BIOL 107 Part H - Proteins and Genes (Questions)
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
Rank prokaryotes, photo-autotrophic eukaryotes, and chemo-heterotrophic eukaryotes in terms of number of genes per cell. Where are the genes found in each?
Most: photo-autotrophic in nucleus, plastids, and mitochondria; 2nd: chemo-heterotophic in nucleus and mitochondria; least: prokaryotes in plastids and prokaryotic chromosomes
Genes on chromosomes can be transcribed in which direction?
Left or right, but must follow 5’ to 3’ on new strand and template strand must reflect this
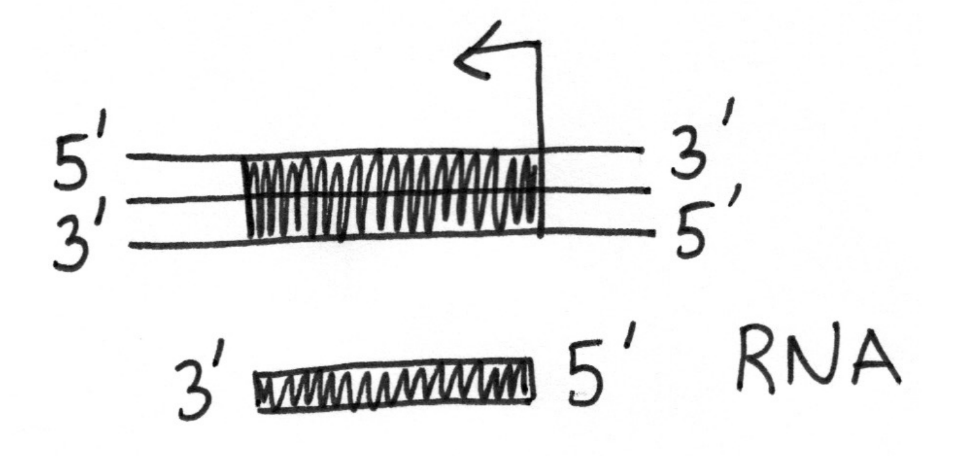
Genes that are transcribed into mRNA are (single/mutliple) copy in the cell. Why?
Single. One gene is sufficient for hundreds of proteins to be made
Genes that are transcribed into tRNA and rRNA are (single/mutliple) copy in the cell. Why?
Multiple, found on several chromosomes. Multiple genes are needed to make enough RNA for millions of ribosomes per cell etc.
Where is genetic material found in the nucleus? Where is tRNA, rRNA, and mRNA genes found?
Nucleolus and euchromatin. Nucleolus contains rRNA genes, euchromatin contains mRNA and tRNA genes. Heterochromatin contains centromeres (not DNA)
Which ribosomes are present in prokaryotes, and which proteins is each responsible for creating? Eukaryotes?
Prokaryotes: 70s ribosomes, create all proteins.
Eukaryotes: 80s ribosomes, 55s ribosomes. 80s ribosomes create vast majority of proteins in cytosol, 55s ribosomes create some proteins in the mitochondria
Describe the mechanism by which amino acids attach to their corresponding tRNA.
The aaRS enzyme for the corresponding amino acid catalyzes the formation of a temporary covalent bond between the tRNA and the amino acid, using ATP hydrolysis as energy
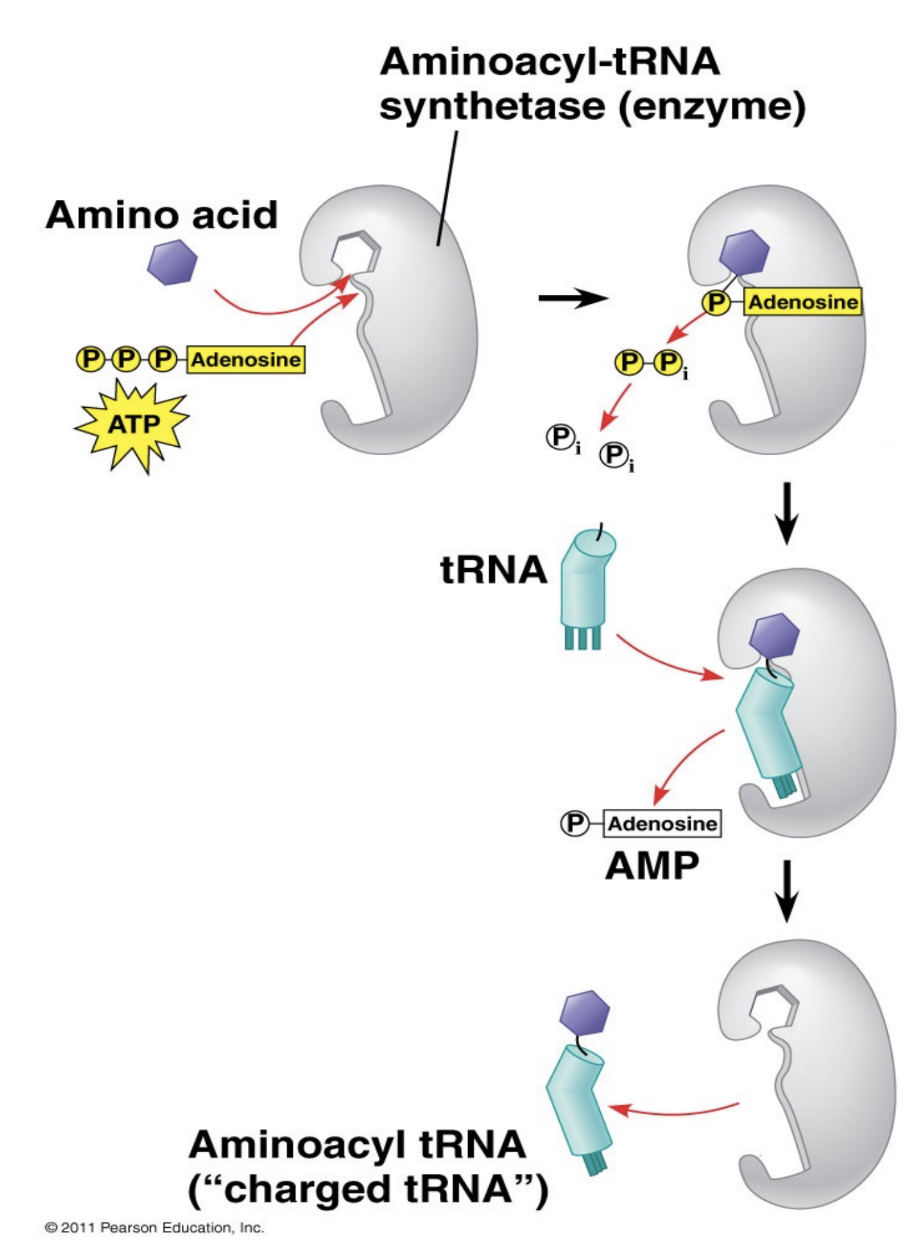
Describe the structure of tRNAphe, including necessary structures and sequences.
At the molecule’s 5’ end is an anticodon for its corresponding amino acid. This flat chain curves up until the 3’ end, where the temporary covalent bond with the amino acid will form
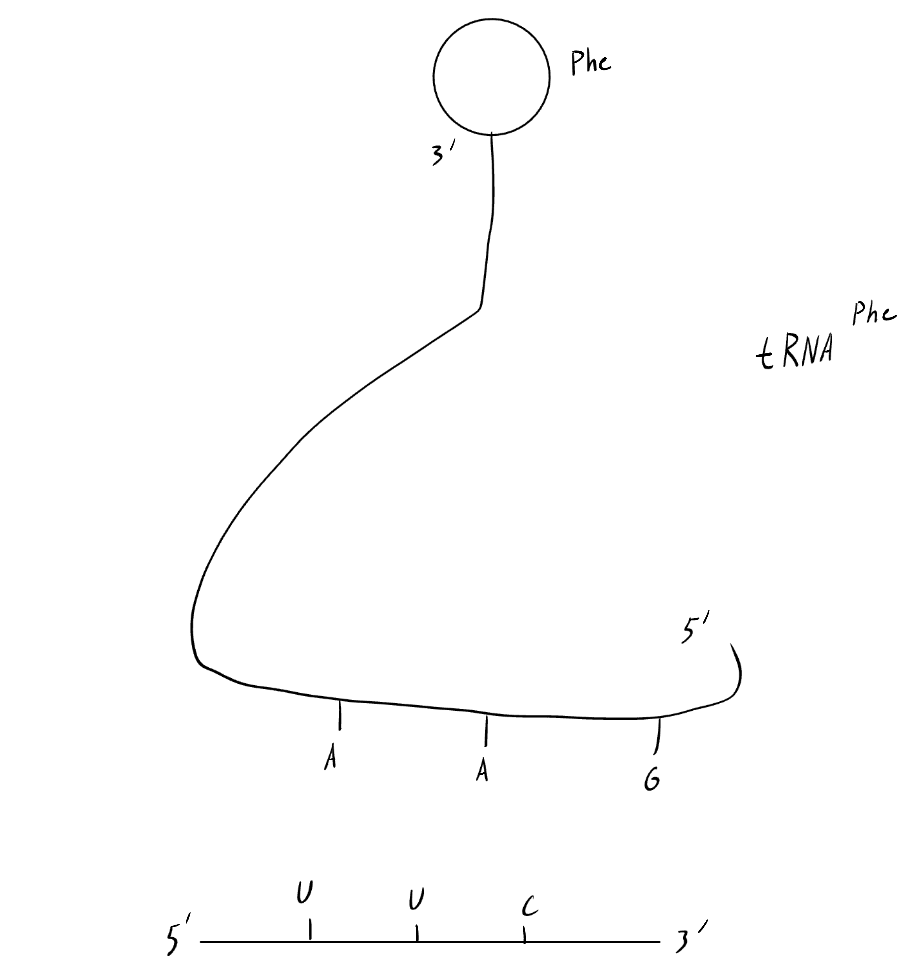
Humans have ~___ tRNA genes in the nucleus.
