Industrial Biotech 2
1/263
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
264 Terms
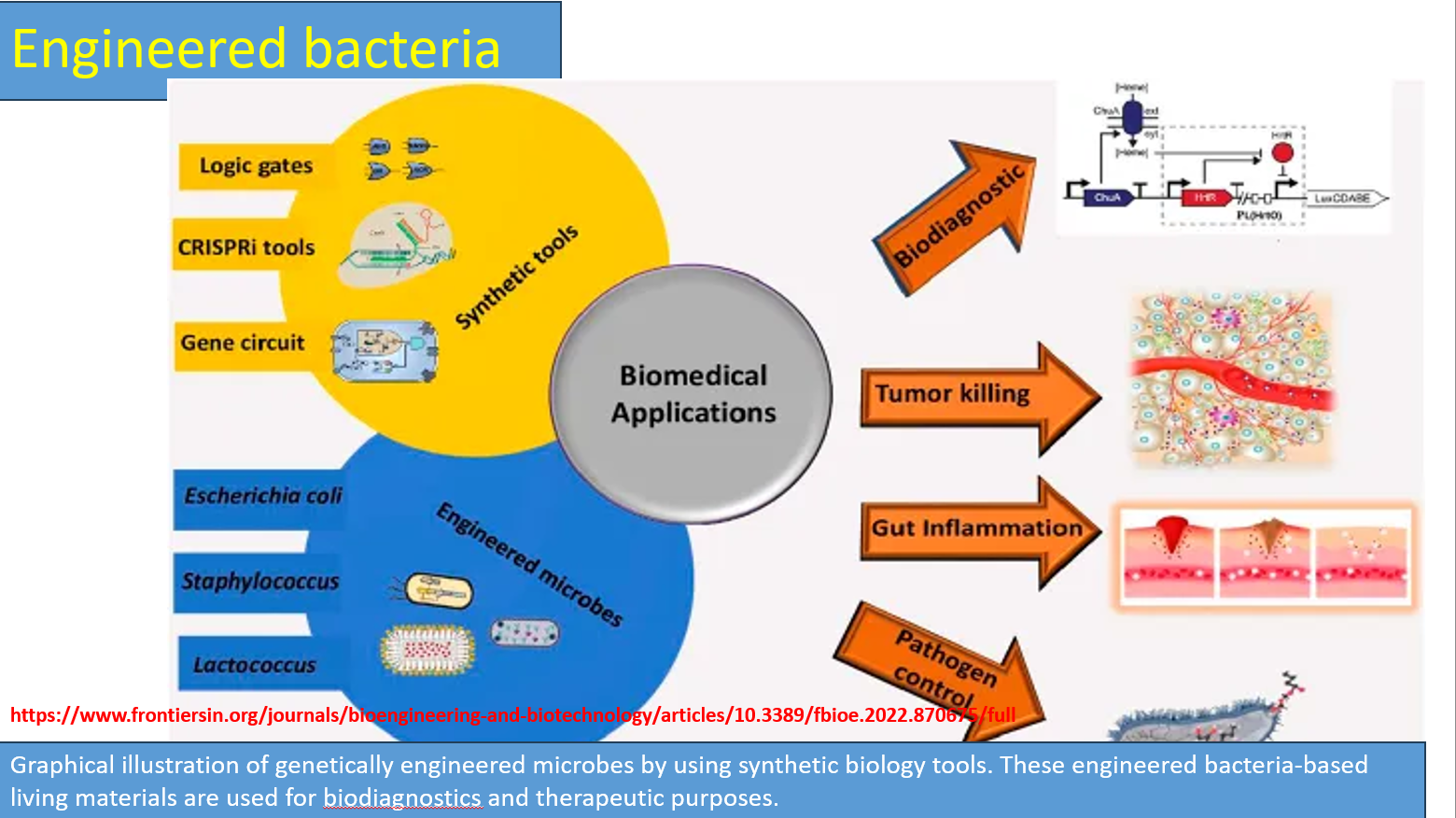
Engineered bacteria diagram: synthetic tools
Logic gates (biological computation)
Cells are engineered to behave like circuits (AND, OR, NOT gates)
Example:
AND gate → bacteria only respond when two signals are present
In tumors: only activate drug release when low oxygen + tumor biomarker
Mechanism:
Promoters + repressors arranged so transcription happens only under specific combinations
CRISPRi tools (gene silencing, not cutting)
Uses dCas9 (dead Cas9) → binds DNA but does not cut
Blocks RNA polymerase → turns genes OFF
Why useful:
Precise control of metabolic pathways
Reduce toxicity by shutting down harmful genes
Gene circuits
Designed DNA networks that control:
timing (delays, oscillations)
thresholds (only activate above certain signal)
Example:
quorum sensing circuit → bacteria activate only when population is high
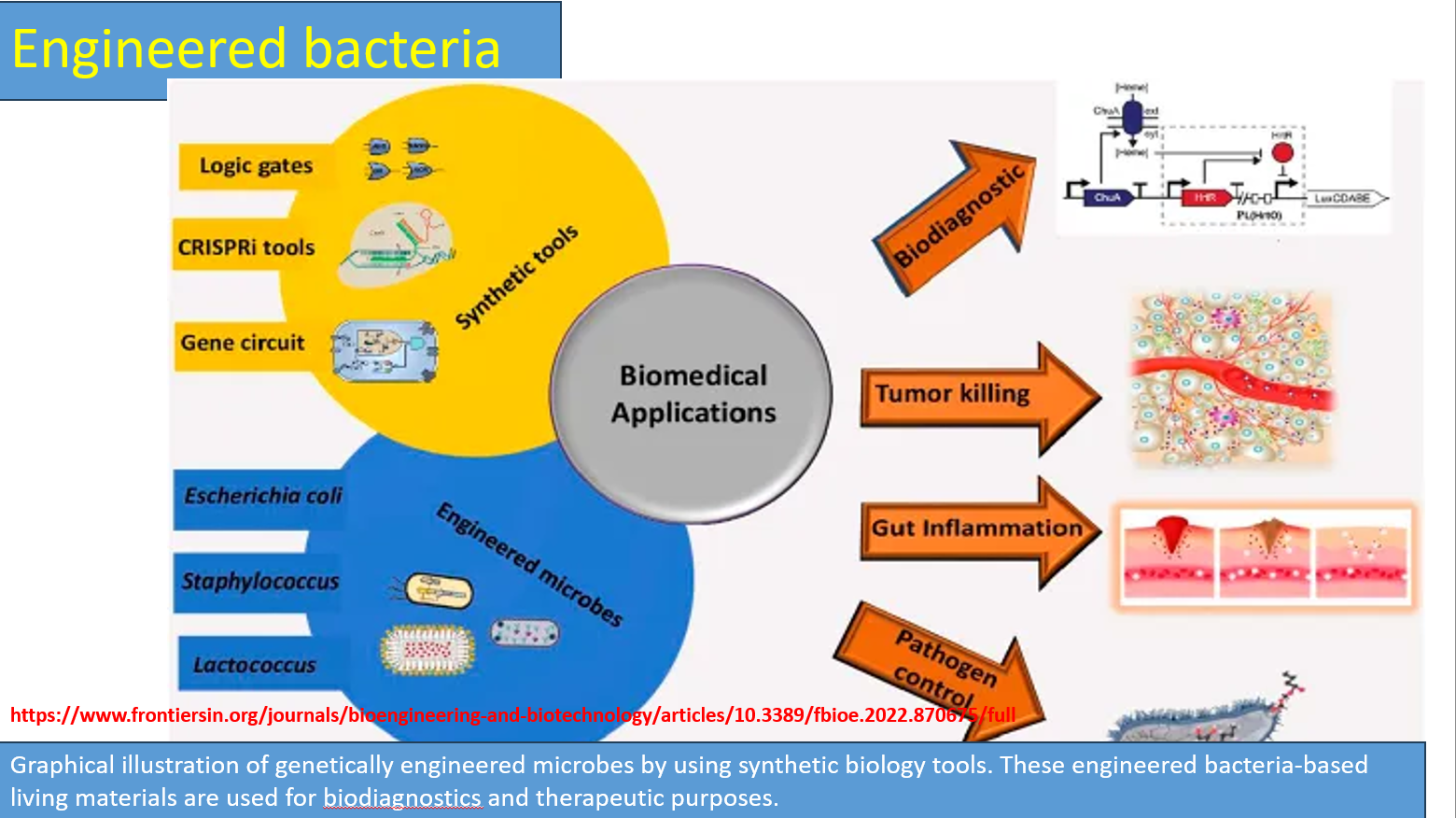
Engineered microbes
Common chassis organisms
E. coli
Easy to engineer
Used for sensing + drug production
Lactococcus / probiotics
Safe for human gut
Used in gut therapy (IBD, inflammation)
Staphylococcus (modified strains)
Used cautiously for skin or targeted delivery
Key idea:
You take a natural bacterium and insert synthetic circuits → it becomes a programmable living machine
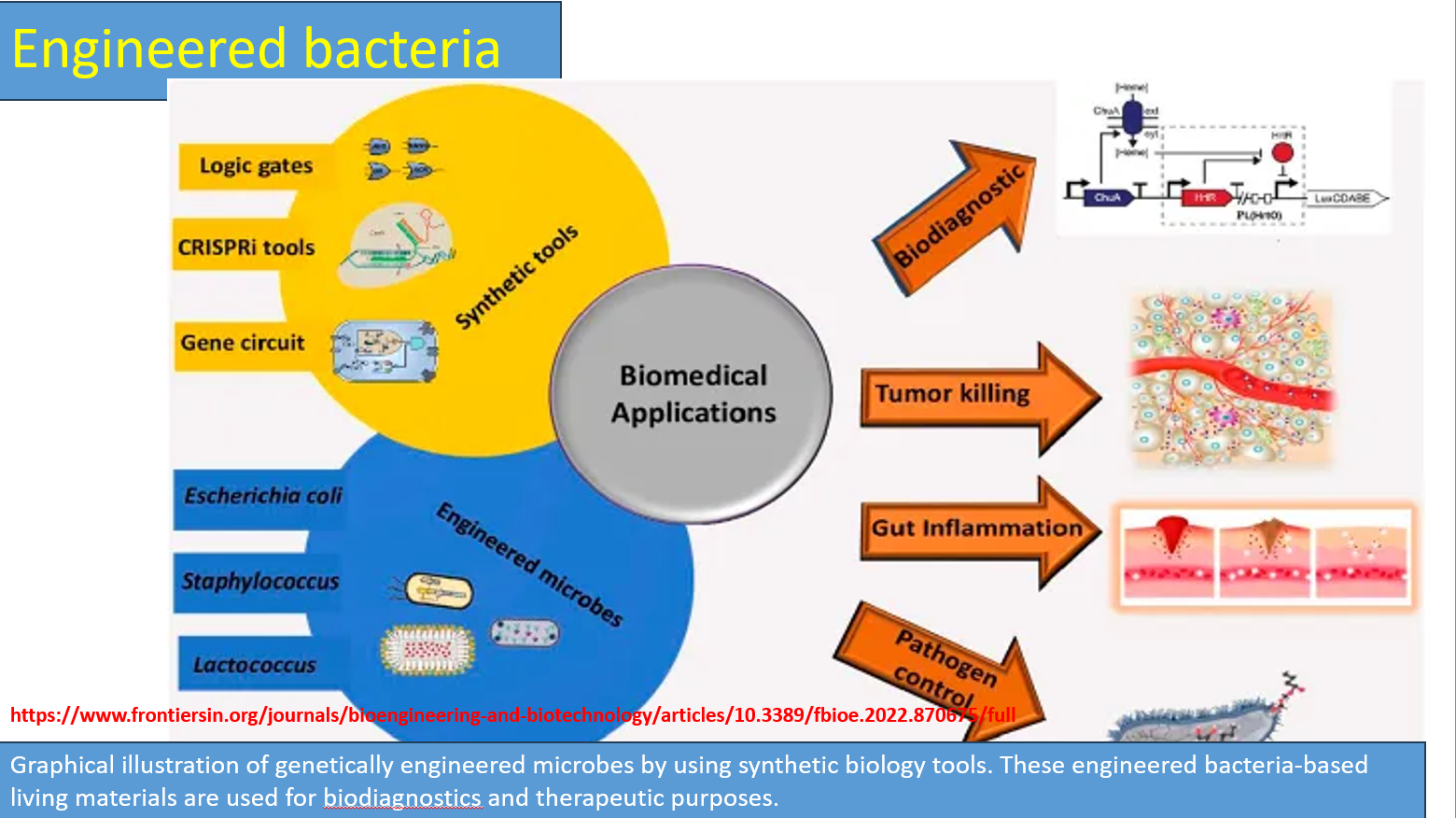
Biomedical applications (Biodiagnostics & Tumor killing)
Biodiagnostics
Engineered bacteria detect biomarkers and produce signals:
fluorescence
color change
electrical signal
Example:
detect gut bleeding → bacteria produce visible pigment in stool
Tumor killing
Tumors have unique environments:
low oxygen (hypoxia)
abnormal nutrients
Bacteria are engineered to:
accumulate in tumors
release:
toxins
immune activators
Advantage:
high specificity vs chemotherapy
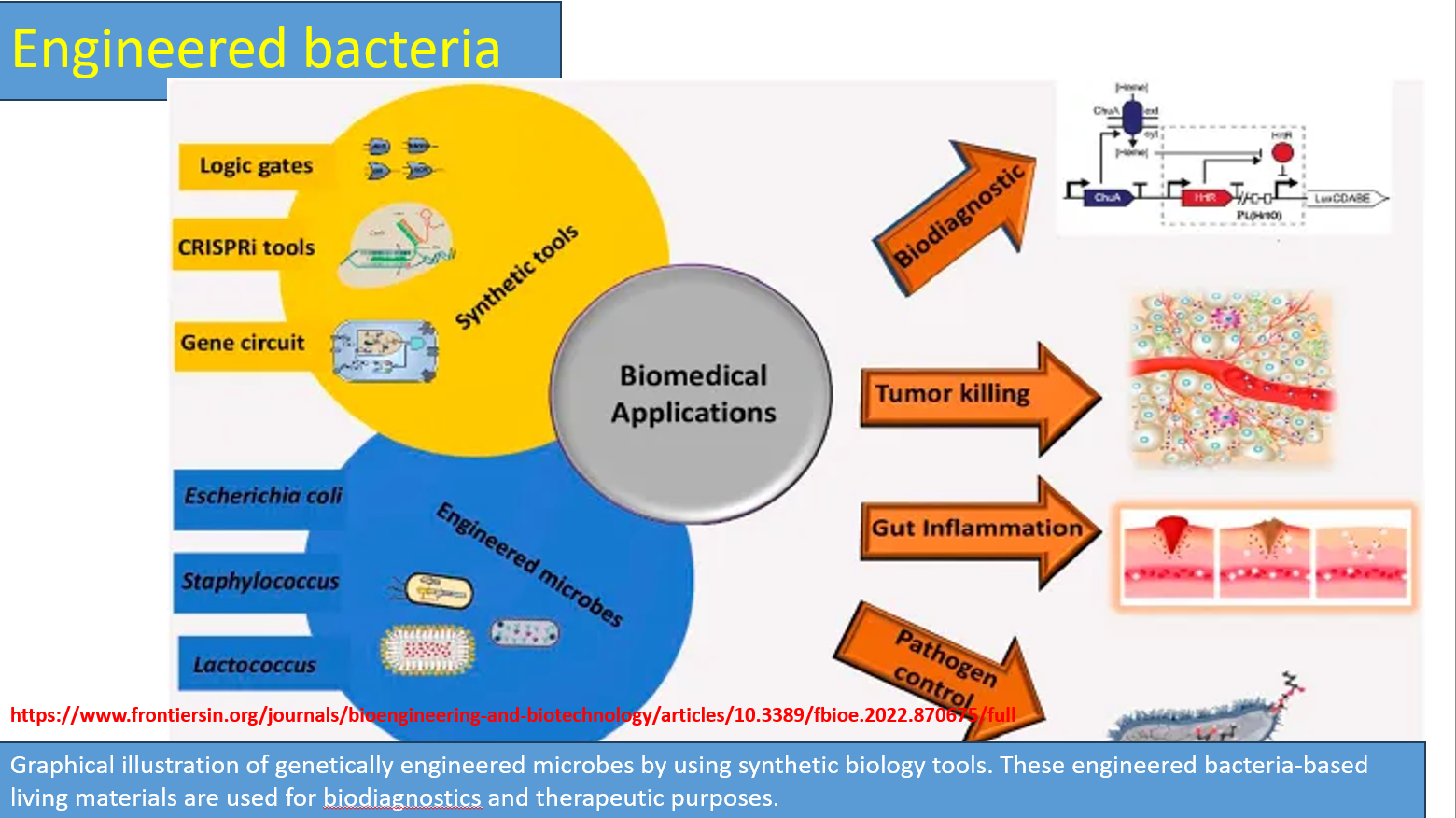
Biomedical applications (Gut inflammation, pathogen control)
Gut inflammation (IBD, Crohn’s)
Probiotic bacteria engineered to:
secrete anti-inflammatory cytokines (e.g., IL-10)
sense inflammation markers and respond dynamically
Acts as a living drug factory inside the gut
Pathogen control
Bacteria detect harmful microbes and:
release antimicrobials
outcompete pathogens
Example:
engineered microbiome to prevent infections
Traditional fermentation
What’s actually happening
Microbes (yeast, bacteria, fungi) metabolize sugars → produce:
lactic acid (yogurt, cheese)
ethanol + CO₂ (bread, alcohol)
Mostly anaerobic metabolism
Key insight (not in slide)
The goal is food transformation, not molecule isolation
Microbes remain in the final product → they shape:
flavor (organic acids, esters)
texture (proteolysis, gas formation)
Limitation
Low control → variability in taste, yield, contamination risk
Biomass fermentation
What’s actually happening
You grow microbes as the product itself
Cells are harvested as:
protein-rich biomass (single-cell protein)
Example mechanism
Fast-growing fungi/bacteria:
convert cheap carbon (glucose, waste) → cell mass
No need to extract specific molecules
Key insight
This is biomass accumulation optimization
maximize growth rate (μ)
maximize yield (Yx/s)
Why important
Efficient protein production:
much lower land + water than livestock
Limitation
Product is less customizable
Texture/flavor need processing
Precision fermentation
What’s actually happening
Microbes are genetically engineered to produce one specific molecule
Example:
insulin
casein (milk protein without cows)
heme (plant-based meat flavor)
Mechanism (important)
Insert gene → microbe expresses enzyme/pathway → produces target compound
Then:
cells are removed
product is purified (high purity step)
Key insight
This is cell as a factory, not food
You control:
metabolic flux
pathway efficiency
product specificity
Precision fermentation: upstream target selection
Target selection
Decide what molecule you want
protein (e.g., casein)
pigment (e.g., beta-carotene)
vitamin (e.g., B12)
This determines entire pathway design
Complex molecules → require multi-enzyme pathways
Precision fermentation: upstream host selection
Host selection & strain development
Choose organism:
bacteria (fast growth)
yeast (better for complex proteins)
fungi (high secretion)
Then engineer:
insert genes
optimize promoters
remove competing pathways
You are optimizing metabolic flux toward your product
Otherwise cells waste energy on growth instead of production
Precision fermentation: feedstock selection upstream
Feedstock selection
Carbon source:
glucose, molasses, agricultural waste
Why it matters:
Feedstock cost = major economic factor
Using waste streams → sustainability advantage
precision fermentation: Fermentation Fermentor design
Fermentor design
Types:
submerged (liquid, most common)
solid-state (for fungi)
membrane systems (retain cells, improve yield)
Engineering goal:
Maximize:
cell density
product formation rate
precision fermentation: Fermentation process optimization
Process optimization
Control key variables:
pH
→ affects enzyme activityTemperature
→ affects growth rate + protein foldingOxygen
→ determines metabolism:aerobic → biomass + ATP
anaerobic → fermentation products
Salt / nutrients
→ stress conditions can increase product yield
Key insight:
Small changes here can double or crash production
precision fermentation: Downstream
Pre-treatment
Break cells (if product is intracellular)
Remove debris
Separation
Centrifugation / filtration
Separate cells from liquid
Concentration
Reduce volume
Increase product concentration
Methods: evaporation, ultrafiltration
Purification
Chromatography, precipitation
Remove impurities → achieve high purity
Important:
This is often the most expensive step
Product formulation
Convert into usable form:
powder
liquid
food ingredient
Fermentationhost selection pros: bacteria
Prokaryotes – easier to manage
→ No nucleus, simple genome → easier gene insertion and control
→ Faster troubleshooting and strain developmentFaster generation time
→ Very short doubling time (~20–30 min)
→ Rapid biomass accumulation → high productivity (g/L/h)Many molecular tools available
→ Strong promoters, plasmids, CRISPR tools
→ Enables high-yield expression systemsHave plasmids
→ Independent DNA vectors → high gene copy number
→ Can overexpress target proteins massivelyLow-cost growth (extra)
→ Grow on simple media (glucose, salts)
→ Industrial scalability is cheaper than eukaryotes
Fermentationhost selection cons: bacteria
Genetic drift – higher mutation rates
→ Fast division = more replication errors
→ Loss of plasmids or mutations → reduced yield over time
→ Requires selection pressure (e.g., antibiotics)Bad public perception
→ Association with pathogens
→ Regulatory + consumer resistance (especially in food)Lack of post-translational modifications (extra)
→ Cannot perform proper glycosylation
→ Complex proteins may be inactive or misfoldedInclusion body formation (extra)
→ Overexpressed proteins aggregate → require refolding steps
Fermentationhost selection pros: yeast
Eukaryotes – more robust than bacteria
→ Can tolerate wider:pH
temperature
osmotic stress
→ Better suited for industrial conditions
High accumulation of compounds
→ Efficient metabolic pathways
→ Used for:vitamins
organic acids
bioethanol
Post-translational modifications (extra)
→ Protein folding + glycosylation
→ Suitable for therapeutic proteinsSecretion capability (extra)
→ Can export proteins → easier downstream processing
Fermentationhost selection cons: yeast
Slower generation time
→ Longer fermentation cycles
→ Lower throughput than bacteriaHigher nutrient requirements
→ More complex media → increased costNon-human glycosylation (extra)
→ Adds incorrect sugar groups
→ Can affect drug safety/function
Fermentation host selection pros: Mold
Eukaryotes – more robust
→ Similar stress tolerance advantages as yeastPowerhouse enzyme producers
→ Naturally secrete enzymes to digest environment
→ Very high yields of:amylases
proteases
cellulases
Extracellular enzymatic degradation
→ Break down complex substrates (e.g., plant biomass)
→ Can use cheap agricultural wasteHigh secretion capacity (extra)
→ Product released into medium → easier purification
→ degrade agricultural waste → cost advantage
Fermentation host selection cons: mold
Slower generation time
→ Slower biomass formationHigher nutrient demand
→ Requires more complex substratesMycotoxin production risk
→ Some species produce toxic compounds
→ Requires strict strain selection and controlNegative impact on rheology
→ Filamentous growth → thick, viscous broth
→ Poor mixing and oxygen transfer → reduced efficiencyScale-up challenges (extra)
→ Hard to maintain uniform growth in large reactors
Fermentation host selection pros: Microalgae
Eukaryotes – more robust than bacteria
→ Handle environmental stress betterAdaptive to existing fermentation infrastructure
→ Can be grown in systems similar to bacteria/fungiHeterotrophic growth
→ Use sugars instead of light → higher cell density
→ More efficient industrial productionWide range of unique molecules
→ Pigments (astaxanthin)
→ Lipids (omega-3 fatty acids)
→ AntioxidantsSustainability potential (extra)
→ Can use CO₂ + sunlight (autotrophic mode)
Fermentation host selection cons: Microalgae
Target extraction challenging
→ Tough cell walls (sometimes silica-based)
→ Requires mechanical/chemical disruption → high costAutotrophic growth limitations
→ Slow growth
→ Low biomass density
→ Light penetration limits scalingPhotobioreactor cost (extra)
→ Expensive infrastructure
Feedtosck selection: Conventional feedstock
Standardized conditions
→ Pure substrates (glucose, sucrose) → consistent composition
→ Predictable microbial growth and product yieldNo food safety issues
→ Clean inputs → low contamination risk
→ Easier regulatory approvalCostly to use
→ Refined sugars require processing → high input cost
→ Competes with human food supplyLess sustainable
→ Uses edible crops → land, water, fertilizer demand
→ Contributes to food vs industry competitionExample: refined sugar
Ideal for:
pharmaceuticals (need consistency)
Not ideal for:
large-scale sustainable food production
Feedtosck selection: alternative feedstock
Not standardized
→ Variable composition (different sugars, inhibitors)
→ Leads to inconsistent fermentation performanceProduction inconsistency
→ Microbes may grow differently batch-to-batch
→ Requires process optimizationRegulatory challenges (food safety)
→ Waste streams may contain:toxins
heavy metals
→ Needs strict purification and validation
More sustainable
→ Uses waste instead of food crops
→ Supports circular bioeconomyExample: food industry side streams
Types of alternative feedstocks: agricultural
Agricultural crops
High carbohydrate content (e.g., sugarcane)
Used for:
bioethanol
alternative proteins
Insight:
Still semi-conventional → not fully sustainable if primary crops are used
Types of alternative feedstocks: Lignocellulosic plant material
Leaves, stems, wood (cellulose + hemicellulose + lignin)
Key mechanism:
Must be pretreated:
lignin removed
cellulose → glucose
Used for:
mycoprotein
biofuels
Limitation (extra):
Pretreatment is expensive and energy-intensive
Types of alternative feedstocks: Food industry by-products
Example: fruit peels, whey, pulp
Advantages:
Cheap or free
Already partially processed
Use:
alternative proteins
fermentation substrates
Types of alternative feedstocks: Plastic waste
Converted via:
pyrolysis → small hydrocarbons
microbes convert → biomass/protein
Insight:
Still emerging technology
Not widely commercial yet
Types of alternative feedstocks: Organic food waste
Mixed waste (household, agriculture)
Use:
fermentation into:
biogas
microbial protein
Challenge:
Highly variable composition
Process optimization: validation of industrial performance step 1
LAB SCALE
What you actually do:
Compare multiple hosts (e.g., bacteria vs yeast)
Compare multiple feedstocks (glucose, waste streams, etc.)
