Lab #6: Bacterial Transformation, Plasmid DNA Isolation and Protein Extraction
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
Prokaryotes
Organisms made up of cells that lack a cell nucleus or any membrane-encased organelles.
Example: bacteria
Plasmids
Small, circular extrachromosomal DNA found in bacteria.
They replicate independently of the chromosome and generally result in phenotypic changes as required for survival in a hostile environment. (eg antibiotic resistance).
Plasmid DNA transfer occurs naturally in bacteria through a process known as transformation.
Transformation is easily performed in a lab and used in Recombinant DNA Technology.
Genome
The complete set of genetic information in an organism.
The genome is stored in form of long DNA molecules called chromosomes.
Genes are short segments of DNA containing the protein coding information and are located in the chromosomal DNA.
In higher organisms, genes are also present on mitochondrial DNA.
DNA
Deoxyribonucleic Acid.
DNA molecules are very long chains composed of repeating subunits known as nucleotides A (Adenine), G (Guanine), C (Cytosine), and T (Thymine). DNA is double-stranded.
The two strands containing complementary. (i) A pairs with T (ii) C pairs with G. The strands are antiparallel (opposite direction to each other). The 2 strands are twisted to form a helix.
DNA is also present in the eukaryotic mitochondria and in bacterial plasmids. Mitochondrial and plasmid DNA is called extra-chromosomal DNA.
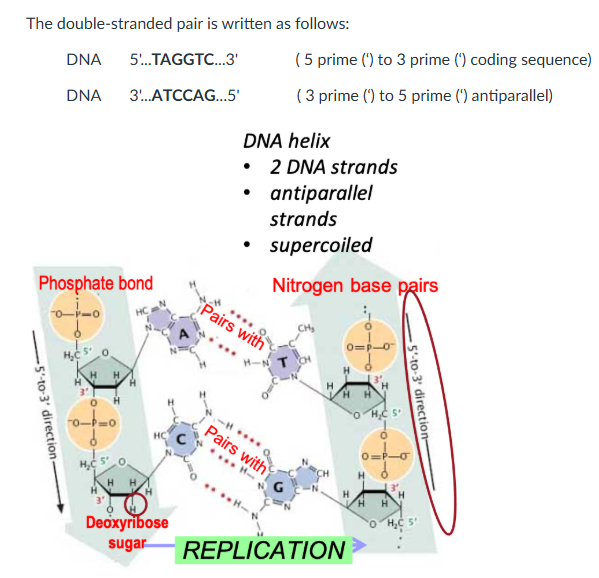
pGLO Plasmid
A recombinant pGLO plasmid.
It contains several genes and DNA sequences that enable replication of the plasmid DNA and differentiate and re pGLO transformed bacteria.
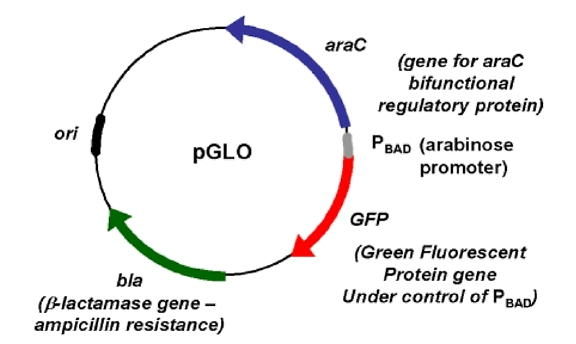
GFP
The jellyfish gene that codes green fluorescent protein and responsible for the green, fluorescent phenotype.
pBAD promoter
A specific DNA sequence upstream from the GFP gene, which binds araC-arabinose and promotes RNA polymerase binding and transcription of the GFP
bla gene
A gene that encodes the enzyme beta-lactamase and responsible for the antibiotic-resistant phenotype.
Beta-lactamase breaks down the antibiotic ampicillin and allows bacteria to grow in presence of ampicillin.
ori
The origin of pGLO plasmid DNA replication.
araC gene
A gene that encodes the regulatory protein that binds to the pBAD promoter.
Only when arabinose binds to the AraC protein is the production of GFP switched on.
araC protein
The regulatory protein encoded by the AraC gene.
The AraC protein binds to the pBAD promoter.
Arabinose
A sugar that binds to the pBAD promoter and displaces the AraC protein from the pBAD promoter so the RNA polymerase can bind and initiate transcription.
Only when arabinose binds to the AraC protein is the production of GFP switched on.
Ampicillin
An antibiotic that kills bacteria. It is destroyed by the enzyme beta-lactamase (encoded by the bla gene).
Genes encoded other antibiotic destroying enzymes may be used when designing the recombinant plasmid.
For example, Neomycin phosphotransferase II gene product, Neomycin phosphotransferase II degrades neomycin. In this case, Neomycin is used for selecting the bacteria transformed with the respective plasmid.
Transformed bacteria
Bacteria which survive/grow in presence of ampicillin.
