Lecture 20: Anterior-Posterior Patterning in the Drosophila Embryo
1/59
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
60 Terms
If a researcher uses mutations to disrupt one binding site for hunchback in the
stripe 2 regulatory element, what do you think will happen to expression of eve at
stripe 2?
Stripe 2 will have normal expression of eve, because bicoid is also an activator.
Stripe 2 will have normal expression of eve, because there are additional intact
binding sites for hunchback in the regulatory element.
Stripe 2 will be larger than normal.
There will be lower expression of eve at stripe 2.
4
What is forward genetics?
A) Starting with a known gene and engineering a specific mutation to study its function B) Randomly mutagenizing organisms and then studying mutants with interesting phenotypes to identify the genes responsible C) Using DNA sequencing to find all genes in a genome before doing any experiments D) Inserting a foreign gene into an organism to observe what happens
B — Forward genetics = phenotype first, gene second. You create random mutations in many flies, find the ones with cool/weird phenotypes, and work backward to figure out which gene caused it. This is how the 120 patterning genes in Drosophila were discovered (Nobel Prize 1995).
What is a syncytium, and why does it matter for Drosophila early development?
A) A cell with two nuclei that divides asymmetrically to create different cell types B) A structure with multiple nuclei sharing a single cytoplasm, which allows morphogens to diffuse freely and regulate gene expression across the embryo C) A specialized membrane that separates the anterior and posterior halves of the embryo D) A type of stem cell that generates all other cell types during development
B — The early Drosophila embryo undergoes ~13 rounds of nuclear division without forming cell membranes, creating a syncytium. This is critical because proteins (morphogens) can diffuse across the entire embryo and form concentration gradients that pattern the body axes.
In the early Drosophila embryo, why do all the nuclei divide synchronously rather than independently?
A) Each nucleus has its own cell cycle checkpoint that coordinates with the others B) The nuclei are physically connected by spindle fibers throughout development C) Since all nuclei share the same cytoplasm in the syncytium, they are all exposed to the same cell cycle signals at the same time D) A master regulator gene forces all nuclei to divide at the same time
Answer: C — Because the syncytium has no cell membranes, all nuclei are bathed in the same cytoplasm with the same cell cycle regulators (like cyclins). There's no way for one nucleus to receive a "go" signal without all the others getting it too.
Maternal genes differ from zygotic genes in that:
A) Maternal genes are expressed only after fertilization, while zygotic genes are expressed before B) Maternal gene mRNAs are deposited into the egg by the mother before fertilization, while zygotic genes are transcribed from the embryo's own genome after cellularization C) Maternal genes encode structural proteins only, while zygotic genes encode all transcription factors D) Maternal genes are found only on the X chromosome
Answer: B — Maternal genes: mom makes the mRNA and loads it into the egg before fertilization. The embryo uses these stored mRNAs early on. After cellularization (~13 nuclear divisions), the embryo starts transcribing its own genes (zygotic genes).
Which of the following correctly lists the three gene classes responsible for anterior-posterior patterning in Drosophila, in the order they act?
A) Hox genes → segmentation genes → egg polarity genes B) Segmentation genes → egg polarity genes → Hox genes C) Egg polarity genes → segmentation genes → Hox genes D) Egg polarity genes → Hox genes → segmentation genes
Answer: C — The hierarchy goes: egg polarity genes establish the anterior-posterior axis → segmentation genes establish body segments with increasing detail → Hox genes give each segment its specific identity. This is the core organizational framework of the entire lecture.
How do Bicoid and Nanos work together to establish the anterior-posterior axis?
A) Both are expressed uniformly throughout the embryo and compete for the same binding sites B) Bicoid mRNA is localized to the posterior and Nanos mRNA to the anterior, creating overlapping gradients C) Bicoid mRNA is localized to the anterior and Nanos mRNA to the posterior; after fertilization both are translated into proteins that diffuse and form opposing morphogen gradients D) Bicoid and Nanos activate each other in a positive feedback loop to amplify positional signals
Answer: C — Bicoid mRNA sits at the anterior end, Nanos mRNA at the posterior. After fertilization and nuclear division, each is translated and diffuses through the syncytium, creating a high-anterior Bicoid gradient and a high-posterior Nanos gradient. These opposing gradients give every nucleus its "address."
A researcher knocks out the bicoid gene in a Drosophila mother. What is the expected phenotype of her offspring?
A) The larvae will have two heads instead of one B) The larvae will have no abdomen or telson C) The larvae will lack all anterior structures and instead have posterior structures (like the telson) at both ends D) The larvae will develop normally because Nanos can compensate
Answer: C — Loss of Bicoid = loss of anterior identity. Without the Bicoid gradient, the anterior end of the embryo defaults to posterior fate, producing a larva with tail structures at both ends (two telsons) and no head.
What would happen if Bicoid mRNA were injected into the posterior end of a wild-type Drosophila embryo?
A) The embryo would develop normally — Bicoid only works at the anterior end B) The posterior end would develop head structures, resulting in a larva with a head at each end C) The embryo would die immediately due to excess Bicoid protein D) The Nanos gradient would be destroyed, causing loss of all posterior structures
Answer: B — This is the classic "move-it" experiment. Since Bicoid protein specifies anterior identity wherever it is present at high concentrations, artificially placing Bicoid mRNA at the posterior end causes head structures to form there — a larva with heads at both ends.
Gap genes are best characterized by which of the following?
A) They are maternal genes that are deposited before fertilization and expressed in every segment B) They are zygotic genes expressed in broad overlapping domains, encoding transcription regulators, and are regulated by egg polarity genes C) They are expressed in alternating segments and their expression persists into adulthood D) They encode signaling proteins like Hedgehog and Wnt that pattern each segment's polarity
Answer: B — Gap genes are zygotic, expressed in broad domains, encode transcription regulators, and are activated/repressed by the egg polarity gene gradients (like Bicoid and Nanos). Their expression is transient. Example: Hunchback, Krüppel, Giant.
Which of the following is NOT a feature of pair-rule genes?
A) They are regulated by egg polarity genes and gap genes B) Their expression pattern consists of 7 stripes in alternating segments C) They encode transcription regulators D) Their expression is maintained throughout adulthood
Answer: D — Pair-rule gene expression is transient, not maintained in the adult. That's segment polarity genes whose expression persists. Pair-rule genes are expressed in 7 stripes (every other segment) and their expression fades away after establishing the segmentation pattern.