500
There are 48 amino acids to 61 amino acids. Is this enough? Are tRNAs specific to one type of codon? Why or why not?
Yes, this is enough. tRNA is not specific to one type of codon, wobble position allows for bonding with multiple codons.
From the 5’ end, the first base of the tRNA anticodon is a “wobble” base, which can form non-standard hydrogen bonds with the third base of the mRNA codon. This wobble base can bond with non-complementary bases, allowing for a tRNA with a specific anticodon to bond with more than one type of codon
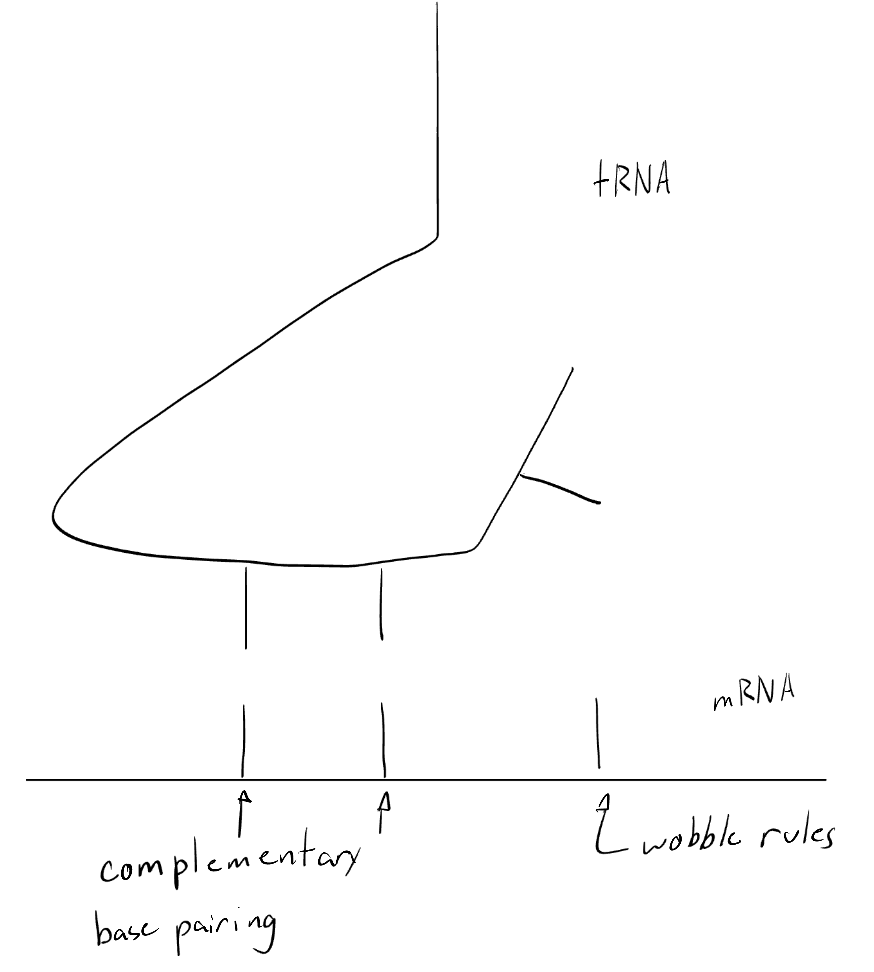
Which tRNA carries the amino acid methionine? Explain in detail why this is needed.
Two different ones: tRNAMeti and tRNAMet. They both carry the anticodon to Methionine’s codon AUG, but since this also codes for the start of a sequence, there needs to be a distinct tRNA for initiating and a distinct tRNA for an internal amino acid so that no confusion occurs
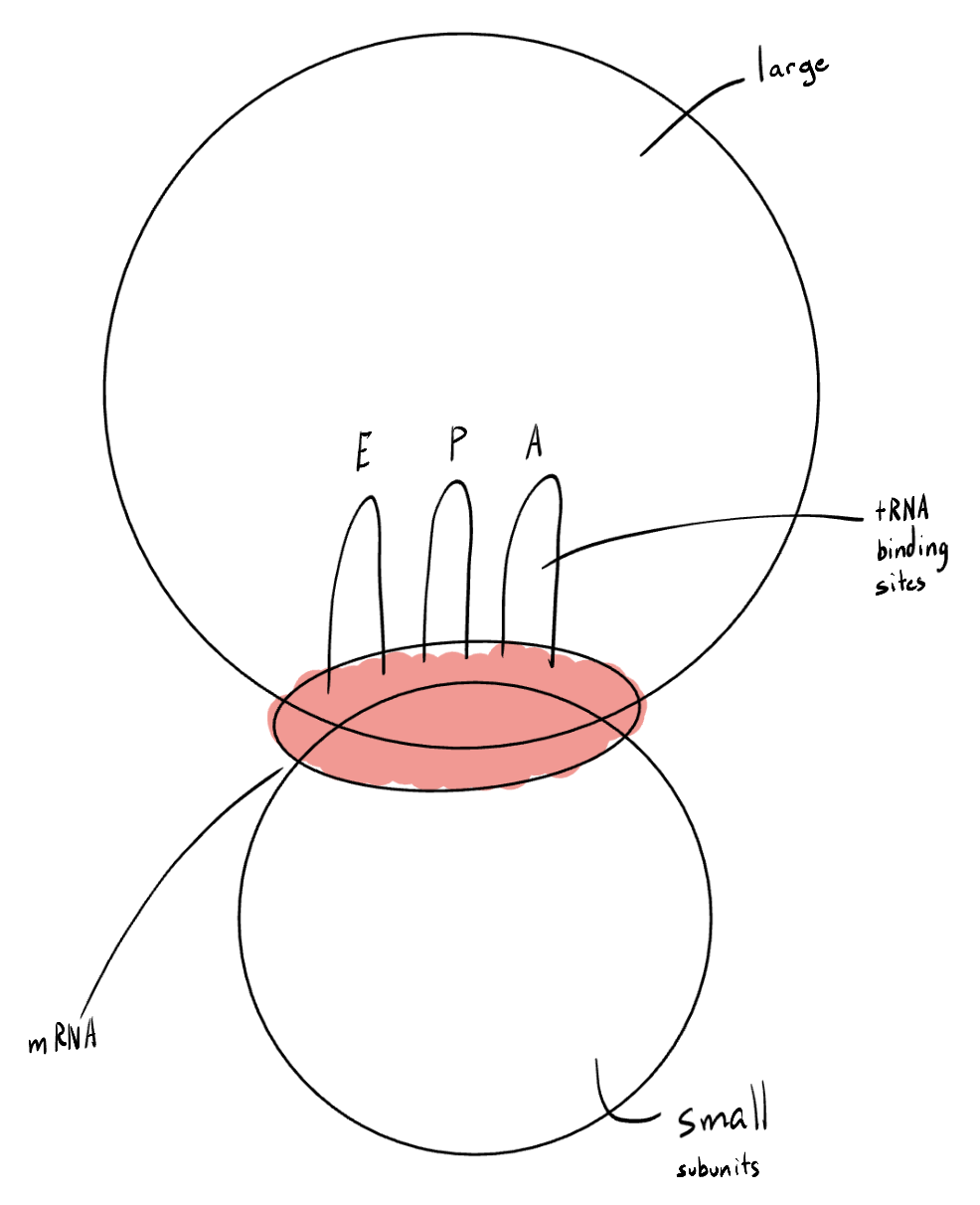
Describe the structure of a ribosome, including subunits and sites.
The ribosome has a small subunit and a large subunit. An mRNA binding site is situated between the two subunits. On top of this, there are 3 tRNA binding sites: E (tRNA exit, where uncharged (empty) tRNA molecules bind), P (polypeptide tRNA site, where a growing polypeptide chain bonds), and A (amino acid tRNA site, where an initially entering tRNA molecule during elongation with only one amino acid attached bonds).