These transformed bacteria can be selected from the non-transformed by growing in medium containing ampicillin.
Recombinant DNA technology
Methods used to modify the genetic properties of an organism. Recombinant as DNA from one organism (foreign) to the bacteria or yeast is inserted into their plasmid (aka DNA Vector)
The recombinant plasmid is transformed in cells.
The cells are grown in media to produce more plasmids or the gene product (proteins).
Recombinant DNA technology includes:
1) Synthesis of a recombinant plasmid - DNA fragments (genes) are cut and recombined to create a plasmid. Besides the gene(s) of interest, a plasmid must contain DNA for replication known as the origin of replication sequence, and an antibiotic-resistant gene for survival in an antibiotic-containing media. Other genes for controlling (regulating) the transcription of the gene products (proteins) may also be included.
2) Transformation of the organism – The plasmid is inserted in bacteria cell.
3) Selection of the transformed bacteria: The transformed cell is grown in an antibiotic-containing media so that only bacteria cells containing the plasmid will grow and non-transformed cells will die.
4) Synthesizing the gene product (protein of interest) – The cells are grown in media that contain molecules required for producing the protein.
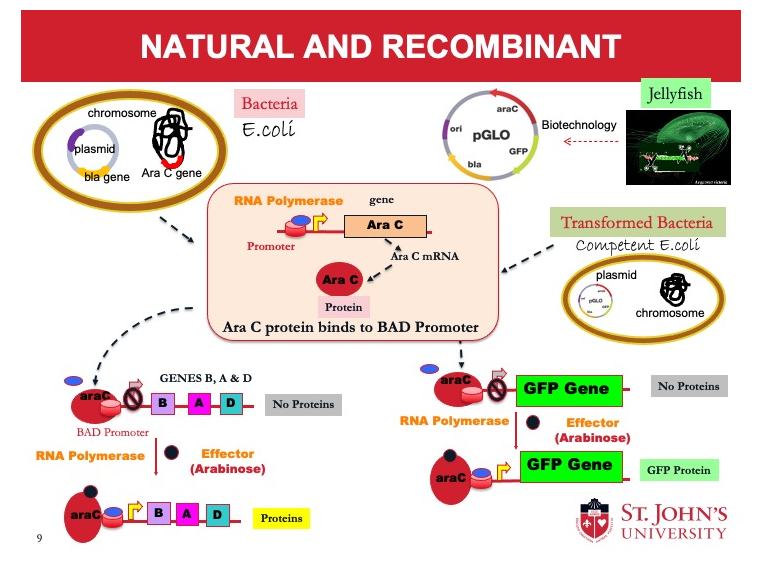
Competent Cells
The cells are capable of taking in the foreign DNA.
They are treated with calcium chloride (CaCl2) to make them competent. The positive charge of Ca++ ions shields the negative charge of DNA phosphates. This makes it easy for DNA to enter cell through the cell membrane.
Competent cells are kept frozen until use. Glycerol prevents formation of ice crystals inside the cells and increases their survival.
E. coli strain used for transformation is derived from pathogenic E. coli and generally non- pathogenic, however they must be handled and disposed similar to other common pathogens.
They are unable to produce enzymes that metabolize arabinose, are not ampicillin resistant and do not have GFP as they DO NOT contain the pGLO plasmid (with bla, and Ara C and GFP genes).
Hence, they cannot grow on media containing ampicillin and exhibit green fluorescence. When L (+) arabinose is added to culture media, the transformed cells, produce GFP and the transformed colonies appear green.
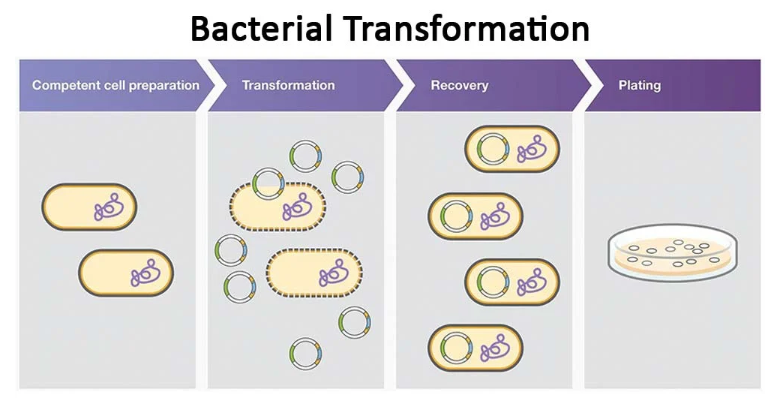
Bacterial Transformation
A method to introduce plasmid DNA in bacteria.
To uptake the foreign DNA, the bacterial cells are made competent to receive the highly negatively charged DNA. Then the bacteria are shocked using hot/cold treatment. For the heat shock, the transformation mixture containing competent cells and plasmid DNA is incubated on ice to make them cold/numb. This slows fluid cell membrane.
