lecture 25: genomic variation and forensic DNA fingerp
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
a true wildtype human genome does not exist
true
we only have refrence sequences which has just as much variability
DNA polymorphisms
two or more variant forms of a specific DNA sequence amongst individuals/populations that are unique to each
even thought theres many variation, most polymorphisms DO NOT influence
phenotype
reasons
codons make up 2% of the human genome (little actually codes for proteins) MEANING most poly.. aren’t in a protein coding region
many mutations in codons don’t change the amino acid (even though these are within protein coding region) MEANING the base changes but it doesn’t alter the amino acid its coding for silent mutation
many deleterious mutations disappear from the population through natural selection (mutation that cause lethality & prevent formation into a baby; doesn’t get to pass on to next generations)
4 categories of genetic variation:
single nucleotide polymorphisms - one base pair changes (within DNA)
deletion-insertion polymorphisms (DIPs) - short insertions or deletions of a single or a few base pairs in genetic material
simple sequence repeats (SSRs) - blocks ranging from 1-10 base sequence repeated fewer than 10 to more than 100 times in tandem
copy number variants (CNVs) - large blocks og genetic material up to 1Mb in length tha are variable in copy number in the genome
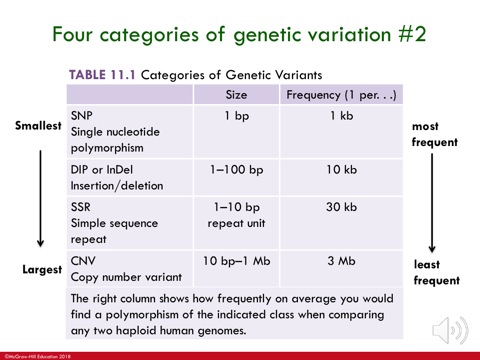
affecting fewest # ——> affecting most # of DNA bases
fewer # of bases that it would affect = the more frequently it is to occur in the genome
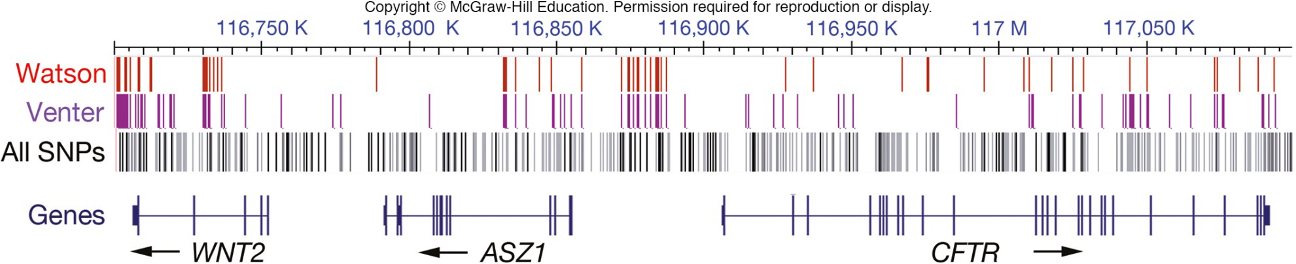
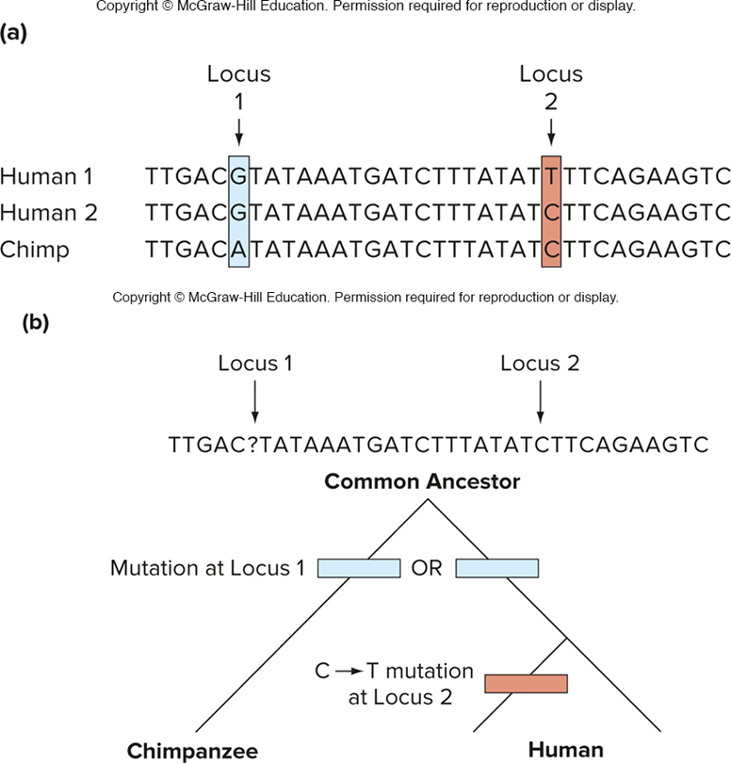
comparison of 2 human genomes to chimpanzee genomes (who evolved into humans at some point) reveals the SNPs that occurred since divergence of these species
the second single base change is polymorphic in humans
the C must be ancestral
the T is derived (derived single nucleotide polymorphism in 1st human genome) occurred AFTER humans evolved from chimps
(look at a closely related species that branched off b4 the group being studied (chimps) this is the outgroup! if the outgroup & some member of the study group share the same gene, the “A” is ANCESTRAL)
how can Deletion-insertion polymorphisms (DIPs) occur? these are biochemical processes that contribute to DIP formation:
problems in DNA replication (DNA got copied wrong).