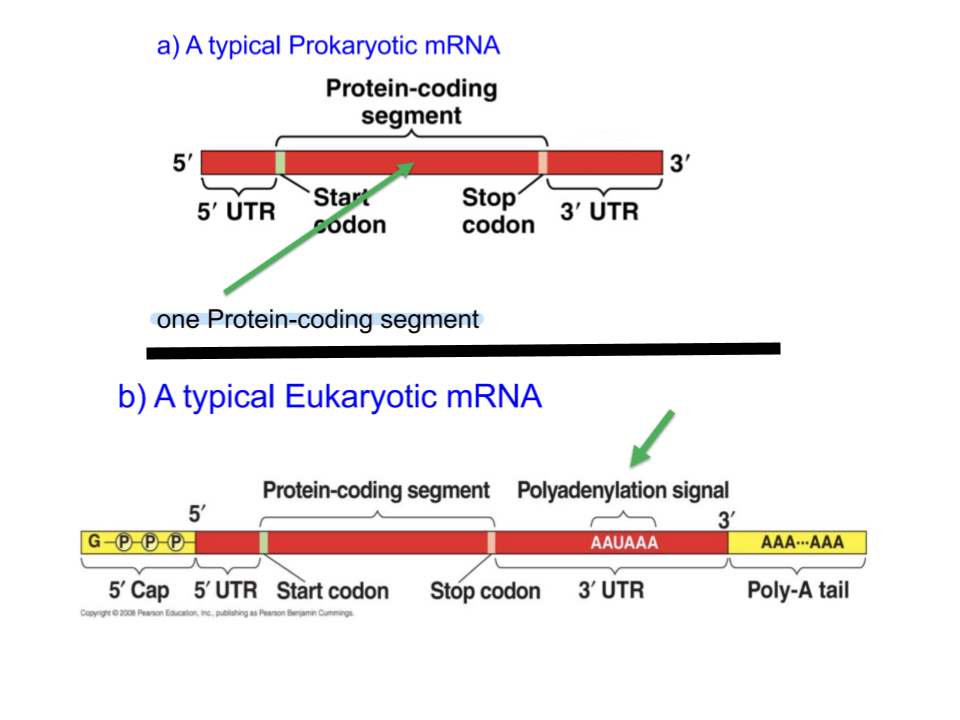
Compare prokaryotic and eukaryotic mRNA in terms of structure.
Both contain protein codiing segment surrounded by two UTRs, but eukaryotic mRNA uniquely contains a 5’ cap, polyadenylation signal, and a poly(A) tail.
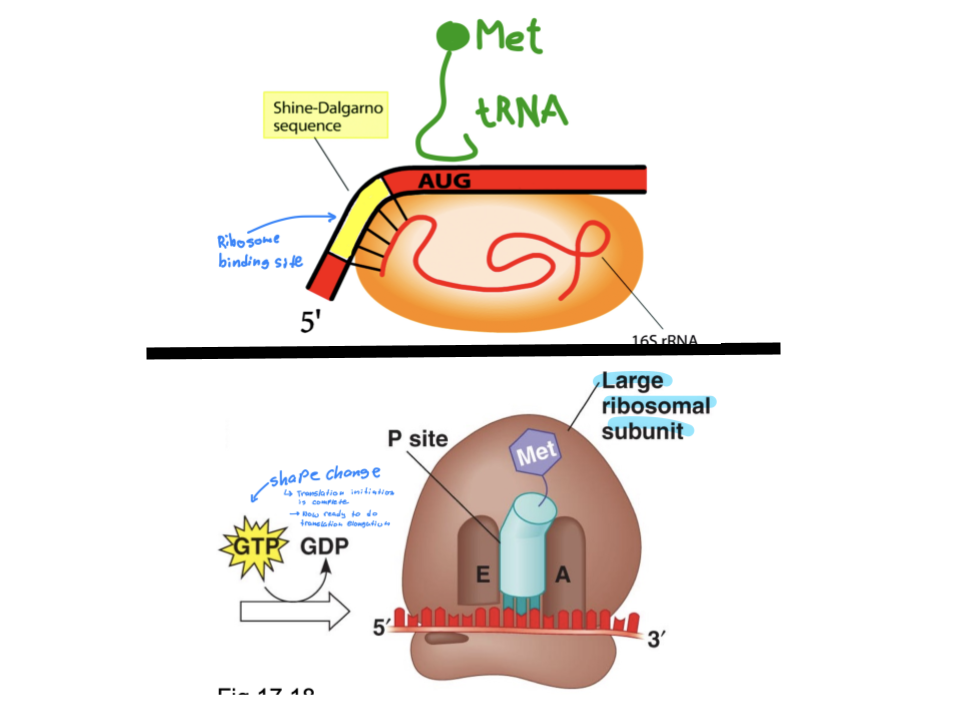
Explain the process of transcription initiation in prokaryotes in both the small and large subunit of the ribosome. What is the ribosome binding site? Which tRNA is used?
Ribosome binding site at the 5’ UTR of the mRNA molecule is located right next to the start codon (start codon is right next to the RBS). The small subunit of the ribosome binds to this site while tRNAMeti binds with the start codon directly.
The hydrolysis of GTP changes the shape of the small subunit, allowing the large subunit to bond. The tRNA and AUG codon bonded complex will fit into the P site of the large subunit, acting as a base ready to form a longer chain of amino acids.
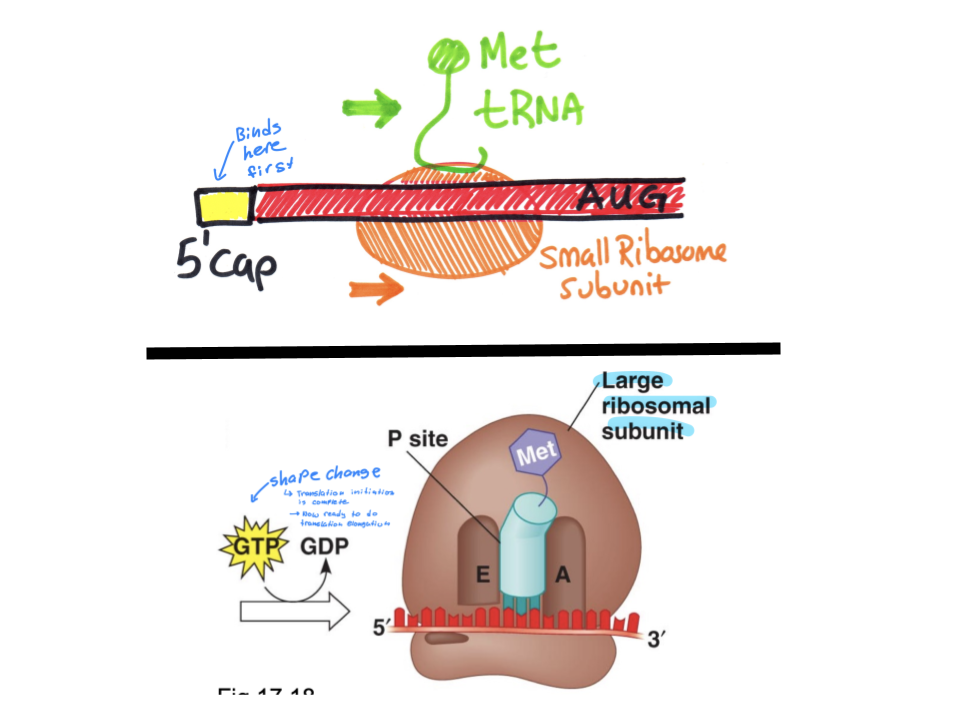
Explain the process of transcription initiation in eukaryotes in both the small and large subunit of the ribosome. What is the ribosome binding site? Which tRNA is used?
Ribosome binding site is the 5’ cap, which is located one UTR away from the start codon (start codon will be the AUG closest to the RBS). The small subunit of the ribosome first binds to the 5’ cap, and the tRNAMeti binds to the small subunit. The subunit and tRNA complex must “slide” across the mRNA, scanning the bases until the start codon (AUG) is reached, and the tRNA will bind to the start codon.
The hydrolysis of GTP changes the shape of the small subunit, allowing the large subunit to bond. The tRNA and AUG codon bonded complex will fit into the P site of the large subunit, acting as a base ready to form a longer chain of amino acids.
Explain the process of transcription elongation. What is the initial situation before this step?
The P-site contains a tRNA molecule bonded to the start codon which acts as a base to create a polypeptide chain. The a-site is empty and ready for new amino acid tRNAs to enter.
The correct aa-tRNA enters and stays in the A-site, bonding with its corresponding codon. The peptide bond between the P-site polypeptide tRNA and its corresponding amino acid/amino acid chain is broken, and a new peptide bond is formed between the existing chain and the a-site amino acid tRNA. The formation of a polypeptide releases a water molecule. The ribosome will slide 3 nucleotides (a codon) forward along the mRNA, causing the now spent P-site tRNA to slide to the E site and exit the ribosome (no longer bonded to mRNA) and the now chained A-site tRNA to slide to the P-site. The A-site is now free and ready for the next correct amino acid to enter.