Measure:
whether the product is made at all
approximate yield
Why this works well:
Everything is well mixed
Oxygen and nutrients are uniformly available
Heat is easily dissipated
Why it is limited:
These conditions are artificially ideal
Cells are not exposed to gradients or stress
A strain that looks “high-performing” here may fail later
Process optimization: validation of industrial performance step 2
LAB SCALE BIOREACTOR
You introduce active control systems:
pH control (acid/base addition)
dissolved oxygen control (aeration + agitation)
temperature regulation
You can monitor variables in real time
What you optimize:
Growth vs production tradeoff
(cells often prioritize growth over product unless tuned)Oxygen supply vs demand
Feeding strategy (batch vs fed-batch)
Why this step matters:
You begin to see process behavior, not just biology
The same organism can behave differently depending on:
oxygen availability
nutrient limitation
stress conditions
Process optimization: validation of industrial performance step
PILOT UNIT
Volume increases significantly (tens → hundreds → thousands of liters).
What fundamentally changes:
Mixing is no longer uniform
→ cells experience microenvironments
(some regions high oxygen, some low)Oxygen transfer becomes limiting
→ oxygen must diffuse through liquid
→ not all cells receive the same amountHeat removal becomes difficult
→ metabolic activity generates heat
→ large volumes retain heat longerShear forces increase
→ agitation needed for mixing can damage cells
Why processes fail here:
A strain optimized for uniform lab conditions may:
stop producing product
shift metabolism
die in low-oxygen zones
What you do at this stage:
Adjust:
agitation speed
aeration rate
reactor geometry
Validate whether the process is stable over time
Process optimization: validation of industrial performance step 4
INDUSTRIAL BIOREACTOR
The focus shifts from “does it work?” to:
Can it run consistently for long periods?
Is it economically viable?
Key constraints:
Even small inefficiencies become expensive
Variability between batches must be minimized
Contamination risk becomes critical
Important requirements:
Strain stability
→ no genetic drift over long runsConsistent feedstock quality
→ variability affects product outputReproducible process control
→ identical conditions across batchesEfficient downstream processing
→ purification cost often dominates total cost
At this scale, optimization is continuous:
small changes in oxygen transfer, feeding, or mixing can significantly affect:
yield
product quality
cost per unit
Precision fermentation: Dairy
Purpose: Produce animal-free milk proteins identical to cow-derived ones
Status: Most advanced and already scaled commercially
What’s produced: Casein and whey proteins with the same amino acid sequence → same functionality (foam, emulsification, texture)
Key advantage: Can replicate real dairy properties because functionality depends mainly on protein structure
Main challenge:
High purification cost (proteins must be isolated from fermentation broth)
Production cost still higher than conventional dairy
Precision fermentation: eggs
Purpose: Replicate functional properties of eggs (not just nutrition)
Status: Commercial products exist; strong growth since ~2021
What’s produced: Egg proteins like ovalbumin → responsible for foaming, binding, emulsifying
Key advantage: Enables plant-based foods to behave like real eggs (e.g., baking, texture)
Main challenges:
Host development → proteins must fold correctly to work
Scale-up → maintaining functionality at industrial volumes
Harder than dairy because eggs rely on multiple interacting properties, not just one protein
Precision fermentation: Fats / Oils
Purpose: Replace animal fats in plant-based foods to improve taste and mouthfeel
Status: Emerging; many companies still scaling
What’s produced: Tailored lipids (specific fatty acid profiles)
Key advantage: Can design fats with:
better nutrition (e.g., omega-3)
specific melting behavior → improves texture
Main challenges:
Controlling fatty acid composition precisely
Small changes in lipid profile → major changes in taste and texture
Scale-up still difficult for high yields
Precision fermentation: Meat / Seafood
Purpose: Produce components that mimic meat/fish
Status: Developing; not fully mature
What’s produced: Key molecules like:
heme proteins (flavor, color)
aroma compounds
Key advantage: Improves realism of plant-based meat alternatives
Main challenges:
Replicating:
texture (muscle structure)
juiciness
full flavor profile
Precision fermentation produces ingredients, not whole tissue
Competes with biomass fermentation approaches
Precision fermentation: Colours / Vitamins / Flavours
Purpose: Replace synthetic additives with natural, microbially produced ones
Status: Strong and growing application
What’s produced:
pigments (carotenoids)
vitamins (e.g., B12)
flavor compounds
Key advantage:
Cleaner label (natural origin)
Avoids chemical synthesis
Main challenges:
Stability issues:
sensitive to heat, pH, light
Maintaining consistency during processing and storage
Scale-up for bulk production
Precision fermentation: Other (e.g., infant formula)
Purpose: Produce specialized high-value compounds
Status: Emerging but commercially promising
What’s produced:
human milk oligosaccharides (HMOs)
specialty nutrients
Key advantage:
High-value products justify higher production cost
Enables formulations closer to human biology
Main challenges:
Product stability
Scale-up and consistent manufacturing
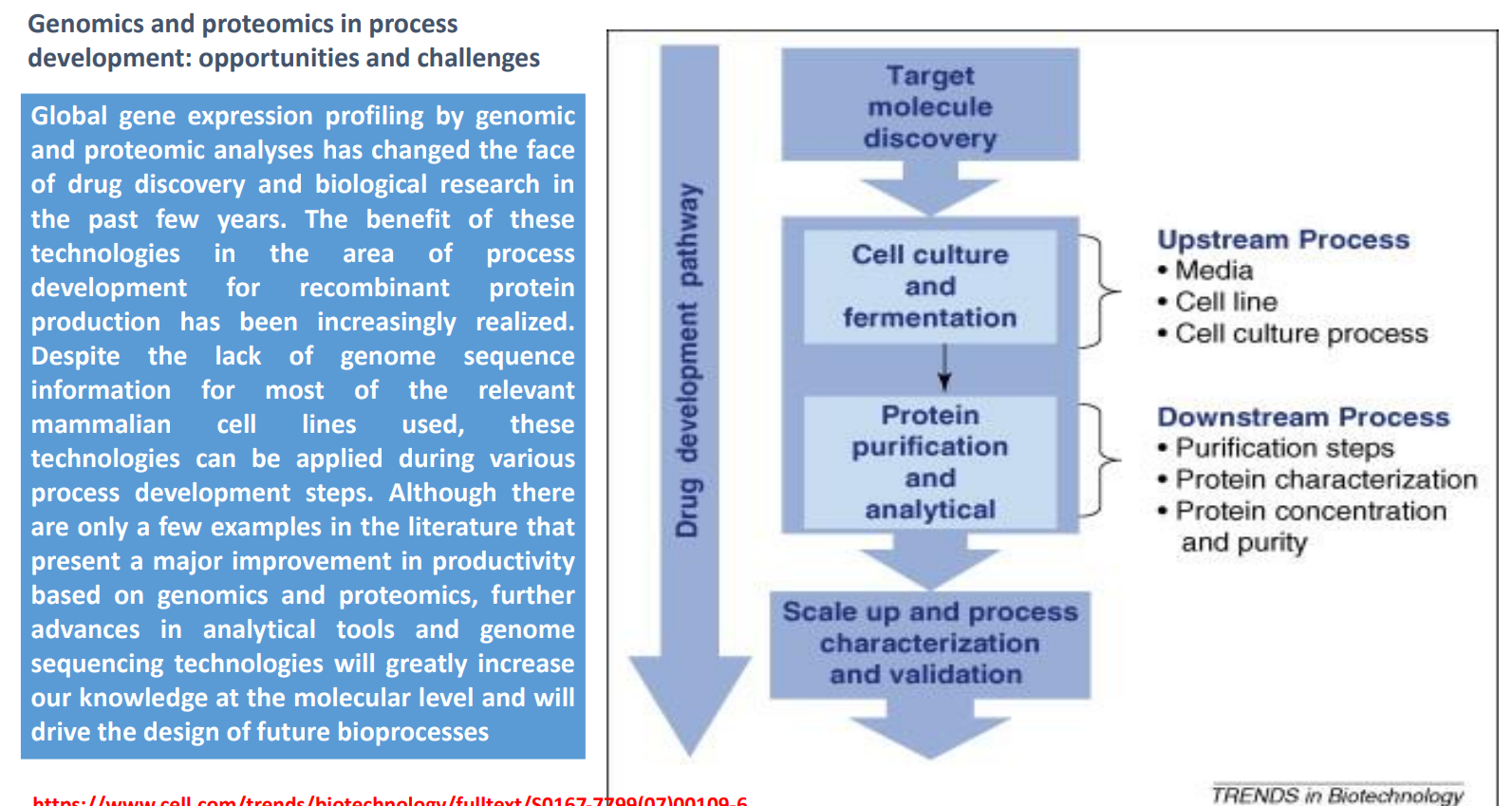
Genomics & Proteomics in Bioprocess Development
Use genomics (DNA-level) and proteomics (protein-level) data to understand how cells behave during production
Enables global gene expression profiling → which genes are ON/OFF under different conditions
Applied to recombinant protein production to improve yield and stability
Limitation: many industrial cell lines are not fully sequenced → incomplete data
Trend: better sequencing + analytical tools → more precise control of bioprocesses
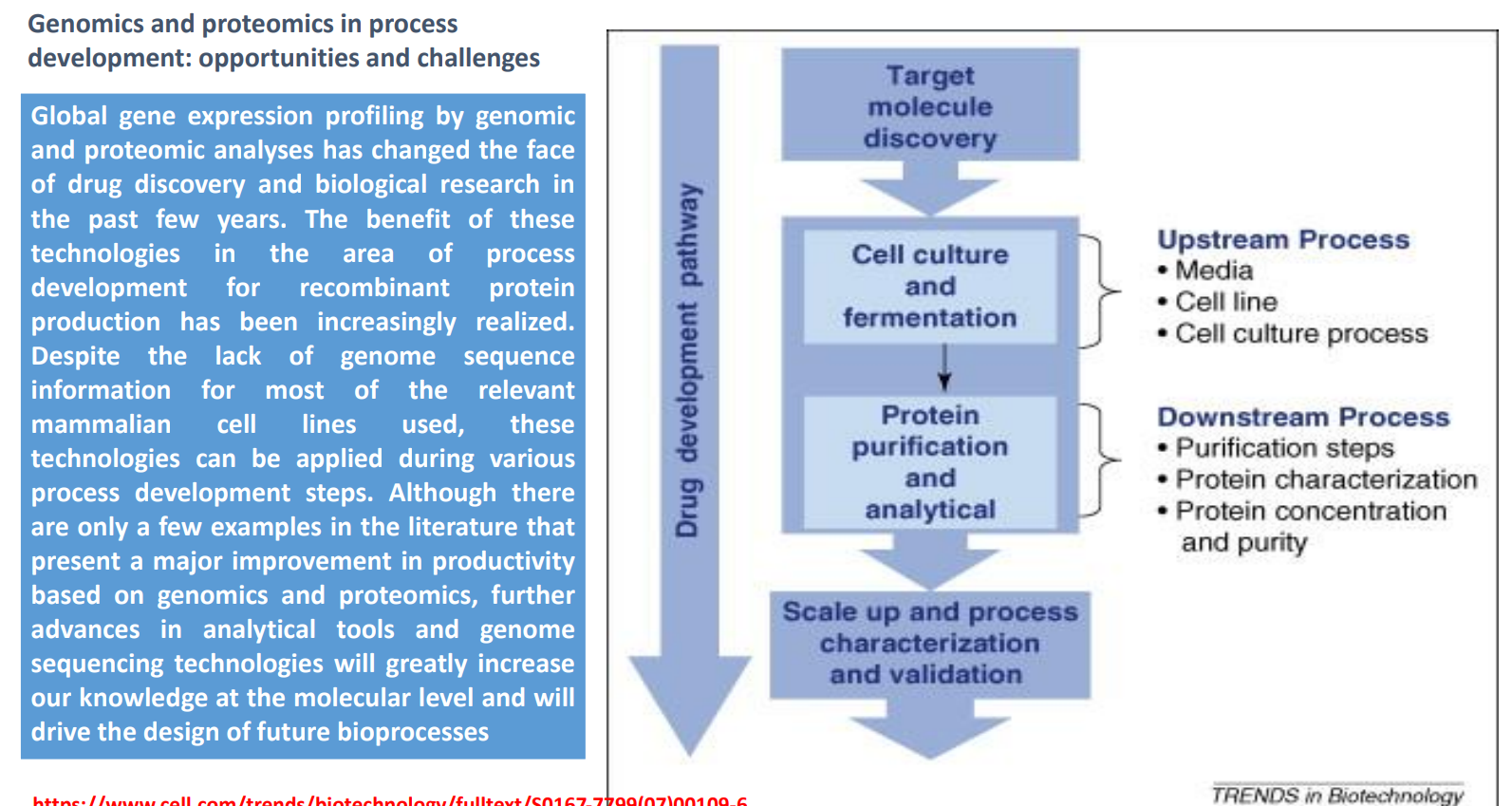
Genomics & Proteomics in Bioprocess Development: Target Molecule Discovery
Identify what protein or molecule to produce
Genomics helps:
find genes encoding useful proteins
Proteomics helps:
understand protein function and interactions
Key idea: choosing the right target early determines:
feasibility
downstream complexity
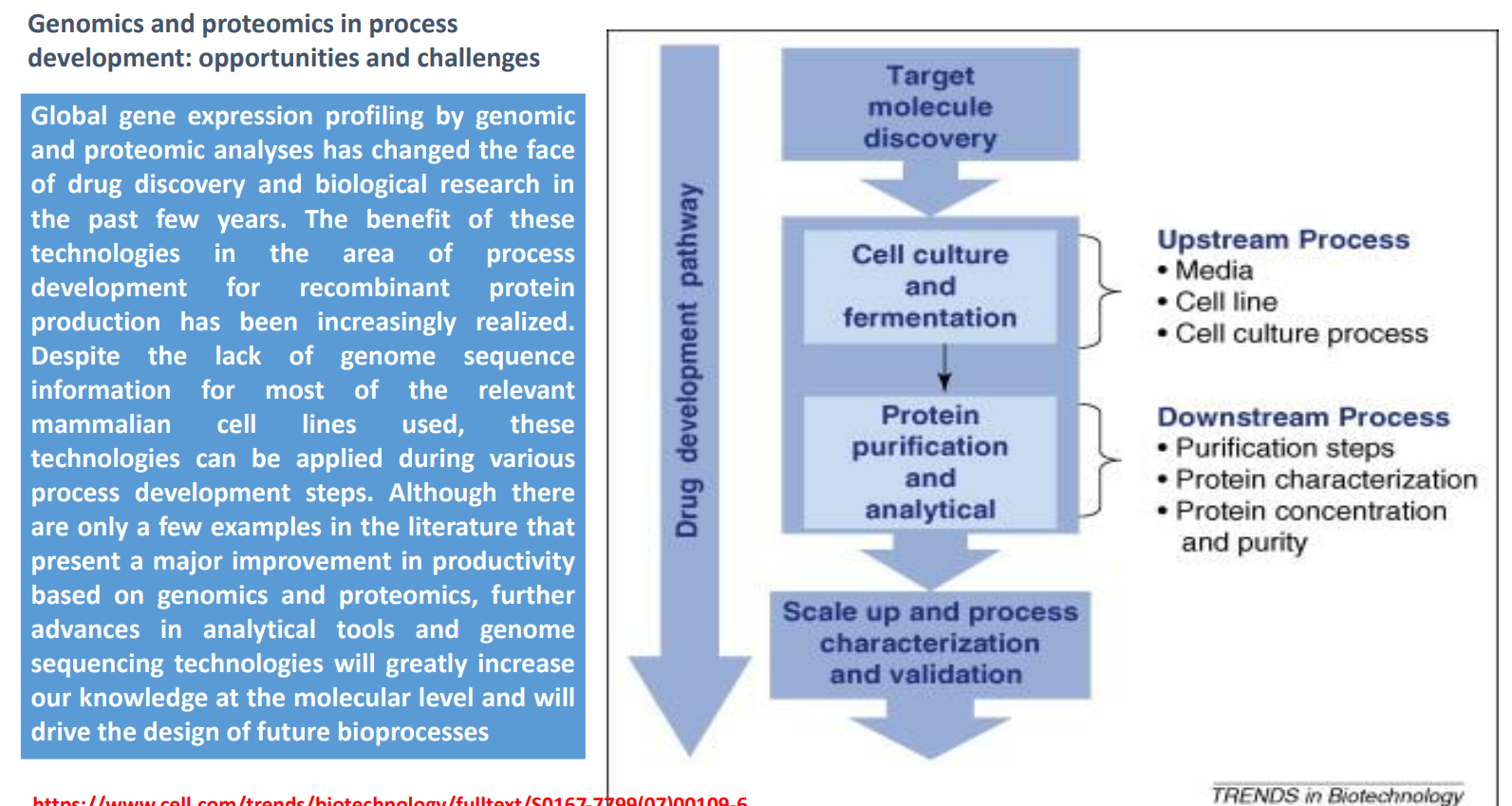
Genomics & Proteomics in Bioprocess Development: Cell Culture & Fermentation (Upstream Process)
Core components:
Media → nutrients affecting growth and expression
Cell line → determines production capability
Culture conditions → pH, oxygen, temperature
Role of genomics/proteomics:
Identify metabolic bottlenecks
Optimize gene expression levels