problems in recombination
mistakes in DNA repair
DIPs cause frameshift mutations unless the number of nucleotides thats inserted or deleted is 3 or a multiple of 3 (delet/insert of 1 or 2 bases has a deletorious affect)
facts abt Simple sequence repeats 9(SSRs) aka microsatellites
-one to a few bases that are repeated fewer than 10 to more than 100 times
most common repeating units are 1,2,or 3 base sequences
SSrs account for about 3% of the total DNA in the human genome (can use as it is repeated, to match its DNA to another DNA sample
arise spontaneously from rare, random events, expanded by slipped mispairing or stuttering
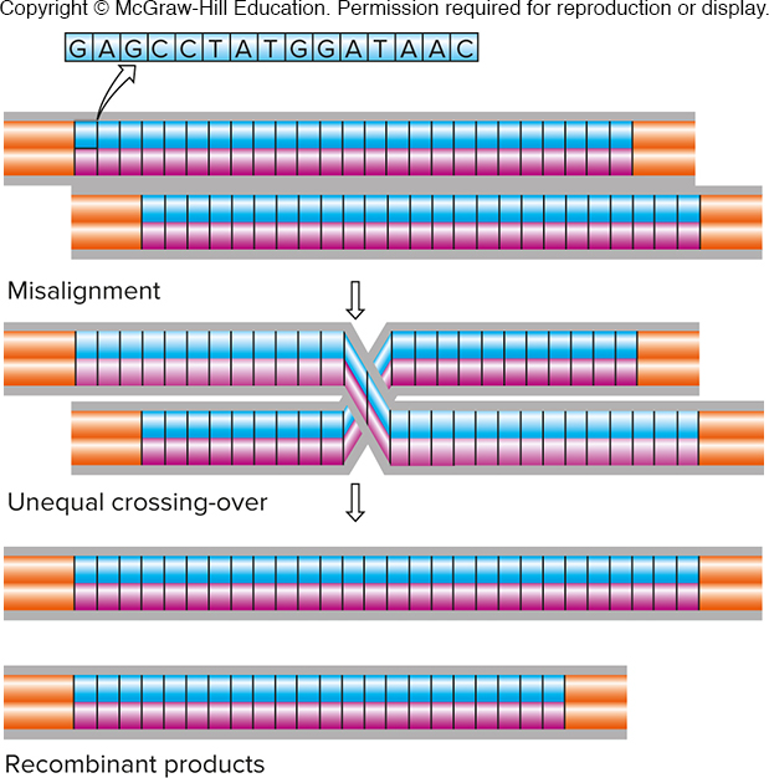
unequal crossing over produces new alleles of
copy number variants (CNVs)
-tandem sequence repeats more than 10 bp long
-misalignment during meiosis leads to unequal crossing-over
not a common evnt, so most CNVs are inherited, rather than being a new mutation
imgae - expansion in 1; contraction in the other
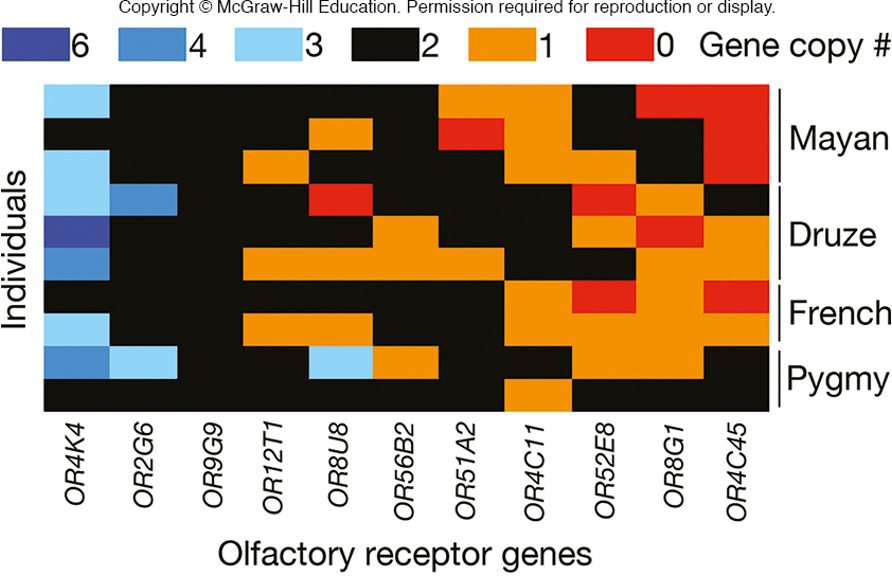
humans have fewer than 1000 olfactory receptor genes at different loci
at each locus, the copy number varies
image - 0 copies of that gene in the Mayan population i.e