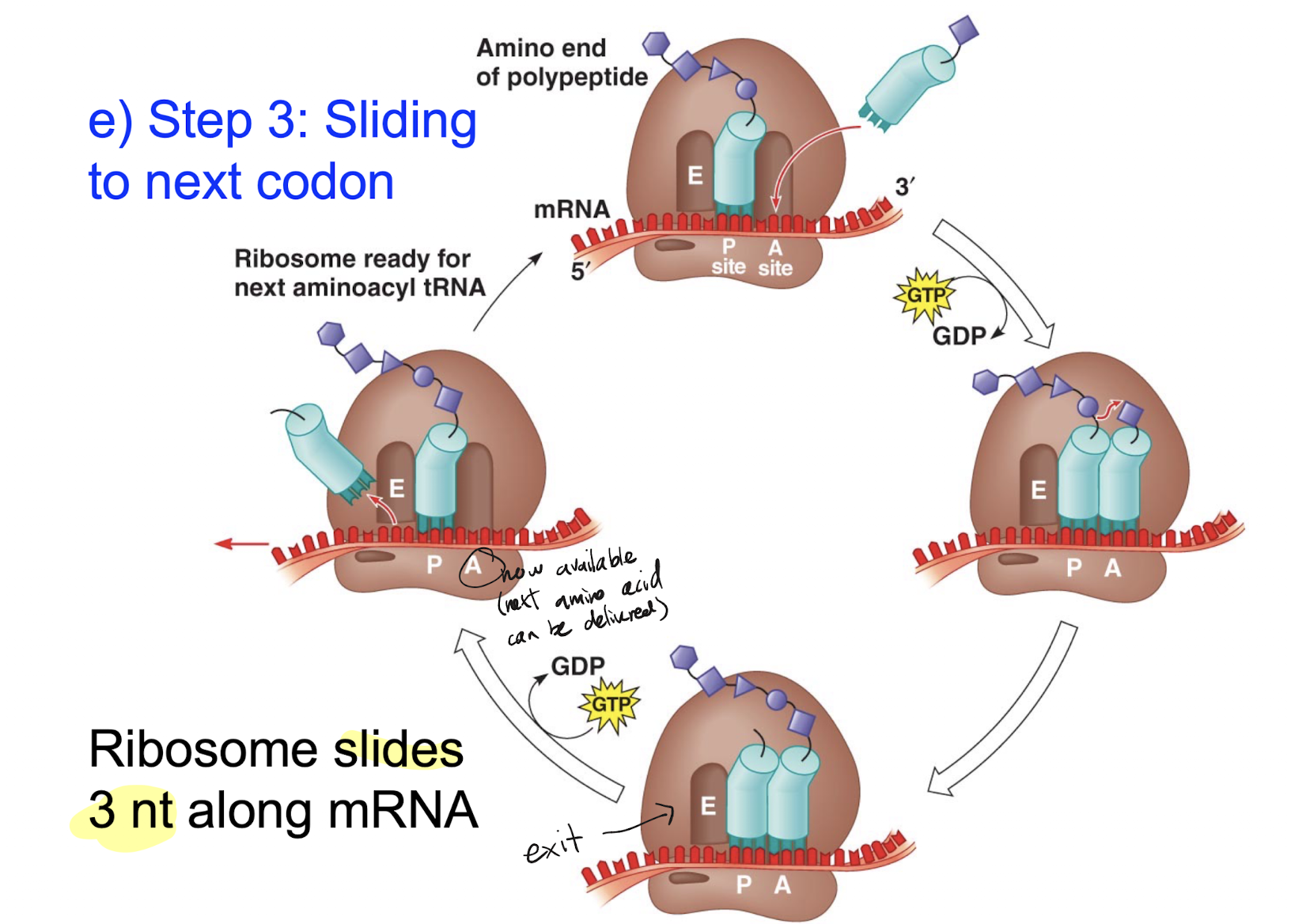
What is the role of GTP and ATP hydrolysis in the breaking and forming of polypeptide bonds between amino acids during translation elongation?
ATP hydrolysis will provide the energy needed to form the polypeptide bonds between the aa and the tRNA as well as the aa with each other. GTP does not provide energy, it changes the ribosome shape to allow for aa-tRNA to bond to the A-site.
Can any amino acid tRNA enter the A-site? How does the ribosome ensure the correct polypeptide sequence is formed?
Yes, any aa-tRNA can enter, but only the correct one can stay in the site and continue to peptide bond formation. The ribosome will only continue if the aa-tRNA is contains the correct anticodon that matches with the corresponding mRNA codon
In translation elongation, the new amino acid is added (above/below) the previous amino acid. Why, given the amount of cuts?
Below. Even though adding the new amino acid above onto the polypeptide chain would only be one cut and one connection, as the chain gets longer the distance is much too far to viable
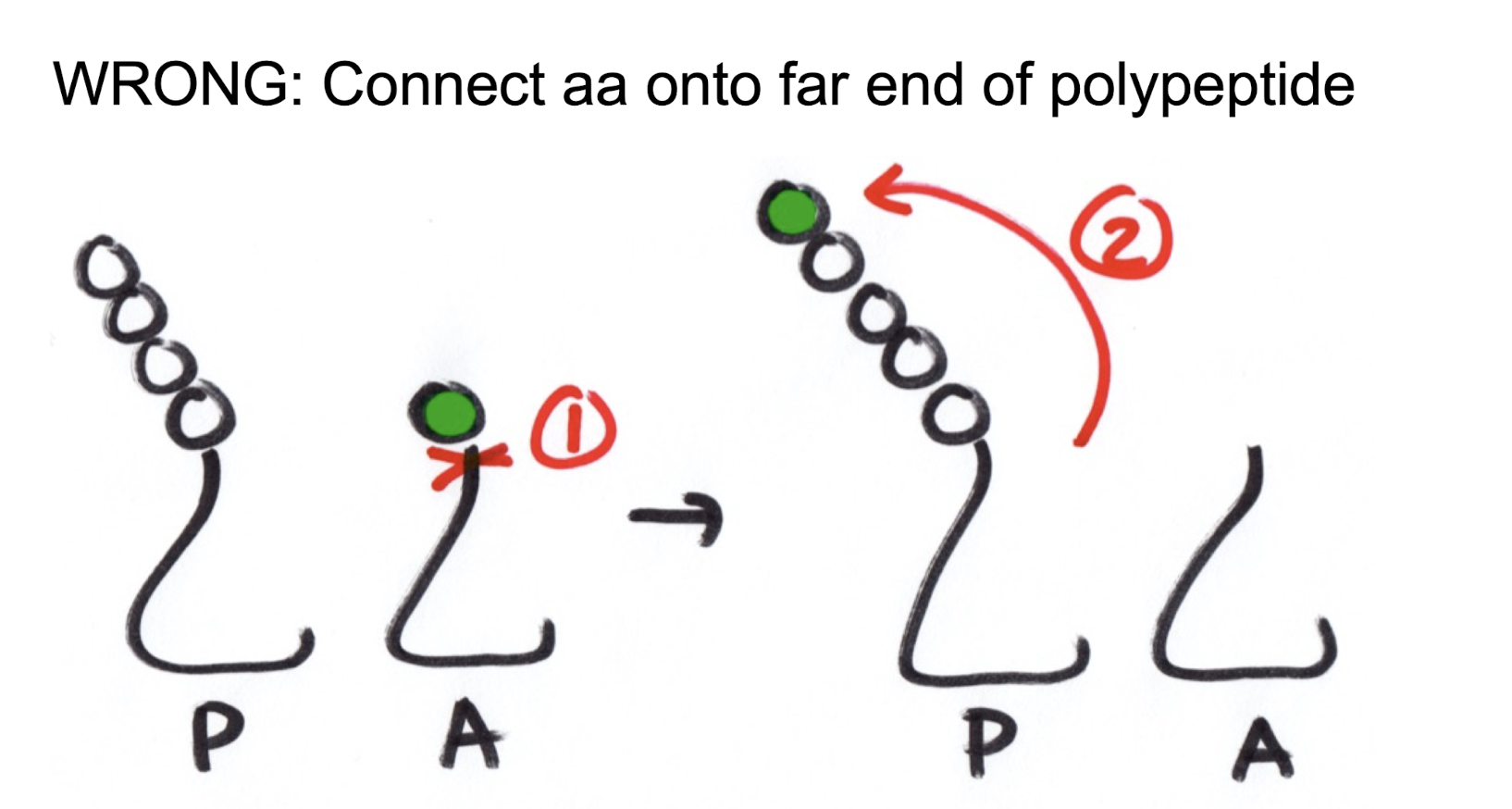
Why is the polypeptide attached onto the aa-tRNA in translation elongation instead of the new amino acid being connected to the polypeptide tRNA?