Detect stress responses that reduce yield
Practical impact:
Improve productivity
Reduce unwanted byproducts
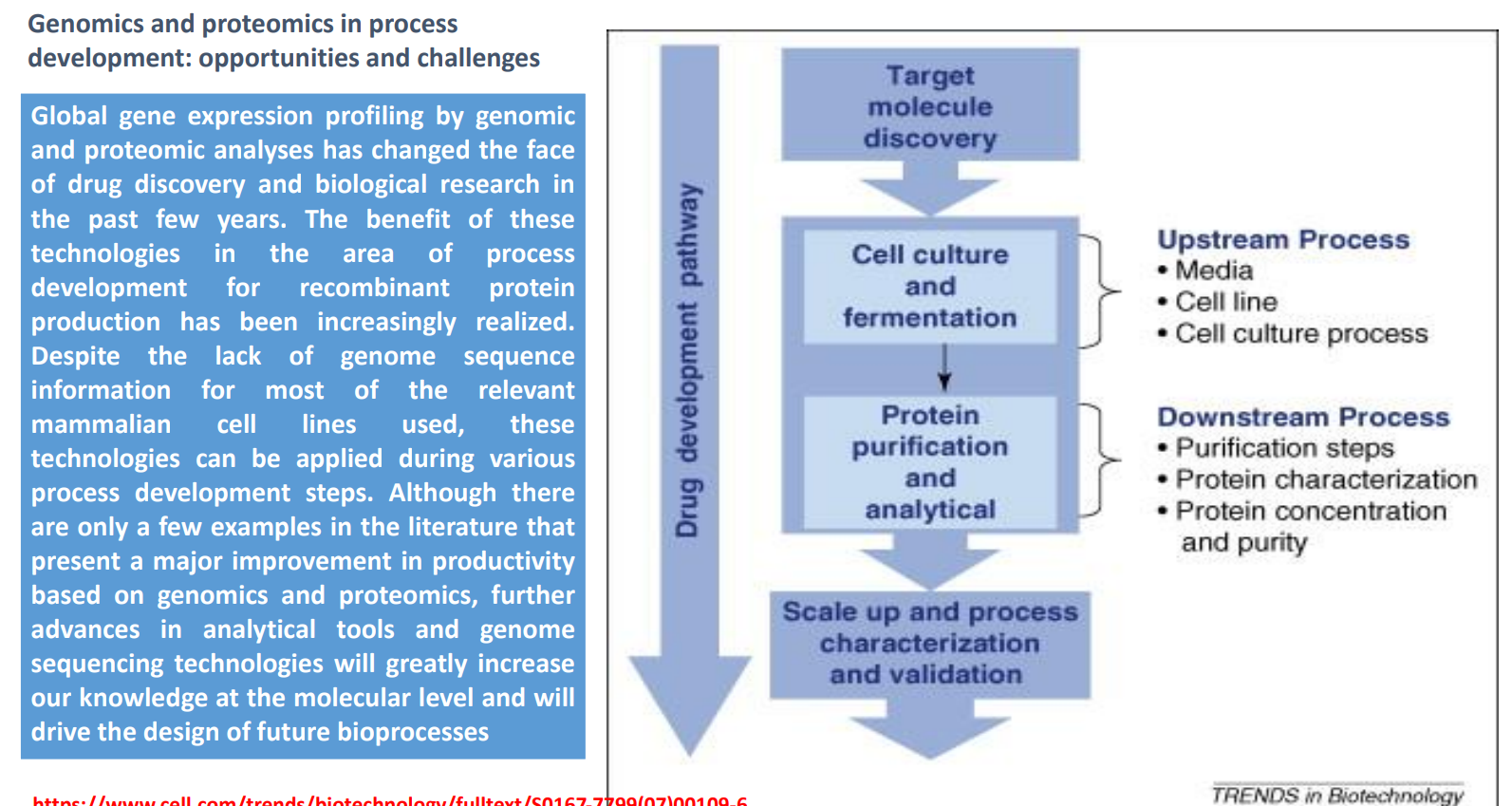
Protein Purification & Analytical (Downstream Process): Protein Purification & Analytical (Downstream Process)
Includes:
Purification steps (filtration, chromatography)
Protein characterization (structure, function)
Concentration & purity analysis
Role of proteomics:
Detect impurities or degraded proteins
Confirm correct folding and modifications
Ensure product quality and consistency
Key constraint:
Downstream processing is often the most expensive stage
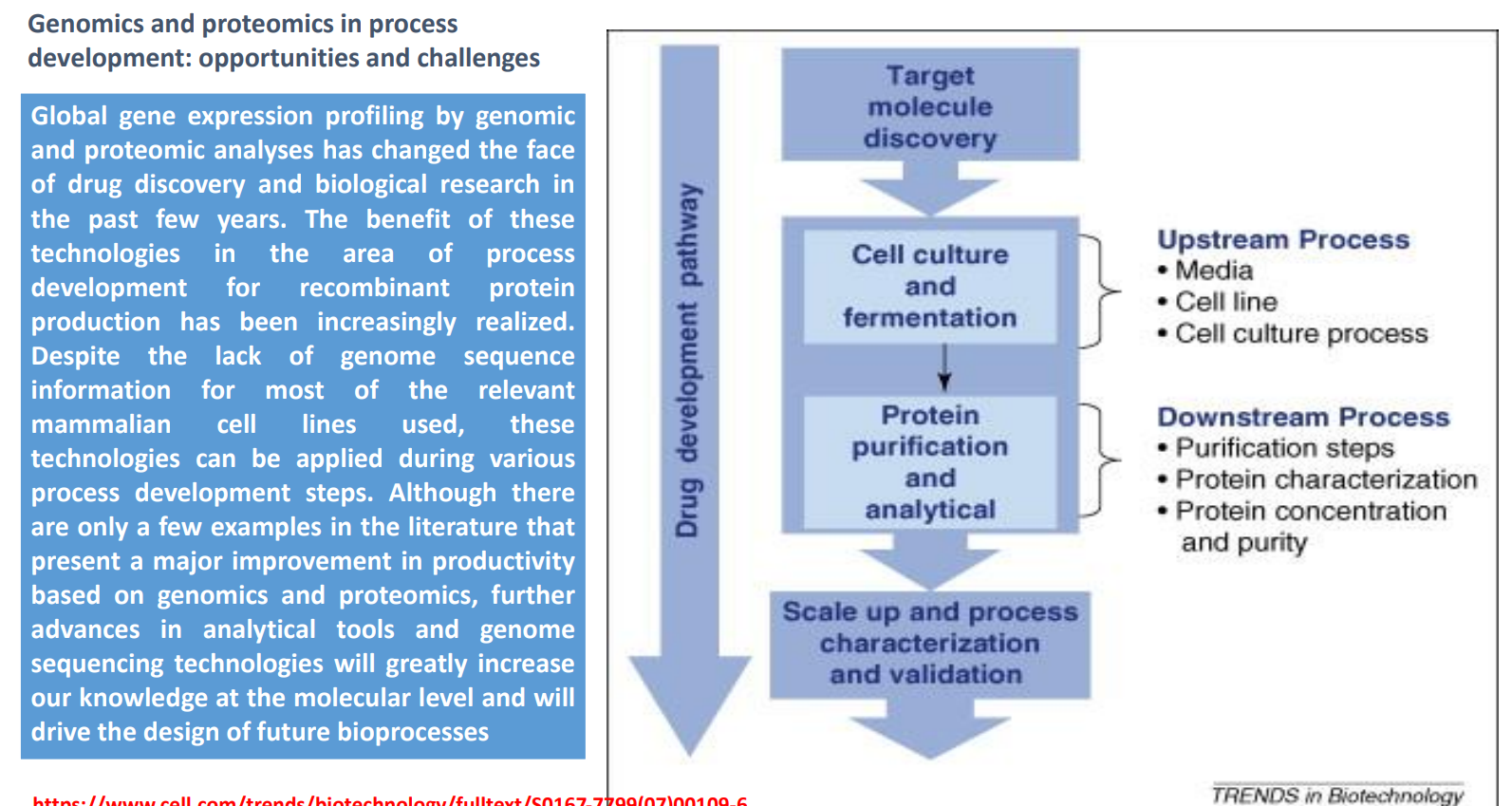
Genomics & Proteomics in Bioprocess Development:
Scale-Up, Characterization & Validation
Transition from lab → industrial production
Requires:
consistent product quality
reproducible performance
Genomics/proteomics role:
Monitor how gene expression changes at larger scale
Identify why yield drops during scale-up
Ensure stability of engineered strains
Critical issue:
Cells behave differently under:
oxygen limitation
nutrient gradients
stress conditions
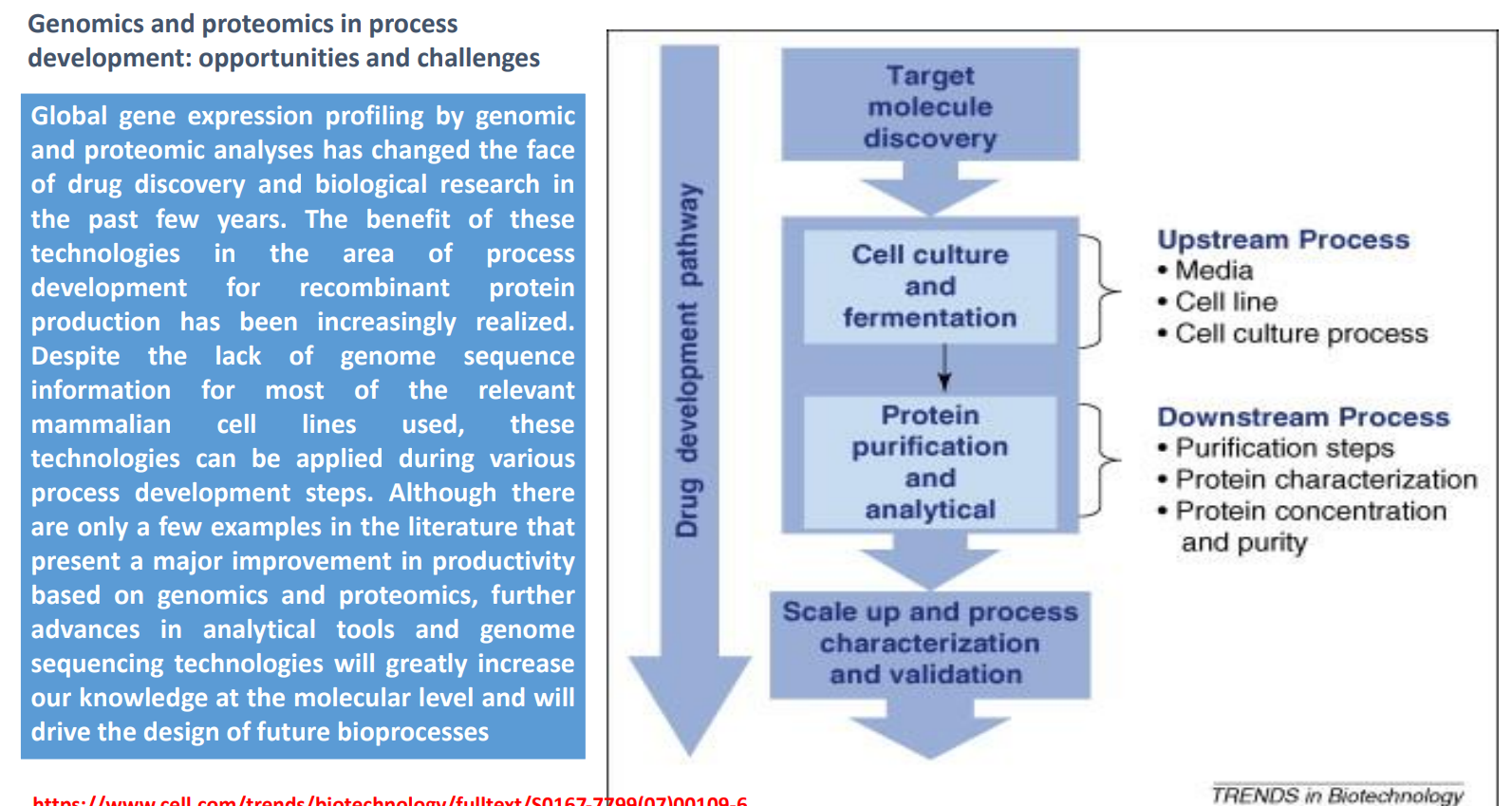
Upstream vs Downstream
Upstream
Media
Cell line
Culture process
→ Focus: maximize production
Downstream
Purification
Characterization
Concentration
→ Focus: ensure purity and functionality
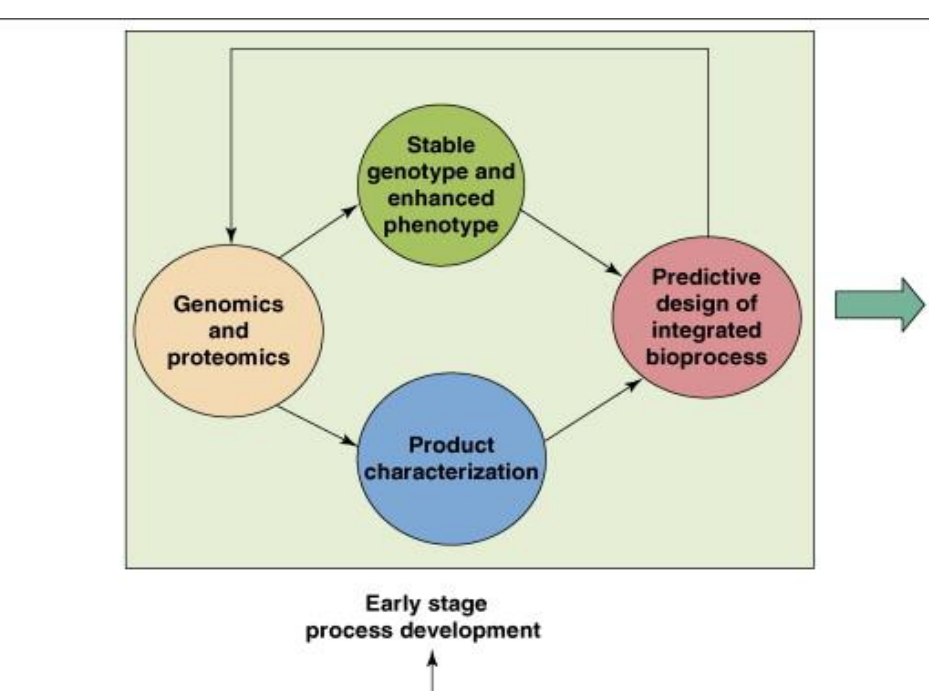
Early Stage of genomic and proteomic tools in process development: Data Integration and System Design step 1
Genomics and proteomics
→ Identify which genes and proteins drive production
→ Detect inefficiencies:low expression pathways
stress-induced protein changes
unwanted byproducts
Stable genotype and enhanced phenotype
→ Ensure the engineered organism:maintains genetic integrity over time
consistently produces high yields
→ Prevents loss of productivity during scale-up
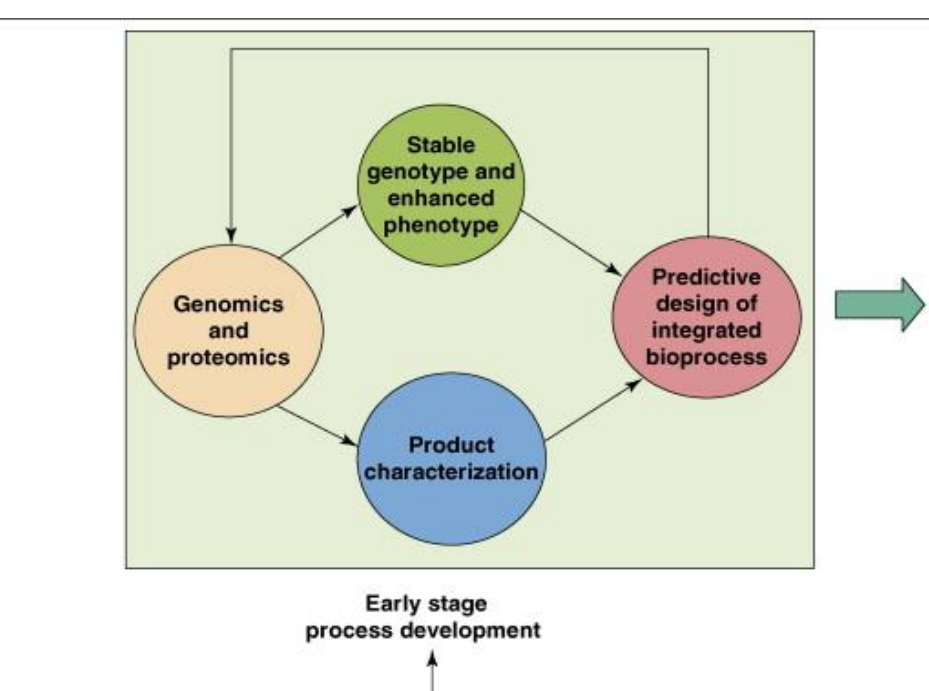
Early Stage of genomic and proteomic tools in process development: Data Integration and System Design step 2
Product characterization
→ Analyze:structure and functionality
impurities and degradation
→ Determines if the product meets quality requirements
Predictive bioprocess design
→ Use collected data to design:optimal strain
optimal growth conditions
compatible downstream strategy
→ Reduces reliance on trial-and-error
Stage of genomic and proteomic tools in process development: Feedback loop
Process is cyclical, not linear
Data from each stage feeds back into:
strain redesign
process adjustment
Outcome:
→ progressively improved performance and stability
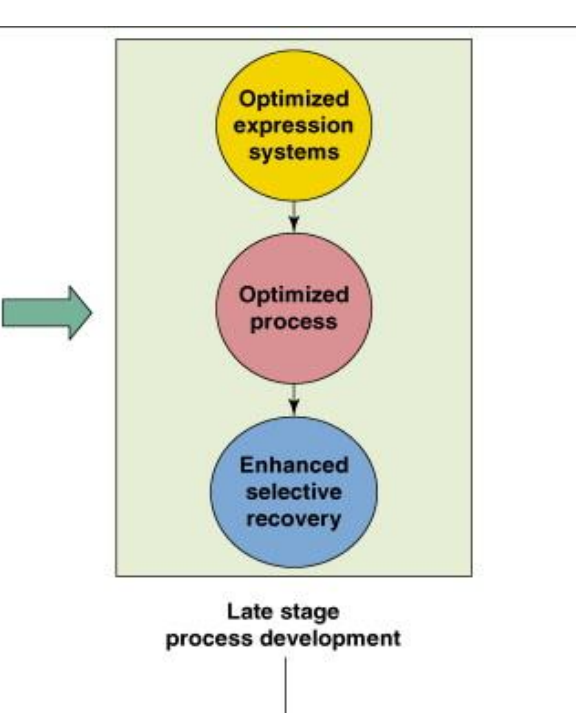
genomic and proteomic tools in process development: late stage
Optimized expression systems
→ Engineered host produces target efficiently with minimal wasteOptimized process
→ Conditions refined for:maximum yield
reproducibility at industrial scale
Enhanced selective recovery
→ Downstream tailored to:isolate product efficiently
improve purity and reduce costgenomic and proteomic tools in process development
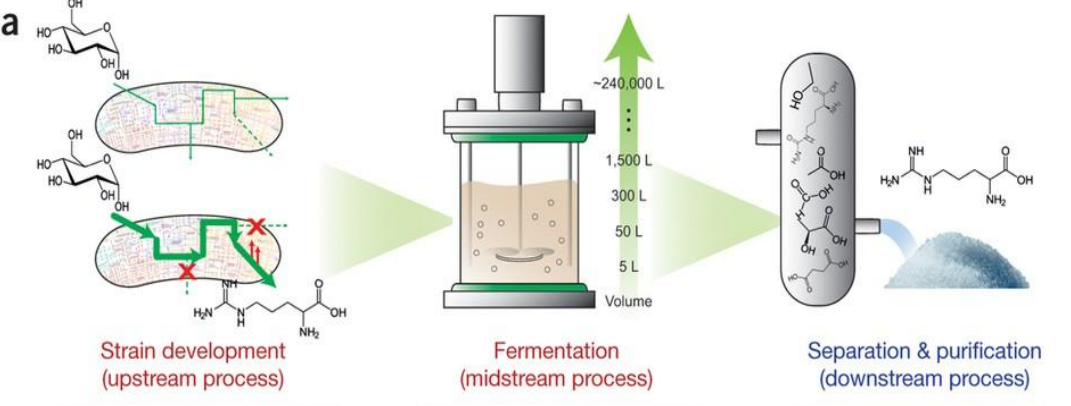
Systems strategies for developing industrial microbial strains: Strain Development (Upstream Process)
Microbial host selection
→ Choose organism based on:growth rate
tolerance to product
ability to express pathway
Biosynthetic pathway construction
→ Introduce or assemble pathways for non-native products
→ Requires inserting multiple genes and coordinating their expressionImprovement of self-tolerance
→ Many products are toxic to the cell
→ Cells are engineered to:resist product toxicity
maintain growth at high concentrations
Removal of negative regulation
→ Eliminate feedback inhibition or repressor systems
→ Prevents shutdown of production pathwaysFlux rerouting (cofactors & precursors)
→ Redirect metabolic intermediates toward target product
→ Balance cofactors (e.g., NADH/NADPH) for efficient synthesisOptimization of metabolic fluxes
→ Maximize carbon flow into product pathway
→ Minimize diversion into:biomass
byproducts
High-throughput systems tools
→ Use omics + screening to rapidly test many variants
→ Enables iterative improvement
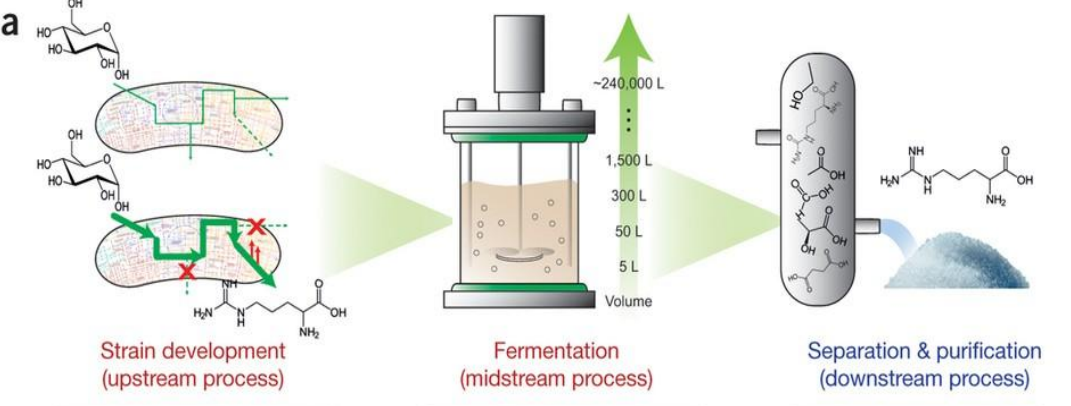
Systems strategies for developing industrial microbial strains: midstream process
Fermentation (Midstream Process)
Carbon source selection
→ Prefer:cheap
abundant
chemically defined substrates
Culture condition optimization
→ Control:pH
temperature
oxygen
→ Directly impacts enzyme activity and pathway efficiency
Feeding strategies
→ Batch vs fed-batch:fed-batch prevents substrate inhibition
maintains optimal growth phase
Performance evaluation
→ Measure:yield
productivity
byproduct formation
Scale-up of bioreactors
→ Transition from lab to industrial volume
→ Must maintain:oxygen transfer
mixing
temperature control
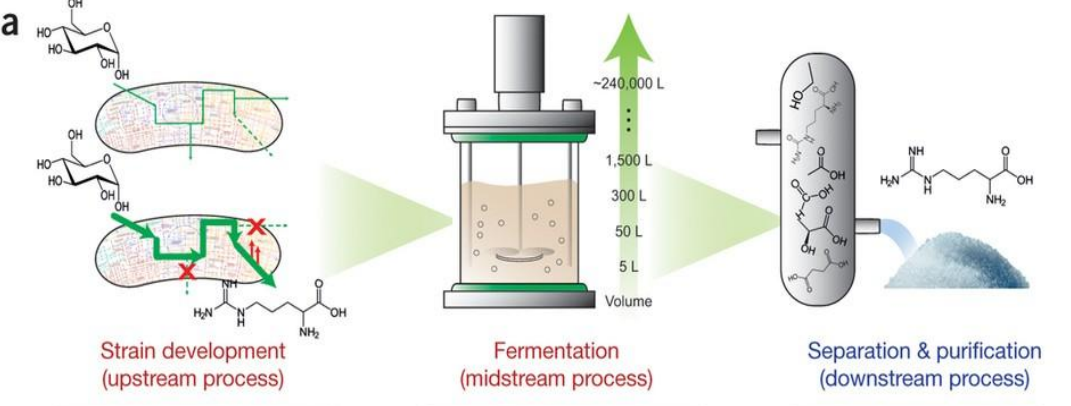
Systems strategies for developing industrial microbial strains: Separation & Purification (Downstream Process)
Separation & Purification (Downstream Process)
Use of defined conditions
→ Simplifies purification by reducing variabilityMinimization of byproducts
→ Easier downstream processing
→ Reduces purification costOptimization of recovery conditions
→ Example: low pH to improve separation
→ Tailored to product chemistry
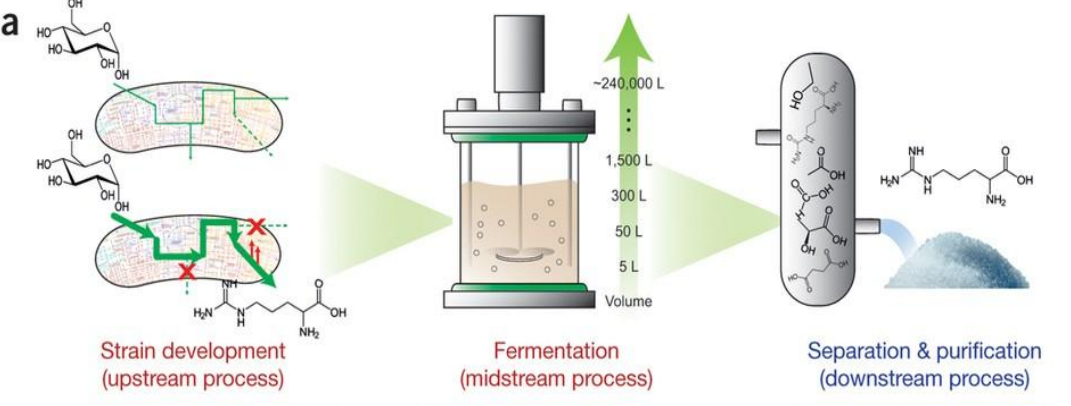
Systems strategies for developing industrial microbial strains: Iterative Design Loop
Arrows indicate continuous feedback between:
strain design
fermentation performance
downstream efficiency
Changes in one stage affect all others:
→ e.g., improving flux may increase byproducts → harder purification
Systems strategies for developing industrial microbial strains: core challenges
Core challenge: biological complexity
Cell behavior depends on interacting networks:
metabolism (carbon flow)
gene regulation (expression control)
signaling (response to environment)
Modifying one part affects others:
→ unpredictable outcomes
→ difficult to design optimal strains directly
Practical constraints
Time-intensive
→ multiple design–test cyclesCost-intensive
→ requires advanced tools, screening, validationLabor-intensive
→ experimental optimization at multiple levels
Systems strategies for developing industrial microbial strains: Demonstrated industrial outputs
Amino acids
L-valine, L-threonine, L-lysine, L-arginine
Bulk chemicals
1,4-butanediol
1,3-propanediol
butanol, isobutanol
succinic acid
Pharmaceuticals
artemisinin
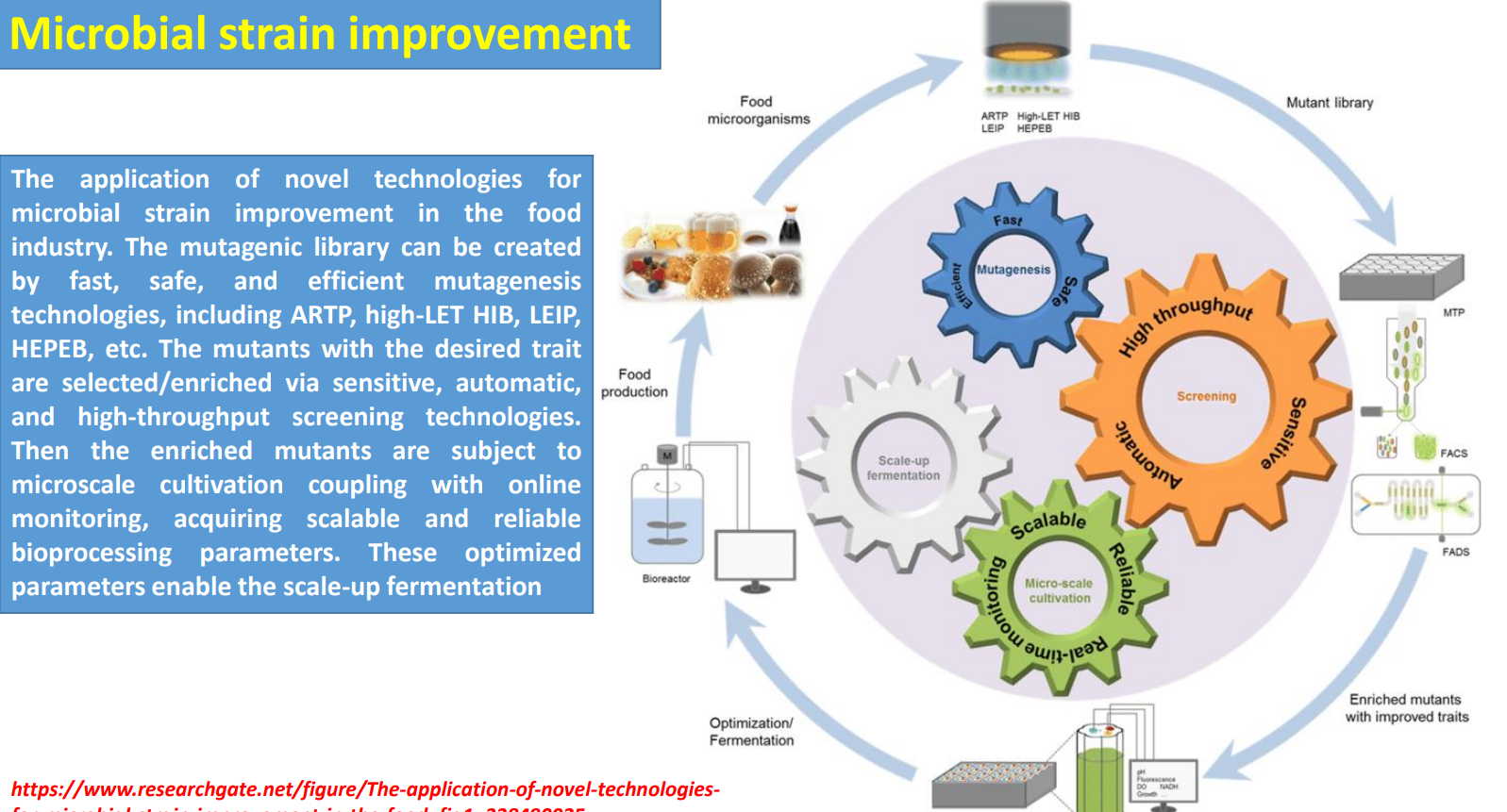
Microbial strain improvement: Mutagenic library → diversity generation under biological complexity
Industrial strains are improved by creating large mutant libraries using fast methods (ARTP, HIB, etc.) because cellular behavior is governed by interconnected metabolic, regulatory, and signaling networks that are hard to model directly.