TRUE
determining/genotyping genomic variations
amplify the gene region by PCR
sequence the gene region to identify polymorphisms (if SSr or CNV ; can look @ size of PCR product & determine genotype that way)
Polymerase chain reaction (PCR) is one of the most common methods for genotyping
method of making many copies of a target region of DNA (first developed in 1985)
fast and extremely efficient - can amplify DNA from a single or hair follicle
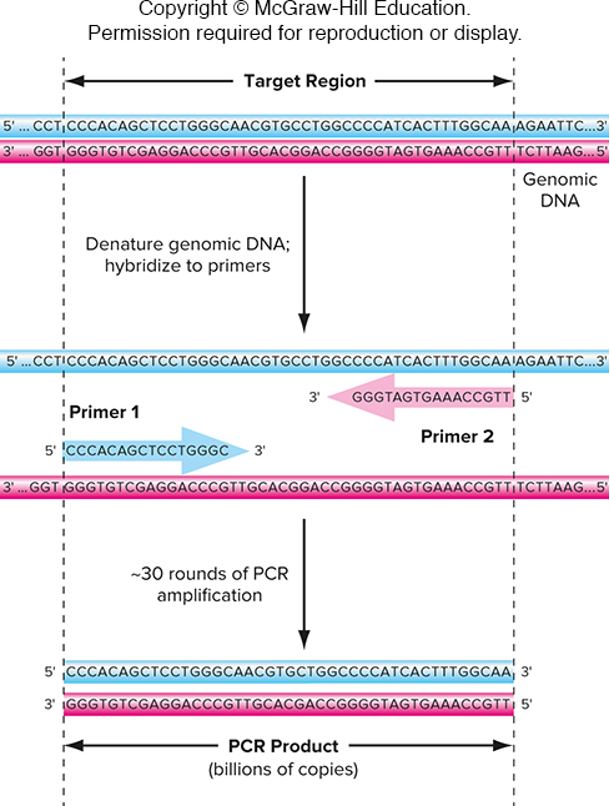
One primer is complementary to one strand of DNA at one end of the target region
The other primer is complementary to the other strand of DNA at the other end of the target region
true
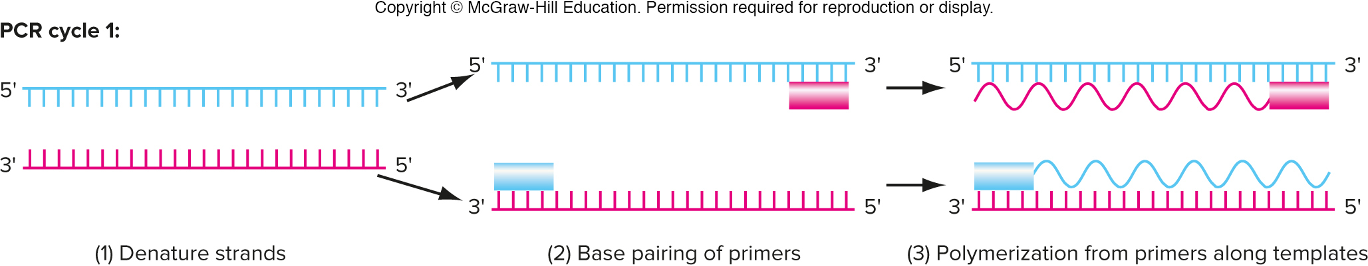
3 steps in PCR (basically replicating DNA)
denature strands (raise temp. to boil to split the 2 strands; this takes the place of DNA helicase) in test tube so we can boil
base pairing of primers (lower temp. to allow primers to anneal to their respective template strands; forward anneals to bottom vice versa) DNA primers takes place of RNA primers
polymerization from primers along templates (add in DNA polymeraze which then synthesizes the complementary strands) same DNA poly.. in PCR and DNA replication