Shifting the polypeptide onto the new amino acid allows for one cut (polypeptide chain from tRNA) and one connection (chain onto new aa), which is more energetically simplistic. Otherwise, it would be required to cut the chain off the polypeptide tRNA, cut the new amino acid from the aa-tRNA, and then connecting the new amino acid to both the polypeptide tRNA and the polypeptide chain. This is 2 breaks and 2 connections, which requires much more energy and is not desired
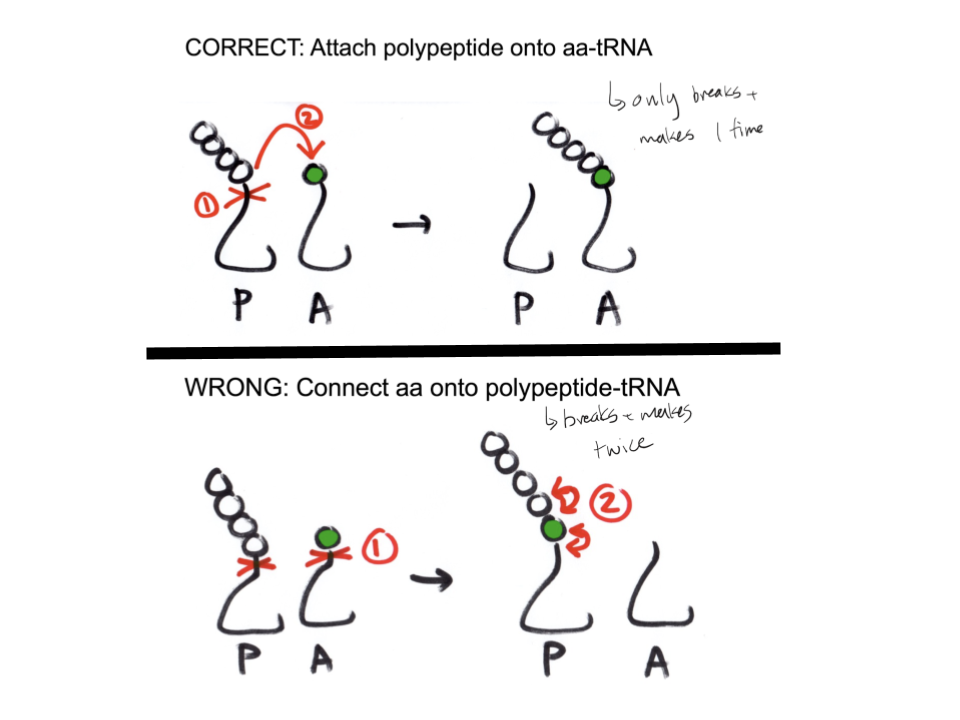
Explain the process of transcription elongation.
Once the A-site reaches a stop codon, a release factor protein will enter the A-site and bind to the mRNA. The release factor facilitates the polypeptide to attach to a water molecule, allowing it to be released from the polypeptide tRNA and ribosome (becomes free-floating). This will cause the mRNA, protein subunits, and release factor to dissassemble
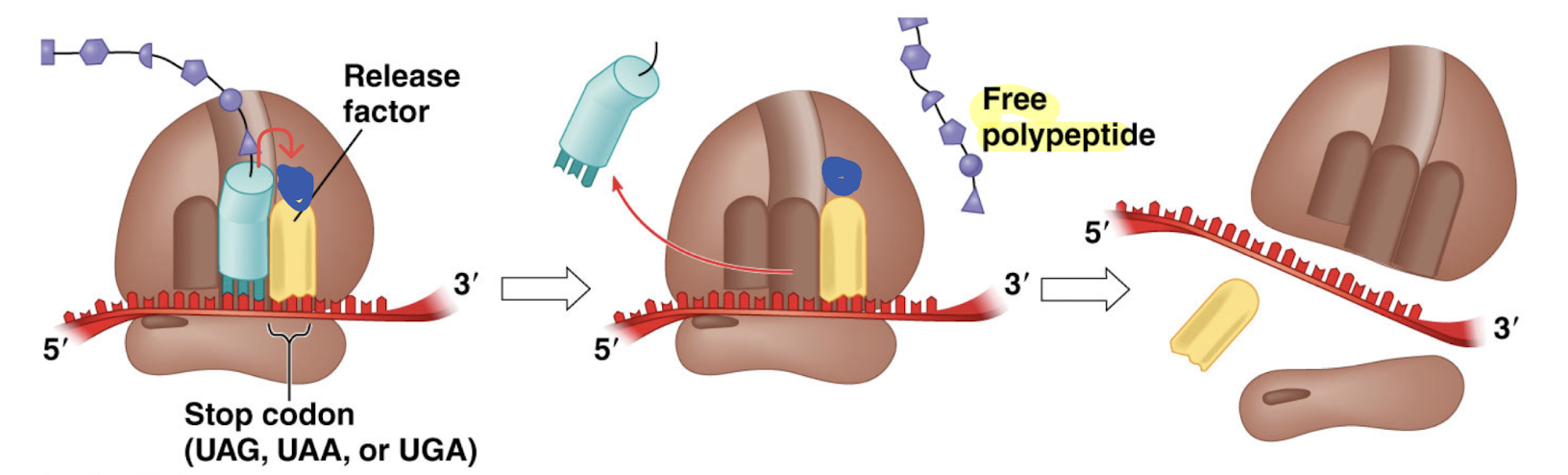
Terminators and promoters are sequences found on ___, used in (protein/RNA) synthesis. Start and stop codons are sequences found on ___, used in (protein/RNA) synthesis.
DNA, RNA. RNA, protein
What is the purpose of having UTRs on an mRNA strand?
Efficiency by helping ribosomes translate mRNA and longevity by protecting the mRNA
Compare the relative sizes of eukaryotic genes, mRNA molecules, and proteins. Explain discrepancies.
Genes are very long (~16000 bp). mRNA genes are much shorter than genes (~1000 nt) as through transcription introns are removed. Translated proteins are relatively small, as not all of the mRNA molecule is actually used in translation (coding sequence is only ~500 nt in the middle of the molecule, the rest makes up poly(A) tail/5’ cap/UTRs)
What determines the primary, secondary, tertiary, and quaternary structure in a mature protein? Explain.
Primary is determined by the mRNA sequence that it is coded from. This determines which amino acids make up the chain
Secondary is determined by primary structure. A specific chain of amino acids determines the folded sections the protein makes (ex. Ala-ala-ala-ala makes alpha helix)
Tertiary is determined by primary structure. The amino acids will determine which ones disulfide bonds are created between, determining the 3D structure (ex. location of a cys-cys will determine where the disulfide bond happens and fold)
Quaternary structure is determined by primary structure. Same explanation as tertiary
Eukaryotes have (one/multiple) protein encoding sections and ribosome binding sites per mRNA. What about prokaryotes?
One, each mRNA codes for a single protein. Prokaryotes have operons, which create mRNAs that each have several protein coding regions and thus multiple ribosome binding sites (one per start codon/protein coding region)
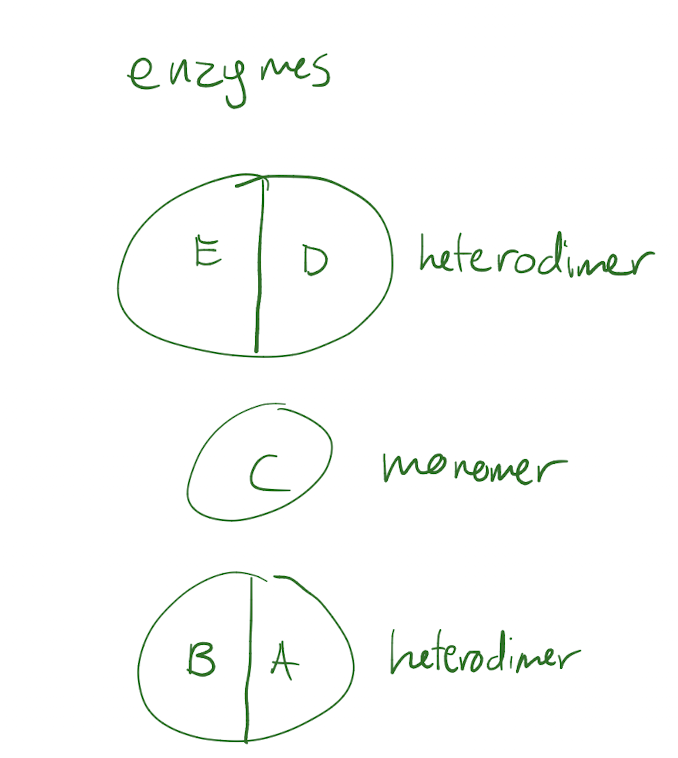
The Trp operon creates 3 enzymes: 2 heterodimer enzymes and one monomer enzyme. How many different polypeptides does it code for?
5
The signal for movement to a specific location of the cell is found in (prokaryotic/eukaryotic) proteins, and the signal for creation in the ER is found in (prokaryotic/eukaryotic) proteins.