How do segment polarity genes differ from gap and pair-rule genes?
A) Segment polarity genes are expressed in broad domains across the whole embryo, while gap and pair-rule genes are expressed in narrow stripes B) Segment polarity genes encode only transcription factors, while gap and pair-rule genes encode only signaling proteins C) Segment polarity genes are expressed in part of every segment and their expression is maintained into adulthood, whereas gap and pair-rule gene expression is transient D) Segment polarity genes are maternal genes, while gap and pair-rule genes are zygotic
Answer: C — Segment polarity genes are unique in that their expression persists into adulthood. They also encode both transcription regulators AND signaling proteins (like Hedgehog and Wingless/Wnt). They're expressed in every segment (not alternating ones like pair-rule genes).
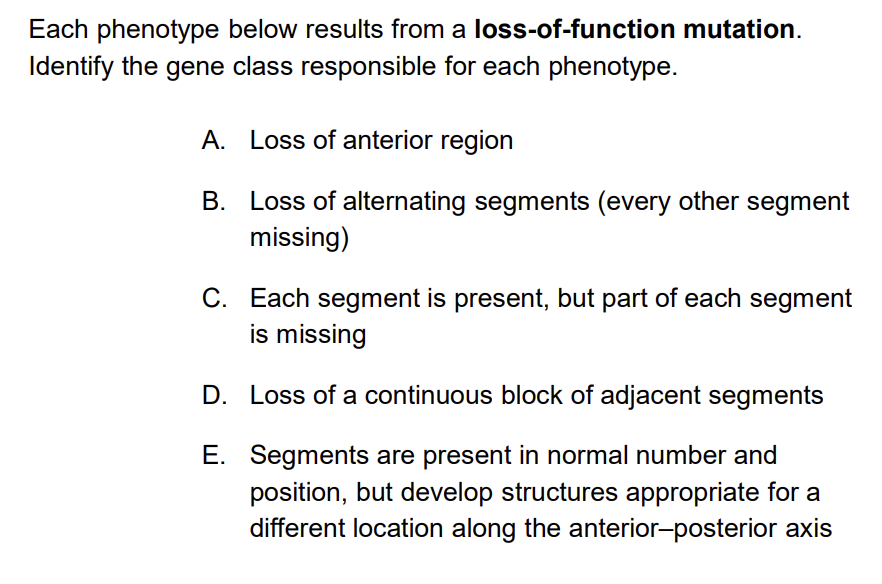
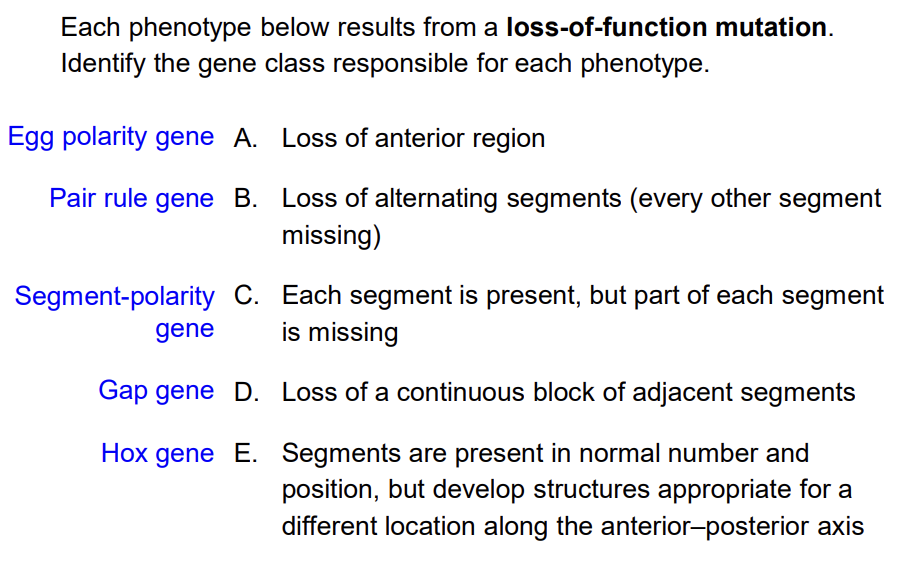
Loss of a gap gene in Drosophila larvae results in which phenotype?
A) Loss of alternating segments across the whole body B) Loss of a large contiguous region of segments where the gap gene is normally expressed C) Loss of anterior polarity within every individual segment D) No phenotype, because pair-rule genes can compensate
Answer: B — Gap genes are expressed in broad contiguous domains. Lose a gap gene → lose a large chunk of consecutive segments in that region. This contrasts with pair-rule mutants (alternating segment loss) and segment polarity mutants (polarity defects within every segment).
In a pair-rule gene loss-of-function mutant, what is the expected larval phenotype?
A) Loss of all segments in the posterior half of the body B) Loss of every other segment throughout the body C) Each segment loses its anterior-posterior polarity but all segments are present D) The larva has no segments at all
Answer: B — Pair-rule genes (like even-skipped) are expressed in every other segment. Losing a pair-rule gene → losing every other segment, resulting in roughly half the normal number of segments. The name "pair-rule" refers to the fact that these mutations affect alternating pairs of segments.
The Even-skipped (Eve) gene is expressed in 7 stripes in the Drosophila embryo. How is the stripe-specific pattern of Eve expression achieved?
A) The Eve protein itself controls where it is expressed by binding its own promoter in a feedback loop B) Each stripe is independently controlled by a different enhancer element in the Eve regulatory region, which integrates inputs from different combinations of transcriptional activators and repressors C) A single enhancer element responds to the overall Bicoid gradient and activates Eve in exactly 7 evenly spaced domains D) Eve expression is triggered by cell-cell contact signaling, with each cell activating Eve in its neighbor
Answer: B — Eve is the classic example of modular enhancer control. The gene has separate enhancers for different stripes (e.g., stripe 2 enhancer, stripe 3 & 7 enhancer, etc.). Each enhancer has binding sites for specific activators and repressors that are present in different combinations at different positions along the A-P axis.
For Eve stripe 2, Bicoid and Hunchback are transcriptional activators, while Giant and Krüppel are transcriptional repressors. A researcher disrupts all binding sites for Giant in the stripe 2 enhancer. What is the most likely outcome?
