GENE221 Mod 1 - Mutations
1/24
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
25 Terms
What is an auxotroph?
a micro-organism that can only grow in the presence of specific growth supplements such as amino acids
In the one gene-one enzyme experiment carried out by Beadle and Tatum, what steps were taken to find loci (genes) in which mutations caused arginine auxotrophy?
They placed a mutant of an organism (Neospora), and determined that it was auxotrophic for arginine by trying to cultivate 20 colonies of Neospora on minimal media + 1 amino acid. The only colony that grew was in minimal media +arginine, indicating the mutant was arginine auxotrophic.
They carried out a detailed study of compounds that would enable the auxotrophs to grow, identifying three gene mutations that caused loss of function of the synthesis of three different compounds in the arginine synthesis pathway, thus having three classes of mutants: arg-1, arg-2, and arg-3.
In the one gene-one enzyme experiment carried out by Beadle and Tatum, what steps were taken to determine what different mutants arg-1, arg-2 and arg-3 caused?
The three mutants were exposed to minimal media + three different compounds, either arginine or one of its precursors in the biosynthesis pathway, and the following results were found.
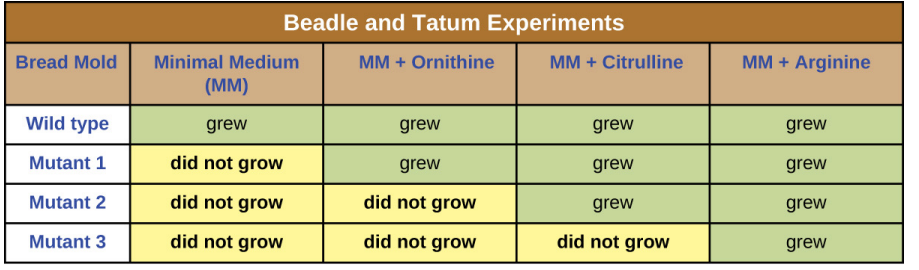
In the one gene-one enzyme experiment carried out by Beadle and Tatum, what did they conclude from their results of growing arg-1, arg-2 and arg-3 mutants in different media?
Each gene arg-1, arg-2 and arg-3 gives rise to a different enzyme that catalyses a different chemical reaction.
This formed the basis of the one-gene-one-enzyme hypothesis: “each gene in an organism corresponds to a single enzyme, and vice-versa ”
how did the fluctuation test show that mutations arise spontaneously in a population, and are not due to the presence of phage inducing some of the bacteria to become resistant (Induced mutation).
If induced mutation was the cause of antibiotic resistance, we would expect resistance to arise naturally within all or many different unique bacterial environments at a reasonably similar rate. If resistance arose spontaneously in a population, it would be a genetic mutation that would be passed on and as such, resistant bacteria may explode in population. This being a small chance, some cultures would have no bacteria, and some would have lots, thus there would be a large fluctuation in bacteria colonies that grow in a given culture.
Upon performing the test, a massive fluctuation in bacteria colony numbers was found, and so spontaneous mutation was ruled as the mechanism of mutation in organisms.
Define Mutagen
an agent that is capable of increasing the mutation rate
How does the Ames test allow detection of mutagens
uses strains of a bacterium Salmonella typhimurium that are AUXOTROPHS - have a mutation in a gene required for synthesis of histidine, an amino acid (His- phenotype; hisC or hisG genes)
• Bacteria are spread onto a growth medium that does not contain histidine
• Any bacteria that can grow have had a second mutation and are revertants - have reverted (gone back) to being able to make Histidine
• If the test compound increases the number of revertants it is likely to be a mutagen
How was the Ames test improved to make it more appropriate for testing mutegenicity on mammals?
many chemicals that are not mutagenic to bacteria are mutagenic in mammals, because these chemicals are converted into an active form in mammals by enzymes in the liver, after they have been ingested
• solution: prepare liver extracts and incorporate them into the medium used in the Ames test
What is way of strengthening the Ames test generally?
Added features of the bacterial test strains:
(i) Mutation in a gene (rfa) that affects the cell envelope - makes the bacteria more permeable to some chemicals
(ii) Defective in a gene (uvrB) encoding a protein that repairs damaged DNA and reduces the frequency of mutations
(iii) Contain a plasmid (pKM101) that enhances the effectiveness of some mutagens
ALL OF THESE INCREASE THE EFFECTIVENESS OF MUTAGENS IN CAUSING MUTATIONS
What are three common types of mutations?