Both, eukaryotic
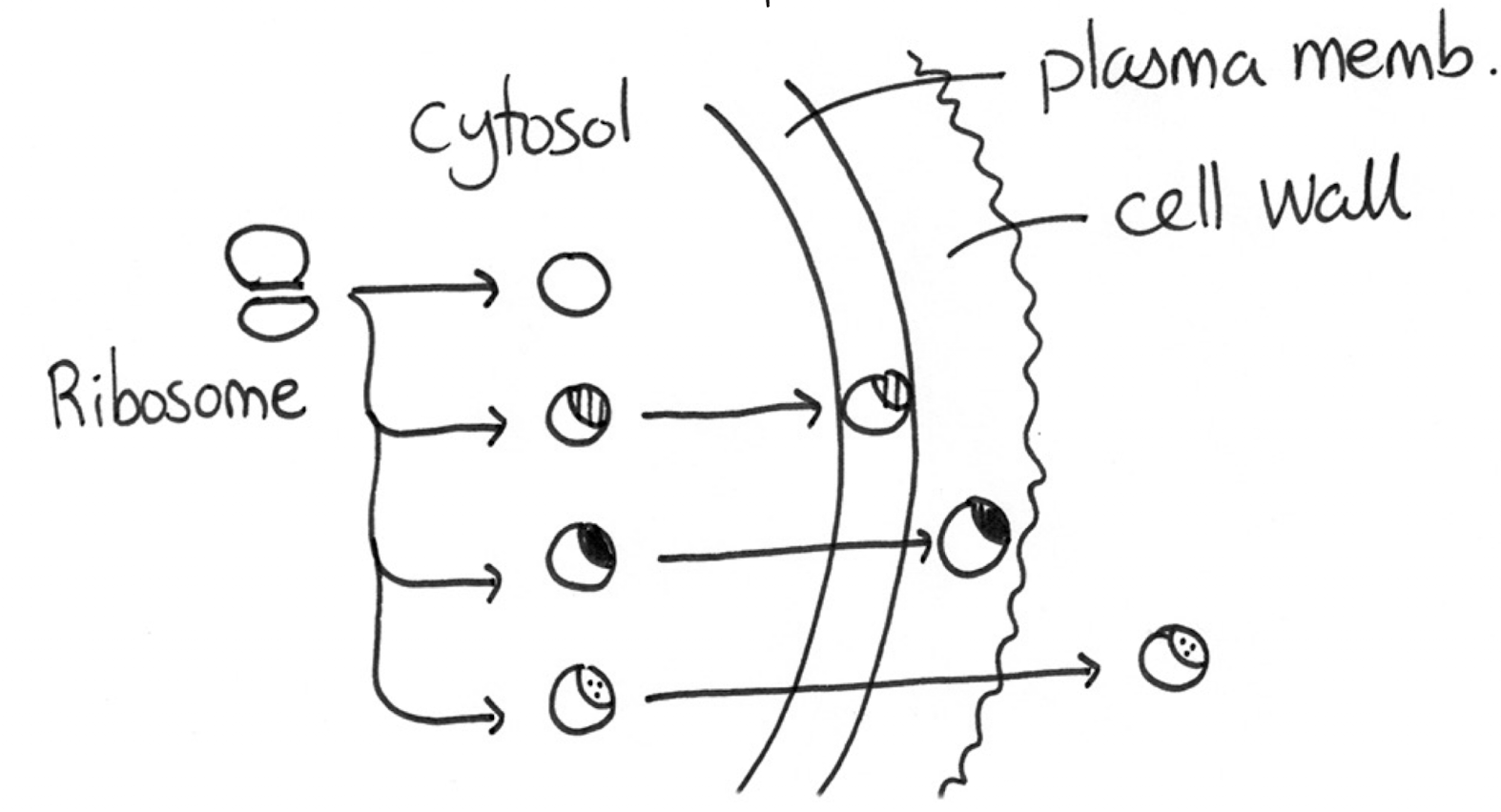
Which signals do prokaryotic cytosol, plasma membrane, cell wall, and export proteins have?
Any proteins that are not needed in the cytosol (plasma membrane, cell wall, export) will have each unique signals that transport them to their correct places, while cytosol proteins do not need signals because they don’t need to be moved elsewhere
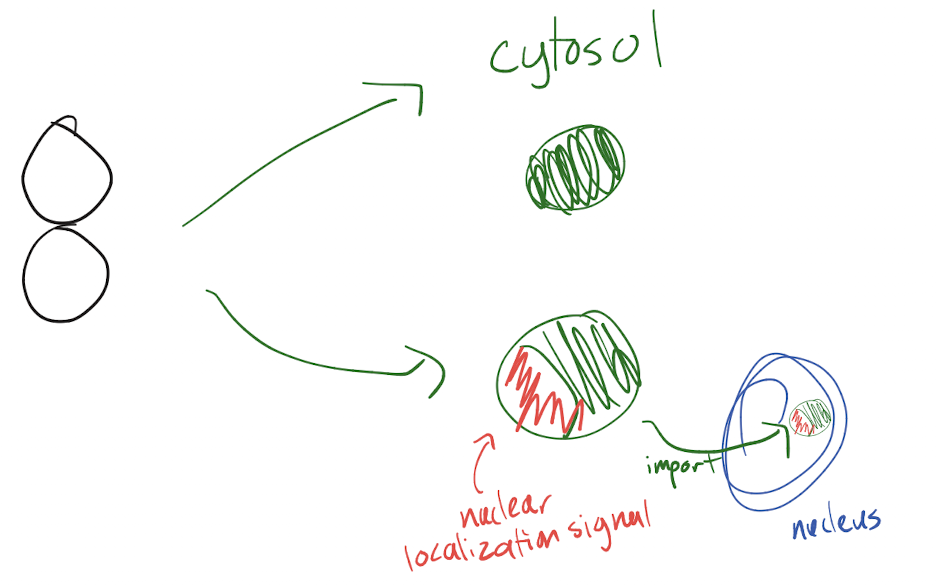
Which eukaryotic proteins are made in free ribosomes in the cytosol? Which signals do each of these proteins have?
Nucleus proteins (ex. DNA polymerase): nuclear localization signal (“put me in” for nucleus)
Cytosol proteins: no signal because no need to be moved
Organelle (mitochondria and chloroplast) proteins: organelle localization signal (“put me in”)
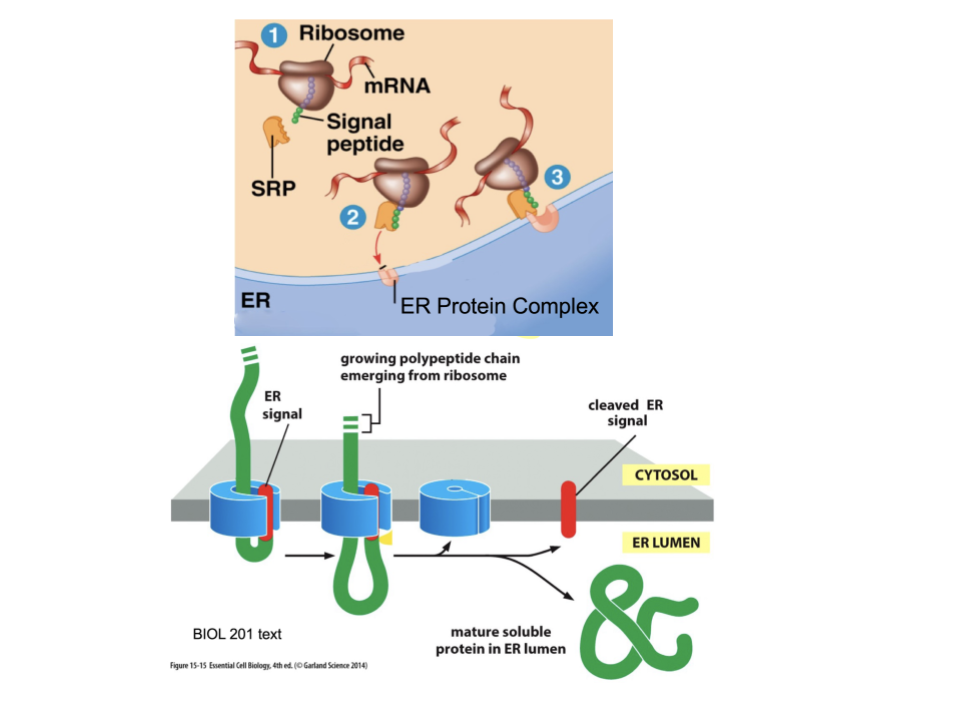
Proteins that need to be made at the ER will have a special signal. Explain this process.
Synthesis will begin in a free ribosome in the cytosol. ER made proteins will have a specific signal, and once synthesized, this signal will recruit an SRP. The SRP grabs the signal peptide sequence and delivers the protein to the ER membrane.
Which eukaryotic proteins are made in the ER? Which signals do each of these proteins have? Explain the process by which each of these molecules move.