Next the transformation mixture is quickly warmed up. The rapid change in temperature shocks the cells and increases permeability of the cell membranes.
Once the plasmid DNA enters the cells, the cell membrane must recover. For this purpose, the cells are kept in nutrient broth for a short period to gently recover the shocked cells. The recovered cell suspension is grown overnight on agar plates.
Plates containing antibiotic are used to select the transformed cells. The bacteria appear as translucent colonies on some of the agar plates. Each colony represented a single transformed cell (clone).
LB agar plates without antibiotic are used as controls to demonstrate that the cells were alive and survived the transformation.
LB agar plates with antibiotic are used to select the transformed cells. These cells are the antibiotic-resistant phenotype and grow in presence of the antibiotic.
QIAprep SPIN MINIPREP KIT
Isolation Buffer Solutions:
Buffer P1: Resuspension buffer. Contains 50 mM of Tris-HCl at pH 8, 10 mM EDTA and 100 µg/ml RNase A. LyseBlue reagent was added.
Tris is a buffering agent used to maintain a constant pH (= 8.0).
LyseBlue is a pH indicator that provides visual identification (color changes from colorless to blue at alkaline pH) of optimum buffer mixing. Solution appears cloudy show the suspension of cells
Buffer P2: Alkaline lysis buffer. Contains 200 mM NaOH and 1% SDS (w/v)
NaOH is alkaline and ruptures the cells and denatures the DNA into single strands.
Buffer N3: Neutralization buffer. Contains guanidine hydrochloride, acetic acid and potassium acetate, pH 4.3
Buffer pH helps to neutralize the alkaline lysed cell solution, allowing the DNA strands to renature (double stranded).
Solution appears milky precipitate of the insoluble DNA and proteins. Plasmid DNA remains in solution due to its small size and better solubility. The color of the solution (given by Lyseblue) will change from blue to colorless as the pH decreases.
The plasmid DNA can be collected in the supernatant after centrifugation. Polysaccharides, proteins and other contaminants that do not adsorb to the column are removed.
Buffer PE: Wash buffer. Contains Tris-HCl pH 7.5 and 80% ethanol
Ethanol helps to wash the chaotropic salts and other contaminants out from the column.
Purified plasmid DNA remains on the silica gel membrane in the column.
Distilled Water:
Distilled water provides a no-salt condition, which recovers the "hydration shells" around plasmid DNA molecules and dislodges plasmid DNA from the silica gel membrane.
Purified plasmid DNA is dissolved in distilled water and eluted.

Transformation And DNA Isolation Technique
pGLO plasmid was transformed into E. coli bacterial cells. Non-transformed E. coli bacteria were also grown on similar agar plates. The bacteria appear as translucent colonies on some of the agar plates. Each colony represented a single transformed cell (clone). Bacterial cell cultures are used to isolate the DNA in the transformed cells based on their phenotype (survival in presence of antibiotic).
The + pGLO Amp culture is prepared by transferring a single transformed E. coli colony from the LB/Amp agar plate and grown in LB broth containing ampicillin.
The +pGLO Amp/Ara culture is prepared by transferring a single transformed E. coli colony from the LB/Amp agar plate and grown in LB broth containing ampicillin and L(+) arabinose. Both cultures are incubated overnight with vigorous shaking at 37 °C. The shaking is essential to provide continuous oxygen for the fast-growing bacteria.
Cell pellets obtained from the +pGLO Amp and +pGLO Amp/Ara using centrifugation. The plasmid pGLO DNA is isolated using QIAprep Spin Miniprep Kitt. The QIAprep Miniprep procedure is based on alkaline lysis of bacterial cells followed by adsorption of DNA onto silica in the presence of high salt.
During the alkaline lysis process, the cell pellet is lysed (causing the cell membrane to break) and releases its contents. The lysate is subsequently neutralized and adjusted to high-salt–binding conditions. DNA is a weak acid and less soluble in acidic environment. Plasmids can range in size from 1-1000 kilo base (or nucleotide) pairs while chromosomal DNA contains several million base pairs. Hence, chromosomal DNA is less soluble and takes longer to renature and precipitates. The chromosomal DNA is removed as a pellet using centrifugation and the supernatant containing soluble plasmid DNA is subsequently transferred to the QIAprep 2.0 column.
The QIAprep 2.0 columns use a silica membrane for selective adsorption of plasmid DNA in high-salt buffer and elution in low-salt buffer. Salts are efficiently removed by a brief wash step with Buffer PE. High-quality plasmid DNA is then eluted from the QIAprep 2.0 column with 50–100 μl of Buffer EB or water.
RNase is an enzyme that rapidly degrades RNA. It is ubiquitous in the environment. In this plasmid DNA extraction, RNase enzyme is added to P1 buffer so as to exclude the RNA in our DNA extract.