A) Stripe 2 disappears because Giant is needed to activate Eve B) Stripe 2 expression expands because the repressor can no longer bind and shut off Eve at the edges of stripe 2 C) Stripe 2 becomes narrower because Giant normally helps define its borders D) None of the other Eve stripes are affected because each has an independent enhancer
Answer: B — Giant normally represses Eve at the anterior border of stripe 2 (where Giant concentration is high). Remove Giant's binding sites → remove that repression → Eve expression expands anteriorly. This is the logic of combinatorial transcriptional regulation: precise boundaries require both activators AND repressors.
A researcher disrupts a single Hunchback binding site in the stripe 2 regulatory element. What is the most likely result?
A) Stripe 2 expression is completely absent because Hunchback is the only activator B) Stripe 2 expression is normal because Bicoid is still present as an activator and multiple intact Hunchback sites remain C) Stripe 2 shifts posteriorly D) All 7 Eve stripes disappear simultaneously
Answer: B — This is directly from your lecture's Poll Everywhere question. Because there are multiple Hunchback binding sites in the stripe 2 enhancer AND Bicoid is also an activator, disrupting just one Hunchback site doesn't eliminate activation entirely. The remaining sites still function. (If ALL Hunchback sites were destroyed, you'd see reduced or absent stripe 2.)
What is the primary function of Hox genes in Drosophila development?
A) Establishing the initial anterior-posterior axis before segmentation occurs B) Creating the physical boundaries between body segments C) Giving each segment its specific identity by controlling which downstream genes are expressed (or repressed) in that segment D) Controlling the number of body segments produced during embryogenesis
Answer: C — Hox genes are master regulators that act AFTER segmentation has already occurred. Their job is to tell each segment what to become — head, thorax, abdomen, etc. They do this primarily by repressing genes for structures that don't belong in that segment.
Answer: C — Hox genes are master regulators that act AFTER segmentation has already occurred. Their job is to tell each segment what to become — head, thorax, abdomen, etc. They do this primarily by repressing genes for structures that don't belong in that segment.
Which of the following correctly describes the chromosomal organization of Hox genes?
A) Hox genes are scattered randomly across all chromosomes B) Hox genes are clustered together on chromosomes, and their order on the chromosome corresponds to their expression pattern along the anterior-posterior axis C) Hox genes are located exclusively on sex chromosomes D) All Hox genes are located on a single chromosome but their expression is determined by which chromosome is active
Answer: B — This is one of the most striking features of Hox genes: their physical order on the chromosome matches the order of where they're expressed in the body, from anterior to posterior. This "spatial colinearity" is conserved across nearly all animals.
Vertebrates have up to four copies of the Hox gene cluster, while Drosophila has only one. What explains this difference?
A) Vertebrates evolved more complex regulatory sequences that duplicated Hox gene functions B) The Hox gene cluster was duplicated (up to four times) during vertebrate evolution C) Vertebrates have more body segments, requiring more Hox genes D) Individual Hox genes in vertebrates are all on different chromosomes due to translocation events
Answer: B — The ancestral Hox cluster was duplicated multiple times during early vertebrate evolution, giving rise to up to four paralogous clusters (HoxA, HoxB, HoxC, HoxD). Drosophila retained only one copy because no such duplications occurred in the insect lineage.
Which of the following is NOT true about Hox genes?
A) They encode transcription regulators B) Their expression is maintained into adulthood C) They are found only in insects and are not conserved in vertebrates D) Misexpression of a Hox gene can result in structures forming in the wrong location
Answer: C — Hox genes are among the most evolutionarily conserved genes known. All animals except sponges have them. The same Hox genes that pattern a fly's body axis have homologs that pattern the vertebrate body axis. This was one of the biggest surprises of evo-devo biology.
When the Antennapedia Hox gene is misexpressed in the fly's head (a "move-it" experiment), legs grow where antennae should be. This happens because:
A) Antennapedia directly destroys antenna precursor cells B) Antennapedia normally represses leg development genes, so its absence allows legs to form everywhere C) Antennapedia is a Hox gene that normally activates leg development in the thorax; when misexpressed in the head, it activates the leg developmental program there instead of the antenna program D) Antennapedia causes head cells to migrate to the thorax where they form extra legs
Answer: C — Antennapedia normally specifies thoracic identity (legs). When artificially expressed in the head, the head precursor cells receive "thorax" instructions and activate the leg developmental program instead of the antenna program. Same cells, different Hox input, completely different outcome.
How are posterior Hox genes prevented from being expressed in the head, where they don't belong?
A) The head cells physically degrade posterior Hox mRNA as soon as it is produced B) Posterior Hox genes are silenced in anterior segments by being packed into heterochromatin via repressive histone modifications C) The Bicoid protein directly binds posterior Hox gene promoters and blocks RNA polymerase D) Posterior Hox genes are located on a different chromosome in head cells than in abdominal cells
Answer: B — This connects back to Lecture 19! Posterior Hox genes are silenced in the head by epigenetic mechanisms — specifically repressive histone modifications (like H3K27me3) that pack the relevant Hox genes into heterochromatin, making them transcriptionally inaccessible. Chromatin remodeling proteins (Polycomb group proteins) maintain this silencing.
Which of the following is the correct sequence describing how the anterior-posterior axis of Drosophila is patterned, from earliest to latest?
A) Hox genes → gap genes → pair-rule genes → segment polarity genes → egg polarity genes B) Egg polarity genes form morphogen gradients → gap genes expressed in broad domains → pair-rule genes expressed in alternating segments → segment polarity genes expressed in every segment → Hox genes give identity to each segment C) Segment polarity genes → pair-rule genes → gap genes → egg polarity genes → Hox genes D) Egg polarity genes → Hox genes → gap genes → pair-rule genes → segment polarity genes
Answer: B — This is the core hierarchy: broad gradients (egg polarity) → broad domains (gap) → alternating stripes (pair-rule) → every segment (segment polarity) → segment identity (Hox). Each level refines the pattern established by the previous one.
All of the following are true about Hox genes EXCEPT:
A) They are master regulators B) They are expressed into adulthood C) They are found in all animals including sponges D) Their order on the chromosome mirrors their expression pattern along the body axis
Answer: C — All animals EXCEPT sponges have Hox genes. Sponges lack true body axes and Hox genes, which is consistent with Hox genes being fundamentally linked to axial patterning.
A researcher conducts a forward genetics screen and finds a Drosophila mutant where the larva has a normal number of segments but every segment appears identical and lacks polarity (no front-to-back differences within each segment). Which class of genes is most likely mutated?