same basic process of DNA replication; just has ways around extra enzymes PCR doesn’t need
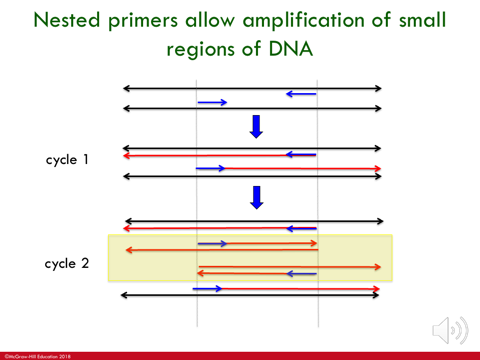
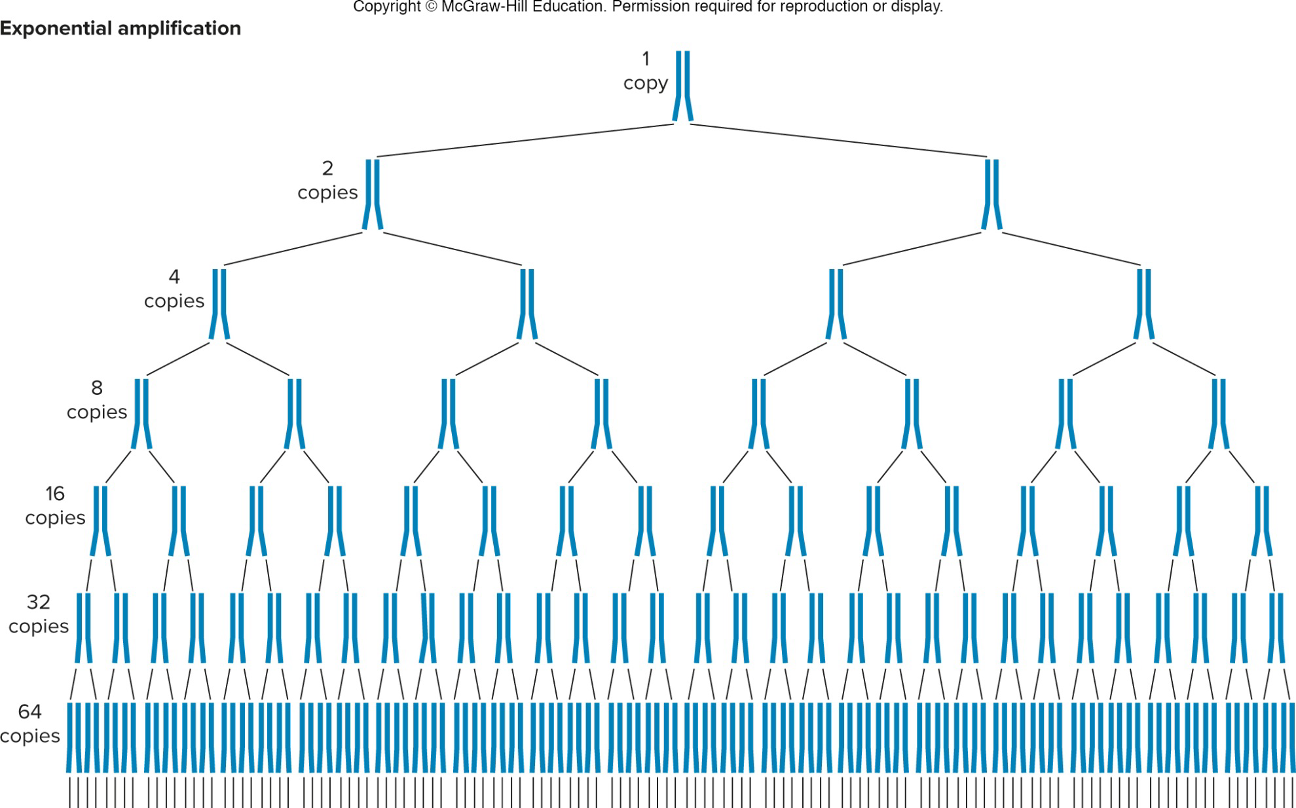
Cycles/rounds within PCR
cycle 1 : primer gets polymerized all the way until it looses steam; in reverse as well (get long products tht go beyond boundary site)
cycle 2 : the 2 newly synthesized strands act as template strands; each split and get a primer opposite to it which stops synthesizing at boundary line
products from cycle 2 are used as templates for next cycle & you get more and more products tht start & stop @ the boundary sites and fewer that goes beyond.
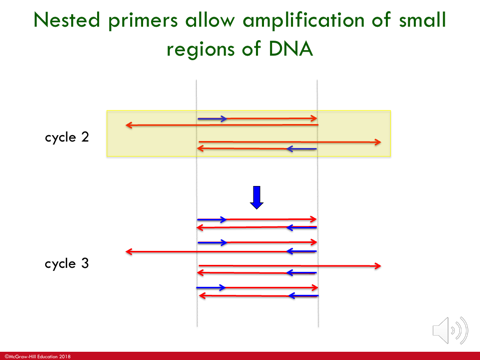
IN PCR, at each round you get amplification where you get double the amount of copies of DNA that serves as templates for next (target DNA increases)
*image- after 1st round you have 2 copies, after 2nd round you have 4 copies…
Using PCR to sequence a particular snip (determine genotype)
sickle cell anemia is caused by a SNP in the Hb beta gene.
genotyping can identify carriers and homozygous individuals.
