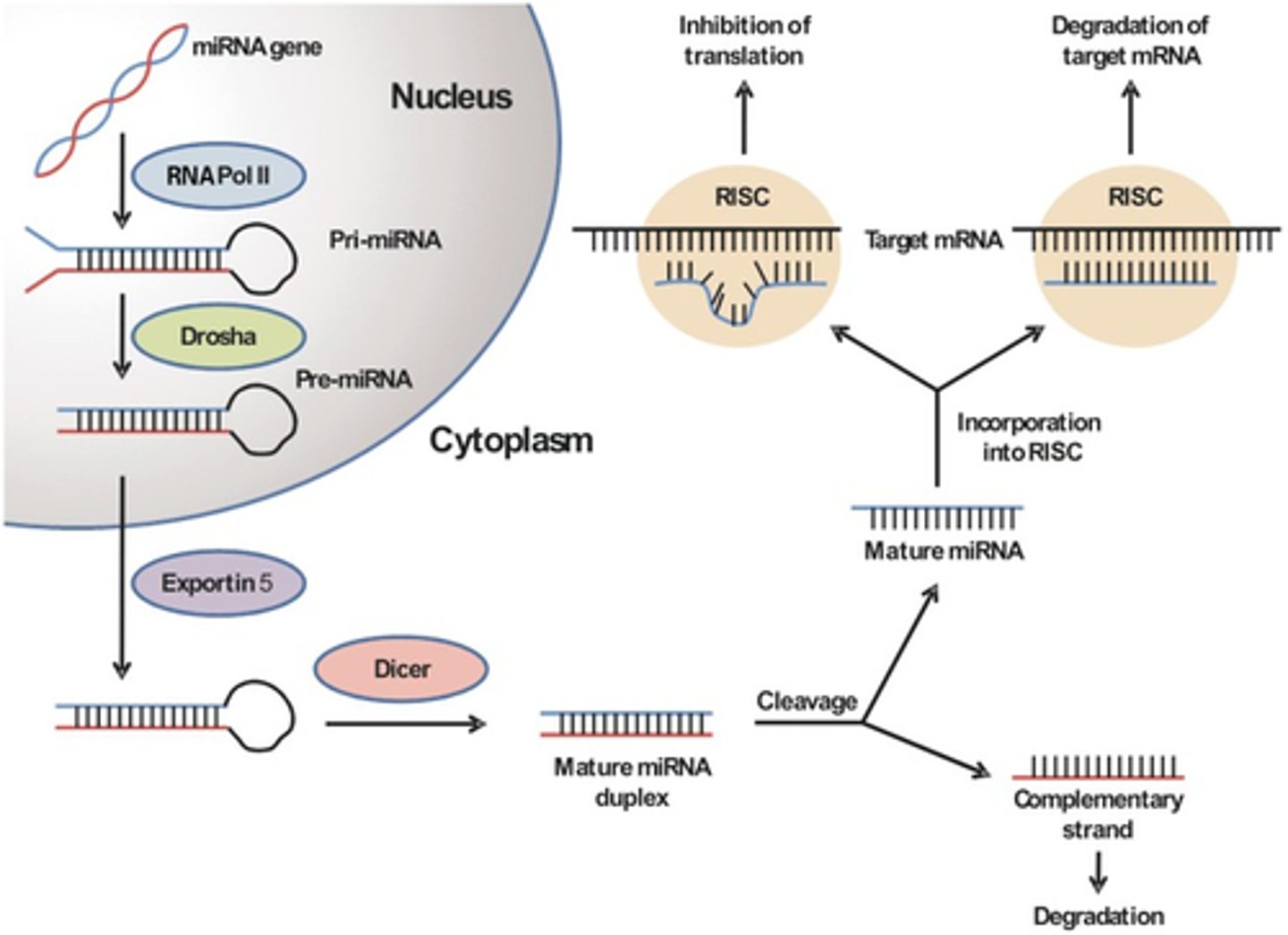
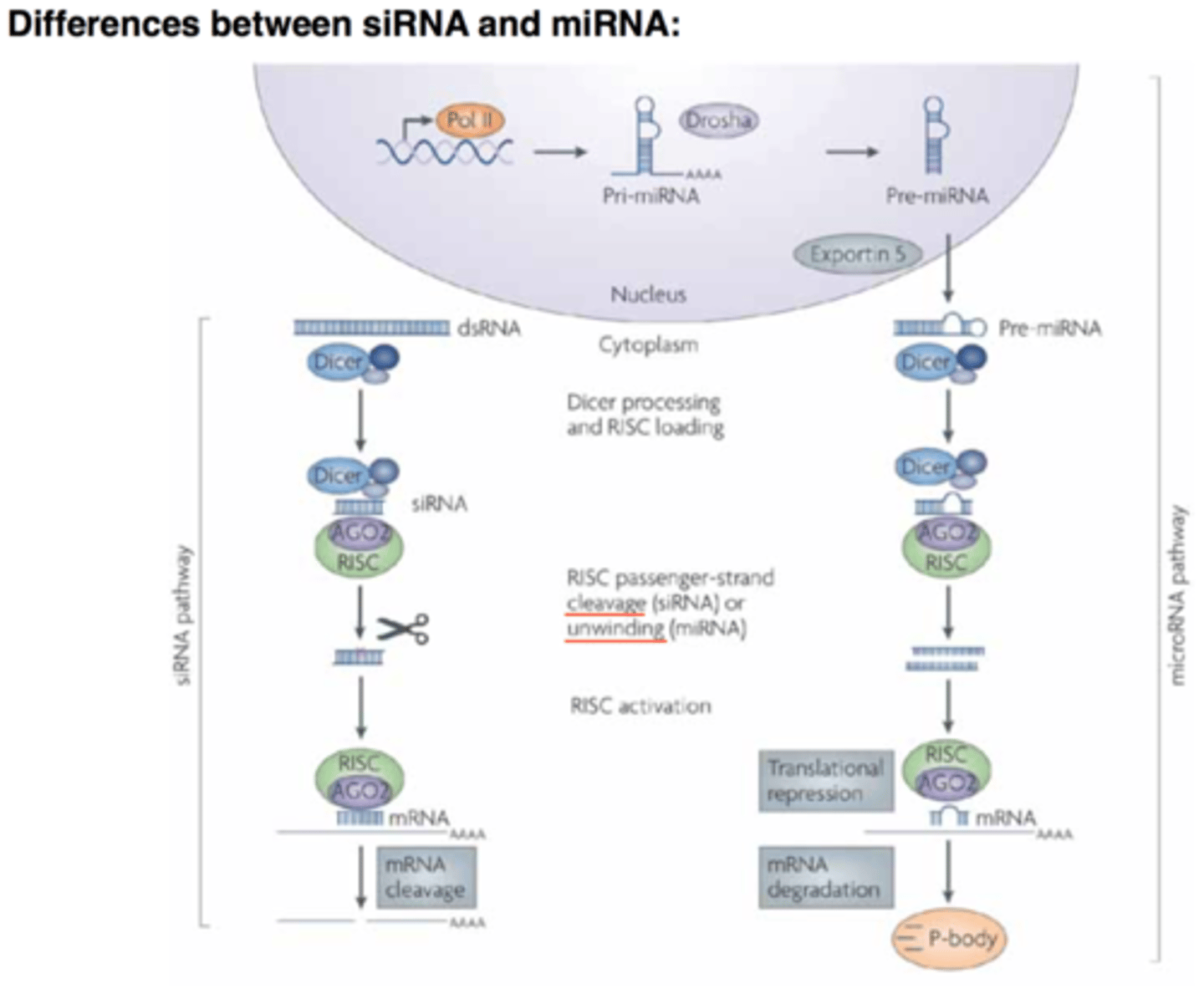
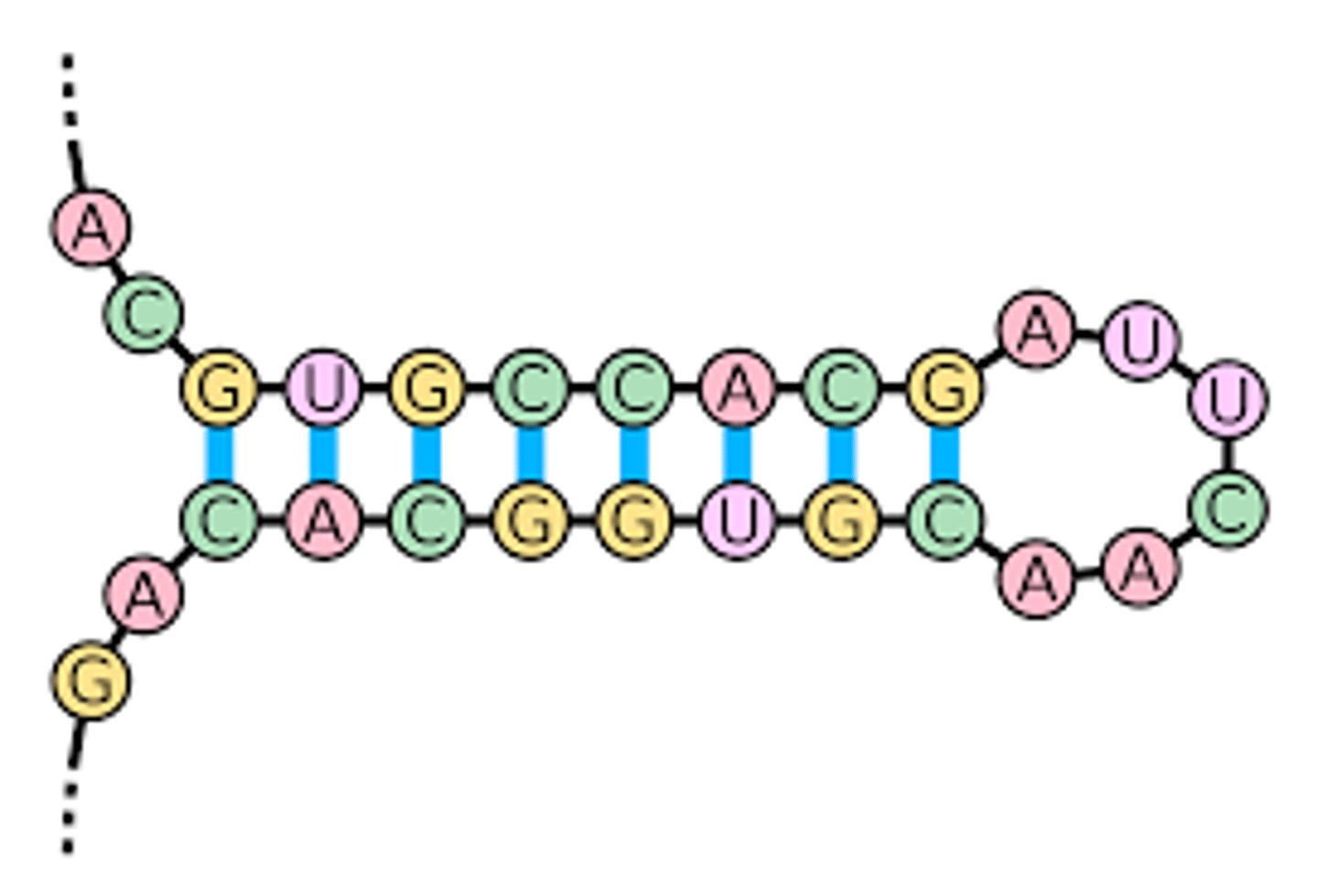
microRNA (miRNA)
1. are always endogenous
2. they originate through TRANSCRIPTION