Random mutagenesis introduces:
enzyme variants with higher activity
altered regulation (e.g., loss of feedback inhibition)
improved tolerance to toxic products
In practice, this often targets traits like:
higher flux through a pathway
resistance to product inhibition (e.g., ethanol, organic acids)
faster substrate uptake
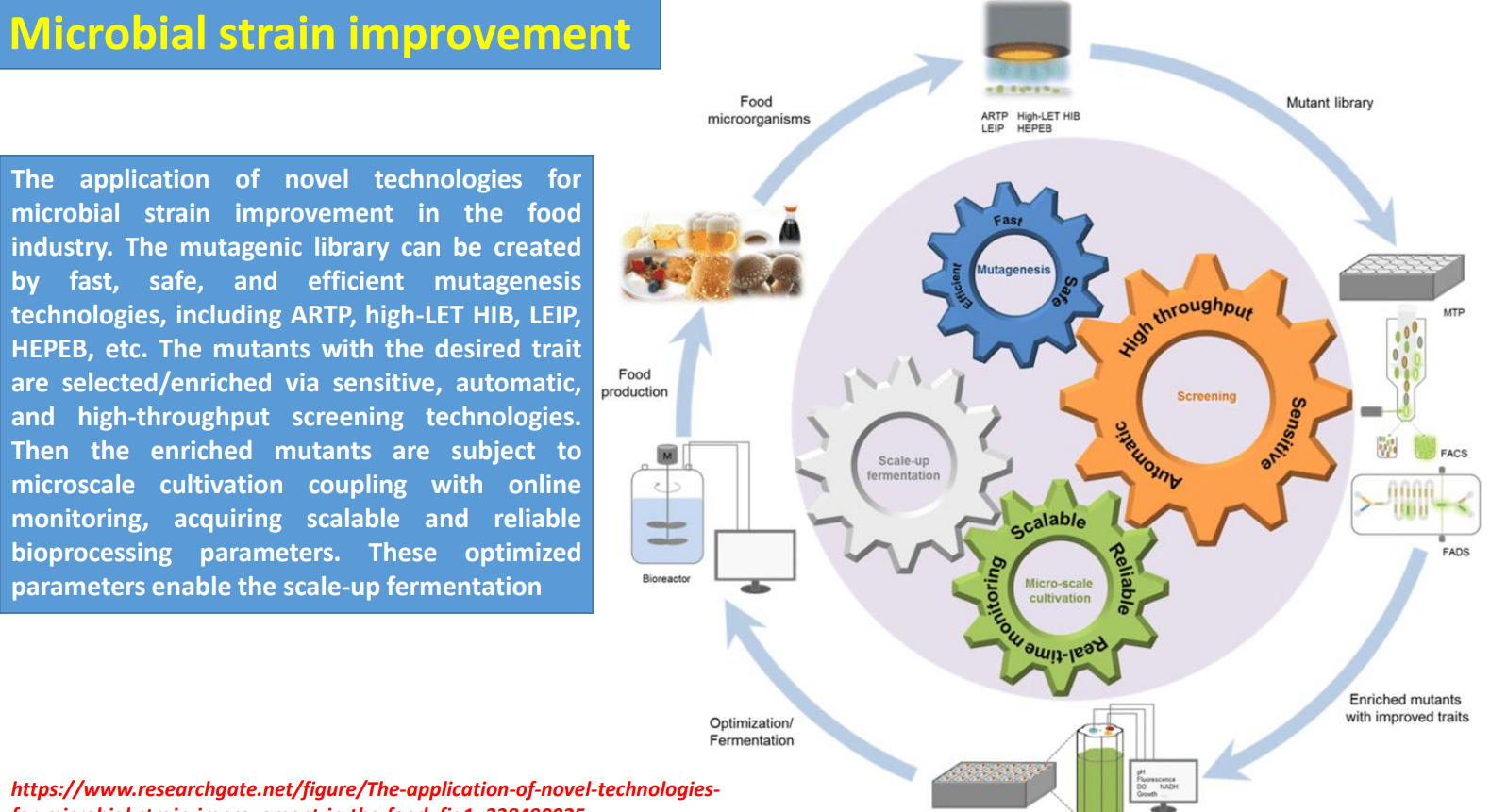
Microbial strain improvement: High-throughput screening → filtering functional phenotypes
The text emphasizes automatic, high-throughput selection, while the diagram shows tools like FACS/FADS—these are necessary because only a tiny fraction of mutants improve performance.
Screening is designed to link phenotype to measurable signal, for example:
fluorescence linked to product concentration
growth under stress (proxy for tolerance)
Critical constraint:
→ a strain that looks good in screening must also maintain:metabolic balance
energy efficiency (ATP usage)
otherwise it fails later during fermentation
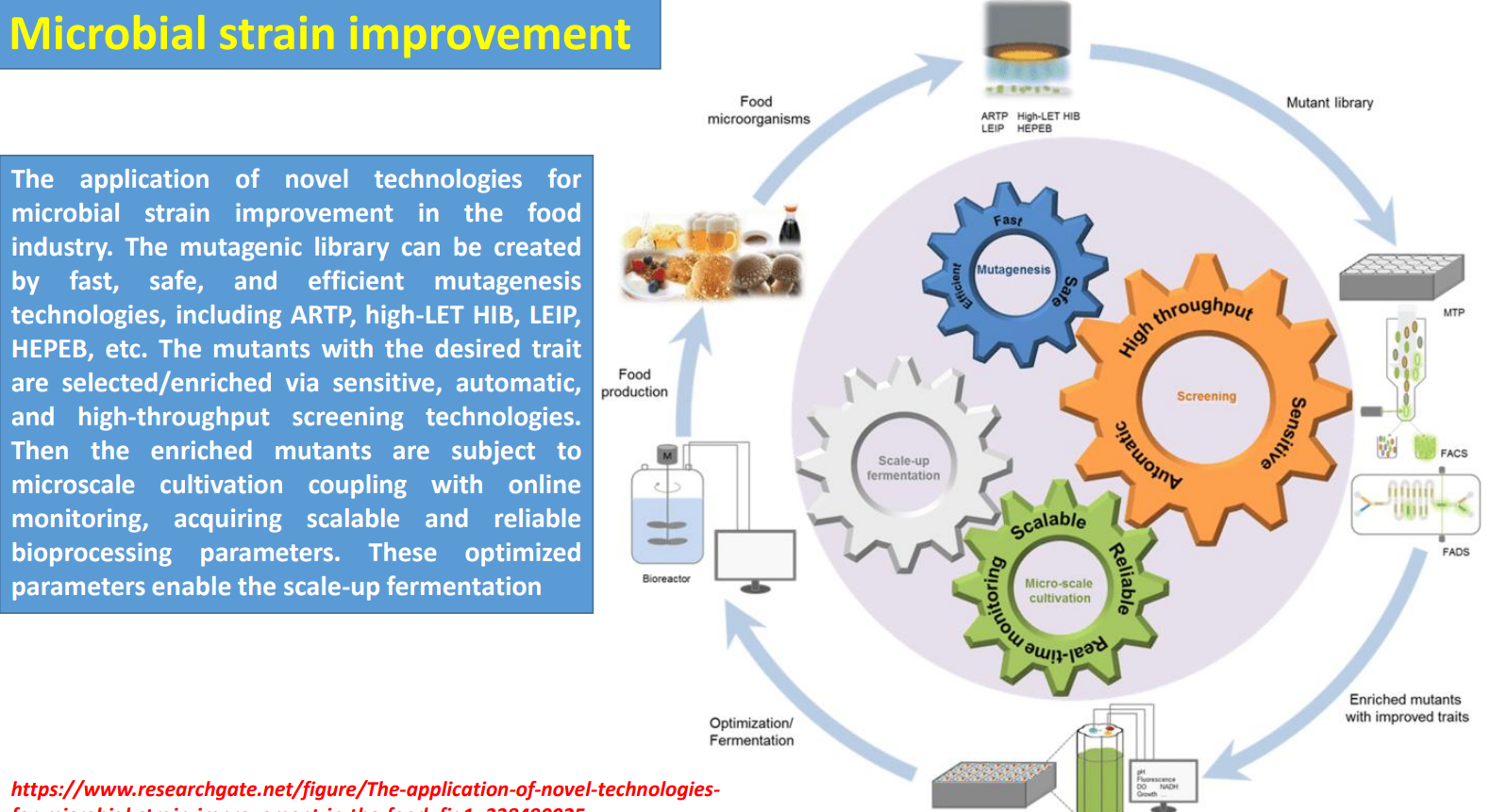
Microbial strain improvement: Microscale cultivation + online monitoring → translating phenotype to process
Selected mutants are moved into microscale cultivation systems where real process variables are introduced:
oxygen limitation
pH shifts
nutrient gradients
Online monitoring (e.g., dissolved oxygen, NADH signals) allows detection of:
metabolic bottlenecks
overflow metabolism (e.g., acetate formation in bacteria)
This step ensures that selected strains are not just high producers, but:
→ robust under dynamic fermentation conditions
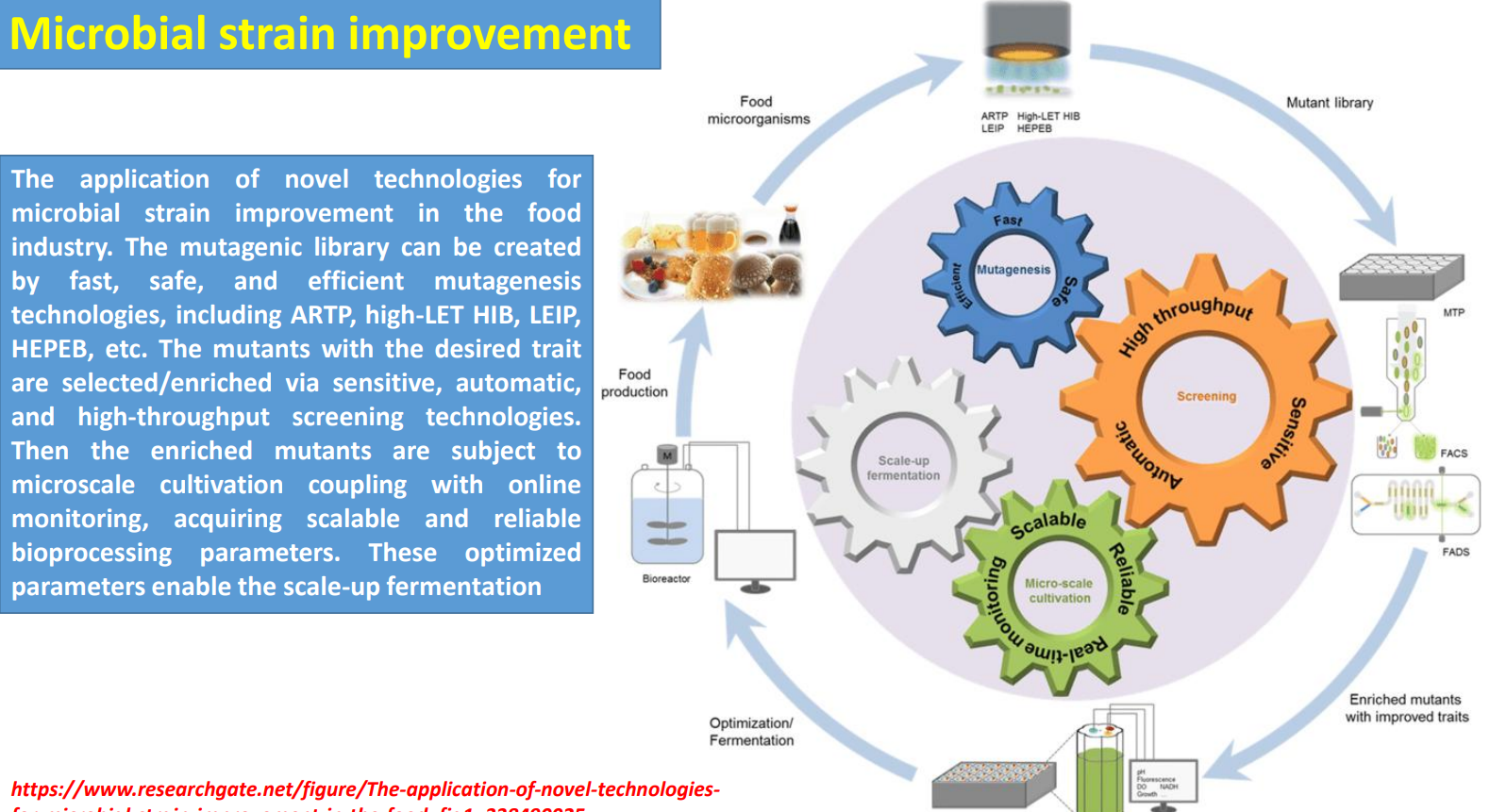
Microbial strain improvement: Parameter extraction → building scalable bioprocess conditions
The text highlights “scalable and reliable bioprocessing parameters”, which come from microscale data:
optimal feeding rate
oxygen demand
growth phase for production
These parameters directly influence:
yield (product per substrate)
productivity (rate of production)
titer (final concentration)
Without this step, scale-up often fails due to:
oxygen transfer limitations
accumulation of toxic intermediates
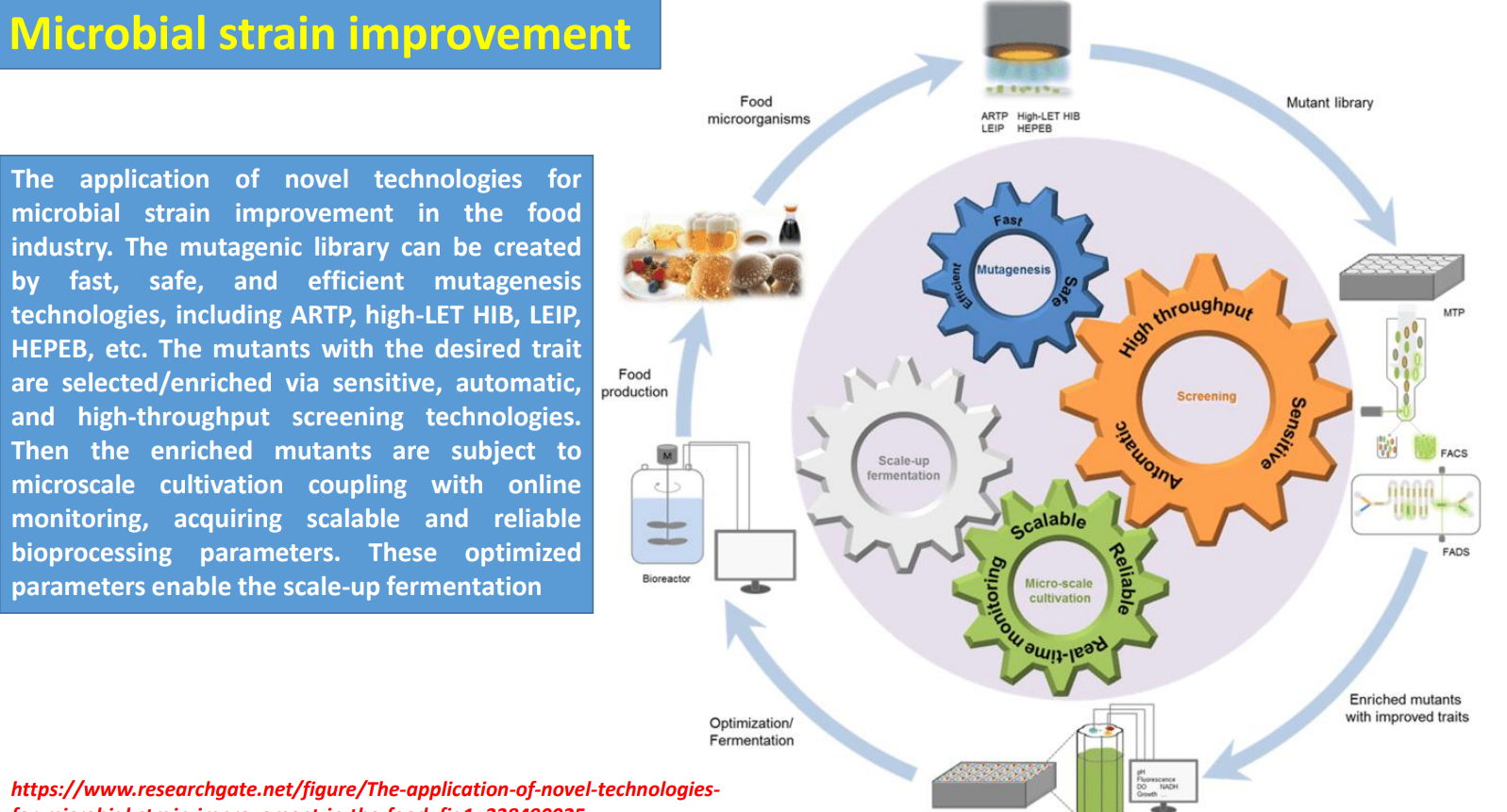
Microbial strain improvement: Scale-up fermentation → validating strain–process compatibility
The diagram’s bioreactor stage represents testing whether:
the strain maintains productivity at larger volume
metabolic behavior remains stable over time
At scale, additional stresses appear:
uneven mixing → nutrient gradients
heat accumulation
shear forces
Many mutants fail here because:
→ improvements at cellular level are not compatible with physical constraints of bioreactors
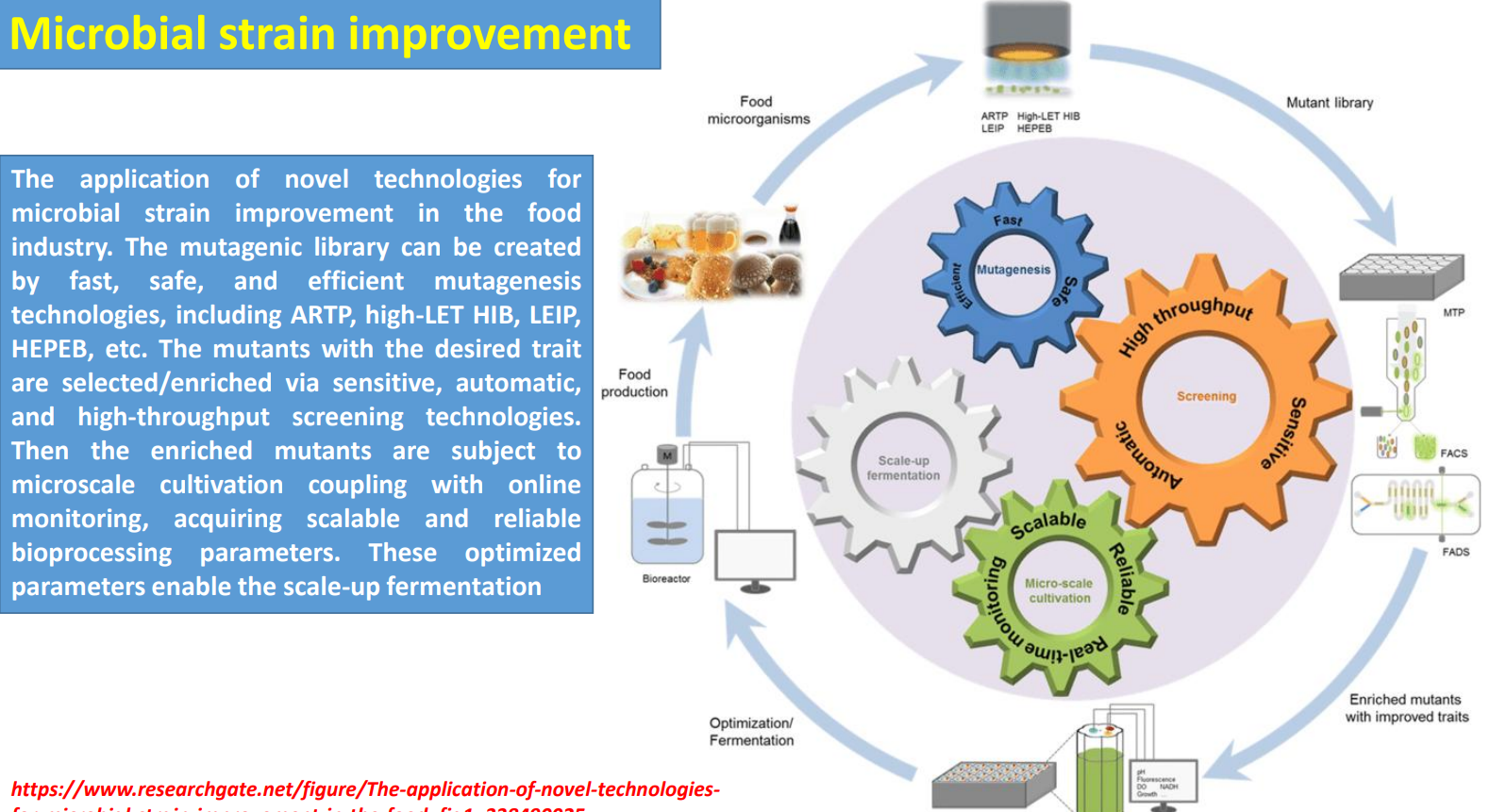
Microbial strain improvement: Iterative optimization loop → compensating for unpredictability
The circular flow in the diagram reflects repeated cycles of:
mutagenesis → screening → cultivation → fermentation
This iterative approach is necessary because:
modifying one pathway often disrupts others
improvements in yield may increase byproducts or reduce growth
Over multiple cycles, strains are tuned for:
metabolic efficiency
stress tolerance
process compatibility
Microbial strain improvement: Mutagenic library creation
Text: “mutagenic library can be created by fast, safe, efficient mutagenesis technologies”
Meaning:
Instead of designing one strain, you generate thousands–millions of variants
Methods (ARTP, HIB, etc.) introduce random mutations across genome
Why:
→ increases probability of discovering improved traits
→ bypasses need to fully understand complex biology
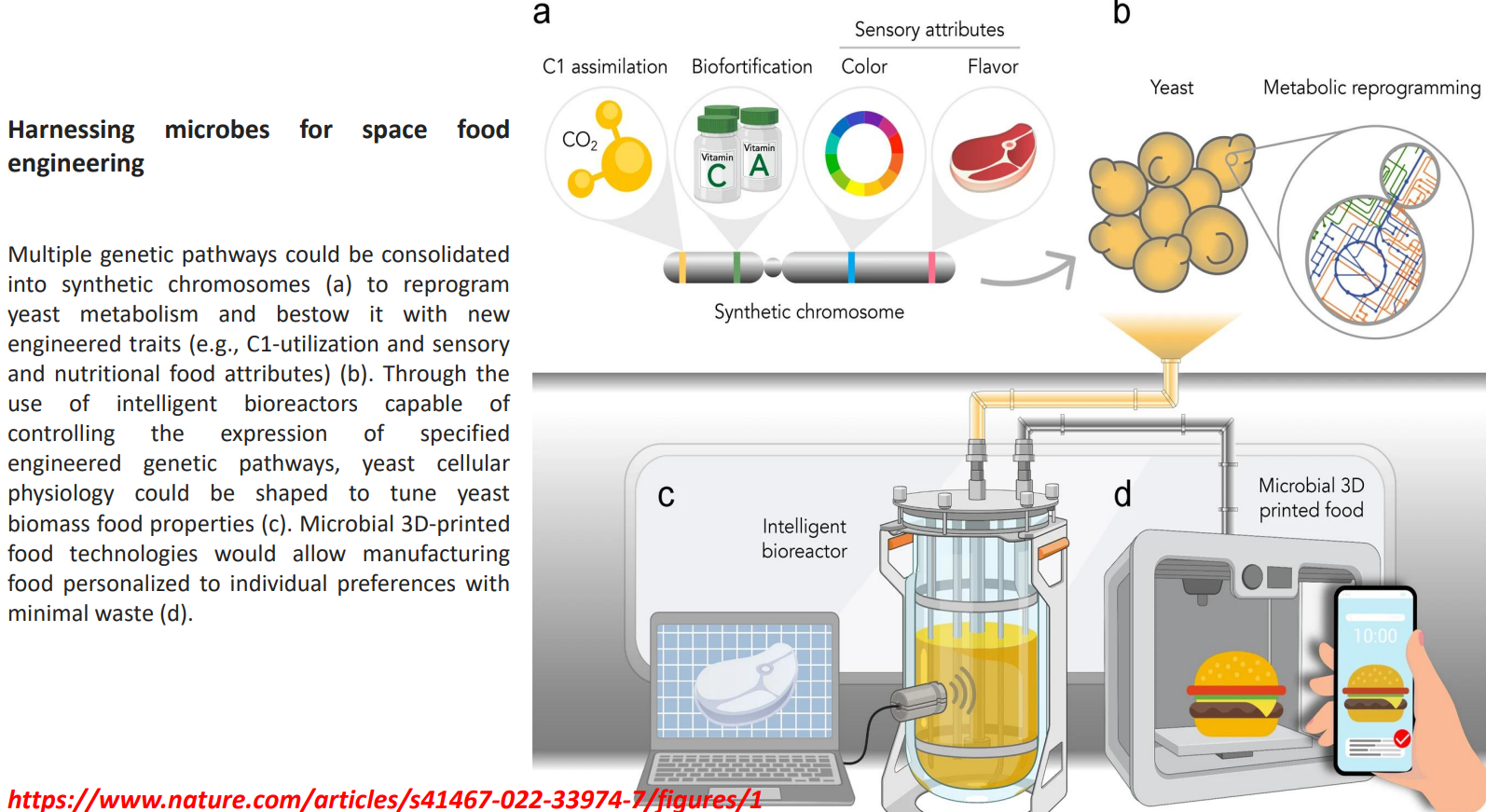
Harnessing microbes for space food engineering: Synthetic chromosomes enable new metabolic functions
Multiple pathways are combined into synthetic chromosomes to reprogram yeast at a systems level
Enables traits not naturally present:
C1 assimilation → using CO₂ or single-carbon substrates instead of sugars
Biofortification → direct production of vitamins (A, C) within biomass
Sensory traits → engineered pigment and flavor pathways
This shifts production from:
→ sugar-dependent fermentation
to
→ resource-efficient metabolism suitable for closed environments (space)