A) Egg polarity genes, because Bicoid and Nanos establish the initial axis B) Gap genes, because loss of gap genes eliminates large regions of the body plan C) Pair-rule genes, because even-skipped patterns alternating segments D) Segment polarity genes, because they establish polarity within each individual segment
Answer: D — The key clue is "normal number of segments, but each segment lacks internal polarity." That's the hallmark of segment polarity gene loss (e.g., hedgehog, wingless/Wnt, engrailed). Losing a gap gene gives you missing segments. Losing a pair-rule gene gives you alternating segment loss. Losing a segment polarity gene leaves all segments but scrambles the anterior-posterior polarity within each one.


Which approach was used to elucidate mechanisms underlying anterior-posterior patterning in Drosophila in the Nobel-prize winning research by Lewis, Nusslein-Volhard, and Weischaus?
A) Descriptive embryology B) Experimental embryology C) Developmental genetics D) Comparative embryology
Answer: C
Which feature best describes the early Drosophila embryo during the first rounds of nuclear divisions?
A) A single nucleus surrounded by thousands of cells B) Alternating bands of differentiated cell types C) Distinct anterior and posterior cells separated by membranes D) Many nuclei sharing a common cytoplasm
Answer: D
Why do early nuclear divisions in the Drosophila embryo occur synchronously?
A) Egg-polarity gene products create different microenvironments along the A-P axis B) Each nucleus independently activates transcription of maternal-effect genes C) All nuclei share a common cytoplasm containing a shared pool of cell-cycle regulators D) Bicoid and Nanos coordinate mitosis in every nucleus
Answer: C
Which statements are TRUE about maternal genes? Select ALL that apply.
A) Maternal mRNAs are deposited in the egg before fertilization B) Maternal gene products can diffuse as morphogens C) Maternal genes remain the primary regulators after cellularization D) Maternal genes often encode transcription factors
Answer: A, B, and D
How do Bicoid and Nanos proteins help establish the anterior-posterior axis?
A) They remain fixed at the poles and activate gene expression only in nearest nuclei B) They are transported by motor proteins to every nucleus to activate the same genes throughout the embryo C) They diffuse through the syncytium from their localized mRNA sources, forming opposing morphogen gradients that regulate gene expression along the A-P axis D) Only their mRNA gradients provide positional information, not the proteins themselves
Answer: C
What would happen if Bicoid mRNA were added to the posterior end of the embryo?
A) Posterior identity remains unchanged B) Posterior structures are duplicated C) An additional anterior region forms D) Posterior identity is lost throughout the embryo
Answer: C
Which region of the embryo would express a gene that is activated by Bicoid but repressed by Nanos?
A) Posterior B) Middle C) Entire embryo D) Anterior
Answer: D
Which statement correctly describes the roles of maternal and zygotic genes in early Drosophila embryogenesis?
A) Maternal mRNAs are not translated until after cellularization, when zygotic transcription begins B) Maternal mRNAs encode key transcription factors that diffuse through the syncytial embryo and regulate early patterning before zygotic genes are activated C) Zygotic genes establish the A-P axis before fertilization, while maternal genes refine patterns later D) Zygotic transcription drives the earliest nuclear divisions, while maternal mRNAs function only in later segmentation
Answer: B
Which gene classes are encoded by the embryo as zygotic genes? Select ALL that apply.
A) Egg polarity genes B) Gap genes C) Pair-rule genes D) Segment polarity genes E) Hox genes
Answer: B, C, D, and E
Which is the correct hierarchical order of A-P patterning genes from earliest to latest acting?
A) Egg-polarity → gap → pair-rule → segment-polarity B) Egg-polarity → pair-rule → gap → segment-polarity C) Segment-polarity → egg-polarity → gap → pair-rule D) Gap → egg-polarity → pair-rule → segment polarity
Answer: A
A mutation eliminates the function of a gap gene in the Drosophila embryo. What is the most likely phenotype?
A) Alternating segments are missing B) A contiguous block of adjacent segments is missing C) Segment boundaries are disrupted within each segment D) Segment identities are transformed into other segment types
Answer: B
Which gene classes, when mutated, would result in fewer total segments? Select ALL that apply.
A) Gap genes B) Pair-rule genes C) Segment polarity genes D) Hox genes
Answer: A and B
Match the expression pattern to the correct gene class. Which gene is expressed as a single anterior gradient in the early embryo?
A) Gap gene (e.g., Hunchback) B) Pair-rule gene (e.g., Even-skipped) C) Egg-polarity gene (e.g., Bicoid) D) Segment-polarity gene (e.g., Engrailed)
Answer: C
Which gene class is expressed in 7 stripes in the early Drosophila embryo?
A) Egg-polarity genes B) Gap genes C) Pair-rule genes D) Segment polarity genes
Answer: C
Which gene class is expressed in 14 stripes and controls the polarity and boundary of each segment?