hetero - 2 peaks at same base (double peak)
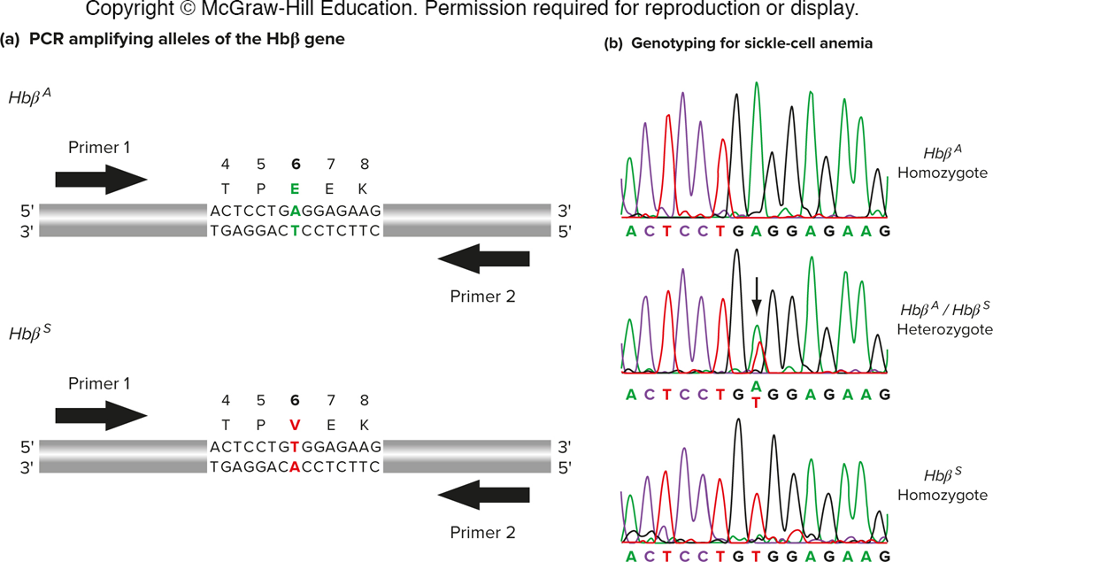
Determining genotype by PCR product size
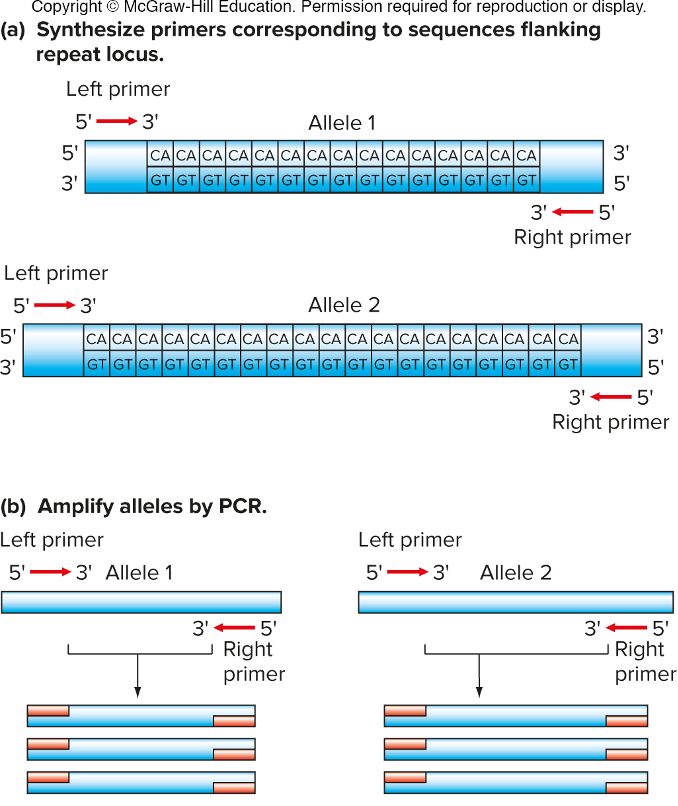
-target regions containing SSRs or DIPs can be amplified by PCR. the PCR products vary by size.
design primers upstream/downstream an amplify region in between that; look @ size of PCR fragment on DNA gel; more repeat is larger PCR

continue..