Export protein: has ER creation signal as it is needs to be matured in the ER. No movement signals as transport vesicles will exocytose them after modification, so it is directly fed into the ER lumen to mature
Plasma membrane protein: has ER creation signal as it is needs to be matured in the ER. Has a plasma membrane localization signal, and so is placed into ER membrane instead of entering ER lumen. Transport vesicle, upon exocytosis, will push its phospholipid membrane (and thus the membrane protein) into the plasma membrane
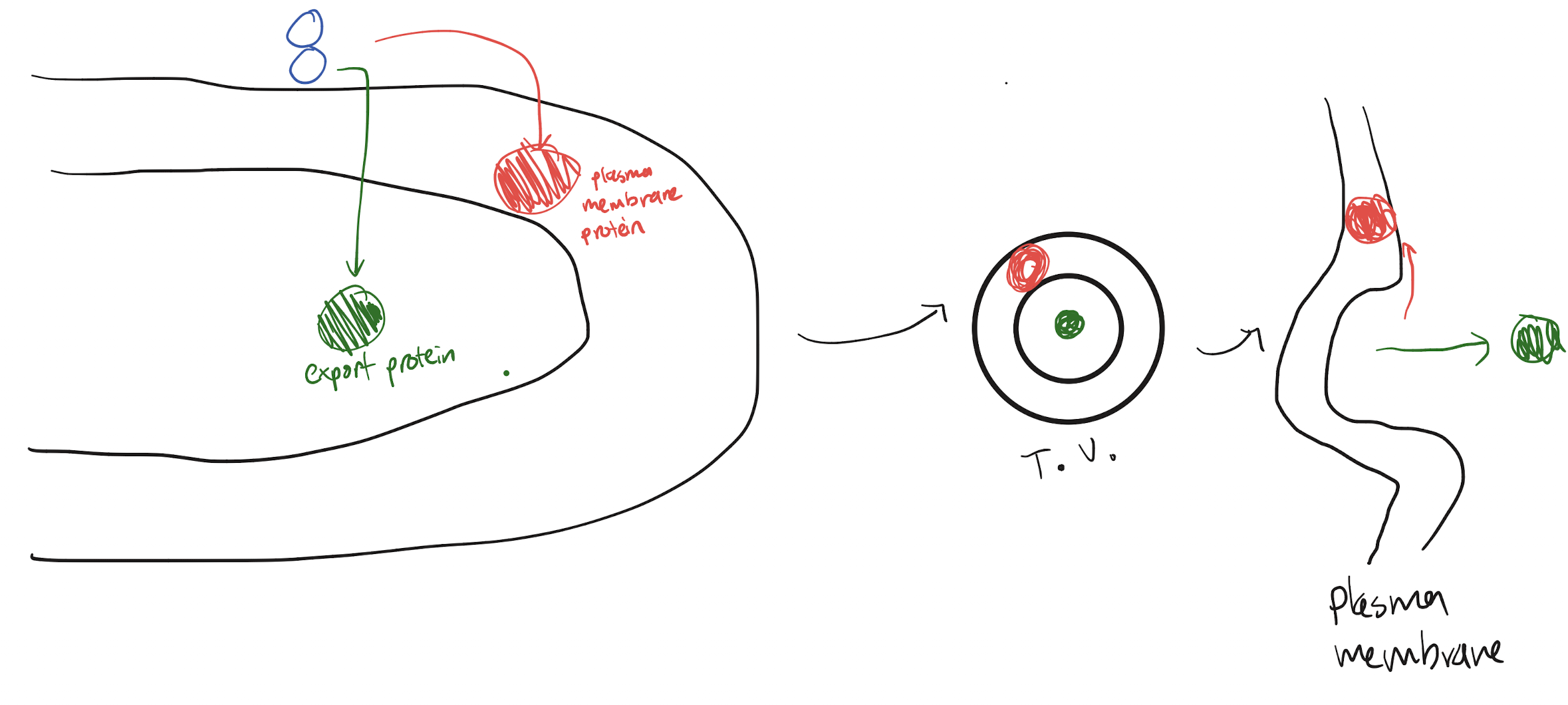
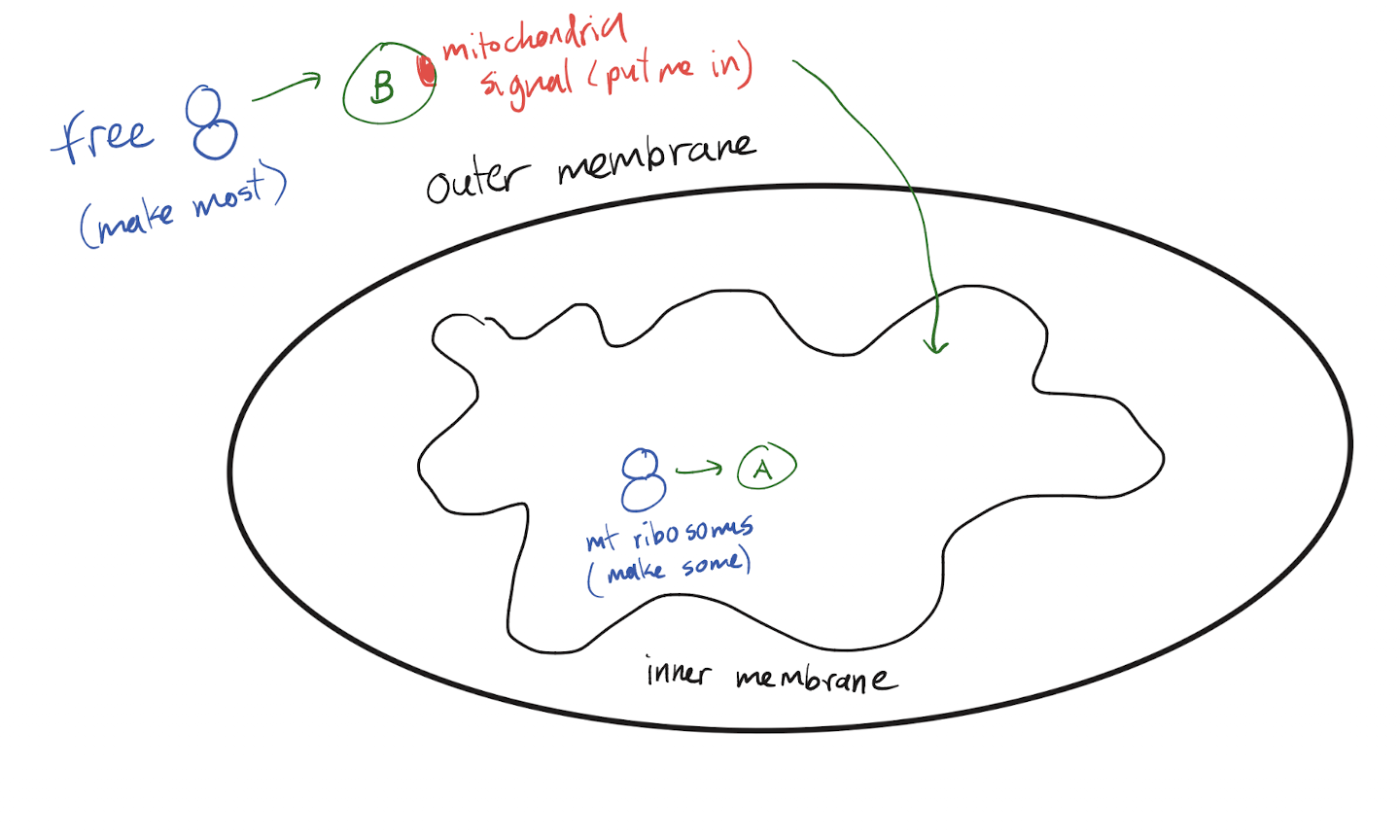
Where are proteins needed in semi-autonomous organelles (mitochondria, chloroplast) created? How do they reach the organelle (include any signals)?
Free ribosomes in the cytosol create most of the proteins needed for these organelles. These proteins have an organelle localization signal so that they can be transported to the correct organelle.
These organelles also have their own ribosomes, which allows them to create some of them that are needed. No signals on these proteins as they don’t need to move.
Why do unicellular organisms regulate genes? What about the cells in multicellular organisms?
Unicellular: Only make proteins needed for the environment so that excess amino acids aren’t wasted
Multicellular: about half the genes are on at a time so that proteins are only created for the specific type of cell that needs it
When can gene regulation occur? When does it typically occur and why?