A) Gap genes B) Pair-rule genes C) Hox genes D) Segment polarity genes
Answer: D
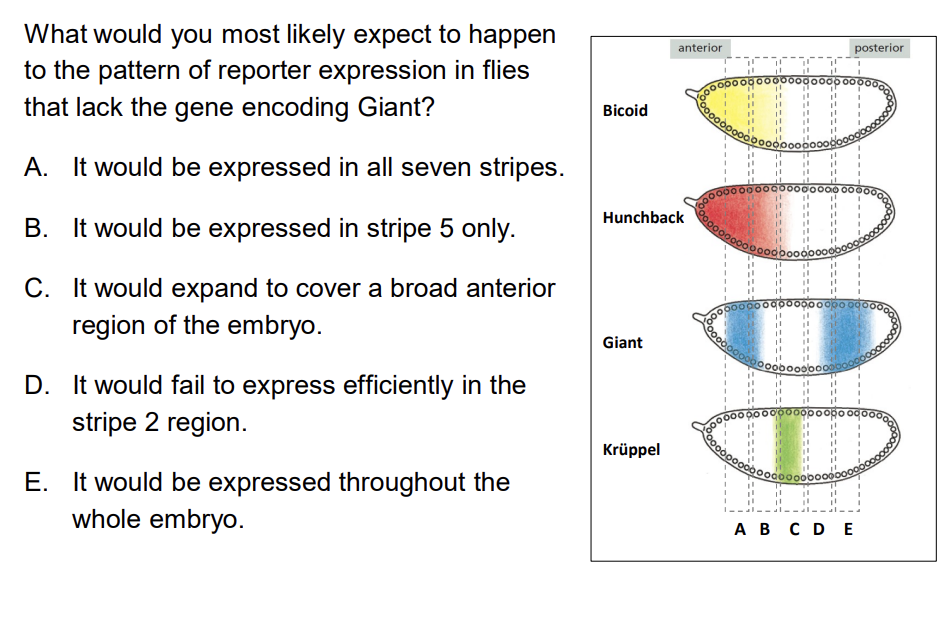
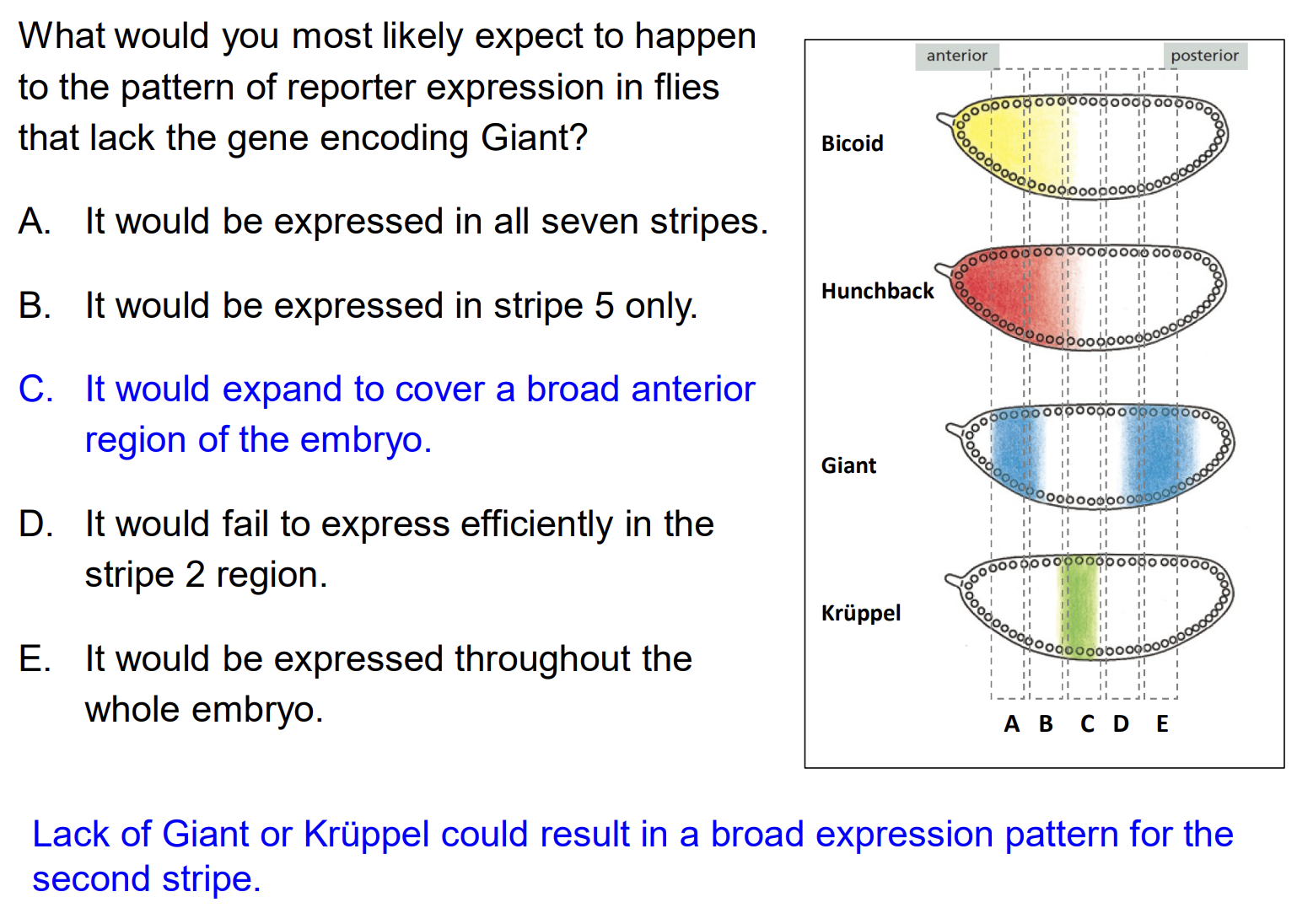
The Eve stripe 2 module has binding sites for Bicoid and Hunchback (activators) and Giant and Krüppel (repressors). What would happen to reporter expression if the gene encoding Giant were lost?