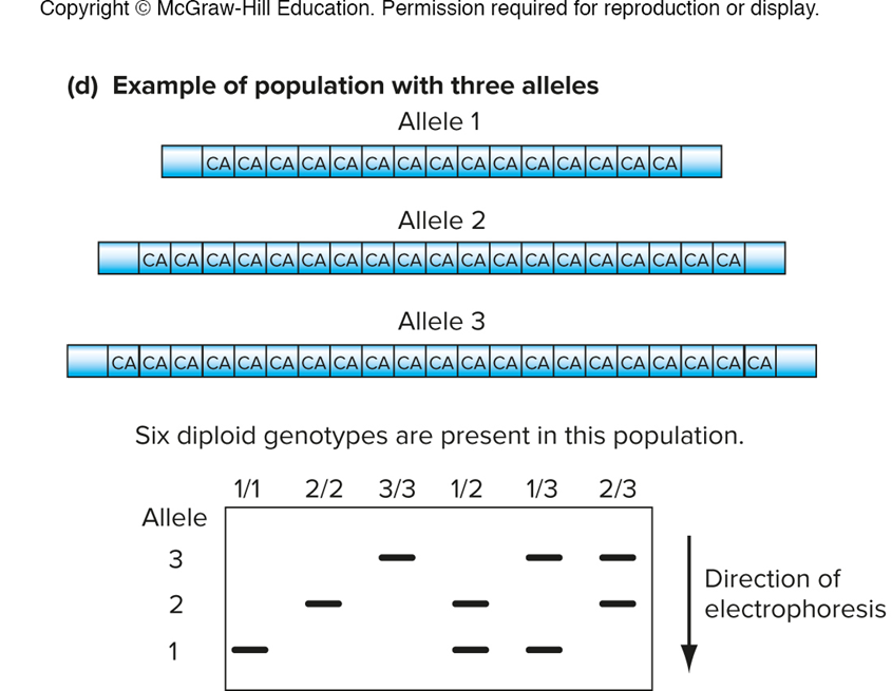
size variants can be detected by gel electrophoresis
1/2, 1/3, 2/3 - all heterozygotes bc they have 2 places on gel
image allele 1 is smallest so its palce in gel is at the bottom
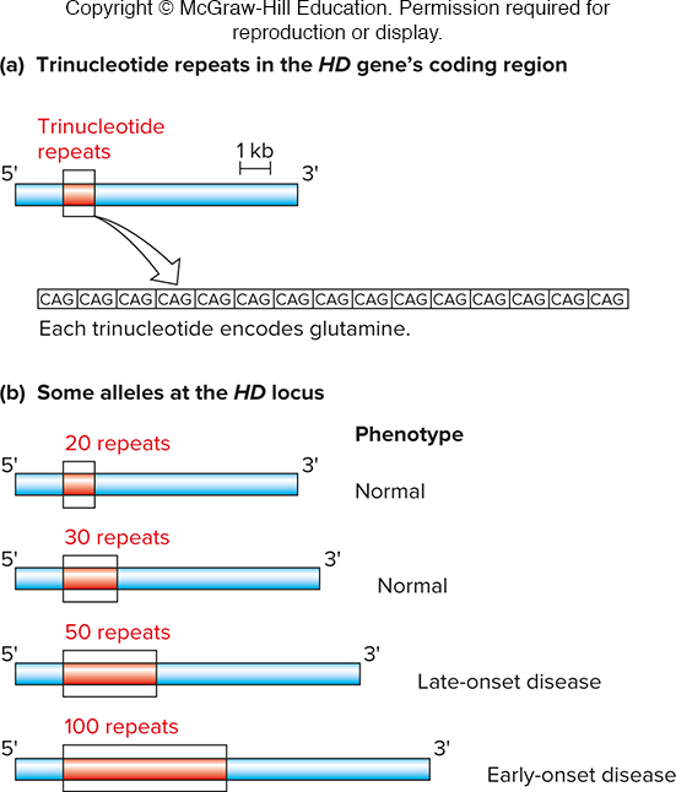
Hunington disease locus by PCR (cause by trinucleotide repeats)
normal alleles have max of 36 CAG repeats…
-however, disease-causing alleles have 36 or more CAG repeats; allels with an expansion with 42 or more repeats are completely penetrant (HT disease)
design primers outside of primers tht contain those repeats, look at size of PCR product to determine if disease allele is present
fetal and embryonic cells can be genotyped usigng PCR (docs did this b4 they implanted embryos)
Prenatal genetic diagnosis
•Genotyping fetal cells
•Cells isolated by amniocentesis—fetal cells in amniotic fluid are extracted using a needle.