3. have a stem-loop structure
miRNAs are transcribed from
1. their own independent genes called MIR genes
2. transcribed from specific regions within protein coding genes
3. transcribed from specific regions within non-protein coding genes
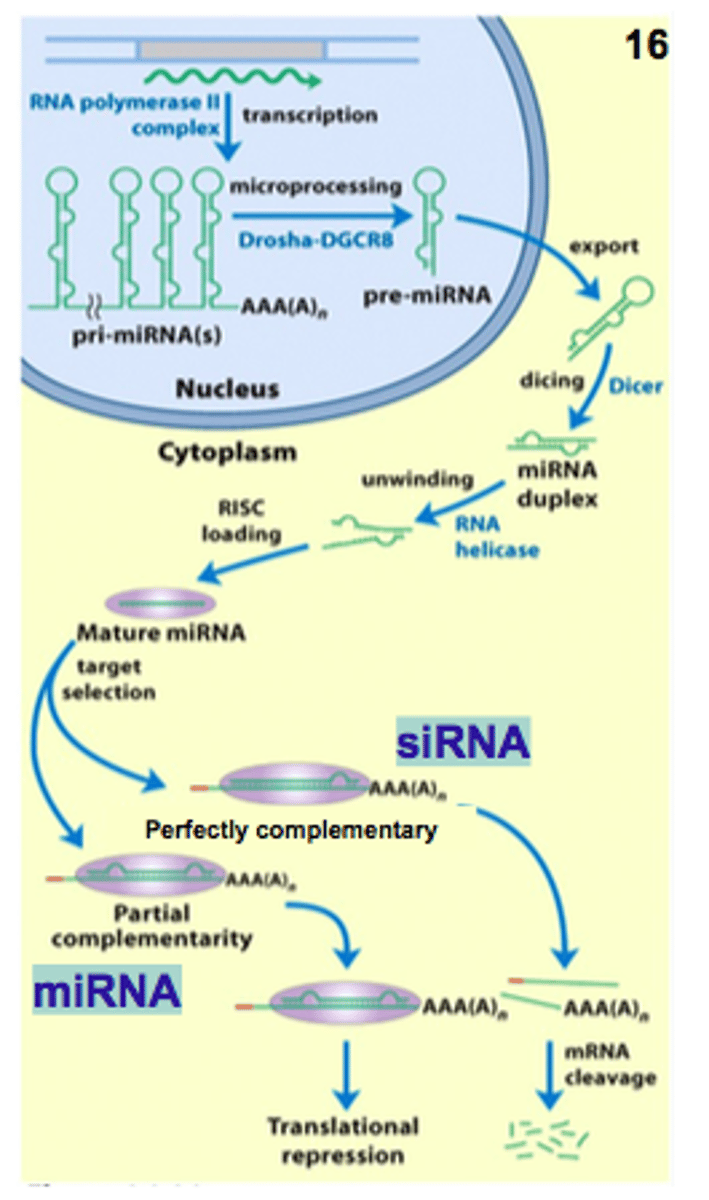
Where are microRNAs synthesized fromin the genome?
Intronic miRNA (nc and coding)
exonic miRNA (nc and coding)
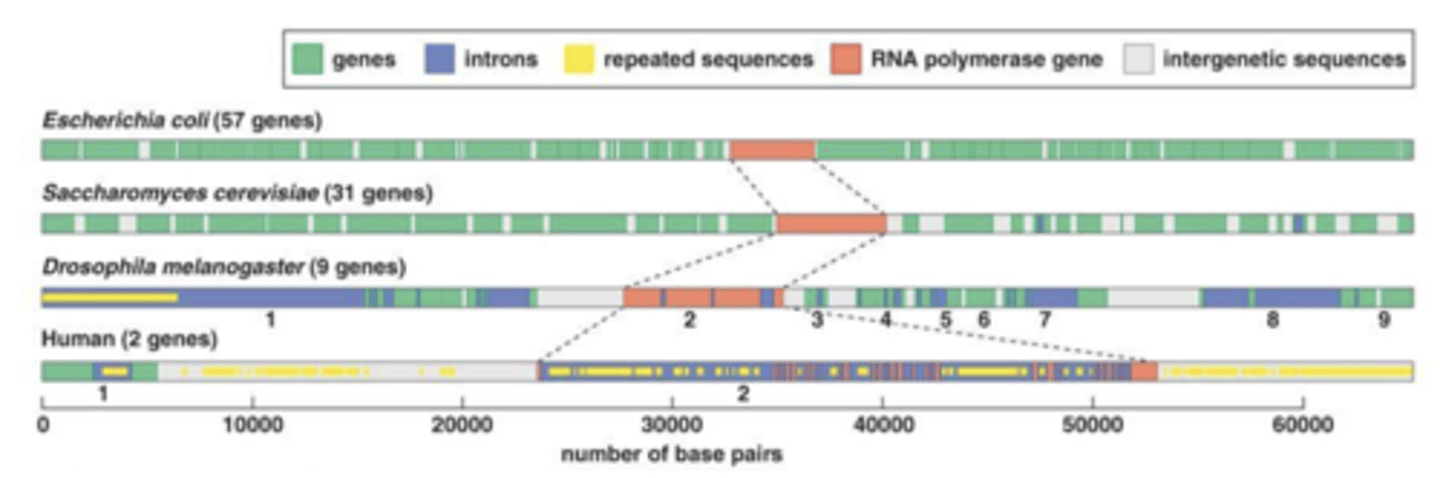