A) It would be expressed in all seven stripes B) It would be expressed in a more posterior stripe C) It would expand to cover a broad anterior region of the embryo D) It would fail to express in the stripe 2 region
Answer: C
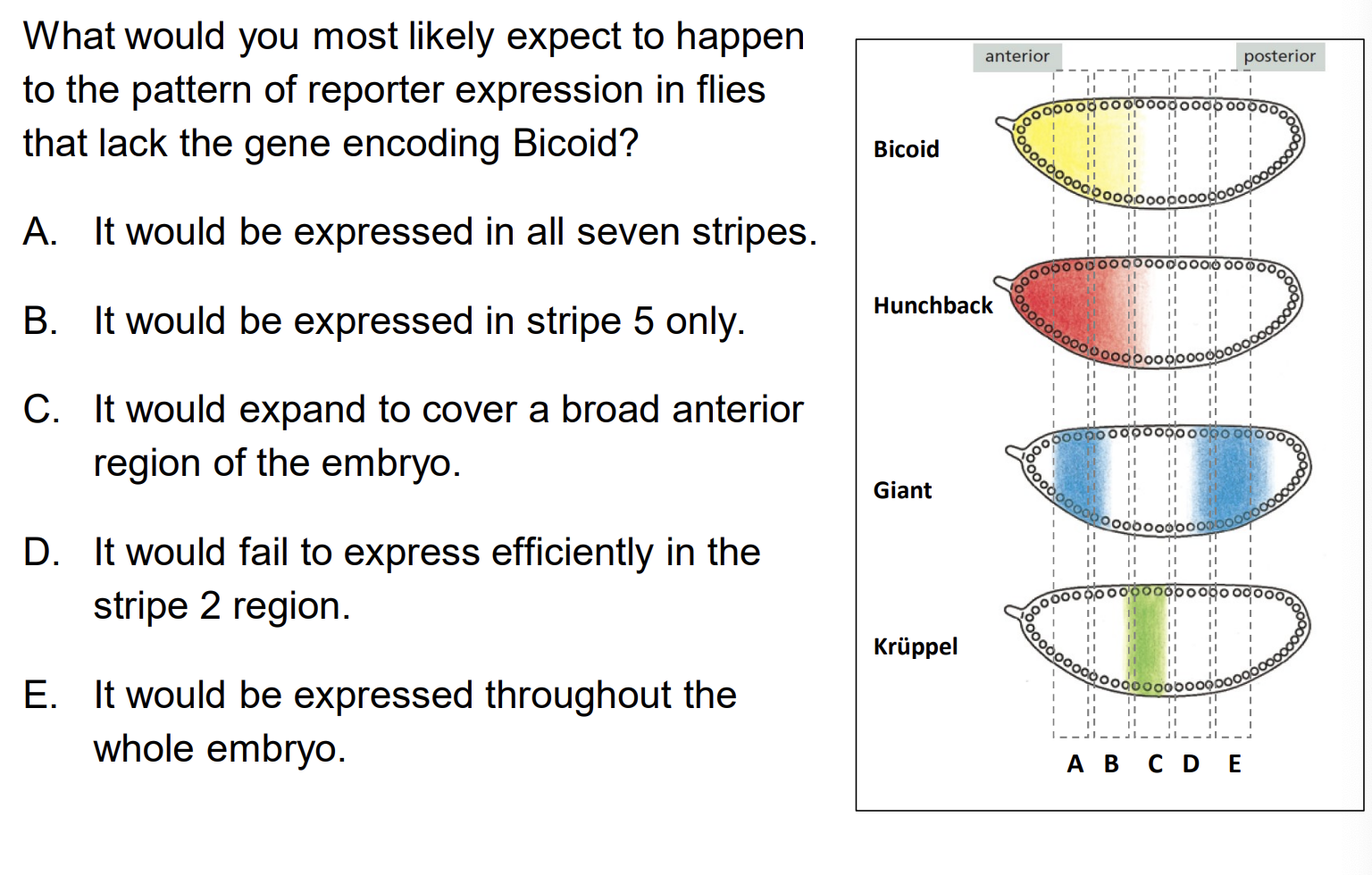
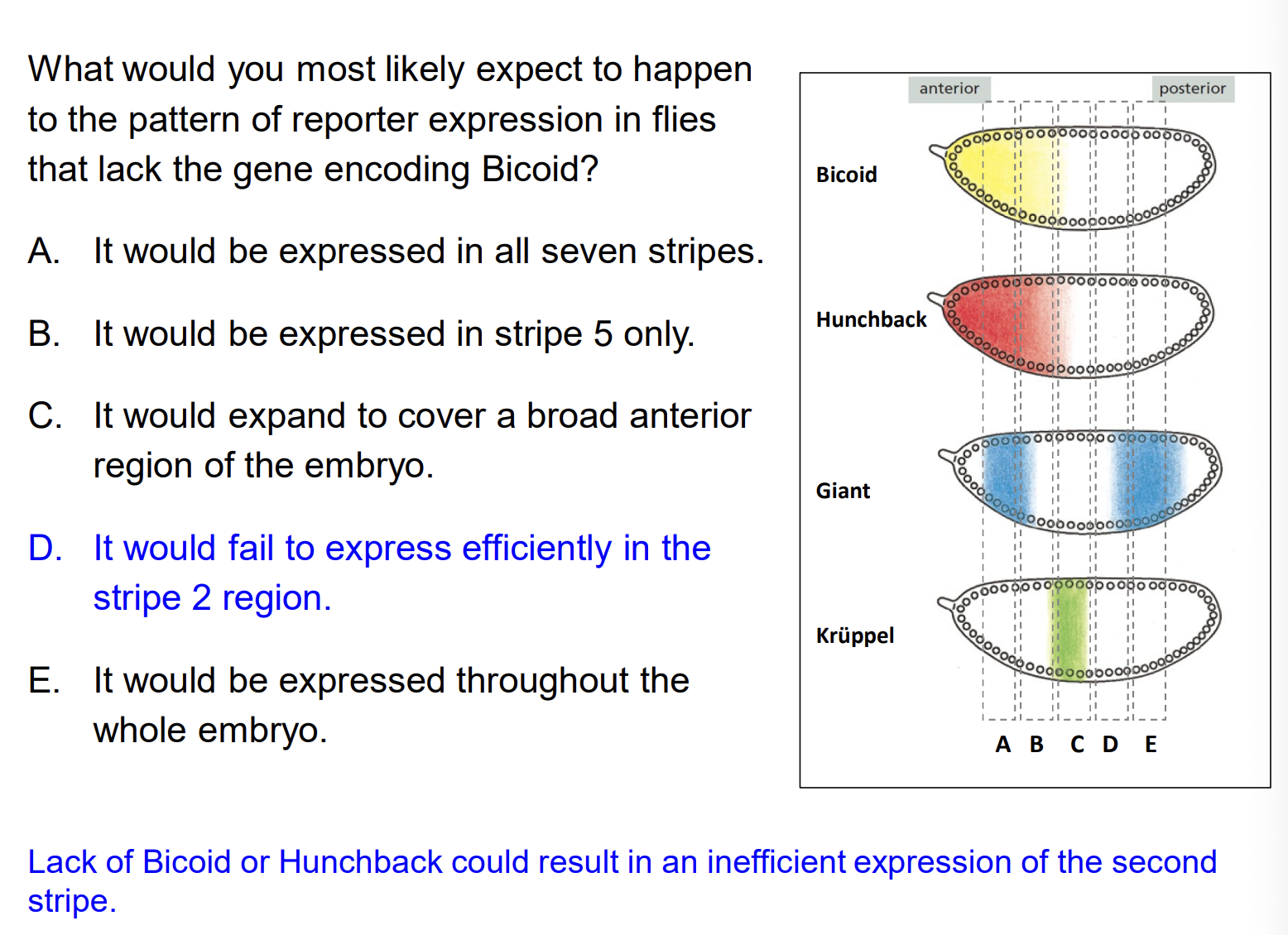
What would happen to Eve stripe 2 reporter expression in flies that lack Bicoid?
A) It would be expressed in all seven stripes B) It would expand to cover a broad anterior region C) It would fail to express efficiently in the stripe 2 region D) It would be expressed throughout the whole embryo
Answer: C
You stain wildtype and gene Z-deficient mutant embryos to examine the distribution of protein A. What can you infer from this experiment?