Preimplantation embryo diagnosis
•Utilizes in vitro fertilization and PCR
Genotype embryos before placing in womb
the process of preimplantation embryo diagnosis
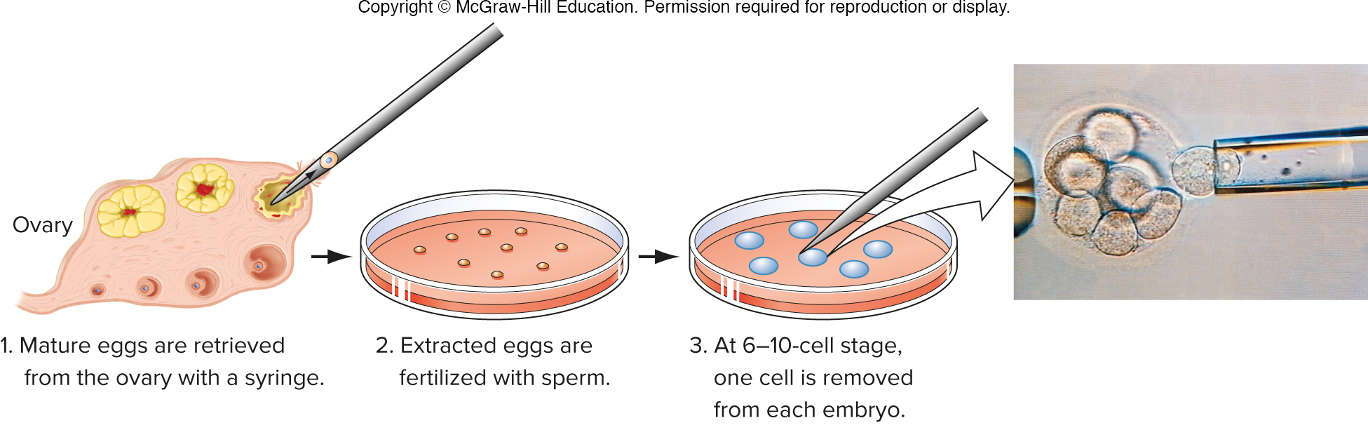
forensic DNA fingerprinting
SSR loci are highly polymorphic
•Many alleles exist in the population
•An individual person carries only two alleles at a given locus.
Genotype is determined by PCR at many SSR loci
•13 pairs of PCR primers are labeled with fluorescent dyes
•The probability that a person has the same alleles at 13 SSR loci is about 1 in 10 trillion (product rule)
CODIS database is maintained by the FBI
•Data from all 13 SSR loci
Data can match DNA from crime scene to a person, or can establish innocence.
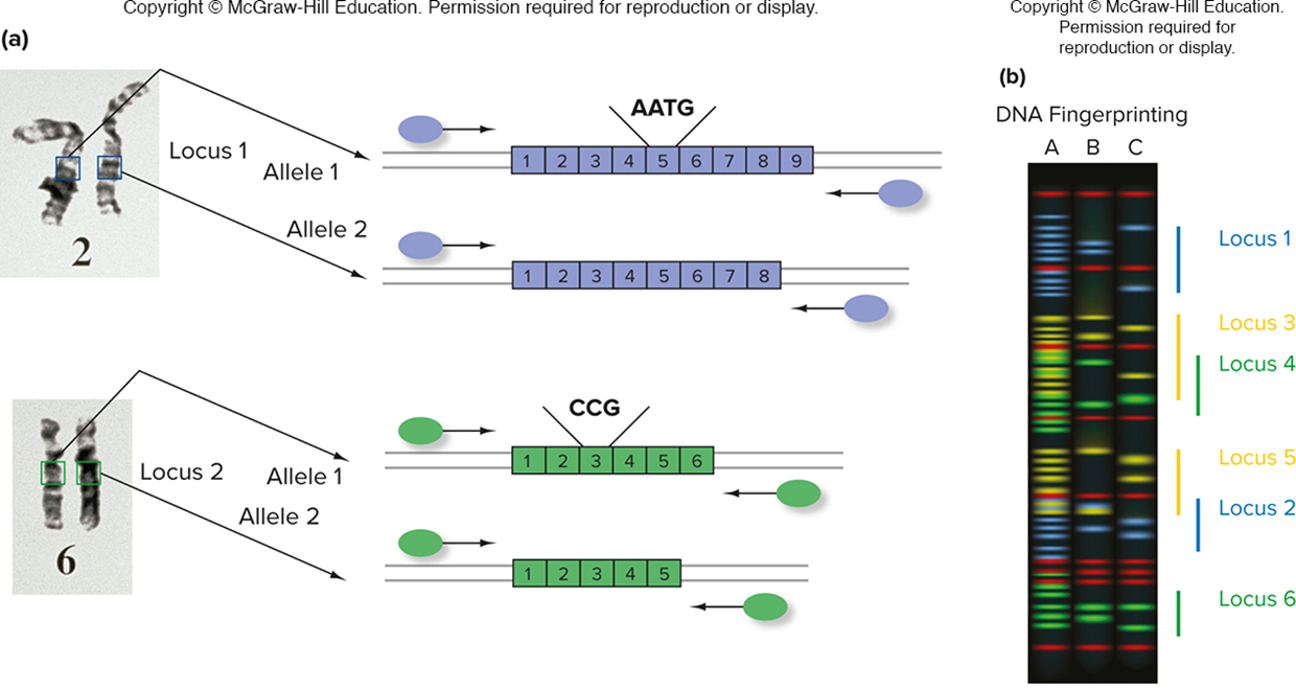
multiplex PCR is used for dna fingerprinting
running multiple and analyzing them; distinguish the multiple primers by the florescent; look for color and size of product to distinguish and get the alleles