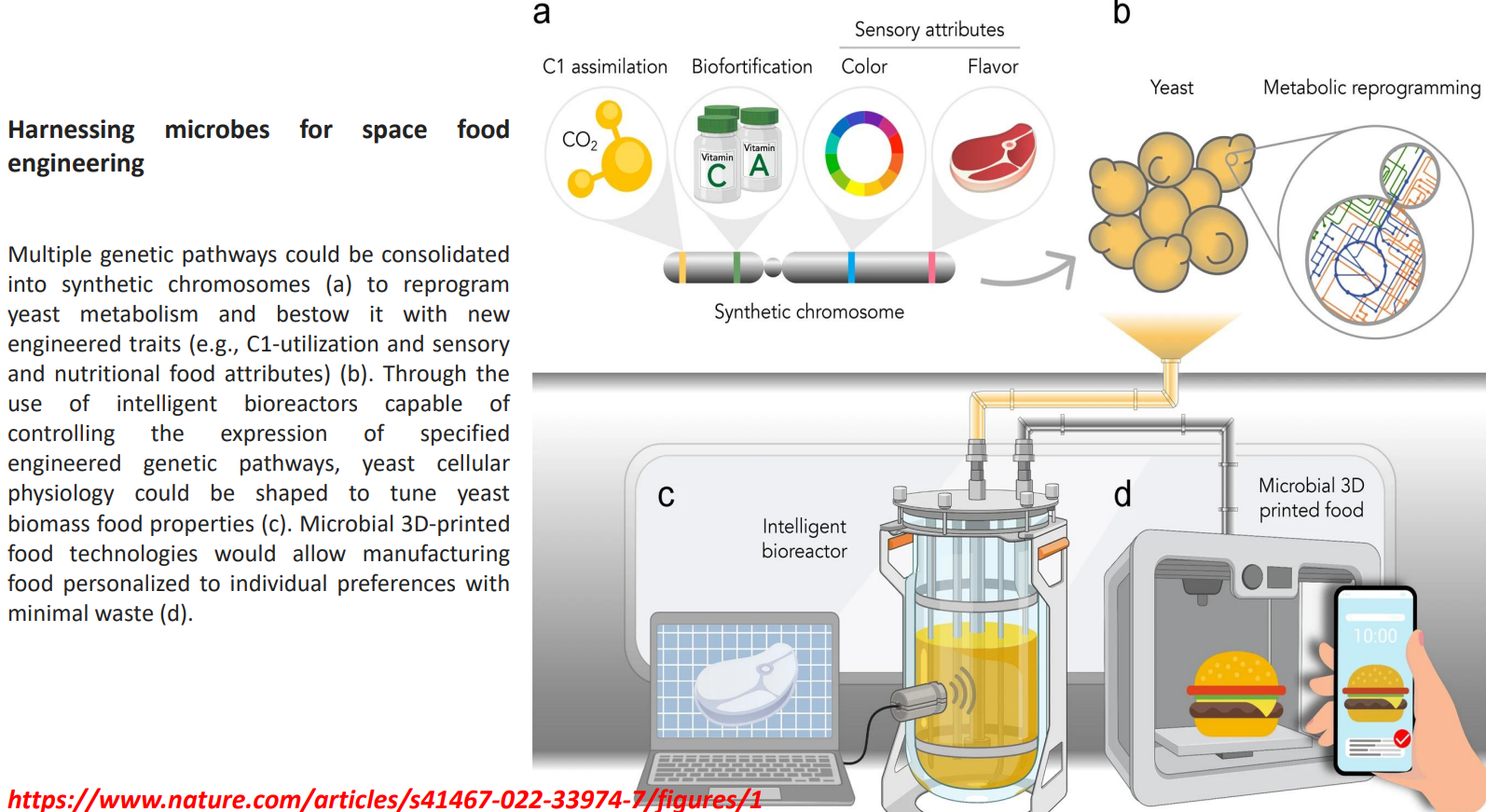
Harnessing microbes for space food engineering: Metabolic reprogramming under cellular constraints
Introducing multiple pathways requires:
redistribution of carbon flux
balancing cofactors (ATP, NADPH)
Without control:
→ growth slows
→ byproducts accumulateEffective designs:
decouple growth and production phases
prioritize flux toward target molecules while maintaining viability
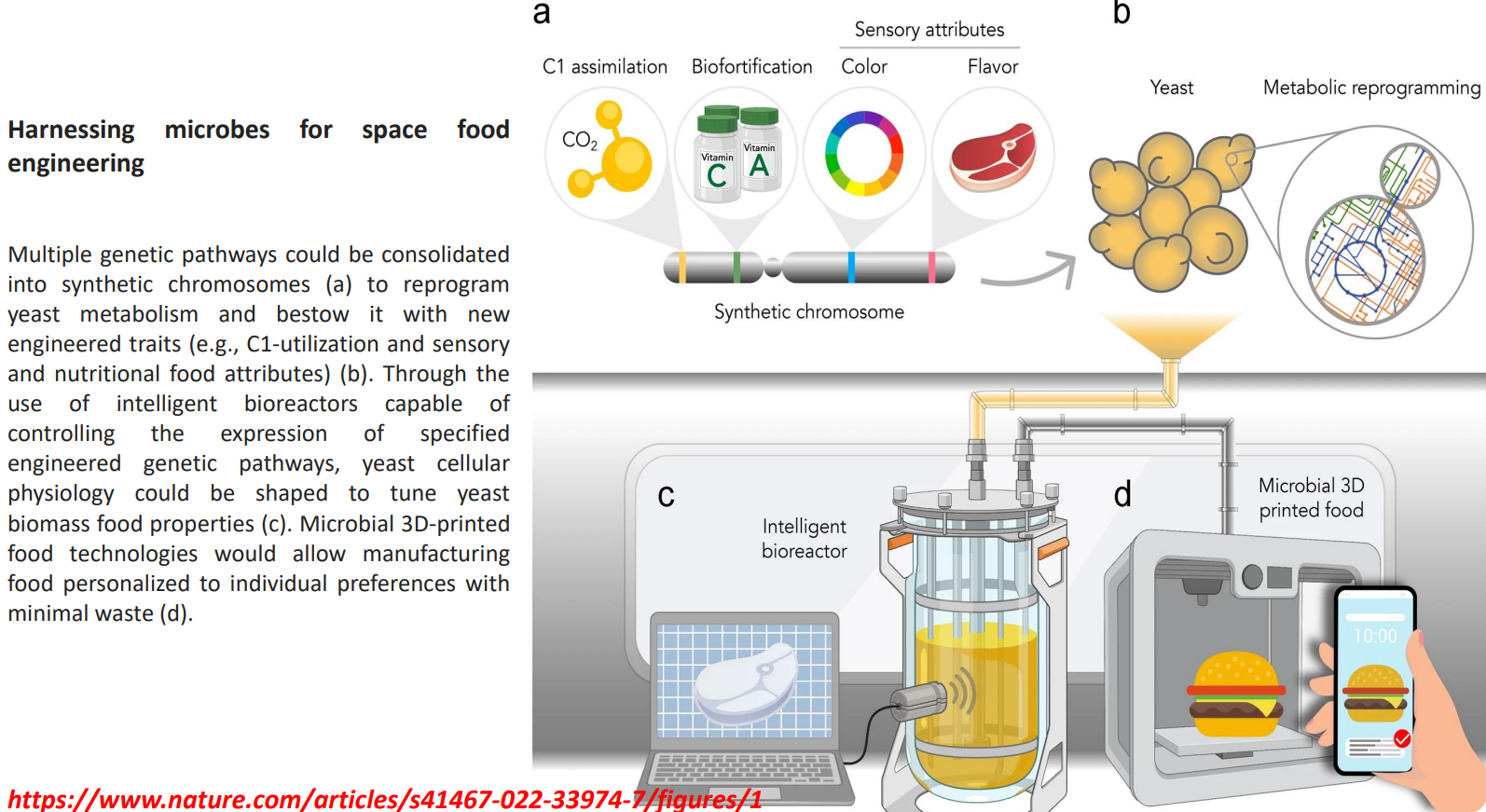
Harnessing microbes for space food engineering: Intelligent bioreactors regulate expression dynamically
Engineered pathways are controlled in real time using:
sensors (oxygen, metabolites, growth signals)
automated feedback systems
Allows:
switching pathways ON/OFF depending on growth stage
maintaining optimal metabolic state
Prevents:
energy waste
toxic intermediate buildup
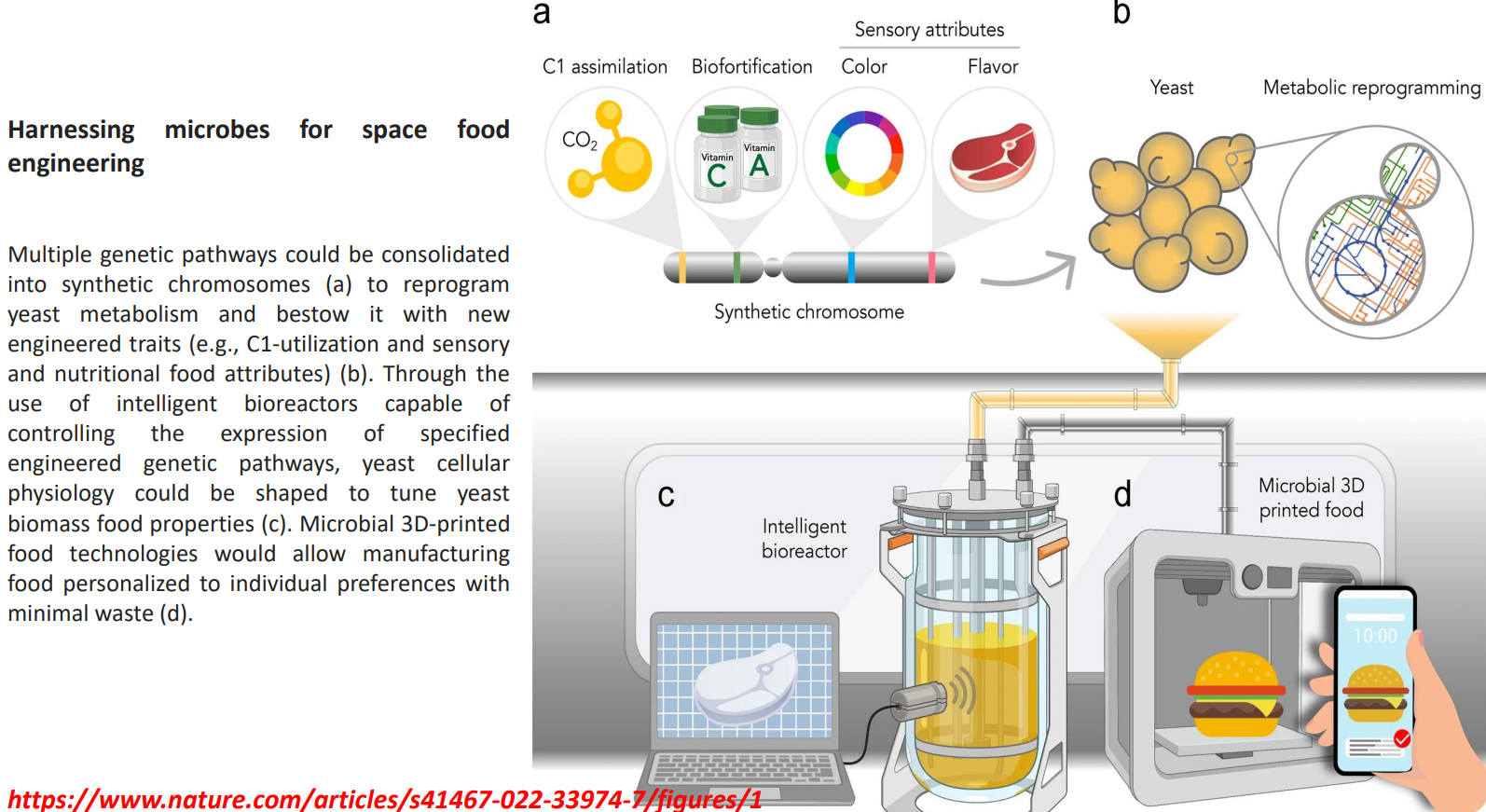
Harnessing microbes for space food engineering: Biomass tuned into functional food material
Yeast biomass is engineered to deliver:
nutritional value (vitamins, amino acids)
sensory properties (flavor, color)
functional traits (texture, binding)
Composition is controlled through:
metabolic pathway design
fermentation conditions
This makes biomass itself the final food ingredient, not just a production system
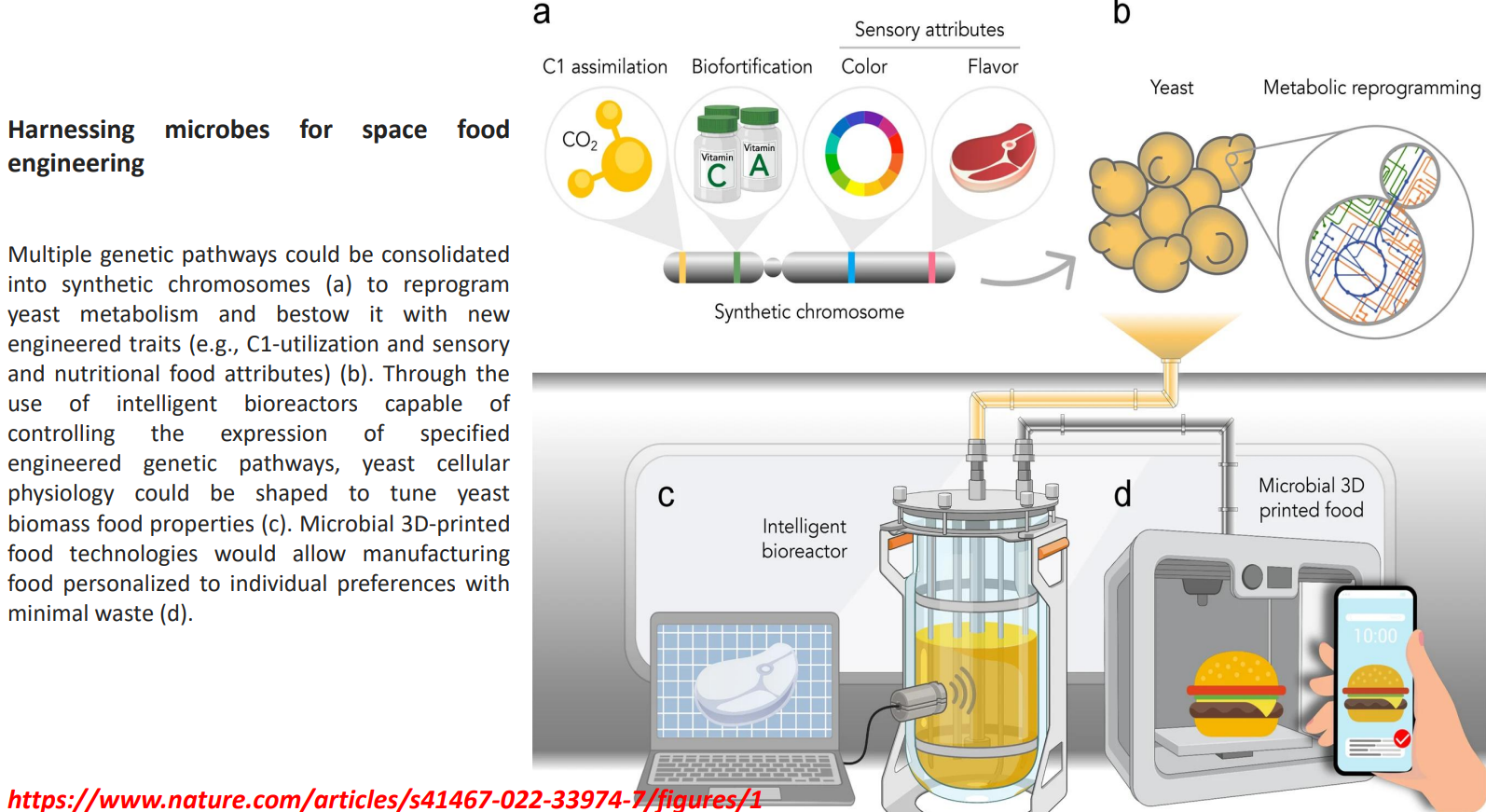
Harnessing microbes for space food engineering: Microbial 3D food structuring
Biomass is processed into structured foods using 3D printing
Enables:
controlled texture (e.g., meat-like structure)
personalized nutrition profiles
minimal waste production
Solves the gap between:
→ biochemical production
→ consumer-ready food form
Design–build–test–learn (DBTL) cycle: Design
Define the biological objective (e.g., produce a protein, metabolite)
Select:
metabolic pathway
genes/enzymes
regulatory elements (promoters, ribosome binding sites)
In practice:
use computational models (flux balance, pathway prediction)
predict bottlenecks before experimentation
Constraint:
→ incomplete knowledge of cellular networks → designs are approximations
Design–build–test–learn (DBTL) cycle: Build
Physically construct the system:
DNA synthesis
assembly of genetic parts
insertion into host organism
Modern methods:
modular cloning (Golden Gate, Gibson assembly)
genome editing (CRISPR)
Key issue:
→ introduced pathways compete with native metabolism for:energy (ATP)
cofactors (NADH/NADPH)
Design–build–test–learn (DBTL) cycle: Test
Evaluate engineered organism:
product yield
growth rate
byproduct formation
Data collected using:
metabolomics
proteomics
fermentation measurements
Important reality:
→ many designs fail due to:metabolic imbalance
toxicity
regulatory interference
Design–build–test–learn (DBTL) cycle: Learn
Analyze test data to identify:
bottlenecks in pathways
inefficient enzyme steps
regulatory constraints
Update:
gene expression levels
pathway structure
host modifications
Increasingly uses:
machine learning
large datasets from previous cycles
Iterative optimization
Cycle repeats:
→ each round improves system performanceOver time:
designs become more predictive
fewer experimental iterations needed
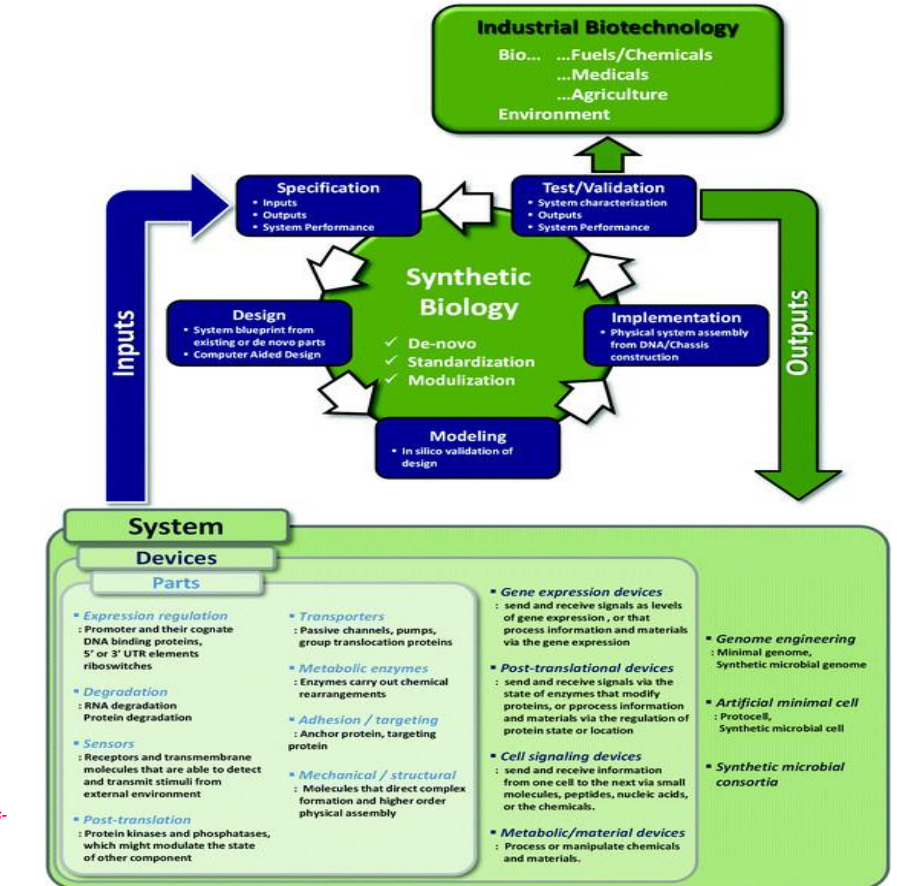
Synthetic biology engineering design pipeline: Specification
Specification → defining system inputs and outputs
Start by defining:
inputs (substrates, signals)
outputs (desired product, behavior)
system performance targets (yield, rate, stability)
This anchors design to industrial requirements (fuels, chemicals, food, medicines) rather than just biological curiosity
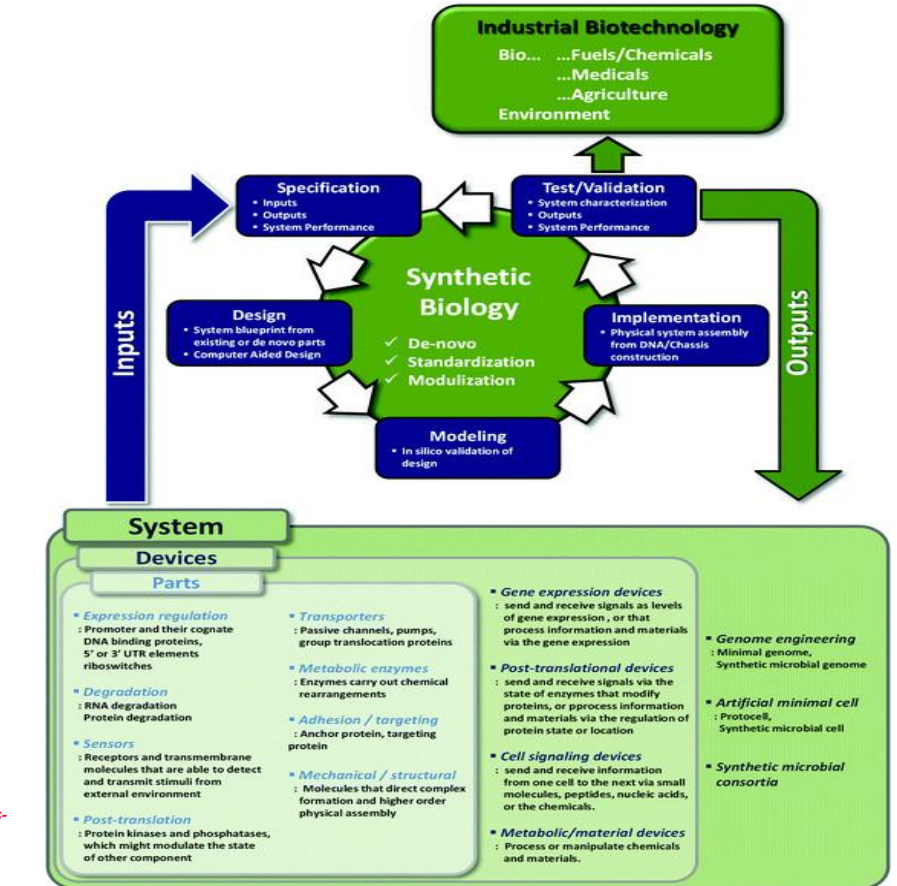
Synthetic biology engineering design pipeline: Design → assembling biological parts into systems
Build a system blueprint using:
existing biological parts
de novo designed components
Parts include:
promoters, riboswitches → control expression
sensors → detect environmental signals
enzymes → drive metabolic reactions
transporters → move substrates/products
These are combined into devices (functional units), then into full systems
Key principle:
→ modularity allows reuse and recombination of parts