miRNAs have specific names
many are conserved across species
- single miRNA (miR-208a) is located inside Intron 27
- Multiple miRNAs (miR-106b, miR-93,miR-25) are located within Intron 13
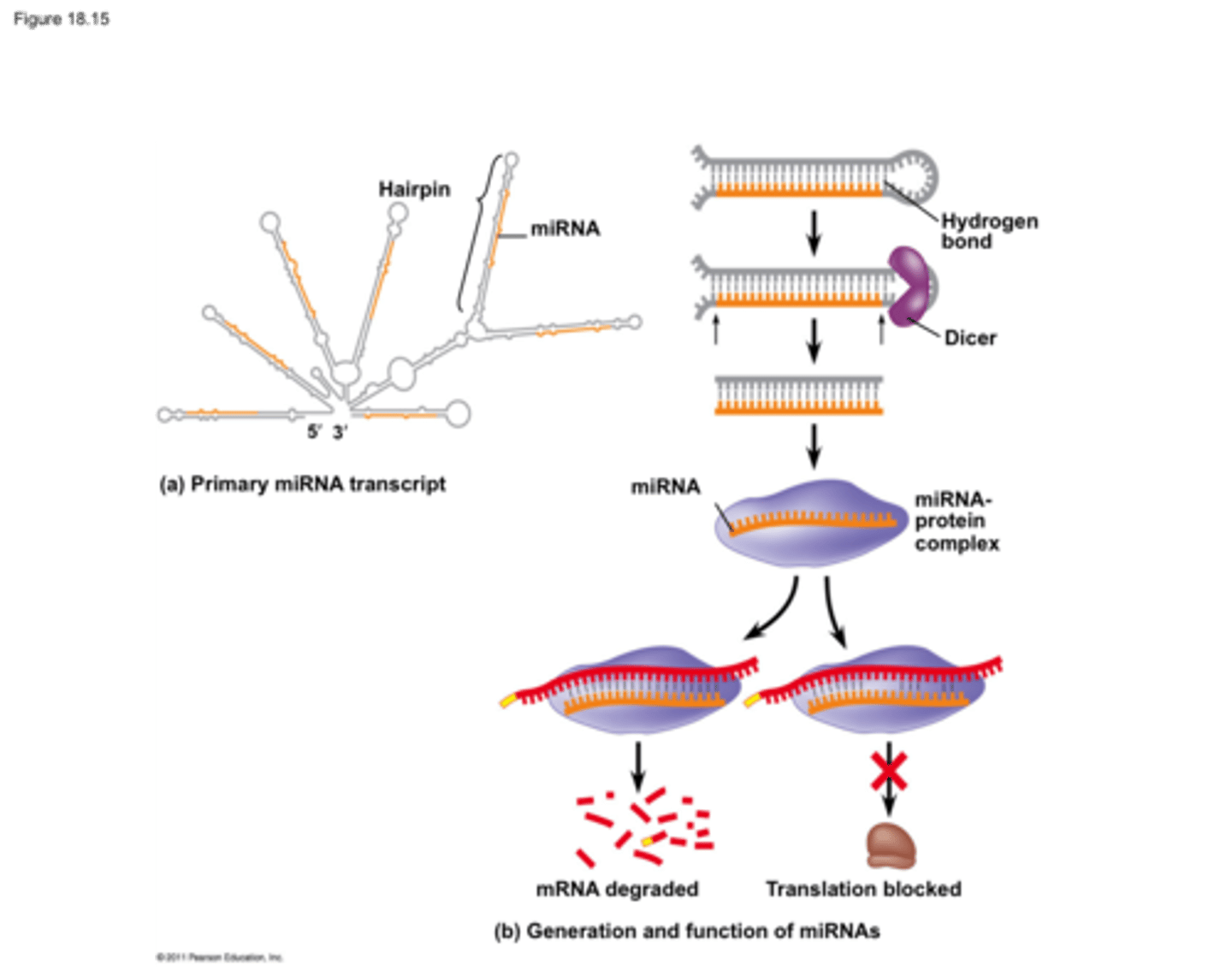
siRNA function
Functions mainly in defense against foreign double-stranded RNA (such as viruses)
Regulate gene expression when too much mRNA is present
Works by binding perfectly to mRNA and destroying it