A) Whether gene A influences expression of gene Z and whether it acts as a repressor or activator B) Whether gene Z influences expression of gene A and whether it acts as a repressor or activator on gene A in each stripe C) Both of the above D) None of the above
Answer: B
Which group of A-P patterning genes encodes both transcription factors AND cell signaling pathway components?
A) Pair-rule genes B) Gap genes C) Segment polarity genes D) Egg polarity genes
Answer: C
Why do segment polarity genes encode cell signaling components, unlike gap and pair-rule genes?
A) Segment polarity genes function before fertilization when signaling is required B) Segment polarity genes begin functioning after cellularization, when cell boundaries require cell-cell signaling C) Segment polarity genes need to activate transcription across long distances in the syncytium D) None of the above
Answer: B
Which groups of genes continue to be expressed through metamorphosis and into adulthood? Select ALL that apply.
A) Egg-polarity genes B) Gap genes C) Pair-rule genes D) Segment polarity genes E) Hox genes
Answer: D and E
Which type of developmental defect results from a mutation in a Hox gene?
A) Several adjacent segments will be missing from the embryo B) Segments will develop into a different type of segment than normal C) The anterior portion of the embryo will not develop D) All of the above
Answer: B
Which statement about Hox genes is FALSE?
A) Hox genes specify the identity of segments B) Hox genes are expressed in patterns that match their order along the chromosome C) Hox genes determine how many segments form D) Hox genes can repress genes for structures in other segments
Answer: C
Which of the following statements regarding Hox genes in Drosophila are TRUE? Select ALL that apply.
A) If all Hox genes are deleted, segmentation still occurs but segments won't take on normal identities B) Hox proteins are homeodomain transcription factors with a conserved DNA-binding domain C) Hox proteins act as master regulators controlling expression of multiple genes D) A mutation in a Hox gene results in a homeotic mutation E) The expression pattern of Hox genes mirrors their chromosomal location
Answer: A, B, C, D, and E — all of the above are TRUE
Which statements about anterior-posterior patterning in Drosophila are TRUE? Select ALL that apply.
A) Maternal gene products are deposited asymmetrically in the egg before fertilization B) Early nuclear divisions occur in a shared cytoplasm C) Maternal gene products can diffuse through the syncytium as morphogens D) Once cellularization occurs, the embryo activates transcription of its own zygotic genes E) Most segmentation genes are zygotic genes F) Segment organization is maintained throughout development into adulthood
Answer: All of the above — A, B, C, D, E, and F
What is forward genetics?
A) Engineering a specific mutation in a known gene and observing the phenotype B) Randomly mutagenizing organisms and studying mutants with interesting phenotypes to learn about the responsible genes C) Comparing gene sequences across species to infer evolutionary relationships D) Injecting mRNA into embryos to observe gain-of-function effects
Answer: B
Which of the following correctly describes a morphogen?
A) A protein that is synthesized uniformly throughout the embryo and activates all genes equally B) A signaling molecule that forms a concentration gradient and specifies different cell fates at different concentrations C) A transcription factor that is only active after cellularization D) A maternal mRNA that is never translated until adulthood
Answer: B
A mutation in which class of genes would produce an embryo missing its anterior or posterior segments?
A) Hox genes B) Pair-rule genes C) Segment polarity genes D) Egg-polarity genes
Answer: D
Which of the following is TRUE about Hox gene conservation across species?
A) Hox genes are unique to Drosophila and not found in vertebrates B) Hox genes are present and regulate anterior-posterior patterning in all bilaterally symmetric animals including humans C) Hox genes in humans regulate segment number, not segment identity D) None of the above
Answer: B
Which of the following best explains why different concentrations of Bicoid activate different target genes along the A-P axis?
A) Bicoid binds different promoters only when it is phosphorylated at high concentrations B) Different target genes have binding sites with different affinities for Bicoid, so genes are activated at different threshold concentrations C) Bicoid is degraded at the anterior and accumulates at the posterior to regulate targets D) Different target genes are only accessible once cellularization separates nuclei from the syncytium
Answer: B
Which approach was used to elucidate mechanisms underlying anterior-posterior patterning in Drosophila in the Nobel-prize winning research by Lewis, Nusslein-Volhard, and Weischaus?
A) Descriptive embryology B) Experimental embryology C) Developmental genetics D) Comparative embryology
C
Which feature best describes the early Drosophila embryo during the first rounds of nuclear divisions?
A) A single nucleus surrounded by thousands of cells B) Alternating bands of differentiated cell types C) Distinct anterior and posterior cells separated by membranes D) Many nuclei sharing a common cytoplasm
D
Why do early nuclear divisions in the Drosophila embryo occur synchronously?
A) Egg-polarity gene products create different microenvironments along the A-P axis B) Each nucleus independently activates transcription of maternal-effect genes C) All nuclei share a common cytoplasm containing a shared pool of cell-cycle regulators D) Bicoid and Nanos coordinate mitosis in every nucleus
C
Which statements are TRUE about maternal genes? Select ALL that apply.
A) Maternal mRNAs are deposited in the egg before fertilization B) Maternal gene products can diffuse as morphogens C) Maternal genes remain the primary regulators after cellularization D) Maternal genes often encode transcription factors
A, B, D
How do Bicoid and Nanos proteins help establish the anterior-posterior axis?
A) They remain fixed at the poles and activate gene expression only in nearest nuclei B) They are transported by motor proteins to every nucleus to activate the same genes throughout the embryo C) They diffuse through the syncytium from their localized mRNA sources, forming opposing morphogen gradients that regulate gene expression along the A-P axis D) Only their mRNA gradients provide positional information, not the proteins themselves
C
What would happen if Bicoid mRNA were added to the posterior end of the embryo?
A) Posterior identity remains unchanged B) Posterior structures are duplicated C) An additional anterior region forms D) Posterior identity is lost throughout the embryo
C
Which region of the embryo would express a gene that is activated by Bicoid but repressed by Nanos?
A) Posterior B) Middle C) Entire embryo D) Anterior
D
Which statement correctly describes the roles of maternal and zygotic genes in early Drosophila embryogenesis?