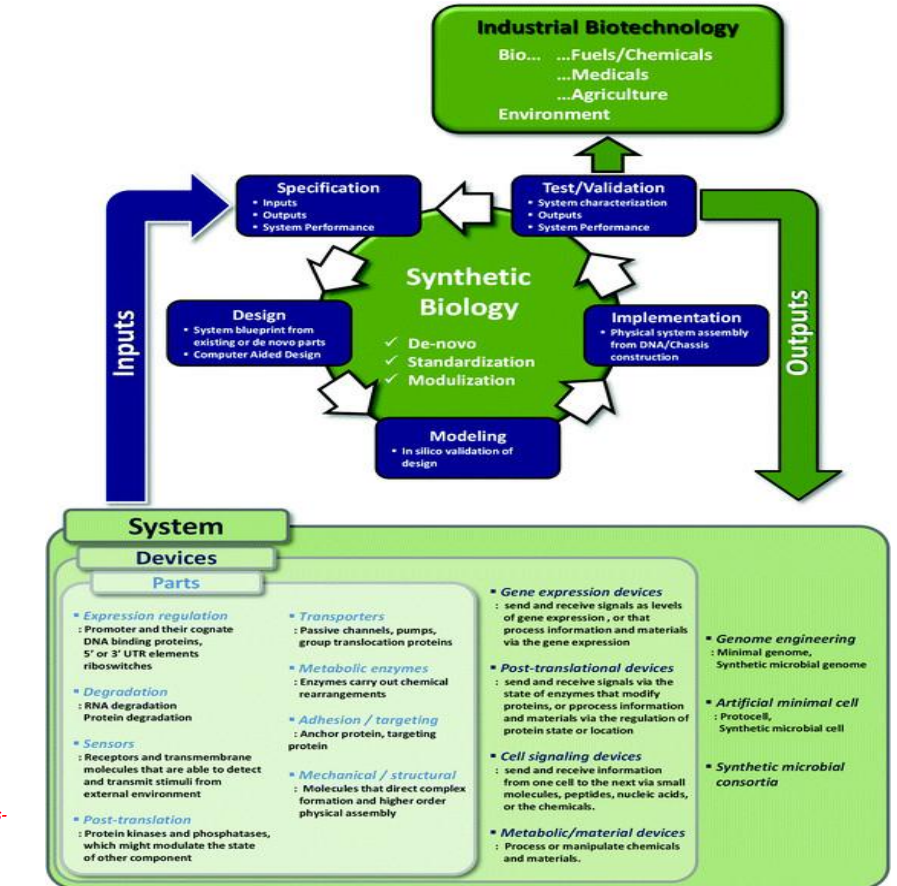
Synthetic biology engineering design pipeline: Modeling → in silico validation
Before building, designs are tested computationally:
metabolic flux modeling
pathway simulation
Purpose:
predict bottlenecks
estimate yields
reduce experimental iterations
Limitation:
→ models approximate reality; cellular networks remain partially unknown
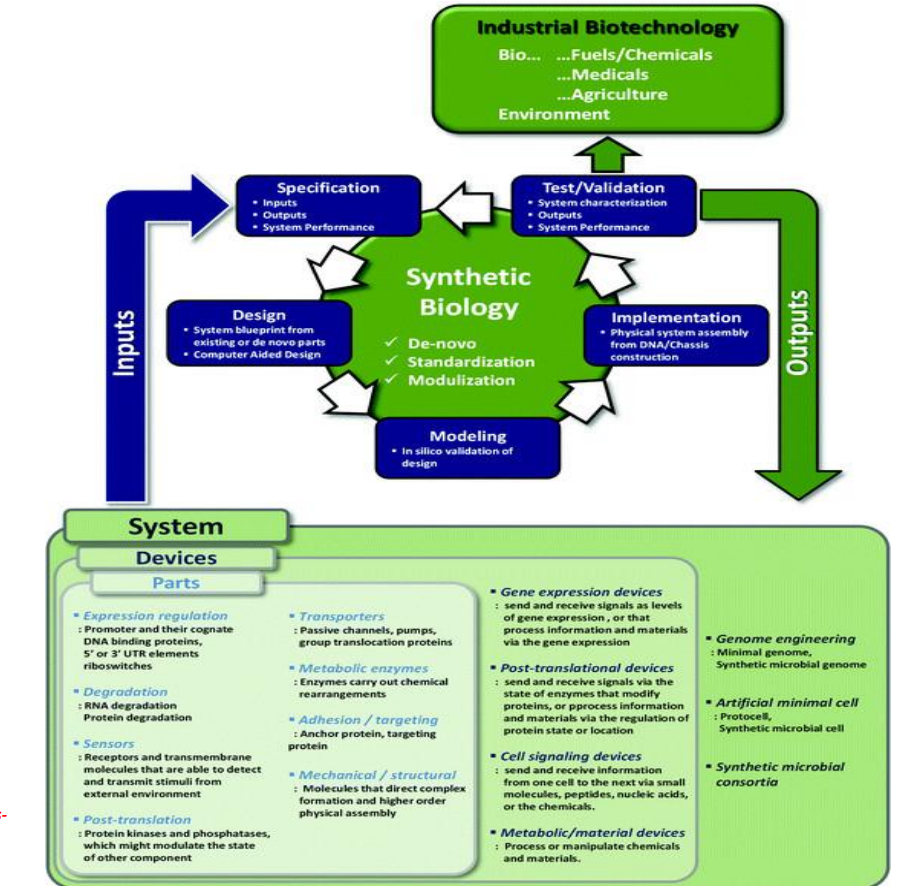
Synthetic biology engineering design pipeline: Implementation → physical system construction
Assemble DNA and introduce into host:
plasmid construction
genome engineering
Includes advanced strategies:
minimal genomes → remove unnecessary genes
synthetic genomes → fully redesigned organisms
synthetic consortia → multiple microbes working together
Constraint:
→ engineered systems must coexist with native cellular processes
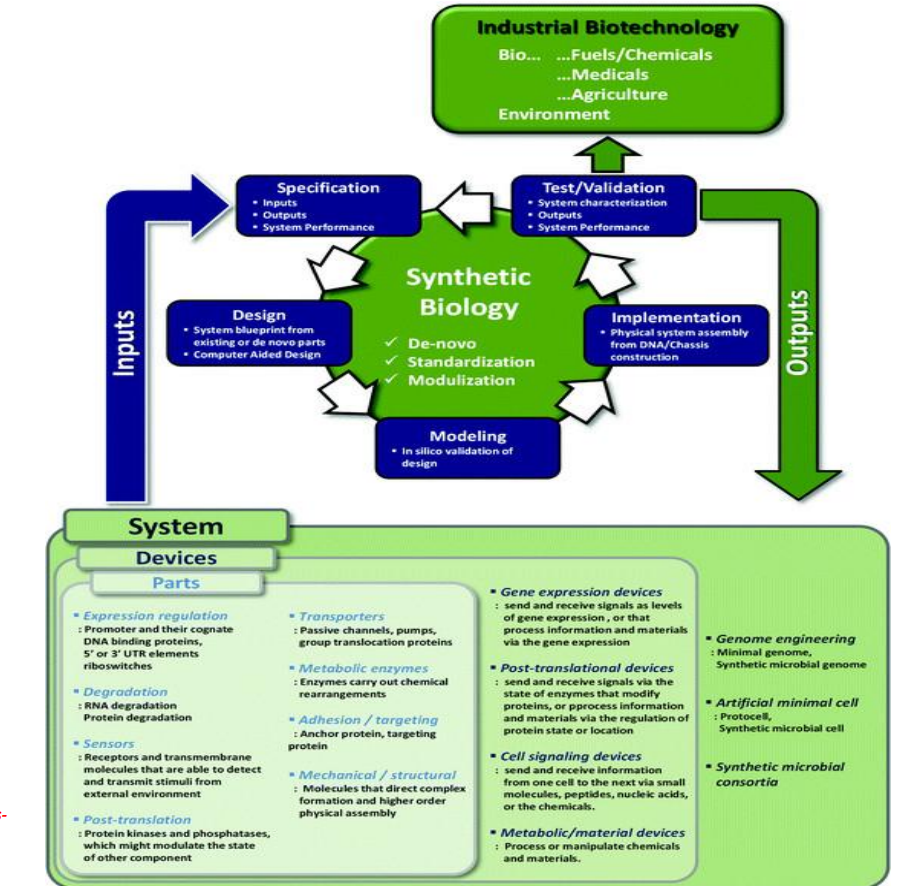
Synthetic biology engineering design pipeline: Test / Validation → system characterization
Evaluate:
product output
system performance
stability
Includes:
protein expression analysis
metabolic profiling
process-level testing
Determines whether system meets specification targets
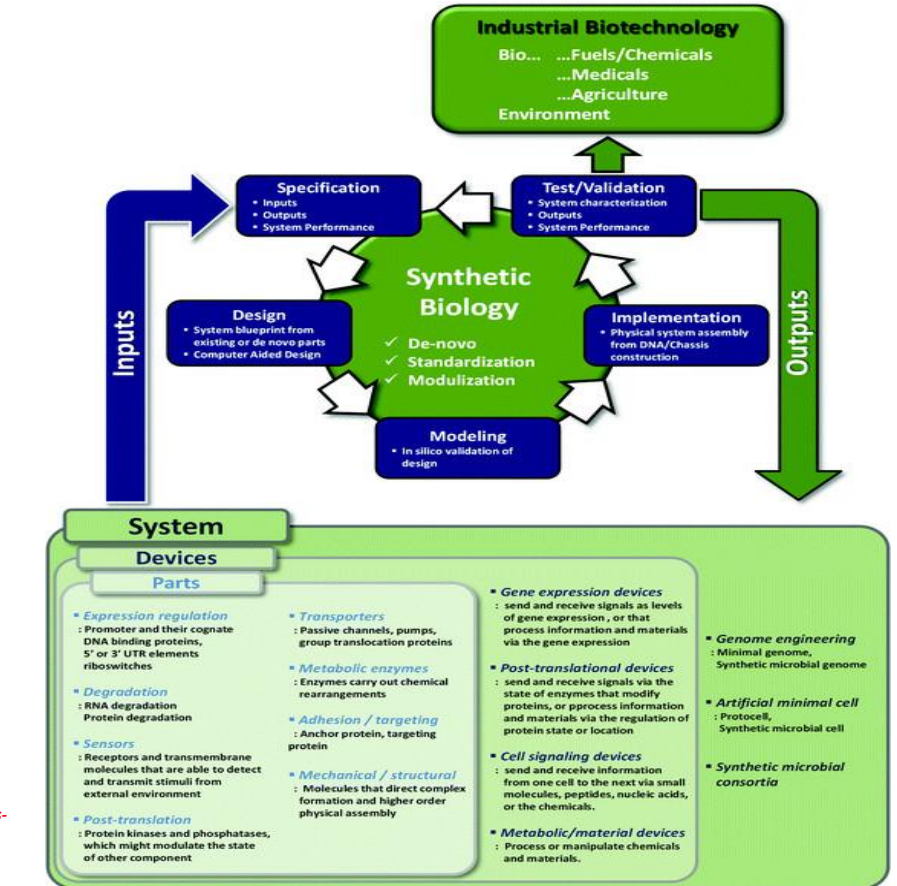
Synthetic biology engineering design pipeline: Parts (basic biological components)
Expression regulation
Promoters, transcription factors, riboswitches, UTRs
→ control how much protein is made
Degradation
RNA/protein degradation tags
→ control how long molecules last → prevents buildup or toxicity
Sensors
Receptors, membrane proteins
→ detect:nutrients
stress signals
environmental changes
Post-translation control
Kinases, phosphatases
→ modify protein activity after synthesis (on/off switching)
Transporters
Channels, pumps
→ move substrates/products across membranes
→ often a bottleneck in production systems
Metabolic enzymes
Catalyze reactions
→ determine pathway efficiency and yield
Adhesion / targeting
Anchor proteins
→ localize proteins or attach cells to surfaces
Mechanical / structural
Scaffold molecules
→ organize multi-enzyme complexes for efficient reactions
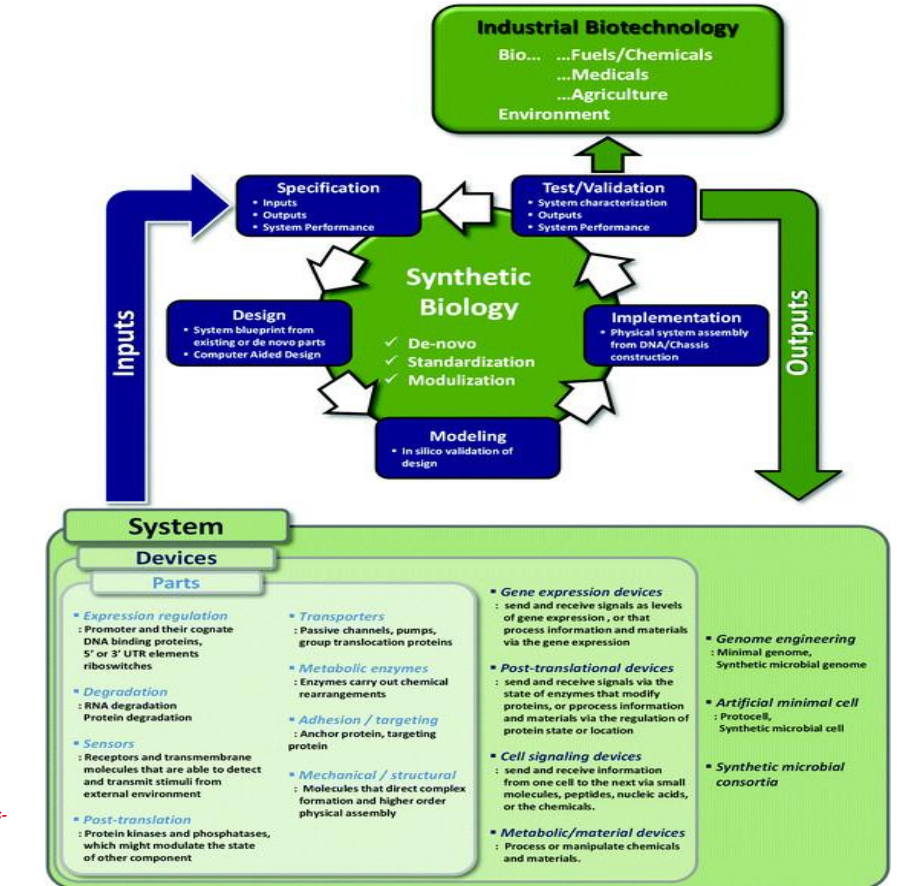
Synthetic biology engineering design pipeline: Devices (functional modules built from parts)
Gene expression devices
→ integrate regulatory parts to control:timing
intensity of expression
→ act like biological “switches” or “circuits”
Post-translational devices
→ regulate activity of proteins dynamically
→ faster control than gene-level regulationCell signaling devices
→ enable communication between cells using:small molecules
peptides
→ useful in multi-cell systems or population control
Metabolic/material devices
→ coordinate multiple enzymes to convert substrates into products
→ core of bioproduction pathways
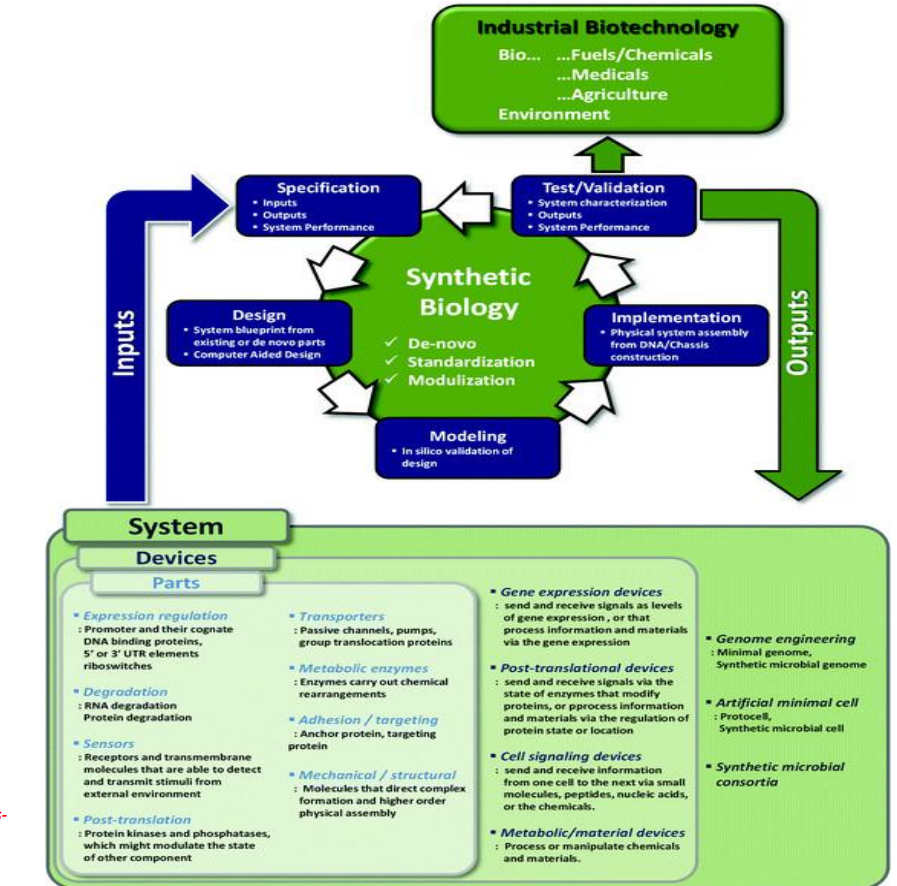
Synthetic biology engineering design pipeline: Systems (fully integrated biological networks)
Combine multiple devices into a coordinated system that can:
sense environment
process information
produce outputs
Advanced system-level designs include:
Genome engineering
→ minimal genomes, synthetic genomes for efficiencyArtificial minimal cells
→ stripped-down systems with only essential functionsSynthetic microbial consortia
→ multiple organisms sharing tasks (division of labor)
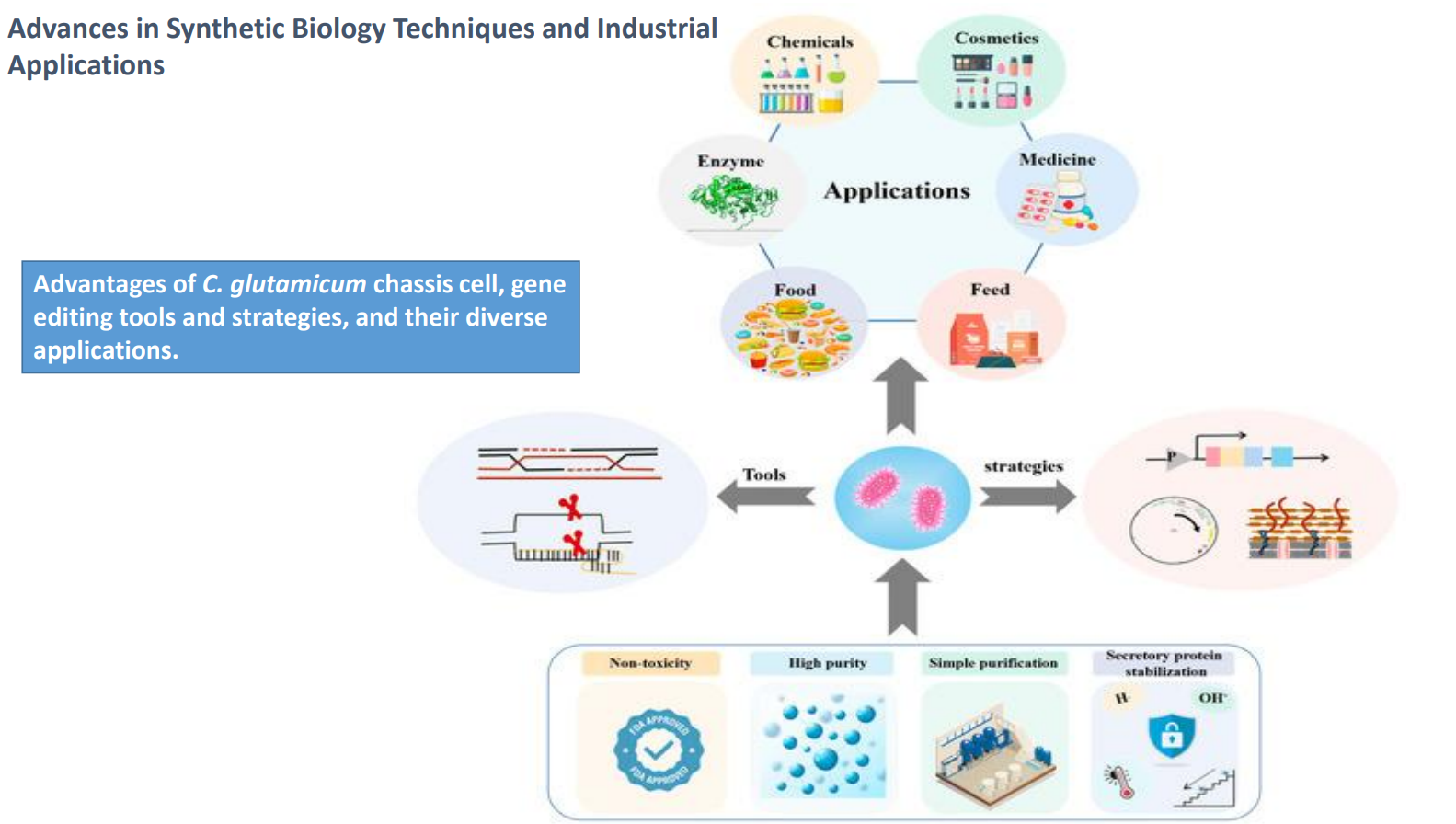
Advances in synthetic biology techniques and industrial applications: C. glutamicum as a chassis cell
Used as a model industrial microbe because it naturally produces amino acids (e.g., glutamate, lysine) at high levels
Advantages shown at the bottom:
Non-toxicity → safe for food and industrial use
High purity → fewer unwanted byproducts
Simple purification → products often secreted, reducing downstream cost
Secretory protein stabilization → maintains protein integrity outside the cell
Integrated implication:
→ reduces one of the biggest industrial bottlenecks: downstream processing cost
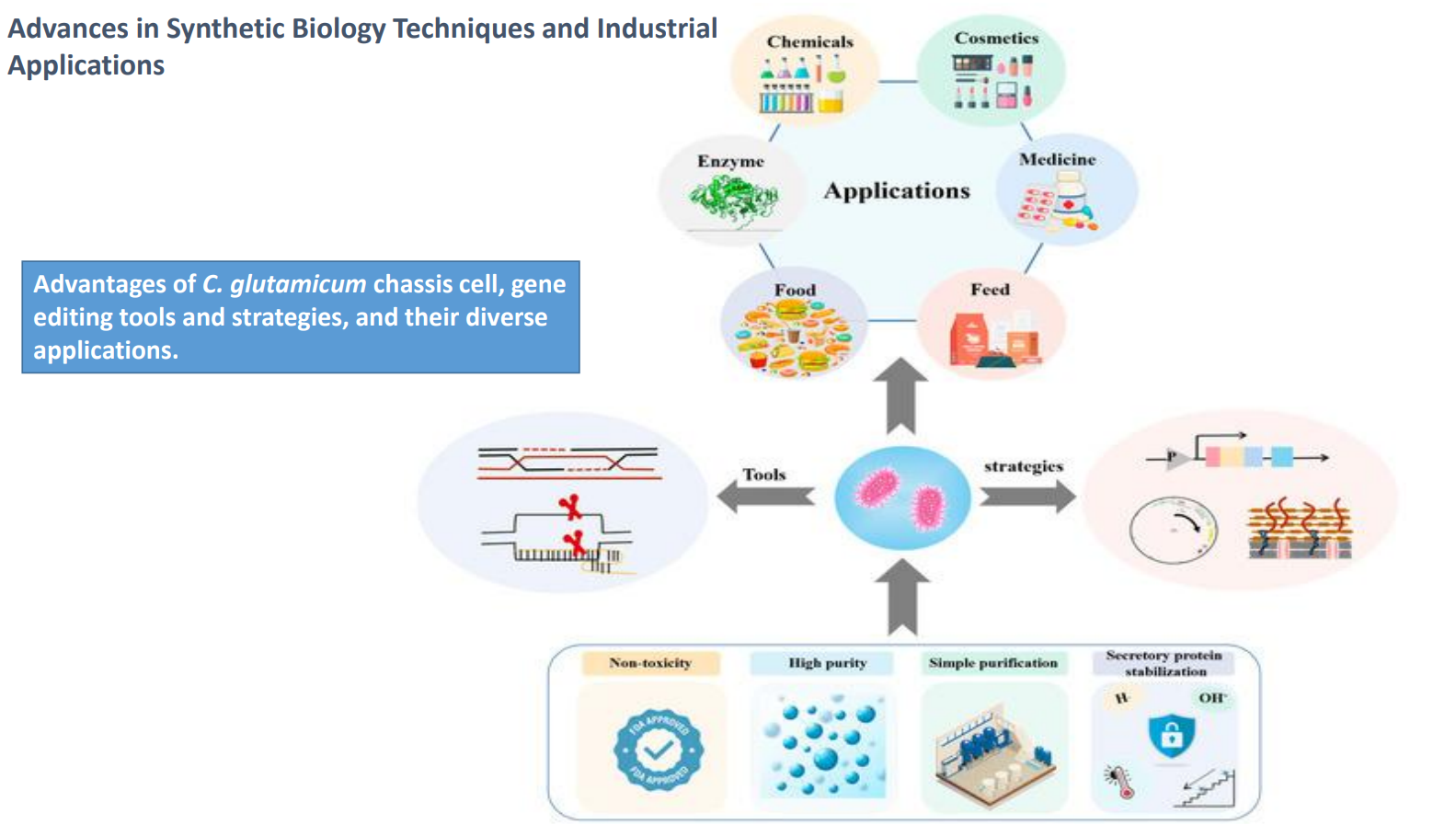
C. glutamicum as a chassis cell: Gene editing tools
Tools (left side) represent:
CRISPR/Cas systems
recombineering
homologous recombination
Enable:
insertion of biosynthetic pathways