1. point mutations = mutations at a single point (!) (most often a single base pair) in a genome – SNPs (Single Nucleotide Polymorphisms)
2. can be substitution mutations or indels (insertion or deletion of base-pairs)
3. can also be deletion (loss) or large-scale rearrangements of DNA or insertion of mobile genetic elements
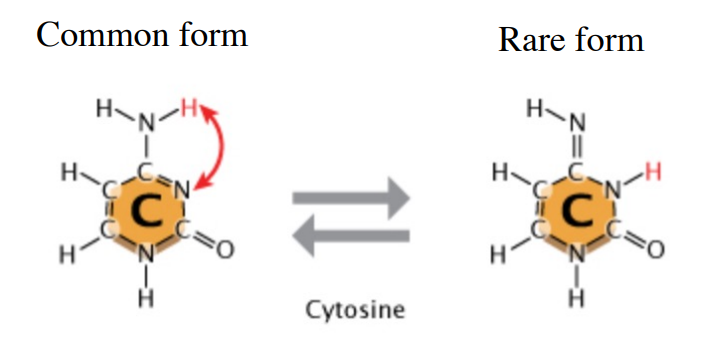
How does tautomerisation affect DNA replication/cause mutations?
Tautomerisation can occur, where a cytosine molecule switches to a rare tautomer. This different kind of cytosine bonds to Adenine instead of Guanine. (Goes from 3 H-bonds to 2). This causes a GC > AT mutation.
How are DNA point mutations fixed during or after DNA replication?
Both DNA pol I and DNA pol III serve a “proofreading” function by excising incorrectly inserted mismatched bases.
Once the mismatched base is removed, the polymerase has another chance to add the correct complementary base.”
What is another mechanism that fixes point mutations in DNA
Mismatch repair (MMR).
The first step in MMR is the detection of mismatches in newly replicated DNA by the MutS protein.
Binding of MutS to distortions in the DNA double helix recruits MutL and MutH.
MutH cuts the newly synthesized strand containing the incorrect base.
Without the ability to discriminate between the parental and newly synthesised strands, the MMR system could not determine which base to excise.
How does the MMR system differentiate between the original and new strand in E.coli?
DNA inside E. coli is chemically modified by methylation of adenine bases in the DNA.
Strand recognition by MutH is directed by adenine methylation at GATC sequences.
Because adenine methylation occurs after DNA synthesis, newly synthesized DNA is temporarily unmodified, and this temporary absence of methylation directs repair to the new strand.
The MutH endonuclease cuts the unmethylated strand
What is a random chemical event that occurs on cytosine, causing a GC > TA mutation?
Deamination of Cytosine.
Deamination = the hydrolytic removal of an amine group
Deamination converts cytosine to uracil. Uracil base pairs with adenine in replication, converting a C · G base pair into a T · A base pair.
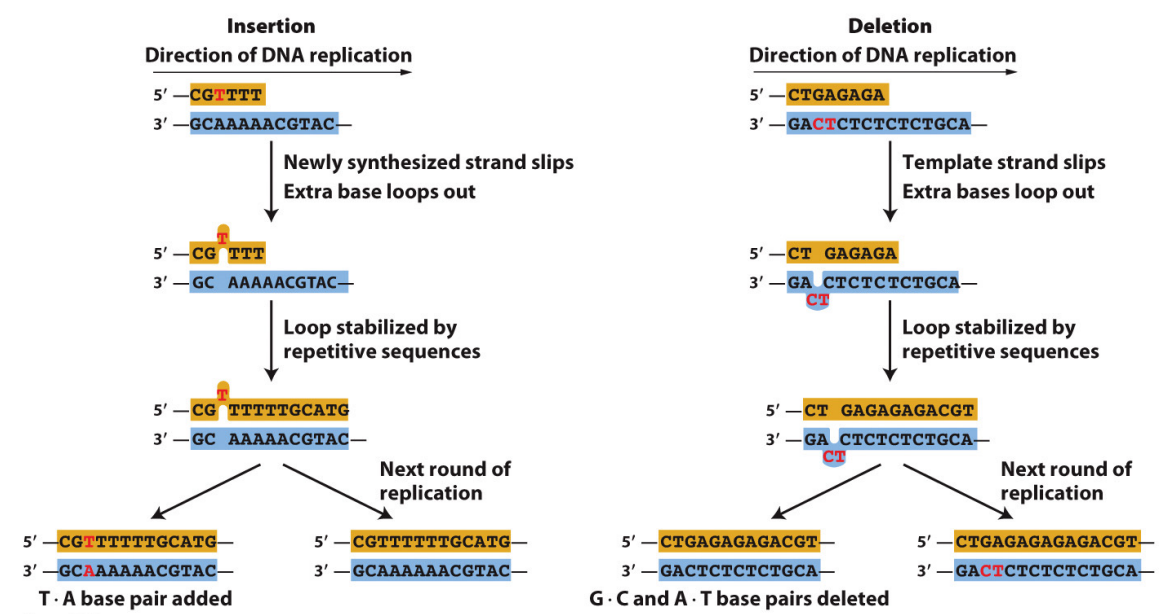
What are the two main ways that InDel mutations can occur?