A) Maternal mRNAs are not translated until after cellularization, when zygotic transcription begins B) Maternal mRNAs encode key transcription factors that diffuse through the syncytial embryo and regulate early patterning before zygotic genes are activated C) Zygotic genes establish the A-P axis before fertilization, while maternal genes refine patterns later D) Zygotic transcription drives the earliest nuclear divisions, while maternal mRNAs function only in later segmentation
B
Which gene classes are encoded by the embryo as zygotic genes? Select ALL that apply.
A) Egg polarity genes B) Gap genes C) Pair-rule genes D) Segment polarity genes E) Hox genes
Answer: B, C, D, and E
B, C, D, E
Which is the correct hierarchical order of A-P patterning genes from earliest to latest acting?
A) Egg-polarity → gap → pair-rule → segment-polarity B) Egg-polarity → pair-rule → gap → segment-polarity C) Segment-polarity → egg-polarity → gap → pair-rule D) Gap → egg-polarity → pair-rule → segment polarity
A
A mutation eliminates the function of a gap gene in the Drosophila embryo. What is the most likely phenotype?
A) Alternating segments are missing B) A contiguous block of adjacent segments is missing C) Segment boundaries are disrupted within each segment D) Segment identities are transformed into other segment types
B
Which gene classes, when mutated, would result in fewer total segments? Select ALL that apply.
A) Gap genes B) Pair-rule genes C) Segment polarity genes D) Hox genes
A and B
Match the expression pattern to the correct gene class. Which gene is expressed as a single anterior gradient in the early embryo?
A) Gap gene (e.g., Hunchback) B) Pair-rule gene (e.g., Even-skipped) C) Egg-polarity gene (e.g., Bicoid) D) Segment-polarity gene (e.g., Engrailed)
C
Which gene class is expressed in 7 stripes in the early Drosophila embryo?
A) Egg-polarity genes B) Gap genes C) Pair-rule genes D) Segment polarity genes
C
Which gene class is expressed in 14 stripes and controls the polarity and boundary of each segment?
A) Gap genes B) Pair-rule genes C) Hox genes D) Segment polarity genes
D
The Eve stripe 2 module has binding sites for Bicoid and Hunchback (activators) and Giant and Krüppel (repressors). What would happen to reporter expression if the gene encoding Giant were lost?
A) It would be expressed in all seven stripes B) It would be expressed in a more posterior stripe C) It would expand to cover a broad anterior region of the embryo D) It would fail to express in the stripe 2 region
C
What would happen to Eve stripe 2 reporter expression in flies that lack Bicoid?
A) It would be expressed in all seven stripes B) It would expand to cover a broad anterior region C) It would fail to express efficiently in the stripe 2 region D) It would be expressed throughout the whole embryo
C
You stain wildtype and gene Z-deficient mutant embryos to examine the distribution of protein A. What can you infer from this experiment?
A) Whether gene A influences expression of gene Z and whether it acts as a repressor or activator B) Whether gene Z influences expression of gene A and whether it acts as a repressor or activator on gene A in each stripe C) Both of the above D) None of the above
B
Which group of A-P patterning genes encodes both transcription factors AND cell signaling pathway components?
A) Pair-rule genes B) Gap genes C) Segment polarity genes D) Egg polarity genes
C
Why do segment polarity genes encode cell signaling components, unlike gap and pair-rule genes?
A) Segment polarity genes function before fertilization when signaling is required B) Segment polarity genes begin functioning after cellularization, when cell boundaries require cell-cell signaling C) Segment polarity genes need to activate transcription across long distances in the syncytium D) None of the above
B
Which groups of genes continue to be expressed through metamorphosis and into adulthood? Select ALL that apply.
A) Egg-polarity genes B) Gap genes C) Pair-rule genes D) Segment polarity genes E) Hox genes
Answer: D and E
D and E
Which type of developmental defect results from a mutation in a Hox gene?
A) Several adjacent segments will be missing from the embryo B) Segments will develop into a different type of segment than normal C) The anterior portion of the embryo will not develop D) All of the above
B
Which statement about Hox genes is FALSE?
A) Hox genes specify the identity of segments B) Hox genes are expressed in patterns that match their order along the chromosome C) Hox genes determine how many segments form D) Hox genes can repress genes for structures in other segments
C
Which of the following statements regarding Hox genes in Drosophila are TRUE? Select ALL that apply.
A) If all Hox genes are deleted, segmentation still occurs but segments won't take on normal identities B) Hox proteins are homeodomain transcription factors with a conserved DNA-binding domain C) Hox proteins act as master regulators controlling expression of multiple genes D) A mutation in a Hox gene results in a homeotic mutation E) The expression pattern of Hox genes mirrors their chromosomal location
All
Which statements about anterior-posterior patterning in Drosophila are TRUE? Select ALL that apply.
A) Maternal gene products are deposited asymmetrically in the egg before fertilization B) Early nuclear divisions occur in a shared cytoplasm C) Maternal gene products can diffuse through the syncytium as morphogens D) Once cellularization occurs, the embryo activates transcription of its own zygotic genes E) Most segmentation genes are zygotic genes F) Segment organization is maintained throughout development into adulthood
all
What is forward genetics?
A) Engineering a specific mutation in a known gene and observing the phenotype B) Randomly mutagenizing organisms and studying mutants with interesting phenotypes to learn about the responsible genes C) Comparing gene sequences across species to infer evolutionary relationships D) Injecting mRNA into embryos to observe gain-of-function effects
B
Which of the following correctly describes a morphogen?
A) A protein that is synthesized uniformly throughout the embryo and activates all genes equally B) A signaling molecule that forms a concentration gradient and specifies different cell fates at different concentrations C) A transcription factor that is only active after cellularization D) A maternal mRNA that is never translated until adulthood
B
A mutation in which class of genes would produce an embryo missing its anterior or posterior segments?
A) Hox genes B) Pair-rule genes C) Segment polarity genes D) Egg-polarity genes
D
Which of the following is TRUE about Hox gene conservation across species?
A) Hox genes are unique to Drosophila and not found in vertebrates B) Hox genes are present and regulate anterior-posterior patterning in all bilaterally symmetric animals including humans C) Hox genes in humans regulate segment number, not segment identity D) None of the above
B
Which of the following best explains why different concentrations of Bicoid activate different target genes along the A-P axis?
A) Bicoid binds different promoters only when it is phosphorylated at high concentrations B) Different target genes have binding sites with different affinities for Bicoid, so genes are activated at different threshold concentrations C) Bicoid is degraded at the anterior and accumulates at the posterior to regulate targets D) Different target genes are only accessible once cellularization separates nuclei from the syncytium
B