deletion of competing pathways
fine-tuning of gene expression
In practice:
→ allows targeted metabolic rewiring instead of random mutagenesis
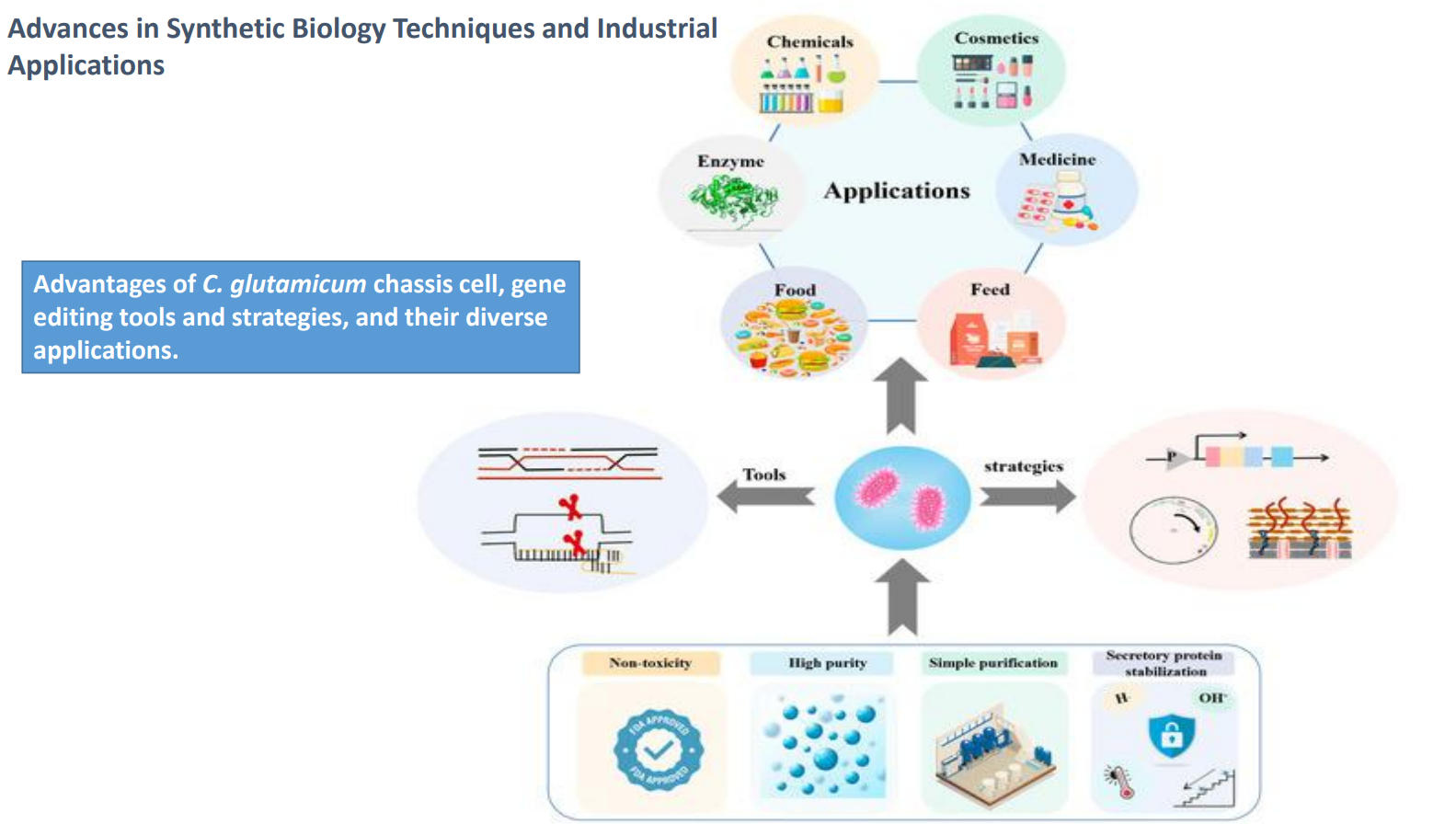
C. glutamicum as a chassis cell: Engineering strategies
Strategies (right side) include:
pathway assembly
promoter engineering
gene copy number control
Also involve:
balancing metabolic flux
optimizing cofactor usage (NADH/NADPH)
reducing accumulation of toxic intermediates
Key constraint:
→ increasing production often creates:metabolic burden
reduced growth rate
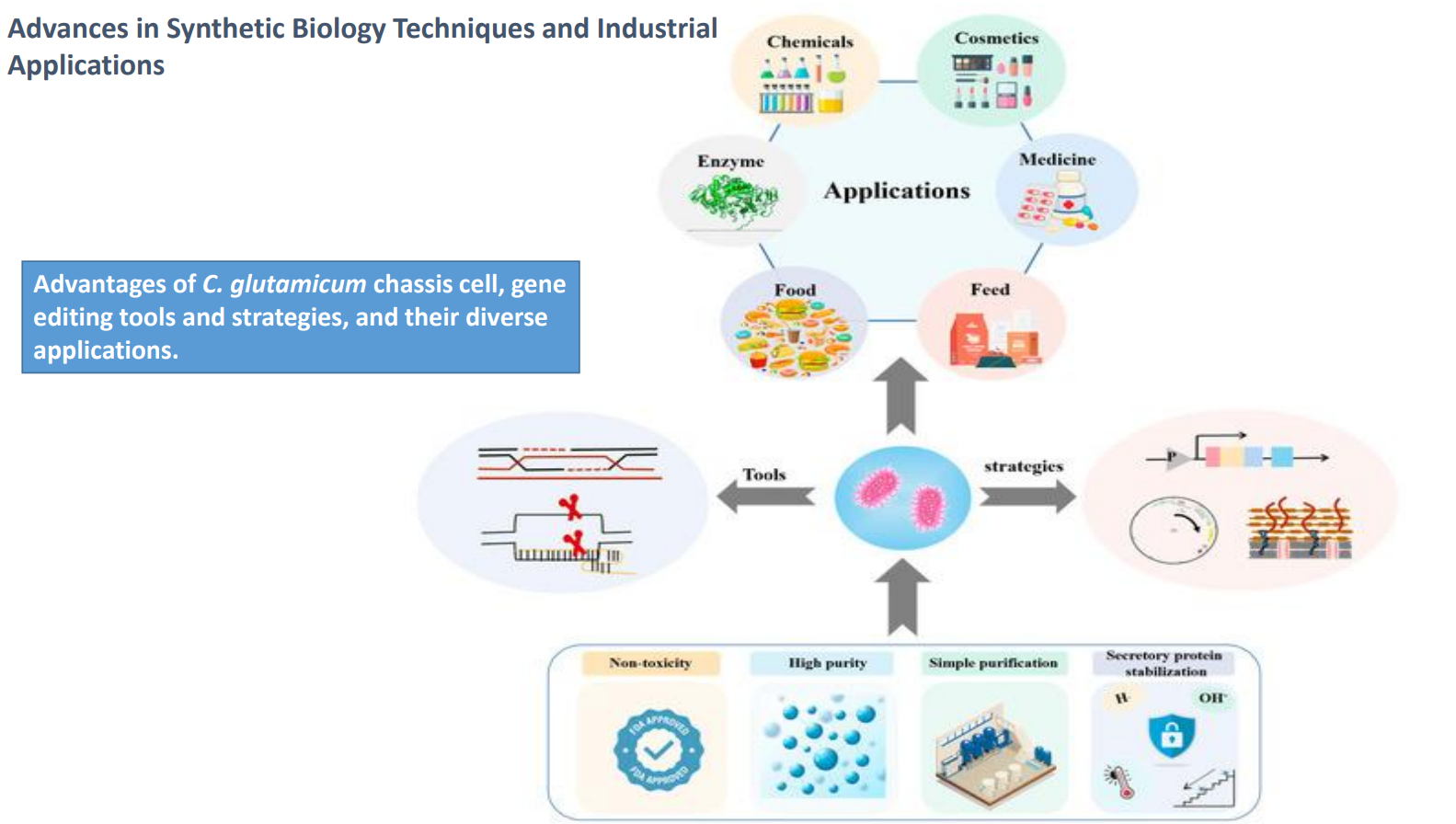
C. glutamicum as a chassis cell: Tools + strategies → engineered microbial system
Central cell represents integration of:
genetic tools
metabolic strategies
Outcome:
→ a strain optimized for:high yield
stability
industrial robustness
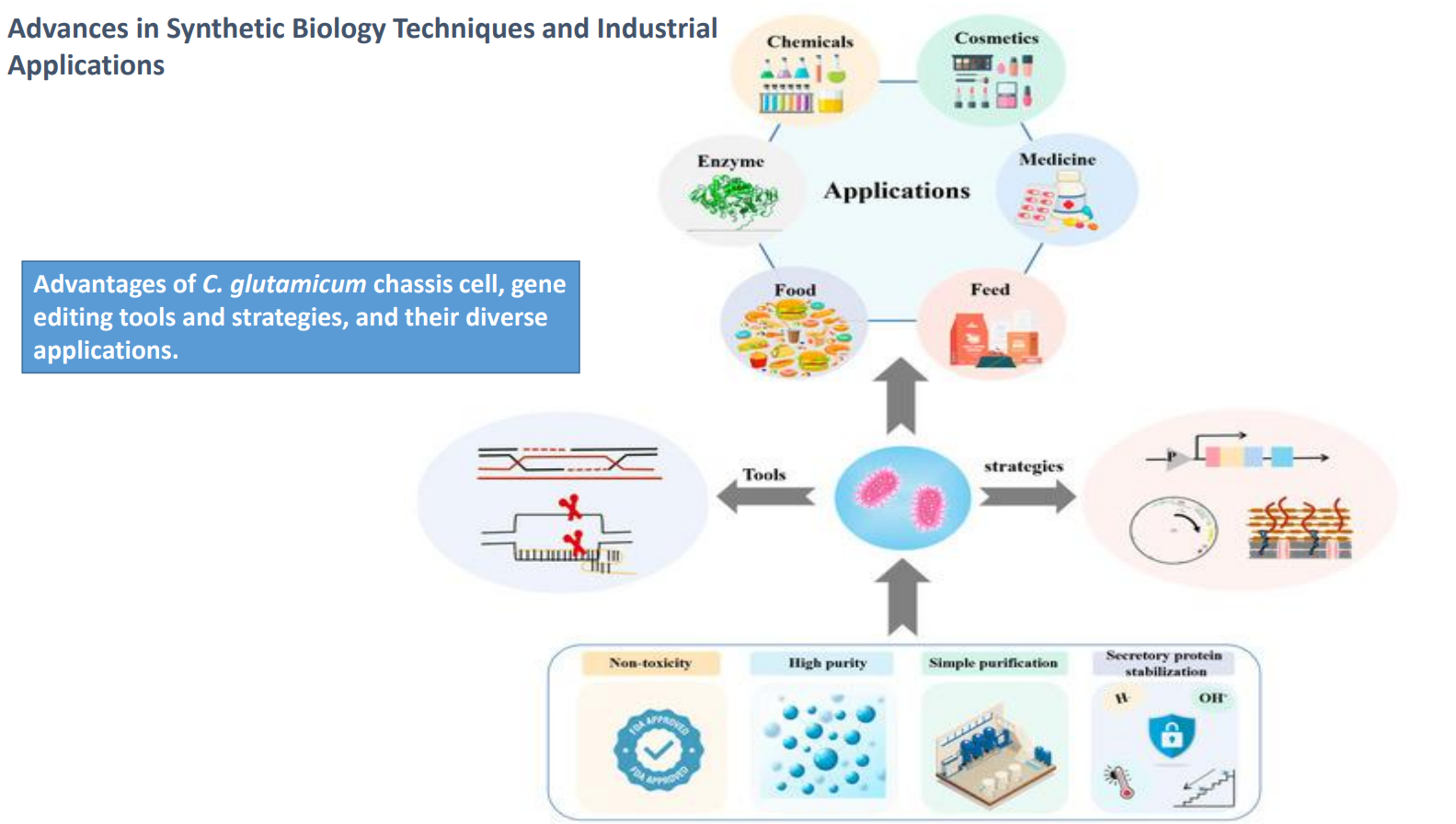
C. glutamicum as a chassis cell: Industrial applications
Amino acids (core strength of C. glutamicum)
L-lysine (animal feed), L-glutamate (MSG), L-threonine, L-valine
Titers often >100 g/L due to strong carbon flux to central metabolism
Detergent enzymes
Proteases, lipases, amylases stable at high pH/temperature
Flavor compounds
Vanillin (from ferulic acid), nootkatone, fruity esters
Drug precursors
Artemisinin precursor (via yeast primarily), statin intermediates, shikimate pathway products
Pigments & antioxidants
Carotenoids (β-carotene, astaxanthin via engineered hosts), coenzyme Q10 intermediates
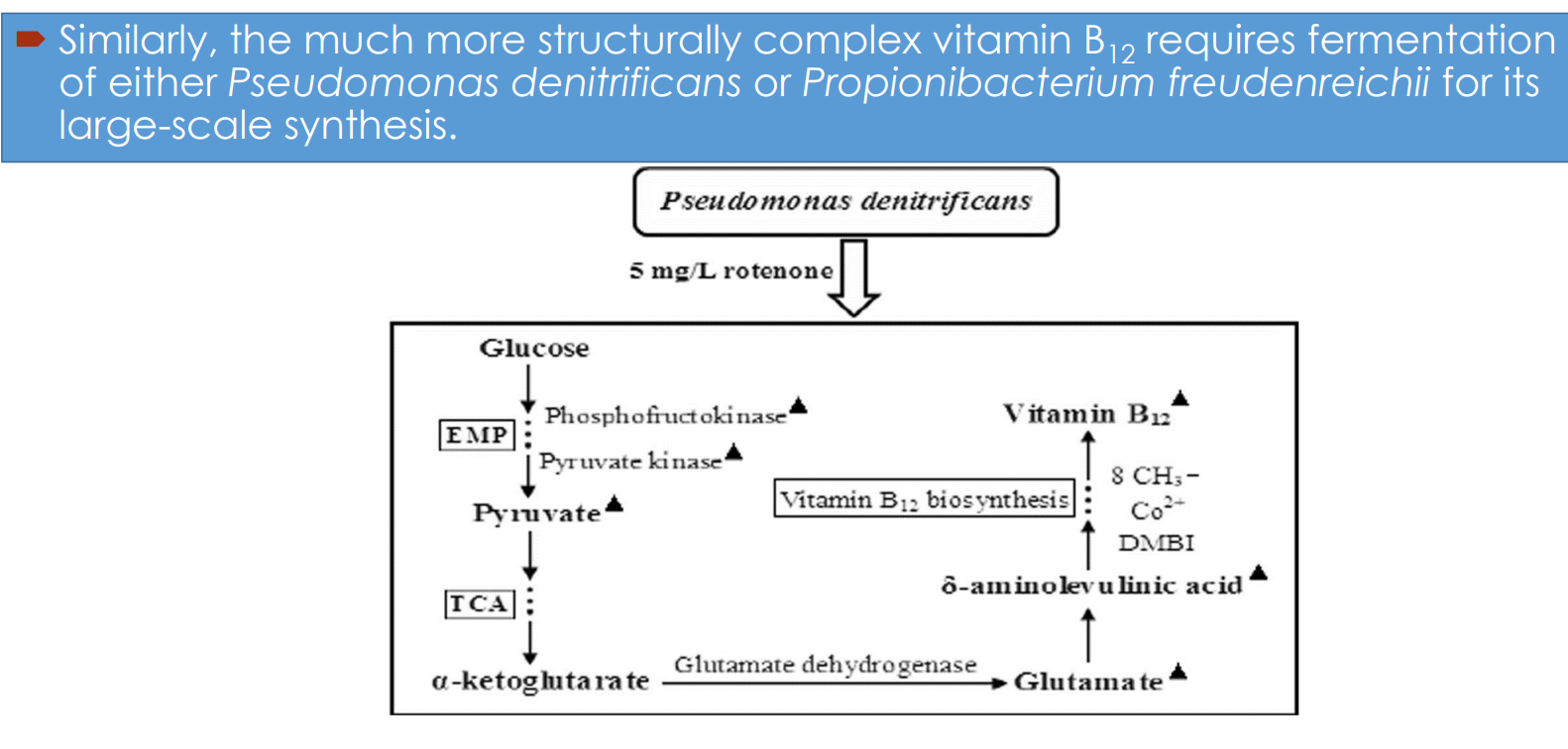
Vitamin B₁₂ fermentation pathway engineering: Microbial hosts
Industrial synthesis relies on:
Pseudomonas denitrificans
Propionibacterium freudenreichii
Reason:
→ B₁₂ has a highly complex corrin ring structure that chemical synthesis cannot produce economicallyTrade-off:
Pseudomonas → faster growth, higher productivity
Propionibacterium → food-grade but slower
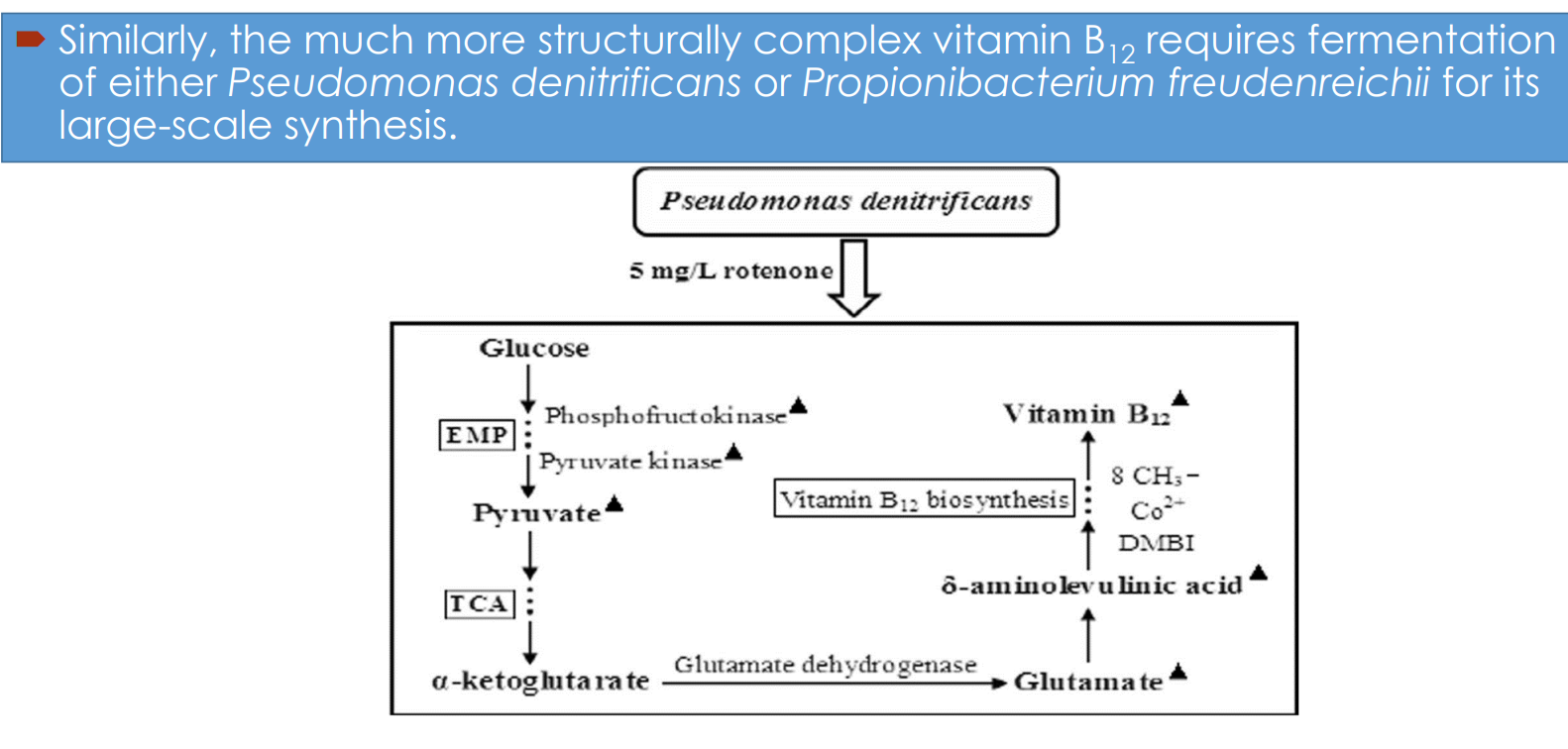
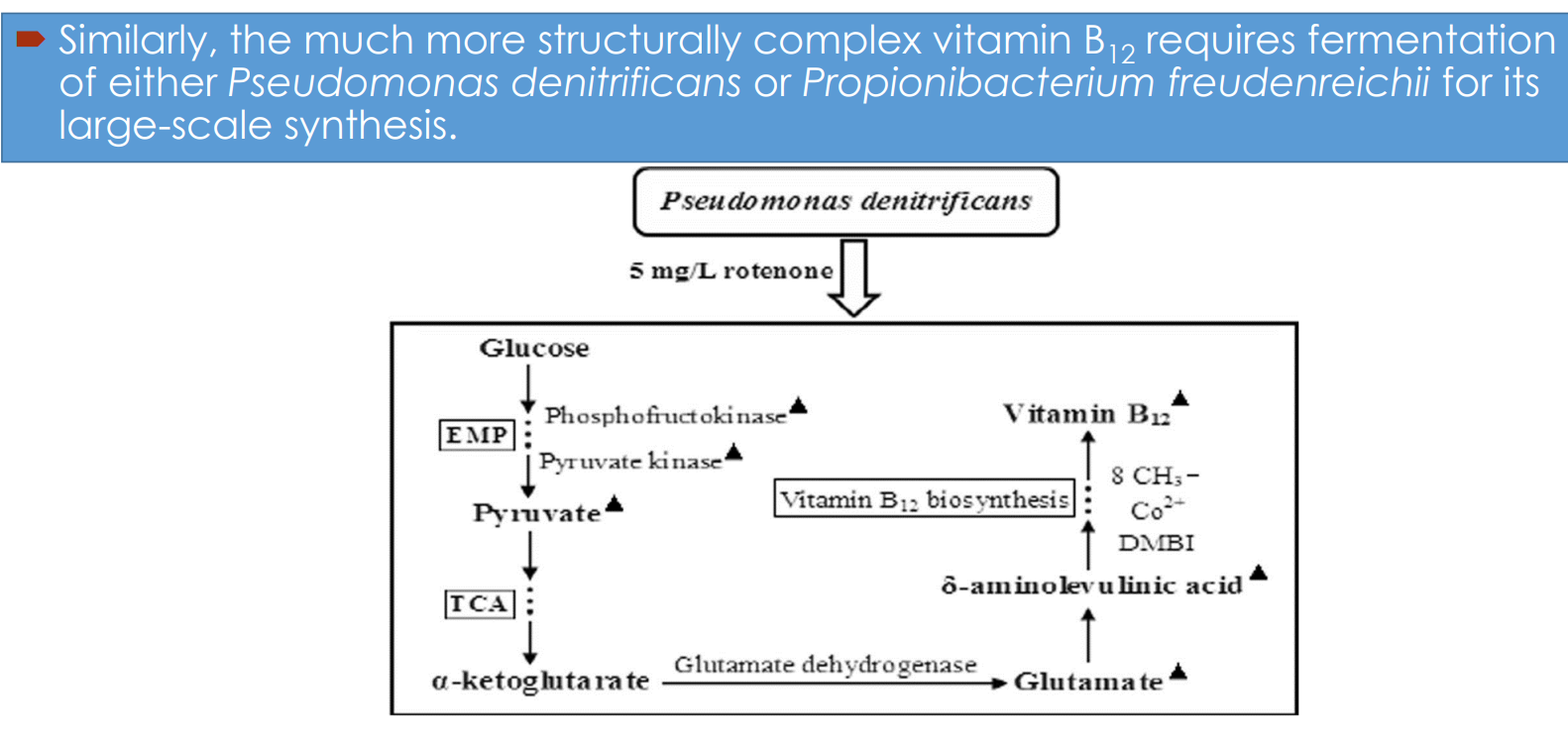
Vitamin B₁₂ fermentation pathway engineering: Central carbon metabolism drives precursor supply
Glucose enters:
EMP (glycolysis) → pyruvate
TCA cycle → α-ketoglutarate
Upregulated enzymes:
phosphofructokinase
pyruvate kinase
Purpose:
→ push carbon flux toward biosynthetic intermediates, not just energy
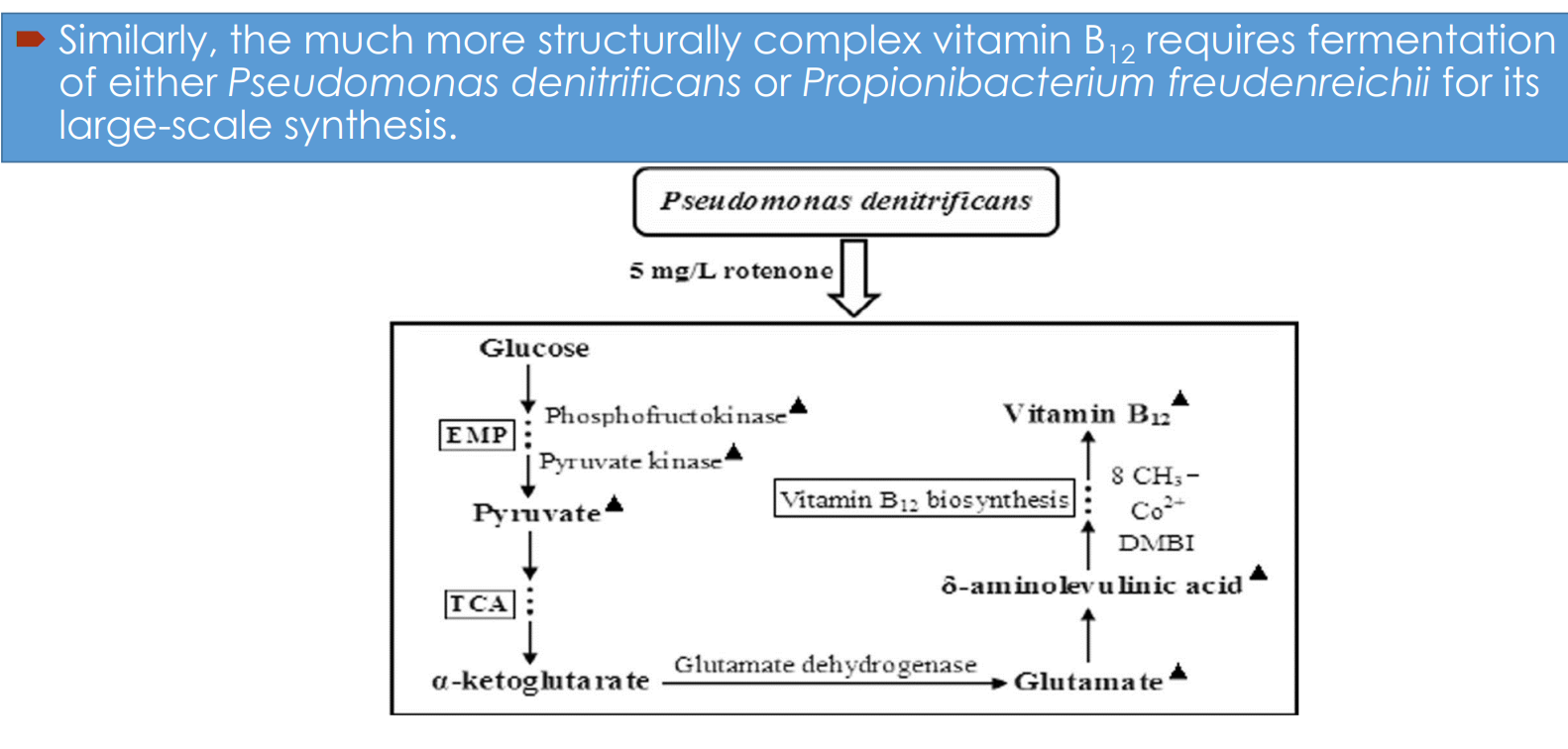
Vitamin B₁₂ fermentation pathway engineering: Link between TCA cycle and B₁₂ synthesis
α-ketoglutarate → glutamate (via glutamate dehydrogenase)
Glutamate feeds into:
→ δ-aminolevulinic acid (ALA)ALA is a key precursor for:
→ tetrapyrrole pathway → ultimately vitamin B₁₂Insight:
→ B₁₂ production depends heavily on amino acid metabolism, not just central carbon
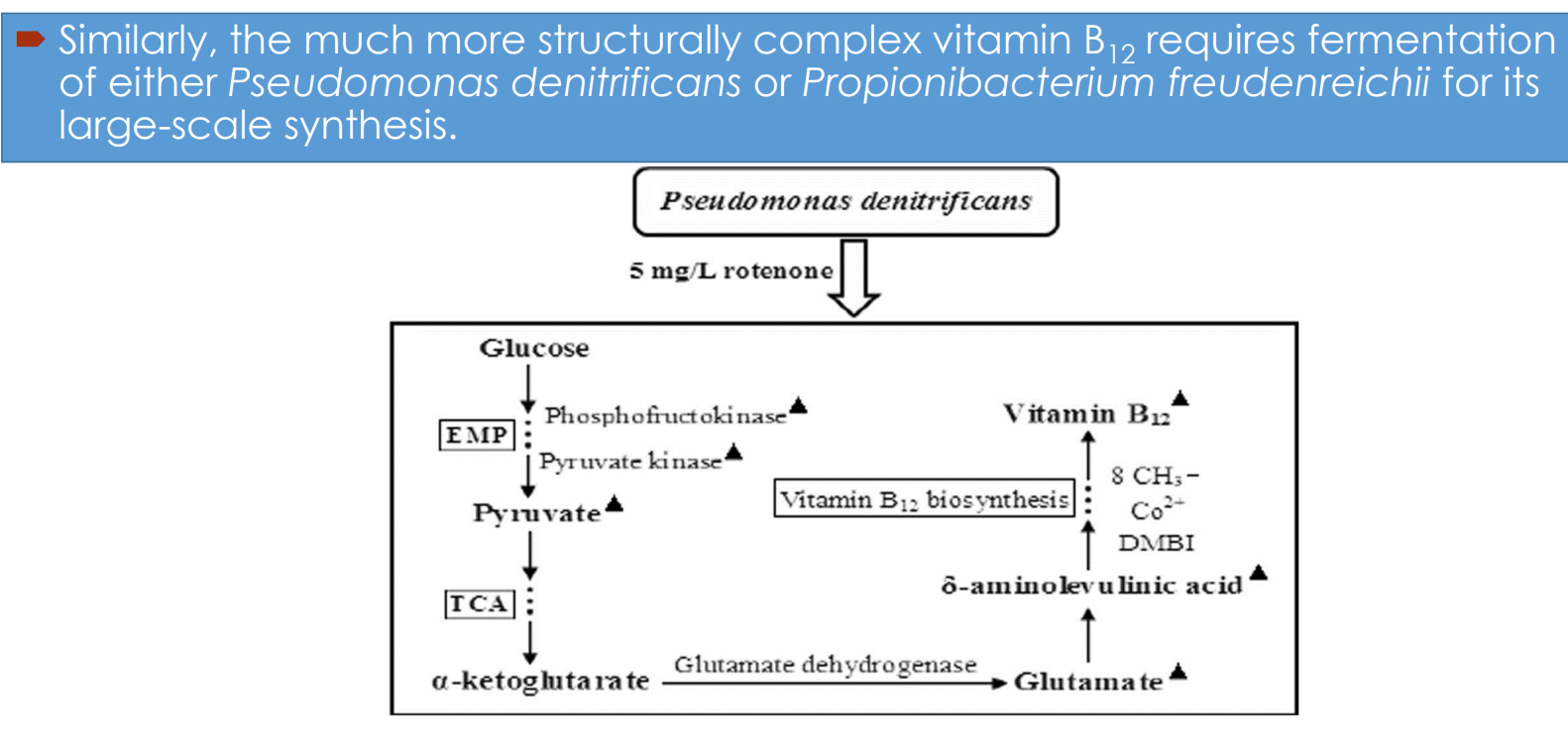
Vitamin B₁₂ fermentation pathway engineering: Tetrapyrrole pathway and corrin ring formation
δ-aminolevulinic acid → multi-step pathway → corrin ring
Requires:
Co²⁺ (cobalt insertion)
methylation steps (CH₃ groups)
DMBI (lower ligand base)
Bottleneck:
→ >30 enzymatic steps → very high metabolic burden
Vitamin B₁₂ fermentation pathway engineering: Cofactor and metal dependence
Cobalt (Co²⁺) is essential:
→ directly incorporated into B₁₂ structureLimitation:
excess cobalt → toxicity
low cobalt → reduced yield
Industrial control:
→ tightly regulated metal feeding during fermentation
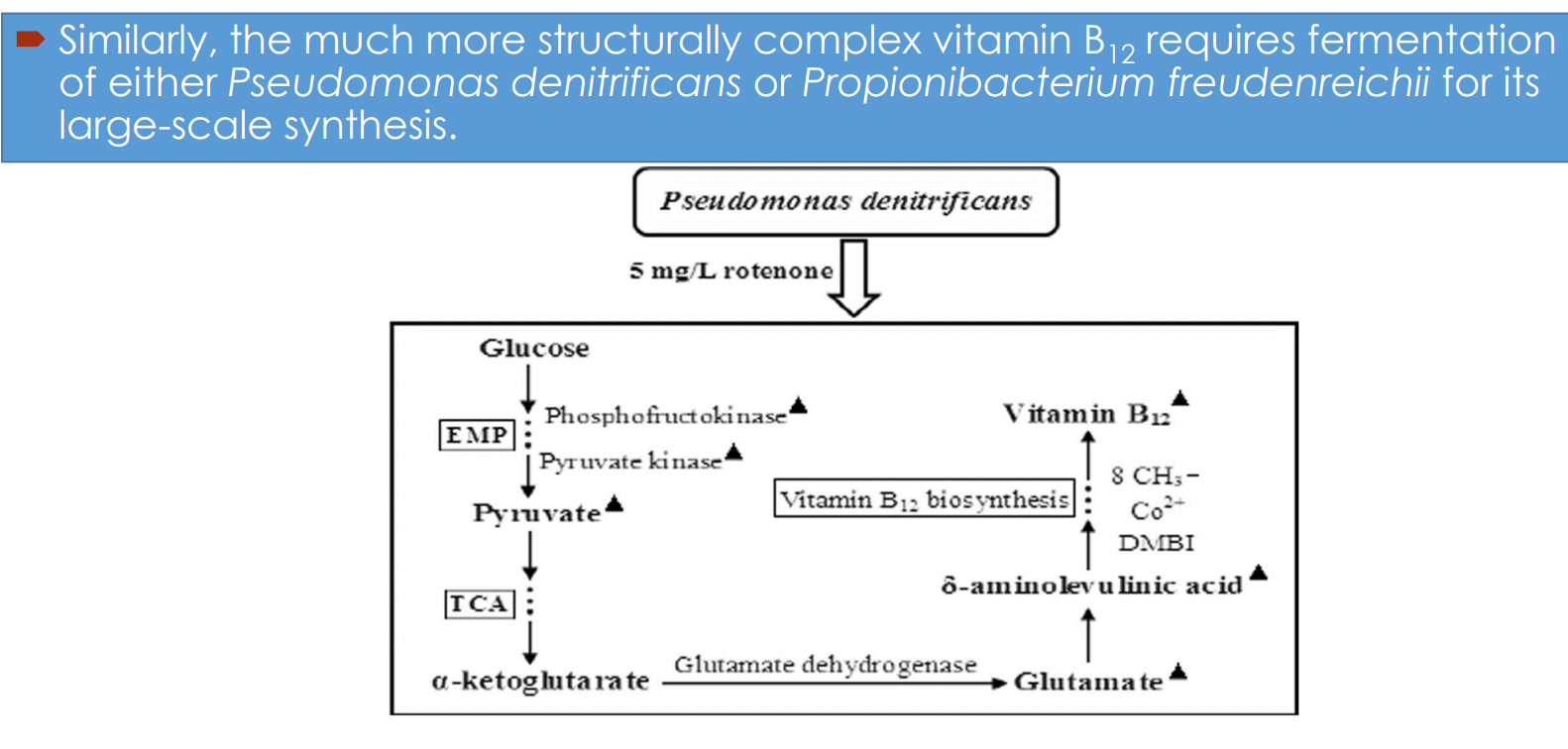
Vitamin B₁₂ fermentation pathway engineering: Process-level intervention (rotenone example)
Addition of compounds like rotenone:
→ inhibits competing respiratory pathwaysEffect:
redirects energy and reducing power toward B₁₂ synthesis
increases precursor availability
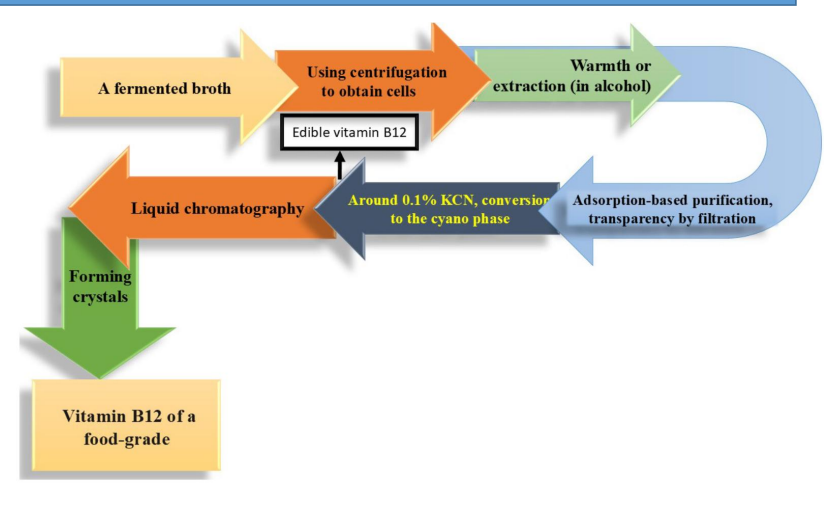
Vitamin B₁₂ downstream processing and purification diagram
Fermented broth → cell recovery
Start with fermentation broth containing cells + B₁₂ (mostly intracellular)
Centrifugation separates biomass from liquid
Extraction (warmth or alcohol)
Cells are treated with:
heat
alcohol (e.g., ethanol)
→ cell lysis + release of B₁₂ into solution
denatures unwanted proteins
reduces viscosity → easier downstream handling
Adsorption-based purification + filtration