Base insertions and deletions (indels) are also caused by DNA replication errors.
Indels arise when loops in single-stranded regions of DNA are stabilized by the “slipped mispairing” of repeated sequences in the course of DNA replication.
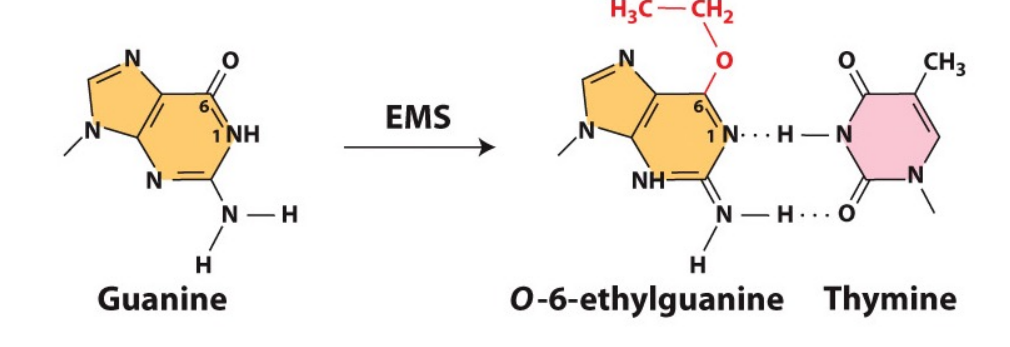
How does the chemical mutagen EMS (ethyl methane sulphonate) cause mutations? And how are these potential mutations corrected?
EMS is an alkylating agent that changes Guanines configuration to bond with Thymine, causing GC → AT mutations.
Repair proteins – alkyltransferases. The (alkyl) ethyl group is transferred from the base in the DNA to the protein
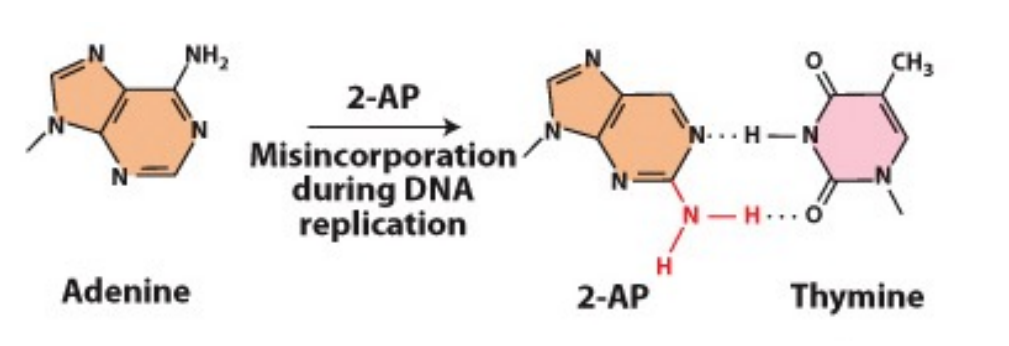
How does 2-AP (2-amino purine) cause mutations?
- Most often pairs with thymine but can also base-pair with cytosine
- If this happens during replication, will lead to an A-T basepair being changed to A-C and subsequently, G-C
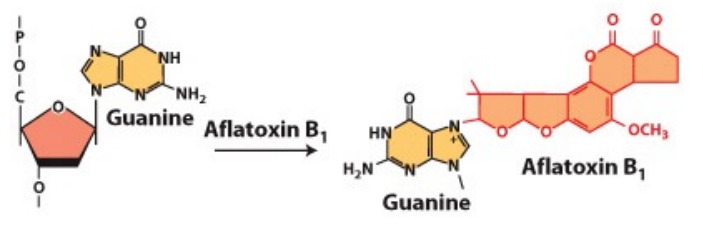
How does Aflatoxin cause mutations?
•produced by fungi
•chemically reacts with guanine (G) bases in DNA, generating apurinic sites. This can lead to mutation
•causes GC →TA mutations
What are apurinic sites and why do they cause mutations?
• Depurination - a purine base (adenine or guanine) is lost from the DNA
• This gives an apurinic site, ie. a site without a purine
• during DNA replication, there is a "blank" where the purine should be
• a base (often an adenine) may be inserted opposite the blank
• this can change the sequence of base-pairs
How does radiation cause mutation?