Can occur at any step of protein synthesis, but usually occurs during transcription initiation (very first step). The cell will usually determine whether a protein is needed before starting the process and stop/start protein synthesis then, as if the protein is not needed there is no point in starting the process at all
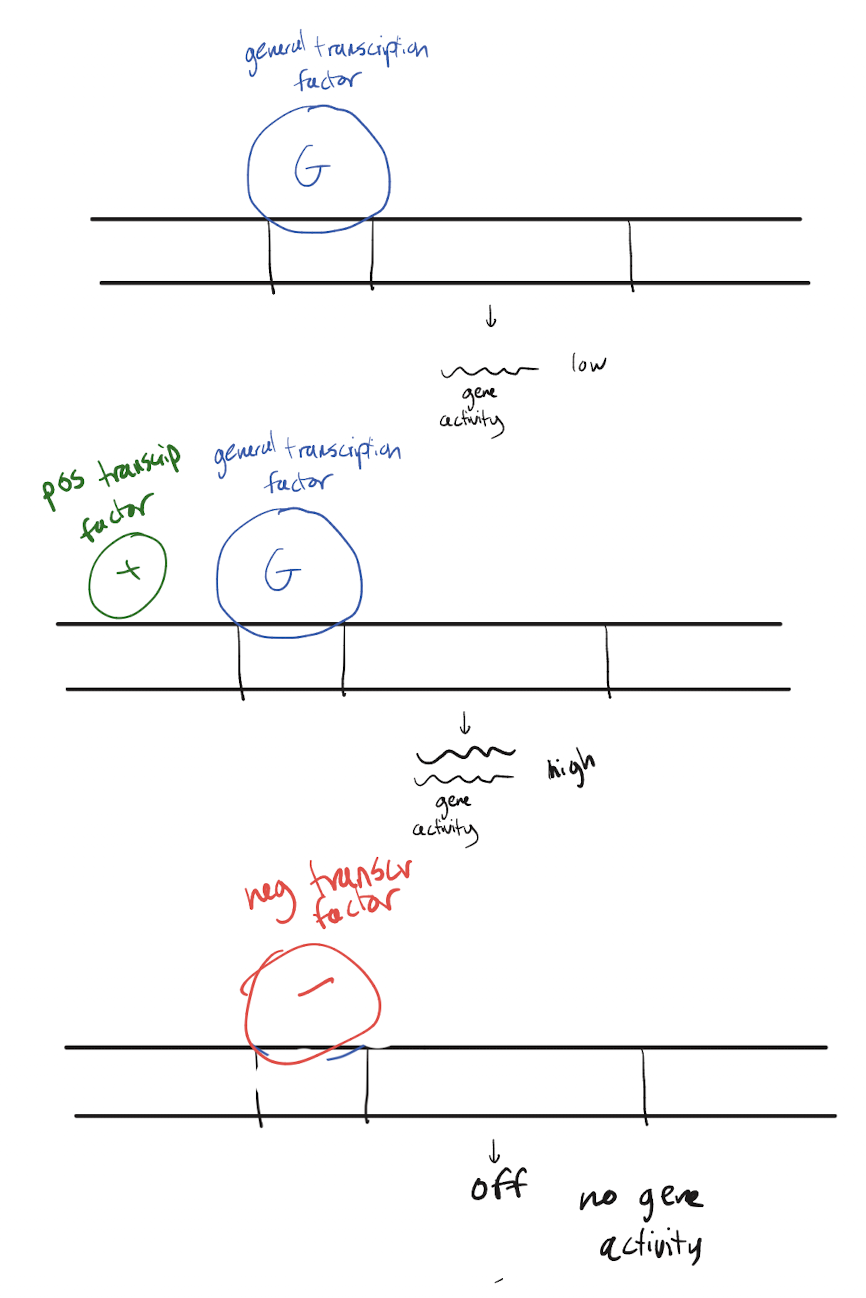
Explain how transcription factors (general, positive, negative) allow for different levels of gene expression.
General TFs allow for the baseline amount of gene activity (low but normal). A GTF along with a positive TF will recruit more RNA polymerase than typical, which increases the amount of gene activity. A negative transcription factor will knock off a general transcription factor and thus turn off transcription entirely, causing no gene activity
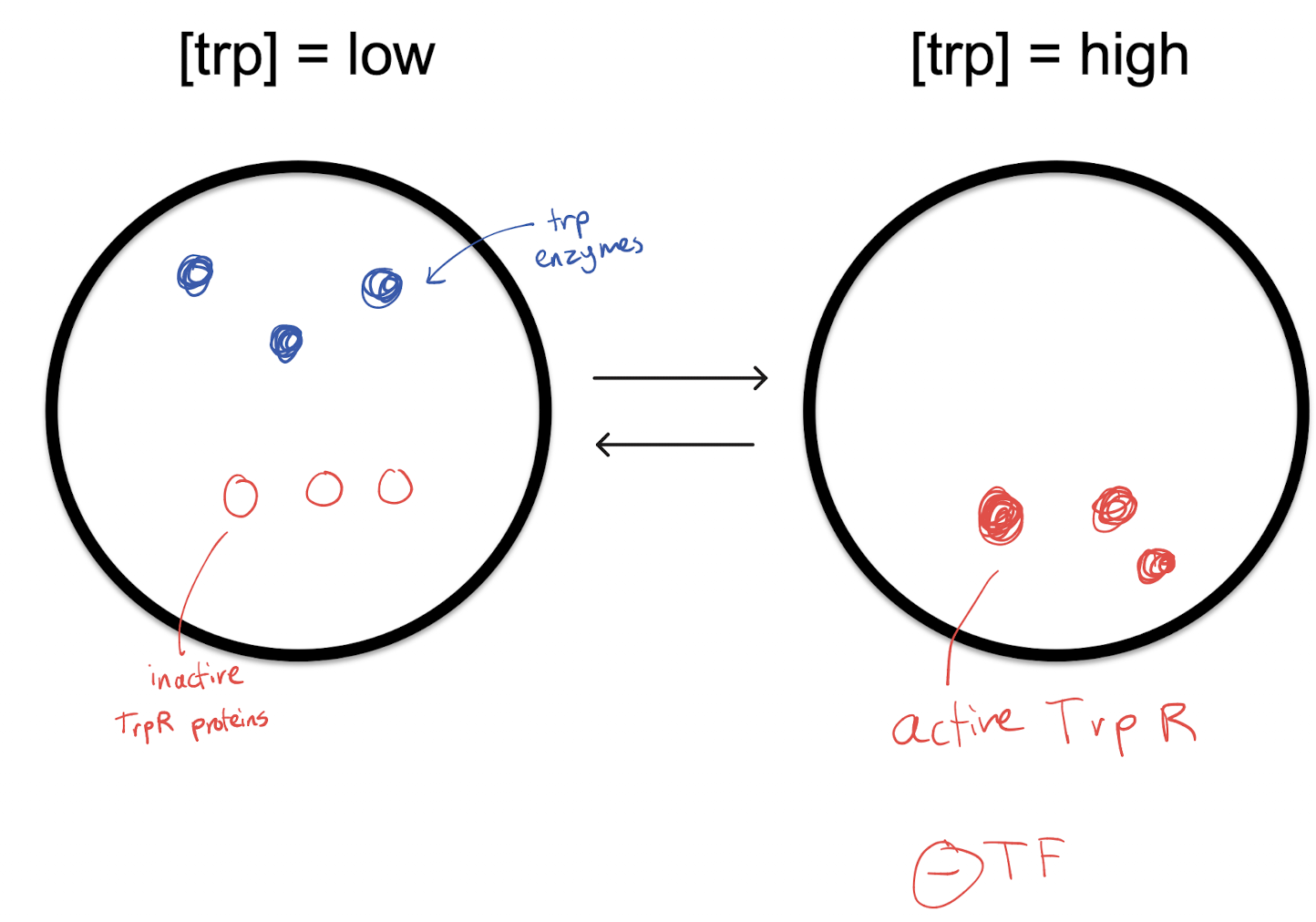
Explain how the Trp operon is regulated through a repressor.
When high levels of tryptophan are present, tryptophan will be able to find a trp repressor protein and bind to it to activate it. The active trp repressor will then temporarily bind to DNA with hydrogen bonds, turning off the operon, stopping mRNA translation and thus trp enzyme creation.
When there is a low concentration of tryptophan, tryptophan will come off of the trp repressors (to be used), deactivating them and causing them to break their temporary bonds with the operon. The trp operon will be active with nothing to repress it, allowing the creation of trp enzymes that will make tryptophan.
How would a mutation in the promoter of a gene affect RNA and protein synthesis?
Mutation would make it harder for RNA polymerase to find and bind with the promoter, decreasing the amount of RNA made and lessening protein synthesis
From silent, missense, nonsense, frameshift, and loss mutations, which proteins would end up functional and which would not? Explain.
Silent: functional. no change in actual protein
Missense: functional or non functional. If wrong amino acid is not integral to protein structure it can still function, but if wrong protein harms or replaces a necessary part it will be nonfunctional
Nonsense: most likely nonfunctional. Missing an entire part of the protein will cause it to lose many parts needed to function
Frameshift: most likely nonfunctional. Entire random sequence of amino acids is likely too large to ignore
Loss: functional or non functional. If missing/added amino acid is not integral to protein structure it can still function, but if missing/added amino acid harms or replaces a necessary part it will be nonfunctional
How would a mutation in the UTR of a gene affect RNA and protein synthesis?
No consequences since the UTR is not translated into the final protein
Explain how a missense mutation that changes a polar amino acid to a nonpolar amino acid in an HBB protein can damage hemoglobins.
Instead of hydrophillic amino acids that are able to be spread evenly about the hemoglobin, the nonpolar amino acids will form chains inside the red blood cells, sticking together to avoid water. This causes deformed (“sickle shaped”) red blood cells that get stuck in capillaries and have a much shorter lifespan