Crude extract contains pigments, proteins, other cofactors
Use adsorption resins:
→ selectively bind B₁₂Followed by filtration:
→ improves clarity (“transparency”)
Conversion to cyano form (KCN treatment)
~0.1% KCN added to convert B₁₂ into:
→ cyanocobalamin (stable form)Why:
native forms (adenosyl-, methyl-B₁₂) are unstable
cyanocobalamin is:
more stable
easier to purify
preferred for supplements
Liquid chromatography → fine purification
Removes remaining impurities:
analogs of B₁₂
incomplete intermediates
Crystal formation
Final purified B₁₂ is:
→ crystallizedBenefits:
high stability
easy storage and transport
precise dosing
Xanthan gum production: Feedstock utilization
Uses moist olive pomace and other agro-wastes (e.g., waste bread + enzymes) as carbon sources
These substrates contain:
sugars
phenolic compounds
Integrated effect:
→ reduces raw material cost and improves sustainability
→ phenolics can also influence polymer structure and antioxidant propertiesConstraint:
→ variability in waste composition → inconsistent fermentation performance
Xanthum production species
Core production organism
Xanthomonas campestris (ATCC 33913)
→ primary industrial strain used for xanthan gumReason:
naturally secretes xanthan extracellularly
high yield and stable production
well-characterized gum structure
Other Xanthomonas species shown
Xanthomonas axonopodis pv. vesicatoria
Xanthomonas hortorum pv. pelargonii
Xanthomonas axonopodis pv. begoniae
These are typically:
→ plant-pathogenic variants used in research or strain screeningRelevance:
alternative strains may produce xanthan with:
different viscosity
different branching patterns
useful for tailoring functional properties
Xanthan gum production: Microbial production system
Produced by Xanthomonas campestris (ATCC 33913)
Other related strains also used depending on application
Typical conditions:
~28°C
agitation (~250 rpm)
~72–76 hours fermentation
During fermentation:
→ bacteria secrete xanthan gum extracellularly into the mediumAdvantage:
→ easier recovery compared to intracellular products
Xanthan gum production: Polymer structure and functional properties
Xanthan is a heteropolysaccharide with:
glucose backbone
side chains containing mannose + glucuronic acid
Phenolic interactions (shown in slide):
→ can enhance:antioxidant activity
functional properties
Key property:
→ extremely high viscosity even at low concentrations
Xanthan gum production: Yield and viscosity optimization
Fermentation parameters strongly affect:
gum yield
viscosity (functional quality)
Variables include:
carbon source concentration
inoculum size
oxygen transfer
agitation
Trade-off:
→ conditions maximizing yield may not maximize viscosity
Xanthan gum production: Measured performance improvements
Slide indicates:
viscosity increase (~395%)
antioxidant activity increase (~179%)
xanthan yield increase (~50%)
Interpretation:
→ substrate choice and process tuning directly impact both quantity and quality
Xanthan gum production: Downstream recovery
After fermentation:
broth contains dissolved xanthan
Typical industrial recovery:
alcohol precipitation (ethanol/isopropanol)
drying to powder
Because xanthan is extracellular:
→ avoids costly cell disruption steps
Industrial applications of xanthan gum
Food industry
thickener (sauces, dressings)
stabilizer (prevents phase separation)
gluten replacement in baking
Oil industry
drilling fluid viscosity control
Pharmaceuticals
controlled drug release matrices
Xanthan gum production: process constraints
High viscosity during fermentation:
→ reduces oxygen transfer
→ creates mixing challengesRequires:
strong agitation
optimized aeration
Genetic engineering to modify antibiotics
Existing antibiotics are chemically altered via engineered biosynthetic pathways
Done by modifying:
polyketide synthases (PKS)
non-ribosomal peptide synthetases (NRPS)
Outcome:
→ new analogs with:improved activity
reduced resistance susceptibility
Example:
modifying erythromycin → azithromycin-like derivatives
altering glycosylation patterns in antibiotics
Constraint:
→ small structural changes can drastically affect:binding affinity
toxicity