When hit by a high-energy UV ray, adjacent thymidine (T) bases in the DNA can become covalently crosslinked - photodimers •
These fail to base-pair properly during DNA replication
• Translesion DNA polymerases (bypass DNA pol) can replicate past these but may incorporate the wrong base as they have significantly less error-correction
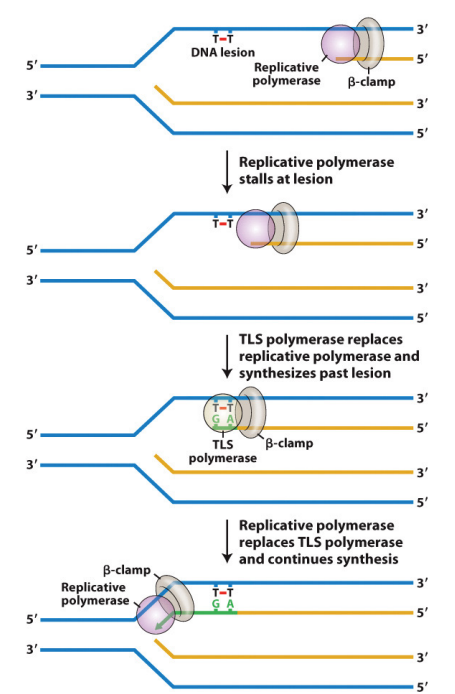
How does Trans-lesion DNA synthesis occur?
It is initiated by stalled DNA polymerase, which triggers the recruitment of a TLS polymerase that synthesizes past the lesion.
Once extension passes the lesion, the TLS polymerase is replaced by the replicative DNA polymerase.”
Conserved from E. coli to humans.
What is a pre-mutagenic lesion and how do they effect mutation?
a change to the DNA that may lead to a mutation.
• At least one round of DNA replication is needed to get from a pre-mutagenic lesion to a mutation
• DNA repair takes place at the pre-mutagenic lesion - once the mutation is "established", it is too late
• DNA repair systems repair “damaged” DNA BEFORE it is replicated
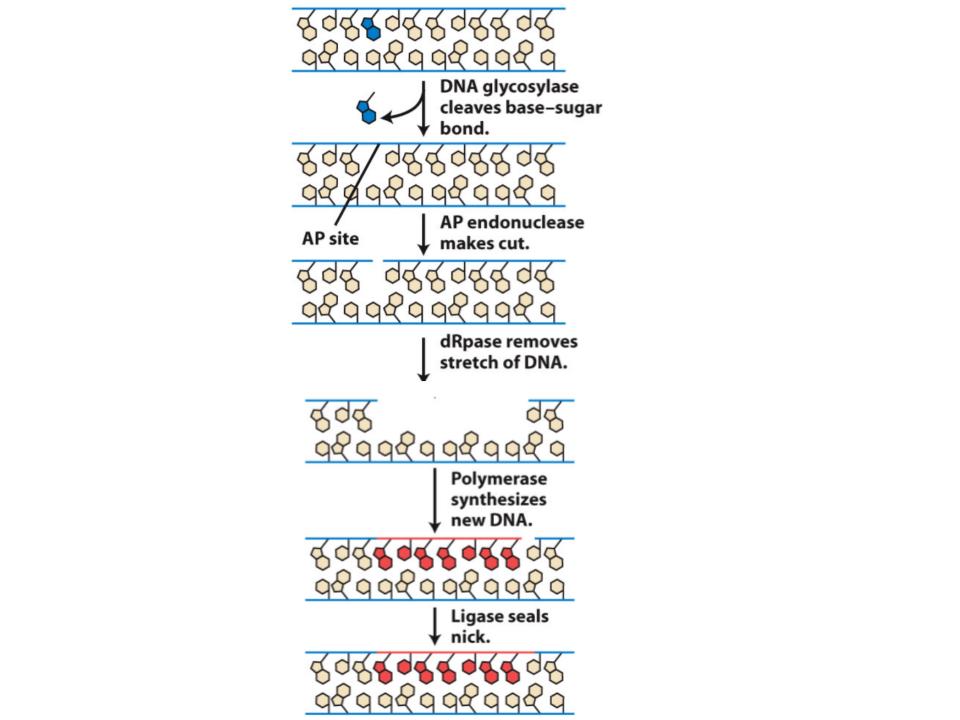
How are apurinic sites in DNA error-corrected?
An enzyme (AP endonuclease) recognises an apurinic site and cuts the strand of DNA that contains it. The defective DNA and some adjacent DNA is then removed by a set of enzymes (excision exonucleases). The gap is filled in by DNA synthesis.
How are photodimers caused by radiation error-corrected so as not to cause mutations?
Multiple mechanisms are known e.g
Nucleotide excision repair similar to repair of apurinic sites (Fig. 15-16 in Griffiths et al 12th ed).
Photolyase enzyme - this uses energy from white light to convert photodimers back to pyrimidines (photorepair)