when looking at gel, to determine relatedness…
each child should have 1 allele from the mother and 1 allele from the father
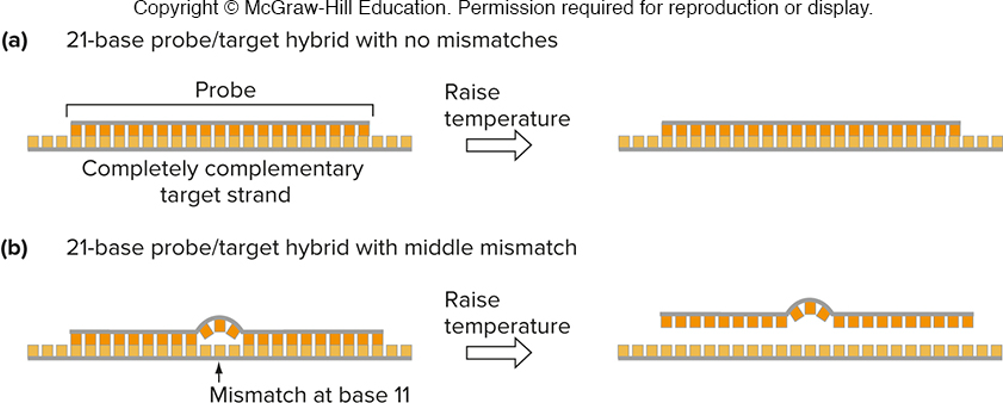
short hybridization probes can distinguish single-base nucleotide mismaches
true
hybridization of short (<40 bases) oligonucleotides to sample (target) DNAs (allele-specific hybridization)
if there is no mismatch between probe and target, hybrid will be stable at high temperature
if there is a mismatch between probe and target, hybrid will not be stable at high temperature
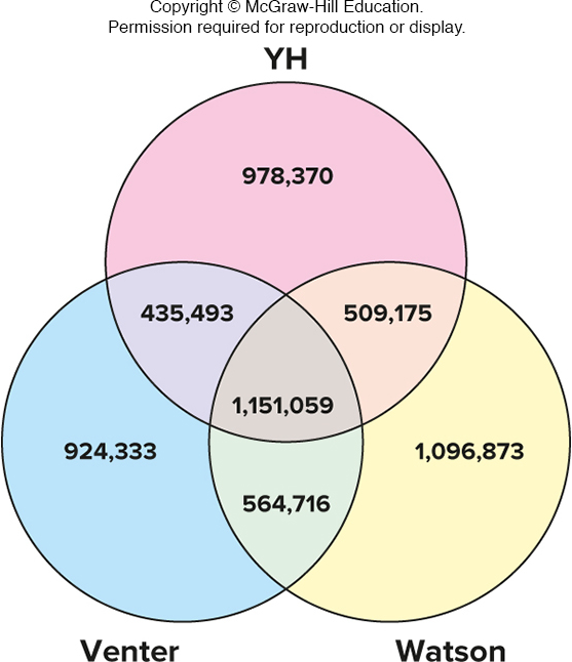
hybridization probes are used on microarrays for genotyping
true
allele-specific oligonucleotides (ASO) are attached to a solid support (like a silicon chip)
two oligonucleotides are shown here, but many can be put on one array
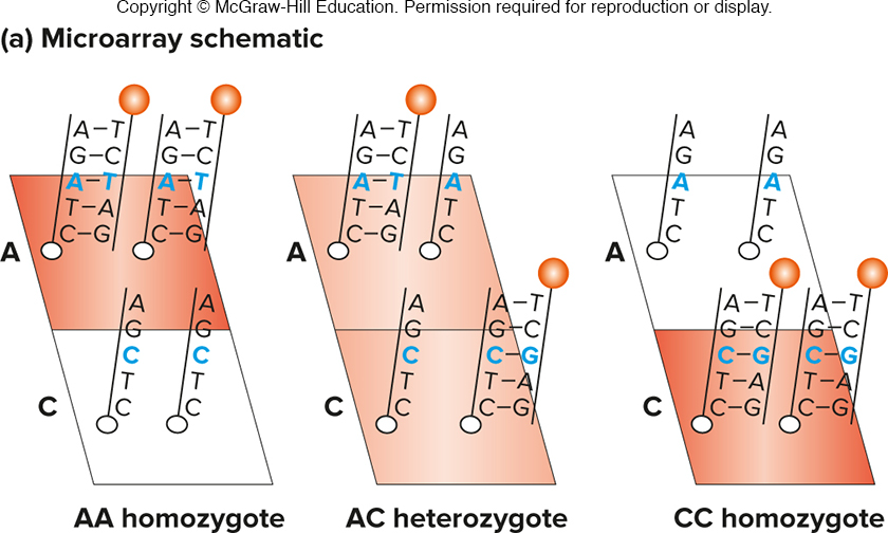
fluorescent output is proportional to the number of copies of each allele
*up to 4 million loci can be genotyped simultaneously"*